Introduction
Cerebral ischemia leads to insufficient oxygen
supply and ischemic stroke (1,2), and
is associated with a number of diseases or disorders (3). Microvascular endothelial cells can be
activated during the hypoxia or ischemia, and by upregulating the
expression levels of various agents, including proinflammatory
mediators and adhesion molecules (4). Currently, there is no effective
treatment or prevention available for the management of cerebral
ischemia, and our knowledge is limited pertaining to brain
microvascular endothelial (bEnd.3) cells during cerebral
ischemia.
Peroxisome proliferator-activated receptors (PPARs)
are ligand-activated transcription factors with three distinct
isoforms (PPARα, PPARγ and PPARδ) (5,6).
PPAR activation serves a role in anti-inflammatory effects in the
brain and may serve as a novel pharmacological target for the
management of neurological diseases (7,8).
Previous studies have revealed that brain ischemic injury enhanced
the expression and activity of PPARγ, and PPARγ agonists may
protect neuron cells against brain ischemic injury (9,10).
However, the function of PPARs in bEnd.3 cells during cerebral
ischemia remains unknown. Baculoviral IAP repeat-containing 5
(BIRC5; also known as survivin) belongs to the inhibitor of
apoptosis (IAP) gene family that is widely expressed in cancer
cells (11). Hypoxic
preconditioning may protect brain endothelium from ischemia-induced
apoptosis by Akt-dependent BIRC5 activation (12), which implied a potential connection
between BIRC5 expression and human brain endothelium injury. BIRC5
was also reported to cooperate with PARP proteins in studies on
cell cycle (13) or on cell
proliferation in bladder cancer cells (14). In short, the potential role of
BIRC5 in cerebral ischemia and its interaction with PPARγ need to
be elucidated (15).
The present study demonstrated that PPARγ may
protect cerebral microvascular endothelium against
ischemia-reperfusion injury, and that BIRC5 may be a novel target
of PPARγ. These results may provide insights for future
investigations considering the crucial role of PPAR regulators and
targets in the pathogenesis of stroke.
Materials and methods
Cell culture
Mouse bEnd.3 cells were obtained from the American
Type Culture Collection (ATCC; Manassas, VA, USA). All cells were
maintained as previously described (16). Briefly, Cells were cultured in
RPMI-1640 (Invitrogen; Thermo Fisher Scientific, Inc., Waltham, MA,
USA) and supplemented with 15% fetal bovine serum (Sigma-Aldrich;
Merck KGaA, Darmstadt, Germany), 100 U/ml penicillin and 100 U/ml
streptomycin (Ameresco, Inc., Framingham, MA, USA). Cells were
grown in a humidified atmosphere of 5% CO2/95% air at
37°C. The growth medium was replaced each day; cells were plated
onto 96-well plates or Petri dishes for further analysis.
Plasmid construction and transfection. The complete
coding sequence of PPARγ (https://www.ncbi.nlm.nih.gov/gene/5468) was amplified
and cloned into pcDNA4.1 vector (Invitrogen; Thermo Fisher
Scientific, Inc.). The bEnd.3 cells (2×105 cells/well)
were seeded into 24-well plates and then pcDNA4.1-PPARγ
overexpression plasmid (7 µg/ml, experimental group) or empty
pcDNA4.1 vector (7 µg/ml, control group) was transfected into
bEnd.3 cells using Lipofectamine 2000 (Invitrogen; Thermo Fisher
Scientific, Inc.), following the manufacturer's instructions. The
working concentration of plasmid was determined by previously study
(17). After incubated at 37°C for
24 h, cells that stably overexpressed PPARγ were selected and moved
to new 24-well plates at a concentration of 2×105
cells/well, after incubated overnight at 37°C, cells were treated
with oxygen-glucose deprivation (OGD) for 12 h to establish
an ischemic cell model, then western blot analysis and
immunofluorescence assay were used to measure the expression of
PPARγ in these cells.
Preparation of OGD model
To mimic ischemic conditions in vitro, bEnd.3
cells (2×105 cells/ml) were exposed to OGD. Cell
cultures were subjected to ischemia-like injury through OGD for 3,
6 and 12 h by placing cultures in a Forma Anaerobic Chamber (Thermo
Fisher Scientific, Inc.) with an atmosphere of O2
tension <0.2% (5% CO2, 5% H2 and 90%
N2) in a deoxygenated glucose-free balanced salt
solution. Cultures were placed in a humidified incubator at 37°C.
Cultured cells and media were harvested by trypsinization and
re-suspended in PBS, and then centrifugation at 16,000 × g for 10
min at 4°C as previously described (18,19)
at different time points for further experiments.
Proliferation assays
The proliferative ability of bEnd.3 cells was
measured using the Cell Counting Kit-8 (CCK-8), according to the
manufacturer's instructions. The bEnd.3 cells were seeded into
24-well plates at a concentration of 2×105 cells/well,
and incubated overnight at 37°C. After treated with OGD for 0, 3, 6
or 12 h, CCK-8 solution (10 µl) was added to 96-well plates
incubated at 37°C for 4 h in 5% CO2, and the absorbance
of each well was detected using a microplate reader (Multiskan
Spectrum; Thermo Fisher Scientific, Inc., Waltham, MA, USA) at a
wavelength of 450 nm.
The 5-ethynyl-29-deoxyuridine (EdU) Cell-Light
Apollo DNA in vitro Imaging kit (Guangzhou RiboBio Co.,
Ltd., Guangzhou, China) was also used to examine proliferative
ability. Cells (1×105 cells/dish) were cultured in Petri
dishes for 24 h at 37°C. Following OGD treatment, 50 µM of EdU was
added to each dish and cells were cultured for an additional 2 h at
37°C, and then cells stained with EdU were analyzed using the
CellQuest Flow Cytometry System version 5.1 (BD Biosciences,
Franklin Lakes, NJ, USA).
RNA extraction and reverse
transcription-quantitative polymerase chain reaction (RT-qPCR)
The assay of qRT-PCR was performed as previously
described (20). Briefly, total
RNA was extracted from the cultured bEnd.3 cells (2×105
cells/ml) using TRIzol reagent (Invitrogen; Thermo Fisher
Scientific, Inc.), according to the manufacturer's instructions.
RNA was reverse transcribed using, and qPCR was performed with an
ABI 9700 PCR Thermal Cycler and an SYBR-Green kit (Bio-Rad
Laboratories, Inc., Hercules, CA, USA) according to the
manufacturer's instructions. Briefly, qRT-PCR was performed using
10 µP 2X SYBR-Green PCR Master Mix (Toyobo Life Science, Osaka,
Japan), with 5 µl of cDNA, 0.5 µl of primers and 4 µN of RNase-free
ddH2O contained in 20 µl of reaction mixture. The
reaction was performed with one cycle of 95°C for 5 min and 40
cycles of 95°C for 15 sec, 65°C for 15 sec and 72°C for 35 sec in
ABI 7300 real-time PCR instrument (Applied Biosystems; Thermo
Fisher Scientific, Inc.). When the reaction proceeded. Ct value was
obtained, and results were analyzed using 2−∆∆Cq
calculation (20). β-actin was
used to normalize the data. The primer sequences were: β-actin
forward, 5′-CATTGCTGACAGGATGCAGA-3′ and reverse,
5′-CTGCTGGAAGGTGGACAGTGA-3′; PPARγ forward,
5′-GGAAGACCACTCGCATTCCTT-3′ and reverse,
5′-GTAATCAGCAACCATTGGGTCA-3′; BIRC5 forward,
5′-GAGGCTGGCTTCATCCACTG-3′ and reverse,
5′-ATGCTCCTCTATCGGGTTGTC-3′. β-actin was used as an internal
control.
