Introduction
Colorectal cancer (CRC) is the third most common
type of human malignancy worldwide and the fourth leading cause of
cancer-associated mortality (1–3). Despite
the significant achievements that have been made in the treatment
of early CRC, the long-term survival rate for advanced CRC remains
low. The survival rate of CRC may benefit from 5-fluorouracil
(5-FU) and oxaliplatin-based adjuvant chemotherapy, which has been
accepted as a standard therapy (4,5). However,
20–40% of patients with advanced-stage CRC relapse following
primary curative surgery (6,7). This is primarily attributed to the
following reasons: Lack of diagnostic markers for early detection,
weak prognostic value of histological indicators, limited
efficiency of current treatment for advanced diseases and lack of
genetic markers utilized for targeted therapy (2). Therefore, the identification of novel
genetic markers for the improvement of diagnostic and prognostic
techniques is required.
With the development of whole genome sequencing
technology, it has been revealed that <2% of the mammalian
genome is in protein-encoded regions and the remainder of the
genome contains non-coding RNAs (ncRNAs) (8). Those ncRNAs >200 nucleotides in
length are termed long non-coding RNAs (lncRNAs) (9). In recent years, lncRNAs have been
regarded as a diagnostic biomarkers and prognostic factors
(10).
Previous studies have identified a series of lncRNAs
with aberrant expression in cancer (11,12).
Upregulated lncRNA HOTAIR relative expression in primary tumors and
in the blood of patients with CRC is associated with unfavorable
prognosis (13). lncRNA
metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) has
been shown to be upregulated in clinical CRC tissue samples
(14). High-level MALAT1 expression
is an independent prognostic risk factor for patients with CRC
(14). One recent study has revealed
that lncRNA colon cancer-associated transcript 2 (CCAT2)
encompasses the rs6983267 single nucleotide polymorphism (15). Furthermore, lncRNA CCAT2 is highly
overexpressed in CRC cell lines and promotes tumor growth,
metastasis and chromosomal instability (15). In addition, it was revealed that CCAT2
promotes MYC expression (15).
To the best of our knowledge, the diagnostic and
prognostic role of lncRNA CCAT2 expression in CRC remains unclear.
In the present study, the levels of CCAT2 expression in clinical
CRC tissues were tested. Subsequently, the association of CCAT2
with disease-free survival and overall survival of patients with
CRC was analyzed.
Materials and methods
Cohort and tissue samples
According to The Code of Ethics of the World Medical
Association, Declaration of Helsinki, the present study was
approved by the Ethics Committee of Peking University (Beijing,
China). All patients involved provided full consent for the study.
All the fresh specimens were obtained from resected colorectal
tissues of 218 patients, who were selected from patients diagnosed
with CRC between January 2005 and December 2007 at the Department
of General Surgery, Peking University First Hospital. A total of
218 patients (113 were male and 105 were female) were included in
the study. The mean age of patients was 64 years (range 23–89
years). A total of 36 noncancerous healthy colon mucosa tissues
served as controls. All specimens were snap-frozen in liquid
nitrogen and kept at −80°C immediately following surgical
resection. All patients with CRC stage II–IV received adjuvant
chemotherapy based on 5-FU. Patients with the following criteria
were subsequently excluded: Received treatment prior to surgery,
including neoadjuvant chemotherapy or neoadjuvant radiotherapy; had
a diagnosis of additional malignant disease; and harvested
insufficient specimens for RNA isolation. Clinicopathological
information and follow-up data of all patients were entered into a
database that was updated with respect to survival status every 3
months. All the specimens were histologically diagnosed by the
Department of Pathology according to the criteria of the World
Health Organization classification and the tumor-node-metastasis
(TNM) stage set out by American Joint Committee on Cancer (16).
Total RNA extraction and reverse
transcription-quantitative polymerase chain reaction (RT-qPCR)
Total RNA from cancer tissues or adjacent normal
tissues or non-cancerous tissues was isolated using
TRIzol® according to the manufacturer's procedure
(Invitrogen; Thermo Fisher Scientific, Inc., Waltham, MA, USA) and
stored at −80°C. RNA concentration and purity was measured using a
Nanodrop spectrophotometer (Thermo Fisher Scientific, Inc.). The
OD260/280 ratios for all samples were between 1.8 and 2.0. RNA
integrity was determined using Agilent 2100 Bioanalyzer, with RNA
6000 Nano Assay Chip (Agilent Technologies Inc., Santa Clara, CA,
USA). The RNA integrity of all samples ranged between 6.0 and 10.0.
cDNA was synthesized from 500 ng total RNA using the Promega A3500
Reverse Transcription System kit (Promega Corporation, Madison, WI,
USA) with random hexamer primers in a final volume of 40 µl,
according to the manufacturer's protocol. The cDNA was stored at
−20°C. RT-qPCR was performed using the SYBR Premix Ex TagTM II
(Takara Biotechnology Co., Ltd., Dalian, China) and ABI 7300 System
(Applied Biosystems; Thermo Fisher Scientific Inc.). The primers
were as follows: CCAT2 forward, 5′-CCCTGGTCAAATTGCTTAACCT-3′ and
reverse, 5′-TTATTCGTCCCTCTGTTTTATGGAT-3′; and GAPDH forward,
5′-CCACATCGCTCAGACACCAT-3′ and reverse, 5′-ACCAGGCGCCCAATACG-3′.
