Introduction
Gastric cancer (GC) is one of the most frequently
occurring malignant tumors worldwide (1). A high proportion of patients with GC are
diagnosed at a late stage, when extensive invasion and lymphatic
metastasis may have already occurred (2,3). Even
following surgical resection, mortality due to recurrent disease as
a result of metastasis and drug resistance is frequent (4,5). To
investigate useful diagnostic biomarkers for early GC diagnosis,
understanding the underlying mechanisms of GC is critical.
MicroRNAs (miRNAs or miRs) are a class of small,
non-coding RNAs (~22 nucleotides in length) that serve key roles in
various biological processes, including cell proliferation,
metabolism, differentiation and apoptosis, at the
post-transcriptional level (6,7). MiRNAs
interact with a messenger RNA (mRNA) target to interfere with its
translation into protein or to promote mRNA degradation, typically
at the 3′-untranslated region (UTR) (8,9). MiRNAs
may be used as molecular biomarkers for the diagnosis of cancer; it
has been reported that numerous miRNAs are associated with the
carcinogenesis of various types of cancer (10).
MiR-203a has been reported to be associated with
various types of tumor, including esophageal squamous cell
carcinoma (11), hepatocellular
carcinoma (12), non-small cell lung
cancer (13) and colorectal carcinoma
(14). However, the function of
miR-203a in GC remains unclear. The aim of the present study was to
explore the function of miR-203a in GC carcinogenesis. The present
study demonstrated that overexpression of miR-203a could suppress
the proliferation of GC cells. In addition, E2F transcription
factor 3 (E2F3) was identified as a direct target of miR-203a by
luciferase activity assay.
Materials and methods
Cell culture
SGC-7901, AGS and HEK293 cells were obtained from
the Cell Bank of Type Culture Collection of the Chinese Academy of
Sciences (Shanghai, China). Cells were grown in Dulbecco's modified
Eagle's medium (HyClone; GE Healthcare Life Sciences, Logan, UT,
USA) supplemented with 10% fetal bovine serum (HyClone; GE
Healthcare Life Sciences). Cells were incubated at 37°C in a
humidified atmosphere of 5% CO2 in air.
Plasmid vector constructs
Synthetic oligonucleotides containing the 3′-UTR
sequences of wild-type (Wt) and mutant (Mut) E2F3 were cloned into
pmirGLO Dual-Luciferase miRNA Target Expression Vector (Promega
Corporation, Madison, WI, USA). The miR-203a inhibitor
(5′-CTAGTGGTCCTAAACATTTCAC-3′); and inhibitor-control
(5′-TGACTGTACTGACTCGACTG-3′) were synthesized by Sangon Biotech
Co., Ltd., Shanghai, China; the miR-203a mimic (sense,
5′-GUGAAAUGUUUAGGACCACUAG-3′; and antisense,
5′-AGUGGUCCUAAACAUUUCACUU-3′), miR-control (sense,
5′-UUCUCCGAACGUGUCACGUTT-3′; and antisense,
5′-ACGUGACACGUUCGGAGAATT-3′). Small interfering (si) RNA targeting
E2F3 (si-E2F3) and a si-control (15)
were purchased from Shanghai GenePharma Co., Ltd., Shanghai,
China).
Luciferase activity assay
RegRNA (regrna.mbc.nctu.edu.tw/html/prediction.html) was
used to predict the potential target genes for miR-203a. To
determine whether E2F3 was a direct target of miR-203a, E2F3 Wt
sense, 5′-CCTGTGGCACCCATCACCATTTCAA-3′ and antisense,
5′-TTGAAATGGTGATGGGTGCCACAGG-3′; and E2F3 Mut sense,
5′-CCTGTGGCACCCATCACCATAAGAA-3′ and antisense,
5′-TTCTTATGGTGATGGGTGCCACAGG-3′. Wt or Mut E2F3 3′-UTR were cloned
into pmirGLO plasmids (Promega Corporation) and Wt and Mut E2F3
3′-UTR pmirGLO plasmids were co-transfected with miR-203a into
HEK293 cells using Lipofectamine 2000 (Thermo Fisher Scientific,
Inc., Waltham, MA, USA). Another group of HEK293 cells were
co-transfected with miR-203a and an empty pmirGLO plasmid as a
control. After 48 h, a luciferase activity assay was performed
using a Dual-Luciferase Reporter (DLR) Assay System (Promega
Corporation), according to the manufacturer's protocol.
