Introduction
Gynecological cancer constitutes >10% of cancer
cases in the female population globally (1). Major gynecological types of cancer,
including cervical, endometrial, ovarian and vulvar cancer, are a
cause of concern in females. Breast cancer accounts for ~23% of all
cancer cases among females (2).
Diagnosis at advanced stages leads to increased morbidity and
mortality, in contrast to an early diagnosis, which increases the
chance of a cure and has correspondingly improved survival rates.
Diagnostic factors, including age, histology, race, histological
grade and stage at diagnosis, along with clinical phenotypes and
the degree of causal association of human papillomavirus (HPV)
infection (3), and the disease
mechanisms of the five aforementioned cancer types appear to be
distinct and precise (1). Extensive
efforts have been made over recent decades to identify diagnostic
genes and develop mechanisms for recognizing cancer in individuals
(1). However, a previous study
looking to identify prognostic genes has highlighted individual
diagnostic genes and their potential function, without scrutinizing
the properties and behavior of these diagnostic genes at the
systemic level (4).
Studies regarding expression profiles and mutation
rates have been conducted to identify common features among various
cancer types, although few studies have identified common molecular
mechanisms across diverse types of cancer (1,5,6). Certain studies employing novel profiling
techniques integrated with existing bioinformatics methods have
revealed systemic-level properties emphasizing on networks that
instigate the interconnection of genes, proteins and metabolites
whose dynamic interactions generate a corresponding function
(7) in various types of cancer
(5,6,8–11). Previously, one study identified four
transcriptional modules associated with the cell cycle and
apoptosis in cervical, endometrial and vulvar cancer (1).
System-level attributes may be better understood by
utilizing biological networks. Protein-protein interaction (PPI)
networks are the most commonly used biological networks for
exploiting information on protein interactions. Utilizing
expression profiles of the five gynecological types of cancer
(cervical, ovarian, vulvar, breast and endometrial), the present
study constructed PPI networks for each cancer type, focusing on
two main questions regarding these five disease systems. First: Are
there genes that are commonly expressed across the five disease
systems? Second: Do these commonly expressed genes share similar
network properties (including inclusion in hub genes and enrichment
in modules) across the five disease systems? The present
comparative study revealed common mechanisms and pathways
associated with the five gynecological types of cancer.
Materials and methods
Expression datasets
Gene expression datasets for five gynecological
types of cancer, breast, cervical, endometrial, vulvar and ovarian
cancer, were retrieved from the Gene Expression Omnibus (https://www.ncbi.nlm.nih.gov/geo/) (12) for the mining of differentially
expressed genes (DEGs) in the disease systems for comparison with
normal systems. The expression dataset GSE63678 (1) incorporated 5 samples of normal cervical
cells and 5 of cervical cancer cells, 5 of normal endometrial cells
and 7 of endometrial cancer cells and 7 of vulvar cancer cells and
6 of normal vulvar cells. The dataset GSE57297 (13) included 7 samples of normal and 25
samples of breast cancer cells. The ovarian cancer dataset GSE26712
(14) included 10 samples of normal
and 185 samples of ovarian cancer cells.
Analyses of differential
expression
Differential expression analyses of the datasets of
different cancer types were performed using the Limma (Linear
Models for Microarray Data) package (15) in R (16). Pre-processing and normalization were
performed to remove noise from the datasets. Adjusted P-values
<0.05 and |fold change| >2 were taken into account for the
identification of DEGs. The P-value was adjusted using the
Benjamini-Hochberg method (17) to
minimize errors due to multiple hypothesis testing. Overlapping
DEGs among the five gynecological types of cancer were selected for
further study.
PPI network construction
DEGs of the five cancer types were individually
mapped to PPI networks, considering a medium confidence in the
Search Tool for the Retrieval of Interacting Genes/Proteins
(STRING) version 10.0 (18) database,
with Homo sapiens as the source species. Interaction data
from high-throughput experiments, genomic contexts, previous
knowledge and conserved co-expression are extensively integrated by
STRING. The PPI networks were constructed and visualized in
Cytoscape (19) version 3.2.
Common module and hubs extraction
The PPI networks for the five gynecological types of
cancer were scrutinized for the presence of common sub-network,
module and hub genes. A common sub-network was identified by
comparing the nodes and edges among the five PPI networks. The
common sub-network was further investigated to identify a common
high-modularity cluster among the five PPI networks using the
Molecular Complex Detection (MCODE) (20) plug-in for Cytoscape. Node score ≥0.2,
k-core=2, degree ≥2 and maximum depth=100 were used as the cut-off
criteria for the extraction of common modules. Hub genes were
selected from the PPI networks with a connectivity degree of
>25, and overlapping hubs among the five cancer networks were
identified.
Enrichment analyses
Pathway and disease enrichment analyses for the
common module genes and common hubs shared between the five
gynecological types of cancer were performed using the Web-based
Gene Set Analysis Toolkit (21). The
cutoff criteria of P<0.05 and number of genes >2 were used
for enrichment. Hyper-geometric distribution was utilized for
enrichment analyses.