The PCRs were performed in a total volume of 30 µl, containing 30
ng of cDNA for each sample. The cycling program was set for initial
hold at 95°C for 10 min, followed by 40 cycles of denaturation at
95°C for 15 sec, annealing and extension at 50°C for 30 sec and
72°C for 15 sec. The relative expression of CCAT2 was analyzed
using the 2−ΔΔCq method (17). All fluorescent data were converted
into relative quantitation (RQ; relative expression obtained by
2−ΔΔCq method) measurements. CCAT2 levels were
normalized to GAPDH. Sequencing of randomly selected qPCR products
was utilized to ensure the quality of qPCR; sequencing services
were provided by BBI Life Sciences Co. (Shanghai, China).
Measurement of disease-free survival
and overall survival
Disease-free survival time was defined as the time
elapsed between surgery and the first occurrence of any of the
following events: Relapse of CRC, distant metastasis of CRC or
mortality from any cause without documentation of a
cancer-associated event. The diagnosis of relapse and distant
metastasis was based on imaging methods, including computed
tomography, magnetic resonance imaging or position emission
tomography, and if possible, biopsy or cytological analysis.
Overall survival time was defined as the time elapsed between
surgery and mortality of patients with CRC.
Statistical analysis
Statistical analysis was performed using SPSS 13.0
(SPSS, Inc., Chicago, IL, USA). Data are presented as the mean ±
standard deviation. The levels of CCAT2 expression from different
tissues were analyzed by one-way analysis of variance followed by
Bonferroni multiple comparisons. Associations between CCAT2
expression and categorical variables were analyzed by Pearson's
χ2 test or Fisher's exact test, as appropriate. Survival
curves were estimated using the Kaplan-Meier method, and
differences in survival distributions were evaluated by the
log-rank test. Cox's proportional hazards modeling of factors
potentially associated with survival was performed in order to
identify factors that may have a significant independent effect on
survival. P<0.05 was considered to indicate a statistically
significant difference.
Results
Characteristics of patients
The characteristics of patients are shown in
Table I. In total, there were 113
male (51.83%) and 105 female (48.17%) patients. There were 136
tumors (62.39%) located in the colon and 82 tumors (37.61%) located
in the rectum. The tumor size of 70 patients (32.11%) with CRC was
<3 cm and that of 148 patients (67.89%) was ≤3 cm. Poorly- and
moderately-differentiated tumors were the most common histological
type (71.10%), followed by well-differentiated (28.90%) tumors.
According to the International TNM Classification, 32 (14.68%), 56
(25.69%), 89 (40.83%) and 41 (18.81%) of 218 patients with CRC were
