Introduction
Laryngeal carcinoma, of which >95% of cases are
laryngeal squamous cell carcinoma (LSCC), is a prevalent malignancy
of the head and neck, with an incidence of 3.5–5.5/100,000 people
and a mortality rate of 2.1–2.4 per 100,000 people worldwide in
2008 (1,2). Although there have been significant
improvements in terms of diagnosis and treatment, the clinical
outcome of therapy for patients with LSCC, particularly those with
advanced stages of the disease, remains poor (3). Therefore, it is necessary to explore the
molecular mechanisms of LSCC, which will contribute to increasing
survival rates and improving prognosis.
Previous studies have revealed that retinoic acid
receptors (RARs) serve important functions in various types of
cancer (4). The RAR family contains
three different subtypes (namely, RARα, RARβ and RARγ). As RARs are
important members of the nuclear receptor superfamily, the three
subtypes function as transcription factors that are essential in
embryonic development, the maintenance of differentiated cellular
phenotypes, metabolism and cell death (5). The three RAR subtypes are encoded by
different genes and they differ in terms of structure and
distribution as well as possessing unique functions. Increasingly,
the evidence has indicated that the aberrant expression of RARα is
present in various types of cancer. RARα was revealed to regulate
the cell differentiation of neurocytoma through the activation of
the transcription of associated target genes (6). The activation of RARα competed with the
fusion protein promyelocytic leukemia gene-RARα, induced by a
mutation to combine with the Fas cell surface death receptor domain
structure, promoting the apoptosis of myeloid leukemia (7). The activation of RARα also promoted cell
cycle arrest and the apoptotic process in lymphoma cells through
the upregulation of reactive oxygen species (8). The protein stability of RARα, enhanced
by ubiquitin, inhibited the proliferation of gastric cancer cells
(9). The contribution of RARα to the
development of hepatocellular carcinoma was associated with its
regulation of cytochrome P450 family 1 subfamily A member 1
expression (10,11). RARα is also involved in the
development of breast cancer and prostate carcinoma (12–14).
However, little is known about the expression and function of RARα
in human LSCC.
In the present study, the function and mechanisms of
RARα in primary LSCC tissues and adjacent normal tissues were
investigated. It was revealed that the overexpression of RARα in
LSCC tissues promoted the proliferation of leukocyte common antigen
(LCA) cells through the regulation of the cell cycle, with
definitive clinical significance. The results of the present study
may provide a reference to gain additional insight into the
potential function of RARα in the development of LCA, suggesting a
potential target for LSCC therapy.
Materials and methods
Patients and tissue samples
Fresh human laryngeal carcinoma and adjacent
noncancerous tissues were obtained from 32 patients who underwent
surgery at the First Affiliated Hospital of Xiamen University
(Xiamen, China). Tumor tissues and adjacent non-tumor tissues were
sampled and immediately soaked in formalin overnight, and then
paraffin embedded. The pathological characteristics were confirmed
by pathological examination. The present study was approved by the
ethics committee of the First Affiliated Hospital of Xiamen
University, and all patients provided written informed consent
prior to participation. None of the patients with LSCC were treated
with radiotherapy, chemotherapy or other types of therapy prior to
surgery. Metastatic tumors from other tissues were excluded from
the study. The clinicopathological characteristics are presented in
Table I.
 | Table I.Associations between RARα and clinical
features of laryngeal squamous cell carcinoma. |
Table I.
Associations between RARα and clinical
features of laryngeal squamous cell carcinoma.
|
|
| RARα |
|
|
|---|
|
|
|
|
|
|
|---|
| Features | N | Low | High | χ2 | P-value |
|---|
| Age, years |
|
|
| 0.0185 | 0.8918 |
|
<59 | 13 | 3 | 10 |
|
|
|
>60 | 19 | 4 | 15 |
|
|
| Sex |
|
|
| 0.0261 | 0.8716 |
| Male | 28 | 6 | 22 |
|
|
|
Female | 4 | 1 | 3 |
|
|
| Pathologic
differentiation |
|
|
| 6.8183 | 0.0331a |
| High | 14 | 6 | 8 |
|
|
|
Middle | 8 | 1 | 7 |
|
|
| Low | 10 | 0 | 10 |
|
|
| T stage |
|
|
| 0.4780 | 0.9237 |
| T1 | 7 | 1 | 6 |
|
|
| T2 | 13 | 3 | 10 |
|
|
| T3 | 9 | 2 | 7 |
|
|
| T4 | 3 | 1 | 2 |
|
|
| Lymph node
metastasis |
|
|
| 1.0135 | 0.3141 |
| N0 | 19 | 3 | 16 |
|
|
| N+ | 13 | 4 | 9 |
|
|
| Distant
metastasis |
|
|
| 0.9874 | 0.3204 |
| M0 | 30 | 6 | 24 |
|
|
| M+ | 2 | 1 | 1 |
|
|
| Clinical stage |
|
|
| 0.5644 | 0.9045 |
| 1 | 6 | 1 | 5 |
|
|
| 2 | 14 | 3 | 11 |
|
|
| 3 | 11 | 3 | 8 |
|
|
| 4 | 1 | 0 | 1 |
|
|
Cell culture
The LSCC cell line AMC-HN-8 purchased from the Cell
Bank of Type Culture Collection of Chinese Academy of Science
(Shanghai, China). Cells were cultured in RPMI-1640 medium (Gibco;
Thermo Fisher Scientific, Inc., Waltham, MA, USA) supplemented with
10% fetal bovine serum (FBS; Hyclone; GE Healthcare Life Sciences,
Logan, UT, USA) and antibiotics (100 U/ml penicillin and 100 mg/ml
streptomycin) at 37°C in a humidified atmosphere of 5%
CO2.