Cell proliferation assay
Cells were plated at a density of 5,000 cells/well
in 96-well plates. miR-203a mimic, miR-control, miR-203a inhibitor,
inhibitor-control, si-E2F3 or si-control plasmids were transfected
with Lipofectamine 2000 into SGC-7901 or AGS cells. An MTT assay
with FLUOstar OPTIMA (BMG Labtech GmbH, Ortenberg, Germany) was
used to measure the extent of cell proliferation. The optical
density was measured at 492 nm at 24, 48 and 72 h
post-transfection.
Colony formation assays
SGC-7901 cells were seeded into 12-well plates at a
density of 1000 cells/ml, 2ml/well. AGS cells were seeded into
6-well plates at adensity of 1000 cells/ml, 2ml/well. The cells
were transfected with miR-203a, miR-control, miR-203a inhibitor,
inhibitor-control, si-E2F3 or si-control. The cells were incubated
in the aforementioned conditions for 14 days and washed with PBS,
then stained with 0.1% crystal violet for 30 min at 37°C. Images of
the colonies were captured by Quantity One (Bio-Rad Laboratories,
Inc., Hercules, CA, USA).
Western blot analysis
SGC-7901 cells were transfected with miR-203a or
miR-control. After 48 h, the cells were lysed in
radioimmunoprecipitation assay buffer (Sigma-Aldrich; Merck KGaA,
Darmstadt, Germany) for 1 h at 4°C. Cell lysates were then
suspended by centrifugation at 12,000 × g at 4°C for 20 min. The
protein concentration was determined with a BCA protein assay kit
(Pierce; Thermo Fisher Scientific, Inc.). Proteins were separated
with 10% SDS-PAGE and transferred to a nitrocellulose membrane. The
membrane was blocked with 5% skimmed milk in TBS containing 0.05%
Tween-20 for 2 h at room temperature. The membrane was then
incubated at 4°C overnight with primary antibodies against E2F3
(1:300; bs-1722R; Beijing Biosynthesis Biotechnology Co., Ltd.,
Beijing, China) and β-actin (١:١,500; sc-47778; Santa Cruz
Biotechnology, Inc., Dallas, TX, USA), followed by incubation with
goat anti-mouse IgG (cat. no. 115-035-003; 1:1,000; Jackson
ImmunoResearch Laboratories, Inc., West Grove, PA, USA) or goat
anti-rabbit antibody IgG (cat. no. 111-035-144; 1:1,000; Jackson
ImmunoResearch Laboratories, Inc.) for 1 h at room temperature.
Immobilon Western Chemiluminescent horseradish-peroxidase substrate
(EMD Millipore, Billerica, MA, USA) were used to visualize the
protein bands.
Statistical analysis
Data are expressed as the mean ± standard error from
three independent experiments. Student's t-tests with SPSS 13.0
software (SPSS Inc., Chicago, IL, USA) were utilized for
statistical analysis. P<0.05 was considered to indicate a
statistically significant difference.