classified as TNM stages I, II, III and IV, respectively (Table I).
 | Table I.Statistical results of CCAT2
expression. |
Table I.
Statistical results of CCAT2
expression.
|
|
| CCAT2 expression, n
(%) |
|
|---|
|
|
|
|
|
|---|
| Parameters | No. (%) | Low | High | P-value |
|---|
| Total | 218 (100.00) | 89 (100.00) | 129 (100.00) |
|
| Sex |
|
|
| 0.7839 |
| Male | 113 (51.83) | 45 (39.82) | 68 (60.18) |
|
|
Female | 105 (48.17) | 44 (41.9) | 61 (58.1) |
|
| Age, years |
|
|
| 0.1592 |
|
<60 | 87 (39.91) | 41 (47.13) | 46 (52.87) |
|
| ≥60 | 131 (60.09) | 48 (36.64) | 83 (63.36) |
|
| Location |
|
|
| 0.7762 |
|
Colon | 136 (62.39) | 57 (41.94) | 79 (58.06) |
|
|
Rectum | 82 (37.61) | 32 (39.02) | 50 (60.98) |
|
| Smoking status |
|
|
| 0.8873 |
| Never
smoked | 135 (61.92) | 56 (41.48) | 79 (58.51) |
|
|
Smoker | 83 (38.08) | 33 (39.76) | 50 (60.24) |
|
| BMI,
kg/m2 |
|
|
| 0.5150 |
|
<25 | 112 (51.38) | 44 (39.29) | 58 (60.71) |
|
|
≥25 | 106 (48.62) | 45 (42.45) | 71 (57.54) |
|
| Diameter, cm |
|
|
| 0.1186 |
|
<3 | 70 (32.11) | 36 (51.43) | 39 (48.57) |
|
| ≥3 | 148 (67.89) | 53 (35.81) | 90 (64.19) |
|
|
Differentiation |
|
|
| 0.0006 |
|
High | 63 (28.90) | 37 (58.73) | 26 (41.27) |
|
|
Poor/moderate | 155 (71.10) | 53 (34.19) | 103 (65.81) |
|
| TNM stage |
|
|
| <0.0001 |
|
I/II | 88 (40.37) | 65 (73.86) | 23 (26.14) |
|
|
III/IV | 130 (59.63) | 24 (18.46) | 106 (81.54) |
|
| Tumor
infiltration |
|
|
| <0.0001 |
|
T1/T2 | 36 (16.51) | 28 (77.78) | 8 (22.22) |
|
|
T3/T4 | 182 (83.49) | 61 (33.52) | 121 (66.48) |
|
| Lymph node
metastasis |
|
|
| <0.0001 |
| N0 | 76 (34.86) | 59 (77.63) | 17 (32.37) |
|
|
N1-N3 | 142 (66.14) | 30 (21.23) | 112 (78.77) |
|
| Distant
metastasis |
|
|
| <0.0001 |
| M0 | 167 (76.61) | 80 (47.90) | 87 (52.10) |
|
| M1 | 51 (23.39) | 9 (17.65) | 42 (82.35) |
|
| Vascular
invasion |
|
|
| <0.0001 |
|
Absent | 112 (51.38) | 61 (54.46) | 51 (45.54) |
|
|
Present | 108 (49.54) | 28 (25.93) | 78 (74.07) |
|
lncRNA CCAT2 is significantly
upregulated in CRC tissues
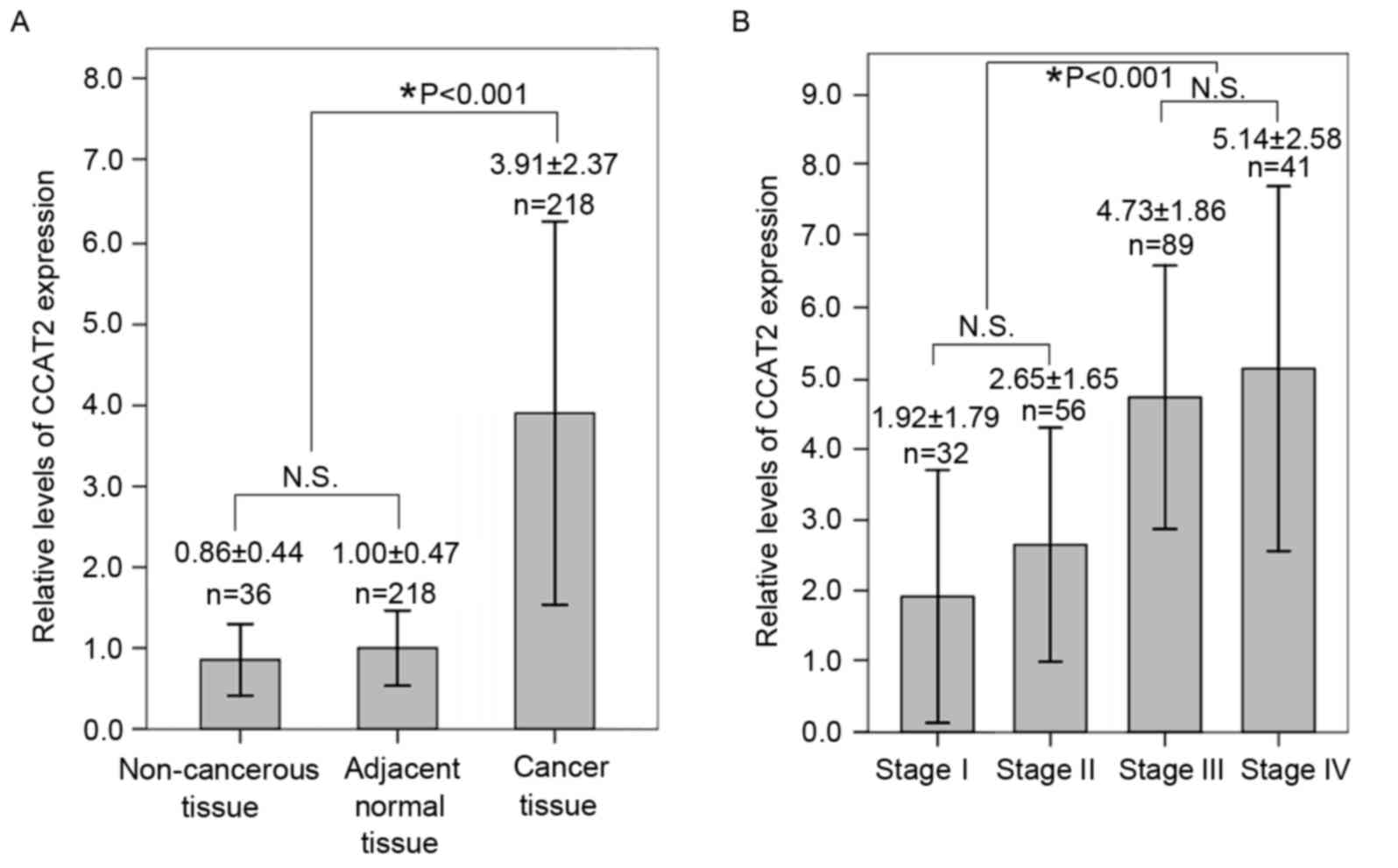
The levels of lncRNA CCAT2 expression were detected
in all human tissues by qPCR. Following normalization to GAPDH
expression, the levels of lncRNA CCAT2 expression in CRC tissues
were significantly increased compared with those of adjacent
non-tumor tissues or noncancerous mucosa tissues (P<0.001;
Fig. 1A). The RQ of lncRNA CCAT2 in
CRC samples was 3.91±2.37, whereas the relative CCAT2 expression
detected in matched adjacent normal tissues was 1.00±0.47, and that
of noncancerous colon mucosa tissues was 0.86±0.44. The CCAT2
expression levels of cancer tissues from patients with stage III
and IV CRC were significantly higher than those from patients with
stage I and II CRC (P<0.001; Fig.
1B).
