Introduction
Lung cancer is the most common type of cancer, with
high mortality rates worldwide (1);
the incidence in the Czech Republic was 86.9 cases in men and 38.0
in women per 100,000 people in 2011 (2). Approximately 85% of all lung cancer
cases are non-small cell lung cancer (NSCLC), which includes two
major histological subtypes: Squamous cell carcinoma (SCC) and
adenocarcinoma. SCC represents ~25–30% of cases of NSCLC (3). The prognosis for patients with advanced
SCC is poorer than that of those with adenocarcinoma (4).
Chemotherapy is an essential modality of palliative
treatment for inoperable SCC at advanced stages. The response rate
to chemotherapy varies widely from patient to patient; therefore,
it is of interest to find biomarkers that predict the effect of
cytostatic therapeutics. The resistance of cancer cells to
chemotherapy can be caused by the increased export of anticancer
drugs out of the cells, improved DNA repair ability or apoptosis
resistance (5). The expression of
genes participating in these processes is regulated by the microRNA
(miRNA/miR) network. miRNAs are small non-coding RNA molecules of
~22 nucleotides that participate in the post-transcriptional
regulation of gene expression (6).
The human genome encodes >2,500 miRNAs (7), which target ~60% of mammalian genes and
are abundant in a number of human cell types (8) (see miRNA database available online at
www.mirbase.org).
The aim of the present study was to evaluate the
association of the expression of miRNAs involved in the processes
resulting in chemotherapy resistance with the overall survival (OS)
time of patients with advanced SCC receiving palliative care. All
patients in the cohort of the present study received palliative
chemotherapy based on platinum derivatives (cisplatin or
carboplatin) in combination with paclitaxel or gemcitabine.
On the basis of previously published literature, the
present study focused on miRNAs whose effect on the processes
involved in chemotherapy resistance may be expected (miR-15b,
miR-21, miR-27a, miR-34a, miR-99a, miR-106a, miR-107, miR-143,
miR-150, miR-192, miR-193, miR-211, miR-218, miR-221, miR-224,
miR-342 and miR-375). A list of the main characteristics of miRNAs
of interest, including references, is included in Table I.
 | Table I.Analyzed miRNAs and their involvement
in pathogenesis and treatment of NSCLC. |
Table I.
Analyzed miRNAs and their involvement
in pathogenesis and treatment of NSCLC.
| Symbol | miRBase accession
no. | Cat. no. 4427975
assay ID | Relation to
NSCLC | (Refs.) |
|---|
| miR-15b | MIMAT0000417 | 000390 | Regulates cisplatin
resistance and metastasis by targeting PEBP4 in lung adenocarcinoma
cells | (35) |
| miR-21 | MIMAT0000076 | 000397 | Regulates NSCLC
cell invasion and chemo-sensitivity through SMAD7 | (36) |
| miR-27a | MIMAT0000084 | 000408 | Higher expression
levels in advanced NSCLC patients resistant to EGFR-TKI | (37) |
| miR-34a | MIMAT0000255 | 000426 | Sensitizes lung
cancer cells to cisplatin via p53/miR-34a/MYCN axis | (38) |
| miR-99a-3p | MIMAT0004511 | 002141 | Promotes
proliferation, migration and invasion of NSCLC cell lines | (39) |
| miR-106a | MIMAT0000103 | 000578 | Confers cisplatin
resistance in non-small cell lung cancer A549 cells | (40) |
| miR-107 | MIMAT0000104 | 000443 | Regulates cisplatin
chemosensitivity of A549 non small cell lung cancer cell lines by
targeting cyclin dependent kinase 8 | (41) |
| miR-143 | MIMAT0000435 | 002249 | Regulates cell
apoptosis in lung cancer by targeting PKCε | (42) |
| miR-150 | MIMAT0000451 | 000473 | Downregulation
induces cell proliferation inhibition and apoptosis in NSCLC by
targeting BAK1 | (43) |
| miR-192 | MIMAT0000222 | 000491 | Regulates
chemo-resistance of lung adenocarcinoma for gemcitabine and
cisplatin combined therapy by targeting Bcl-2 | (44) |
| miR-193a-3p | MIMAT0000459 | 002250 | Suppresses the
metastasis of NSCLC by downregulating the ERBB4/PIK3R3/mTOR/S6K2
signaling pathway | (45) |
| miR-211 | MIMAT0000268 | 000514 | Promotes NSCLC
proliferation by targeting SRCIN1 | (46) |
| miR-218 | MIMAT0000275 | 000521 | Regulates cisplatin
chemosensitivity in NSCLC by targeting RUNX2 | (47) |
| miR-221 | MIMAT0000278 | 000524 | Overexpressed in
aggressive NSCLC and regulates TRAIL resistance through PTEN and
TIMP3 | (48) |
| miR-224 | MIMAT0000281 | 002099 | Is implicated in
lung cancer pathogenesis through targeting caspase-3 and
caspase-7 | (19) |
| miR-342-3p | MIMAT0000753 | 002260 | Suppresses
proliferation and invasion of NSCLC by targeting RAPB2 | (21) |
| miR-375 | MIMAT0000728 | 000564 | Predictive for
response for non-small cell lung cancer treated with
cisplatin-vinorelbine A | (49) |
| RNU6B | – | 001093 | Reference gene | (50) |
Patients and methods
Ethics statement
This study was approved by the Ethics Committee of
the University Hospital in Pilsen (Pilsen, Czech Republic). Written
informed consent was obtained from all the subjects. Anonymized
data were used to conduct the present study.
Patients
The present study was retrospective. The study group
consisted of 81 patients with late-stage (3B or 4) and the SCC
histological subtype with an Eastern Cooperative Oncology Group
(ECOG) performance status (PS) of 0–2 of NSCLC treated between
January 2000 and June 2014 at the Department of Pneumology and
Phthisiology of the University Hospital in Pilsen. Stage of disease
was determined using the TNM (Tumor Nodus Metastasis) system of the
International Union Against Cancer (IUCC; 7th edition) (9). The median patient age was 62.4 years
(range, 32.7–79.3 years), and there were 74 males and 7 females.
All patients underwent palliative chemotherapy using platinum
derivatives in combination with paclitaxel or gemcitabine. The use
of sequential radiotherapy was permitted for patients with stage 3B
disease; patients with concurrent radiotherapy were excluded from
the present study. In certain patients with stage 3B disease,
radiotherapy was not indicated due to poor PS. The exclusion
criteria for entering the study were >80 years of age, other
malignancy and high cardiopulmonary risk. Clinicopathological data,
including age at the time of diagnosis, smoking status, clinical
disease stage, radiotherapy status and chemotherapy regimen, are
listed in Table II.
 | Table II.Clinicopathological characteristics
of patients with squamous cell carcinoma of the lung (n=81). |
Table II.
