1. Introduction
Technological advances have driven an improved
understanding of protein-coding genes; however, the functional
roles of non-coding (nc)RNAs are relatively less well understood.
ncRNAs account for >90% of the human genome, whereas
protein-coding genes account for only 1.5% (1,2). Based
on transcript size, ncRNAs are divided into two groups: Small
ncRNAs with transcripts <200 nucleotides and long ncRNAs
(lncRNAs) with transcripts >200 nucleotides in length (3). lncRNAs, first discovered in the
sequencing of cDNA libraries in mouse cells (4), are mRNA-like transcripts that are
likely transcribed by RNA polymerase II (RNA pol II), but which
lack a stable open reading frame (5). Initially, these non-coding RNAs were
viewed as by-products and noise of the transcription process
(4). However, with the continuous
development of gene technologies, a large number of studies have
found that lncRNAs are involved in various physiological and
pathological processes.
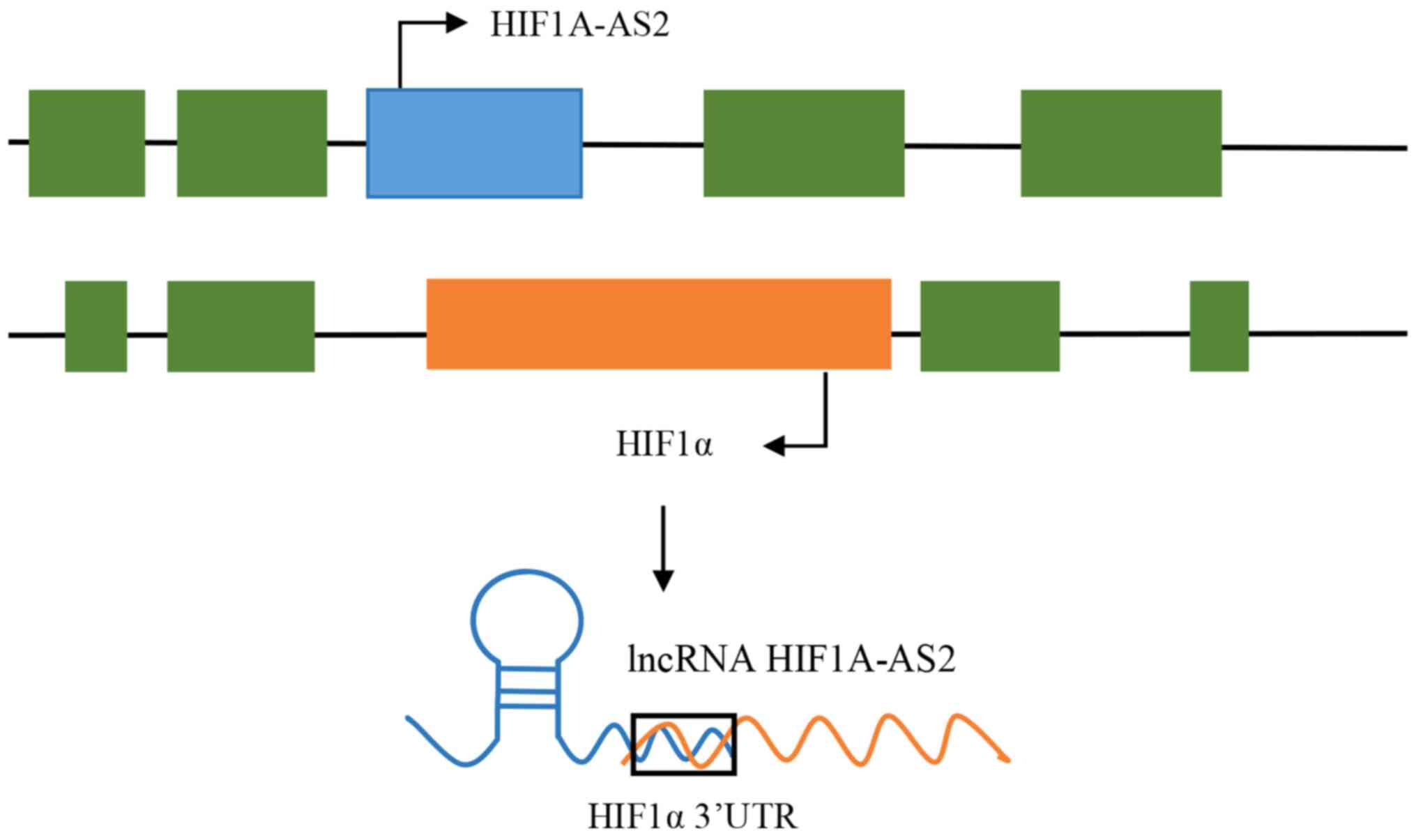
lncRNA HIF1A-AS2, also known as HIF1A-AS2, is the
endogenous antisense transcript of hypoxia-inducible factor 1α
(HIF1α), and 3'aHIF, termed HIF1α antisense RNA 2 (HIF1A-AS2), is
localized at chromosome 14q23.2, and is 2,052 nucleotides in
length. In 1999, it was first discovered to be abnormally expressed
in clear cell renal carcinoma by Thrash-Bingham and Tartof
(6), and was identified as the
endogenous antisense transcript, which could bind to the 3'
untranslated region (3'UTR) of HIF1α mRNA in a complementary manner
(Fig. 1), and this bound form is
referred to as aHIF. In 2002, Rossignol et al (7) reported that HIF1A-AS2 was expressed in
several human tissues, both physiologically, and when the tissues
had become cancerous. These findings attracted increased focus on
HIF1A-AS2. Further studies demonstrated that HIF1A-AS2 was
aberrantly expressed in various human diseases, including
preeclampsia (PE), epithelial ovarian cancer (EOC), colorectal
cancer (CRC), gastric cancer (GC), breast cancer (BC), bladder
cancer, osteosarcoma (OS), renal cell carcinoma, non-small cell
lung cancer (NSCLC) and glioblastoma (GBM). Chen et al
(8) reported that the expression
levels of HIF1A-AS2 were upregulated in GC tissues and cells, and
this upregulated expression was correlated with
Tumor-Node-Metastasis stage, tumor invasion, lymph node metastasis
and a poor prognosis. Lin et al (9) also demonstrated upregulated expression
of HIF1A-AS2 in 60 OS tissues compared with the adjacent healthy
tissues. Thus, HIF1A-AS2 may serve as a promising target for
treatment of several types of cancer.
