Introduction
In 2012, there were an estimated 951,600 new gastric
cancer cases and 723,100 gastric cancer-associated mortalities
worldwide (1). Gastric cancer rates
are typically twice as high in males as in females, and China has
one of highest incidence rates (1).
Cancer is a complex disease that results from interactions between
multiple genetic and environmental factors (2,3). Even
between individuals with similar environmental risk factors,
including Helicobacter pylori infection (4,5), smoking
(6) and obesity (7), not all individuals will develop gastric
carcinoma (GC). This suggests that genetic factors may serve an
important function in the development and progression of GC
(8).
Galectin-3, a 31 kDa oncogenic protein, is a member
of the β-galactoside-binding gene family that regulates cell
growth, proliferation, apoptosis, adhesion, metastasis and
angiogenesis (9,10). In GC, galectin-3 expression has been
identified to be associated with metastasis; galectin-3 expression
was significantly increased in metastatic lymph nodes, compared
with that in the primary gastric tumor; therefore, it may be used
as a biomarker for peritoneal metastasis and as a prognostic factor
for gastric cancer (11–14). The silencing of galectin-3 alters the
expression of a range of cancer-associated genes, including cyclin
D1 and survivin, and increases the sensitivity of gastric cancer
cells to chemotherapeutic agents (15). Human breast carcinoma cells that
overexpress galectin-3 exhibit increased resistance to apoptosis
induced by the DNA-cross-linking agent cisplatin, the anthracycline
adriamycin, the anti-metabolite fluorouracil (5-FU) or the
topoisomerase II inhibitor etoposide (9).
Rs4644 and rs4652 are the most common single
nucleotide polymorphism (SNP) sites in galectin-3, and are
associated with galectin-3 protein expression levels (16). Rs4644 and rs4652 are located in exon 3
of galectin-3. Rs4644 C>A at position 191 results in
substituting a proline for histidine (17). When rs4652 occurs at position 292,
there is a threonine to proline substitution (18). Rs4644 C>A overexpression increases
the rate of gastric cancer cell growth and resistance to cisplatin
and 5-FU (17). The rs4644 variant of
galectin-3 may exert a protective effect in prostate cancer cells
(19). Furthermore, rs4644 is
associated with the incidence of breast cancer, resulting in
susceptibility to matrix metalloproteinase cleavage and the
acquisition of resistance to drug-induced apoptosis (20). Rs4652 has also been identified to
serve a function in glioma (21).
Rs4644 and rs4652 regulate lectin galactoside-binding soluble 3,
affecting immunoglobulin binding.
Studies have demonstrated the biological and drug
response alterations associated with galectin-3 SNPs in cancer
(22); however, only a limited number
of studies have investigated the function of galectin-3 rs4644 and
rs4652 polymorphisms in the susceptibility and chemotherapeutic
drug resistance of GC. Galectin-3 functional genetic variations may
contribute to the development and chemotherapy response of GC. The
objective of the present study was to evaluate the association
between galectin-3 rs4644 and rs4652 genotypes with the risk of
developing GC and its chemotherapy drug resistance in a
hospital-based case-control study.
Materials and methods
Patients and controls
The study population consisted of 479 patients with
primary GC undergoing total gastrectomy and 458 cancer-free
individuals. The 479 patients with GC were recruited from Fujian
Cancer Hospital (Fujian, China) between January 2007 and December
2009. All diagnoses were histopathologically confirmed as primary
gastric cancer. The exclusion criteria were as follows: i) Patients
who had previously been diagnosed with cancer; ii) any metastasis;
and iii) patients who had received radiotherapy, chemotherapy or
immunotherapy prior to surgery. The 458 cancer-free controls were
randomly selected from a pool of healthy volunteers following a
routine medical check during the same period at the Fujian cancer
hospital; all volunteers had no history of personal or familial
malignancy or other serious diseases. All participants provided
written informed consent to participate in the study and the study
was approved by the Ethics Committee of Fujian Cancer Hospital.
DNA extraction and genotyping
analyses
The DNA of the patients with GC was isolated from
paraffin-embedded normal stomach tissue adjacent to the tumor site
(distance, >5 cm) using a QIAamp DNA FFPE Tissue kit (Qiagen
GmbH, Hilden, Germany). The DNA of cancer-free individuals was
extracted from venous blood samples (2 ml) using a Whole Blood
Genome DNA Isolation kit (BioTeke Corporation, Beijing, China). The
concentration of DNA was determined using a NanoDrop-1000
ultraviolet spectrophotometer, with a A260/A280 ratio between 1.8
and 2.0.