Cell transfection
RARα-specific small interfering (si)RNA (siRARα),
5′-AUCUCUUCAGAACUGCUGCUCUGGG-3′ and the Stealth RNAi™ siRNA
negative control (siCtrl), 5′-UAUCCUUACGAGUACCUUGCGGCUG-3′ were
designed and obtained from Invitrogen; Thermo Fisher Scientific,
Inc. LSCC cells were seeded in 6 well plates at a density of
0.5×106 cells per well. siRARα and siCtrl were
transfected into cells using the LipoRNAiMAX transfection reagent
(Invitrogen; Thermo Fisher Scientific, Inc.), according to the
manufacturer's protocol. Following transfection at 37°C in a
humidified atmosphere with 5% CO2 for 24 h, cells were
collected for subsequent experiments.
Cell proliferation assay
Cell proliferation was analyzed using an MTT assay
(Sigma-Aldrich; Merck KGaA, Darmstadt, Germany). The cells were
seeded at a density of 5×103 cells per well into 96-well
plates overnight. Following the transfection of cells with siRARα
and siCtrl, 20 µl MTT (5 mg/ml) was added to each well, and the
cells were cultured for a further 4 h at 37°C. Then the formazan
crystals formed were dissolved in DMSO. The absorbance was measured
at 490 nm using an ELISA microplate reader. All experiments were
performed in triplicate.
Colony formation
A total of 600 AMC-HN-8 cells were cultured in
6-well plates for 14 days and then fixed and stained with 0.005%
crystal violet for 30 min at room temperature. Colonies >100 µm
in diameter were counted under a light microscope (Olympus BH-2,
Olympus Corporation, Tokyo, Japan).
Reverse transcription quantitative
RT-(q)PCR
Total RNA was extracted from cells using an RNA
Extraction kit (cat. no. DP405-02; Tiangen, Beijing, China)
according to manufacturer's protocol. Reverse transcription was
carried out using the SuperScript III First-Strand Synthesis system
(Invitrogen; Thermo Fisher Scientific, Inc.) for qPCR. The qPCR was
carried out by ABI 7500 Fast Real-Time PCR system (Applied
Biosystems; Thermo Fisher Scientific, Inc.) with SYBR-Green
(Promega Corporation, Madison, WI, USA) as a fluorophore. GAPDH was
used as a control. The primer sequences are listed in Table II. All experiments were performed in
triplicate, and the data were relatively calculated using the
2−ΔΔCq method (15).
 | Table II.Primer sequences for reverse
transcription-quantitative polymerase chain reaction analysis. |
Table II.
Primer sequences for reverse
transcription-quantitative polymerase chain reaction analysis.
| Gene | Sense (5′-3′) | Antisense
(5′-3′) |
|---|
| RARα |
AATACACTACGAACAACAGC |
CGAACTCCACAGTCTTAATG |
| P21 |
CTAGTTCTACCTCAGGCAGCT |
GTCGCTGGACGATTTGAGG |
| P27 |
GTTAGCGGAGCAATGCGC |
CAGGCTTCTTGGGCGTCTG |
| Cyclin A |
AAGCACTCCCTGACTGTGG |
ACTGATGTTTCTTGGTGAC |
| Cyclin B |
CCTCAACCATCCTGGCTGCG |
CTGTTCTTGGCCTCAGTCC |
| Cyclin D |
CCGAGTCACCAGGAACTCGA |
AGGACAGACTCCGCTGTGC |
| Cyclin E |
AATGTCAAGACGAAGTAGCC |
ATTTCCTCAAGTTTGGCTGCA |
Western blotting
Whole-cell lysates were prepared in lysis buffer (50
mM Tris-HCl pH 7.5, 100 mM NaCl, 50 mM NaF, 1 mM Na3VO4, 30 mM
sodium pyrophosphate, 0.5% NP-40 and 0.5 mM PMSF (Sigma-Aldrich;
Merck KGaA) supplemented with EDTA-free protease inhibitor cocktail
(Roche Diagnostics, Basel, Switzerland), and insoluble debris was
pelleted by centrifugation for 20 min at 4°C and 13,000 × g and
removed. Protein concentration was determined by Bradford assay
(Bio-Rad Laboratories, Inc., Hercules, CA, USA), and lysate
proteins denatured by boiling for 5 min in reducing SDS-sample
buffer. Lysate proteins (15 µg total protein/lane) were separated
by 12% SDS-PAGE and transferred to polyvinylidene fluoride
membranes (PVDF, Millipore, Billerica, MA, USA). Membranes were
blocked in 7% bovine serum albumin in TBS and Tween-20 (TBST) for 1
h at room temperature, and probed overnight at 4°C in primary
antibody solutions (RARα, RARβ and RARγ; Abcam, Cambridge, UK;
dilution, 1:1,000) [cyclin-dependent kinase inhibitor 1A (p21),
proliferating cell nuclear antigen (PCNA), protein kinase B (AKT),
phosphorylated (p)-AKT and GAPDH (dilution, 1:500; Santa Cruz
Biotechnology, Inc., Dallas, TX, USA) in TBST. They were then
washed in TBST 3 times for 5 min, and incubated in TBST solution
containing 25% (v/v) non-fat dry milk powder and goat anti-rabbit
(cat. no., 31430) and goat anti-mouse (cat. no. 31460) secondary
antibodies (1:300) conjugated to horseradish peroxidase
(Invitrogen; Thermo Fisher Scientific, Inc.) for 1 h at room
temperature. Finally, the membranes were washed in TBST 3 time for
5 min, and bands imaged using Western Lightning Plus-ECL
(PerkinElmer, Inc., Waltham, MA, USA) and HyBlot ES autoradiography
film (Denville Scientific Inc.). The band intensities for western
blot analysis were quantified using ImageJ 1.48 software (National
Institutes of Health, Bethesda, MD, USA) and normalized to GAPDH.