However, several studies demonstrated that the
expression levels of HIF1A-AS2 in tumor tissues was abnormal,
indicating the potential correlation between HIF1A-AS2 and cancer.
Therefore, this review summarizes the current body of knowledge
regarding the aberrant expression of this lncRNA (Table I), its function and the regulatory
mechanisms of HIF1A-AS2 in several types of cancer.
 | Table IExpression and function of long
non-coding RNA HIF1α antisense RNA in PE and various types of
cancer. |
Table I
Expression and function of long
non-coding RNA HIF1α antisense RNA in PE and various types of
cancer.
| Disease | Change in
expression | Role | Biological
function | Related genes | Refs. |
|---|
| PE | Down | Pathogenic | Proliferation,
migration, invasion, pro-apoptosis, cell cycle arrest | LSD1, PHLDA1 | (14) |
| Epithelial ovarian
cancer | Up | Oncogenic | Proliferation,
migration and invasion | Bax, caspase-7,
caspase-9, BCL-2, caspase-3 | (18) |
| Colorectal
cancer | Up | Oncogenic | Proliferation,
migration and invasion | miR-129-5p,
miR-33b-5p DNMT3A | (23) |
| Gastric cancer | Up | Oncogenic | Proliferation,
migration and invasion | - | (8) |
| Breast cancer | Up | Oncogenic | Proliferation,
migration and invasion | miR-548c-3p, HIF1α,
VEGF | (30) |
| Bladder cancer | Up | Oncogenic | Proliferation,
migration, invasion and anti-apoptosis | - | (35) |
| Osteosarcoma | Up | Oncogenic | Proliferation,
migration, invasion and anti-apoptosis | miR-33b-5p, SIRT6,
miR-129-5p | (9,40) |
| Glioblastoma | Up | Oncogenic | Neurosphere
formation | IGF2BP2, DHX9,
HMGA1 | (43) |
| Renal cancer | Up | Oncogenic | Proliferation,
migration, invasion and anti-apoptosis | HIF1α,
miR-130-5p | (6,50) |
| Non-small cell lung
cancer | Up | Oncogenic | Proliferation,
migration, invasion and anti-apoptosis | miR-153b-5p,
S100A14 | (54) |
2. Expression and function of lncRNA
HIF1A-AS2 in several types of cancer
PE
PE is one of the leading causes of maternal death
and a pregnancy-specific disease, affecting 3-14% of parturients
worldwide (10). Although PE has
been extensively studied (11), the
underlying pathogenesis of PE remains elusive. However, it is
hypothesized that inadequate trophoblastic invasion may cause PE
(12,13). Wu et al (14) reported that HIF1A-AS2 expression was
significantly downregulated in the tissues of 52 patients with PE
compared with the adjacent normal samples. Knockdown of HIF1A-AS2
expression significantly inhibited proliferation, migration and
invasion, as well as inducing G0/G1 cell cycle arrest and increased
cell apoptosis in two trophoblast cell lines (HTR/SVneo and JAR).
In contrast, overexpression of HIF1A-AS2 exerted the opposite
effect. Mechanistically, a subcellular localization assay indicated
that HIF1A-AS2 was primarily localized in the cell nucleus; thus,
it may play a role in regulation of transcription. Further
experiments showed that HIF1A-AS2 inhibited the transcription of
pleckstrin homology like domain, family A, member 1 (PHLDA1), which
plays a significant role in the activation-induced apoptosis
following binding to lysine-specific demethylase (LSD1) at the
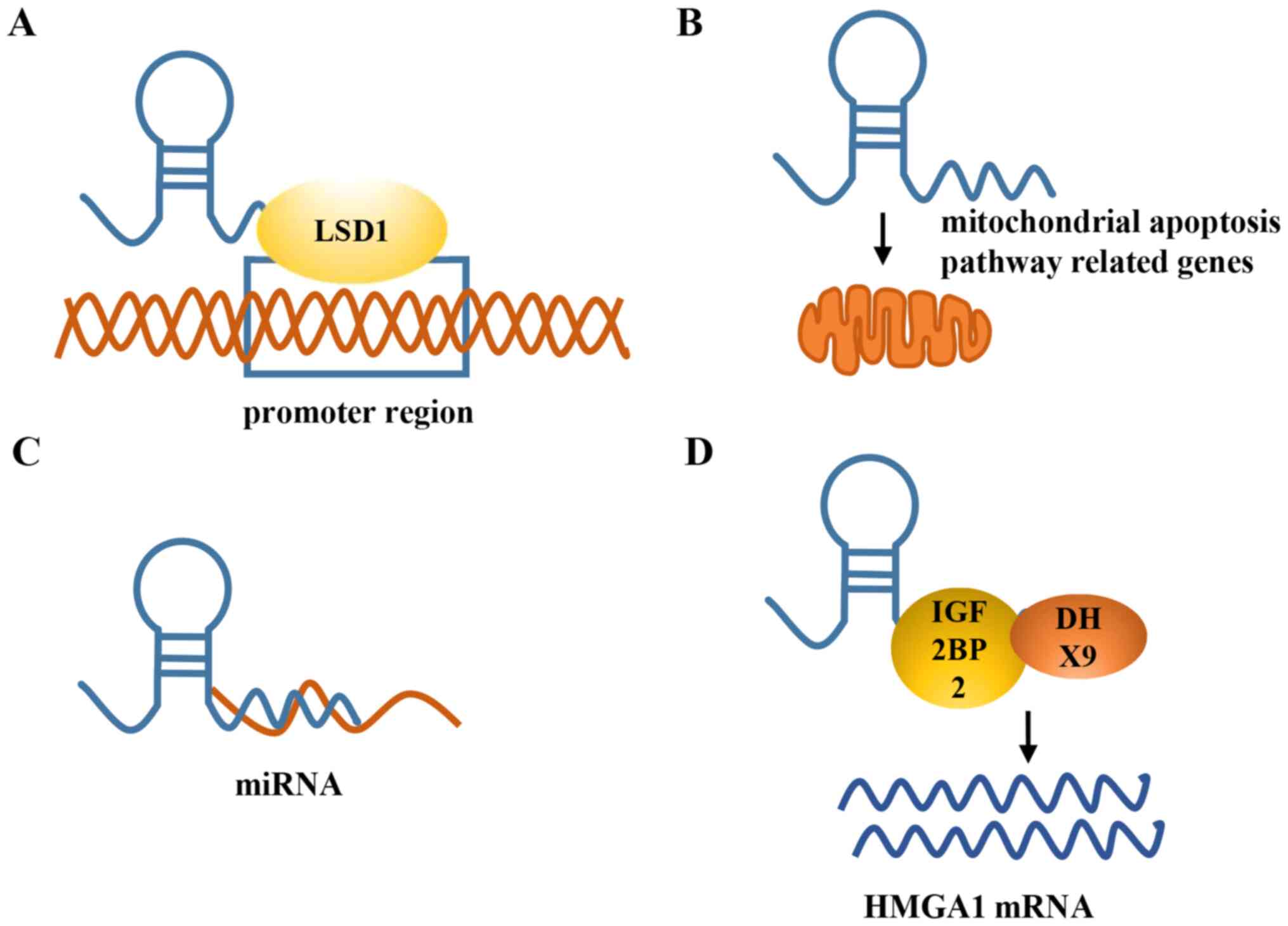