Genotyping analyses for SNPs were performed using
Matrix-Assisted Laser Desorption/Ionization Time of Flight Mass
Spectrometry (MALDI-TOF MS) as previously described (23). Briefly, polymerase chain reaction
(PCR) was conducted to amplify the target sequence using
HotStarTaq DNA Polymerase (Qiagen GmbH, Hilden, Germany).
Primer sequences used for rs4644 detection were as follows: 1st-PCR
primer, ACG TTG GAT GTC CGG GAT AAG CTC CAG GT; 2nd-PCR primer, ACG
TTG GAT GGC TTA TCC TGG ACA GGC AC; UEP_SEQ reverse primer, CTC CAG
GCG CCT ACC. For rs4652 detection, 1st-PCR primer, ACG TTG GAT GAA
GGA ATG CCA TCT CAC CAG; 2nd-PCR primer, ACG TTG GAT GCT ACC CAT
CTT CTG GAC AGC; UEP_SEQ reverse primer, GGA CAG CCA AGT GCC.
The excess dNTPs were removed and base extension
reactions were conducted using ThermoSequenase (GE Healthcare,
Chicago, IL, USA). Extended products were treated with SpectroCLEAN
resin (Sequenom, San Diego, CA, USA) to remove salts. Subsequently,
10 µl of the reaction was dispensed into a SpectroCHIP microarray
(Sequenom). A MALDI-TOF mass spectrometer was used to acquire the
SNP profile data, which was analyzed by TYPER 4.0 software
(Sequenom).
Quality control
A total of 3 control types were included: Negative
control, internal standard control and repeat control. The number
of negative controls and internal standard controls was 1% of the
total sample number, whereas the number of repeat controls was 5%
of the total sample number. For the negative control, sample DNA
was replaced with water to monitor the contamination of product in
the experiment. ‘Chinese number one’ cell line DNA was used as the
internal standard control template to detect the primer design
reliability of each SNP site. The ‘Chinese number one’ cell line is
cultivated by Beijing Genomics Institute (Shenzhen, China) and the
SNP site data can be accessed from the ‘Chinese number one’
database (24). For the repeat
control, randomly selected samples were used as repeat controls to
validate the stability of the experiment.
Immunohistochemical analysis
Immunohistochemical analyses were performed for 207
samples using the BenchMark XT platform (Ventana Medical Systems,
Tucson, AZ, USA). Primary antibodies included P53 (Kit-0010-2,
clone: DO-7), Ki67 (Kit-0005-2, clone: MIB-1), glutathione
S-transferase (GST; MAB-0583, clone: LW29), P-glycoprotein (Pgp;
MAB-0237, clone: C494), topoisomerase II (TOPII; MAB-0588, clone:
3F6) and thymidylate synthase (TS; MAB-0171, clone: TS106), (all
ready-to-use dilution; Fuzhou Maixin Biotech, Fuzhou, Fujian,
China). Each slide was subjected to heat treatment in CC1 retrieval
solution (Ventana Medical Systems, Tucson, AZ, USA) for 30 min and
subsequently incubated with all six antibodies at 37°C for 30 min.
Immune complexes were detected with the use of an UltraView
3′3-diaminobenzidine detection kit (Ventana Medical Systems,
Tucson, AZ, USA) according to the manufacturer's protocol.
The sections were scored semi-quantitatively, using
light microscopy, by two pathologists without any knowledge of the
clinical history of the patients. The intensity of staining was
evaluated according to the following scale: 0, no staining; 1, weak
staining; 2, moderate staining; and 3, strong staining. The extent
of staining was scored as follows: 0, <10%; 1, ≥10% and <25%;
2, ≥25% and <50%; 3, ≥50% and <75%; 4, ≥75%. The final
results were the product of the intensity and extent of staining.
The grading system was divided into four parts according to the
product of the intensity and the extent of staining: 0–3, negative;
4–6, +; 7–9, 2+; and 10–12, 3+.
Statistical analysis
Univariate analysis was performed with a
χ2 test. The computation of odds ratio (OR) and 95%
confidence intervals (95% CI) were conducted using SPSS software
(version 17.0; SPSS, Inc., Chicago, IL, USA). The tests included an
evaluation of the association between genotypic distributions and
the development of GC, clinicopathological features (including
tumor size, depth of invasion, differentiation and the status of
the lymph node metastasis) and the protein expression levels from
the immunohistochemistry data. Haplotypes were estimated using the
Expectation Maximization algorithm (25). Haplotype association with GC was
determined using SHEsis, as previously described (26).
Results
Characteristics of the study
population
The fundamental background data for patient cases
and patient controls are summarized in Table I, and demonstrate they were adequately
matched for age and sex. In the control patients, the frequency of
the genotypes of the two polymorphisms were determined using the
Hardy-Weinberg test, which revealed that the P-values for rs4644
and rs4652 were 0.81 and 0.50, respectively. These results
demonstrated that the genotype distribution conforms to a
Hardy-Weinberg equilibrium.
 | Table I.Baseline data for patients with
gastric carcinoma and healthy controls. |
Table I.