Apoptosis assay
Apoptotic rates were examined by flow cytometric
analysis using Annexin V staining kit (BD Pharmingen; BD
Biosciences). The transfected bEnd.3 cells (1×106
cells/ml) or untransfected bEnd.3 cells (1×106 cells/ml)
were collected by trypsinization and the suspensions centrifuged at
16,000 × g for 10 min at 4°C. Cells (1×106 cells/ml)
were resuspended in 1X binding buffer (BD Biosciences).
Subsequently, 100 µl of this solution (~1×105 cells) was
transferred to a 5-ml culture tube. Annexin V (5 µl) and propidium
iodide (5 µl; BD Biosciences), used for apoptosis signal detection,
were added to the samples, and then incubated for 15 min at room
temperature in the dark. A total of 400 ml 1X binding buffer was
added to each tube and the samples were immediately analyzed by BD
FACSCanto II flow cytometry (BD Biosciences). The data were
analyzed by FlowJo software version 8.8.6 (FlowJo LLC, Ashland, OR,
USA). For the Hoechst staining, treated or control cells were
seeded in 24-well plates at a concentration of 1×106
cells/well and incubated overnight at 37°C, and the DNA content of
cells in each well were stained with 100 µl of Hoechst 33342 for 30
min followed by DAPI staining for 10 min at room temperature and
visualized under a fluorescence microscope (Olympus, Tokyo, Japan).
Experiments were performed in triplicate.
Western blotting
Cell samples (2×106 cells/ml) were lysed
in 4°C for at least 30 min by radioimmunoprecipitation assay buffer
(Beyotime Institute of Biotechnology, Haimen, China) to obtain
total cell lysates. Protein concentrations were determined using
the Bicinchoninic Acid Protein assay kit (Thermo Fisher Scientific,
Inc.). Similar amounts of protein (40 µg) from each sample were
separated by 10% SDS-PAGE and transferred to polyvinylidene
fluoride membranes (Merck KGaA, Darmstadt, Germany). Membranes were
incubated with primary rabbit polyclonal antibodies against PPARγ
(ab59256, 1:1,000) and BIRC5 (ab469, 1:1500) (both from Abcam,
Cambridge, MA, USA) overnight at 4°C, followed by incubation with
horseradish peroxidase-conjugated monoclonal goat anti-rabbit
immunoglobulin (Ig)G (BA1054, 1:2,000; Boster Biological
Technology, Pleasanton, CA, USA) at room temperature for 2 h.
Membranes were stripped and reprobed with a primary monoclonal
mouse anti-rabbit antibody against GAPDH (KF703, 1:1,000; Nanjing
Jiancheng Bioengineering Institute, Nanjing, China,). Protein bands
were quantified by densitometry using the gel analysis software
ImageJ (National Institutes of Health, Bethesda, MD, USA).
Dual-luciferase reporter assay
The BIRC5 promoter binding site sequence (gene ID,
11799) for PPARγ was predicted using Ensembl (http://www.ensembl.org/Multi/Tools/Blast?db=core)
and NCBI (https://blast.ncbi.nlm.nih.gov/Blast.cgi?PROGRAM=blastn&PAGE_TYPE=BlastSearch&LINK_LOC=blasthome).
Wild-type (WT) BIRC5 promoter and a mutated (MUT) promoter sequence
containing the predicted target sites were synthesized and cloned
into the XbaI and FseI restriction sites of a pGL3
control vector (Promega, Madison, WI, USA); the constructs were
termed pGL3-promoter-WT and pGL3-promoter-MUT. In the reporter
assay experiment, bEnd.3 cells (1×103 cells/well) were
seeded onto 24-well plates and transfected with either
pGL3-promoter-WT or pGL3-promoter-MUT, and co-transfected with the
pcDNA4.1-PPARγ or pcDNA4.1 control vectors using Lipofectamine 2000
(Invitrogen; Thermo Fisher Scientific, Inc.), following the
manufacturer's instructions at 4°C for 2 h. A Renilla
luciferase vector, pRL-SV50 (Promega), was also co-transfected into
the cells and used to normalize the differences in firefly
luciferase activities. Following 48 h transfection, cells were
harvested and luciferase activities were measured with the
Dual-Luciferase Reporter Assay System (Promega), according to the
manufacturer's instructions. Transfections were repeated in
triplicate in three independent experiments.
Immunofluorescence assay
PPARγ protein expression levels were also evaluated
by immunofluorescence. Treated bEnd.3 cells (1×105
cells/ml) were seeded into 6-well plates overnight at 37°C. Then,
the cells were fixed by 4% paraformaldehyde for 24 h, and blocked
with 1% bovine serum albumin for 2 h at room temperature and
incubated with anti-PPARγ antibody (cat. no. ab209350, 1:200;
Abcam, Cambridge, UK) overnight at 4°C and then incubated with 1
µg/ml DAPI dihydrochloride; D9542; Sigma-Aldrich; Merck KGaA) at
room temperature for 10 min. Fluorescence images of six random
fields were captured on a Nikon Eclipse Ti-U fluorescence
microscope (Nikon, Tokyo, Japan) equipped with a SPOT-RTTM digital
camera (Diagnostic Instruments, Inc., Sterling Heights, MI, USA).
The fluorescent images were visualized with a Leica fluorescence
microscope (Leica Microsystems GmbH, Wetzlar, Germany). Each
experimental was replicated three times.
Electrophoretic mobility shift assay
(EMSA) and supershift assay
Nuclear proteins from bEnd.3 cells (1×106
cells/ml) were extracted using the NE-PER® Nuclear
Extraction Reagent (Thermo Fisher Scientific, Inc.) according to a
previously described protocol (21). The treated cells were washed for
three times with cold phosphate-buffered saline (PBS; pH 7.4), the
cells were then scraped and centrifuged at 15,000 × g for 5 min, to
collect the pellets. Biotin-labeled PPARγ specific oligonucleotides
(Invitrogen; Thermo Fisher Scientific, Inc.), with the following
sequence, 5′-AAAGGAGGTTAGAGGGGAAGGGGCGTAG-′3, were prepared as
labeled probes, according to the manufacturer's instructions
(Promega). Double-stranded oligonucleotides for PPARγ were
end-labeled with adenosine-5′-triphosphate (ATP)-γ-32P
using the T4 polynucleotide kinase (Promega), according to the
manufacturer's instructions. Biotin end-labeled double-stranded DNA
and the nuclear extracts were incubated at room temperature for 20
min, and then 10 µl protein-DNA complex was subjected to 6.5% PAGE
at 100 V for 1 h at 4°C and transferred onto a nylon membrane. The
radiolabeled probes were purified by spin columns (Roche Applied
Science, Penzberg, Germany). Nuclear protein extracts (5 µg) from
bEnd.3 cells were incubated with 100,000 cpm 32P-labeled
oligonucleotide probe in 25 mM HEPES (pH 7.4), 50 mM KCl, 10%
glycerol (v/v), 5 mM dithiothreitol and 1 µg of poly
(deoxyinosinic-deoxycytidylic) acid (GE Healthcare Life Sciences,
Shanghai, China) for 30 min at room temperature in a final volume
of 20 µl. Following binding, protein-DNA complexes were separated
on a 6% non-denaturing polyacrylamide gel at 120 V in 0.5X
Tris/borate/EDTA buffer. Gels were analyzed with a PhosphorImager
Gel Imaging System (Bio-Rad Laboratories, Inc.). For the antibody
supershift analysis, 1 µg of antibody against PPARγ (ab59256;
Abcam) was added to the nuclear extracts at a dilution of 1:1,000
for 16 h prior to addition of the radiolabeled
oligonucleotides.