epigenetic level. Furthermore, chromatin immunoprecipitation assays
showed LSD1 and H3K4 me2 enrichment in the promoter region of the
PHLDA1 gene (Fig. 2A) after
transfection with small-interfering (si)-HIF1A-AS2. Thus, HIF1A-AS2
may be a useful diagnostic biomarker for PE.
EOC
Ovarian cancer (OC) is one of the most common types
of malignant tumors in females, with EOC being the most common,
accounting for 80-90% of OC cases (15,16) .
Although EOC treatments have improved notably, even in developed
countries, such as the United States and Canada, the overall
survival remains at only 47% 5 years after diagnosis (17). Therefore, investigating the
molecular mechanism and finding effective therapeutic targets for
management of EOC is of great importance. Qiu et al
(18) reported that the expression
of HIF1A-AS2 in EOC tissues was significantly higher compared with
the normal controls, and HIF1A-AS2 was a lncRNA that was
upregulated under hypoxic conditions. Thus, the following assays
were performed under hypoxic conditions. Functional assays revealed
that knockdown of HIF1A-AS2 promoted cell apoptosis and weakened
tumorigenesis in nude mice. In contrast, overexpression of
HIF1A-AS2 inhibited EOC cell apoptosis and enhanced cell
proliferation.
Further mechanistic experiments showed that
HIF1A-AS2 functions by regulating the mitochondrial apoptosis
pathway-related genes (Fig. 2B).
Briefly, HIF1A-AS2 knockdown resulted in increased expression of
Bax, Bcl-2, caspase-7, and caspase-9 at the mRNA level under
hypoxic conditions. Thus, overexpression of HIF1A-AS2 may serve as
a diagnostic biomarker for EOC.
CRC
CRC is the third most common type of cancer and the
fourth leading cause of cancer-associated death globally (19,20).
At present, chemotherapy is an essential treatment for CRC;
however, both the incidence and death rate of CRC is increasing
rapidly (21,22). Thus, it is crucial to identify novel
critical genes involved in the pathogenesis of CRC to develop
effective treatments. Lin et al (23) observed upregulated expression of
HIF1A-AS2 in CRC tissues and cells compared with the healthy
controls. Moreover, high expression of HIF1A-AS2 was strongly
associated with a poor prognosis and advanced TNM stages in
patients with CRC. Functionally, knockdown of HIF1A-AS2 inhibited
the proliferation, invasion and epithelial-mesenchymal
transformation (EMT) of CRC cells in-vitro. HIF1A-AS2
mechanistically functioned as a competing endogenous (ce)RNA
binding to microRNA (miR)-129-5p (Fig.
2C), a tumor suppressor. Consistent with this, DNMT3A was
identified to be a target of miR-129–5p. The critical role of the
HIF1A-AS2/miR-129-5p/DNMT3A axis in the proliferation, invasion and
EMT of CRC cells was further confirmed by reverse
transcription-quantitative PCR and dual- luciferase reporter
assays. Thus, due to its oncogenic role and clinical significance
in colorectal cancer, HIF1A-AS2 may be considered a diagnostic
biomarker and prognostic indicator for CRC.
GC
GC is the third leading cause of cancer-associated
death worldwide, with ~1,000,000 newly diagnosed cases each year,
and a higher rate of occurrence in East Asia (24,25).