Baseline data for patients with
gastric carcinoma and healthy controls.
| Characteristic | Cases, n (%) | Controls, n (%) | χ2
value | P-value |
|---|
| Sex |
|
|
|
|
| Male | 347 (72.44) | 335 (73.14) | 0.33 | 0.56 |
|
Female | 132 (27.56) | 117 (25.55) |
|
|
| Age, years |
|
|
|
|
| ≤60 | 156 (32.57) | 147 (32.10) | 0.17 | 0.68 |
|
>60 | 323 (67.43) | 287 (62.66%) |
|
|
| Lauren
classification |
|
|
|
|
|
Intestinal type | 175 (36.53) |
|
|
|
| Diffuse
type | 304 (63.47) |
|
|
|
| Tumor-node-metastasis
stage |
|
|
|
|
| 0-I | 60 (12.53) |
|
|
|
| II | 100 (20.88) |
|
|
|
|
III | 298 (62.21) |
|
|
|
| IV | 21 (4.38) |
|
|
|
| Total, n | 479 | 458 |
|
|
Quality control results
The present study was based on a large number of
clinical samples. A total of 10 negative controls were used and
analyzed to validate that the results were not false positives. The
data of the 10 internal standard controls were all matched in the
‘Chinese number one’ database. In addition, the conformity of 47
repeat controls was >98%. Therefore, the results of the
genotyping were deemed reliable.
Patients with rs4652 CA/AA are more
likely to develop GC
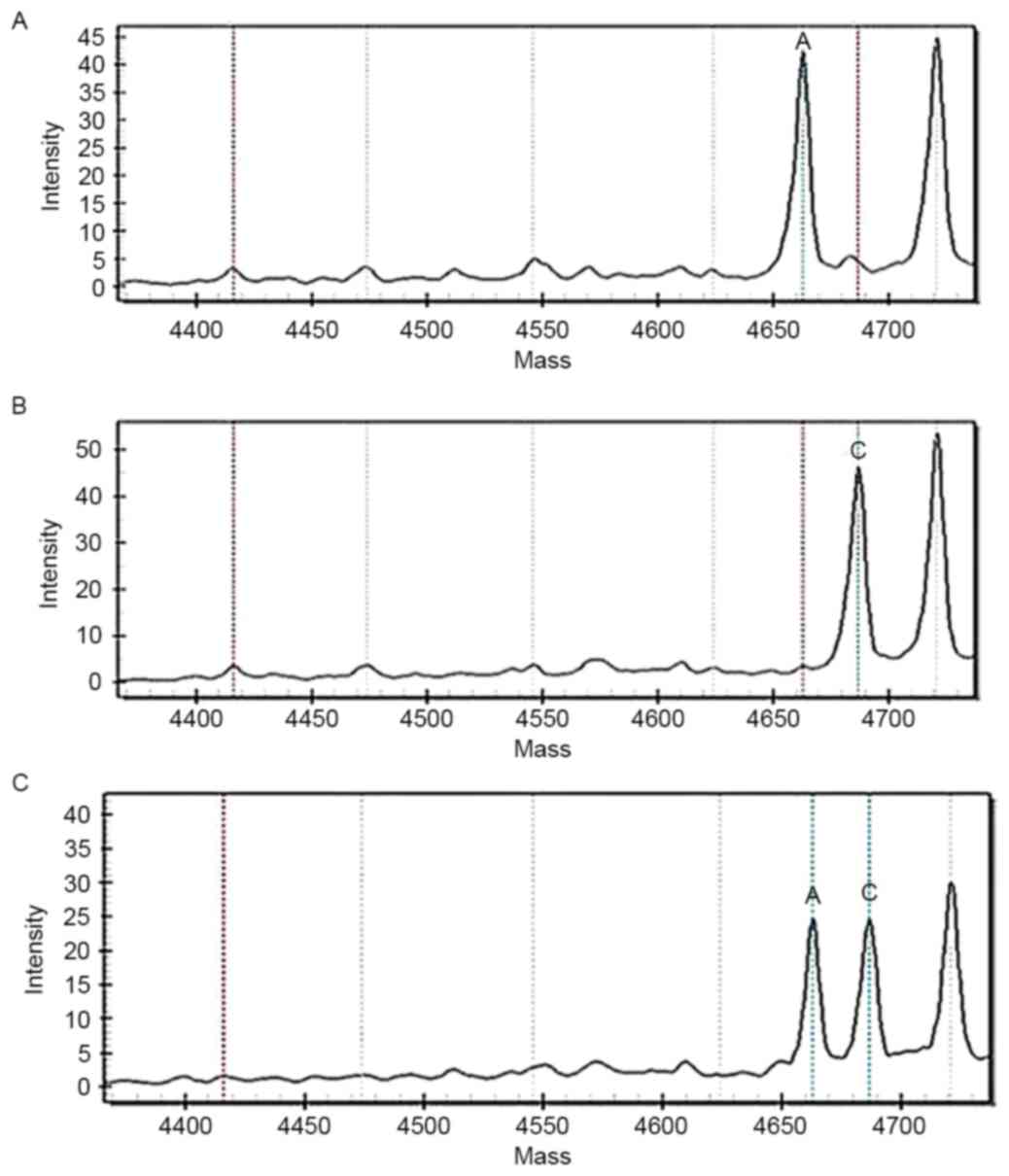
The genotyping results of 479 patients and 458
controls are presented in Table II,
and the MALDI-TOF MS spectra are presented in Fig. 1. Compared with the controls, the
patients' rs4652 allelic genotype and its genotypic distribution
frequency differed significantly; the rs4652 CA/AA carriers
exhibited a significant increased risk of GC (adjusted-OR, 1.51;