Association between lncRNA CCAT2
expression and clinicopathological characteristics of CRCs
On the basis of these aforementioned data, the mean
expression level of lncRNA CCAT2 in CRC tissues (3.91) was used as
a cutoff point to classify all 218 cancerous tissues into two
groups: Low expression (n=98) and high expression (n=120). The
associations between CCAT2 expression levels and different
clinicopathological factors are shown in Table I. No statistically significant
associations were observed between CCAT2 expression and sex, age at
diagnosis, tumor location, smoking status, body mass index (BMI)
status and tumor size. Statistically significant associations were
observed between high CCAT2 expression and low degree of tumor cell
differentiation (P=0.0006), advanced TNM staging and deeper tumor
infiltration, high incidence of lymph node metastasis, distant
metastasis and vascular invasion (all P<0.0001).
Association between lncRNA CCAT2
expression and disease-free survival of patients with CRC
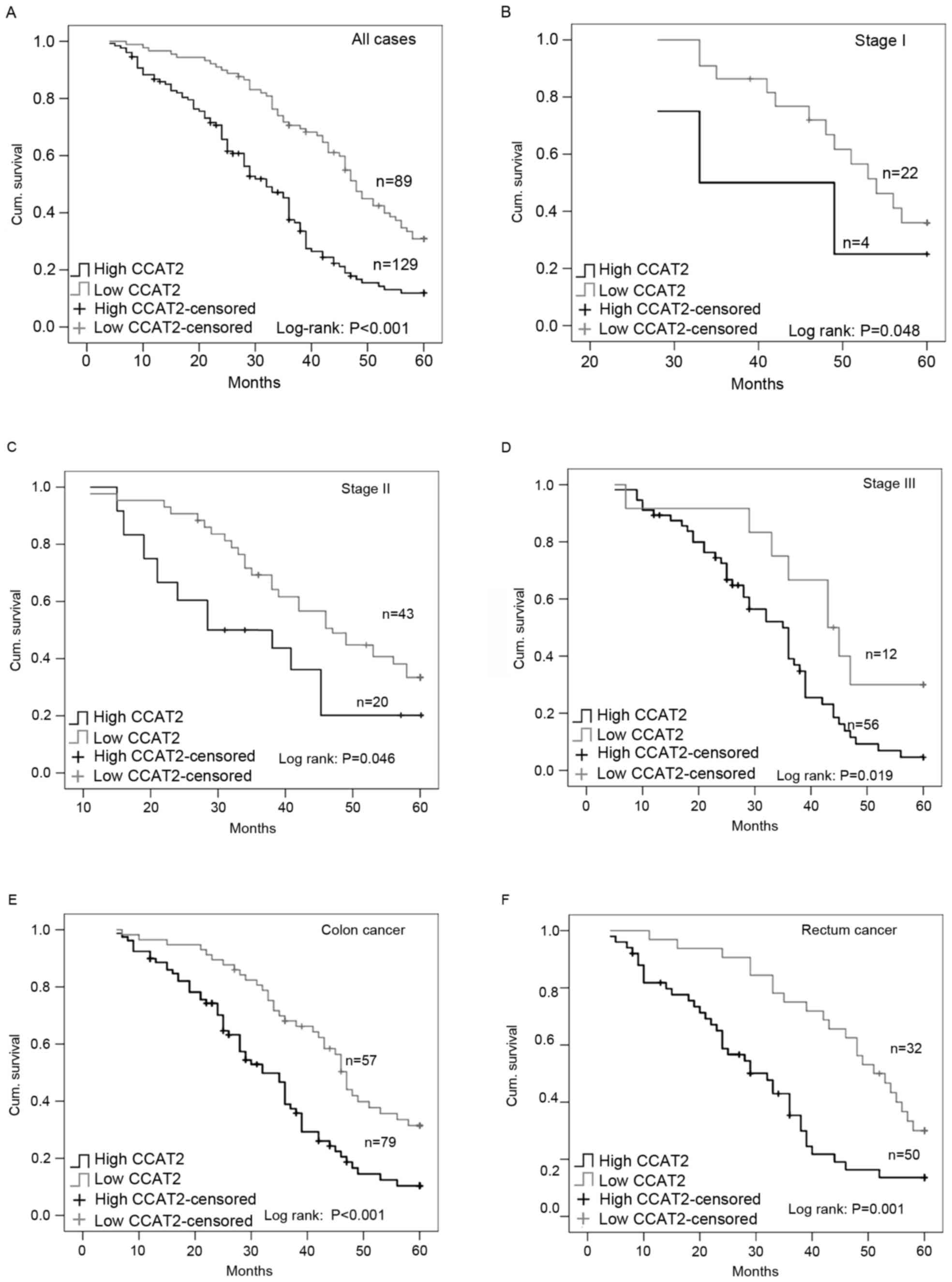
Kaplan-Meier analysis revealed that patients with
low lncRNA CCAT2 expression had increased disease-free survival
times compared with those with high lncRNA CCAT2 expression
(P<0.001; Fig. 2A). In addition,
patients with poor differentiation, tumor diameter >3 cm,
vascular invasion, deeper tumor infiltration, lymph node
metastasis, distant metastasis or advanced TNM stage had shorter
disease-free survival times and an increased risk of relapse
compared with those without. However, other clinical parameters had
no prognostic value on disease-free survival of patients with CRC.
Unadjusted risk ratios (RR) are shown in Table II.
 | Table II.Disease-free risk ratio. |
Table II.
