Introduction
Gastric cancer is the second most common malignancy
in China with highest mortality (1).
Despite advanced techniques introduced in surgery, medications and
targeted therapy, the overall survival rate and prognosis remains
unsatisfactory in patients with gastric cancer (2,3). This may
be caused by anti-apoptosis of cancer cells (4–7). In order
to enhance prognosis and improve the outcome of treatment,
effective suppression of the anti-apoptotic cells in gastric
carcinoma may be needed. Therefore, it can be meaningful to
identify new genes involved in apoptosis in gastric cancer. LMO1
gene has been reported to have a close relation with
lelakemogenesis (8) with
characteristics of proto-oncogene. However, little research has
been done to explore LMO1 expression and also the mechanism of LMO1
in gastric cancer apoptosis remains ambiguous. In this study, in
order to investigate the clinical value and the mechanism of LOM1
in gastric carcinoma, we examined the LOM1 expression level in
gastric cancer tissues and cell lines, also the relation between
LMO1 expression and clinicopathological features in patients with
gastric cancer was investigated and the correlation between LOM1
gene and genes associated with apoptosis in terms of Bcl-2 and Bax
was analyzed. Moreover, variations of cell viability, apoptosis
rate, and Bcl-2, Bax genes were measured to determine the molecular
mechanism of LMO1 in gastric cancer after LMO1-siRNA was
synthesized and transfected to MKN45 gastric cancer cell line.
Materials and methods
Patients and tissue samples
Gastric cancer cases were enrolled from Hebei
General Hospital (Shijiazhuang, China) between January 2011 and
December 2011, diagnosed of gastric cancer and underwent surgery. A
total of 96 patients were recruited consisting of 66 males and 30
females, with median age of 58 years (range, 32–81 years). Patients
were considered eligible for the present study if they i) had a
primary gastric cancer and underwent a tumorectomy; ii) had gastric
cancer with no distant metastasis; and iii) had not be treatment of
radiation/chemotherapy or targeted therapy before the tumorectomy.
Patients were excluded if they met one of these criteria: i) had
recurrent gastric cancer; ii) had gastric cancer with other
malignancies; or iii) dissented to their participation. All
paticipants had complete clinicopathological data and follow-ups
for 60 months after surgery. The patient follow-ups ended in
December 2015. Samples were obtained from tumor tissues and
adjacent normal tissues (3 cm distance from the tumor margin
without traces of cancerous cells under microscope).
Paraffin-embedded samples were prepared for immunohistochemistry
(IHC) after fixed with 10% polyoxymethylene. Fresh cancer and
adjacent normal tissues of 20 gastric cancer patients, including 14
males and 6 females (median age of 56.5 years; range, 41–72 years),
were collected from February 2017 to May 2017. Informed consent was
obtained from participants, and approval from the Institutional
Review Board at our hospital was attained.
Cell lines and reagents
Human gastric cancer cell lines including: Hixed
gastric adenocarcinoma cells MKN28 (which are a derivative of MKN74
cells) (9), MKN45, SGC-7901, BGC823,
MGC803 and human normal gastric epithelial cell lines GES-1 were
purchased from Academy of Military Medical Science (Beijing,
China). Rabbit anti-human LMO1, Bcl-2, Bax and GAPDH polyclonal
antibody were the products of Santa Cruz Co. (Dallas, TX, USA).
Horseradish peroxidase-labeled IgG antibodies and the BCA Protein
Assay kit were purchased from Zhong Shan-Golden Bridge Biological
Technology (Beijing, China). Fetal bovine serum (FBS), PRMI1640
medium, Lipofectamine™ 2000, TRIzol and Real-time PCR
kits were supplied by Invitrogen (Carlsbad, CA, USA), while
LMO1-siRNA, negative control siRNA and PCR primers were synthesized
by GenePharma Co. (Shanghai, China). Cell Counting kit-8 (CCK8) was
purchased from Boster Biological Technology (Wuhan, China).
IHC staining for detection of LMO1,
Bcl-2 and Bax proteins in gastric cancer tissues and adjacent
tissues
Paraffin-embedded tissue sections were dewaxed and
rehydrated followed by incubation in 3% H2O2,
washing and antigen-retrieval methods. Then IHC assay was applied
according to the manufacturer's instruction which includes
processes such as blocking, primary antibody incubating, washing
and secondary antibody incubating followed by addition of
horseradish enzyme labeled chain enzyme working solutions, and
finally blotting, re-staining, dehydrating and mounting. Results
were evaluated by pathological professionals. Five arbitrary visual
fields under magnification, ×400 were selected with 100 cells
counted for each field. IHC reactivity for LMO1, Bcl-2 and Bax was
read by a scoring system based on both the quantity and quality of
brown pigmentation in the plasma of cells. The presence of cells
with yellow or brownish-yellow staining was identified as positive.
If the positive rate was: <10%=0; 0–30%=1; 30–50%=2; >50%=3.
The quality of staining of IHC was also scored if it was: clear=0;
light yellow=1; yellow=2; brownish-yellow=3. By multiplication of
two scores for each, the final score was calculated, of which if
0–1=(−); 2–3=(+); 4–6=(++); >6=(+++).
Cell culture
All cell lines were cultured in RPMI 1640 media
supplemented with 10% FBS, 1% penicillin streptomycin combination
in a 5% CO2 at 37°C. Media were changed every second
day. Cells growing exponentially were selected for experiment.
Experimental samples were divided into three groups according to
the transfection status. The three groups were LMO1-siRNA
transfection (LMO1-siRNA group), NS-siRNA transfection
(control-siRNA group), and Lipofectamine™ 2000 (blank
group). Each treatment repeated 3 triple.
LMO1-siRNA transfection and
grouping
The sequence of LMO1-siRNA was synthesized as below:
5′-GGGCCCGAGACAAUGUGUAUtt-3′. Meanwhile, one fluorescent-labeled
siRNA (NS-siRNA) was used as the negative control, and the sequence
was as follows: 5′-UUCUCCGAACGUGUCACGUtt-3′. The MKN45 cells
undergoing exponential growth phase were transplanted in 6-well
plates with density of 2×105 cells/ml. Transfection was
performed using the Lipofectamine™ 2000. Efficiency of
transfection was tested under fluorescence microscope in 24 h after
transfection. After 48 h, the cells were collected for
experiment.
Cell viability of each group (CCK8
cell proliferation assay)
Cells (1×105 cell/well) were planted in
96-well plates and divided into 3 groups which were transfected
with LMO1-siRNA, NS-siRNA and Lipofectamine™ 2000
reagent, respectively after 24-hour incubation. 48 h after the
interference, 10 µl of CCK8 solution was added to each well of the
plates. Bubbles should not to be introduced to the wells. The
absorbance (optical density, OD) was measured at 450 nm with
microplate reader. The assay was repeated 3 times.