All experiments were performed in triplicate.
Hematoxylin and eosin staining
(H&E) and immunohistochemistry (IHC)
LSCC and the adjacent noncancerous specimens were
fixed in 10% formalin for 24 h at room temperature, paraffin
embedded and cut into 3-µm thick sections for H&E or IHC. The
H&E staining were performed according to manufacturer's
protocol of H&E Staining kit (Beijing Solarbio Science &
Technology Co., Ltd., Beijing, China). IHC staining using anti-RARα
antibody (1:300; cat. no. ab117728; Abcam) was detected using the
EliVisionTM Plus kit (Fuzhou Maixin Biotech Co., Ltd., Fuzhou,
China) according to the manufacturer's protocol. PBS without
primary antibody was used as the negative control. The IHC staining
result was evaluated based on the proportion of stained tumor
cells. RARα-positive tumor cells were counted in 10 randomly
selected high power fields under a light microscope (Olympus BH-2;
Olympus Corporation) at a magnification of ×400 and was evaluated
according to the following criteria: <5% was defined as -; 5–25%
as +; 25–50% as ++; >50% as +++. Expression of RARα protein was
categorized as low (− to +) or high (++ to +++).
Cell cycle analysis
To detect the cell cycle distribution, cells were
collected following transfection with siRARα or siCtrl at 37°C for
48 h and washed twice with ice cold PBS, and then fixed in ice-cold
70% ethanol at 4°C overnight. Following centrifugation for 5 min at
4°C and 2,000 × g, the cells were resuspended in 100 µg/ml RNase A
at 37°C for 30 min and subsequently stained with 50 µg/ml propidium
iodide (Sangon Biotech Co., Ltd., Shanghai, China) at 4°C for 30
min in the dark. The cells were analyzed using a FACScan flow
cytometer (BD Biosciences, Franklin Lakes, NJ, USA) at 488 nm and
the data was analyzed using ModFit 3.3 (Verity Software House,
Topsham, ME, USA) software.
Statistical analysis
Data analysis was conducted using SPSS software
(version 16.0; SPSS, Inc., Chicago, IL, USA). Continuous data was
expressed as the mean ± the standard error of the mean and analyzed
using one-way analysis of variance with a Tukey's post hoc test,
whilst χ2 or Fisher's exact tests were used to examine
the association between RARα and the clinicopathological parameters
of LSCC. P<0.05 was considered to indicate a statistically
significant difference.