Disease-free risk ratio.
|
| Univariate | Multivariate |
|---|
|
|
|
|
|---|
| Parameters | Unadjusted RR (95%
CI) | P-value | Adjusted RR (95%
CI) | P-value |
|---|
| CCAT2 | 2.340
(1.677–3.264) | <0.001 | 1.561
(1.045–2.331) | 0.030 |
| Sex | 1.162
(0.849–1.592) | 0.348 | 1.111
(0.798–1.548) | 0.533 |
| Age, years | 1.159
(0.840–1.598) | 0.369 | 0.994
(0.696–1.419) | 0.972 |
| Location | 0.901
(0.655–1.238) | 0.518 | 1.013
(0.721–1.424) | 0.939 |
| Smoking status | 0.913
(0.689–1.427) | 0.891 | 0.987
(0.616–1.299) | 0.808 |
| BMI status,
kg/m2 | 1.038
(0.758–1.422) | 0.815 | 1.003
(0.713–1.412) | 0.984 |
| Tumor diameter,
cm | 1.462
(1.036–2.064) | 0.031 | 1.380
(0.918–2.073) | 0.121 |
|
Differentiation | 1.445
(1.022–2.042) | 0.037 | 1.119
(0.725–1.727) | 0.613 |
| TNM stage | 1.482
(1.065–2.064) | 0.020 | 0.616
(0.375–1.011) | 0.045 |
| Tumor
infiltration | 1.707
(1.111–2.625) | 0.015 | 0.954
(0.524–1.739) | 0.878 |
| Lymph node
metastasis | 2.351
(1.658–3.332) | <0.001 | 1.767
(1.128–2.769) | 0.013 |
| Distant
metastasis | 2.578
(1.792–3.710) | <0.001 | 2.376
(1.557–3.625) | <0.001 |
| Vascular
invasion | 1.766
(1.286–2.426) | <0.001 | 1.573
(1.125–2.200) | 0.008 |
Kaplan-Meier analysis was also performed with
stratification by TNM stage. Since TNM stage IV tumors were defined
as tumors with distant metastasis, and itself was a marker of high
risk, patients were classified into groups of TNM I, II and III
tumors, respectively. Results demonstrated that the levels of
lncRNA CCAT2 expression were associated with disease-free survival
in three groups of patients with TNM I, II and III CRC.
Disease-free survival was significantly shorter in patients with
high CCAT2 expression vs. low CCAT2 expression (Fig. 2B-D). In addition, both colon cancer
and rectal cancer patients with low CCAT2 expression had more
favorable disease-free survival compared with those with high CCAT2
expression (Fig. 2E and F).
To verify the independent prognostic value of CCAT2
expression, the Cox proportional hazards model adjusted for the
presence of other clinical parameters was utilized to control for
other prognostic factors. As a result, the level of CCAT2
expression was shown to be an independent prognostic factor,
subsequent to controlling for all other clinical parameters.
Adjusted RR was 1.00 (as a reference) in low CCAT2 expression
patients, while the adjusted RR of patients with CRC with a high
level of CCAT2 expression was 1.561 (P=0.030; Table II). Thus, CCAT2 maybe an independent
prognostic factor of disease-free survival for patients with CRC.
In addition, vascular invasion, lymph node metastasis, distant
metastasis and TNM stages were also revealed to be independent
prognostic factors for disease-free survival of patients with CRC
(Table II).
Association between lncRNA CCAT2
expression and overall survival of patients with CRC
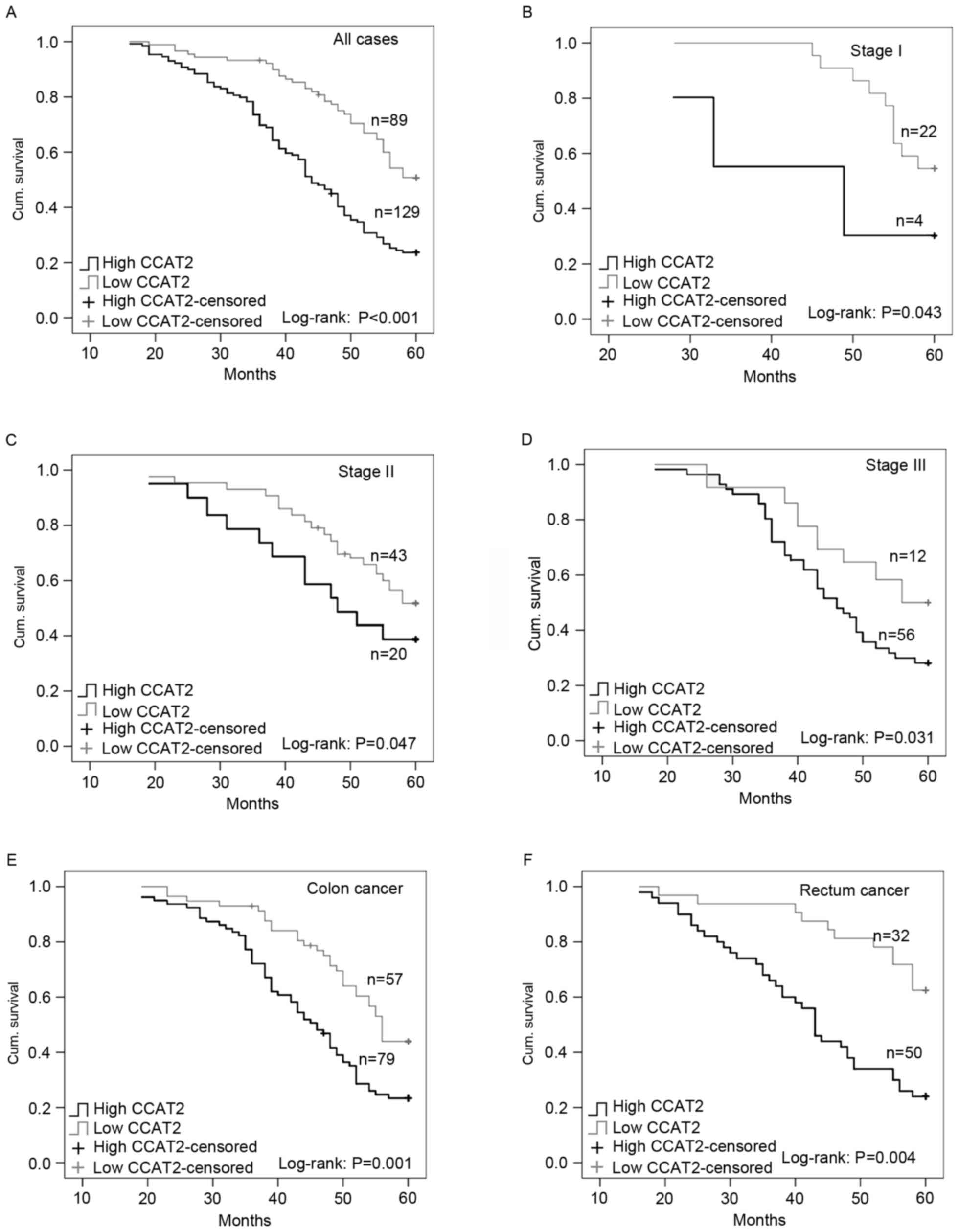
A statistically significant association between high
level of CCAT2 expression and poor overall survival times was
observed in patients with CRC (P<0.001; Fig. 3A). Kaplan-Meier analysis revealed that
patients with CRC with high CCAT2 expression had shorter overall