Results
DEGs identification
P<0.05 and |fold change| >2 were used as the
demarcating parameters for the identification of DEGs in the five
disease systems. A total of 3,521 DEGs for breast cancer, 1,086
DEGs for cervical cancer, 478 DEGs for endometrial cancer, 2,028
DEGs for ovarian cancer and 728 DEGs for vulvar cancer were
identified, all compared with normal samples. The mined DEGs
included upregulated and downregulated DEGs. Next, overlapping DEGs
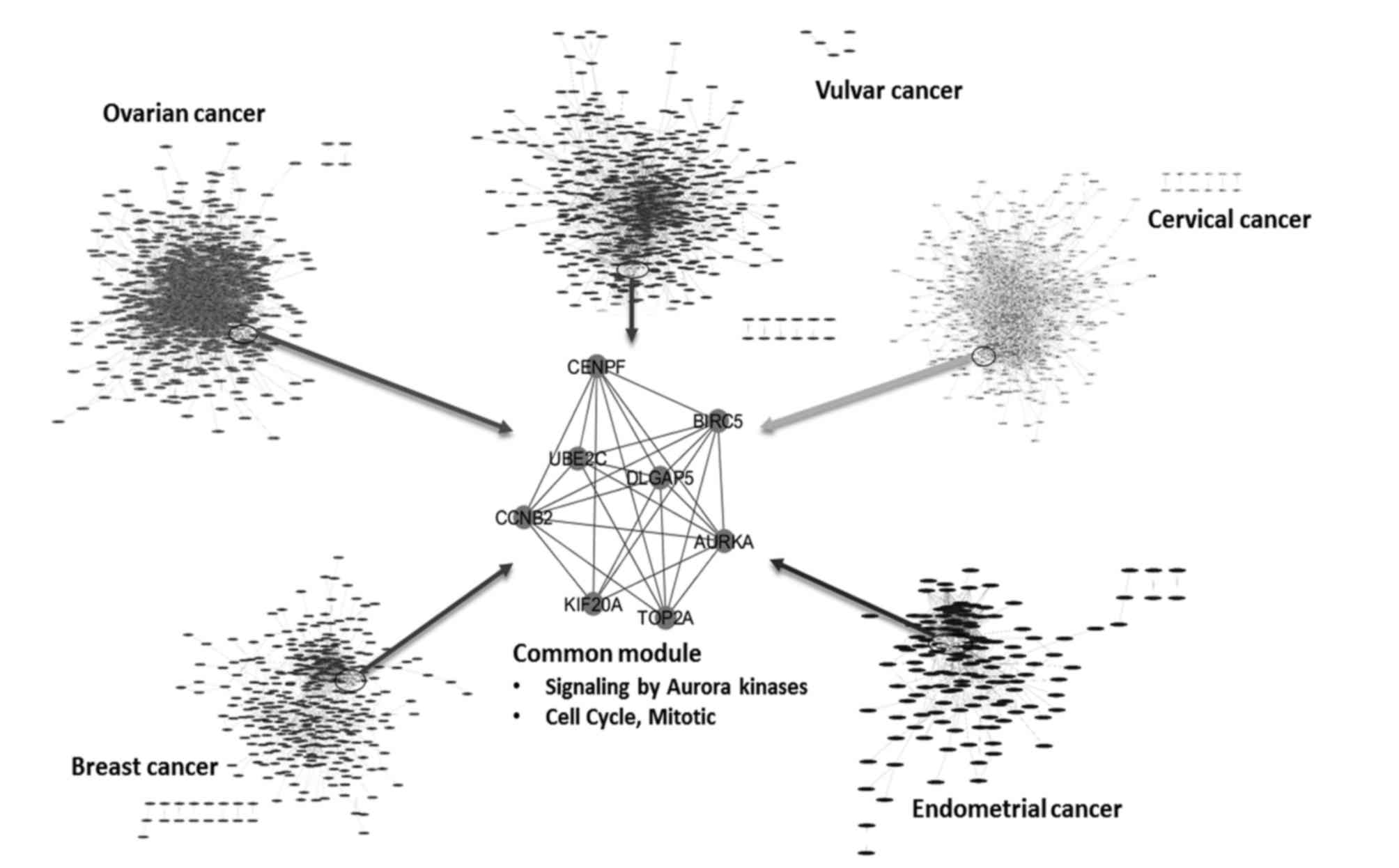
among the five cancer disease systems were identified (Fig. 1). A total of 10 DEGs, namely, cyclin
B2 (CCNB2), ubiquitin-conjugating enzyme E2C (UBE2C), topoisomerase
(DNA) II-α (TOP2A), centromere protein F (CENPF), baculoviral IAP
repeat containing 5 (BIRC5), aurora kinase A (AURKA), discs, large
(Drosophila) homolog-associated protein 5 (DLGAP5),
transforming growth factor-β receptor III (TGFBR3), Krüppel-like
factor 11 (KLF11) and kinesin family member 20A (KIF20A), were
found to be differentially expressed in all five gynecological
types of cancer. In total, 29 DEGs were revealed to overlap among
cervical, endometrial, breast and ovarian cancer.
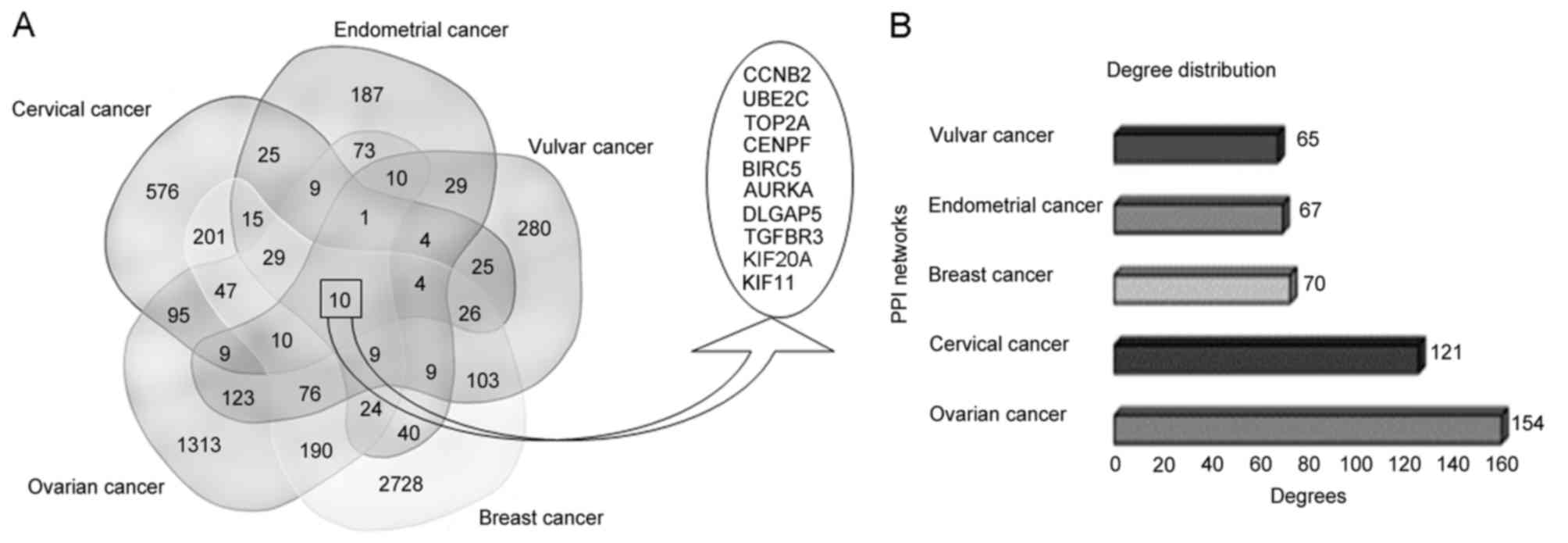 | Figure 1.(A) Venn diagram representing the 10
common differently expressed genes among the five female-specific
cancer types. (B) Degree distribution of the five cancer PPI
networks. Number adjacent to the horizontal represents the maximum
degree of the nodes (proteins) in the PPI network. PPI,
protein-protein interaction; CCNB2, cyclin 2; UBE2C,
ubiquitin-conjugating enzyme E2 C; TOP2A, DNA topoisomerase II-α;
CENPF, centromere protein F; BIRC5, baculoviral IAP repeat
containing 5; AURKA, aurora kinase A; DLGAP5, DLG associated
protein 5; TGFBR3, transforming growth factor-β receptor 3; KIF20A,
kinesin family member 20A; KIF11, kinesin family member 11. |
PPI network construction
Extracted DEGs from the five cancer datasets were
mapped to create PPI networks by utilizing gene-coding protein
interaction information. PPI networks for cervical cancer, breast
cancer, endometrial cancer, ovarian cancer and vulvar cancer were
formed with 689 nodes (4,505 edges), 364 nodes (1,494 edges), 155
nodes (1,005 edges), 733 nodes (5,721 edges) and 542 nodes (1,772
edges), respectively where each node represents a protein and the
edge between the nodes represents the interaction between the
proteins. These PPI networks were visualized in Cytoscape for
further analysis.