Fluorescence quantitative real-time
PCR (qRT-PCR) for the detection of targeted gene expression
Single-step RNA isolation method with TRIzol was
applied to extract total RNA from cell or tissue samples. The RNA
samples were used for synthesis of the complementary DNA (cDNA)
template for qRT-PCR with 2 µg reverse transcription product. Gene
expression was detected using SYBR-Green PCR mix (Toyoko) and 1 µg
of the template. Reverse transcription PCR (RT-PCR) was performed
with 2 µg reverse transcription product for the quantification of
target mRNA levels, while GAPDH was employed as an internal
standard reference. PCR was performed in a total volume of 20 µl
containing 2 µl reverse transcription product, 10 µl of SYBR-Green
Mix (Applied Biosystems, Foster City, CA) and 0.5 µl of sense an
anti-sense primer each (10 µmol/l). PCR thermal cycling condition
was optimized as following: 5 min initial denaturation at 95°C,
followed by 45 cycles of 30 sec denaturation at 94°C, 30 sec
annealing at 60°C and 30 sec extension at 70°C. The PCR primers
used for experiment were designed using Primer 5.0 software with
interpretation of the BLAST result to improve the specificity of
the designed primers. The template sequences were as follows: LMO1
forward, 5′-TCTACACCAAGGCCAACCTC-3′ and reverse,
5′-CTGCCCTTCCTCATAGTCCA-3′; Bcl-2 forward,
5′-TGTGTGGAGAGCGTCAACC-3′ and reverse, 5′-TGGATCCAGGTGTGCAGGT-3′;
Bax forward, 5′-TTTCTGACGGCAACTTCAAC-3′ and reverse,
5′-AGTCCAATGTCCAGCCCAT-3′; GAPDH forward,
5′-GACCCCTTCATTGACCTCAAC-3′ and reverse,
5′-CGCTCCTGGAAGATGGTGAT-3′. The RT-PCR result was examined adopting
1.5% agarose gel electrophoresis, while the result of real-time
fluorescent quantitative PCR was analyzed using the
2−ΔΔCt method with GAPDH as internal reference of
standard.
Western blot detection of protein
expression of target genes
Tissue samples or cells were placed in ice with 100
µl lysis solution for 30 min, followed by 30 min spinning in a 4°C
centrifuge at 12,000 × g. After that, bicinchoninic acid (BCA)
assay was used for protein quantitation. Total 50 µg protein was
loaded in 10% dodecyl sulfate, sodium salt-polyacrylamide gel
electrophoresis (SDS-PAGE) and transferred to the poly vinylidene
fluoride (PVDF) membrane. After 2 h blocking, LMO1, Bcl-2 and Bax
antibodies were added followed by overnight incubation at 4°C. The
blot was rinsed with TBST before and after the addition of HRP-Goat
anti-mouse IgG with 1 h incubation at room temperature.
Chemiluminescence was applied for color causing and imaging. The
band integrated optical density value (IOD) of loaded proteins was
then read. The ratios of IOD of each target proteins to that of the
internal reference (GAPDH) were represented as the density of
target proteins.
Statistical analysis
All assay results were analyzed using SPSS version
16.0 (SPSS, Inc., Chicago, IL, USA). The individual relations of
each protein in various tissues between their related
clinicopathological characteristics were measured using
χ2 test. The association of the proteins with each other
was measured by the Spearman's rank-order correlation. Kaplan-Meier
(K-M) survival analysis and COX multivariate model were employed
for analysis of the relation between LMO1 expression and patient
prognosis. Comparison of the cell viability, level of protein was
evaluated with analysis of variance (ANOVA) or t-test. P<0.05
was considered to indicate a statistically significant
difference.
Results
Expression level of LMO1, Bcl-2 and
Bax in gastric cancerous and adjacent tissues
In gastric cancer tissues, the positive expression
rates of LMO1, Bcl-2 and Bax protein were 75.00% (72/96), 71.88%
(69/96) and 38.54% (37/96), respectively, compared to the rates of
that in adjacent tissue samples at 28.00% (14/50), 30.00% (15/50),
76.00% (38/50), respectively. The result showed that LMO1 and Bcl-2
proteins had higher positive rates in gastric cancer tissue than
that in adjacent tissue (P<0.05), whereas Bax saw a lower rate
in gastric cancer tissue than it was in adjacent benign tissue
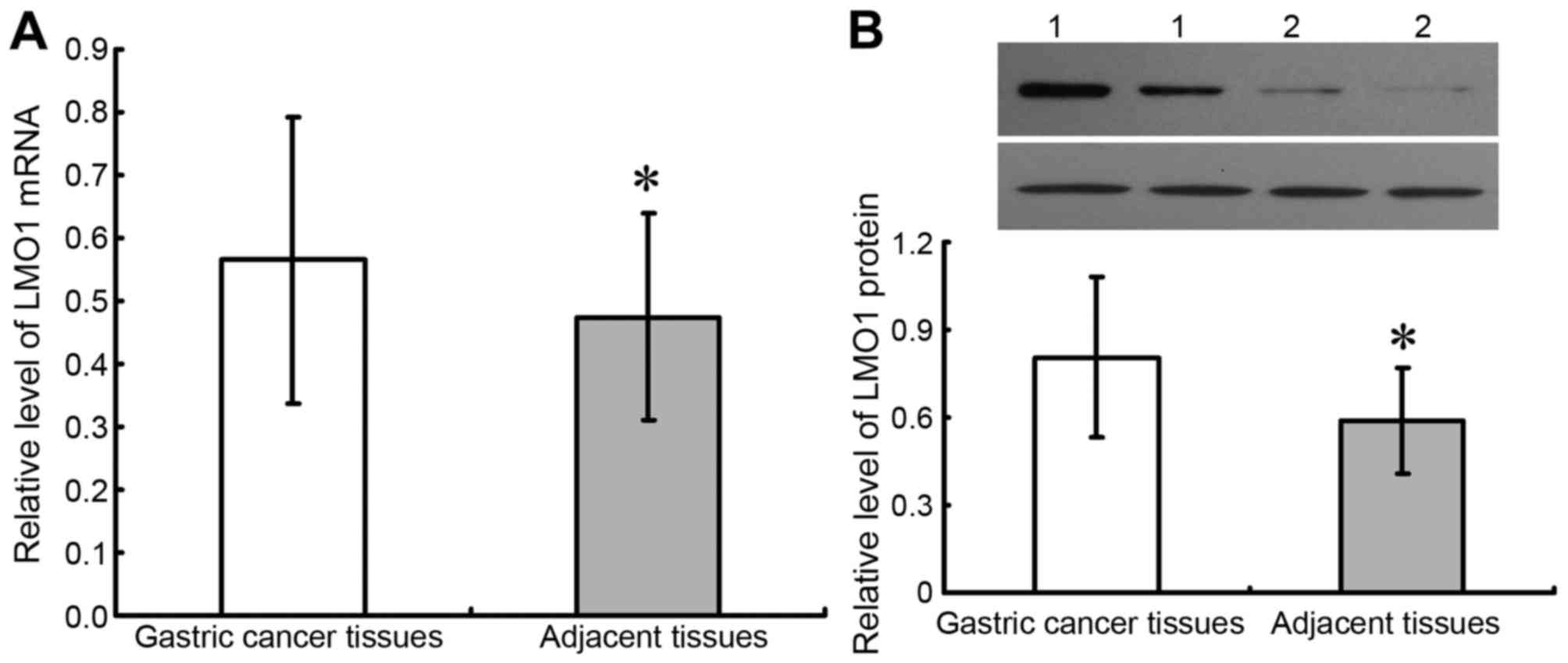
(Table I; Fig. 1). Results of fresh tissues from
qRT-PCR and western blot were demonstrated as Fig. 2, which showed that expressions of LMO1
mRNA and protein were higher in cancer tissues than in adjacent
normal tissues (P<0.05).