survival times compared with patients with a low level of CCAT2
expression (Fig. 3A; P<0.001).
Similar to results of disease-free survival, compared with other
patients, patients with poor differentiation, deeper tumor
infiltration, increased incidence of lymph node metastasis, distant
metastasis, vascular invasion and advanced TNM stage experienced
shorter overall survival times compared with those without.
However, other clinical factors had no prognostic value on overall
survival of patients with CRC. Unadjusted RR values are shown in
Table III.
 | Table III.Overall risk ratio. |
Table III.
Overall risk ratio.
|
| Univariate | Multivariate |
|---|
|
|
|
|
|---|
| Parameters | Unadjusted RR (95%
CI) | P-value | Adjusted RR (95%
CI) | P-value |
|---|
| CCAT2
expression | 2.387
(1.663–3.424) | <0.001 | 1.584
(1.013–2.477) | 0.044 |
| Sex | 0.920
(0.660–1.281) | 0.620 | 0.886
(0.627–1.252) | 0.493 |
| Age, years | 1.207
(1.106–1.481) | 0.085 | 1.196
(0.821–1.744) | 0.351 |
| Location | 0.877
(0.620–1.239) | 0.456 | 0.872
(0.612–1.241) | 0.446 |
| Smoking status | 0.891
(0.658–1.347) | 0.869 | 0.831
(0.601–1.278) | 0.727 |
| BMI, kg/m2 | 1.194
(0.858–1.661) | 0.293 | 1.071
(0.747–1.535) | 0.708 |
| Tumor diameter,
cm | 0.991
(0.701–1.402) | 0.960 | 0.872
(0.612–1.241) | 0.253 |
|
Differentiation | 1.727
(1.173–2.542) | 0.006 | 1.468
(0.927–2.327) | 0.102 |
| TNM stage | 1.501
(1.265–1.948) | 0.048 | 0.413
(0.249–0.684) | 0.001 |
| Tumor
infiltration | 2.819
(1.905–4.174) | <0.001 | 1.691
(0.860–3.327) | 0.128 |
| Lymph node
metastasis | 2.819
(1.905–4.174) | <0.001 | 2.136
(1.314–3.472) | 0.002 |
| Distant
metastasis | 2.533
(1.761–3.645) | <0.001 | 2.032
(1.328–3.108) | 0.001 |
| Vascular
invasion | 1.605
(1.151–2.239) | 0.005 | 1.334
(0.942–1.889) | 0.104 |
With regard to TNM stage, significant association
with overall survival was observed in patients with high CCAT2
expression vs. those with low CCAT2 expression with TNM I, II and
III tumors (P<0.05; Fig. 3B-D). In
all three groups, patients with high expression of CCAT2 had
shorter overall survival times than patients with low expression of
CCAT2 (Fig. 3B-D). With regard to
tumor location, patients with colon cancer and patients with rectal
cancer with high CCAT2 expression had shorter overall survival
times compared with those with high CCAT2 expression (Fig. 3E and F). Multivariate analysis
revealed that CCAT2 expression may be a prognostic factor for
overall survival of patients with CRC, independent of other
clinical parameters. The adjusted RR of CRC patients with high
level CCAT2 expression was 1.584 (P=0.044; Table III). In addition, TNM stages, lymph
node metastasis and distant metastasis were also shown to be
independent prognostic factors, subsequent to controlling for all
other clinical parameters (Table
III).
Discussion
The incidence of CRC has been increasing in the last
few decades, particularly in China; CRC has >340,000 newly
diagnosed cases and leads to >80,000 mortalities each year in
China, and ranks as the fifth cause of cancer-associated
mortalities among all malignant diseases (18).