Results
miR-203a suppresses the proliferation
of GC cells
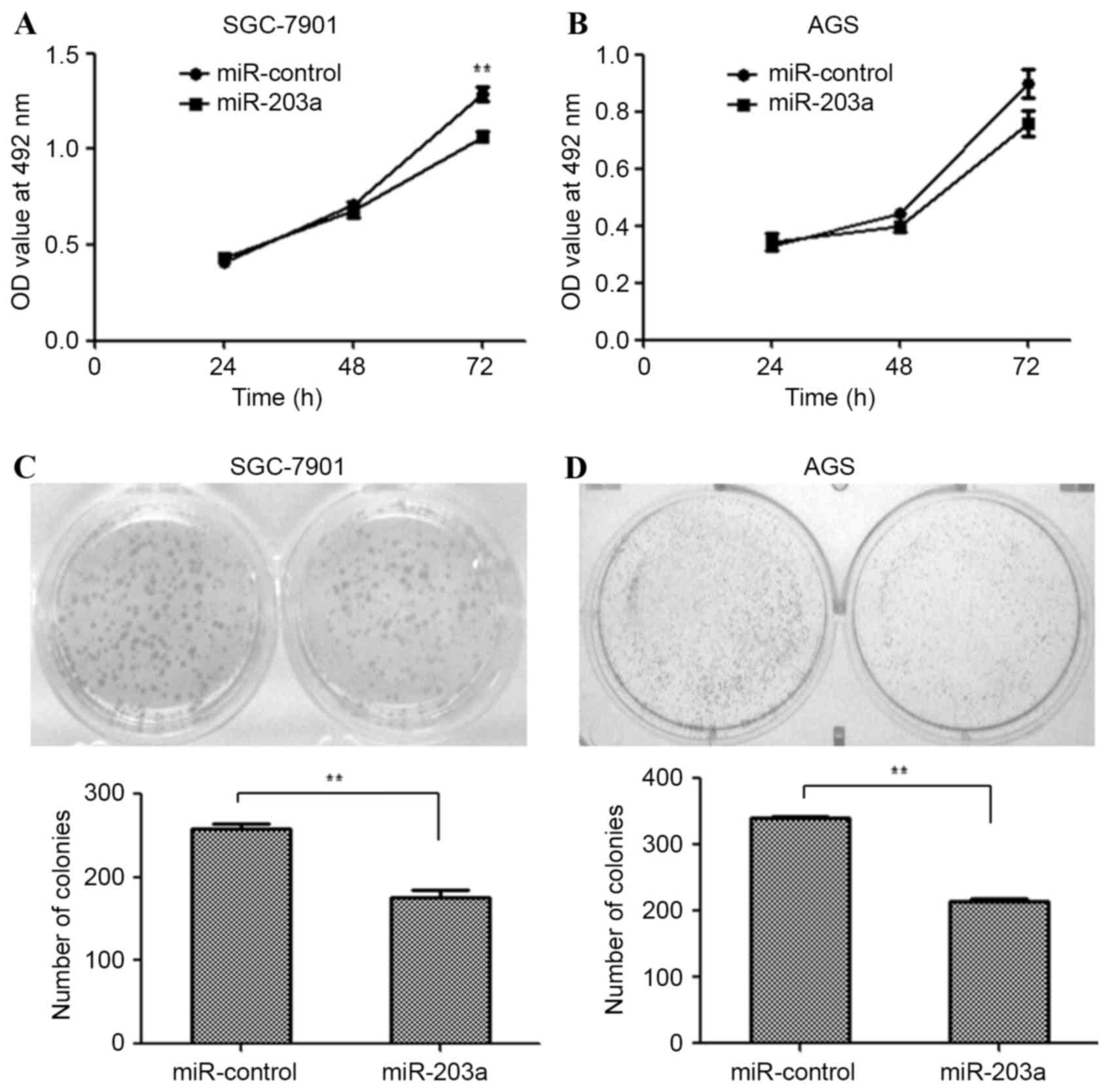
SGC-7901 and AGS cells were transfected with a
miR-203a mimic or miR-control plasmid and an MTT assay was used to
examine the proliferation of these cells. The MTT assay revealed
that the proliferation of SGC-7901 cells was significantly
inhibited by miR-203a overexpression (Fig. 1A; P<0.01), whereas the
proliferation of AGS cells was not significantly affected (Fig. 1B). Similarly, colony formation ability
was assessed for SGC-7901 and AGS cells transfected as described
above. Fewer colonies were observed in miR-203a mimic-transfected
SGC-7901 (Fig. 1C; P<0.01) and AGS
(Fig. 1D; P<0.01) cells, compared
with miR-control-transfected cells. This result demonstrates that
miR-203a inhibits the proliferation of GC cells.