Chromatin immunoprecipitation (ChIP)
assay
Cells (5×107) were fixed with 4%
paraformaldehyde for 10 min at 37°C and a ChIP assay was performed
with the EZ-ChIP assay kit (Sigma-Aldrich; Merck KGaA), as
previously described (22).
Briefly, Pellet cells were resuspended by SDS lysis buffer with 1%
protease inhibitor cocktail set III EDTA-free (Calbiochem; Merck
KGaA) and incubated for 10 min on ice. Sonicate lysate to shear DNA
to lengths between 500–1,000 bp which were detected by 1% ethidium
bromide gel electrophoresis. The conditions have been optimized
following steps number of bursts: 8, length of bursts: 10 sec,
interval time: 10 sec, output control setting: 30%, duty cycle:
constant. The lysates were incubated with anti-PPARγ antibody
(ab59256, 1:10; Abcam) or a rabbit control IgG (ab6789, 1:500;
Abcam) for 24 h at 4°C, and the complexes were isolated using
protein A-agarose/salmon sperm DNA (EMD Millipore, Billerica, MA,
USA). Immunoprecipitates were added in 1 ml of high-salt wash
buffer for ChIP to all samples and rotated for 10 min at room
temperature; the samples were centrifuged at 4500 × g for 2 min at
room temperature; the supernatants were carefully aspirated and
added 1 ml of high-salt wash buffer for ChIP; the samples were
rotated for 10 min at room temperature; the above two steps were
repeated twice in a total of four high-salt washes; the
supernatants were aspirated and washed twice with TE as above.
Immunoprecipitates were subsequently eluted with freshly prepared
1% SDS + 0.1 M NaHCO3 buffer. To the immune complexes
were added 20 µl 5 M NaCl and histone-DNA crosslinks reversed by
heating at 65°C for 4 h. Following the reversing of crosslinking,
DNA was purified with the QIAquick PCR Purification kit (Qiagen
GmbH, Hilden, Germany) followed by PCR amplification. The
amplification reactions were performed using Amplitaq DNA
polymerase, GeneAmp dNTPs (deoxynucleoside triphosphates) with
dUTP, and AmpErase UNG (all from PerkinElmer, Inc., Waltham, MA,
USA). The thermocycling conditions were a thermal cycler preheated
to 94°C; and then 94°C for 1 min, 55°C for 1 min, 72°C for 2 min
for 30 cycles and a final extension at 72°C for 7 min. The
PCR-amplified products were examined by electrophoresis in a 1.5%
agarose gel, stained with a 1% solution of ethidium bromide, and
examined under ultraviolet illumination. Primers used to amplify
the PPARγ binding site area were: Forward,
5′-TCCCTTCCAACCTCCCAAT-3′ and reverse, 5′-AGCCCAGTATCCCAAATCAAC-3′,
which resulted in a 98 bp fragment. PCR products were resolved by
2% agarose gel electrophoresis, visualized by ethidium bromide
staining and analyzed by densitometry using ImageJ software version
1.37 (National Institutes of Health, Bethesda, MD, USA). To ensure
the specificity of each assay, DNA binding in normal IgG
immunoprecipitates was regarded as the background control.
Statistical analysis
All experiments were performed at least three times,
and all samples were tested in triplicate. Experimental data are
displayed as the mean ± standard deviation. All analyses were
performed using one-way analysis of variance or an unpaired
Student's t-test performed on SPSS software, version 12.0 (SPSS,
Inc., Chicago, IL, USA). P<0.05 was considered to indicate a
statistically significant difference.
Results
OGD reduces proliferation and enhances
apoptosis of bEnd.3 cells
To establish a cerebral microvascular endothelial
cell injury model with cerebral ischemia, bEnd.3 cells were exposed
to OGD conditions to mimic ischemia-like conditions in
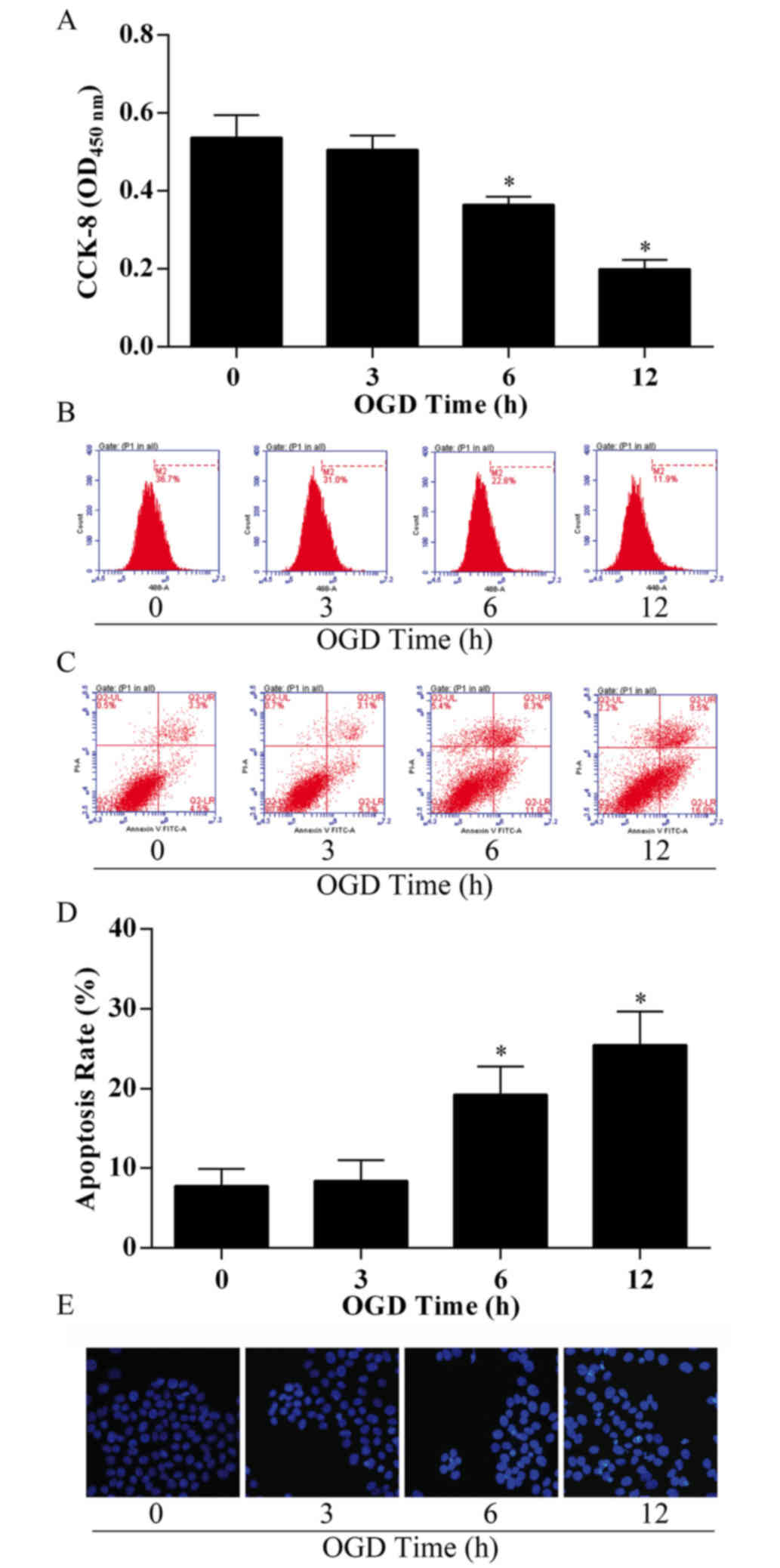
vitro. Following OGD treatment, bEnd.3 cells exhibited a
significant decrease in proliferation capacity in a time-dependent
manner (Fig. 1A), with the group
treated for 12 h exhibiting ~60% loss in cell viability compared
with the control group (treated for 0 h). To confirm cell
viability, the proliferation capacity of bEnd.3 cells
post-treatment was examined by EdU incorporation and flow
cytometry; the M2 phase rate was 11.9%, much lower compared with
the control group (36.7%; Fig.