Results
RARα was elevated in laryngeal
carcinoma tissues
IHC was performed to detect the expression levels of
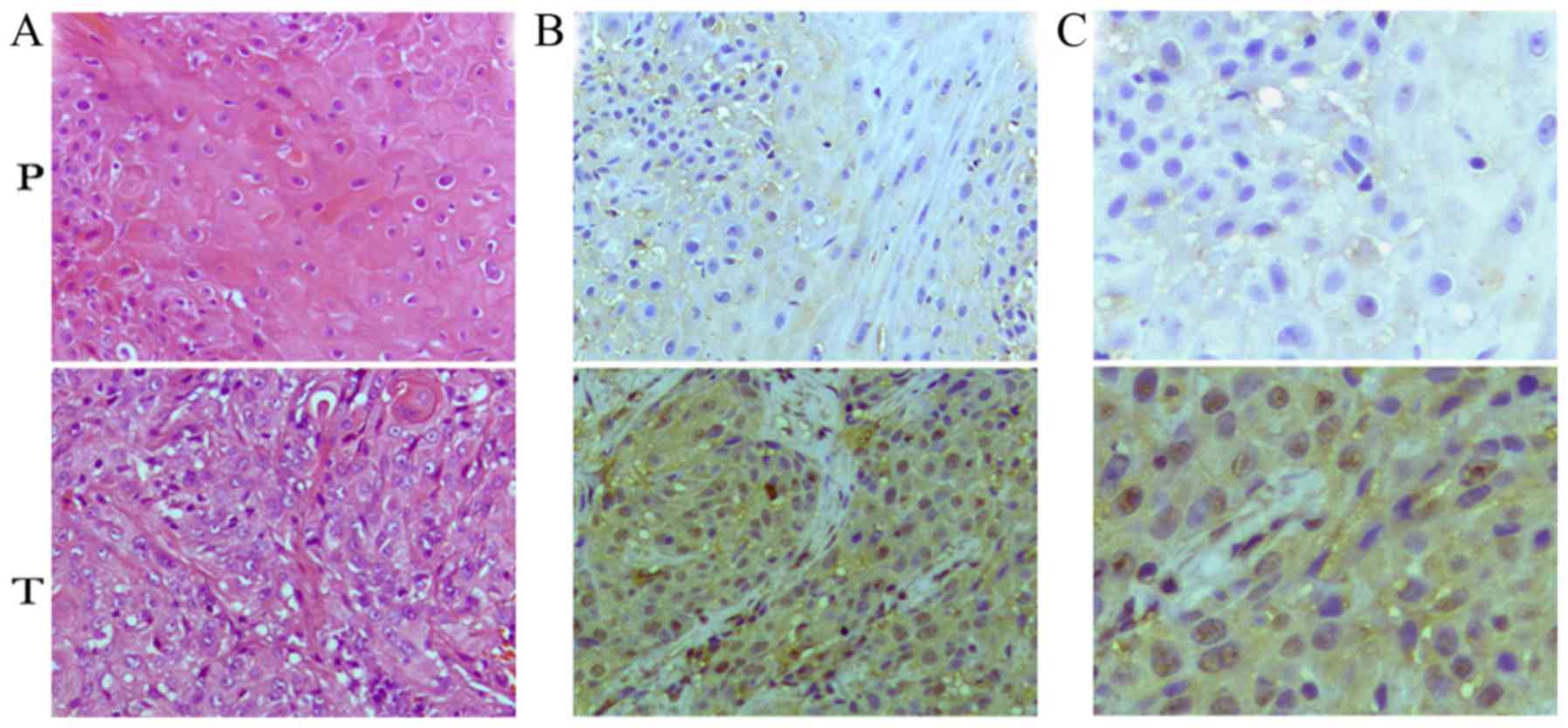
RARα in LSCC tissues and the adjacent noncancerous tissues. As
presented in Fig. 1, the expression
of RARα in the paired adjacent paratumor tissues was weak or
negative, but was strong in the LSCC tissues. In addition, RARα
exhibited high expression levels in the cytoplasm of cancer cells,
with comparatively low expression levels in the nucleus.
Furthermore, in order to compare the expression level of RARα
protein in LSCC tissues and the adjacent noncancerous tissues, the
expression level was classified into two groups (low expression, -
to +; high expression, ++ to +++), which were presented in Table III. The rate of high expression in
LSCC tissues was 78.1%, and the rate of high expression in paired
adjacent paratumor tissues was 6.3%. The χ2 test
statistics demonstrated that the expression of RARα was
significantly higher in LSCC tissues compared with adjacent tissues
(P<0.05).
 | Table III.Expression of RARα in LSCC and paired
adjacent paratumor tissues. |
Table III.
Expression of RARα in LSCC and paired
adjacent paratumor tissues.
|
| RARα
expression |
|---|
|
|
|
|---|
| Tissues | Low | High | P-value |
|---|
| Paratumor
(N=32) | 30 | 2 | <0.0001 |
| LSCC (N=32) | 7 | 25 |
The clinical significance of RARα
overexpression in laryngeal carcinoma
The association between RARα and the
clinicopathological characteristics of LSCC was further analyzed.
There was no significant association between RARα expression and
age, sex, TNM stage or clinical stage (P>0.05). However, high
RARα expression was significantly associated with advanced
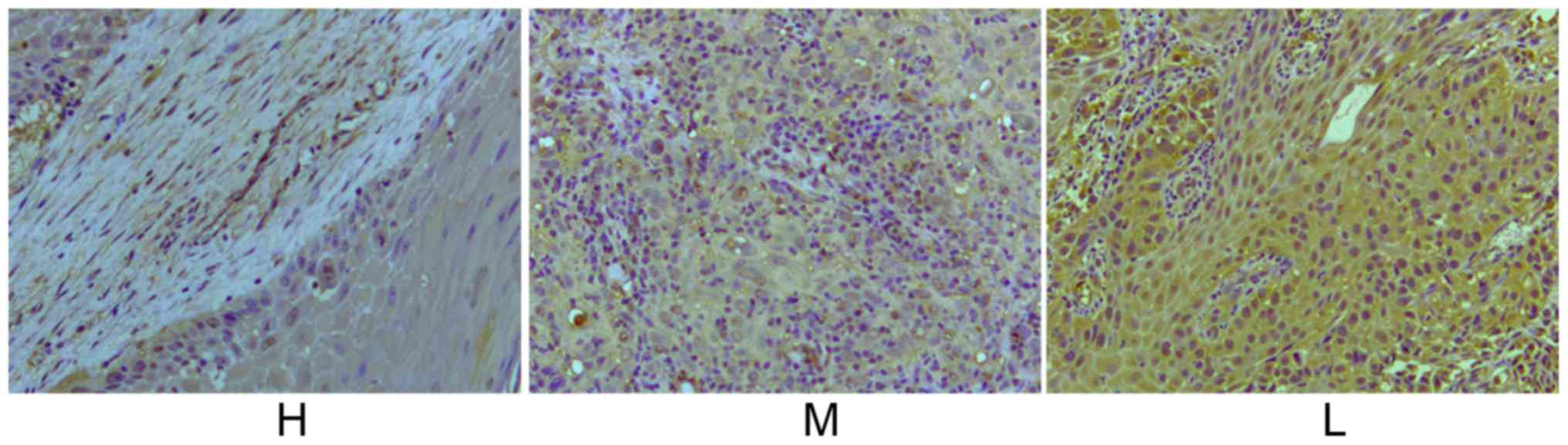
pathological differentiation (P<0.05). As presented in Fig. 2, the expression levels of RAR protein
in poorly differentiated LSCC tissues was visibly higher than that
in moderately differentiated and highly differentiated LSCC
tissues. The results of the present study indicate that RARα may be
involved in the regulation of the differentiation of LSCC
cells.