Inhibition of miR-203a increases the
proliferation of GC cells
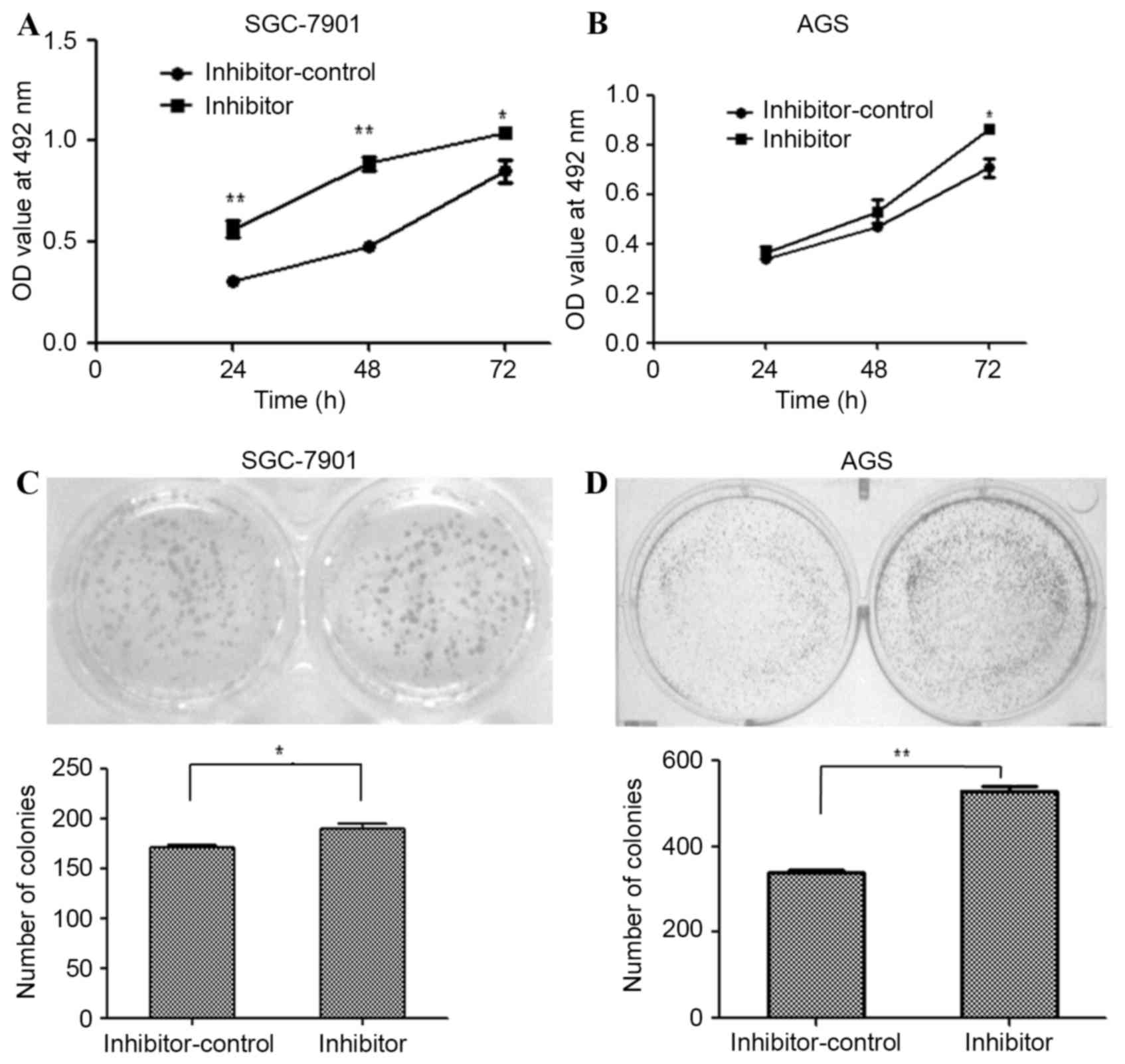
miR-203a inhibitor oligonucleotides were used to
silence endogenous miR-203a expression in SGC-7901 and AGS cells.
The effect of this inhibitor on SGC-7901 and AGS proliferation was
determined by an MTT assay at 24, 48 and 72 h post-transfection.
The miR-203a inhibitor increased the proliferation of SGC-7901
(Fig. 2A; P<0.01) and AGS cells
(Fig. 2B; P>0.05), as determined
by the MTT assays. The formation of colonies was upregulated by
transfection with the inhibitor in SGC-7901 (Fig. 2C; P<0.01) and AGS (Fig. 2D; P<0.01) cells. Taken together,
these results demonstrate that inhibition of miR-203a contributes
to the proliferation of GC cells.