The majority of patients are diagnosed with advanced stage GC, and
thus, GC has a high mortality rate (26). Therefore, it has been a central
issue to study the pathogenic mechanisms of GC and identify
effective tumor markers to improve early diagnosis. Chen et
al (8) reported that HIF1A-AS2
was upregulated in 38 GC samples and four human GC cell lines
compared with the matched paracarcinoma tissues or a normal GC cell
line (GES-1), respectively. The high expression of HIF1A-AS2 was
significantly associated with a more advanced TNM stage, tumor
invasion, lymph node metastasis and a poor prognosis. Functionally,
knockdown of HIF1A-AS2 suppressed the proliferative ability of GC
cells in-vitro and restrained tumor weight and volume in
nude mice. In addition, it was found that HIF1A-AS2 had value in
the early diagnosis of GC and could be used as a potential
diagnostic marker for detection of GC. Therefore, HIF1A-AS2 is a
potential tumorigenic gene in GC, but its molecular mechanisms have
not been studied, to the best of our knowledge.
BC
BC is the most common malignancy and the leading
cause of cancer-related death in women (27). Breast cancer tumors usually express
a combination of the following receptors: Estrogen receptor (ER),
progesterone receptor (PR) and human epidermal growth factor
receptor (HER2). Cases that lack expression of these three
receptors are termed triple-negative breast cancer (TNBC). TNBC
accounts for ~20% of all breast cancer cases, ad is most common in
women >40(28). TNBC is highly
invasive, with high mortality and recurrence rates. Current
treatments for TNBC include surgery, chemotherapy, radiotherapy and
targeted therapy. However, the median overall survival rarely
extends beyond 18 months in patients with advanced BC (29). Therefore, it is essential to study
the molecular mechanism and identify novel biomarkers for
management of TNBC. Guo et al (30) showed that HIF1A-AS2 was
significantly overexpressed in four BC cell lines compared with a
normal mammary epithelial cell line. Knockdown of HIF1A-AS2 levels
effectively suppress proliferation, invasion, EMT and senescence of
MCF-7 cell lines in-vitro. In vivo studies also
showed that tumor growth was reduced after the knockdown of
HIF1A-AS2 by short hairpin (sh)RNA targeting HIF1A-AS2
in-vivo, thus indicating that HIF1A-AS2 functions as an
oncogene. Mechanistically, a HIF1A-AS2/miR-548c-3p/HIF1a/VEGF axis
was confirmed to regulate the proliferation, invasion, migration
and EMT of BC cells. Jiang et al (31) also reported that expression of
HIF1A-AS2 was increased in 33 TNBC tissues compared with the
adjacent normal breast tissues. Knockdown of HIF1A-AS2 functionally
suppressed TNBC cell proliferation. These results indicated that
HIF1A-AS2 was involved in the pathogenesis of TNBC, suggesting that
it could be a prognostic indicator or therapeutic target for
TNBC.
Bladder cancer
Bladder cancer is one of the most common
malignancies of the urinary system worldwide, posing a severe
threat to human health (32).
Surgery, radiotherapy and chemotherapy are the primary modes of
treatment for bladder cancer; however, the 5-year overall survival
rate is only 50-60% (33). Although
several studies have demonstrated novel biomarkers for the early
detection and diagnosis of bladder cancer, the survival rate of
patients with bladder cancer remains very low (34). Therefore, it is necessary to
identify novel biomarkers to improve the early diagnosis and
prognosis of bladder cancer. Chen et al (35) revealed that the expression of
HIF1A-AS2 was significantly upregulated in 44 bladder cancer
samples and cancer cell lines (5637 and T24) compared with the
matched normal peritumoral tissues or the SVHUC-1 normal bladder
cell line. In addition, the upregulated HIF1A-AS2 expression was
closely related to histological grade, tumor invasion depth and TNM
stage. These results indicated that lncRNA HIF1A-AS2 may function
as an oncogene in bladder cancer. Functionally, knockdown of
HIF1A-AS2 significantly inhibited bladder cancer cell proliferation
and migration, and increased apoptosis. Conversely, overexpression
of HIF1A-AS2 had the opposite effect.
Furthermore, a tetracycline-induced shRNA using
medical synthetic biology techniques was designed, which could
effectively inhibit the expression of HIF1A-AS2 in a dose-dependent
manner, and in turn inhibited cell growth and migration, and
induced apoptosis in bladder cancer cells. It also indicated that
tetracycline-induced shRNA may be a novel approach for
quantitatively controlling specific targets in human cancers, and
may be an effective treatment method for bladder cancer. Thus,
HIF1A-AS2 may serve as a target for the treatment of bladder
cancer; however, the exact molecular regulatory mechanisms in
bladder cancer require further study.
OS
OS is a skeletal system primary malignant tumor,
common amongst the younger population, particularly children and
adolescents (36,37). OS accounts for 60% of all sarcoma
cases, characterized by early metastasis, high aggressiveness, a
high rate of disability and a high recurrence rate (38). Despite advances in OS treatment, the
overall survival of patients has not substantially increased, the
5-year overall survival still remains only 20% over the past 30
years (39). Thus, understanding
the molecular mechanism of OS and identifying novel therapeutic
targets is of great clinical significance to improve early
diagnosis and survival rates of patients with OS. Lin et al
(9) observed increased HIF1A-AS2
expression in 60 OS samples and four OS cell lines when compared
with the 60 adjacent normal samples or the hFOB 1.19 cells,
respectively. In addition, high expression of HIF1A-AS2 was
significantly associated with a larger tumor size, higher tumor
grade, advanced stage disease and distance of metastasis.