Clinicopathological characteristics
of patients with squamous cell carcinoma of the lung (n=81).
| Characteristic | Patients, n
(%) |
|---|
| Sex |
|
|
Male | 74 (91.4) |
|
Female | 7 (8.6) |
| Age, years |
|
|
<55 | 11 (13.6) |
|
55–65 | 41 (50.6) |
|
>65 | 29 (35.8) |
| Smoking status |
|
|
Non-smoker | 0 (0) |
|
Ex-smoker | 42 (51.9) |
|
Smoker | 39 (48.1) |
| Clinical stage |
|
| 3B | 42 (51.9) |
| 4 | 39 (48.1) |
| Eastern Cooperative
Oncology |
|
| Group performance
status |
|
| 0 | 2 (2.5) |
| 1 | 58 (71.6) |
| 2 | 18 (22.2) |
| 3 | 3 (3.7) |
| Radiotherapy |
|
|
Yes | 25 (30.9) |
| No | 56 (69.1) |
| Chemotherapy |
|
|
Paclitaxel and
carboplatin | 35 (43.2) |
|
Gemcitabine and cisplatin | 46 (56.8) |
Tissue samples and RNA isolation
Biopsy tissue samples were obtained using
bronchoscopy for diagnostic purposes prior to chemotherapy and were
processed by standard laboratory techniques at the Department of
Pathology of the University Hospital in Pilsen. Formalin-fixed
paraffin-embedded (FFPE) tissue samples were stored at room
temperature until use. Paraffinized sections were stained with
hematoxylin and eosin, microscopically verified by pathologists and
examined to identify sites with cancer cells for macrodissection.
Total RNA (including miRNA) was extracted from 15-µm thick FFPE
sections following the macrodissection of tumor tissue using the
miRNeasy FFPE kit (Qiagen, Hilden, Germany) as described previously
(10).
Quantitative estimation of microRNA
expression
A quantitative estimation of 17 selected miRNAs
(Table I) was performed by the
reverse transcription-quantitative polymerase chain reaction
(RT-qPCR) method using TaqMan® MicroRNA assays (Applied
Biosystems; Thermo Fisher Scientific, Inc., Waltham, MA, USA) in
technical duplicates on the Stratagene Mx3005P apparatus (Agilent
Technologies, Santa Clara, CA, USA) according to the manufacturer's
protocol. The two-step protocol included reverse transcription with
a miRNA-specific primer, followed by qPCR with TaqMan®
probes. Briefly, 5 ng of RNA was reverse transcribed in a 20-µl
reaction containing 2.5 µl of primers specific to particular miRNA
(Applied Biosystems; Thermo Fisher Scientific, Inc.). The 20-µl PCR
reactions included 2.5 µl of RT product. The reactions were
incubated in 96-well plates at 95°C for 15 min and then followed by
48 cycles of 95°C for 15 sec and 60°C for 60 sec. The
2−ΔΔCq method was used for the quantification of qPCR
data, as described previously (11);
the expression was normalized to RNU6B (U6snRNA). Details of all
kits for estimation of all miRNAs and RNU6B are included in
Table I.
Statistical analysis
SAS version 9.3 statistical software (SAS Institute,
Inc., Cary, NC, USA) was used to perform all statistical
calculations. Although the group of patients in the present study
was as homogenous as possible (SCC subtype of NSCLC, stages 3B and
4), certain patients underwent radiotherapy, which may have caused
confounders. The potential effect of treatment inconsistency was
mitigated by evaluating miRNA expression in the subgroups of
patients.
The evaluation of prognostic significance (the
association between markers and time to recurrence) was performed
as a univariate analysis of maximum likelihood estimates using the
Cox regression hazard model. For markers significant in the Cox
model, an optimal cut off was identified. There is no standard
method for biomarker cut-off determination to split continuous
variables into two groups; the simplest approach used in
exploratory studies is to set the median as a cut-off value.
However, in the case of unequal distribution of events in the
studied group, this approach is far from optimal. The present study
used continuous search for the cut-off value by searching for the
lowest P-value of the log-rank test, as described previously
(12,13).
miRNAs identified as significant by univariate
analysis were incorporated into a multivariate analysis and the
Kaplan-Meier survival distribution functions were generated for
combinations of miRNA expression levels (clustering) with the
cut-off values from univariate analysis and later, the median.
P<0.05 was considered to indicate a statistically significant
difference. DIANA-TarBase v7.0 and DIANA-miRPath v3.0 bioinformatic
tools were used to identify overlapped target genes of the miRNAs
of interest (14,15).
Results
Effect of treatment modalities on OS
time
Prior to the analysis of the association between
gene expression and OS, the outcomes for subgroups of patients
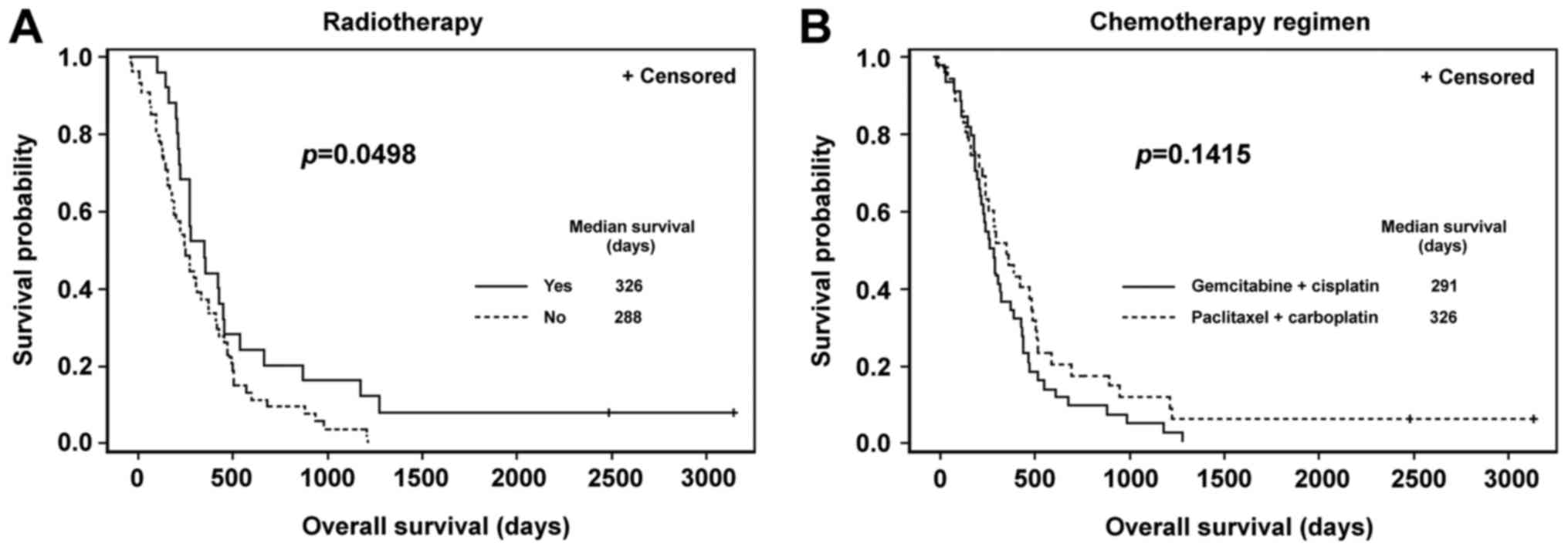
undergoing different treatment were compared. A significantly
longer OS time was identified in the subgroup of patients who
underwent chemotherapy combined with radiotherapy in comparison
with patients who underwent chemotherapy alone (P=0.0498; Fig. 1A). There were no significant
differences in OS between subgroups of patients with different
chemotherapy regimens (Fig. 1B).