E2F3 is a direct target of
miR-203a
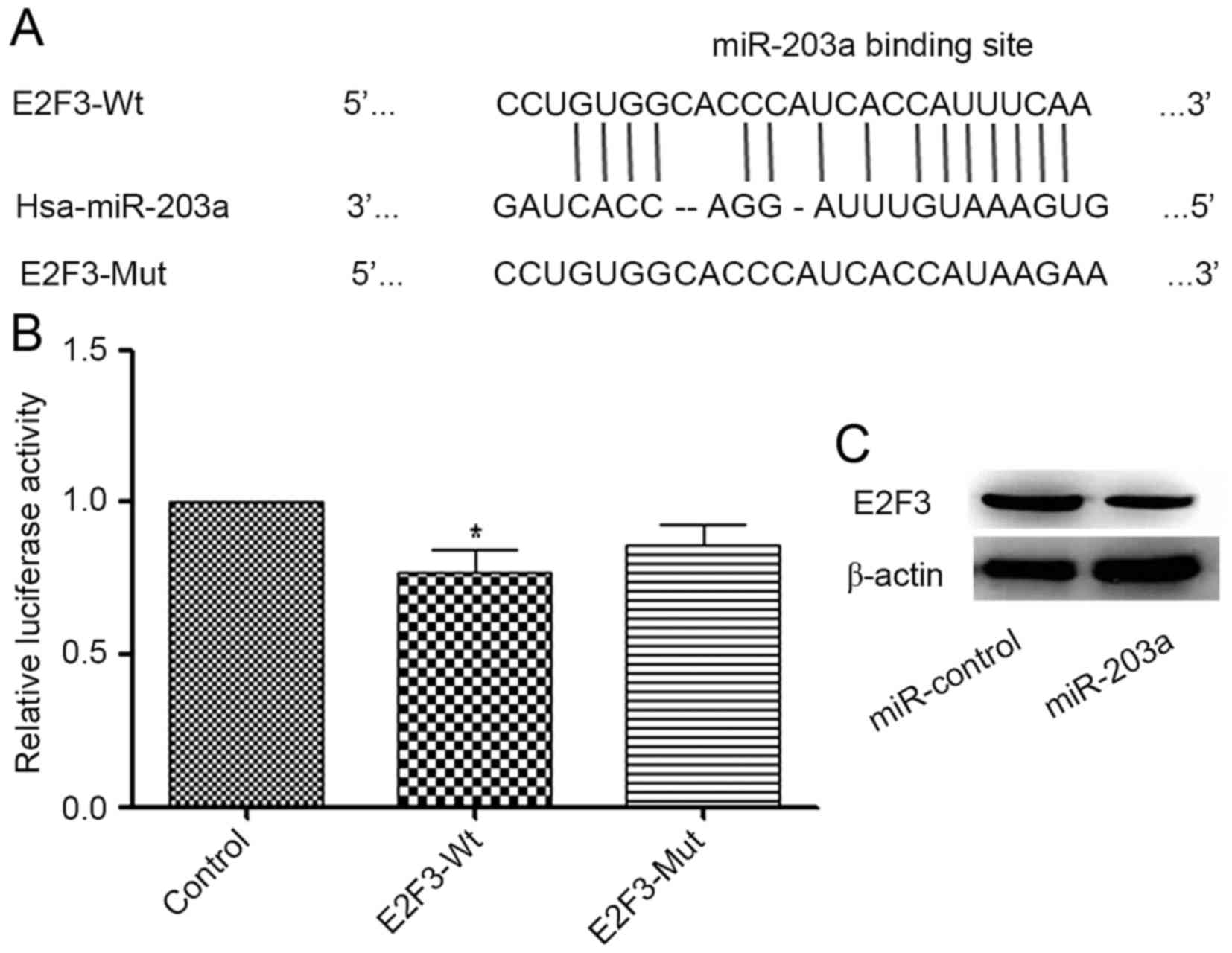
RegRNA (regrna.mbc.nctu.edu.tw/html/prediction.html) was
used to identify potential target genes of miR-203a. A binding site
for miR-203a in the E2F3 3′-UTR was identified (Fig. 3A). A luciferase reporter assay
demonstrated that the addition of miR-203a decreased the luciferase
activity of E2F3-Wt significantly when compared with that of the
control group, whereas the change in the luciferase activity of
E2F3-Mut was not significant (Fig.
3B). To further investigate the association between miR-203a
and E2F3, a western blot analysis was performed. The result
demonstrated that the protein expression of E2F3 was decreased in
the miR-203a-overexpressing group compared with that in the control
group (Fig. 3C). Collectively, these
data indicate that E2F3 is a direct target of miR-203a.