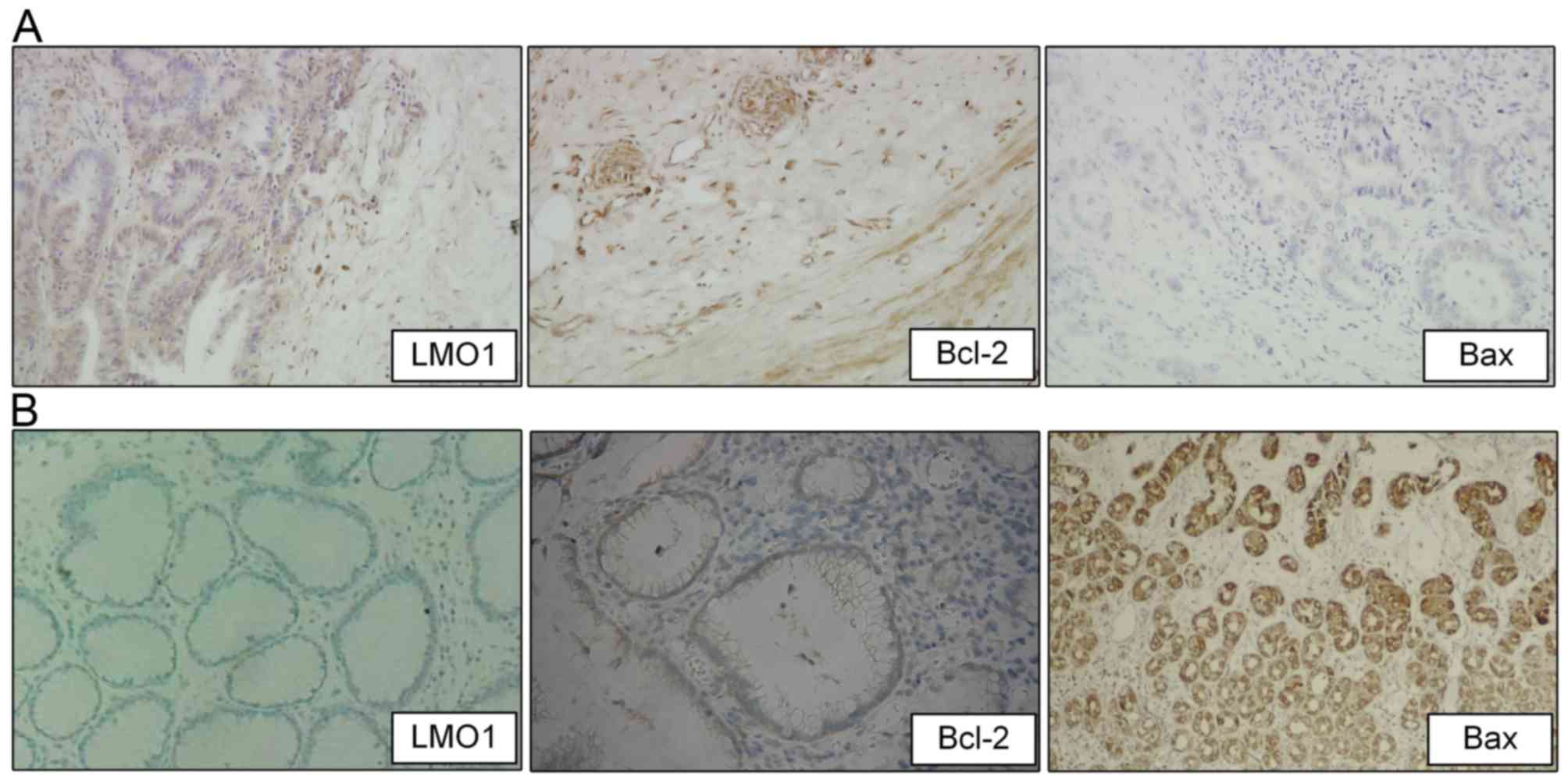 | Figure 1.Expression of LMO1, Bcl-2, and Bax
proteins in gastric cancer and adjacent tissues
(Immunohistochemistry). Immunohistochemistry (magnification ×200)
was utilized to detect expression of Bcl-2, and Bax protein in
gastric cancer and adjacent tissues, and positive staining of LMO1,
Bcl-2, and Bax proteins were located in cytoplasm. Expressions of
LMO1, Bcl-2, and Bax proteins in cancer tissues were illustrated as
(A), and in adjacent tissues were illustrated as (B). LMO1, LIM
domain only 1; Bcl-2, apoptosis regulator Bcl-2; Bax, apoptosis
regulator Bax. |
 | Table I.Expression of LMO1, Bcl-2, Bax
proteins in gastric cancer and tumor-adjacent tissues. |
Table I.
Expression of LMO1, Bcl-2, Bax
proteins in gastric cancer and tumor-adjacent tissues.
|
|
| LMO1 | Bcl-2 | Bax |
|---|
|
|
|
|
|
|
|---|
| Tissue type | n | Negative | Positive | Negative | Positive | Negative | Positive |
|---|
| Gastric cancer | 96 | 24 | 72 | 27 | 69 | 59 | 37 |
| Tumor-adjacent
tissues | 50 | 36 | 14 | 35 | 15 | 12 | 38 |
| χ2 |
| 30.001 | 23.596 | 18.466 |
| P-value |
| <0.001 | <0.001 | <0.001 |
Correlation of LMO1, Bcl-2 and Bax
protein expression in gastric cancer tissues
The Spearman's rank-order correlation was
applied to analyze protein expression of LMO1, Bcl-2 and BAX in
gastric cancer cells. The results indicated a positive correlation
between LMO1 and Bcl-2 protein (r=0.3024; P=0.003), whereas
a negative one was shown between LMO1 and Bax protein
(r=−0.2334; P=0.022). Bax protein expression was also
negatively correlated to that of Bcl2 (r=−0.3445;
P<0.001).
Association between LMO1, Bcl-2 and
Bax protein and the clinicopathological characteristics in gastric
cancer
The χ2 test result showed that LMO1
protein was associated with TNM cancer staging and lymphatic
metastasis. Bcl-2 protein was shown in relation to the depth of
tumor infiltration, while Bax protein was associated with lymphatic
metastasis (P<0.05) (Table
II).
 | Table II.Relationship between LMO1, Bcl-2, Bax
proteins and biological characteristics of gastric cancer
patients. |
Table II.