Association of miRNA expression with
OS time
The Cox regression hazard model was used to
determine the association between the levels of miRNA expression
with OS time. From the 17 miRNAs of interest, in the subgroup of
smokers, the low expression of miR-342 (P=0.0500) and high
expression level of miR-34a and miR-224 (P=0.0338 and P=0.0400,
respectively) were associated with a shorter OS time. High
expression levels of miR-34a were associated with shorter OS time
in the subgroup of patients treated with platinum derivate-based
chemotherapy in combination with gemcitabine (P=0.0364). High
expression levels of miR-224 were associated with shorter OS time
in the subgroup of patients who underwent chemotherapy combined
with radiotherapy (P=0.0250).
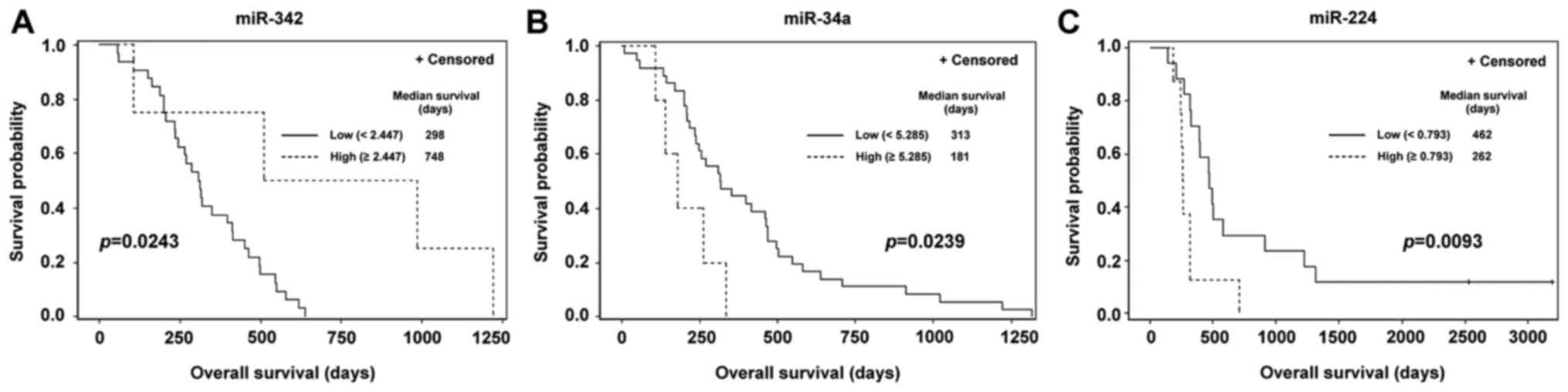
For the statistically significant miRNA markers,
optimal cut-off values were identified and Kaplan-Meier survival
distribution functions for OS were generated. Statistically
significant differences in OS time between subgroups with marker
expression levels below and above the cut-off value were obtained
for miR-342 in the subgroup of smokers (P=0.0243; Fig. 2A), miR-34a in a subgroup of patients
that were treated with gemcitabine in chemotherapy regimen
(P=0.0239; Fig. 2B) and miR-224 in
the subgroup of patients that underwent chemotherapy combined with
radiotherapy (P=0.0093; Fig. 2C).
Statistical values obtained from the Kaplan-Meier analyses are
summarized in Table III.
 | Table III.Association between the level of miRs
and overall survival time as determined by Kaplan-Meier
estimation. |
Table III.
Association between the level of miRs
and overall survival time as determined by Kaplan-Meier
estimation.
|
| Below cut-off | Above cut-off |
|
|---|
|
|
|
|
|
|---|
| Patient group | Treatment | Marker | Patients, n | Cut-off | n | Median, days | n | Median, days | P-value |
|---|
| Smokers only | Chemotherapy | miR-342 | 36 | 2.447 | 32 | 298 | 4 | 748 | 0.0243 |
| Smokers and
ex-smokers | Gemcitabine and
cisplatin | miR-34a | 41 | 5.285 | 36 | 313 | 5 | 181 | 0.0239 |
| Smokers and
ex-smokers | Chemotherapy and
radiotherapy | miR-224 | 25 | 0.793 | 17 | 462 | 8 | 262 | 0.0093 |
miRNAs associated with OS (miR-34a, −224 and −342)
were the subject of the subsequent cluster analysis. Pairs of these
miRNAs (miR-34a and −224, miR-34a and −342, and miR-224 and −342)
were analyzed for their association with OS time. For each pair of
miRNAs, patients were stratified into groups according to the miRNA
expression being above (high) or below (low) the cut-off value:
Group A (high miR-224 and high miR-342); group B (high miR-224 and
low miR-342); group C (low miR-224 and high miR-342); and group D
(low miR-224 and low miR-342). Initially, the cut-off value
obtained from univariate analysis was used also for cluster
analysis; however, this led to a highly disproportional
distribution of patients among subgroups. Therefore, a median was
used as a cut off value for cluster analysis.