Furthermore, Kaplan-Meier survival analysis showed
that the 5-year survival rate of the high HIF1A-AS2 expression
group was lower than the low HIF1A-AS2 expression group. Knockdown
of HIF1A-AS2 resulted in decreased cell proliferation, migration
and invasion, increased cell cycle arrest in the G0/G1-phase and an
increased percentage of apoptotic cells. In in-vivo
experiments, knockdown of HIF1A-AS2 resulted in reduced tumor size
in nude mice. Mechanistically, HIF1A-AS2/miR-33b-5p/SIRT6 was
confirmed to regulate OS cell proliferation, migration and
apoptosis. Wang et al (40)
also confirmed increased expression of HIF1A-AS2 in 30 OS samples
and four OS cell lines compared with the adjacent normal tissues
and osteoblast cell lines, respectively. Moreover, high HIF1A-AS2
expression was associated with poor survival rates. Functional
assays revealed that HIF1A-AS2 overexpression promoted osteosarcoma
cell proliferation, cell cycle progression and invasion. HIF1A-AS2
mechanistically served as a ceRNA to negatively regulate miR-129–5p
(Fig. 2C). Thus, HIF1A-AS2 may be
an effective diagnostic and prognostic indicator of OS.
GBM
GBM is the most common and aggressive primary
malignant brain tumor, with a median patient survival time of 14-16
months (41). GBM is a highly
proliferative and invasive tumor with a poor prognosis. Despite
advances in GBM treatment, patients are still likely to face a poor
prognosis (42). Therefore, new
therapeutic methods and targets are required. lncRNAs are involved
in the development of GBM. Mineo et al (43) reported that HIF1A-AS2 contributes to
the formation of stem-like glioma cells (GSCs) in the tumor
microenvironment and their adaptation to hypoxia. Based on
characterization of the GBM genome and transcriptome, GBM can be
divided into several cellular subtypes, including mesenchymal (M),
proneural (P), neural (N), and classical (C) (44). Patients with the aggressive and
predominant M subtype exhibit a particularly high degree of tumor
necrosis (45). It was observed
that HIF1A-AS2 expression was significantly increased in the GSCs
of patients with the M subtype. Moreover, knockdown of HIF1A-AS2
led to reduced growth, decreased cellular activity and decreased
neurosphere-forming capacity of M GSC cells (43).
Furthermore, the HIF1A-AS2 expression is increased
under hypoxic conditions. In order to clarify the pro-oncogenic
function of HIF1A-AS2, researchers revealed that knockdown of
HIF1A-AS2 by shRNA resulted in smaller tumor sizes in nude mice.
Mechanistic experiments showed that HIF1A-AS2 could bind to IGF2BP2
and DHX9 to directly modulate the expression of HMGA1 (Fig. 2D) and maintain the growth of M GSCs
under hypoxic conditions (43). In
addition, Liao et al (46)
showed that the upregulated HIF1A-AS2 expression could mediate
radiation resistance of the glioma, leading to tumor recurrence
following radiotherapy by regulating expression of apoptotic
proteins. Knockdown of HIF1A-AS2 increased the expression of the
pro-apoptotic protein caspase 7 and the number of apoptotic cells.
Thus, HIF1A-AS2 may be a novel diagnostic indicator and potential
therapeutic target for the management of GBM.
RCC
RCC is one of the most common malignancies of the
urinary system, and accounts for 2-3% of all malignancies (47,48).
The estimated number of new cases and deaths worldwide in 2018 were
403,262 and 175,098, respectively (49). Relatively fewer biomarkers for RCC
have been identified when compared with other types of cancer.
Thus, it is essential to identify novel and sensitive biomarkers to
predict the progress and prognosis of the disease. In 1999,
Thrash-Bingham and Tartof (6) first
discovered a natural antisense transcript that could bind to the
3'UTR of HIF1α mRNA in non-papillary kidney cancer and termed it
aHIF, for which the official gene symbol is now HIF1A-AS2.
Expression of HIF1A-AS2 is increased in non-papillary renal
carcinoma cells compared with the control cells, but not in
papillary renal carcinoma cells. It is hypothesized that decreased
HIF1a mRNA expression through HIF1A-AS2 may serve an important role
in regulating P53 to regulate progression of cancer, but this
mechanism requires further investigation (6). Zhu et al (50) also reported increased expression of
HIF1A-AS2 in kidney cancer tissues and RCC cells compared with the
non-cancerous tissues. In addition, knockdown of HIF1A-AS2
inhibited renal cancer cell proliferation, invasion and migration,
whilst accelerating cell apoptosis. Overexpression of HIF1A-AS2
resulted in the opposite effect. HIF1A-AS2 mechanistically
functions as a ceRNA, binding to miR-130a-5p (Fig. 2C) to modulate renal carcinoma
progression. Thus, HIF1A-AS2 may be a promising diagnostic
biomarker and a potential therapeutic target for management of
renal cancer.
NSCLC
Lung cancer is the most common type of cancer and
the leading cause of cancer-associated death worldwide. The
majority of patients are diagnosed with advanced stage disease in
the first instance, and NSCLC accounts for nearly 85% of patients
with lung cancer (51). Despite
advances in cancer treatment, lung cancer has a high mortality
rate, accounting for 18.4% of all cancer deaths (52,53).
Thus, understanding the molecular mechanism of NSCLC and
identifying novel therapeutic targets is of great clinical
significance. Zhang et al (54) reported elevated expression levels of
HIF1A-AS2 in NSCLC tissues and cell lines, and this increased
expression was associated with a poor prognosis. However, knockdown
of HIF1A-AS2 resulted in decreased cell proliferation, migration
and invasion, and an increased percentage of apoptotic cells.
Mechanistically, a HIF1A-AS2/miR-153–5p/S100A14 axis was confirmed
to regulate NSCLC cell proliferation, migration and apoptosis
(Fig. 2C). Thus, HIF1A-AS2 may be
an effective diagnostic and prognostic indicator for NSCLC.
3. Conclusions and future perspective
A wealth of studies have shown that lncRNAs exert
their functions through various mechanisms, such as associating
with transcription factors, chromatin modifiers, signaling
adapters, enzymes and miRNAs, to influence gene expression,
post-translational modifications and protein activities (55).