1B). The results demonstrated that viability in bEnd.3 cells
significantly decreased following OGD treatment in a time-dependent
manner.
The effects of OGD treatment on apoptosis in bEnd.3
cells were examined using the apoptosis assay by flow cytometry
analysis and Annexin V/PI staining. OGD-treated cells exhibited an
increase in the proportion of early apoptotic cells (5.3% in 3 h
treatment group, 11.0% in 6 h treatment group and 16.0% in 12 h
treatment group) and late apoptotic cells (3.1% in 3 h treatment
group, 8.3% in 6 h treatment group, 9.5% in 12 h treatment group),
compared with 4.5% (early) and 3.3% (late) in the 0 h control group
(Fig. 1C), which resulted in an
overall threefold enhancement in total cell apoptosis in the 12 h
treatment group compared with the control (P<0.05; Fig. 1D). Following incubation and
treatment, bEnd.3 cell nuclei were stained with Hoechst, and
notable changes in cell apoptosis were observed, as the Hoechst
nuclear staining became increasingly bright with the longer
durations of OGD treatment (Fig.
1E).
PPARγ expression is inhibited by OGD
treatment
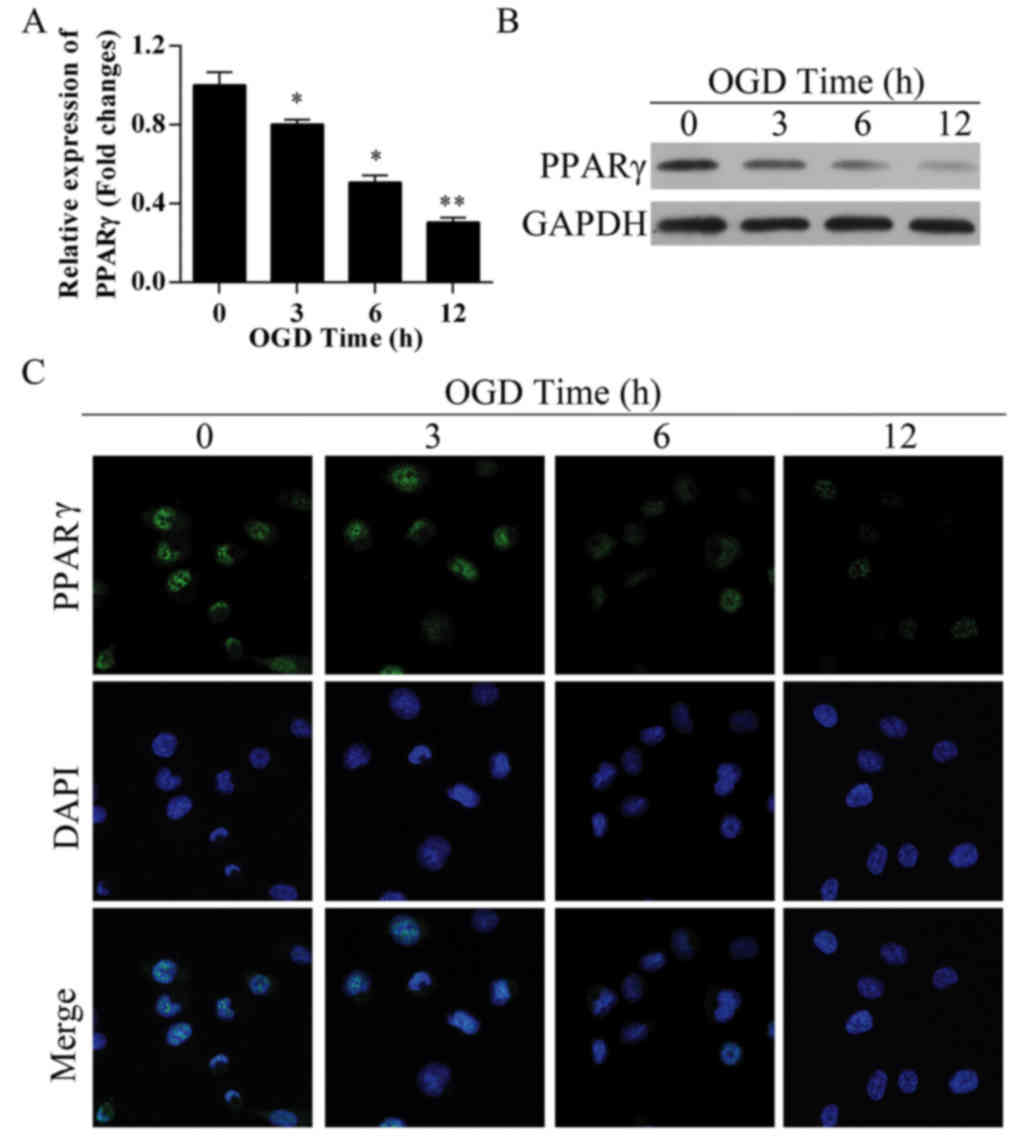
To evaluate the status of PPARγ expression following
OGD treatments of bEnd.3 cells, PPARγ mRNA and protein expression
levels were detected. RT-qPCR analysis and western blotting
revealed that the OGD treatment led to reduction in PPARγ mRNA and
protein expression levels (Fig. 2A and
B, respectively), and this decrease occurred in a
time-dependent manner. To examine the PPARγ protein expression
status in situ, immunofluorescence staining was performed
(Fig. 2C). Untreated bEnd.3 cells
(0 h control) exhibited uniform distribution of PPARγ expression
compared with OGD-treated cells, which obtained exhibited a faint
staining pattern. These results confirmed that PPARγ expression was
inhibited in bEnd.3 cells treated with OGD.