Relationship between LMO1, Bcl-2, Bax
proteins and biological characteristics of gastric cancer
patients.
|
| LMO1 | Bcl-2 | Bax |
|---|
|
|
|
|
|
|---|
| Biological
characteristics (n) | Negative (24) | Positive (72) | χ2
(P-value) | Negative (27) | Positive (69) | χ2
(P-value) | Negative (59) | Positive (37) | χ2
(P-value) |
|---|
| Sex |
|
|
|
|
|
|
|
|
|
| Male
(66) | 17 | 49 | 0.065 (0.799) | 17 | 49 | 0.586 (0.444) | 39 | 20 | 1.393 (0.238) |
| Female
(30) | 7 | 23 |
| 10 | 20 |
| 20 | 17 |
|
| Age, years |
|
|
|
|
|
|
|
|
|
| ≥60
(55) | 15 | 40 | 0.355 (0.551) | 17 | 38 | 0.494 (0.482) | 36 | 19 | 0.868 (0.351) |
| <60
(41) | 9 | 32 |
| 10 | 31 |
| 23 | 18 |
|
| Diameter, cm |
|
|
|
|
|
|
|
|
|
| ≥5
(58) | 14 | 44 | 0.058 (0.810) | 14 | 44 | 1.152 (0.283) | 39 | 19 | 2.069 (0.150) |
| <5
(38) | 10 | 28 |
| 13 | 25 |
| 20 | 18 |
|
| Serosal
invasion |
|
|
|
|
|
|
|
|
|
|
Negative (31) | 9 | 22 | 0.397 (0.529) | 13 | 18 | 4.320 (0.038) | 20 | 11 | 0.181 (0.671) |
|
Positive (65) | 15 | 50 |
| 14 | 51 |
| 39 | 26 |
|
| TNM staging |
|
|
|
|
|
|
|
|
|
| I–II
(28) | 11 | 17 | 4.303 (0.038) | 10 | 18 | 1.126 (0.289) | 15 | 13 | 1.038 (0.308) |
| III
(68) | 13 | 55 |
| 17 | 51 |
| 44 | 24 |
|
|
Differentiation |
|
|
|
|
|
|
|
|
|
| Well
(67) | 16 | 51 | 0.148 (0.700) | 15 | 52 | 3.611 (0.057) | 40 | 27 | 0.289 (0.591) |
| Poor
(29) | 8 | 21 |
| 12 | 17 |
| 19 | 10 |
|
| Lymphatic
metastasis |
|
|
|
|
|
|
|
|
|
|
Positive (71) | 13 | 58 | 6.508 (0.011) | 18 | 53 | 1.037 (0.309) | 48 | 23 | 4.350 (0.037) |
|
Negative (25) | 11 | 14 |
| 9 | 16 |
| 11 | 14 |
|
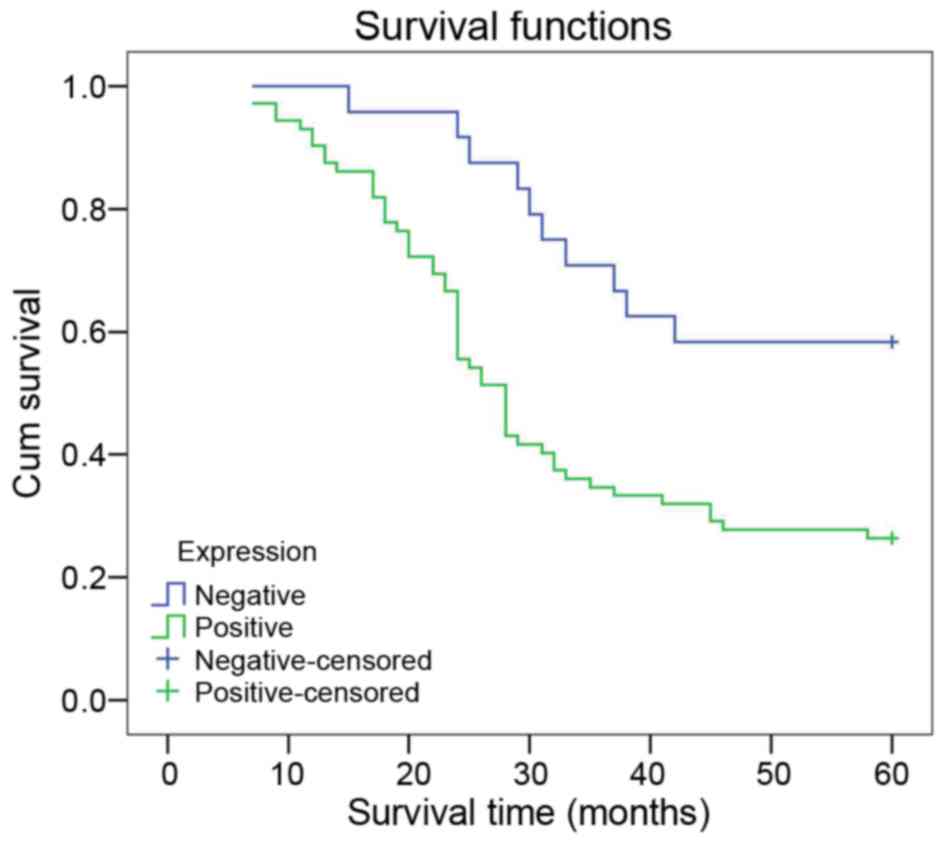
The relation between LMO1 protein and
patient prognosis in gastric cancer
In the K-M survival analysis, patients with positive
LMO1 expression had a lower survival rate than those with negative
expression (χ2=9.025; P=0.003) (Fig. 3). Cox multivariate analysis suggests
that LMO1 expression and TNM staging were independent risk factors
of gastric cancer prognosis (P<0.05) (Table III).
 | Table III.Results of Cox risk model for
survival of gastric cancer patients. |
Table III.
Results of Cox risk model for
survival of gastric cancer patients.
|
|
|
|
|
|
|
| 95.0% CI for
Exp(B) |
|---|
|
|
|
|
|
|
|
|
|
|---|
| Characteristic | B | SE | Wald | df | Sig | Exp(B) | Lower | Upper |
|---|
| LMO1
expression | 1.082 | 0.425 | 6.477 | 1 | 0.011 | 2.952 | 1.283 | 6.794 |
| Lymphatic
metastasis | −0.112 | 0.317 | 0.125 | 1 | 0.723 | 0.894 | 0.480 | 1.664 |
| TNM stages | 0.698 | 0.345 | 4.105 | 1 | 0.043 | 2.010 | 1.023 | 3.949 |
| Invasion | 0.143 | 0.306 | 0.218 | 1 | 0.640 | 1.153 | 0.634 | 2.099 |
| Sex | −0.264 | 0.314 | 0.707 | 1 | 0.400 | 0.768 | 0.415 | 1.420 |
| Age | −0.250 | 0.336 | 0.551 | 1 | 0.458 | 0.779 | 0.403 | 1.506 |
|
Differentiation | 0.274 | 0.277 | 0.981 | 1 | 0.322 | 1.316 | 0.765 | 2.264 |
| Diameter | 0.090 | 0.313 | 0.082 | 1 | 0.775 | 1.094 | 0.592 | 2.019 |
Expression of LMO1 mRNA and protein in
gastric cell lines
Expressions of LMO1 mRNA and protein were detected
in 5 gastric cancer cell lines and gastric epithelial cell line
GES-1, in which higher expressions of LMO1 mRNA and protein were
demonstrated in gastric cancer cell lines than in GES-1 cells, and
highest expressions of LMO1 mRNA and protein were detected in MKN45
cells (P<0.05) (Fig. 4). Then
MKN45 cell line was selected for further studies.