Downregulation of RARα inhibited
laryngeal carcinoma cell proliferation in vitro
To assess the effect of downregulating RARα on the
proliferation of LSCC cells, the expression of RARα was knocked
down using siRARα, and the control group siCtrl was transfected
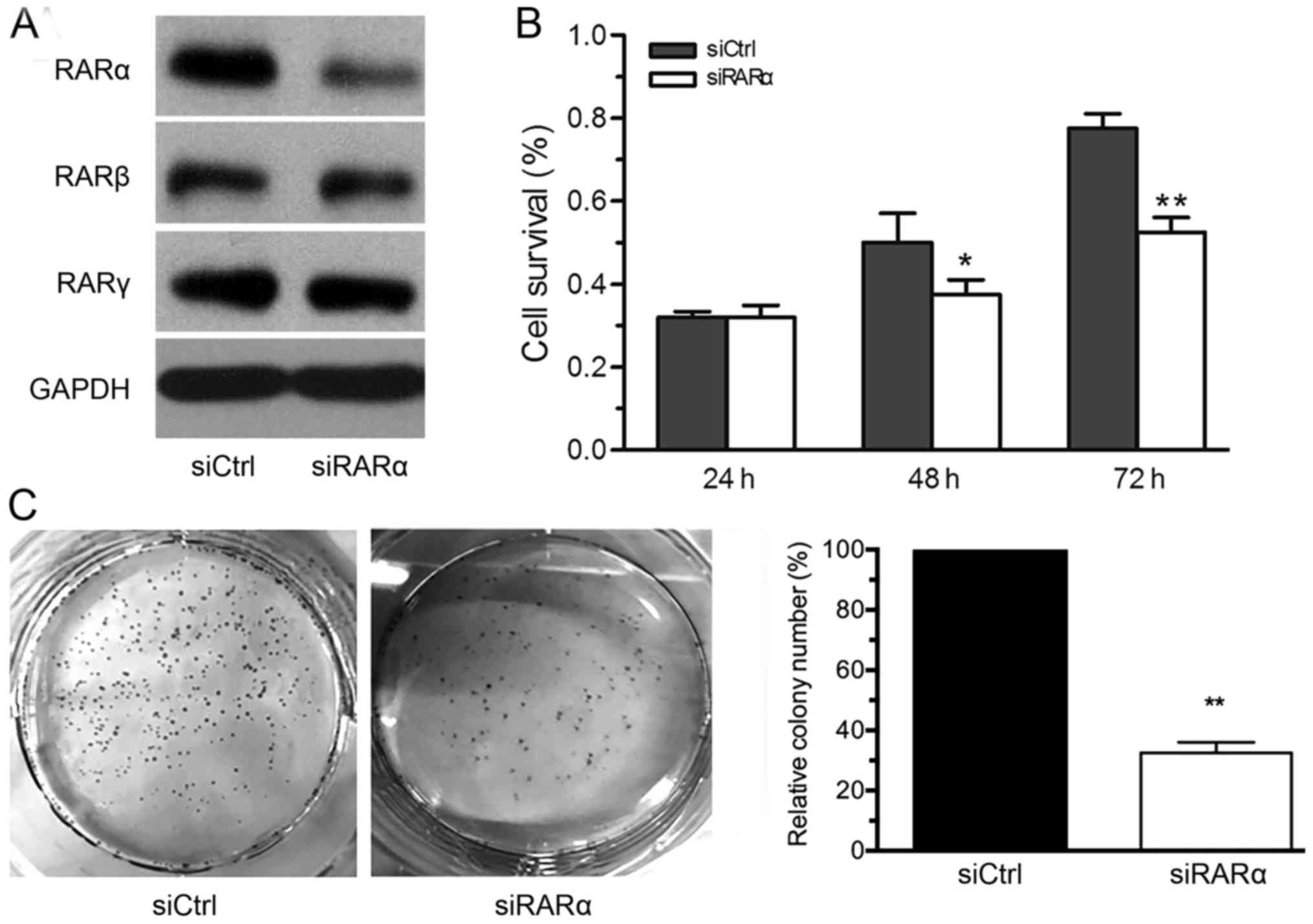
with non-specific fragments. The specificity of siRNA was confirmed
by the observation that the expression of RARα, but not RARβ or
RARγ, was downregulated using siRARα (Fig. 3A). The results from the MTT assay
revealed that the proliferation of siRARα-AMC-HN-8 cells were
significantly inhibited from 48 h following transfection, compared
with the siCtrl-AMC-HN-8 cells (Fig.
