Introduction
Rheumatoid arthritis (RA) is a common systemic
autoimmune disorder that influences ~1% of the general population.
Advanced RA can be critically morbid with a decisively fatal rate
that reduces survival by 10 years (1). RA is an irreversible inflammatory
disorder that is identified based on increased inflammation,
chronic synovitis, advanced joint devastation and general
extra-articular manifestations, such as pericarditis, neuropathy,
rheumatoid nodules and vasculitis. Despite several years of
research, the pathophysiology of RA remains obscure to a great
extent, although it has been determined that a range of
inflammatory cytokines, such as tumor necrosis factor-α (TNF-α) and
interleukin-6 (IL-6) participate in RA pathogenesis (2,3).
Several biological processes, such as the growth and function of
immune cells are regulated by long noncoding RNAs (lncRNAs). More
recently, several lncRNAs have bene identified to be involved in
the pathophysiology of autoimmune inflammatory disorders (4), such as RA (5), autoimmune thyroid disease and systemic
lupus erythematosus (SLE) (6).
Growth arrest-specific 5 (GAS5), a type of lncRNA,
is a 650-nt broad-spectrum growth repressor. lncRNA GAS5 is
associated with an increased risk of SLE in mice (7). Furthermore, genetic analysis has
revealed that 1q25, the chromosomal site of GAS5, is associated
with the pathogenesis of SLE in humans (8).
LncRNA GAS5 is associated with RA, and it functions
as an effective inhibitor of the glucocorticoid receptor (GR)
through its RNA glucocorticoid response element (9). Several studies have detected lncRNA
GAS5 in patients with autoimmune diseases and reported that GAS5
levels decrease markedly in these patients when compared with
healthy individuals (10).
Recently, genetic studies have reported several single-nucleotide
polymorphisms (SNPs) that are associated with susceptibility to RA
(11). Certain SNPs are present
within exons; however, several other SNPs are present within
introns, involving lncRNAs (12).
Subsequently, lncRNA genetic polymorphisms may be associated with
the molecular pathophysiology of RA, and investigating these
variants may be valuable to reveal genetic aspects linked with the
development of RA.
The aim of the present study was to detect the serum
levels of lncRNA GAS5 in patients with RA and healthy controls and
establish the association between rs2067079, rs6790 and rs17359906
SNPs of the lncRNA GAS5 gene with RA risk in the Egyptian
population.
Patients and methods
Patients
In the present study, 350 Egyptian participants were
recruited between January 2019 and June 2019 from the Rheumatology
and Rehabilitation Department, Fayoum University Hospital, Fayoum
University, Egypt. The participants were divided into two groups:
200 patients with RA (152 females and 48 males; mean age,
42.45±0.89 years) and 150 age- and sex-matched healthy controls
(112 females and 38 males; mean age, 40.85±0.99 years).
The patients were diagnosed via clinical
examination, radiological outcomes and laboratory screenings, and
in accordance with the 2010 Rheumatoid Arthritis Classification
Criteria (13). Exclusion criteria
included other autoimmune diseases and other rheumatic,
inflammatory or cardiovascular diseases. Written informed consent
was obtained from all participants, and this study was approved by
the Ethics Committee of the Faculty of Medicine, Fayoum University.
The study procedures were performed in accordance with the ethical
code of the World Medical Association Declaration of Helsinki for
human experiments (14).
Sample collection
Fresh venous blood (5 ml) from the antecubital vein
of all participants was collected by venipuncture into EDTA-coated
plain tubes to evaluate the erythrocyte sedimentation rate (ESR)
using the Wintrobe method (15) and
to extract the DNA for reverse transcription-quantitative
(RT-q)PCR. In addition, 3-ml venous blood was collected and
incubated at 37˚C for 15 min and centrifuged at 3,000 x g for 10
min at room temperature to separate the serum, which was then
stored in aliquots at -80˚C.
Biochemical investigations
Serum was used to determine the levels of C-reactive
protein (CRP), anti-cyclic citrullinated peptide (anti-CCP)
antibodies and lncRNA GAS5 levels. CRP and anti-CCP levels were
determined using a commercially available Human C-Reactive Protein
ELISA kit (cat. no. RAB0096; Sigma-Aldrich; Merck KGaA) and an
anti-CCP ELISA kit (Orgentec Diagnostika GmbH).
DNA extraction
Genomic DNA was segregated from whole peripheral
blood using a QIAamp DNA blood mini-kit extraction kit (Qiagen
GmbH) according to the manufacturer's protocol. Purified samples of
DNA were stored at 4˚C until required for PCR (16).