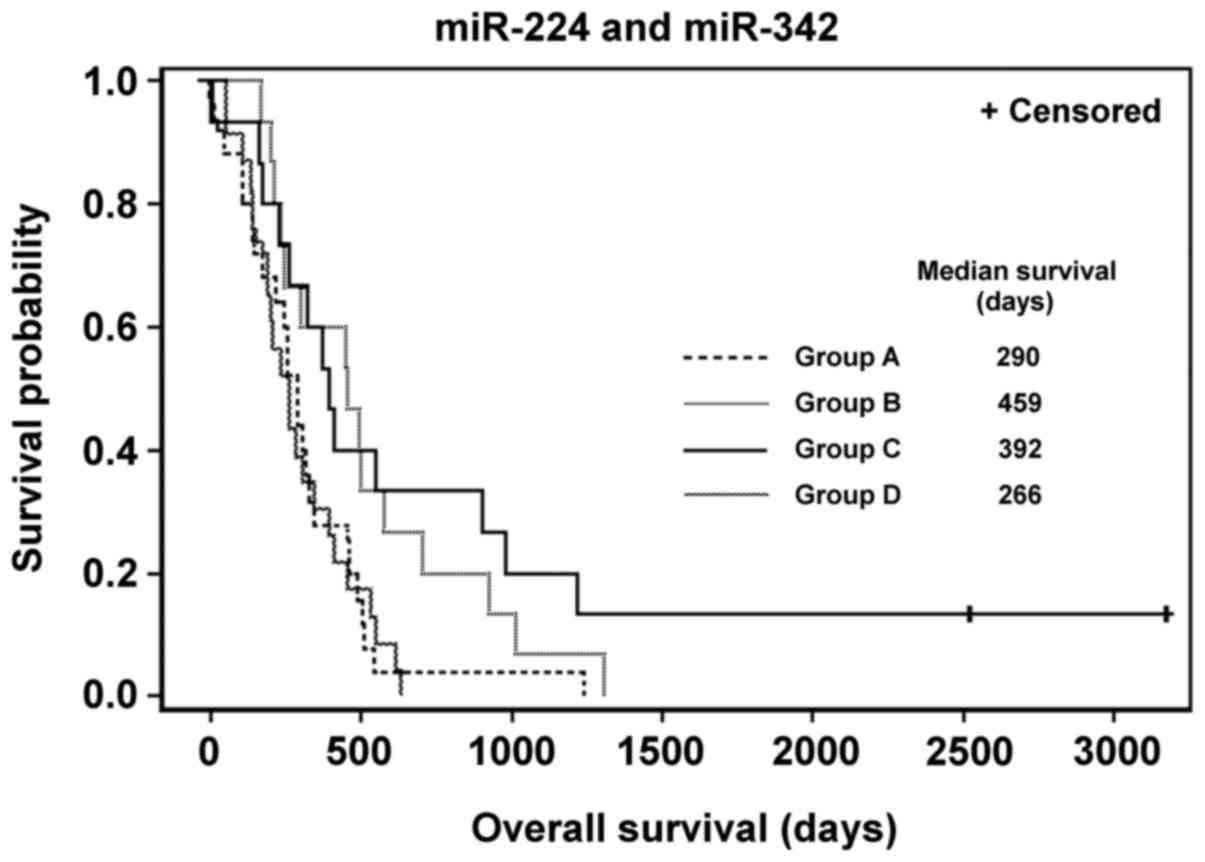
Fig. 3 demonstrates
the Kaplan-Meier survival distribution functions of patients
stratified into groups according to the expression of miR-224 and
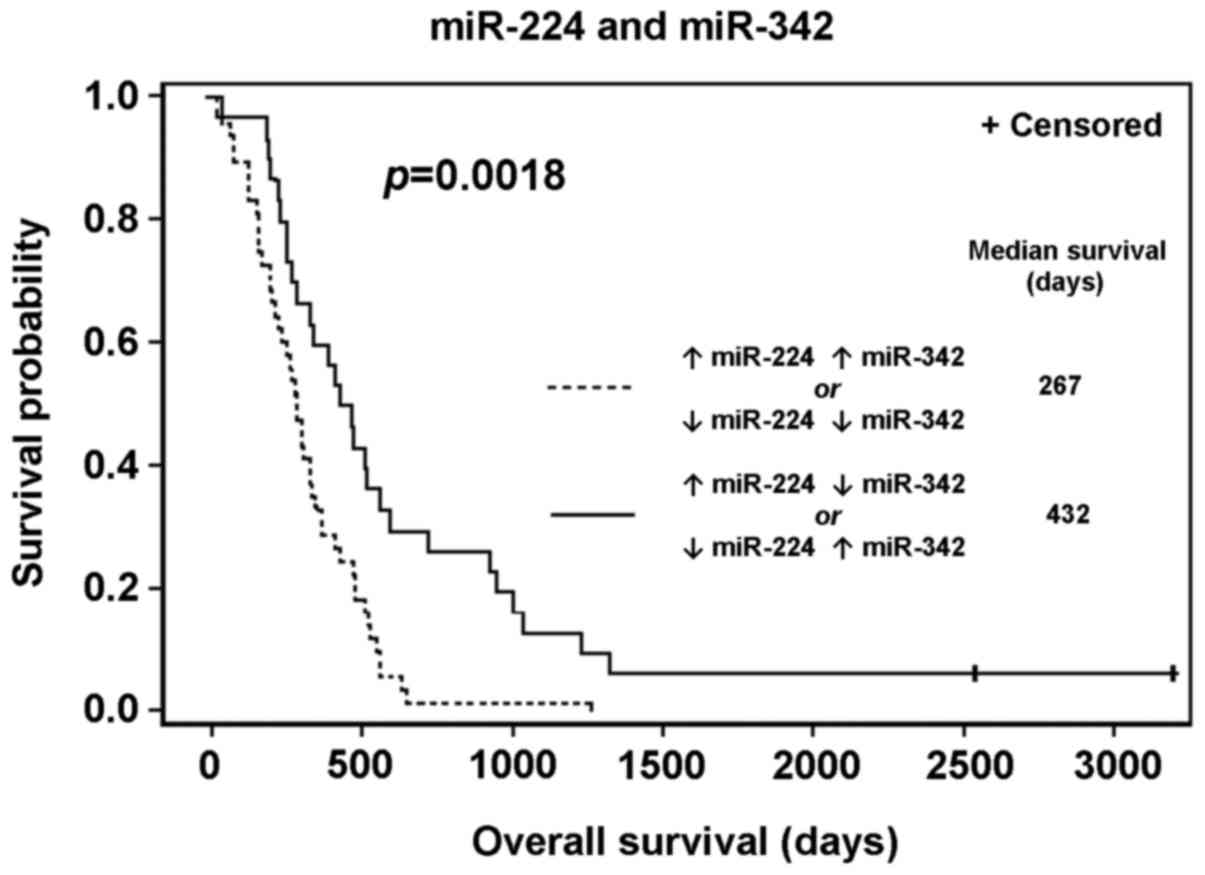
miR-342. There are two pairs of groups with similar OS
distributions; Fig. 4 includes a
comparison of the OS of two groups of patients created by combining
the groups from Fig. 3 with similar
survival outcomes (group A/D vs. group B/C). There was a
significant difference in survival between these groups (P=0.0018),
as detailed in Table IV. The same
approach was used to analyze the other pairs of miRNAs; however, no
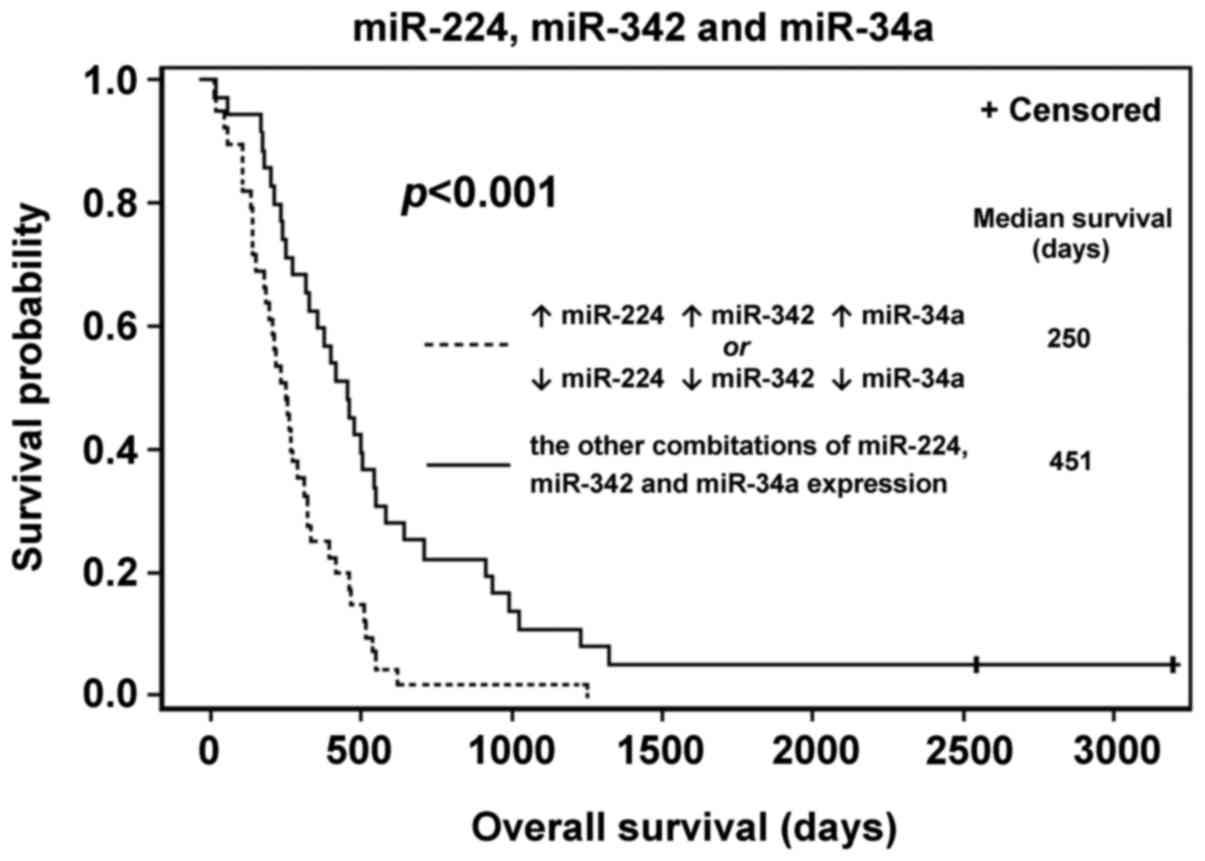
significance was identified. All three miRNAs were analyzed
together in the same manner (miR-34a, −224 and −342). Patterns of
expression associating patients with significantly shorter survival
times were identified (Fig. 5;
Table IV).
 | Table IV.Association between combinations of
miRs and OS (Kaplan-Meier estimation). |
Table IV.
Association between combinations of
miRs and OS (Kaplan-Meier estimation).
| Expression
pattern | Patients, n | Median OS,
days |
|---|
| miR-342 and
−224 |
|
|
| High
miR-224 and −342, or low miR-224 and −342 | 48 | 267 |
| High
miR-224 and low miR-342, or low miR-224 and high miR-342 | 30 | 432 |
| miR-342, −224 and
−34a |
|
|
| High
miR-224, −342 and −34a, or low miR-224, −342 and −34a | 39 | 250 |
| Other
combinations | 35 | 451 |
Identification of potential target
genes for miR-34a, −224 and −342
Using DIANA-TarBase v7.0 and DIANA-miRPath v3.0
bioinformatic tools (11,12), 6 overlapping target genes with
P<0.05 were identified between miR-34a, −224 and −342. These
genes, including GNAS complex locus (GNAS), insulin like growth
factor 1 receptor (IGF1R), cyclin D1 (CCND1), cyclin G2 (CCNG2),
serpin family E member 1 (SERPINE1) and ribonucleotide reductase
regulatory subunit M2 (RRM2), are associated with cell cycle
regulation, p53 signaling and DNA repair.
Discussion
miRNAs may have the potential to become accurate,
easily measurable biomarkers, with features convenient for
diagnostic testing methods, including stability in FFPE tissue
blocks, blood, and potentially, other bodily fluids (16). The present study focused on patients
with the NSCLC SCC subtype with an advanced-stage SCC. The included