Common module and hubs
Overlapping nodes and edges among the five PPI
networks were identified to construct a common sub-network among
the five gynecological types of cancer. A common module with 8
nodes (MCODE score=7.429; with 26 edges) was extracted from the
common sub-network by MCODE. The common modules of the five cancer
networks included 8/10 overlapping DEGs: CCNB2, UBE2C, TOP2A,
CENPF, BIRC5, AURKA, DLGAP5 and KIF20A (Fig. 2). Additionally, the PPI networks for
the five gynecological types of cancer were analyzed for the
identification of hub genes exhibiting connectivity degrees of
>25 (Table I). In total, 4 common
hub genes (TOP2A, BIRC5, AURKA and CCNB2) were mined from the five
gynecological types of cancer.
 | Table I.Hub genes of five female-specific
cancer PPI networks. |
Table I.
Hub genes of five female-specific
cancer PPI networks.
| Cancer type | Hub genes |
|---|
| Ovarian cancer | AKT1, GAPDH, CDK1,
JUN, TOP2A, CCNB1, RAC1, FOS, ACTB, AURKA, HSPA5, ACTA2, BIRC5,
CCNB2, CCNA2, CDC20, MMP2, TBP, SMARCA4, ERBB2, MAD2L1, YWHAZ,
VEGFA, CDKN2A, TGFB1, BUB1, CDK16, IL8, MCM3, BUB1B, ACTG2, YWHAE,
H2AFX, GNB2L1, ISG15, ENO1, TUBA1B, COL1A1, RHOB, TUBA4A, APOE,
RACGAP1, MYBL2, TUBB4B, BUB3, RRM2, THBS1, TUBA1C, EIF4G1, PRKDC,
HSPB1, MCM7, UBE2C, JUND, COL1A2, TUBB3, NCBP2, SERPINE1, CCNA1,
TUBB, KIF2C, PRC1, CCT5, RPL23, TYMS, CALR, HSPD1, CCNE1, PSMB2,
PSMB3, TXN, PKM, MYH9, RAD23A, RPL27A, RAD21, SDHC, NEK2, HNRNPC,
RPL38, RPL37A, CCT2, HMGB1, FOXM1, OAS1, MMP14, RALA, CALM1, YWHAH,
CDCA8, VWF, SDC1, MCL1, ACTN1, PSMC2, SMC4, SNRPE, KIF20A, RBBP4,
SNRPF, RPS7, MCM4, KIF4A, SNRPB, OAS3, ACTN4, CFL1, MMP7, CENPF,
OASL, ASF1A, RPL36, SMARCB1, BAK1, HSPH1, CTGF, MRPL13, JUNB,
COL18A1, SPTAN1, HSPG2, PTBP1, PLK2, RANGAP1, MCM2, ATF3, HSF1,
MELK, PPP1CA, SNRPD2, POLR2K, P4HB, COL4A1, SF3A2, ASF1B, PABPN1,
DLGAP5, SNCA, CEBPB, PDIA4, PAK2, CLU, PRKCD, COL4A2, ARF1, NUSAP1,
POLR2J, RPL36A, SMARCC1, UBA1, RAB8A, COX6A1, POLR2I, TSC2, CEP55,
PRPF4 |
| Cervical
cancer | CDK1, TOP2A, PCNA,
CDK2, CCNB1, BUB1, MAD2L1, BIRC5, CCNB2, MCM5, AURKA, CDC6, MCM3,
CHEK1, MCM4, CENPA, MCM7, KIF11, BUB1B, MCM2, RAD51, NDC80, BRCA1,
ESR1, ATR, XPO1, STAT1, RFC4, CDKN2A, KIF2C, NUF2, KIF23, CDC25A,
BUB3, PRC1, CENPF, RRM2, RBBP4, MMP9, MCM6, TYMS, SMC4, IL8, RFC3,
CCNE1, PIK3CA, KIF20A, FEN1, SMC2, RECQL, KNTC1, TOPBP1, KIF4A,
NEK2, MSH2, PRKDC, MCM8, CDC7, CDC25C, WEE1, STAG1, CASC5, RACGAP1,
CCNE2, EGR1, ZWILCH, CENPI, MCM10, KIF18A, RFC5, NCAPG, NUP107,
ACTA2, MSH6, E2F3, NDE1, PDS5B, DLGAP5, CDC25B, GMPS, WDHD1,
NUSAP1, PLK4, FOXM1, ISG15, RBL1, ECT2, CKS2, CENPN, CENPQ, CENPK,
TTK, ZWINT, ITGB1, AHCTF1, DSN1, CEP55, ANLN, UBE2C, BARD1, GEN1,
DDX58, COL1A1, SGOL2, POLE2, MELK, TRIP13, APOE, DNA2, ASPM, CKS1B,
GINS1, EZH2, MMP1, FANCD2, DBF4, ESCO2, PRIM1, YWHAH |
| Endometrial
cancer | CDK1, TOP2A, CCNB1,
CDC20, CCNB2, AURKA, BIRC5, MAD2L1, CCNA2, KIF11, BUB1B, PRC1,
CENPE, KIF23, KIF4A, KIF2C, CENPA, CDC6, DLGAP5, CENPF, NEK2,
ESPL1, KIF20A, RACGAP1, NUSAP1, RRM2, UBE2C, MELK, CEP55,
NCAPG |
| Breast cancer | CDK1, TOP2A, PLK1,