Current clinical TNM staging systems often fail to
discriminate the biological features of a number of tumors
(19). Patients with the same stage
of disease even exhibited a marked discrepancy in prognosis and
survival (19). Genetic alterations
involved in cancer recurrence and outcome may serve as biomarkers
for early detection of metastasis and as a measure for therapeutic
intervention (1). In previous years,
a number of molecules have been used for the prediction of the
prognosis of patients with CRC, but their roles in determining the
individual risk level of the patient are limited (20). Therefore, identification of new
prognostic markers remains important for the prevention and
treatment of CRC (21).
More than 50% of patients have a diagnosis of stage
II or III tumors (1,22). Following curative surgery, patients
with stage III CRC experience 50–60% chance of developing
recurrence (23). 5-FU-based adjuvant
chemotherapy has been accepted as a standard therapy for patients
with stage III CRC, and it has been demonstrated that the overall
survival rate of these patients benefit from it (7,24).
Considering that the role of neoadjuvant chemotherapy for CRC is
also controversial, and neoadjuvant therapy may affect the level of
CCAT2 expression, patients who received neoadjuvant therapy were
excluded.
Early studies proposed that lncRNAs may be simply
transcriptional noise (25). However,
recent studies have revealed that numerous lncRNAs perform
important roles in the regulation of gene transcription, cell
differentiation, genetic and epigenetic and other cellular
activities (10,25,26).
lncRNAs are considered to be new tumor biomarkers for cancer
diagnosis and prognosis (8). The
latest studies have demonstrated that the lncRNA CCAT2 promotes the
MYC and Wnt signaling pathways in CRC in a positive feedback loop
model (15). Furthermore, it is
considered that MYC induces 5-FU-based chemoresistance (27). Regarding these findings, CCAT2
overexpression may induce chemoresistance.
The present study focused on the association of
lncRNA CCAT2 expression with clinicopathological features and
prognosis of patients with CRC. Results of the present study
uncovered that lncRNA CCAT2 expression in CRC tissues was
significantly increased compared with that in adjacent non-tumor
tissues or in noncancerous healthy colon tissues. The qPCR results
revealed that CCAT2 expression progressively increased between TNM
stage I and IV. These results supported the hypothesis that CCAT2
facilitates carcinogenesis and is associated with CRC
progression.
In addition, statistical analysis demonstrated that
high CCAT2 expression was associated with poor cell
differentiation, deeper tumor infiltration, vascular invasion,
lymph node metastasis and distance metastasis, which indicated that
CCAT2 upregulation performs an important role in CRC progression.
However, CCAT2 expression was not associated with age, sex, BMI
status, smoking status or tumor size.
The primary goal of the present study was to
determine whether the CCAT2 expression level in primary CRC could
predict disease relapse. In the present cohort, it was demonstrated
that high levels of CCAT2 expression were associated with
unfavorable disease-free survival and overall survival. The
prognostic value of CCAT2 expression for disease-free survival and
overall survival was statistically significant in univariate and
multivariate analyses.
Kaplan-Meier analysis revealed that the patients
with CRC with a high level of CCAT2 expression had shorter
disease-free survival and overall survival times compared with
patients with CRC with a low level of CCAT2 expression.
Multivariate analysis revealed that patients with a high level of
CCAT2 expression had an increased risk of relapse and mortality.
Although survival patterns of colon cancer and rectum cancer were
found to be different, sub-analysis stratified by primary cancer
location showed that these findings for overall CRC could also be
applied to colon cancer and rectal cancer separately. The
Kaplan-Meier analysis stratified by TNM stage demonstrated that
CCAT2 expression was associated with disease-free and overall
survival in patients with TNM stage I–III tumors.
Thus far, there is no definitive approach to predict
which patients will develop recurrent disease. Prolongation of
disease-free survival means prevention or delay of recurrence or
metastasis. In this regard, results of the present study
demonstrated that measurements of CCAT2 expression may be helpful
to identify patients who were at high risk of early recurrence or
metastasis. Therefore, this may contribute to a tailored treatment
regime for individual patients, thus preventing patients from
receiving insufficient or excessive adjuvant treatment.
In summary, the present study demonstrated that
lncRNA CCAT2 expression may be an independent biomarker for CRC
diagnosis. Furthermore, CCAT2 may be a prognostic marker to
evaluate recurrence, early metastasis and prognosis of CRC, and it
also may be a potential therapeutic target in molecular
therapy.
Acknowledgements
The present study was supported by grants from the
National Natural Science Foundation of China (grant no. 81641098),
the Chinese Society of Clinical Oncology-Merck Serono Oncology
Research Fund (grant no. YMX2015-002) and the Research Foundation
of Peking University First Hospital (grant no. 2016QN20).