95% CI, 1.05–2.18; adjusted-P=0.03), which indicated that the
patients with rs4652 CA/AA were more likely to develop GC. There
was no significant difference identified in the genotypic
distribution of rs4644 between patients and controls, indicating
that this polymorphism was not associated with the development of
GC.
 | Table II.Association between galectin-3
polymorphism with the occurrence of gastric carcinoma. |
Table II.
Association between galectin-3
polymorphism with the occurrence of gastric carcinoma.
| Genotype | Cases, n (%) | Controls, n
(%) | χ2
value |
P-valuea |
|---|
| Total | 479 | 458 |
|
|
| rs4644 C>A |
|
|
| 0.26 |
| AA | 11 (2.33) | 16 (3.59) |
|
|
|
AC/CC | 462 (97.67) | 430 (96.41) | 1.28 |
|
| rs4652 A>C |
|
|
| 0.03a |
| CC | 58 (12.24) | 79 (17.40) |
|
|
|
CA/AA | 416 (87.76) | 375 (82.60) | 4.92 |
|
Galectin-3 polymorphisms are not
associated with the clinicopathological features of GC
The present study aimed at identifying associations
between the genotypic distributions of galectin-3 SNPs and the
overall clinicopathological features of the disease, including
tumor size, depth of invasion, differentiation and the status of
lymph node metastasis. No associations were determined between
rs4644 and rs4652 genotypic distribution with the overall
clinicopathological features of patients with GC.
Rs4652 genotypic distributions are
associated with Pgp expression
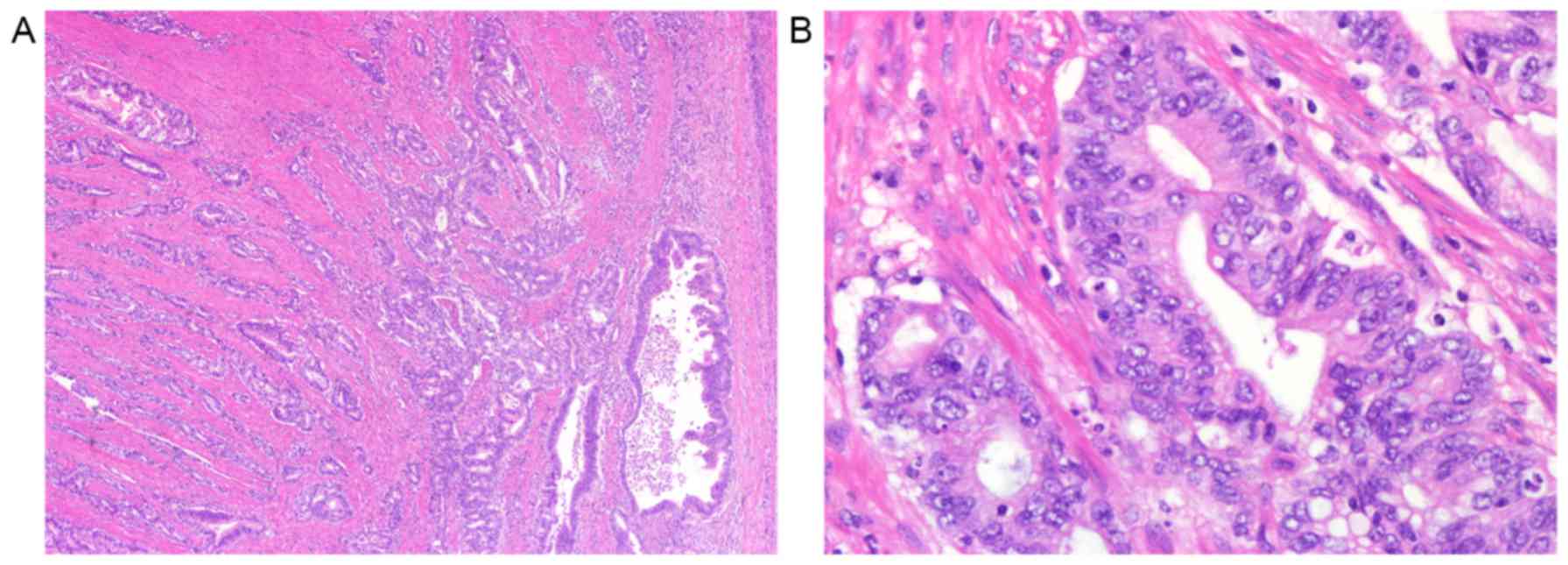
Samples from 207 patients with GC were analyzed by
immunohistochemistry. Representative hematoxylin and eosin staining
images of GC is presented in Fig. 2.