Silencing the expression of E2F3
inhibits the proliferation of GC cells
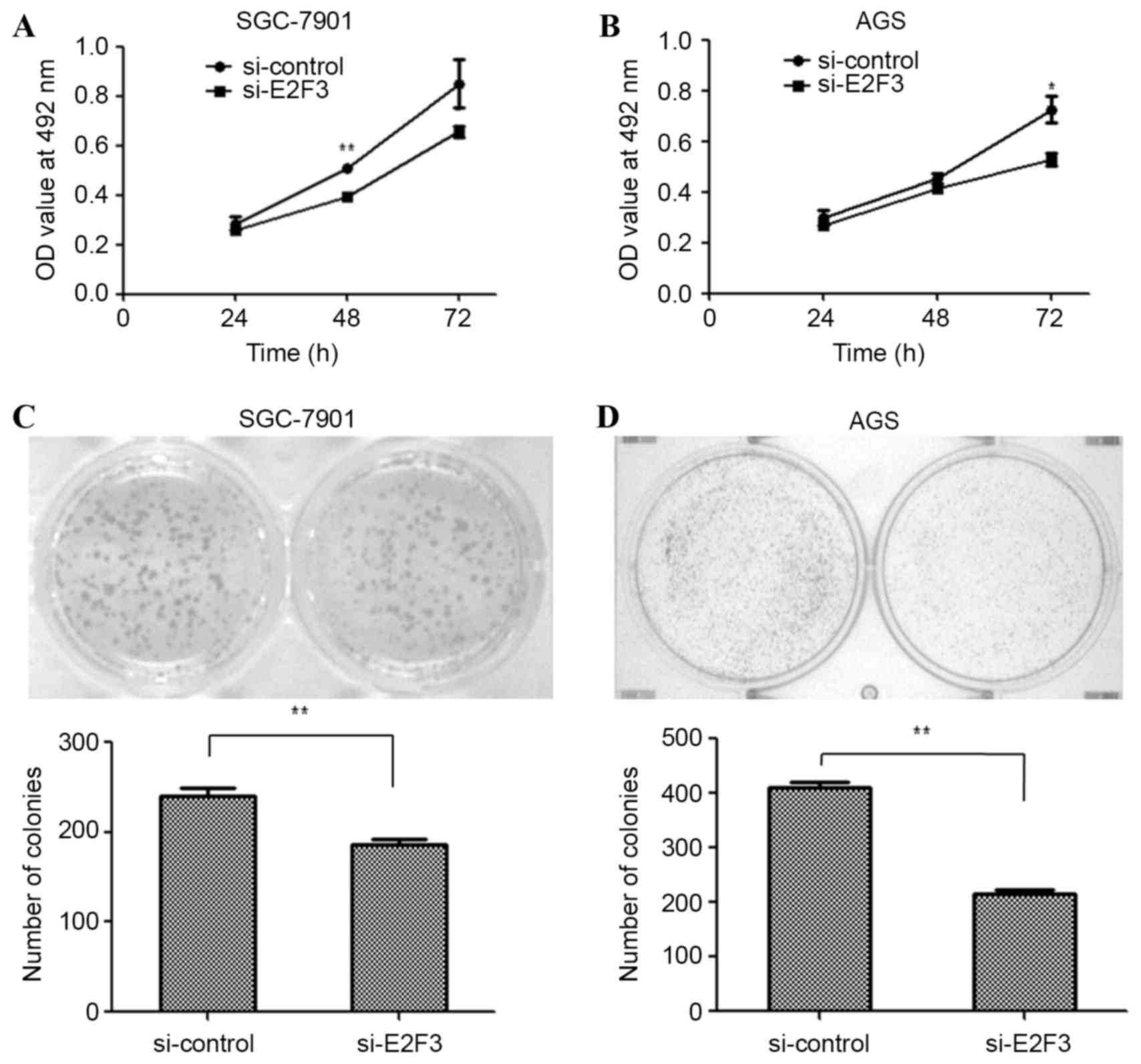
To examine the effects of E2F3 on the proliferation
of SGC-7901 and AGS cells, the cells were transfected with si-E2F3
or a si-control. An MTT assay was performed to examine the
proliferation of SGC-7901 and AGS cells. The MTT assay revealed
that cell proliferation was inhibited in si-E2F3-transfected
SGC-7901 (Fig. 4A; P<0.01 at 48 h)
and AGS (Fig. 4B; P<0.05 at 72 h)
cells. Similarly, the colony formation assay was employed in
si-E2F3- and si-control-transfected SGC-7901 and AGS cells. The
results revealed that fewer colonies were formed by
si-E2F3-transfected SGC-7901 (Fig.