CCNB2, AURKA, BUB1, BIRC5, CENPA, MMP9, BUB1B, AGT, NDC80, PRC1,
CENPE, KIF2C, CDC42, KIF4A, NUF2, CDCA8, ESR1, KIF20A, STAT1,
CENPF, DLGAP5, NUSAP1, ANLN |
| Vulvar cancer | BCL2, VEGFA,
NOTCH1, FYN, AR, STAT1, CREB1, PIK3R1, MMP9, KIT, IL8, TOP2A,
ACACB, IGF1, VWF, MET, BIRC5, MMP1, FOXO3, GATA3, DCN, YWHAZ,
CCNB2, AURKA, IL1B |
Enrichment analyses
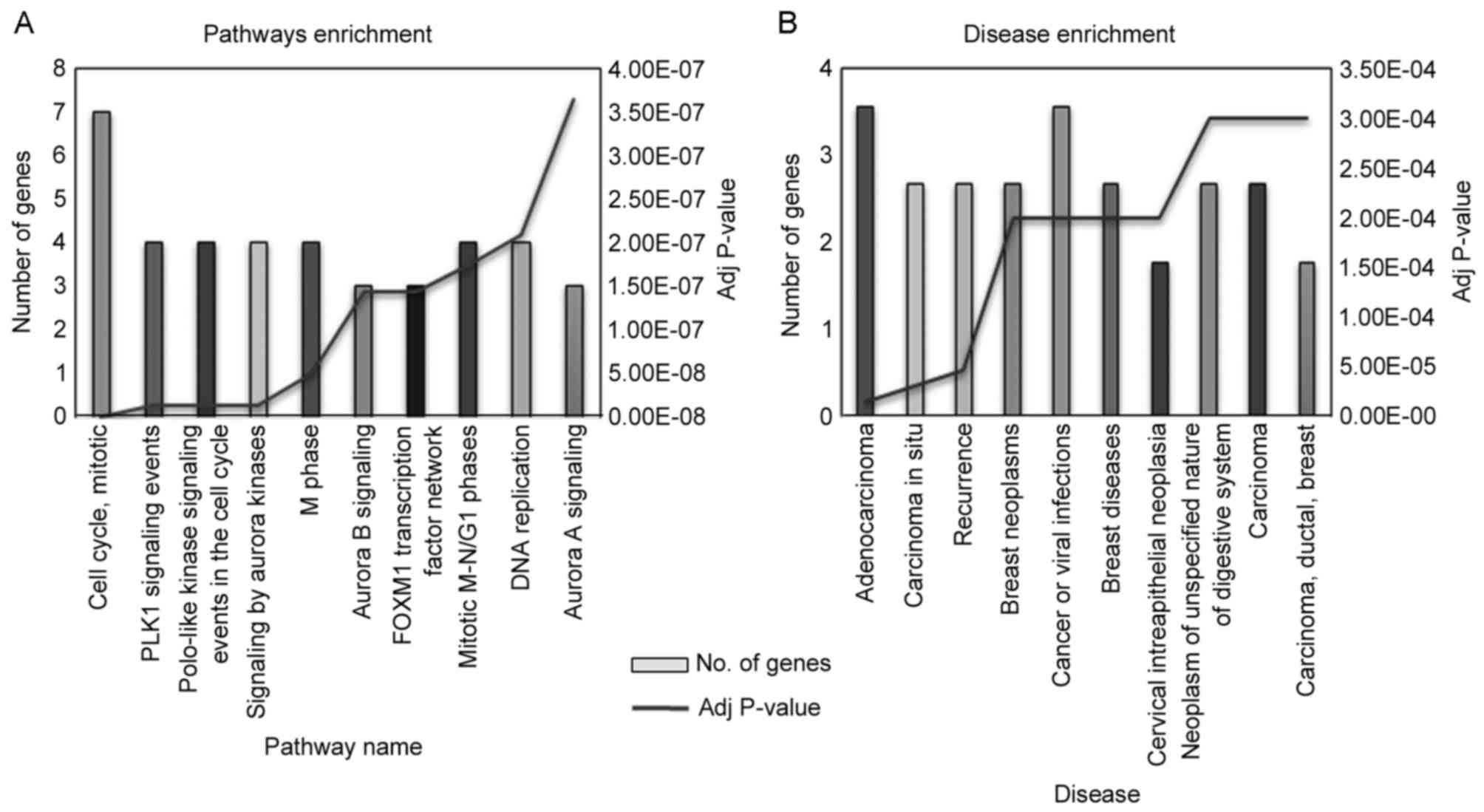
Pathway enrichment analyses of the common module
genes (Fig. 3A) revealed their
association with the cell cycle, mitotic and aurora kinase
signaling pathways. Overall, 7 genes from the common module were
found to be associated with the cell cycle and mitotic pathways.
There were 4 genes, KIF20A, UBE2C, BIRC5 and CENPF, enriched in the
mitotic phase-associated pathways. KIF20A, BIRC5, AURKA and DLGAP5
were involved in multiple pathways, including the aurora kinase
signaling pathway and the polo-like kinase signaling event in the
cell cycle. Moreover, 3/4 common hub genes (TOP2A, BIRC5 and AURKA)
were revealed to have a function in the aurora kinase signaling
pathway. The genes of the common module were enriched in certain
conditions, including cancer, viral infection and adenocarcinoma
(Fig. 3B).
Discussion
In the present study, overlapping modules and DEGs
among five gynecological types of cancer (breast, cervical,
endometrial, vulvar and ovarian cancer) were mined to identify
common pathways, which may indicate a common method of
tumorigenesis. An integrative approach utilizing differential
expression and network analyses was employed for this purpose. It
has been hypothesized that cervical, endometrial and vulvar cancer
may share common molecular and biochemical features, and the
resulting pathways present during cancer development in these three
cancer types could shed light on disease development and
embryogenesis aspects (1). The
present study involved the construction of PPI networks for the
five gynecological types of cancer from the gene-encoded protein
interaction information, which are differentially expressed at the
malignant stage compared with normal tissues. There were 10 DEGs
found to be overlapping among the five cancer systems, with 4 DEGs
identified as common hubs of the five PPI networks. The PPI
networks were investigated to identify overlapping nodes and edges,
and to reveal a common sub-network shared by all five
female-specific cancer types. There were 4 genes (AURKA, KIF20A,
DLGAP5 and BIRC5) involved in aurora kinase signaling pathway
activation. Aurora kinase activity peaks during the G2
phase to mitotic phase transition in the cell cycle. KIF20A, UBE2C,
BIRC5 and CENPF were enriched in the mitotic phase-associated
pathways, thus assisting in the dysregulation of the aurora kinase
pathway. AURKA, TOP2A, KIF20A, CCNB2, UBE2C, BIRC5 and CENPF were
enriched in the dysregulation of the cell cycle and mitotic
pathways, which in turn could affect the normal function of the
aurora kinase signaling pathways. In total, 7/8 genes from the
common module were involved in the dysregulation of the aurora
kinase signaling pathway.