3B). Furthermore, the downregulation of RARα significantly
decreased the forming foci of AMC-HN-8 cells (Fig. 3C). This data indicated that the
downregulation of RARα inhibited the growth of LSCC cells.
Downregulation of RARα induced
laryngeal carcinoma cell cycle arrest via inhibition of the protein
kinase B (AKT) signaling pathway
The development of LSCC is associated with disorder
of the cell cycle (16). Therefore,
flow cytometry was performed to evaluate the effect of RARα
downregulation on the cell cycle distribution of LSCC cells. The
results demonstrated that the down-regulation of RARα in AMC-HN-8
cells reduced the proportion of cells in the S and G2/M phases, and
a higher number of cells were arrested in the G0/G1 phase compared
with cells in the control group (Fig.
4A). Furthermore, RT-qPCR was performed to detect the mRNA
expression of cell cycle associated genes. As presented in Fig. 4B, the expression levels of P21 mRNA in
the siRARα-AMC-HN-8 cells were >2× higher than those of the
siCtrl cells, but the mRNA expression of p27, cyclin A, cyclin B,
cyclin D, cyclin E was not significantly altered. Furthermore, the
results from the western blot analysis confirmed the upregulation
of the P21 protein in siRARα-AMC-HN-8 cells (Fig. 4C), which may be associated with the
inhibition of the AKT signaling pathway. Thus, this data indicated
that the downregulation of RARα resulted in P21 upregulation and
induced G1 phase arrest via inhibition of the AKT signaling
pathway.
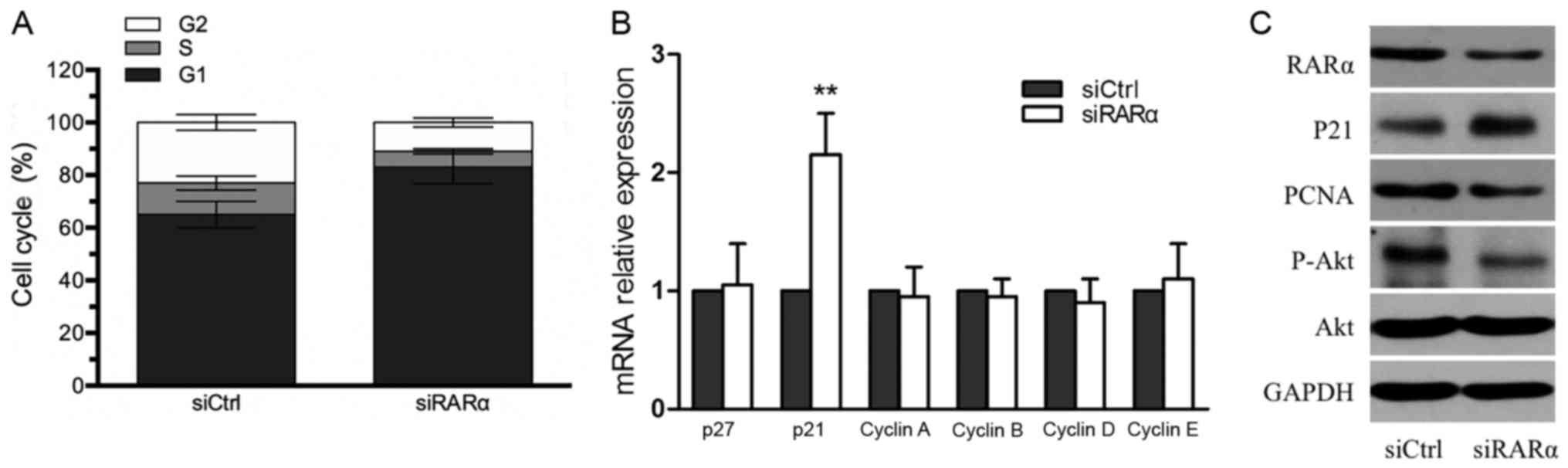 | Figure 4.Effect of RARα knockdown on the cell
cycle of AMC-HN-8 cells. (A) Flow cytometry analysis revealed that
RARα knockdown increased the proportion of cells in G0/G1 phase and
decreased the number in the G2 phase, compared with the control
group. (B) The mRNA expression of several cell cycle associated
genes were assessed using reverse transcription-quantitative
polymerase chain reaction, following transfection with siRARα or
siCtrl for 24 h. (C) The protein expression of RARα, p21, PCNA,
p-AKT and AKT in siCtrl and siRARα-AMC-HN-8 cells were detected
using western blotting analysis. **P<0.01 vs. control group.