PPARγ protects brain endothelium from
ischemic apoptosis
A previous report suggested that PPARγ activation
may protect neural cells following cerebral ischemia (23); therefore, the present study
hypothesized that the upregulation of PPARγ gene expression may
also relieve cerebral microvascular endothelial cells from cerebral
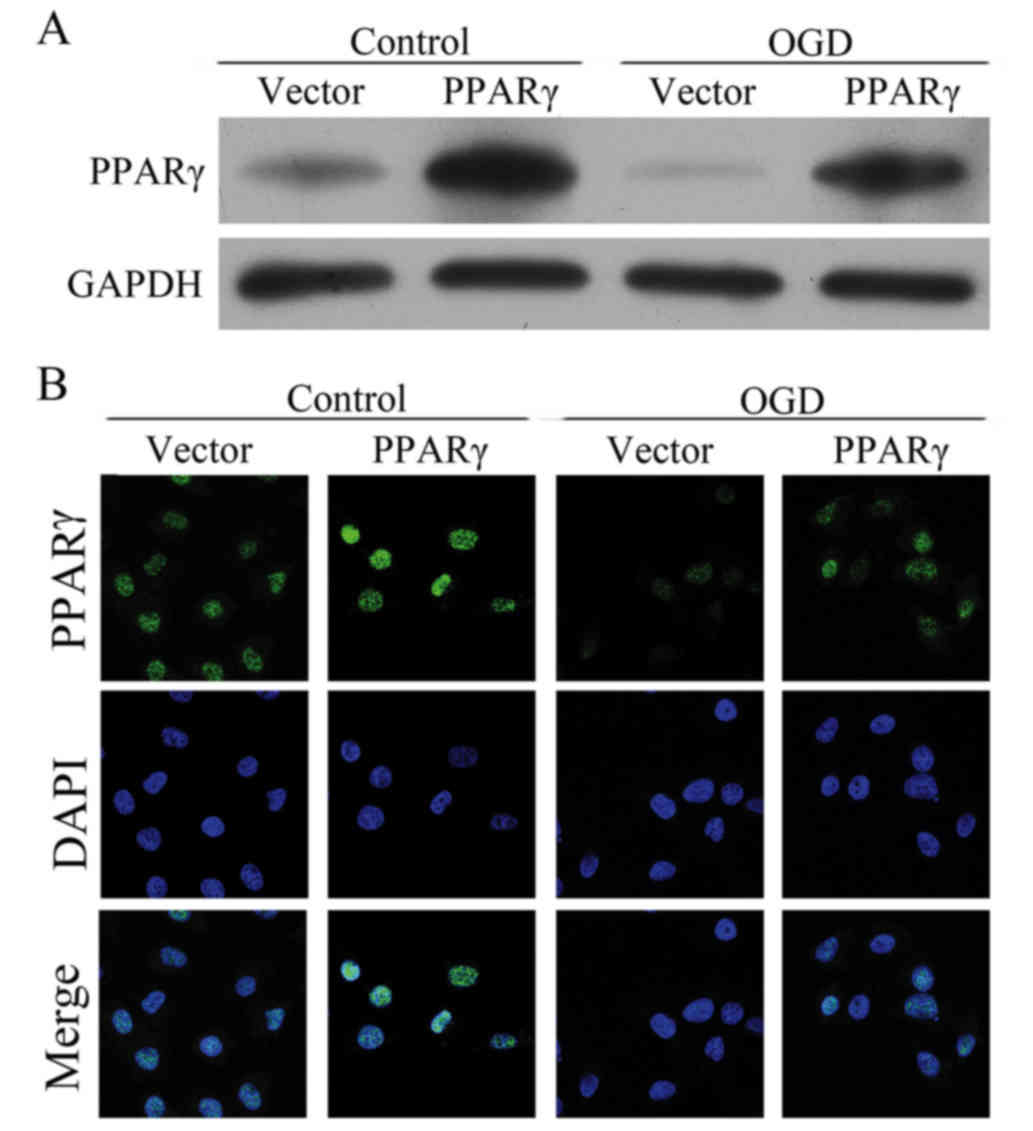
ischemia injury. To examine this, bEnd.3 cells were transfected
with pcDNA4.1-PPARγ overexpression plasmid, which exhibited a
notable increase in PPARγ protein expression levels compared with
cells transfected with the pcDNA4.1-empty vector control, in the
presence or absence of OGD treatment (Fig. 3A). No significant difference was
observed between bEnd.3 cells transfected with pcDNA4.1-PPARγ
overexpression plasmid and pcDNA4.1-empty vector control without
the treatment of OGD. However, in the presence of OGD treatment,
the PPARγ expression was significantly higher in the PPARγ
overexpressed bEnd.3 cells compared with the vector group (Fig. 3B).
PPARγ overexpression alleviates bEnd.3
cell death caused by 12 h OGD exposure
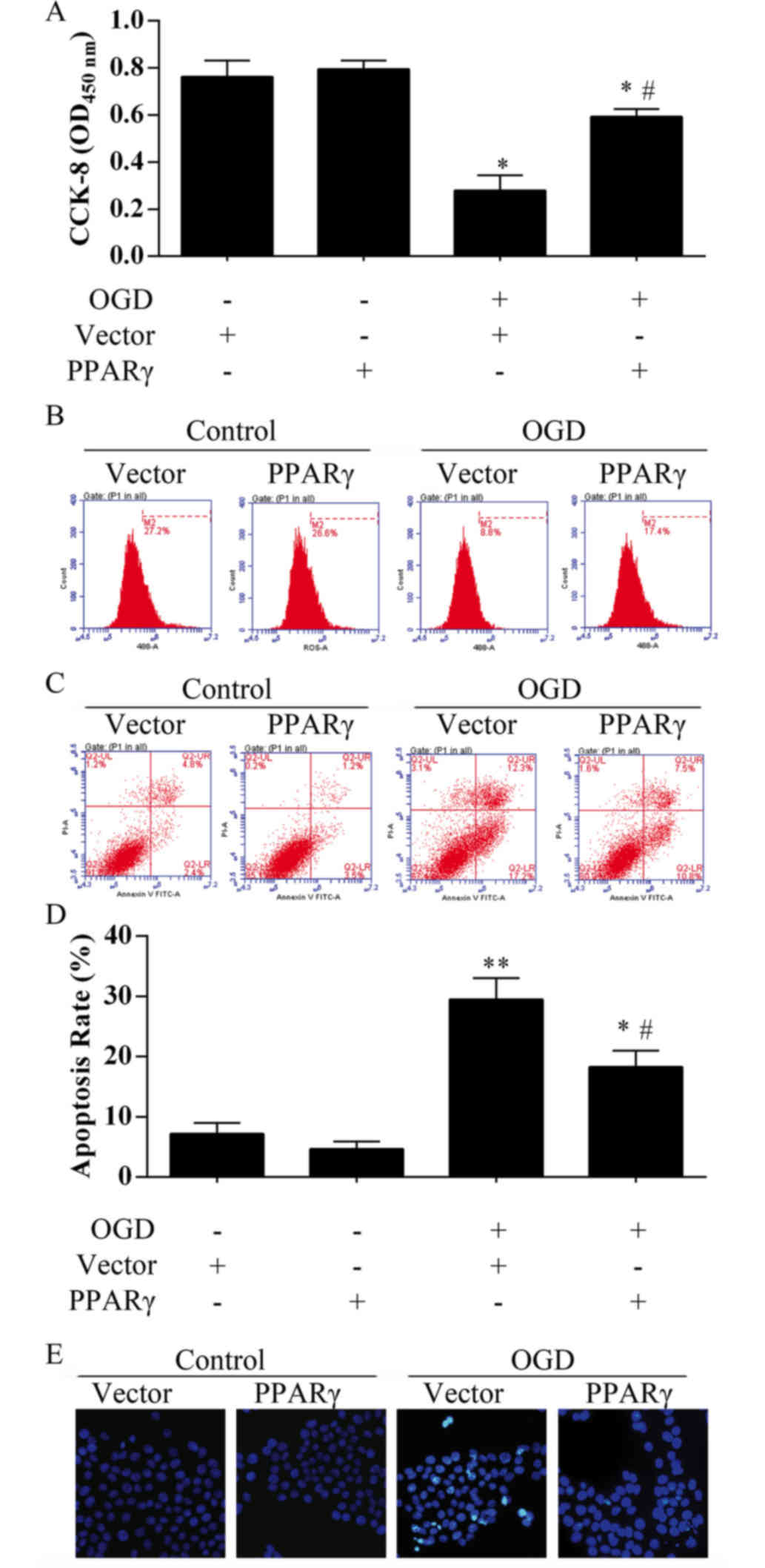
Results from live cell counts demonstrated that the
cell viability recovered more than 50% in the bEnd.3 cells
transfected with pcDNA4.1-PPARγ overexpression plasmid compared
with those cells transfected with the empty vector (Fig. 4A). Proliferation capacity of bEnd.3
cells was also analyzed by EdU incorporation assay (Fig. 4B); M2 phase rate was notably higher
(~17%) in OGD-treated cells that were transfected with the PPARγ
overexpression plasmid compared with OGD-treated cells transfected
with the empty vector (8.8%). No significant differences in
viability were identified in bEnd.3 cells transfected with
pcDNA4.1-PPARγ plasmid compared with the control groups (Fig. 4A and B). In the Annexin V/PI
apoptosis assay, the proportion of total apoptotic cells was
observed to be 7.2% in the control group, 4.7% in the PPARγ
overexpression group, 29.5% in the OGD treatment group and 18.3% in
the OGD-treated cells that overexpressed PPARγ (Fig. 4C and D), which indicated that PPARγ
may significantly inhibit apoptosis under ischemia-like conditions.