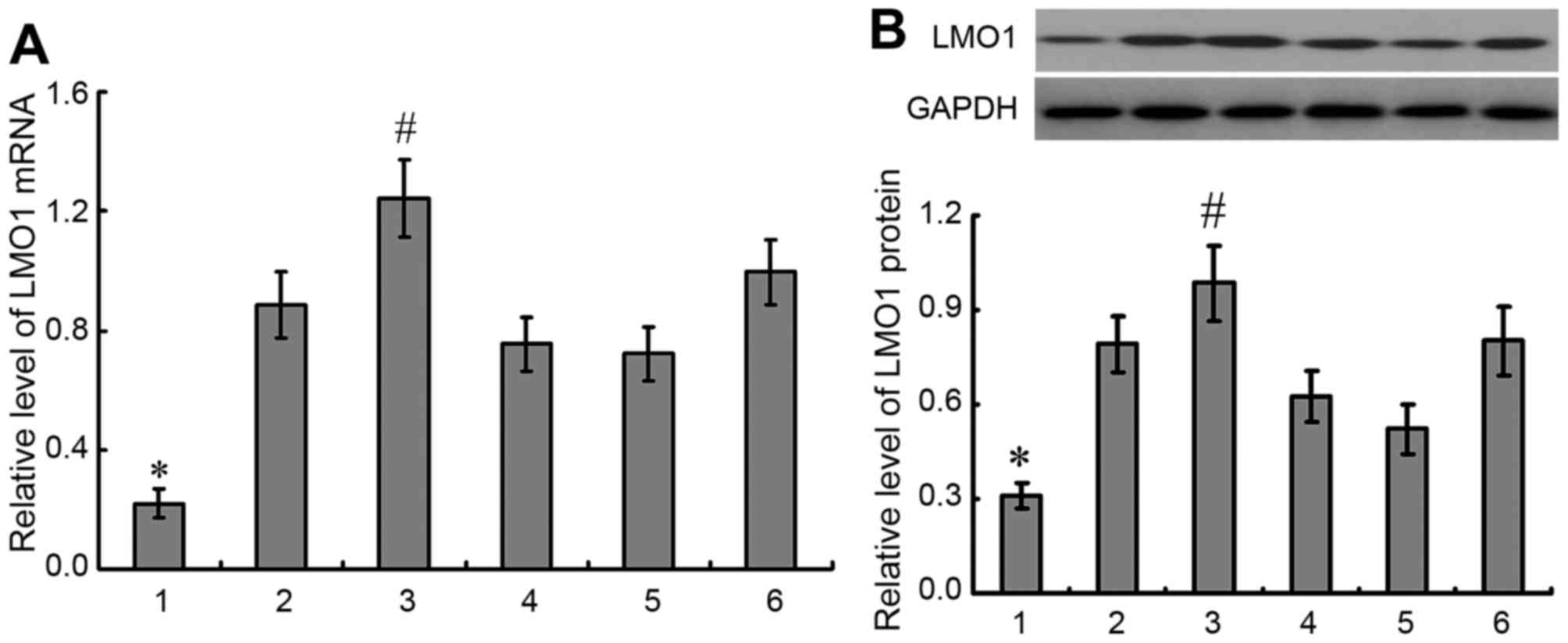 | Figure 4.Expression of LMO1 mRNA and protein in
gastric cell lines. A total of 5 gastric cancer cell lines and
gastric epithelial cell line GES-1 were selected to detected
expression of LMO1 gene with qRT-PCR (A) and western blotting (B).
It was demonstrated that there was higher expression of LMO1 in
gastric cancer cell lines than in GES-1 cells, and highest
expression of LMO1 gene was detected in MKN45 cells. 1, GES-1; 2,
mixed gastric adenocarcinoma cells MKN28 (which are a derivative of
MKN74 cells); 3, MKN45; 4, SGC-7901; 5, BGC823; 6, MGC803.
*P<0.01, compared with gastric cancer cell lines;
#P<0.01, compared with other gastric cell lines.
LMO1, LIM domain only 1. |
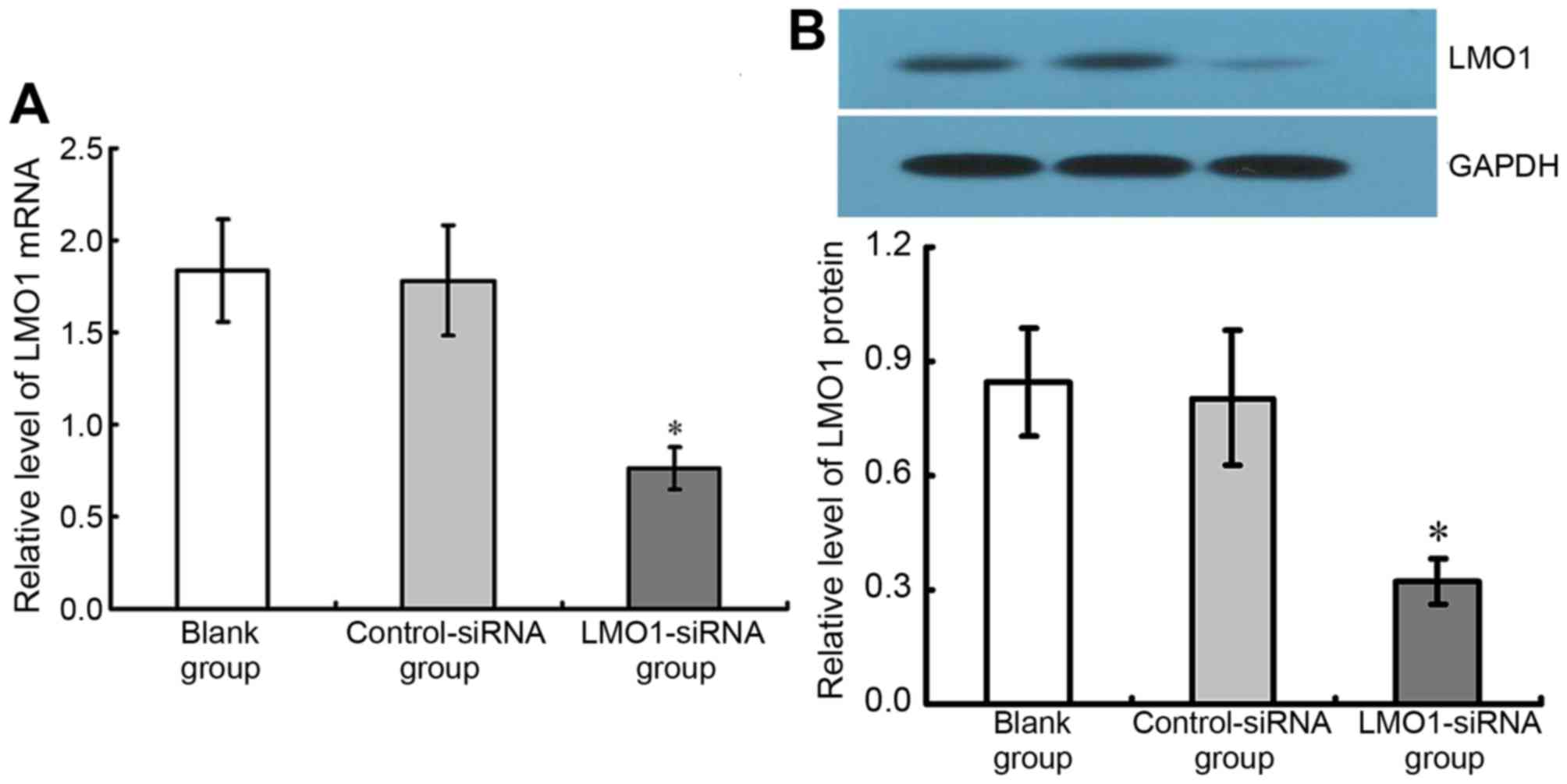
Inhibitory effect of LMO1-siRNA on
LMO1 in MKN45 cells
48 h after transfection of LMO1-siRNA (40 µmol/l),
mRNA and protein expression levels of LMO1 in MKN45 cells, with
high expression of LMO1 protein, were detected using RT-qPCR
(Fig. 4A) and western blotting
(Fig. 4B). The results show that the
group with LMO1-siRNA transfection had significant lower LMO1 mRNA
and protein expression rates than Control-siRNA group and Blank
group (P<0.05). The Control-siRNA group and Blank group
presented no significant difference in their LMO1 expression
(P<0.05) (Fig. 5).