RARα, retinoic acid receptor-α; siRARα, RARα-specific small
interfering RNA; siCtrl, small interfering RNA negative control;
p21, cyclin dependent kinase inhibitor 1A; PCNA, proliferating cell
nuclear antigen; AKT, protein kinase B; p-, phosphorylated. |
Discussion
Laryngeal carcinoma is one of the most aggressive
malignant tumors out of all head and neck squamous cell carcinoma,
and generally has a poor prognosis (2). The survival time of laryngeal cancer has
not increased, despite substantial progress in the developments of
surgery and radiotherapy (3,17). Revealing the molecular mechanism
underlying the development of LSCC is important for early diagnosis
and targeted therapy. The development of LSCC is a complex process
which is associated with changeable expression of multiple genes
(18). Here, it was revealed that
RARα was elevated in the LSCC tissues and was associated with poor
pathological differentiation. Knockdown of RARα significantly
inhibited the proliferation of LSCC cells through the induction of
cell cycle arrest.
RARα, regarded as an important member of the
retinoic acid receptor family, is involved in the regulation of
cell differentiation, proliferation, apoptosis and metabolism.
Accumulating evidence has indicated that RARα serves an important
function in the development of malignant tumors. Notably, the
functions of RARα in the development of tumors is dependent on the
origin and the microenvironment of tumor cells. RARα served as a
tumor suppressor gene in the development of gastric cancer,
non-small cell lung cancer, melanoma tumor and prostate cancer.
However, RARα may also serve as an oncogene in the development of
hepatocellular carcinoma, leukemia and breast cancer. In the
present study, it was identified that the expression of RARα in
LSCC tissues was significantly higher than that in adjacent
noncancerous tissues, suggesting that RARα may serve as an
oncogene, participating in the process of the development of LSCC.
Further analysis revealed that overexpression of RARα was
associated with poor pathologic differentiation. Despite the small
sample size of the present study, it was indicated that RARα may be
involved in the regulation of LSCC cell differentiation.
Rapid proliferation, resistance to apoptosis,
indefinite reproduction, angiogenesis, invasion and metastasis are
the main features in the development of a tumor (19). These are also the biological functions
of various types of tumor oncogenes and suppressor genes involved
in the development of malignant tumors. In the present study, the
downregulation of RARα significantly inhibited the proliferation of
AMC-HN-8 cells through inducing cell cycle arrest. It was indicated
that the overexpression of RARα served an important function in the
development of LSCC. Consistent with the present study, previous
studies have reported that RARα was also involved in the regulation
of the cell cycle in breast cancer (13). Retinoic acid, a ligand of RARα,
inhibited the growth of breast cancer cells through the induction
of cell cycle arrest at the G1 phase and cell apoptosis (13). The mechanism of RARα in the regulation
of the development of LSCC is the focus of a number of studies; it
has been reported that there are genomic and non-genomic influences
(20). During genomic regulation, the
conformation of RARα changes and recruits a coactivator protein
once it is combined with its ligand. The complex forms the retinoic
acid response element in the promoter region of the target gene,
and then activates the transcription of target genes, resulting in
the promotion of cell differentiation, apoptosis and other
biological processes (20). In
previous years, there was a greater focus on the non-genomic
regulation of RARα. Previous studies have identified that RARα may
regulate the phosphoinositide 3-kinase, AKT, c-Jun N-terminal
kinase, mitogen-activated protein kinase 14 and protein kinase C
signaling pathways following abnormal translocation into the
cytoplasm (10,21–23). The
present study identified that the overexpression of RARα was mainly
located in the cytoplasm of LSCC cells and its downregulation
significantly inhibited the activation of the AKT signaling
pathway. Therefore, it was speculated that RARα promoted the
development of LSCC through non-genomic transcriptional
regulation.
Altogether, the present study has demonstrated that
RARα expression was significantly elevated in patients with LSCC,
and the downregulation of RARα induced cell cycle arrest and
inhibited the proliferation of LSCC cells via the inhibition of the
AKT signaling pathway. The present study may offer a potential
molecular basis for the further development of RARα as a novel
therapeutic target for patients with LSCC.
Acknowledgements
The present study was supported by the Youth
Foundation of the Fujian Health Department, Fujian, China (grant
no. 2013-2-83), the Project of Young and Middle-Aged Backbone
Talent Cultivation, Fujian, China (grant no. 2013-ZQN-JC-32) and
the Science and Technology Bureau of Xiamen, China (grant no.
3502Z20144004). The present study was also supported by the
National Nature Science Foundation of China (grant no.
81572394).
References
|
1
|
Jemal A, Bray F, Center MM, Ferlay J, Ward
E and Forman D: Global cancer statistics. CA Cancer J Clin.