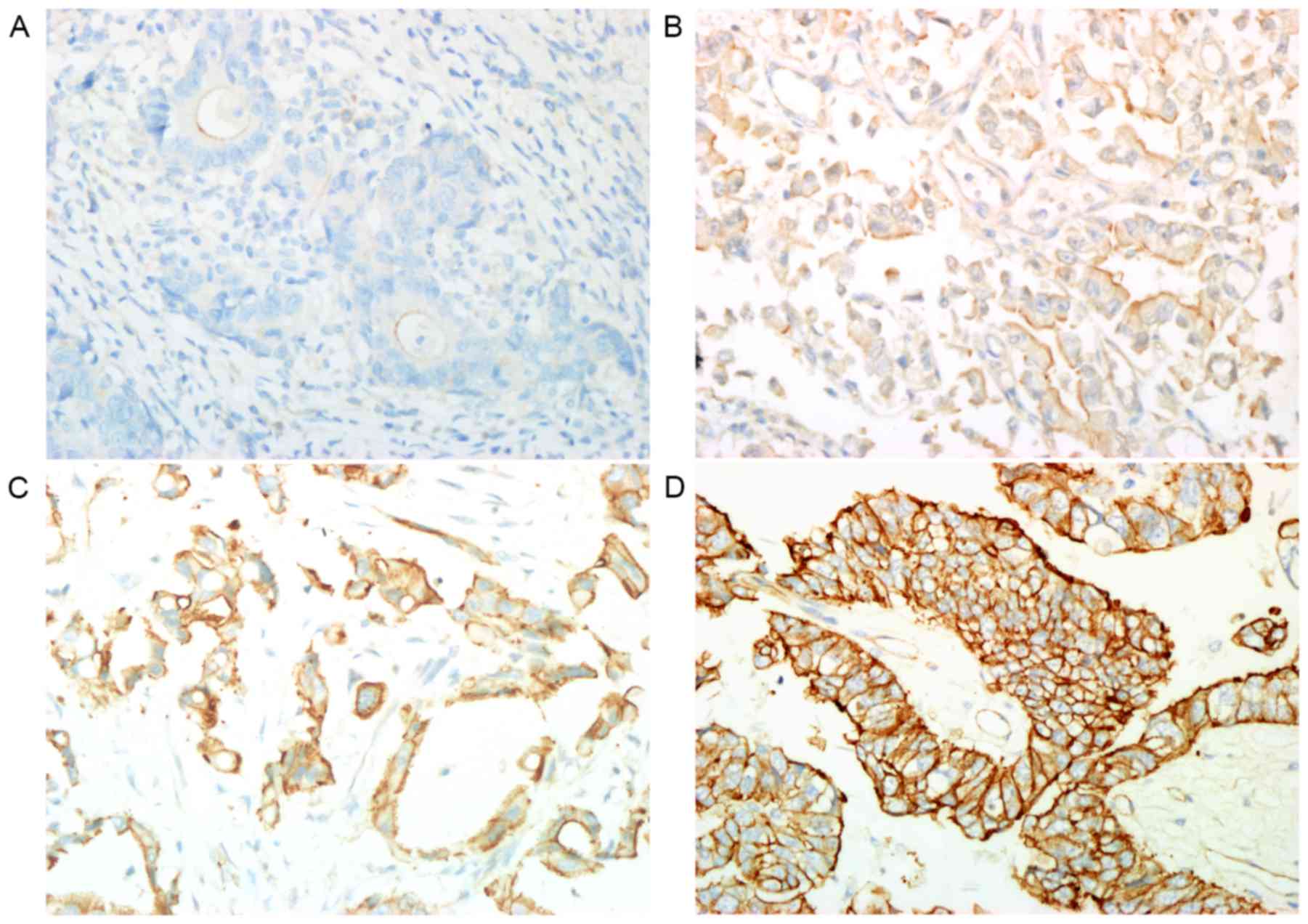
The Pgp protein expression in the tumor tissues was predominantly
homogenous and was primarily identified in the cytoplasm or at the
cell membrane. With the exclusion of 4 patients who failed SNPs
genotyping analyses due to low sample quality, the Pgp protein
expression in the remaining 203 patients was as follows: Negative,
41.9% (85/203); +, 15.8% (32/203); 2+, 35.0% (71/203); 3+, 7.4%
(15/203; Fig. 3). The association
between the rs4652 genotypic distribution and Pgp protein
expression is presented in Table
III. The results indicated that there were significant
associations between rs4652 genotypic distributions and Pgp
expression level (χ2=9.063; P=0.028), although the
association was not linear. There was no significant association
identified between the genotypic distribution of galectin-3 SNPs
and other proteins including P53, Ki67, GST, Pgp, TOPII and TS.