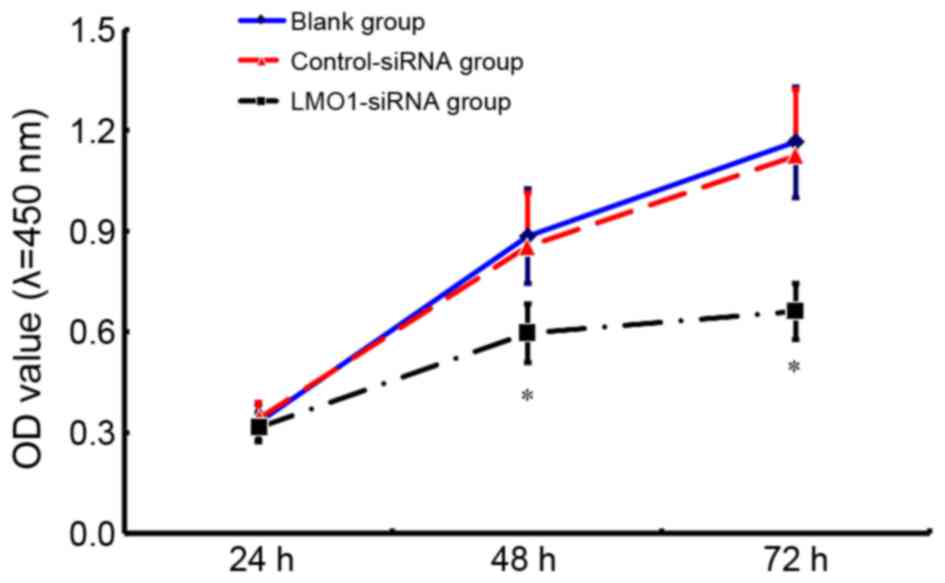
Change in cell viability of MKN45
cells after LMO1-siRNA transfection
The results of CCK8 assay indicate that the cell
viability in the group treated with LMO1-siRNA transfection (40
µmol/l) was significantly lower than that in Control-siRNA group
and Blank group (Fig. 6). 48 h after
interference, LMO1-siRNA group saw a relative OD value at
0.598±0.097, while the value of Control-siRNA group and Blank group
was 0.977±0.162 and 1.162±0.140, respectively.
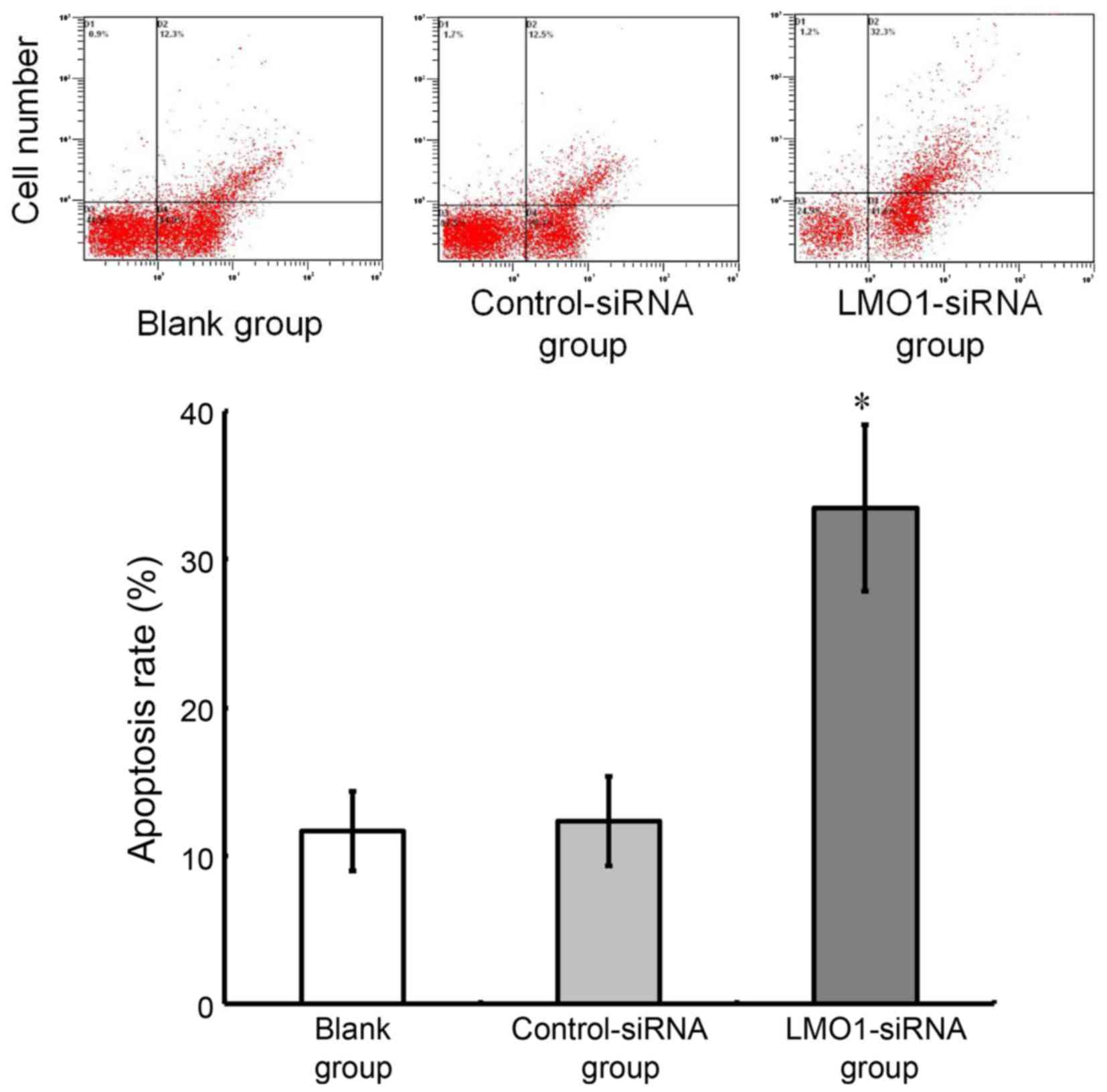
Variation in cell apoptosis of MKN45
cells after LMO1-siRNA transfection
As analyzed using flow cytometry, the cell apoptotic
rate was 31.46±5.22% in LMO1-siRNA group, while the rate in
Control-siRNA group and Blank group was 13.45±1.51% and
12.68±1.32%, respectively. The cell apoptotic rate was higher in
the LMO1-siRNA group than that in Control-siRNA group and Blank
group (P>0.05). The Control-siRNA group and Blank group showed
no significant difference in the cell apoptotic rate (P>0.05)
(Fig. 7).