The aurora kinase family is comprised of 3
serine-threonine protein kinases: Aurora kinase A, aurora kinase B
and aurora kinase C, and the aurora kinase A protein is encoded by
the AURKA gene. This family of proteins are regulators of cell
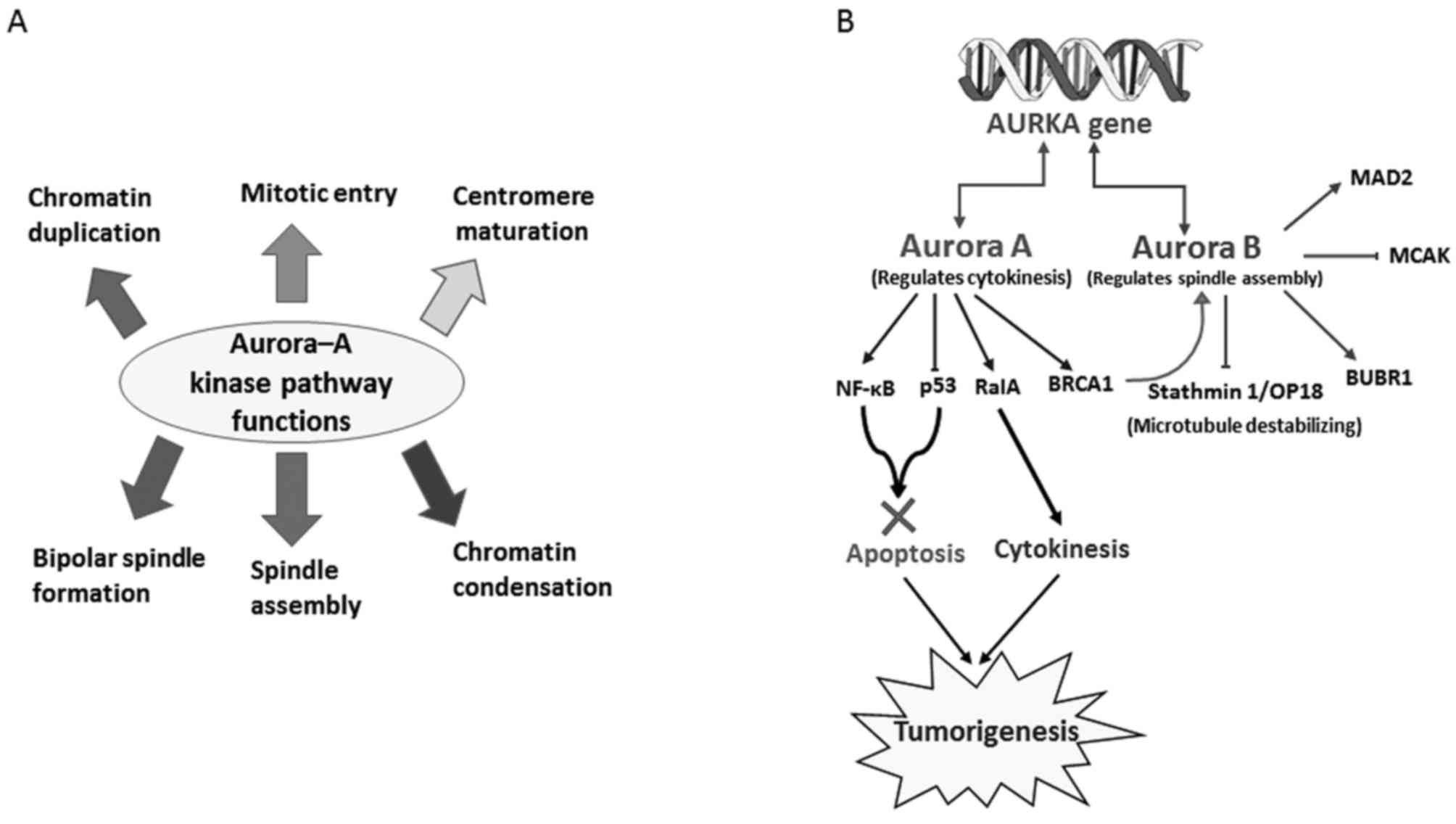
division (22). Since the aurora
kinase pathway is involved in chromatin duplication, mitotic entry,
centromere maturation, chromatin condensation, spindle assembly and
bipolar spindle formation (Fig. 4A)
(23), overexpression or
dysregulation of the associated pathways may lead to an increased
rate of cell division and then to tumorigenesis.
Overexpression of aurora kinase has been reported in
breast, liver, pancreatic, bladder, thyroid and gastrointestinal
cancer types (24). AURKA
overexpression has been proposed to be associated with aneuploidy
in breast cancer (25). Polymorphisms
in the AURKA gene have been shown to increase the risk of primary
breast cancer (26). Studies have
revealed that the increased expression of AURKA is associated with
laryngeal, nasopharyngeal and breast cancer metastasis (27–30). AURKA
has been demonstrated to be accountable for the phosphorylation of
Breast Cancer 1, Early Onset (BRCA1) (31). Tanaka et al (32) investigated the immunohistochemical
analysis of invasive ductal adenocarcinomas of the breast, that
revealed the overexpression of aurora kinases in a majority of
cases. Miyoshi et al (33)
investigated the correlation of aurora kinase A mRNA expression
with numerous clinic pathological factors and CIN in breast cancer.
Nadler et al (34)
investigated the correlation of a population of patients with a
decreased survival rate based on aurora kinase A expression,
whereas aurora kinase B does not interfere with the survival of
patients. Thus, aurora kinase A was proposed to be a preferred drug
target in breast cancer. Studies have indicated that aurora kinase
A and aurora kinase B are significantly overexpressed in carcinoma
and in cervical intraepithelial neoplasia 3 (CIN3), compared with
in the normal cervix (35). A
previous study found that the overexpression of aurora kinase A is
crucial for survival of HPV-transformed cervical cancer cells
(36). Overexpression of aurora
kinase A is observed in 83% of human epithelial ovarian carcinoma
cases (37). It has been proposed
that AURKA regulates cell migration and adhesion in epithelial
ovarian cancer (38). Human
chromosome 20q13.2, which contains AURKA, is usually amplified in
ovarian cancer (39). Furthermore,
Aurora kinase A has been revealed to be associated with Federation
Internationale de Gynecologie et d'Obstetrique stage, tumor grade
and survival (37). AURKA and AURKB
are overexpressed in endometrial cancer, in comparison with normal
proliferative tissues, as determined by Kurai et al
(40). Moreno-Bueno et al
(41) revealed the presence of high
AURKA expression levels in type-II adenocarcinoma via the
microarray analysis of endometrial cancer tissue. Few studies are
available that highlight the potential mechanisms for vulvar cancer
(42). It has been revealed to
originate from two distinct pathways: HPV-dependent and
HPV-independent (43).
KIF20A belongs to the kinesin superfamily, which is
involved in essential cellular functions including migration,
mitosis and intracellular transport though interaction with
microtubules (44). KIF20A has been
proposed to be a potential immunotherapeutic target for cancer
(45) and is often overexpressed in
breast cancer, lung cancer, pancreatic cancer, gastric cancer,
bladder cancer, melanoma and other malignancies. (46–48).
KIF20A small interfering RNA (siRNA), along with six other siRNAs,
has been reported to kill HeLa cervical cancer cells (49). Another study revealed KIF20A to be a
hub of the gene co-expression network of endometrial cancer
(50).
BIRC5 is a member of the inhibitor of apoptosis
family, which is involved in mitosis and protection against
apoptotic cell death (51). Signaling
networks required for tumor maintenance are expected to be
compromised by disabling BIRC5 and survivin, thereby enhancing its
utility as a cancer drug target (52). Genomic copy number variations in BIRC5
have been proposed to be involved in the progression of breast
cancer (53). Furthermore, BIRC5
expression may result in breast tumor proliferation by promoting
genetic instability (54). BIRC5 has
been reported to be overexpressed in cervical cancer (55). Overexpression of BIRC5 promotes
cervical cancer progression and metastasis (55). BIRC5 has been reported to be
overexpressed in various human malignancies, including ovarian
cancer (56) and endometrial cancer
(57). Human telomerase reverse
transcriptase is an enzyme that enables cells to divide
indefinitely and overcome replicative senescence (58); this enzyme has been reported to be
upregulated in vulvar intraepithelial neoplasia, with the emergence
of resistance to apoptosis and cellular longevity via survivin
activation (58).
Silencing of DLGAP suppresses tumorigenicity and
inhibits cellular proliferation by inducing cell cycle arrest at
the G2/M phase (59).
DLGAP5 has been proposed to be a target of the neurogenic locus
notch homolog protein 3 signaling pathway (59), which is associated with embryogenesis
and breast cancer tumorigenesis (60). DLAGP5 is reportedly a member of the
cervical cancer interaction network (61), and has been revealed to be
overexpressed in ovarian carcinoma compared with in normal ovarian
epithelium (59). DLGAP5 is
overexpressed in endometrial cancer tissues, in comparison with
normal endometrial tissues (62).