 | Table III.Association between rs4652 genotypic
distributions and P-glycoprotein expression. |
Table III.
Association between rs4652 genotypic
distributions and P-glycoprotein expression.
|
| Genotypic
distribution |
|
|---|
|
|
|
|
|---|
| Intensity | CC | CA | AA | CA/AA |
P-valuea |
|---|
| Negative | 61 | 25 | 0 | 25 | 0.028a |
| + | 23 | 7 | 1 | 8 |
|
| 2+ | 49 | 22 | 0 | 22 |
|
| 3+ | 12 | 3 | 0 | 3 |
|
Galectin-3 haplotype is not associated
with gastric cancer risk
The haplotype of galectin-3 rs4644 and rs4652 were
analyzed using SHEsis software and three haplotypes were identified
in patients and controls. No association was revealed between the
galectin-3 haplotype and the risk of gastric cancer.
Discussion
Previous studies have demonstrated that the
expression level of galectin-3 is associated with the development
and progression of gastric cancer (11–14).
Galectin-3 SNPs are associated with galectin-3 expression levels
(16). In addition, an association
has been identified between rs4644 and the incidence of prostate
and breast cancer (19,20). In the present study, no association
was identified between rs4644 C>A and the susceptibility or
chemotherapeutic drug resistance to GC. However, the results of the
present study indicated that individuals with an rs4652 A allele
(CA or AA genotype) exhibited a 1.5-fold increased risk of
developing gastric cancer compared with individuals carrying the CC
genotype.
Galectin-3 is a galactose-binding protein that
belongs to the lectin family. It has three binding domains, as
follows: A NH2 ligand binding domain (for target cells);
a carbohydrate recognition domain and a glycine, proline, tyrosine
repetitive sequence binding domain (for matrix metalloproteinase)
(27). Rs4652 is located in the
carbohydrate recognition domain, which is the most critical domain
of galectin-3 as it activates fibroblasts via the transforming
growth factor-β pathway by ligating to fibroblast surface
receptors. Activated fibroblasts (myofibroblasts) interact with
cancer cells to promote angiogenesis, required for sustained cancer
cell growth (28,29). Therefore, we hypothesized that the
rs4652 A>C alteration to the structure of the carbohydrate
recognition domain strengthens its interaction with fibroblast
surface receptors, resulting in the myofibroblast promotion of
angiogenesis and the growth of gastric cancer cells. This
hypothesis requires further study and validation.
The other principal result of the present study was
that rs4652 genotypic distribution was significantly associated
with the Pgp protein expression level, indicating that patients
with GC with different genotypes may exhibit a distinct reaction to
chemotherapeutic drugs. The resistance of gastric cancer cells to
chemotherapeutics following surgery remains a challenge in the
treatment of patients with gastric cancer. The expression of Pgp,
TOPII, GST and TS are recognized as classic resistance indexes, and
are associated with a poor prognosis (30). Pgp is a transmembrane protein encoded
by the gene, MDR1. Pgp consists of 1,280 amino acids, with a
molecular mass of 170 kDa, 12 transmembrane domains and 2 ATP
binding domains. The Pgp protein binds chemotherapeutic molecules
and pumps the drug out of the cell in an ATP-dependent reaction,
resulting in a decreased intracellular concentration of the drug
and facilitating drug resistance (31). Therefore, the increased expression of
the Pgp protein can be used as a marker for multi-drug resistance.
The level of Pgp protein expression has an effect on the
distribution of a range of natural hydrophobic drugs, including
doxorubicin, pharmorubicin, taxol and docetaxel (32). Therefore, drugs should be carefully
selected for patients according to Pgp expression level. Although
Pgp expression level was associated with rs4652 genotypic
distribution in the present study, no linear relationship was
identified; therefore, it cannot be concluded that a particular
genotype is associated with increased Pgp protein expression or
chemotherapeutic resistance.