patients were unable to undergo surgical resection and received
palliative treatment only. For these patients, there were multiple
treatment modalities. The main clinical concern in such cases is
deciding which therapeutic regimen is indicated. However, in the
group of patients in the present study, it was only possible to
analyze the potential predictors for the treatment response to
platinum base derivatives in combination with either paclitaxel or
gemcitabine, with or without the application of radiotherapy.
Initially, the present study focused on a univariate
analysis of the association between miRNA expression and OS time.
Subsequently, a multivariate analysis was performed that included
the miRNAs that had been identified to exhibit associations with
OS. On the basis of the results of the present study, we
hypothesize that the effect of a single miRNA may depend on the
level of expression of other members of the miRNA network, to be
further discussed.
Higher levels of miR-224 indicated shorter OS times
for patients with chemotherapy combined with radiotherapy in the
present study. Cui et al (17)
reported that miR-224 expression was significantly upregulated in
NSCLC tissues and suggested it performed its oncogenic role in lung
cancer pathogenesis through targeting caspase-3 and −7. Wang et
al (18) identified through
microarray analysis that miR-224 expression was upregulated in
cisplatin-resistant cell lines, and demonstrated that miR-224 could
promote cisplatin resistance via regulating the G1/S
cell cycle transition and apoptosis by targeting p21. These
findings indicated the association of miR-224 with the effect of
chemotherapy based on DNA damage, and its potential as a predictor
for the response to treatment. However, Zhu et al (19) reported that miR-224 expression levels
were downregulated in NSCLC compared with non-cancerous lung
tissue. These authors also observed that decreased miR-224
expression was significantly associated with lymph node metastasis,
an advanced tumor-node-metastasis stage and a reduced OS time
(19). Furthermore, Wang et al
(20) recently identified that
miR-224 was significantly upregulated in NSCLC tissues and
hypothesized that miR-224 expression promotes NSCLC cell
proliferation by downregulating Ras association domain family
member 8 expression; the inconsistency of these studies will be
discussed in the following paragraph.