4C; P<0.01) and AGS cells (Fig.
4D; P<0.01), compared with si-control-transfected cells. The
data suggest that miR-203a suppresses GC cell proliferation by
downregulating E2F3.
Discussion
E2F3 is a member of the E2F family, which serves a
key function in the control of cell cycle progression (16). It has been reported that E2F3
regulates cell cycle-dependent gene expression during the G1/S
transition (17). E2F3 may function
as an oncogene in GC (18). Previous
studies have reported that overexpression of E2F3 is a frequent
oncogenic event in human tumorigenesis and that this may be
regulated by miRNA (19,20). In the present study, E2F3 was
confirmed to be a direct target of miR-203a with luciferase
activity and western blot assays. In previous studies, a number of
miRNAs have been categorized as oncogenes or tumor-suppressor
genes; and may contribute to cancer progression by interfering with
mRNA stability or protein translation to regulate the gene
expression of oncogenic or tumor-suppressive proteins (21).
miR-203 has been reported to be involved in several
types of cancer (22,23) and may function as a tumor suppressor
involved in proliferation, apoptosis, invasion and migration
(24). For example, miR-203a
suppresses the development of hepatocellular carcinoma cells by
targeting homeobox D3 (25).
Additionally, miR-203 inhibits proliferation and migration of lung
cancer cells by targeting PKCα (26).
Therefore, miR-203a may serve a critical function in the
development of cancer. The present study suggested that miR-203a
may serve an important role in suppressing GC proliferation and may
regulate the expression of E2F3. The results of the present study
provide an improved understanding of the tumor suppressive role of
miR-203a during GC progression.
In summary, the present study demonstrated that the
overexpression of miR-203a suppresses GC cell proliferation by
targeting and reducing the relative expression level of the cell
cycle-regulatory protein E2F3. The effect on GC cell proliferation
following transfection with si-E2F3 was comparable to the effect of
miR-203a overexpression. Furthermore, a miR-203a inhibitor
upregulated the proliferation of GC cells. The tumor-suppressive
function of miR-203a may provide new insight for potential
therapies for GC. In the future, investigation of the
E2F3-associated signaling pathway is to be performed by our group
in order to explore more deeply the mechanism by which miR-203a
suppresses the proliferation of GC cells.
Acknowledgements
The present study was supported by the Natural
Science Foundation of Xinjiang Uygur Autonomous Region (grant no.
2013211A114).
References
|
1
|
Jemal A, Bray F, Center MM, Ferlay J, Ward
E and Forman D: Global cancer statistics. CA Cancer J Clin.
61:69–90. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Dassen A, Lemmens VE, van de Poll-Franse
LV, Creemers GJ, Brenninkmeijer SJ, Lips DJ, Wurff Vd AA, Bosscha K
and Coebergh JW: Trends in incidence, treatment and survival of
gastric adenocarcinoma between 1990 and 2007: A population-based
study in the Netherlands. Eur J Cancer. 46:1101–1110. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Wu HH, Lin WC and Tsai KW: Advances in
molecular biomarkers for gastric cancer: miRNAs as emerging novel
cancer markers. Expert Rev Mol Med. 16:e12014. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Kim SJ, Wang YG, Lee HW, Kang HG, La SH,
Choi IJ, Irimura T, Ro JY, Bresalier RS and Chun KH: Up-regulation
of neogenin-1 increases cell proliferation and motility in gastric
cancer. Oncotarget. 5:3386–3398. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Ohtsu A: Chemotherapy for metastatic
gastric cancer: Past, present, and future. J Gastroenterol.