HIF1A-AS2 has been reported to regulate cellular
pathological processes, but is primarily focused on tumors.
HIF1A-AS2 is primarily functions as a protein scaffold, protein
decoy and a ceRNA. Mineo et al (43) reported that HIF1A-AS2 acts as a
protein scaffold to bind both IGF2BP2 and DHX9 to modulate the
expression of HMGA1. Wu et al (14) reported that HIF1A-AS2 functions as a
protein decoy to inhibit the transcription of PHLDA1 by binding to
LSD1, a histone demethylase. Additionally, HIF1A-AS2 acts as a
molecular sponge to bind miRNAs to further affect expression of
other genes (23,40,50,54,56).
Although significant achievements have been obtained with regard to
understanding the role of HIF1A-AS2 in various types of cancer,
further studies are still required with regard to its regulatory
function, as lncRNAs often exhibit several complex regulatory
functions/mechanisms.
Studies have shown that HIF1A-AS2 may serve as a
novel biomarker for the clinical diagnosis of several types of
cancer. These data demonstrate that upregulated expression of
HIF1A-AS2 is associated with poor overall survival and an
unfavorable prognosis, such as in TNBC, OS and CRC. Nevertheless,
the clinical diagnostic value of HIF1A-AS2 in these types of cancer
needs to be validated using large-scale multicenter cohorts.
Acknowledgements
Not applicable.
Funding
This study was supported by grants from The National Natural
Science Fund of China (grant no. 81601736); Science and Technology
Planning Project of Panyu District, China (grant no. 2019-Z04-85);
Medical Scientific Research Foundation of Guangdong Province, China
(grant no. A2020560); Basic and Applied Research Project of
Guangzhou Research Program, China (grant no. 202102080539).
Availability of data and materials
Not applicable.
Authors' contributions
YLi conceived and designed the study. YLiu, YZ and
CC participated in drafting and revising the article. All authors
have read and approved the final manuscript. Data authentication is
not applicable.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Slack FJ and Chinnaiyan AM: The role of
non-coding RNAs in oncology. Cell. 179:1033–1055. 2019.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Esteller M: Non-coding RNAs in human
disease. Nat Rev Genet. 12:861–874. 2011.PubMed/NCBI View
Article : Google Scholar
|
|
3
|
Khandelwal A, Bacolla A, Vasquez KM and
Jain A: Long non-coding RNA: A new paradigm for lung cancer. Mol
Carcinog. 54:1235–1251. 2015.PubMed/NCBI View
Article : Google Scholar
|
|
4
|
Mercer TR, Dinger ME and Mattick JS: Long
non-coding RNAs: Insights into functions. Nat Rev Genet.
10:155–159. 2009.PubMed/NCBI View
Article : Google Scholar
|
|
5
|
Bhat SA, Ahmad SM, Mumtaz PT, Malik AA,
Dar MA, Urwat U, Shah RA and Ganai NA: Long non-coding RNAs:
Mechanism of action and functional utility. Noncoding RNA Res.
1:43–50. 2016.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Thrash-Bingham CA and Tartof KD: aHIF: A
natural antisense transcript overexpressed in human renal cancer
and during hypoxia. J Natl Cancer Inst. 91:143–151. 1999.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Rossignol F, Vaché C and Clottes E:
Natural antisense transcripts of hypoxia-inducible factor 1alpha
are detected in different normal and tumour human tissues. Gene.
299:135–140. 2002.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Chen WM, Huang MD, Kong R, Xu TP, Zhang
EB, Xia R, Sun M, De W and Shu YQ: Antisense long noncoding RNA
HIF1A-AS2 is upregulated in gastric cancer and associated with poor
prognosis. Dig Dis Sci. 60:1655–1662. 2015.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Lin H, Zhao Z, Hao Y and He J and He J:
Long noncoding RNA HIF1A-AS2 facilitates cell survival and
migration by sponging miR-33b-5p to modulate SIRT6 expression in
osteosarcoma. Biochem Cell Biol. 98:284–292. 2020.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Wu P, Haththotuwa R, Kwok CS, Babu A,
Kotronias RA, Rushton C, Zaman A, Fryer AA, Kadam U, Chew-Graham
CA, et al: Preeclampsia and future cardiovascular health: A
systematic review and meta-analysis. Circ Cardiovasc Qual Outcomes.
10(e003497)2017.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Phipps EA, Thadhani R, Benzing T and
Karumanchi SA: Pre-eclampsia: Pathogenesis, novel diagnostics and
therapies (vol 15, pg 275, 2019). Nat Rev Nephrol. 15:386.
2019.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Irminger-Finger I, Jastrow N and Irion O:
Preeclampsia: A danger growing in disguise. Int J Biochem Cell
Biol. 40:1979–1983. 2008.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Ma Y, Liang X, Wu H, Zhang C and Ma Y:
Long non coding RNA NR_002794 is upregulated in pre eclampsia and
regulates the proliferation, apoptosis and invasion of trophoblast
cells. Mol Med Rep. 20:4567–4575. 2019.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Wu D, Yang N, Xu Y, Wang S, Zhang Y,
Sagnelli M, Hui B, Huang Z and Sun L: lncRNA HIF1A antisense RNA 2
modulates trophoblast cell invasion and proliferation through
upregulating PHLDA1 expression. Mol Ther Nucleic Acids. 16:605–615.