Overexpression of aurora kinases not only
dysregulates the cell cycle but also facilitates interactions with
several cancer-associated proteins, including tumor protein p53,
nuclear factor κ-light-chain enhancer of activated B cells (NF-κB),
v-Myc, Ras and pathways including the mitogen-activated protein
kinase/extracellular-related kinase and phosphoinositide-3
kinase/protein kinase B signaling pathways (Fig. 4B) (63,64). In
the present study, the overexpression of these oncogenes was also
observed. Aurora kinase A is essential for cytokinesis as it
activates NF-κB, an anti-apoptotic gene that is involved in growth
regulation of cells, the overexpression of which leads to cancer
(65). Additionally, p53 induces
apoptosis; the overexpression of aurora kinase A phosphorylates p53
and inhibits its function. (22).
Aurora kinase A may phosphorylate RalA, thus influencing Ras
signaling and leading to tumorigenesis (66). Myc was also overexpressed in the
present analysis. Myc is a quintessential oncogene, which is
overexpressed in cancer cells and functions as an angiogenic switch
(67). Myc has been revealed to be
present in number of cancer types, including colorectal, prostate
and breast cancer (68). Aurora
kinase B overexpression leads to chromosomal segregation error and
abnormal cytokinesis, potentially causing carcinogenesis (63).
Aurora kinase A has been revealed to have a vital
function in increasing chemotherapy resistance, cell exhibiting
stem-cell-like properties and mesenchymal phenotypes (69). Thus, the inhibition of aurora kinase A
has demonstrated an increased sensitivity and decreased
proliferation of breast cancer to chemotherapy and hormonal
treatments (70). The aurora kinase B
inhibitor ZM447439 has already been proven to suppress the growth
of cervical cancer cells, thus enhancing chemosensitivity (71). A previous study has confirmed the
potential utility of the aurora kinase A inhibitor alisertib, when
used in combination with taxanes, as a therapeutic strategy for
ovarian cancer (72). To verify the
clinical significance of AURKA in improving the therapy of
endometrial cancer, a combination treatment utilizing AURKA
inhibitors and paclitaxel is proposed (73).
DEGs identified from the microarray analysis of the
five gynecological types of cancerous cell sample were utilized for
the construction of PPI networks in the present study.
Identification and analysis of the common hub genes and modules
suggested the presence of the aurora kinase pathway in all five of
the gynecological types of cancer. Aurora kinases are a potentially
promising area of research, as they are overexpressed in five
gynecological types of cancer and are key regulators of cell
division, as well as acting together with a number of oncogenes.
The findings of the present study may be utilized for the
development of exhaustive models of carcinogenesis, which may lead
to the development of novel therapeutic approaches that target
mechanisms frequently altered across various cancer types. However,
further experimental validations are required to confirm these
findings.
Acknowledgements
Computing facilities at the Indian Institute of
Information Technology (Allahabad, India) are acknowledged.
References
|
1
|
Pappa KI, Polyzos A, Jacob-Hirsch J,
Amariglio N, Vlachos GD, Loutradis D and Anagnou NP: Profiling of
discrete gynecological cancers reveals novel transcriptional
modules and common features shared by other cancer types and
embryonic stem cells. PLoS One. 10:e01422292015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Mohankumar K, Pajaniradje S, Sridharan S,
Singh VK, Ronsard L, Banerjea AC, Benson CS, Coumar MS and
Rajagopalan R: Mechanism of apoptotic induction in human breast
cancer cell, MCF-7, by an analog of curcumin in comparison with
curcumin-an in vitro and in silico approach. Chem Biol Interact.
210:51–63. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
US Cancer Statistics Working Group, .
United States Cancer Statistics: 1999–2011 incidence and mortality
web-based report. U.S. Department of Health and Human Services,
Center for Disease Control and Prevention and National Cancer
Institute; Atlanta, GA: 2014, https://www.cdc.gov/uscs3–March. 2015
|
|
4
|
Yang Y, Han L, Yuan Y, Li J, Hei N and
Liang H: Gene co-expression network analysis reveals common
system-level properties of prognostic genes across cancer types.
Nat Commun. 5:32312014.PubMed/NCBI
|
|
5
|
Peng L, Bian XW, Li DK, Xu C, Wang GM, Xia
QY and Xiong Q: Large-scale RNA-seq transcriptome analysis of 4043
cancers and 548 normal tissue controls across 12 TCGA cancer types.
Sci Rep. 5:134132015. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Martinez E, Yoshihara K, Kim H, Mills GM,
Treviño V and Verhaak RG: Comparison of gene expression patterns
across 12 tumor types identifies a cancer supercluster
characterized by TP53 mutations and cell cycle defects. Oncogene.
34:2732–2740. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Alberghina L, Höfer T and Vanoni M:
Molecular networks and system-level properties. J Biotechnol.
144:224–33. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Alisoltani A, Fallahi H, Ebrahimi M,
Ebrahimi M and Ebrahimie E: Prediction of potential cancer-risk
regions based on transcriptome data: towards a comprehensive view.
PLoS One. 9:e963202014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Mine KL, Shulzhenko N, Yambartsev A,
Rochman M, Sanson GF, Lando M, Varma S, Skinner J, Volfovsky N,
Deng T, et al: Gene network reconstruction reveals cell cycle and
antiviral genes as major drivers of cervical cancer. Nat Commun.
4:18062013. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Hoadley KA, Yau C, Wolf DM, Cherniack AD,
Tamborero D, Ng S, Leiserson MDM, Niu B, McLellan MD, Uzunangelov
V, et al: Multiplatform analysis of 12 cancer types reveals
molecular classification within and across tissues of origin. Cell.
158:929–944. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Fruzangohar M, Ebrahimie E, Ogunniyi AD,
Mahdi LK, Paton JC and Adelson DL: Correction: Comparative GO: A
web application for comparative gene ontology and gene
ontology-based gene selection in bacteria. PLoS One.
10:e01255372015. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Barrett T, Wilhite SE, Ledoux P,
Evangelista C, Kim IF, Tomashevsky M, Marshall KA, Phillippy KH,
Sherman PM, Holko M, et al: NCBI GEO: Archive for functional
genomics data sets-update. Nucleic Acids Res. 41:D991–D995. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Fu J, Allen W, Xia A, Ma Z and Qi X:
Identification of biomarkers in breast cancer by gene expression
profiling using human tissues. Genom Data. 2:299–301. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Bonome T, Levine DA, Shih J, Randonovich
M, Pise-Masison CA, Bogomolniy F, Ozbun L, Brady J, Barrett JC,
Boyd J and Birrer MJ: A gene signature predicting for survival in
suboptimally debulked patients with ovarian cancer. Cancer Res.
68:5478–5486. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Gautier L, Cope L, Bolstad BM and Irizarry
RA: Affy-analysis of affymetrix genechip data at the probe level.