Hoechst nuclear staining also demonstrated that PPARγ
overexpression may rescue bEnd.3 cell death following OGD treatment
(Fig. 4E).
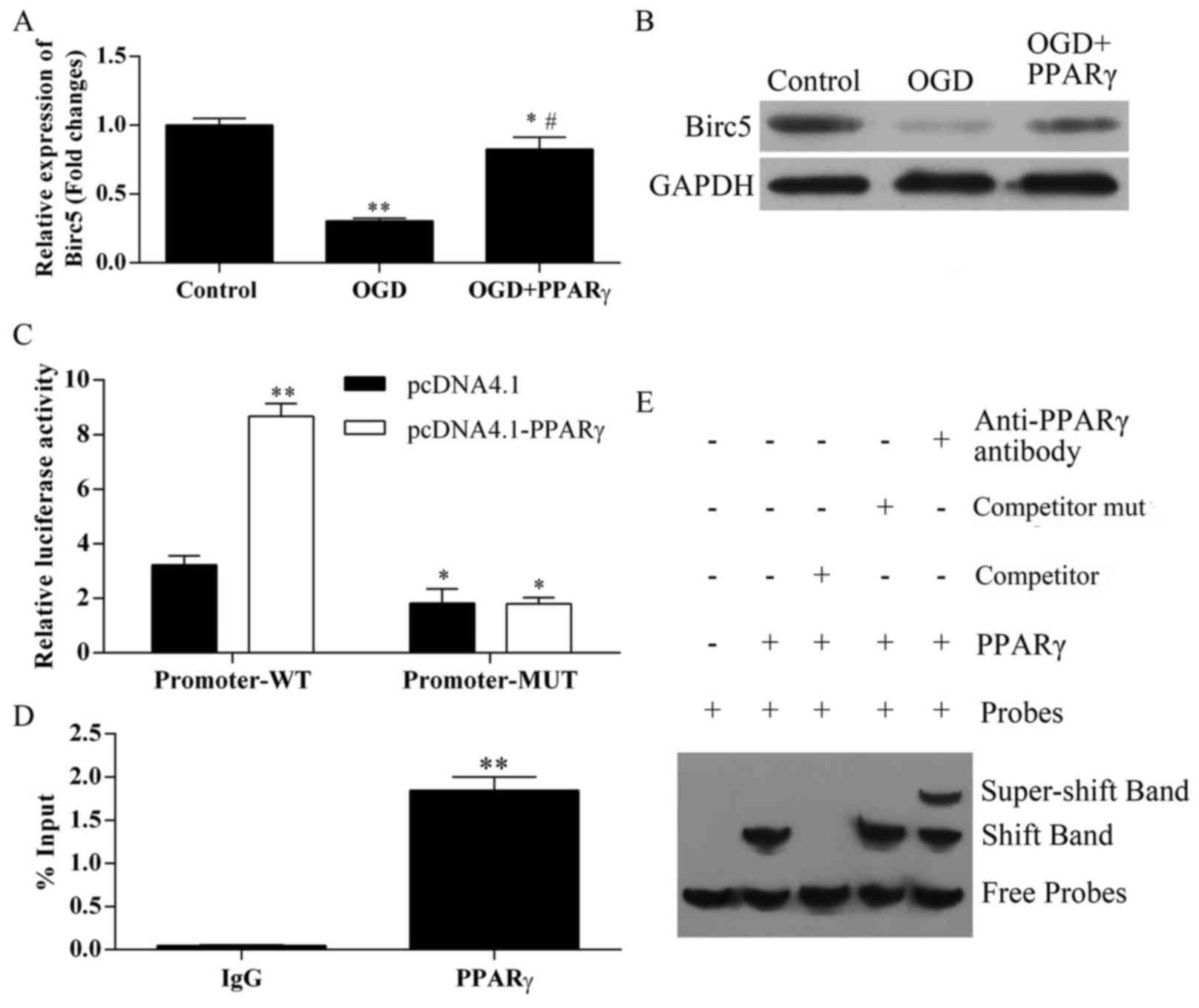
BIRC5 expression is regulated by PPARγ
during ischemia
To further determine the molecular mechanisms
responsible for PPARγ-mediated protective roles in the OGD
treatment process, the expression of another critical factor that
also serves important roles in ischemic apoptosis, BIRC5, was
examined. BIRC5 mRNA expression was significantly decreased in
bEnd.3 cells following OGD treatment, compared with control cells
(Fig. 5A); this reduced expression
was recovered in OGD-treated cells that overexpressed PPARγ.
Similar results were obtained in western blot analyses of BIRC5
protein expression (Fig. 5B).
These results indicated that BIRC5 expression may be regulated at
both the mRNA and the protein level by PPARγ during ischemic
conditions.
BIRC5 is a target of PPARγ
regulation
To elucidate the mechanisms of PPARγ regulation on
BIRC5, a functional analysis was performed to verify the potential
PPARγ binding sites in the BIRC5 promoter. The transcriptional
responses of the BIRC5 pGL-promoter-WT and pGL-promoter-MUT
plasmids were analyzed using an in vitro luciferase
transcriptional assay (2).
pcDNA4.1-PPARγ overexpression significantly increased the
transcriptional activity of the pGL-promoter-WT, compared with
cells co-transfected with the empty pcDNA4.1 vector (Fig. 5C). Conversely, no significant
differences were identified in the luciferase activities of the
pGL-promoter-MUT group co-transfected with the PPARγ overexpression
vector.
ChIP analysis was applied to verify the interaction
between PPARγ and BIRC5 promoter. Consistent with previously
reported transcriptional activity, a significant increase in PPARγ
binding to the BIRC5 promoter site was detected, with isotypic IgG
antibody used as a negative immunoprecipitation control (P<0.01;
Fig. 5D).
EMSA supershift assay was applied to determine
whether PPARγ was able to bind to the BIRC5 promoter. The PPARγ
protein formed a complex band (shift band) using probes (Fig. 5E). By contrast, the PPARγ protein
competitor prevented the formation of the shift band, which
indicated that it interfered DNA binding. Specificity of binding
was examined with a mutated competitor, which failed to elicit
competition, as demonstrated in the unaltered band formation. The
specificity of the complex was reconfirmed using a PPARγ antibody
that supershifted the PPARγ + BIRC5 band. These data suggested that
BIRC5 may be a novel target of PPARγ transcriptional
regulation.
Discussion
Cerebral microvascular endothelial cells serve a
major role in ischemic insult of the brain, and regulate the
trafficking of cells, substrates and other molecules across the
blood-brain barrier, vasomotor reactivity and homeostasis at the
interface of the blood/vascular wall. Neurovascular protection is
considered as an effective part of stroke therapy (6,24).
Elucidation of the underlying mechanism of different regulators and
bEnd.3 cells may provide new insights into the cerebral vasculature
and provide a novel therapeutic strategy for the treatment of
diseases such as stroke.