The present study included certain limitations.
First, data on environmental factors, including Helicobacter
pylori or Epstein-Barr virus infection, were not available;
therefore, it was not possible to explore gene-environment
interactions. In addition, there was a lack of follow-up data;
analyses to determine the function of galectin-3 polymorphisms in
GC prognosis could not be performed.
The results of the present study suggested that
rs4652 SNPs of the galectin-3 gene contribute to the susceptibility
to and chemotherapeutic drug resistance of GC. Thus, galectin-3
rs4652 SNPs may be a biomarker for predicting the development and
prognosis of patients with GC.
Acknowledgements
The present study was supported by the National
Natural Science Foundation (grant no. 81470111), the Natural
Science Foundation of Fujian Province (grant no. 2014J01300) and
the National Clinical Key Specialty Construction Program of
China.
Glossary
Abbreviations
Abbreviations:
|
GC
|
gastric carcinoma
|
|
SNP
|
single nucleotide polymorphisms
|
|
MALDI-TOF MS
|
matrix-assisted laser
desorption/ionization time-of-flight mass spectrometry
|
|
PCR
|
polymerase chain reaction
|
|
Pgp
|
P-glycoprotein
|
|
GST
|
glutathione S-transferase
|
|
TOPII
|
topoisomerase II
|
|
TS
|
thymidylate synthase
|
References
|
1
|
Torre LA, Bray F, Siegel RL, Ferlay J,
Lortet-Tieulent J and Jemal A: Global cancer statistics, 2012. CA
Cancer J Clin. 65:87–108. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Liu L, Zhong R, Wei S, Yin JY, Xiang H,
Zou L, Chen W, Chen JG, Zheng XW, Huang LJ, et al: Interactions
between genetic variants in the adiponectin, adiponectin receptor 1
and environmental factors on the risk of colorectal cancer. PLoS
One. 6:e273012011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Reeves GK, Pirie K, Green J, Bull D and
Beral V: Million Women Study Collaborators: Comparison of the
effects of genetic and environmental risk factors on in situ and
invasive ductal breast cancer. Int J Cancer. 131:930–937. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Parkin DM: The global health burden of
infection-associated cancers in the year 2002. Int J Cancer.
118:3030–3044. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Plummer M, Franceschi S, Vignat J, Forman
D and de Martel C: Global burden of gastric cancer attributable to
Helicobacter pylori. Int J Cancer. 136:487–490. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Bertuccio P, Chatenoud L, Levi F, Praud D,
Ferlay J, Negri E, Malvezzi M and La Vecchia C: Recent patterns in
gastric cancer: A global overview. Int J Cancer. 125:666–673. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Jemal A, Center MM, DeSantis C and Ward
EM: Global patterns of cancer incidence and mortality rates and
trends. Cancer Epidemiol Biomarkers Prev. 19:1893–1907. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Lin XD, Chen SQ, Qi YL, Zhu JW, Tang Y and
Lin JY: Polymorphism of THBS1 rs1478604 A>G in 5-untranslated
region is associated with lymph node metastasis of gastric cancer
in a Southeast Chinese population. DNA Cell Biol. 31:511–519. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Fukumori T, Kanayama HO and Raz A: The
role of galectin-3 in cancer drug resistance. Drug Resist Updat.
10:101–108. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Nakahara S, Oka N and Raz A: On the role
of galectin-3 in cancer apoptosis. Apoptosis. 10:267–275. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Jeong GA, Kim HC, Kim HK and Cho GS:
Perigastric lymph node metastasis from papillary thyroid carcinoma
in a patient with early gastric cancer: The first case report. J
Gastric Cancer. 14:215–219. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Lee KB, Park DJ, Choe G, Kim HH, Kim WH
and Lee HS: Protein expression status in mucosal and submucosal
portions of early gastric cancers and their predictive value for
lymph node metastasis. APMIS. 121:926–937. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Yang ZM, Wu XT, He T, Da MX, Luo T and
Qian K: Study on association between the expression of galectin- 3
and the peritoneal metastasis in gastric cancer. Zhonghua Wei Chang
Wai Ke Za Zhi. 8:151–154. 2005.(In Chinese). PubMed/NCBI
|
|
14
|
Miyazaki J, Hokari R, Kato S, Tsuzuki Y,
Kawaguchi A, Nagao S, Itoh K and Miura S: Increased expression of
galectin-3 in primary gastric cancer and the metastatic lymph
nodes. Oncol Rep. 9:1307–1312. 2002.PubMed/NCBI
|
|
15
|
Cheong TC, Shin JY and Chun KH: Silencing
of galectin-3 changes the gene expression and augments the
sensitivity of gastric cancer cells to chemotherapeutic agents.