Bioinformatics. 20:307–315. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Team RCore, . R: A language and
environment for statistical computing. Vienna, Austria: R
Foundation for Statistical Computing; pp. 3–36. 2014, http://www.r-project.org/
|
|
17
|
Benjamini Y and Hochberg Y: Controlling
the false discovery rate: A practical and powerful approach to
multiple testing. Journal of the Royal Statistical Society. J R
Statics Soc B. 1–300. 1995.
|
|
18
|
Von Mering C, Huynen M, Jaeggi D, Schmidt
S, Bork P and Snel B: STRING: A database of predicted functional
associations between proteins. Nucleic Acids Res. 31:258–261. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Smoot ME, Ono K, Ruscheinski J, Wang PL
and Ideker T: Cytoscape 2.8: New features for data integration and
network visualization. Bioinformatics. 27:431–432. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Bader GD and Hogue CW: An automated method
for finding molecular complexes in large protein interaction
networks. BMC Bioinformatics. 4:22003. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wang J, Duncan D, Shi Z and Zhang B:
Web-based gene set analysis toolkit (WebGestalt): Update 2013.
Nucleic Acids Res. 41:W77–W83. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Dar AA, Goff LW, Majid S, Berlin J and
El-Rifai W: Aurora kinase inhibitors-rising stars in cancer
therapeutics? Mol Cancer Ther. 9:268–278. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2015. CA Cancer J Clin. 65:5–29. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Carvajal RD, Tse A and Schwartz GK: Aurora
kinases: New targets for cancer therapy. Clin Cancer Res.
12:6869–75. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Xia LP, Zhou FF, Yang MT and Liu Q: Roles
of Aurora-A in tumorigenesis and prognosis of breast cancer. Ai
Zheng. 28:668–672. 2009.(In Chinese). PubMed/NCBI
|
|
26
|
Cox DG, Hankinson SE and Hunter DJ:
Polymorphisms of the AURKA (STK15/Aurora Kinase) gene and breast
cancer risk (United States). Cancer Causes Control. 17:81–83. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Yang LY, He CY, Chen XH, Su LP, Liu BY and
Zhang H: Aurora kinase A revives dormant laryngeal squamous cell
carcinoma cells via FAK/PI3K/Akt pathway activation. Oncotarget.
7:48346–48359. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Lo Iacono ML, Monica V, Saviozzi S, Ceppi
P, Bracco E, Papotti M and Scagliotti GV: Aurora Kinase A
expression is associated with lung cancer histological-subtypes and
with tumor de-differentiation. J Transl Med. 9:1002011. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Staff S, Isola J, Jumppanen M and Tanner
M: Aurora-A gene is frequently amplified in basal-like breast
cancer. Oncol Rep. 23:307–312. 2010.PubMed/NCBI
|
|
30
|
Zou Z, Yuan Z, Zhang Q, Long Z, Chen J,
Tang Z, Zhu Y, Chen S, Xu J, Yan M, et al: Aurora kinase A
inhibition-induced autophagy triggers drug resistance in breast
cancer cells. Autophagy. 8:1798–1810. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Ouchi M, Fujiuchi N, Sasai K, Katayama H,
Minamishima YA, Ongusaha PP, Deng C, Sen S, Lee SW and Ouchi T:
BRCA1 phosphorylation by Aurora-A in the regulation of G2 to M
transition. J Biol Chem. 279:19643–19648. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Tanaka T, Kimura M, Matsunaga K, Fukada D,
Mori H and Okano Y: Centrosomal kinase AIK1 is overexpressed in
invasive ductal carcinoma of the breast. Cancer Res. 59:2041–2044.
1999.PubMed/NCBI
|
|
33
|
Miyoshi Y, Iwao K, Egawa C and Noguchi S:
Association of centrosomal kinase STK15/BTAK mRNA expression with
chromosomal instability in human breast cancers. Int J Cancer.
92:370–373. 2001. View
Article : Google Scholar : PubMed/NCBI
|
|
34
|
Nadler Y, Camp RL, Schwartz C, Rimm DL,
Kluger HM and Kluger Y: Expression of Aurora A (but not Aurora B)
is predictive of survival in breast cancer. Clin Cancer Res.
14:4455–4462. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Twu NF, Yuan CC, Yen MS, Lai CR, Chao KC,
Wang PH, Wu HH and Chen YJ: Expression of Aurora kinase A and B in
normal and malignant cervical tissue: High Aurora A kinase
expression in squamous cervical cancer. Eur J Obstet Gynecol Reprod
Biol. 142:57–63. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Gabrielli B, Bokhari F, Ranall MV, Oo ZY,
Stevenson AJ, Wang W, Murrell M, Shaikh M, Fallaha S, Clarke D, et
al: Aurora A is critical for survival in HPV-transformed cervical
cancer. Mol Cancer Ther. 14:2753–2761. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Campos SM and Ghosh S: A current review of
targeted therapeutics for ovarian cancer. J Oncology.
2010:1493622010. View Article : Google Scholar
|
|
38
|
Do TV, Xiao F, Bickel LE, Klein-Szanto AJ,
Pathak HB, Hua X, Howe C, O'Brien SW, Maglaty M, Ecsedy JA, et al:
Aurora kinase A mediates epithelial ovarian cancer cell migration
and adhesion. Oncogene. 33:539–549. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Katsha A, Belkhiri A, Goff L and El-Rifai
W: Aurora kinase A in gastrointestinal cancers: Time to target. Mol
Cancer. 14:1062015. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Kurai M, Shiozawa T, Shih HC, Miyamoto T,
Feng YZ, Kashima H, Suzuki A and Konishi I: Expression of Aurora
kinases A and B in normal, hyperplastic and malignant human
endometrium: Aurora B as a predictor for poor prognosis in
endometrial carcinoma. Hum Pathol. 36:1281–1288. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Moreno-Bueno G, Sánchez-Estévez C, Cassia
R, Rodríguez-Perales S, Díaz-Uriarte R, Domínguez O, Hardisson D,
Andujar M, Prat J, Matias-Guiu X, et al: Differential gene
expression profile in endometrioid and nonendometrioid endometrial
carcinoma STK15 is frequently overexpressed and amplified in
nonendometrioid carcinomas. Cancer Res. 63:5697–5702.
2003.PubMed/NCBI
|
|
42
|
van de Nieuwenhof HP, van der Avoort IA
and de Hullu JA: Review of squamous premalignant vulvar lesions.
Crit Rev Oncol Hematol. 68:131–156. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Ueda Y, Enomoto T and Kimura T, Yoshino K,
Fujita M and Kimura T: Two distinct pathways to development of
squamous cell carcinoma of the vulva. J Skin Cancer.
2011:9512502011. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Miki H, Setou M, Kaneshiro K and Hirokawa
N: All kinesin superfamily protein, KIF, genes in mouse and human.