61:69–90. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Marur S and Forastiere AA: Head and neck
squamous cell carcinoma: Update on epidemiology, diagnosis, and
treatment. Mayo Clin Proc. 91:386–396. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Belcher R, Hayes K, Fedewa S and Chen AY:
Current treatment of head and neck squamous cell cancer. J Surg
Oncol. 110:551–574. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
di Masi A, Leboffe L, De Marinis E, Pagano
F, Cicconi L, Rochette-Egly C, Lo-Coco F, Ascenzi P and Nervi C:
Retinoic acid receptors: From molecular mechanisms to cancer
therapy. Mol Aspects Med. 41:1–115. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Altucci L, Leibowitz MD, Ogilvie KM, de
Lera AR and Gronemeyer H: RAR and RXR modulation in cancer and
metabolic disease. Nat Rev Drug Discov. 6:793–810. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Shiohira H, Kitaoka A, Shirasawa H, Enjoji
M and Nakashima M: Am80 induces neuronal differentiation in a human
neuroblastoma NH-12 cell line. Int J Mol Med. 26:393–399.
2010.PubMed/NCBI
|
|
7
|
Casey NP and Woods GM: Anti-PML-RARα shRNA
sensitises promyelocytic leukaemia cells to all-trans retinoic
acid. J RNAi Gene Silencing. 8:464–469. 2012.PubMed/NCBI
|
|
8
|
Singh AT, Evens AM, Anderson RJ, Beckstead
JA, Sankar N, Sassano A, Bhalla S, Yang S, Platanias LC, Forte TM,
et al: All trans retinoic acid nanodisks enhance retinoic acid
receptor mediated apoptosis and cell cycle arrest in mantle cell
lymphoma. Br J Haematol. 150:158–169. 2010.PubMed/NCBI
|
|
9
|
Wu Q, Lin XF, Ye XF, Zhang B, Xie Z and Su
WJ: Ubiquitinated or sumoylated retinoic acid receptor alpha
determines its characteristic and interacting model with retinoid X
receptor alpha in gastric and breast cancer cells. J Mol
Endocrinol. 32:595–613. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Hoshikawa Y, Kanki K, Ashla AA, Arakaki Y,
Azumi J, Yasui T, Tezuka Y, Matsumi Y, Tsuchiya H, Kurimasa A, et
al: c-Jun N-terminal kinase activation by oxidative stress
suppresses retinoid signaling through proteasomal degradation of
retinoic acid receptor α protein in hepatic cells. Cancer Sci.
102:934–941. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Sano K, Takayama T, Murakami K, Saiki I
and Makuuchi M: Overexpression of retinoic acid receptor alpha in
hepatocellular carcinoma. Clin Cancer Res. 9:3679–3683.
2003.PubMed/NCBI
|
|
12
|
Schneider SM, Offterdinger M, Huber H and
Grunt TW: Activation of retinoic acid receptor alpha is sufficient
for full induction of retinoid responses in SK-BR-3 and T47D human
breast cancer cells. Cancer Res. 60:5479–5487. 2000.PubMed/NCBI
|
|
13
|
Lu M, Mira-y-Lopez R, Nakajo S, Nakaya K
and Jing Y: Expression of estrogen receptor alpha, retinoic acid
receptor alpha and cellular retinoic acid binding protein II genes
is coordinately regulated in human breast cancer cells. Oncogene.
24:4362–4369. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Gyftopoulos K, Perimenis P,
Sotiropoulou-Bonikou G, Sakellaropoulos G, Varakis I and Barbalias
GA: Immuno-histochemical detection of retinoic acid receptor-alpha
in prostate carcinoma: Correlation with proliferative activity and
tumor grade. Int Urol Nephrol. 32:263–269. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Pignataro L, Sambataro G, Pagani D and
Pruneri G: Clinico-prognostic value of D-type cyclins and p27 in
laryngeal cancer patients: A review. Acta Otorhinolaryngol Ital.
25:75–85. 2005.PubMed/NCBI
|
|
17
|
Li P, Hu W, Zhu Y and Liu J: Treatment and
predictive factors in patients with recurrent laryngeal carcinoma:
A retrospective study. Oncol Lett. 10:3145–3152. 2015.PubMed/NCBI
|
|
18
|
Ni RS, Shen X, Qian X, Yu C, Wu H and Gao
X: Detection of differentially expressed genes and association with
clinicopathological features in laryngeal squamous cell carcinoma.
Oncol Lett. 4:1354–1360. 2012.PubMed/NCBI
|
|
19
|
Hanahan D and Weinberg RA: Hallmarks of
cancer: The next generation. Cell. 144:646–674. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Hart SM: Modulation of nuclear receptor
dependent transcription. Biol Res. 35:295–303. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Srinivas H, Xia D, Moore NL, Uray IP, Kim
H, Ma L, Weigel NL, Brown PH and Kurie JM: Akt phosphorylates and
suppresses the transactivation of retinoic acid receptor alpha.
Biochem J. 395:653–662. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Boskovic G, Desai D and Niles RM:
Regulation of retinoic acid receptor alpha by protein kinase C in
B16 mouse melanoma cells. J Biol Chem. 277:26113–26119. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Piskunov A and Rochette-Egly C: A retinoic
acid receptor RARα pool present in membrane lipid rafts forms
complexes with G protein αQ to activate p38MAPK. Oncogene.
31:3333–3345. 2012. View Article : Google Scholar : PubMed/NCBI
|