Cancer Sci. 101:94–102. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
de Boer RA, Verweij N, van Veldhuisen DJ,
Westra HJ, Bakker SJ, Gansevoort RT, Muller Kobold AC, van Gilst
WH, Franke L, Mateo Leach I and van der Harst P: A genome-wide
association study of circulating galectin-3. PLoS One.
7:e473852012. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Kim SJ, Shin JY, Cheong TC, Choi IJ, Lee
YS, Park SH and Chun KH: Galectin-3 germline variant at position
191 enhances nuclear accumulation and activation of β-catenin in
gastric cancer. Clin Exp Metastasis. 28:743–750. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Hu CY, Chang SK, Wu CS, Tsai WI and Hsu
PN: Galectin-3 gene (LGALS3) +292C allele is a genetic
predisposition factor for rheumatoid arthritis in Taiwan. Clin
Rheumatol. 30:1227–1233. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Mazurek N, Byrd JC, Sun Y, Ueno S and
Bresalier RS: A galectin-3 sequence polymorphism confers TRAIL
sensitivity to human breast cancer cells. Cancer. 117:4375–4380.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Balan V, Nangia-Makker P, Schwartz AG,
Jung YS, Tait L, Hogan V, Raz T, Wang Y, Yang ZQ, Wu GS, et al:
Racial disparity in breast cancer and functional germ line mutation
in galectin-3 (rs4644): A pilot study. Cancer Res. 68:10045–10050.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Lee YH and Song GG: Genome-wide pathway
analysis in glioma. Neoplasma. 62:230–238. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Song L, Tang JW, Owusu L, Sun MZ, Wu J and
Zhang J: Galectin-3 in cancer. Clin Chim Acta. 431:185–191. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Justenhoven C, Hamann U, Pesch B, Harth V,
Rabstein S, Baisch C, Vollmert C, Illig T, Ko YD, Brüning T and
Brauch H: ERCC2 genotypes and a corresponding haplotype are linked
with breast cancer risk in a German population. Cancer Epidemiol
Biomarkers Prev. 13:2059–2064. 2004.PubMed/NCBI
|
|
24
|
Wang J, Wang W, Li R, Li Y, Tian G,
Goodman L, Fan W, Zhang J, Li J, Zhang J, et al: The diploid genome
sequence of an Asian individual. Nature. 456:60–65. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Li Z, Zhang Z, He Z, Tang W, Li T, Zeng Z,
He L and Shi Y: A partition-ligation-combination-subdivision EM
algorithm for haplotype inference with multiallelic markers: Update
of the SHEsis. http://analysis.bio-x.cnCell Res. 19:519–523.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Shi YY and He L: SHEsis, a powerful
software platform for analyses of linkage disequilibrium, haplotype
construction, and genetic association at polymorphism loci. Cell
Res. 15:97–98. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Zhou JP, Yang ZL, Liu DC and Zhou JP:
Expression of galectin 3 and Sambucus nigra agglutinin and their
clinicopathological significance in benign and malignant lesions of
stomach. Zhonghua Wei Chang Wai Ke Za Zhi. 12:297–300. 2009.(In
Chinese). PubMed/NCBI
|
|
28
|
Miyazono K, Ehata S and Koinuma D:
Tumor-promoting functions of transforming growth factor-β in
progression of cancer. Ups J Med Sci. 117:143–152. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Webber JP, Spary LK, Sanders AJ, Chowdhury
R, Jiang WG, Steadman R, Wymant J, Jones AT, Kynaston H, Mason MD,
et al: Differentiation of tumour-promoting stromal myofibroblasts
by cancer exosomes. Oncogene. 34:290–302. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Shi H, Lu D, Shu Y, Shi W, Lu S and Wang
K: Expression of multidrug resistance-related proteins
p-glycoprotein, glutathione-s-transferases, topoisomerase-II and
lung resistance protein in primary gastric cardiac adenocarcinoma.
Hepatogastroenterology. 55:1530–1536. 2008.PubMed/NCBI
|
|
31
|
Sui H, Zhou S, Wang Y, Liu X, Zhou L, Yin
P, Fan Z and Li Q: COX-2 contributes to P-glycoprotein-mediated
multidrug resistance via phosphorylation of c-Jun at Ser63/73 in
colorectal cancer. Carcinogenesis. 32:667–675. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Roy S, Kenny E, Kennedy S, Larkin A,
Ballot J, Perez De Villarreal M, Crown J and O'Driscoll L:
MDR1/P-glycoprotein and MRP-1 mRNA and protein expression in
non-small cell lung cancer. Anticancer Res. 27:1325–1330.
2007.PubMed/NCBI
|