In the present study, low levels of miR-342
indicated a poorer outcome in patients with a history of smoking,
independent of treatment modality. Xie et al (21) demonstrated that miR-342 was
downregulated in NSCLC and acted as a tumor suppressor through the
repression of RAP2B, member of RAS oncogene family. Similarly, Tai
et al (22) identified that
miR-342 was capable of indirectly regulating MYC activity via the
direct repression of E2F transcription factor 1. Takahashi et
al (23) investigated how
cigarette smoking altered plasma miRNA profiles; they identified
that there was a decrease in plasma miR-342 in subjects who quit
smoking, compared with smokers.
miR-34a is a member of the miR-34 family that is
associated with the p53 pathway, and is implicated in cell
death/survival signaling (24). The
miR-34 family is transcriptionally activated by p53; in turn, p53
is a direct miR-34a target. However, the effect of miR-34a on p53
depends on the cellular context (25). miR-34a can have also a positive effect
on p53 transcriptional activity and protein stability by targeting
multiple p53 inhibitor genes (including MDM4, p53 regulator,
sirtuin 1, metastasis associated 1 family member 2, histone
deacetylase 1 and YY1 transcription factor) (26). A previous study identified that
miR-34a inhibits cell proliferation (27). Expression of the miR-34 family was
downregulated in tumor tissue compared with normal tissue, and low
levels of miR-34a expression were associated with a higher
probability of relapse in surgically resected NSCLC (28). However, higher levels of circulating
miR-34a were observed in patients with NSCLC compared with healthy
controls (29). Higher levels of
miR-34a indicated a shorter OS time in patients receiving
palliative platinum derivate-based chemotherapy in combination with
gemcitabine in the present study.
Multivariate analysis was performed with the miRNAs
(miR-34a, −224 and −342) that were identified as associated with
OS. The most notable finding was that patients with the high
expression of miR-224 and −342 exhibited similar outcomes to those
with low expression of miR-224 and −342, which was significantly
shorter than that of patients with high expression of either
miR-224 or miR-342 and the low expression of the other (Figs. 3 and 4).
We hypothesize that the effect of a single miRNA is
dependent on the level of expression of the other members of the
miRNA network. It has been established that an miRNA can have a
predominantly oncogenic role in one type of cancer and a tumor
suppressive role in another; for instance, miR-224 was identified
to be a tumor suppressor in prostate cancer (30), whereas in other types of malignancy,
including gastric (11,31) and colorectal cancer (32), an oncogenic role for miR-224 was
described. The ambiguous role of miR-224 was also observed within
the SCC histological subtype of NSCLC in the present study. Tumor
progression occurs as a result of the dysregulation of a number of
protein-coding genes and epigenetic processes, including the
deregulation of a number of miRNAs. Therefore, to understand the
role of one particular miRNA, it is necessary to determine the
levels of the other ‘co-players’. In the present study, OS time was
influenced by the mutual association of miR-224 and −342. The high
level of miR-224 can be associated with adverse or favorable
outcomes, depending on the simultaneous level of miR-342. These
findings could explain the inconsistent results of previously
published studies on miR-224 expression in NSCLC. In 2014, Zhu
et al (19) reported that
miR-224 was significantly downregulated in NSCLC and that a
decrease in miR-224 expression was significantly associated with
shorter OS time (19). Also in NSCLC,
Cui et al (33) identified
that miR-224 was significantly upregulated, with the increased
expression of miR-224 promoting cell migration, invasion, and
proliferation. As aforementioned, the present study also identified
that the high expression levels of miR-224 were associated with
shorter OS time in one subgroup of patients, specifically those who
underwent chemotherapy combined with radiotherapy.
Using bioinformatic tools, the present study
identified overlapping experimentally validated target genes for
miR-34a, −224 and −342. Notably, all overlapping target genes
identified in the present study (GNAS, IGF1R, CCND1, CCNG2,
SERPINE1, and RRM2) are involved in processes associated with
carcinogenesis, including cell cycle regulation, p53 signaling and
DNA repair. This may explain the complicated mutual dependency of
those miRNAs in relation to tumor progression and the effectiveness
of treatment. We hypothesize that these molecules could be involved
in competing endogenous RNA crosstalk, where RNA transcripts
co-regulate each other by competing for shared miRNAs, thereby
titrating miRNA availability (34).
However, one limitation of the present study is the absence of
immunoprecipitation data and reporter assays, which are methods
that may confirm the interactions among the set of 3 miRNAs and 6
target genes. Nevertheless, the results of the present study may
provide a stimulus for further research in this area.
With cluster analysis, novel associations between
miR-34a, −224 and −342 that affected patient survival time were
identified in the present study. The result may demonstrate that
the effort to find a particular miRNA as a perfect marker for a
particular event may be fruitless due to the complex interactions
between RNA transcripts. In order to understand all aspects of the
effect of miRNAs on the regulation of gene expression in cancer and
their associations with phenotype and treatment outcome, miRNA
profiling and deep bioinformatic analysis will be necessary. Only
this approach can facilitate the future application of miRNAs in
clinical practice. miRNAs can generally be assessed with more
precision and ease than the mRNAs of coding genes, as miRNA
analysis is less demanding in terms of the quality and quantity of
isolated RNA, features that may be problematic in RNA samples
extracted from FFPE tissue (16).
FFPE tissue samples are routinely taken and analyzed during
standard lung cancer management, which is why miRNAs may become
clinically applicable predictors of the effectiveness of palliative
treatment in patients with lung cancer. Nevertheless, the findings
of the present study demonstrated that, due to the complex network
of interactions, this objective could be achieved by combining more
markers together rather than by using individual miRNAs. On the
basis of the results of the current study, miR-224, −342 and −34a
could be members of this panel of predictors of treatment
efficacy.
Acknowledgements
The present study was supported by grants from the
Ministry of Health of the Czech Republic Conceptual Development of
Research Organization (grant nos. 00669806 and AZV 17-30748A), and
by the project of Faculty of Medicine in Pilsen (grant no.
SVV-2017-260693).