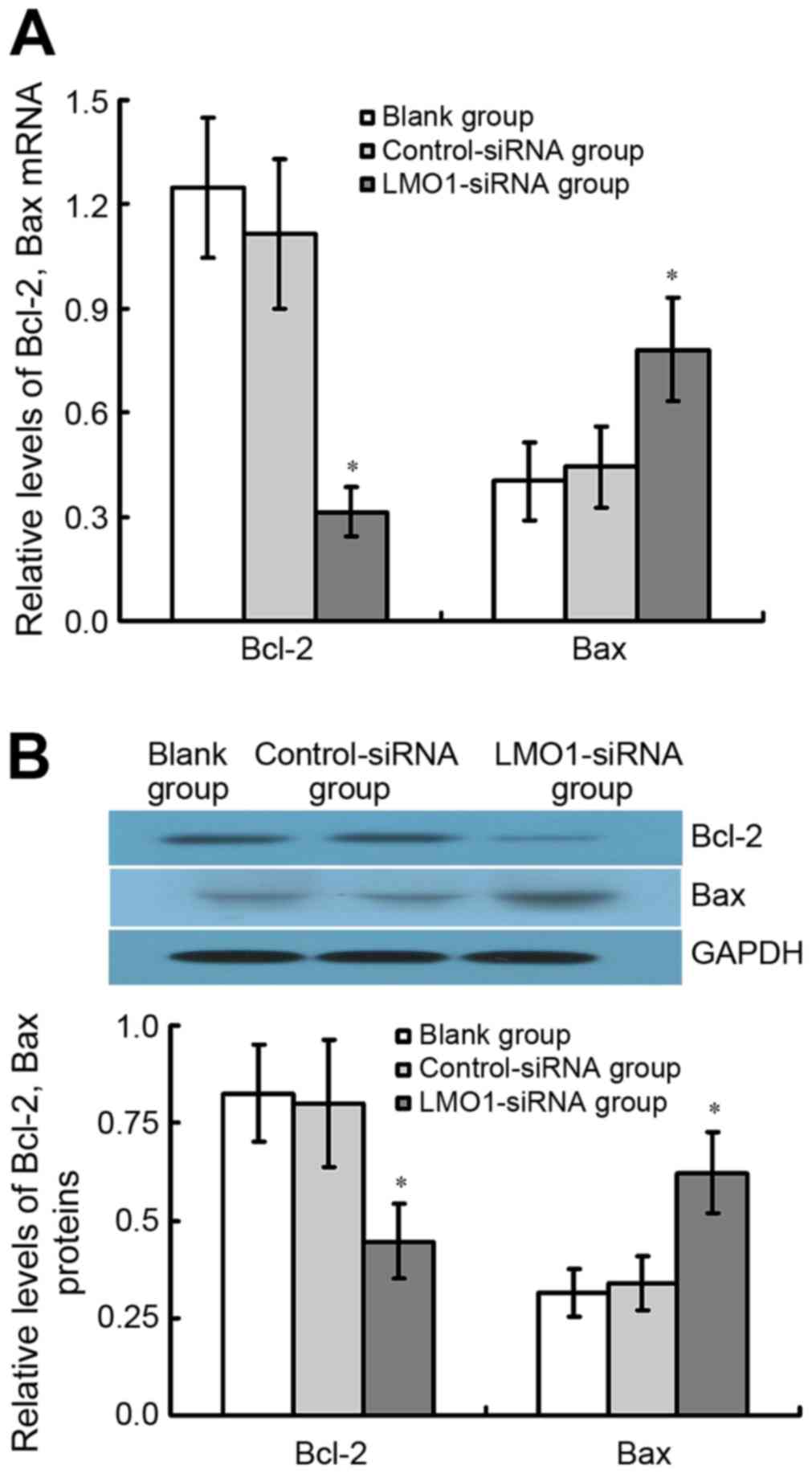
Expression of Bcl-2 and Bax mRNA and
protein in MKN45 cells transfected with LMO1-siRNA
The results of qRT-PCR and western blot assay
demonstrate that Bcl-2 protein expression considerably decreased in
LMO1-siRNA treated group comparted with Control-siRNA group and
Blank group, whereas Bax was found the contrary (P<0.05). There
was no significant difference between Control-siRNA group and Blank
group (P<0.05) (Fig. 8).
Discussion
One major cause of rapid gastric cancer
aggressiveness is the strong anti-apoptotic ability of the tumor
cells which increase survival of gastric cells and therefore
promote tumor progression and metastasis (10,11). Thus,
in order to improve the outcome of treatment, the anti-apoptotic
ability may need to be inhibited and this may promote apoptosis in
gastric cancer cells resulting in decreased tumor progression. As a
variety of genes and channels are involved in the anti-apoptotic
process (12,13), identifying the major functional genes
is crucial. Although to search and identify novel genetic markers
has become a hot topic for gastric carcinoma research, no major
breakthrough has been achieved.
LMO1 is one of the novel genes which has been
reported in recent studies that is associated with a variety of
malignancies such as leukemia, colorectal cancer and lung cancer in
terms of tumor progression, metastasis and apoptosis (8,14,15). However, only a few studies focused on
the relation between LMO1 and gastric cancer. The present study
found higher LMO1 expression in gastric cancer tissue comparing
with that in adjacent mucosa. Similarly, in MKN45 gastric cancer
cell line, the expression level was higher compared with that in
GES-1 gastric epithelial cell line and other gastric cancer cell
lines. These results suggested that LMO1 may have effect on
progression of gastric carcinoma. Furthermore, the results
indicated that LMO1 protein expression is associated with TNM
staging and lymph node metastasis. In addition to that, LMO1
positive patients have a lower survival rate and the positive
expression is an independent risk factor in prognosis. Thus, LMO1
can be suggested as one of the prognostic marker in gastric
carcinoma, although additional experiments of a larger number of
samples are needed.
In order to understand the potential mechanism of
LMO1 in gastric cancer apoptosis, Bcl-2 and Bax protein were
included in the present study. Bcl-2 which plays a vital role in
the mitochondrial pathways of apoptosis can suppress cell apoptosis
(16,17), whereas Bax functions as a
pro-apoptotic gene which can induce cell apoptosis if overexpressed
(18,19). A heterodimer of Bcl-2 and Bax can
determine the cell apoptotic status by altering its binding ratio
(20). Some studies report that
inhibiting Bcl-2 expression but promoting Bax expression may induce
tumor cell apoptosis in gastric carcinoma (21,22). The
present study which compares the expression level of Bcl-2 and Bax
in gastric cancer tissues to those in adjacent normal tissues,
finds that Bcl-2 had higher expression level in the former tissues
while Bax was seen an opposite result. Because of the association
between Bcl-2 protein expression and gastric cancer filtration in
addition to the relation of Bax protein expression with gastric
cancer lymph node metastasis, Bcl-2 and Bax are thus suggested in
relation to the progression of gastric cancer. In this study, LMO1
was found in positive correlation to Bcl-2, but negative
correlation to Bax. LMO1 probably promote gastric cancer invasion
and metastasis through regulation of Bcl-2 and Bax protein. With
regard to better understand the mechanism of LMO1 in gastric cancer
cell death, the present study adopted RNA interference technique
(23) to suppress endogenous
expression of LMO1 in MKN45 gastric cancer cell line. The results
showed that the effective inhibition of LMO1 in gastric cancer
cells leads to a decrease of tumor cell activity and an increase of
cell apoptosis. Furthermore, the expression of Bcl-2 in MKN45
decreases while the expression of Bax increases significantly. All
of these results indicated that LMO1 in gastric cancer may promote
Bcl-2 expression and inhibit Bax expression although more
experiments in vivo at molecular level are needed.