43:256–264. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Wienholds E and Plasterk RH: MicroRNA
function in animal development. FEBS Lett. 579:5911–5922. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
He L and Hannon GJ: MicroRNAs: Small RNAs
with a big role in gene regulation. Nat Rev Genet. 5:522–531. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Fabian MR, Sonenberg N and Filipowicz W:
Regulation of mRNA translation and stability by microRNAs. Annu Rev
Biochem. 79:351–379. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Shah PP, Hutchinson LE and Kakar SS:
Emerging role of microRNAs in diagnosis and treatment of various
diseases including ovarian cancer. J Ovarian Res. 2:112009.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Liu Y, Dong Z, Liang J, Guo Y, Guo X, Shen
S, Kuang G and Guo W: Methylation-mediated repression of potential
tumor suppressor miR-203a and miR-203b contributes to esophageal
squamous cell carcinoma development. Tumour Biol. 37:5621–5632.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Liu D, Wu J, Liu M, Yin H, He J and Zhang
B: Downregulation of miRNA-30c and miR-203a is associated with
hepatitis C virus core protein-induced epithelial-mesenchymal
transition in normal hepatocytes and hepatocellular carcinoma
cells. Biochem Biophys Res Commun. 464:1215–1221. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Lin QH, Zhang KD, Duan HX, Liu MX, Wei WL
and Cao Y: ERGIC3, which is regulated by miR-203a, is a potential
biomarker for non-small cell lung cancer. Cancer Sci.
106:1463–1473. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Kara M, Yumrutas O, Ozcan O, Celik OI,
Bozgeyik E, Bozgeyik I and Tasdemir S: Differential expressions of
cancer-associated genes and their regulatory miRNAs in colorectal
carcinoma. Gene. 567:81–86. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Yang Y, Chang S, Zhao Z, et al:
MicroRNA-214 suppresses the proliferation of human hepatocellular
carcinoma cells by targeting E2F3. Oncol Lett. 10:3779–3784.
2015.PubMed/NCBI
|
|
16
|
Julian LM, Vandenbosch R, Pakenham CA,
Andrusiak MG, Nguyen AP, McClellan KA, Svoboda DS, Lagace DC, Park
DS, Leone G, et al: Opposing regulation of Sox2 by cell-cycle
effectors E2f3a and E2f3b in neural stem cells. Cell Stem Cell.
12:440–452. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Sharma N, Timmers C, Trikha P, Saavedra
HI, Obery A and Leone G: Control of the p53-p21CIP1 Axis by E2f1,
E2f2, and E2f3 is essential for G1/S progression and cellular
transformation. J Biol Chem. 281:36124–36131. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Li X, Li H, Zhang R and Liu J and Liu J:
MicroRNA-449a inhibits proliferation and induces apoptosis by
directly repressing E2F3 in gastric cancer. Cell Physiol Biochem.
35:2033–2042. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Oeggerli M, Tomovska S, Schraml P,
Calvano-Forte D, Schafroth S, Simon R, Gasser T, Mihatsch MJ and
Sauter G: E2F3 amplification and overexpression is associated with
invasive tumor growth and rapid tumor cell proliferation in urinary
bladder cancer. Oncogene. 23:5616–5623. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Miles WO, Tschop K, Herr A, Ji JY and
Dyson NJ: Pumilio facilitates miRNA regulation of the E2F3
oncogene. Genes Dev. 26:356–368. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wiemer EA: The role of microRNAs in
cancer: No small matter. Eur J Cancer. 43:1529–1544. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Lee SA, Kim JS, Park SY, Kim HJ, Yu SK,
Kim CS, Chun HS, Kim J, Park JT, Go D, et al: miR-203 downregulates
Yes-1 and suppresses oncogenic activity in human oral cancer cells.
J Biosci Bioeng. 120:351–358. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Xu L, Shen B, Chen T and Dong P: miR-203
is involved in the laryngeal carcinoma pathogenesis via targeting
VEGFA and Cox-2. Onco Targets Ther. 9:4629–4637. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Hu G, Lai P, Liu M, Xu L, Guo Z, Liu H, Li
W, Wang G, Yao X, Zheng J, et al: miR-203a regulates proliferation,
migration, and apoptosis by targeting glycogen synthase kinase-3β
in human renal cell carcinoma. Tumour Biol. 35:11443–11453. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Wang L, Sun H, Wang X, Hou N, Zhao L, Tong
D, He K, Yang Y, Song T, Yang J, et al: EGR1 mediates miR-203a
suppress the hepatocellular carcinoma cells progression by
targeting HOXD3 through EGFR signaling pathway. Oncotarget.
7:45302–45316. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Wang C, Wang X, Liang H, Wang T, Yan X,
Cao M, Wang N, Zhang S, Zen K, Zhang C, et al: miR-203 inhibits
cell proliferation and migration of lung cancer cells by targeting
PKCα. PloS One. 8:e739852013. View Article : Google Scholar : PubMed/NCBI
|