Proc Natl Acad Sci USA. 98:pp. 7004–7011. 2001; View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Tomita Y, Yuno A, Tsukamoto H, Senju S,
Kuroda Y, Hirayama M, Irie A, Kawahara K, Yatsuda J, Hamada A, et
al: Identification of promiscuous KIF20A long peptides bearing both
CD4+ and CD8+ T-cell epitopes: KIF20A-specific CD4+ T-cell immunity
in patients with malignant tumor. Clin Cancer Res. 19:4508–4520.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Imai K, Hirata S, Irie A, Senju S, Ikuta
Y, Yokomine K, Harao M, Inoue M, Tomita Y, Tsunoda T, et al:
Identification of HLA-A2-restricted CTL epitopes of a novel
tumour-associated antigen, KIF20A, overexpressed in pancreatic
cancer. Br J Cancer. 104:300–307. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Yamashita J, Fukushima S, Jinnin M, Honda
N, Makino K, Sakai K, Masuguchi S, Inoue Y and Ihn H: Kinesin
family member 20A is a novel melanoma-associated antigen. Acta Derm
Venereol. 92:593–597. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Claerhout S, Lim JY, Choi W, Park YY, Kim
K, Kim SB, Lee JS, Mills GB and Cho JY: Gene expression signature
analysis identifies vorinostat as a candidate therapy for gastric
cancer. PLoS One. 6:e246622011. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Groth-Pedersen L, Aits S, Corcelle-Termeau
E, Petersen NH, Nylandsted J and Jäättelä M: Identification of
cytoskeleton-associated proteins essential for lysosomal stability
and survival of human cancer cells. PloS one. 7:e453812012.
View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Chou WC, Cheng AL, Brotto M and Chuang CY:
Visual gene-network analysis reveals the cancer gene co-expression
in human endometrial cancer. BMC genomics. 15:3002014. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Marivin A, Berthelet J, Plenchette S and
Dubrez L: The inhibitor of apoptosis (IAPs) in adaptive response to
cellular stress. Cells. 1:711–737. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Sawyers C: Targeted cancer therapy.
Nature. 432:294–297. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Ghaffari K, Hashemi M, Ebrahimi E and
Shirkoohi R: birc5 genomic copy number variation in early-onset
breast cancer. Iran Biomed J. 20:241–245. 2016.PubMed/NCBI
|
|
54
|
Boidot R, Vegran F, Jacob D, Chevrier S,
Gangneux N, Taboureau J, Oudin C, Rainville V, Mercier L and
Lizard-Nacol S: The expression of BIRC5 is correlated with loss of
specific chromosomal regions in breast carcinomas. Genes,
Chromosomes Cancer. 47:299–308. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Zhou XL and Wang M: Expression levels of
survivin, Bcl-2 and KAI1 proteins in cervical cancer and their
correlation with metastasis. Genet Mol Res. 14:17059–17067. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Ferrandina G, Legge F, Martinelli E,
Ranelletti FO, Zannoni GF, Lauriola L, Gessi M, Gallotta V and
Scambia G: Survivin expression in ovarian cancer and its
correlation with clinico-pathological, surgical and
apoptosis-related parameters. Br J Cancer. 92:271–277. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Zahedi P, Aminimoghaddam S, Sayahpour FA,
Haghpanah V, Amiri P, Fereidoni F, Mahrampour E, Larijani B,
Tavakkoly-Bazzaz J and Amoli MM: Association of survivin gene
polymorphism with endometrial cancer. Int J Gynecol Cancer.
22:35–37. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Wellenhofer A and Brustmann H: Expression
of human telomerase reverse transcriptase in vulvar intraepithelial
neoplasia and squamous cell carcinoma: An immunohistochemical study
with survivin and p53. Arch Pathol Lab Med. 136:1359–1365. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Chen X, Thiaville MM, Chen L, Stoeck A,
Xuan J, Gao M, Shih IeM and Wang TL: Defining NOTCH3 target genes
in ovarian cancer. Cancer Res. 72:2294–2303. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Smalley M, Piggott L and Clarkson R:
Breast cancer stem cells: Obstacles to therapy. Cancer Lett.
338:57–62. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Luo Y, Wu Y, Peng Y, Liu X, Bie J and Li
S: Systematic analysis to identify a key role of CDK1 in mediating
gene interaction networks in cervical cancer development. Ir J Med
Sci. 185:231–239. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Risinger JI, Allard J, Chandran U, Day R,
Chandramouli GV, Miller C, Zahn C, Oliver J, Litzi T, Marcus C, et
al: Gene expression analysis of early stage endometrial cancers
reveals unique transcripts associated with grade and histology but
not depth of invasion. Front Oncol. 3:1392013. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Katayama H and Sen S: Aurora kinase
inhibitors as anticancer molecules. Biochim Biophy Acta.
1799:829–839. 2010. View Article : Google Scholar
|
|
64
|
Lai CH, Tseng JT, Lee YC, Chen YJ, Lee JC,
Lin BW, Huang TC, Liu YW, Leu TH, Liu YW, et al: Translational
up-regulation of Aurora-A in EGFR-overexpressed cancer. J Cell Mol
Med. 14:1520–1531. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Sun C, Chan F, Briassouli P and
Linardopoulos S: Aurora kinase inhibition downregulates NF-κB and
sensitises tumour cells to chemotherapeutic agents. Biochem Biophys
Res Commun. 352:220–225. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Lim KH, Brady DC, Kashatus DF, Ancrile BB,
Der CJ, Cox AD and Counter CM: Aurora-A phosphorylates, activates
and relocalizes the small GTPase RalA. Mol Cell Biol. 30:508–523.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Nilsson JA and Cleveland JL: Myc pathways
provoking cell suicide and cancer. Oncogene. 22:9007–9021. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Cole MD: MYC association with cancer risk
and a new model of MYC-mediated repression. Cold Spring Harb
Perspect Med. 4:a0143162014. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
D'Assoro AB, Liu T, Quatraro C, Amato A,
Opyrchal M, Leontovich A, Ikeda Y, Ohmine S, Lingle W, Suman V, et
al: The mitotic kinase Aurora-a promotes distant metastases by
inducing epithelial-to-mesenchymal transition in ERα(+) breast
cancer cells. Oncogene. 33:599–610. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Opyrchal M, Sangameswaran KD, Khoury T,
Boland P, Galanis E and Haddad TC: Aurora Kinase inhibitors in
breast cancer treatment. American Journal of Hematology/Oncology.
11:2015.
|
|
71
|
Zhang L and Zhang S: ZM447439, the Aurora
kinase B inhibitor, suppresses the growth of cervical cancer SiHa
cells and enhances the chemosensitivity to cisplatin. J Obstet
Gynaecol Res. 37:591–600. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Do TV, Xiao F, Bickel LE, Klein-Szanto AJ,
Pathak HB, Hua X, Howe C, O'Brien SW, Maglaty M, Ecsedy JA, et al:
Aurora kinase A mediates epithelial ovarian cancer cell migration
and adhesion. Oncogene. 33:539–549. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Umene K, Yanokura M, Banno K, Irie H,
Adachi M, Iida M, Nakamura K, Nogami Y, Masuda K, Kobayashi Y, et
al: Aurora kinase A has a significant role as a therapeutic target
and clinical biomarker in endometrial cancer. Int J Oncol.
46:1498–1506. 2015. View Article : Google Scholar : PubMed/NCBI
|