In conclusion, LMO1 was found increased in gastric
cancer tissues and cell lines, probably affecting gastric cancer
apoptosis through regulation of Bcl-2 and Bax. However, due to a
limited number sample, only a partial experiment in vitro
was accomplished. We consider enlarging the scale of experiment and
completing the follow-up data in the urther study. Meanwhile, more
work should be employed for further investigation of LMO1 in
relation to other genetic molecule regulation. Animal experiments
are also favored to ensure the accuracy of the study results.
References
|
1
|
Chen W, Zheng R, Baade PD, Zhang S, Zeng
H, Bray F, Jemal A, Yu XQ and He J: Cancer statistics in China,
2015. CA Cancer J Clin. 66:115–132. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Somi MH, Ghojazadeh M, Bagheri M and
Tahamtani T: Clinicopathological factors and gastric cancer
prognosis in the iranian population: A meta-analysis. Asian Pac J
Cancer Prev. 16:853–857. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Son T, Hyung WJ, Kim JW, Kim HI, An JY,
Cheong JH, Choi SH and Noh SH: Anatomic extent of metastatic lymph
nodes: Still important for gastric cancer prognosis. Ann Surg
Oncol. 21:899–907. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Marin JJ, Al-Abdulla R, Lozano E, Briz O,
Bujanda L, Banales JM and Macias RI: Mechanisms of resistance to
chemotherapy in gastric cancer. Anticancer Agents Med Chem.
16:318–334. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Kang HC, Kim IJ, Park JH, Shin Y, Ku JL,
Jung MS, Yoo BC, Kim HK and Park JG: Identification of genes with
differential expression in acquired drug-resistant gastric cancer
cells using high-density oligonucleotide microarrays. Clin Cancer
Res. 10:272–284. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Kumar R, Kaur M and Silakari O:
Physiological modulation approaches to improve cancer chemotherapya
review. Anticancer Agents Med Chem. 14:713–749. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Frenzel LP, Patz M, Pallasch CP, Brinker
R, Claasen J, Schulz A, Hallek M, Kashkar H and Wendtner CM: Novel
X-linked inhibitor of apoptosis inhibiting compound as sensitizer
for TRAIL-mediated apoptosis in chronic lymphocytic leukaemia with
poor prognosis. Br J Haematol. 152:191–200. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Beuten J, Gelfond JA, Piwkham D, Pollock
BH, Winick NJ, Collier AB III and Tomlinson GE: Candidate gene
association analysis of acute lymphoblastic leukemia identifies new
susceptibility locus at 11p15 (LMO1). Carcinogenesis. 32:1349–1353.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Capes-Davis A, Theodosopoulos G, Atkin I,
Drexler HG, Kohara A, MacLeod RA, Masters JR, Nakamura Y, Reid YA,
Reddel RR and Freshney RI: Check your cultures! A list of
cross-contaminated or misidentified cell lines. Int J Cancer.
127:1–8. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Sugita H, Iida S, Inokuchi M, Kato K,
Ishiguro M, Ishikawa T, Takagi Y, Enjoji M, Yamada H, Uetake H, et
al: Methylation of BNIP3 and DAPK indicates lower response to
chemotherapy and poor prognosis in gastric cancer. Oncol Rep.
25:513–518. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Hu X, Zhang F, Luo D, Li N, Wang Q, Xu Z,
Bian H, Liang Y, Lu Y, Zheng Q and Gu J: URI promotes gastric
cancer cell motility, survival, and resistance to adriamycin in
vitro. Am J Cancer Res. 6:1420–1430. 2016.PubMed/NCBI
|
|
12
|
Lee JH, Soung YH, Lee JW, Park WS, Kim SY,
Cho YG, Kim CJ, Seo SH, Kim HS, Nam SW, et al: Inactivating
mutation of the pro-apoptotic gene BID in gastric cancer. J Pathol.
202:439–445. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Mashima T, Oh-hara T, Sato S, Mochizuki M,
Sugimoto Y, Yamazaki K, Hamada J, Tada M, Moriuchi T, Ishikawa Y,
et al: p53-defective tumors with a functional apoptosome-mediated
pathway: A new therapeutic target. J Natl Cancer Inst. 97:765–777.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Liu J, Yan P, Jing N and Yang J: LMO1 is a
novel oncogene in colorectal cancer and its overexpression is a new
predictive marker for anti-EGFR therapy. Tumour Biol. 35:8161–8167.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Zhang Y, Yang J, Wang J, Guo H and Jing N:
LMO1 is a novel oncogene in lung cancer, and its overexpression is
a new predictive marker for anti-EGFR therapy. Med Oncol.
31:992014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Cheng Y, Zhao G, Zhang S, Nigim F, Zhou G,
Yu Z, Song Y, Chen Y and Li Y: AS1411-induced growth inhibition of
glioma cells by up-regulation of p53 and downregulation of Bcl-2
and Akt1 via nucleolin. PLoS One. 11:e01670942016. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Reddy TL, Garikapati KR, Reddy SG, Reddy
BV, Yadav JS, Bhadra U and Bhadra MP: Simultaneous delivery of
Paclitaxel and Bcl-2 siRNA via pH-Sensitive liposomal nanocarrier
for the synergistic treatment of melanoma. Sci Rep. 6:352232016.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Gil J, Ramsey D, Szmida E, Leszczynski P,
Pawlowski P, Bebenek M and Sasiadek MM: The BAX gene as a candidate
for negative autophagy-related genes regulator on mRNA levels in
colorectal cancer. Med Oncol. 34:162017. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Xu L, Wang Z, He SY, Zhang SF, Luo HJ,
Zhou K, Li XF, Qiu SP and Cao KY: Bax-interacting factor-1 inhibits
cell proliferation and promotes apoptosis in prostate cancer cells.
Oncol Rep. 36:3513–3521. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Adefolaju GA, Theron KE and Hosie MJ:
BAX/BCL-2 mRNA and protein expression in human breast MCF-7 cells
exposed to drug vehicles-methanol and dimethyl sulfoxide (DMSO) for
24 hrs. Niger Med J. 56:169–174. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Kim EM, Kim J, Park JK, Hwang SG, Kim WJ,
Lee WJ, Kang SW and Um HD: Bcl-w promotes cell invasion by blocking
the invasion-suppressing action of Bax. Cell Signal. 24:1163–1172.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zhu P, Zhang J, Zhu J, Shi J, Zhu Q and
Gao Y: MiR-429 induces gastric carcinoma cell apoptosis through
Bcl-2. Cell Physiol Biochem. 37:1572–1580. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Baker M: RNA interference: From tools to
therapies. Nature. 464:12252010. View
Article : Google Scholar
|