References
|
1
|
Torre LA, Siegel RL and Jemal A: Lung
Cancer Statistics. Adv Exp Med Biol. 893:1–19. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Institute of Health Information and
Statistics of the Czech Republic, . Czech Health Statistics Year
Book 2013UZIS CR. Prague: 2014
|
|
3
|
Travis WD: Pathology of lung cancer. Clin
Chest Med. 32:669–692. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Socinski MA, Obasaju C, Gandara D, Hirsch
FR, Bonomi P, Bunn P, Kim ES, Langer CJ, Natale RB, Novello S, et
al: Clinicopathologic features of advanced squamous NSCLC. J Thorac
Oncol. 11:1411–1422. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Olaussen KA and Postel-Vinay S: Predictors
of chemotherapy efficacy in non-small-cell lung cancer: A
challenging landscape. Ann Oncol. 27:2004–2016. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Inamura K and Ishikawa Y: MicroRNA in lung
cancer: Novel biomarkers and potential tools for treatment. J Clin
Med. 5:pii: E362016. View Article : Google Scholar
|
|
7
|
Griffiths-Jones S, Saini HK, van Dongen S
and Enright AJ: miRBase: Tools for microRNA genomics. Nucleic Acids
Res. 36(Database Issue): D154–D158. 2008.PubMed/NCBI
|
|
8
|
Friedman RC, Farh KK, Burge CB and Bartel
DP: Most mammalian mRNAs are conserved targets of microRNAs. Genome
Res. 19:92–105. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Sobin LH, Gospodarowicz MK and Wittekind
Ch: TNM Classification of Malignant Tumours. 7th. Wiley-Blackwell;
Chichester: 2010
|
|
10
|
Kalfert D, Pesta M, Kulda V, Topolcan O,
Ryska A, Celakovsky P, Laco J and Ludvikova M: MicroRNA profile in
site-specific head and neck squamous cell cancer. Anticancer Res.
35:2455–2463. 2015.PubMed/NCBI
|
|
11
|
Smid D, Kulda V, Srbecka K, Kubackova D,
Dolezal J, Daum O, Kucera R, Topolcan O, Treska V, Skalicky T and
Pesta M: Tissue microRNAs as predictive markers for gastric cancer
patients undergoing palliative chemotherapy. Int J Oncol.
48:2693–2703. 2016.PubMed/NCBI
|
|
12
|
Mazumdar M and Glassman JR: Categorizing a
prognostic variable: Review of methods, code for easy
implementation and applications to decision-making about cancer
treatments. Stat Med. 19:113–132. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Faraggi D and Simon R: A simulation study
of cross-validation for selecting an optimal cutpoint in univariate
survival analysis. Stat Med. 15:2203–2213. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Vlachos IS, Paraskevopoulou MD, Karagkouni
D, Georgakilas G, Vergoulis T, Kanellos I, Anastasopoulos IL,
Maniou S, Karathanou K, Kalfakakou D, et al: DIANA-TarBase v7.0:
Indexing more than half a million experimentally supported
miRNA:mRNA interactions. Nucleic Acids Res. 43(Database Issue):
D153–D159. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Vlachos IS, Zagganas K, Paraskevopoulou
MD, Georgakilas G, Karagkouni D, Vergoulis T, Dalamagas T and
Hatzigeorgiou AG: DIANA-miRPath v3.0: Deciphering microRNA function
with experimental support. Nucleic Acids Res. 43:W460–W466. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Khan J, Lieberman JA and Lockwood CM:
Variability in, variability out: Best practice recommendations to
standardize pre-analytical variables in the detection of
circulating and tissue microRNAs. Clin Chem Lab Med. 55:608–621.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Cui R, Kim T, Fassan M, Meng W, Sun HL,
Jeon YJ, Vicentini C, Tili E, Peng Y and Scarpa A: MicroRNA-224 is
implicated in lung cancer pathogenesis through targeting caspase-3
and caspase-7. Oncotarget. 6:21802–21815. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Wang H, Zhu LJ, Yang YC, Wang ZX and Wang
R: MiR-224 promotes the chemoresistance of human lung
adenocarcinoma cells to cisplatin via regulating G1/S transition
and apoptosis by targeting p21(WAF1/CIP1). Br J Cancer.
111:339–354. 2014. View Article : Google Scholar
|
|
19
|
Zhu D, Chen H, Yang X, Chen W, Wang L, Xu
J and Yu L: Decreased microRNA-224 and its clinical significance in
non-small cell lung cancer patients. Diagn Pathol. 9:1982014.
View Article : Google Scholar
|
|
20
|
Wang L, Liu W, Zhang YP and Huang XR: The
miR-224 promotes non-small cell lung cancer cell proliferation by
directly targeting RASSF8. Eur Rev Med Pharmacol Sci. 21:3223–3231.
2017.
|
|
21
|
Xie X, Liu H, Wang M, Ding F, Xiao H, Hu
F, Hu R and Mei J: miR-342-3p targets RAP2B to suppress
proliferation and invasion of non-small cell lung cancer cells.
Tumour Biol. 36:5031–5038. 2015. View Article : Google Scholar
|
|
22
|
Tai MC, Kajino T, Nakatochi M, Arima C,
Shimada Y, Suzuki M, Miyoshi H, Yatabe Y, Yanagisawa K and
Takahashi T: miR-342-3p regulates MYC transcriptional activity via
direct repression of E2F1 in human lung cancer. Carcinogenesis.
36:1464–1473. 2015.
|
|
23
|
Takahashi K, Yokota SI, Tatsumi N, Fukami
T, Yokoi T and Nakajima M: Cigarette smoking substantially alters
plasma microRNA profiles in healthy subjects. Toxicol Appl
Pharmacol. 272:154–160. 2013. View Article : Google Scholar
|
|
24
|
Rokavec M, Li H, Jiang L and Hermeking H:
The p53/miR-34 axis in development and disease. J Mol Cell Biol.
6:214–230. 2014. View Article : Google Scholar
|
|
25
|
Okada N, Lin CP, Ribeiro MC, Biton A, Lai
G, He X, Bu P, Vogel H, Jablons DM, Keller AC, et al: A positive
feedback between p53 and miR-34 miRNAs mediates tumor suppression.
Genes Dev. 28:438–450. 2014. View Article : Google Scholar
|
|
26
|
Navarro F and Lieberman J: miR-34 and p53:
New insights into a complex functional relationship. PLoS One.
10:e01327672015. View Article : Google Scholar
|
|
27
|
Ma ZL, Hou PP, Li YL, Wang DT, Yuan TW,
Wei JL, Zhao BT, Lou JT, Zhao XT, Jin Y and Jin YX: MicroRNA-34a
inhibits the proliferation and promotes the apoptosis of non-small
cell lung cancer H1299 cell line by targeting TGFβR2. Tumour Biol.
36:2481–2490. 2015. View Article : Google Scholar
|
|
28
|
Gallardo E, Navarro A, Viñolas N, Marrades
RM, Diaz T, Gel B, Quera A, Bandres E, Garcia-Foncillas J, Ramirez
J and Monzo M: miR-34a as a prognostic marker of relapse in
surgically resected non-small-cell lung cancer. Carcinogenesis.