2019.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Tang X, Liu S, Liu Y, Lin X, Zheng T, Liu
X, Qiu J and Hua K: Circulating serum exosomal aHIF is a novel
prognostic predictor for epithelial ovarian cancer. Onco Targets
Ther. 12:7699–7711. 2019.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Guler E, Smith DA, Somarouthu B, Gujrathi
R, Ramaiya NH and Tirumani SH: Overview of imaging findings
associated with systemic therapies in advanced epithelial ovarian
cancer. Abdom Radiol (NY). 45:828–841. 2020.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Lheureux S, Braunstein M and Oza AM:
Epithelial ovarian cancer: Evolution of management in the era of
precision medicine. CA Cancer J Clin. 69:280–304. 2019.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Qiu JJ, Lin XJ, Zheng TT, Tang XY and Hua
KQ: Natural antisense transcript of hypoxia-inducible factor 1
regulates hypoxic cell apoptosis in epithelial ovarian cancer. Onco
Targets Ther. 11:9101–9110. 2018.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Yang J, McDowell A, Kim EK, Seo H, Lee WH,
Moon CM, Kym SM, Lee DH, Park YS, Jee YK, et al: Development of a
colorectal cancer diagnostic model and dietary risk assessment
through gut microbiome analysis. Exp Mol Med. 51:1–15.
2019.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Arnold M, Sierra MS, Laversanne M,
Soerjomataram I, Jemal A and Bray F: Global patterns and trends in
colorectal cancer incidence and mortality. Gut. 66:683–691.
2017.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Yuan W, Peng S, Wang J, Wei C, Ye Z, Wang
Y, Wang M, Xu H, Jiang S, Sun D, et al: Identification and
characterization of circRNAs as competing endogenous RNAs for
miRNA-mRNA in colorectal cancer. PeerJ. 7(e7602)2019.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Dong Y, Zhang Y, Kang W, Wang G, Chen H,
Higashimori A, Nakatsu G, Go M, Tong JH, Zheng S, et al: VSTM2A
suppresses colorectal cancer and antagonizes Wnt signaling receptor
LRP6. Theranostics. 9:6517–6531. 2019.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Lin J, Shi Z, Yu Z and He Z: lncRNA
HIF1A-AS2 positively affects the progression and EMT formation of
colorectal cancer through regulating miR-129-5p and DNMT3A. Biomed
Pharmacother. 98:433–439. 2018.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Fan H, Jin X, Liao C, Qiao L and Zhao W:
MicroRNA-301b-3p accelerates the growth of gastric cancer cells by
targeting zinc finger and BTB domain containing 4. Pathol Res
Pract. 215(152667)2019.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Wan P, Bai X, Yang C, He T, Luo L, Wang Y,
Fan M, Wang Z, Lu L, Yin Y, et al: miR-129-5p inhibits
proliferation, migration, and invasion in rectal adenocarcinoma
cells through targeting E2F7. J Cell Physiol. 235:5689–5701.
2020.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Wang LL, Zhang L and Cui XF:
Downregulation of long noncoding RNA LINC01419 inhibits cell
migration, invasion, and tumor growth and promotes autophagy via
inactivation of the PI3K/Akt1/mTOR pathway in gastric cancer. Ther
Adv Med Oncol. 11(1758835919874651)2019.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Lee KL, Kuo YC, Ho YS and Huang YH:
Triple-negative breast cancer: Current understanding and future
therapeutic breakthrough targeting cancer Stemness. Cancers
(Basel). 11(1334)2019.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Khan MA, Jain VK, Rizwanullah M, Ahmad J
and Jain K: PI3K/AKT/mTOR pathway inhibitors in triple-negative
breast cancer: A review on drug discovery and future challenges.
Drug Discov Today. 24:2181–2191. 2019.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Song N, Zhao L, Zhu M and Zhao J:
99mTc-Labeled LyP-1 for SPECT Imaging of Triple Negative Breast
Cancer. Contrast Media Mol Imaging. 2019(9502712)2019.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Guo X, Lee S and Cao P: The inhibitive
effect of sh-HIF1A-AS2 on the proliferation, invasion, and
pathological damage of breast cancer via targeting miR-548c-3p
through regulating HIF-1α/VEGF pathway in vitro and vivo. Onco
Targets Ther. 12:825–834. 2019.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Jiang YZ, Liu YR, Xu XE, Jin X, Hu X, Yu
KD and Shao ZM: Transcriptome Analysis of triple-negative breast
cancer reveals an integrated mRNA-lncRNA signature with predictive
and prognostic value. Cancer Res. 76:2105–2114. 2016.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Yu G, Zhou H, Yao W, Meng L and Lang B:
lncRNA TUG1 promotes cisplatin resistance by regulating CCND2 via
epigenetically silencing miR-194-5p in bladder cancer. Mol Ther
Nucleic Acids. 16:257–271. 2019.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Miao L, Liu HY, Zhou C and He X: LINC00612
enhances the proliferation and invasion ability of bladder cancer
cells as ceRNA by sponging miR-590 to elevate expression of PHF14.
J Exp Clin Cancer Res. 38(143)2019.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Zhuang C, Ma Q, Zhuang C, Ye J, Zhang F
and Gui Y: lncRNA GClnc1 promotes proliferation and invasion of
bladder cancer through activation of MYC. FASEB J. 33:11045–11059.
2019.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Chen M, Zhuang C, Liu Y, Li J, Dai F, Xia
M, Zhan Y, Lin J, Chen Z, He A, et al: Tetracycline-inducible shRNA
targeting antisense long non-coding RNA HIF1A-AS2 represses the
malignant phenotypes of bladder cancer. Cancer Lett. 376:155–164.
2016.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Qi X, Yu XJ, Wang XM, Song TN, Zhang J,
Guo XZ, Li GJ and Shao M: Knockdown of KCNQ1OT1 suppresses cell
invasion and sensitizes osteosarcoma cells to CDDP by upregulating
DNMT1-mediated Kcnq1 expression. Mol Ther Nucleic Acids.