Real-time PCR
This phase included DNA amplification and
identification of rs2067079 (C>T), rs6790 (G>A) and
rs17359906(G>A) SNPs (Table I),
(selected from the lncRNASNP2 database (bioinfo.life.hust.edu.cn/lncRNASNP#!/) and
situated in the chromosome locus transcribed into lncRNA GAS5
utilizing Real MOD™ RT-PCR MasterMix (cat. no. 44248; iNtRON
Biotechnology). The quantity of DNA template used for SNP
genotyping was 20 ng/µl in the reaction mix. DNA concentration was
detected by estimating the absorption at 260 nm on a Nanodrop
ND-1000 spectrophotometer. The sites of genetic information for the
studied SNPs are substantiated by the National Center for
Biotechnology Information database (ncbi.nlm.nih.gov).
 | Table ICharacteristics of the studied
SNPs. |
Table I
Characteristics of the studied
SNPs.
| SNP | Chromosome | Position | Regulatory
feature | Localization |
|---|
| rs2067079 | 1 | 173866073 |
Promoter/Enhancer | Intron |
| rs6790 | 1 | 173865494 |
Promoter/Enhancer | Exon |
| rs17359906 | 1 | 173867056 |
Promoter/Enhancer | Intron |
RNA extraction and RT-qPCR
Total RNA (including small RNAs) were extracted from
serum using an miRNeasy mini kit (Invitrogen; Thermo Fisher
Scientific, Inc.), and RNA concentration was estimated on a
NanoDrop™ 2000 spectrophotometer (Thermo Fisher Scientific, Inc.).
Total RNA was reverse-transcribed into cDNA using a miScript II RT
kit (Qiagen GmbH). Serum expression levels of lncRNA GAS5 were
assessed using GAPDH as the internal control. The sequences of the
primers were: GAS5 forward, 5'-AGGTRGGAGYGGGAGCATTCGG-3' and
reverse, 5'-CCTGAACTGGAGCCACCAGCAGG-3'; and GAPDH forward,
5'-TAATRGGAGCCCGAGCGGGATA-3' and reverse
5'-TTTGCCCTGGAGGGACCAGCAAA-3'. All primers were part of the
RT2 qPCR Primer assay kit (Qiagen GmbH). A Maxima SYBR
Green PCR kit (Thermo Fisher Scientific, Inc.) was used for
amplification according to the manufacturer's protocol, using 0.4
µl of each primer (forward and reverse). The thermocycling
conditions were follows: 95˚C for 1 min; followed by 42 cycles at
95˚C for 10 sec, 60˚C for 30 sec and 72˚C for 1 min. The
2-∆∆Cq method was used normalized to the internal
control to calculate the relative expression levels of lncRNA
GAS5(17).
Statistical analysis
Data were analyzed using SPSS version 25 (IBM,
Corp.). Plain descriptive analysis as numbers and percentages was
performed for qualitative data, using standard deviations as a
measurement of variation of quantitative parametric data and an
inferential statistical tests. A Student's t-test was used to
compare the differences between two groups. Pearson's correlation
analysis was used to assess the correlation between quantitative
variables; the value of the Pearson correlation coefficient (r)
describes the strength and direction of a correlation ranging from
-1 to +1. Receiver operating curve analysis were plotted to
determine the optimal cutoff levels of serum lncRNA GAS5 to
differentiate between cases and controls. The area under the curve
(AUC) was calculated to evaluate the accuracy of the diagnostic
tests. LncRNA GAS5 rs2067079, rs6790 and rs17359906 SNPs in the
patient and control groups were in Hardy-Weinberg equilibrium.
P<0.05 was considered to indicate a statistically significant
difference.
Results
The present study included 350 participants: 200
patients with RA and 150 healthy controls. The mean age did not
differ significantly between the two groups (P=0.245). The number
of females was significantly higher than that of males in each
group; however, there was no significant difference between the two
groups in terms of sex (P=0.774; Table
II).
 | Table IICharacteristics of the study
groups. |
Table II
Characteristics of the study
groups.
| Characteristics | Controls, n=150 | Patients, n=200 | P-value |
|---|
| Agea | 40.85±0.99 | 42.45±0.89 | 0.245 |
| Sex, n (%) | | | 0.774 |
|
Female | 112 (74.6%) | 152 (76%) | |
|
Male | 38 (25.4%) | 48 (24%) | |
The mean levels of ESR, CRP and anti-CCP antibodies
were significantly higher in the patients with RA (54.02±2.31,
2.56±0.10 and 62.40±26.51, respectively) than the controls
(8.05±1.04, 0.59±0.06 and 7.11±4.40, respectively) (P<0.0001).