Cell culture models of cerebrovascular endothelium
are essential for exploring the molecular mechanisms of ischemic
injury (18,25–27).
The present study established an OGD-induced apoptotic injury model
using the mouse microvascular endothelial cell line bEnd.3 and
found the most suitable duration time (12 h) of this model, which
may facilitate the future study of ischemic injury. Similar to
previous studies (3,12,28),
the present study revealed that PPARγ protein expression was
reduced in this established model. In order to further investigate
the biological functions of PPARγ, a PPARγ overexpressed cell model
was established by transfecting bEnd.3 cells with PPARγ
overexpression plasmid. Cell proliferation and apoptosis assay
demonstrated that PPARγ may act as a vascular protective agent in
the OGD treated cell model. PARγ activation has been reported to
stimulate proliferation and attenuate apoptosis in endothelial
progenitor cells through PPARγ-dependent signaling cascades
(8,29,30).
Results from another study also indicated that PPARγ may inhibit
H2O2-induced apoptosis of bEnd.3 cells by
upregulating the expression level of 14-3-3 (5,10).
It is hypothesized that PPARγ and its ligands may serve as
protective agents in the brain during stroke (31).
BIRC5 is a known survival factor in studies on
embryogenesis and oncology (11,32),
and was recently revealed to serve an important role in cerebral
microvascular endothelial cell injury (33,34).
However, the regulatory function of PPARγ on BIRC5 was not widely
recognized. The present study demonstrated that PPARγ upregulation
regulated the expression of BIRC5 both transcriptionally and
post-translationally in OGD-treated cells. In addition, the
upregulation of BIRC5 may be involved in the acquired resistance
from various of noxious stimulations, such as ischemia and other
lesions (35). The present study
was the first, to the best of the authors' knowledge, to reveal
that BIRC5 can be regulated by PPARγ via directly binding in
ischemia. In conclusion, the present results demonstrated a
cerebrovascular protective role of PPARγ in an ischemia model and
identified BIRC5 as a novel target in the pathogenesis of ischemic
vascular injury. Therefore, pharmacological activation of either
PPARγ or BIRC5 expression may provide potentially therapeutic
options for vascular damage induced by ischemia in clinical
treatment. The mechanisms of PPARγ-mediated protection in
endothelial cell damage explained in this study may aid in further
understanding the pathogenesis and therapy of cerebral
ischemia.
Previous studies have demonstrated that IRC5 may be
a pejorative prognostic marker in stage II/III breast cancer
(36), that silencing of BIRC5
induces neuroblastoma apoptosis (37). BIRC5 is involved in the biological
processes of colorectal cancer (38) and can serve as a serum diagnostic
and prognostic biomarker of colorectal cancer (39). Silencing BIRC5 promotes hepatoma
cell apoptosis (40). However, the
expression level of BIRC5 in endothelial cells treated with OGD
following PPARγ treatment and the relationship between BIRC5 and
PPARγ remain to be elucidated. The present study identified that
OGD treatment markedly decreased BIRC5 expression in bEnd.3 cells,
and that overexpression of PPARγ recovered this reduction mediated
by OGD. In addition, it was identified that PPARγ increased the
transcriptional activity of BIRC5, which suggested that BIRC5 is a
target of PPARγ regulation. ChIP assay also indicated that there
was a significant increase in PPARγ binding to the BIRC5 promoter
site, suggesting that PPARγ can interact with BIRC5 promoter. EMSA
supershift assay further showed that PPARγ can bind to the BIRC5
promoter. Therefore, it was demonstrated that BIRC5 may be a novel
target of PPARγ transcriptional regulation.
References
|
1
|
Pan Q, He C, Liu H, Liao X, Dai B, Chen Y,
Yang Y, Zhao B, Bihl J and Ma X: Microvascular endothelial
cells-derived microvesicles imply in ischemic stroke by modulating
astrocyte and blood brain barrier function and cerebral blood flow.
Mol Brain. 9:632016. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Xu F, Li J, Ni W, Shen YW and Zhang XP:
Peroxisome proliferator-activated receptor-γ agonist
15d-prostaglandin J2 mediates neuronal autophagy after cerebral
ischemia-reperfusion injury. PLoS One. 8:e550802013. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Wu JS, Tsai HD, Cheung WM, Hsu CY and Lin
TN: PPAR-γ ameliorates neuronal apoptosis and ischemic brain injury
via suppressing NF-κB-Driven p22phox transcription. Mol Neurobiol.
53:3626–3645. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Zhang Y, Zhang X, Park TS and Gidday JM:
Cerebral endothelial cell apoptosis after ischemia-reperfusion:
Role of PARP activation and AIF translocation. J Cereb Blood Flow
Metab. 25:868–877. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Fong WH, Tsai HD, Chen YC, Wu JS and Lin
TN: Anti-apoptotic actions of PPAR-gamma against ischemic stroke.
Mol Neurobiol. 41:180–186. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Ketsawatsomkron P and Sigmund CD:
Molecular mechanisms regulating vascular tone by peroxisme
proliferator activated receptor gamma. Curr Opin Nephrol Hyperten.
24:123–130. 2015. View Article : Google Scholar
|
|
7
|
Chu SF, Zhang Z, Zhang W, Zhang MJ, Gao Y,
Zuo W, Huang HY and Chen NH: Upregulating the expression of
survivin-HBXIP complex contributes to the protective role of
IMM-H004 in transient global cerebral ischemia/reperfusion. Mol
Neurobiol. 54:524–540. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Zhou Y, Jia S, Wang C, Chen Z, Chi Y, Li
J, Xu G, Guan Y and Yang J: FAM3A is a target gene of peroxisome
proliferator-activated receptor gamma. Biochim Biophysica Acta.
1830:4160–4170. 2013. View Article : Google Scholar
|
|
9
|
Romera C, Hurtado O, Mallolas J, Pereira
MP, Morales JR, Romera A, Serena J, Vivancos J, Nombela F, Lorenzo
P, et al: Ischemic preconditioning reveals that GLT1/EAAT2
glutamate transporter is a novel PPARgamma target gene involved in
neuroprotection. J Cereb Blood Flow Metab. 27:1327–1338. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Wu JS, Cheung WM, Tsai YS, Chen YT, Fong
WH, Tsai HD, Chen YC, Liou JY, Shyue SK, Chen JJ, et al:
Ligand-activated peroxisome proliferator-activated receptor-gamma
protects against ischemic cerebral infarction and neuronal
apoptosis by 14-3-3 epsilon upregulation. Circulation.
119:1124–1134. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Fukuda S and Pelus LM: Survivin, a cancer
target with an emerging role in normal adult tissues. Mol Cancer
Ther. 5:1087–1098. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Zhang Y, Park TS and Gidday JM: Hypoxic
preconditioning protects human brain endothelium from ischemic
apoptosis by Akt-dependent survivin activation. Am J Physiol Heart
Circ Physiol. 292:H2573–H2581. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Lu M, Kwan T, Yu C, Chen F, Freedman B,
Schafer JM, Lee EJ, Jameson JL, Jordan VC and Cryns VL: Peroxisome
proliferator-activated receptor gamma agonists promote
TRAIL-induced apoptosis by reducing survivin levels via cyclin
D3 repression and cell cycle arrest. J Biol Chem.