References
|
1
|
Brenner H, Kloor M and Pox CP: Colorectal
cancer. Lancet. 383:1490–1502. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Fedirko V, Romieu I, Aleksandrova K,
Pischon T, Trichopoulos D, Peeters PH, Romaguera-Bosch D,
Bueno-de-Mesquita HB, Dahm CC, Overvad K, et al: Pre-diagnostic
anthropometry and survival after colorectal cancer diagnosis in
Western European populations. Int J Cancer. 135:1949–1960. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Torre LA, Bray F, Siegel RL, Ferlay J,
Lortet-Tieulent J and Jemal A: Global cancer statistics, 2012. CA
Cancer J Clin. 65:87–108. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Gustavsson B, Carlsson G, Machover D,
Petrelli N, Roth A, Schmoll HJ, Tveit KM and Gibson F: A review of
the evolution of systemic chemotherapy in the management of
colorectal cancer. Clin Colorectal Cancer. 14:1–10. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Teufel A, Gerken M, Hartl J, Itzel T,
Fichtner-Feigl S, Stroszczynski C, Schlitt HJ, Hofstädter F and
Klinkhammer-Schalke M: Benefit of adjuvant chemotherapy in patients
with T4 UICC II colon cancer. BMC Cancer. 15:4192015. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Barton MK: Oxaliplatin in the adjuvant
treatment of colon cancer. CA Cancer J Clin. 62:3–4. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Wolpin BM, Meyerhardt JA, Mamon HJ and
Mayer RJ: Adjuvant treatment of colorectal cancer. CA Cancer J
Clin. 57:168–185. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Angrand PO, Vennin C, Le Bourhis X and
Adriaenssens E: The role of long non-coding RNAs in genome
formatting and expression. Front Genet. 6:1652015. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Esteller M: Non-coding RNAs in human
disease. Nat Rev Genet. 12:861–874. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Xie X, Tang B, Xiao YF, Xie R, Li BS, Dong
H, Zhou JY and Yang SM: Long non-coding RNAs in colorectal cancer.
Oncotarget. 7:5226–5239. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Deng Q, He B, Gao T, Pan Y, Sun H, Xu Y,
Li R, Ying H, Wang F, Liu X, et al: Up-regulation of 91H promotes
tumor metastasis and predicts poor prognosis for patients with
colorectal cancer. PLoS One. 9:e1030222014. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Yu H, Xu Q, Liu F, Ye X, Wang J and Meng
X: Identification and validation of long noncoding RNA biomarkers
in human non-small-cell lung carcinomas. J Thorac Oncol.
10:645–654. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Svoboda M, Slyskova J, Schneiderova M,
Makovicky P, Bielik L, Levy M, Lipska L, Hemmelova B, Kala Z,
Protivankova M, et al: HOTAIR long non-coding RNA is a negative
prognostic factor not only in primary tumors, but also in the blood
of colorectal cancer patients. Carcinogenesis. 35:1510–1515. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Zheng HT, Shi DB, Wang YW, Li XX, Xu Y,
Tripathi P, Gu WL, Cai GX and Cai SJ: High expression of lncRNA
MALAT1 suggests a biomarker of poor prognosis in colorectal cancer.
Int J Clin Exp Pathol. 7:3174–3181. 2014.PubMed/NCBI
|
|
15
|
Ling H, Spizzo R, Atlasi Y, Nicoloso M,
Shimizu M, Redis RS, Nishida N, Gafà R, Song J, Guo Z, et al:
CCAT2, a novel noncoding RNA mapping to 8q24, underlies metastatic
progression and chromosomal instability in colon cancer. Genome
Res. 23:1446–1461. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Edge SB and Compton CC: The American Joint
Committee on Cancer: The 7th edition of the AJCC cancer staging
manual and the future of TNM. Ann Surg Oncol. 17:1471–1474. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Zheng R, Zeng H, Zhang S, Chen T and Chen
W: National estimates of cancer prevalence in China, 2011. Cancer
Lett. 13:00620–00625. 2015.
|
|
19
|
Brenner H, Kloor M and Pox CP: Colorectal
cancer. Lancet. 383:1490–1502. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Li W, Qiu T, Ling Y, Guo L, Li L and Ying
J: Molecular pathological epidemiology of colorectal cancer in
Chinese patients with KRAS and BRAF mutations. Oncotarget.
6:39607–39613. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Carethers JM: Biomarker-directed Targeted
therapy in colorectal cancer. J Dig Cancer Rep. 3:5–10.
2015.PubMed/NCBI
|
|
22
|
Tsikitis VL, Larson DW, Huebner M, Lohse
CM and Thompson PA: Predictors of recurrence free survival for
patients with stage II and III colon cancer. BMC cancer.
14:3362014. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Rodriguez-Moranta F, Saló J, Arcusa A,
Boadas J, Piñol V, Bessa X, Batiste-Alentorn E, Lacy AM, Delgado S,
Maurel J, et al: Postoperative surveillance in patients with
colorectal cancer who have undergone curative resection: A
prospective, multicenter, randomized, controlled trial. J Clin
Oncol. 24:386–393. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Schmoll HJ, Tabernero J, Maroun J, de
Braud F, Price T, Van Cutsem E, Hill M, Hoersch S, Rittweger K and
Haller DG: Capecitabine plus oxaliplatin compared with
fluorouracil/folinic acid as adjuvant therapy for stage III colon
cancer: Final results of the NO16968 randomized controlled phase
III trial. J Clin Oncol. 33:3733–3740. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Schmitt AM and Chang HY: Long Noncoding
RNAs in Cancer Pathways. Cancer Cell. 29:452–463. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Meseure D, Drak Alsibai K, Nicolas A,
Bieche I and Morillon A: Long noncoding RNAs as new Architects in
cancer epigenetics, prognostic biomarkers, and potential
therapeutic targets. Biomed Res Int. 2015:3202142015. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Kugimiya N, Nishimoto A, Hosoyama T, Ueno
K, Enoki T, Li TS and Hamano K: The c-MYC-ABCB5 axis plays a
pivotal role in 5-fluorouracil resistance in human colon cancer
cells. J Cell Mol Med. 19:1569–1581. 2015. View Article : Google Scholar : PubMed/NCBI
|