17:804–818. 2019.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Jiang N, Wang X, Xie X, Liao Y, Liu N, Liu
J, Miao N, Shen J and Peng T: lncRNA DANCR promotes tumor
progression and cancer stemness features in osteosarcoma by
upregulating AXL via miR-33a-5p inhibition. Cancer Lett. 405:46–55.
2017.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Zhang CL, Zhu KP and Ma XL: Antisense
lncRNA FOXC2-AS1 promotes doxorubicin resistance in osteosarcoma by
increasing the expression of FOXC2. Cancer Lett. 396:66–75.
2017.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Zhong L, Liao D, Li JJ, Liu WQ, Wang JX,
Zeng CL, Wang X, Cao ZL, Zhang RH, Li M, et al: Rab22a-NeoF1 fusion
protein promotes osteosarcoma lung metastasis through its secretion
into exosomes. Signal Transduct Target Ther. 6(59)2021.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Wang X, Peng L, Gong X, Zhang X and Sun R:
lncRNA HIF1A-AS2 promotes osteosarcoma progression by acting as a
sponge of miR-129-5p. Aging (Albany NY). 11:11803–11813.
2019.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Tang C, Wang Y, Zhang L, Wang J, Wang W,
Han X, Mu C and Gao D: Identification of novel lncRNA targeting
Smad2/PKCα signal pathway to negatively regulate malignant
progression of glioblastoma. J Cell Physiol. 235:3835–3848.
2020.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Ji J, Xu R, Ding K, Bao G, Zhang X, Huang
B, Wang X, Martinez A, Wang X, Li G, et al: Long Noncoding RNA
SChLAP1 forms a growth-promoting complex with HNRNPL in human
glioblastoma through stabilization of ACTN4 and activation of NF-κB
signaling. Clin Cancer Res. 25:6868–6881. 2019.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Mineo M, Ricklefs F, Rooj AK, Lyons SM,
Ivanov P, Ansari KI, Nakano I, Chiocca EA, Godlewski J and Bronisz
A: The Long Non-coding RNA HIF1A-AS2 facilitates the maintenance of
mesenchymal glioblastoma stem-like cells in hypoxic niches. Cell
Rep. 15:2500–2509. 2016.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Wang L, Babikir H, Müller S, Yagnik G,
Shamardani K, Catalan F, Kohanbash G, Alvarado B, Di Lullo E,
Kriegstein A, et al: The phenotypes of proliferating glioblastoma
cells reside on a single axis of variation. Cancer Discov.
9:1708–1719. 2019.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Bhat KPL, Balasubramaniyan V, Vaillant B,
Ezhilarasan R, Hummelink K, Hollingsworth F, Wani K, Heathcock L,
James JD, Goodman LD, et al: Mesenchymal differentiation mediated
by NF-κB promotes radiation resistance in glioblastoma. Cancer
Cell. 24:331–346. 2013.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Liao K, Ma X, Chen B, Lu X, Hu Y, Lin Y,
Huang R and Qiu Y: Upregulated AHIF-mediated radioresistance in
glioblastoma. Biochem Biophys Res Commun. 509:617–623.
2019.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Kovacs G, Wilkens L, Papp T and de Riese
W: Differentiation between papillary and nonpapillary renal cell
carcinomas by DNA analysis. J Natl Cancer Inst. 81:527–530.
1989.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Korenaga Y, Naito K, Okayama N, Hirata H,
Suehiro Y, Hamanaka Y, Matsuyama H and Hinoda Y: Association of the
BCRP C421A polymorphism with nonpapillary renal cell carcinoma. Int
J Cancer. 117:431–434. 2005.PubMed/NCBI View Article : Google Scholar
|
|
49
|
Zhang C, Huang D, Liu A, Xu Y, Na R and Xu
D: Genome-wide screening and cohorts validation identifying novel
lncRNAs as prognostic biomarkers for clear cell renal cell
carcinoma. J Cell Biochem. 121:2559–2570. 2020.PubMed/NCBI View Article : Google Scholar
|
|
50
|
Zhu Y, Yang Z, Chen H, Pan Y, Gong L, Chen
F, Jin X, Wen S, Li Y and Chen G: lncRNAHIF1A-AS2 promotes renal
carcinoma cell proliferation and migration via miR-130a-5p/ERBB2
pathway. Onco Targets Ther. 13:9807–9820. 2020.PubMed/NCBI View Article : Google Scholar
|
|
51
|
The Lancet: Lung cancer: Some progress,
but still a lot more to do. The Lancet. 394(1880)2019.PubMed/NCBI View Article : Google Scholar
|
|
52
|
Ramalingam SS, Owonikoko TK and Khuri FR:
Lung cancer: New biological insights and recent therapeutic
advances. CA Cancer J Clin. 61:91–112. 2011.PubMed/NCBI View Article : Google Scholar
|
|
53
|
Dawson Q: NELSON trial: Reduced
lung-cancer mortality with volume CT screening. Lancet Respir Med.
8:236. 2020.PubMed/NCBI View Article : Google Scholar
|
|
54
|
Zhang W, Liu K, Pei Y, Tan J, Ma J and
Zhao J: Long noncoding RNA HIF1A-AS2 promotes non-small cell lung
cancer progression by the miR-153-5p/S100A14 axis. Onco Targets
Ther. 13:8715–8722. 2020.PubMed/NCBI View Article : Google Scholar
|
|
55
|
Wang P: The Opening of Pandora's Box: An
emerging role of long noncoding RNA in viral infections. Front
Immunol. 9(3138)2019.PubMed/NCBI View Article : Google Scholar
|
|
56
|
Mu L, Wang Y, Su H, Lin Y, Sui W, Yu X and
Lv Z: HIF1A-AS2 promotes the proliferation and metastasis of
gastric cancer cells through miR-429/PD-L1 axis. Dig Dis Sci Feb 8,
2021 (Epub ahead of print). doi: 10.1007/s10620-020-06819-w.
|