280:6742–6751. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wang Y, Tan H, Xu D, Ma A, Zhang L, Sun J,
Yang Z, Liu Y and Shi G: The combinatory effects of PPAR-γ agonist
and survivin inhibition on the cancer stem-like phenotype and cell
proliferation in bladder cancer cells. Int J Mol Med. 34:262–268.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Yamashita-Sugahara Y, Tokuzawa Y, Nakachi
Y, Kanesaki-Yatsuka Y, Matsumoto M, Mizuno Y and Okazaki Y: Fam57b
(family with sequence similarity 57, member B), a novel peroxisome
proliferator-activated receptor γ target gene that regulates
adipogenesis through ceramide synthesis. J Biol Chem.
288:4522–4537. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Cao G, Jiang N, Hu Y, Zhang Y, Wang G, Yin
M, Ma X, Zhou K, Qi J, Yu B and Kou J: Ruscogenin attenuates
cerebral ischemia-induced blood-brain barrier dysfunction by
suppressing TXNIP/NLRP3 inflammasome activation and the MAPK
pathway. Int J Mol Sci. 17:E14182016. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Gentil BJ, Minotti S, Beange M, Baloh RH,
Julien JP and Durham HD: Normal role of the low-molecular-weight
neurofilament protein in mitochondrial dynamics and disruption in
charcot-marie-tooth disease. FASEB J. 26:1194–1203. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Ma X, Zhang H, Pan Q, Zhao Y, Chen J, Zhao
B and Chen Y: Hypoxia/aglycemia-induced endothelial barrier
dysfunction and tight junction protein downregulation can be
ameliorated by citicoline. PLoS One. 8:e826042013. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Li YN, Pan R, Qin XJ, Yang WL, Qi Z, Liu W
and Liu KJ: Ischemic neurons activate astrocytes to disrupt
endothelial barrier via increasing VEGF expression. J Neurochem.
129:120–129. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Hussain AR, Ahmed SO, Ahmed M, Khan OS, Al
Abdulmohsen S, Platanias LC, Al-Kuraya KS and Uddin S: Cross-talk
between NF-κB and the PI3-kinase/AKT pathway can be targeted in
primary effusion lymphoma (PEL) cell lines for efficient apoptosis.
PLoS One. 7:e399452012. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Yin KJ, Deng Z, Hamblin M, Xiang Y, Huang
H, Zhang J, Jiang X, Wang Y and Chen YE: Peroxisome
proliferator-activated receptor delta regulation of miR-15a in
ischemia-induced cerebral vascular endothelial injury. J Neurosci.
30:6398–6408. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Arsenijevic D, de Bilbao F, Plamondon J,
Paradis E, Vallet P, Richard D, Langhans W and Giannakopoulos P:
Increased infarct size and lack of hyperphagic response after focal
cerebral ischemia in peroxisome proliferator-activated receptor
beta-deficient mice. J Cereb Blood Flow Metab. 26:433–445. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Zhang Y, Wang T, Yang K, Xu J, Ren L, Li W
and Liu W: Cerebral microvascular endothelial cell apoptosis after
ischemia: Role of enolase-phosphatase 1 activation and
aci-reductone dioxygenase 1 translocation. Front Mol Neurosci.
9:792016. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Sun H, Xiong W, Arrick DM and Mayhan WG:
Low-dose alcohol consumption protects against transient focal
cerebral ischemia in mice: Possible role of PPARγ. PLoS One.
7:e417162012. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Deng Y, Xiong D, Yin C, Liu B, Shi J and
Gong Q: Icariside II protects against cerebral ischemia-reperfusion
injury in rats via nuclear factor-κB inhibition and peroxisome
proliferator-activated receptor up-regulation. Neurochem Int.
96:56–61. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Green DE, Sutliff RL and Hart CM: Is
peroxisome proliferator-activated receptor gamma (PPARγ) a
therapeutic target for the treatment of pulmonary hypertension?
Pulm Circ. 1:33–47. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Mysiorek C, Culot M, Dehouck L, Derudas B,
Staels B, Bordet R, Cecchelli R, Fenart L and Berezowski V:
Peroxisome-proliferator-activated receptor-alpha activation
protects brain capillary endothelial cells from oxygen-glucose
deprivation-induced hyperpermeability in the blood-brain barrier.
Curr Neurovasc Res. 6:181–193. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Chen YC, Wu JS, Tsai HD, Huang CY, Chen
JJ, Sun GY and Lin TN: Peroxisome proliferator-activated receptor
gamma (PPAR-γ) and neurodegenerative disorders. Mol Neurobiol.
46:114–124. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Zeng Y, Xie K, Dong H, Zhang H, Wang F, Li
Y and Xiong L: Hyperbaric oxygen preconditioning protects cortical
neurons against oxygen-glucose deprivation injury: Role of
peroxisome proliferator-activated receptor-gamma. Brain Res.
1452:140–150. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Gupta D, Jetton TL, Mortensen RM, Duan SZ,
Peshavaria M and Leahy JL: In vivo and in vitro studies of a
functional peroxisome proliferator-activated receptor gamma
response element in the mouse pdx-1 promoter. J Biol Chem.
283:32462–32470. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
He F, Qu F and Song F: Aspirin upregulates
the expression of neuregulin 1 and survivin after focal cerebral
ischemia/reperfusion in rats. Exp Ther Med. 3:613–616. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Liu ZJ, Liu HQ, Xiao C, Fan HZ, Huang Q,
Liu YH and Wang Y: Curcumin protects neurons against oxygen-glucose
deprivation/reoxygenation-induced injury through activation of
peroxisome proliferator-activated receptor-γ function. J Neurosci
Res. 92:1549–1559. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Liu N, Zhang Y, Fan L, Yuan M, Du H, Cheng
R, Liu D and Lin F: Effects of transplantation with bone
marrow-derived mesenchymal stem cells modified by Survivin on
experimental stroke in rats. J Transl Med. 9:1052011. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Fiatte C, Huin C, Collet P, Plenat F,
Dauca M and Schohn H: Expression of PPARgamma is reduced by medium
supplementation with L-glutamine in human colorectal Caco-2 cells.
Int J Mol Med. 22:825–832. 2008.PubMed/NCBI
|
|
36
|
Hamy AS, Bieche I, Lehmann-Che J, Scott V,
Bertheau P, Guinebretiere JM, Matthieu MC, Sigal-Zafrani B, Tembo
O, Marty M, et al: BIRC5 (survivin): A pejorative prognostic marker
in stage II/III breast cancer with no response to neoadjuvant
chemotherapy. Breast Cancer Res Treat. 159:499–511. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Lamers F, Van der Ploeg I, Schild L, Ebus
ME, Koster J, Hansen BR, Koch T, Versteeg R, Caron HN and Molenaar
JJ: Knockdown of survivin (BIRC5) causes apoptosis in neuroblastoma
via mitotic catastrophe. Endocr Relat Cancer. 18:657–668. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Li PL, Zhang X, Wang LL, Du LT, Yang YM,
Li J and Wang CX: MicroRNA-218 is a prognostic indicator in
colorectal cancer and enhances 5-fluorouracil-induced apoptosis by
targeting BIRC5. Carcinogenesis. 36:1484–1493. 2015.PubMed/NCBI
|
|
39
|
Wang H, Zhang X, Wang L, Zheng G, Du L,
Yang Y, Dong Z, Liu Y, Qu A and Wang C: Investigation of cell free
BIRC5 mRNA as a serum diagnostic and prognostic biomarker for
colorectal cancer. J Surg Oncol. 109:574–579. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Wang Q, Shu R, He H, Wang L, Ma Y, Zhu H,
Wang Z, Wang S, Shen G and Lei P: Co-silencing of Birc5 (survivin)
and Hspa5 (Grp78) induces apoptosis in hepatoma cells more
efficiently than single gene interference. Int J Oncol. 41:652–660.
2012. View Article : Google Scholar : PubMed/NCBI
|