30:1903–1909. 2009. View Article : Google Scholar
|
|
29
|
Franchina T, Amodeo V, Bronte G, Savio G,
Ricciardi GR, Picciotto M, Russo A, Giordano A and Adamo V:
Circulating miR-22, miR-24 and miR-34a as novel predictive
biomarkers to pemetrexed-based chemotherapy in advanced non-small
cell lung cancer. J Cell Physiol. 229:97–99. 2014.
|
|
30
|
Goto Y, Nishikawa R, Kojima S, Chiyomaru
T, Enokida H, Inoguchi S, Kinoshita T, Fuse M, Sakamoto S, Nakagawa
M, et al: Tumour-suppressive microRNA-224 inhibits cancer cell
migration and invasion via targeting oncogenic TPD52 in prostate
cancer. FEBS Lett. 588:1973–1982. 2014. View Article : Google Scholar
|
|
31
|
Zhang Y, Li CF, Ma LJ, Ding M and Zhang B:
MicroRNA-224 aggrevates tumor growth and progression by targeting
mTOR in gastric cancer. Int J Oncol. 49:1068–1080. 2016.
|
|
32
|
Adamopoulos PG, Kontos CK, Rapti SM,
Papadopoulos IN and Scorilas A: miR-224 overexpression is a strong
and independent prognosticator of short-term relapse and poor
overall survival in colorectal adenocarcinoma. Int J Oncol.
46:849–859. 2015. View Article : Google Scholar
|
|
33
|
Cui R, Meng W, Sun HL, Kim T, Ye Z, Fassan
M, Jeon YJ, Li B, Vicentini C, Peng Y, et al: MicroRNA-224 promotes
tumor progression in nonsmall cell lung cancer. Proc Natl Acad Sci
USA. 112:pp. E4288–E4297. 2015; View Article : Google Scholar
|
|
34
|
Tay Y, Rinn J and Pandolfi PP: The
multilayered complexity of ceRNA crosstalk and competition. Nature.
505:344–352. 2014. View Article : Google Scholar
|
|
35
|
Zhao Z, Zhang L, Yao Q and Tao Z: miR-15b
regulates cisplatin resistance and metastasis by targeting PEBP4 in
human lung adenocarcinoma cells. Cancer Gene Ther. 22:108–114.
2015. View Article : Google Scholar
|
|
36
|
Lin L, Tu HB, Wu L, Liu M and Jiang GN:
MicroRNA-21 regulates non-small cell lung cancer cell invasion and
chemo-sensitivity through SMAD7. Cell Physiol Biochem.
38:2152–2162. 2016. View Article : Google Scholar
|
|
37
|
Wang S, Su X, Bai H, Zhao J, Duan J, An T,
Zhuo M, Wang Z, Wu M, Li Z, et al: Identification of plasma
microRNA profiles for primary resistance to EGFR-TKIs in advanced
non-small cell lung cancer (NSCLC) patients with EGFR activating
mutation. J Hematol Oncol. 8:1272015. View Article : Google Scholar
|
|
38
|
Song C, Lu P, Sun G, Yang L and Wang Z and
Wang Z: miR-34a sensitizes lung cancer cells to cisplatin via
p53/miR-34a/MYCN axis. Biochem Biophys Res Commun. 482:22–27. 2017.
View Article : Google Scholar
|
|
39
|
Chen C, Zhao Z, Liu Y and Mu D:
microRNA-99a is downregulated and promotes proliferation, migration
and invasion in non-small cell lung cancer A549 and H1299 cells.
Oncol Lett. 9:1128–1134. 2015.
|
|
40
|
Ma Y, Li X, Cheng S, Wei W and Li Y:
MicroRNA-106a confers cisplatin resistance in non-small cell lung
cancer A549 cells by targeting adenosine triphosphatase-binding
cassette A1. Mol Med Rep. 11:625–632. 2015. View Article : Google Scholar
|
|
41
|
Zhang Z, Zhang L, Yin ZY, Fan XL, Hu B,
Wang LQ and Zhang D: miR-107 regulates cisplatin chemosensitivity
of A549 non small cell lung cancer cell line by targeting cyclin
dependent kinase 8. Int J Clin Exp Pathol. 7:7236–7241. 2014.
|
|
42
|
Zhang N, Su Y and Xu L: Targeting PKCε by
miR-143 regulates cell apoptosis in lung cancer. FEBS Lett.
587:3661–3667. 2013. View Article : Google Scholar
|
|
43
|
Gu XY, Wang J, Luo YZ, Du Q, Li RR, Shi H
and Yu TP: Down-regulation of miR-150 induces cell proliferation
inhibition and apoptosis in non-small-cell lung cancer by targeting
BAK1 in vitro. Tumour Biol. 35:5287–5293. 2014. View Article : Google Scholar
|
|
44
|
Cao J, He Y, Liu HQ, Wang SB, Zhao BC and
Cheng YS: MicroRNA 192 regulates chemo-resistance of lung
adenocarcinoma for gemcitabine and cisplatin combined therapy by
targeting Bcl-2. Int J Clin Exp Med. 8:12397–12403. 2015.
|
|
45
|
Yu T, Li J, Yan M, Liu L, Lin H, Zhao F,
Sun L, Zhang Y, Cui Y, Zhang F, et al: MicroRNA-193a-3p and −5p
suppress the metastasis of human non-small-cell lung cancer by
downregulating the ERBB4/PIK3R3/mTOR/S6K2 signaling pathway.
Oncogene. 34:413–423. 2015. View Article : Google Scholar
|
|
46
|
Ye L, Wang H and Liu B: miR-211 promotes
non-small cell lung cancer proliferation by targeting SRCIN1.
Tumour Biol. 37:1151–1157. 2016. View Article : Google Scholar
|
|
47
|
Xie J, Yu F, Li D, Zhu X, Zhang X and Lv
Z: MicroRNA-218 regulates cisplatin (DPP) chemosensitivity in
non-small cell lung cancer by targeting RUNX2. Tumour Biol.
37:1197–1204. 2016. View Article : Google Scholar
|
|
48
|
Garofalo M, Di Leva G, Romano G, Nuovo G,
Suh SS, Ngankeu A, Taccioli C, Pichiorri F, Alder H, Secchiero P,
et al: miR-221&222 regulate TRAIL resistance and enhance
tumorigenicity through PTEN and TIMP3 downregulation. Cancer Cell.
16:498–509. 2009. View Article : Google Scholar
|
|
49
|
Berghmans T, Ameye L, Willems L, Paesmans
M, Mascaux C, Lafitte JJ, Meert AP, Scherpereel A, Cortot AB,
Cstoth I, et al: Identification of microRNA-based signatures for
response and survival for non-small cell lung cancer treated with
cisplatin-vinorelbine A ELCWP prospective study. Lung Cancer.
82:340–345. 2013. View Article : Google Scholar
|
|
50
|
Fiedler SD, Carletti MZ and Christenson
LK: Quantitative RT-PCR methods for mature microRNA expression
analysis. Methods Mol Biol. 630:49–64. 2010. View Article : Google Scholar
|