The mean levels of lncRNA GAS5 were significantly lower in the
patients with RA (0.60±0.031) than in the controls (1.16±0.03)
(P<0.0001; Table III).
 | Table IIILaboratory parameters and lncRNA GAS5
levels in the study groups. |
Table III
Laboratory parameters and lncRNA GAS5
levels in the study groups.
| Factor | Controls,
n=150 | Patients,
n=200 | P-value |
|---|
| Erythrocyte
sedimentation rate, mm/h | 8.05±1.04 | 54.02±2.31 |
<0.0001a |
| C-reactive protein,
mg/dl | 0.59±0.06 | 2.56±0.10 |
<0.0001a |
| Anti-cyclic
citrullinated peptide antibodies, U/ml | 7.11±4.40 | 62.40±26.51 |
<0.0001a |
| lncRNA GAS5 | 1.16±0.03 | 0.60±0.031 |
<0.0001a |
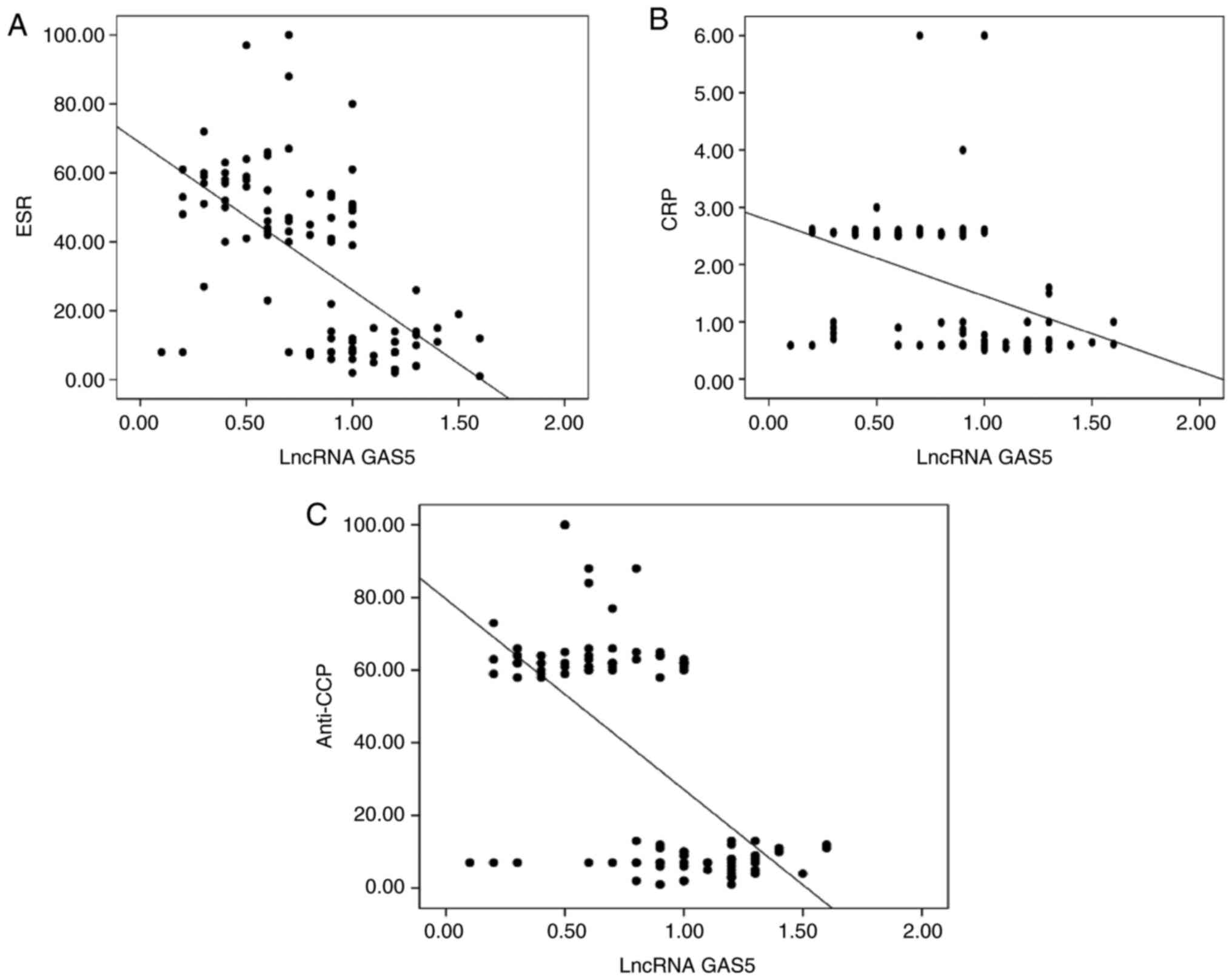
The serum levels of lncRNA GAS5 were significantly
negatively correlated with ESR (r=-0.64, P<0.001), CRP (r=-0.38,
P<0.001), and anti-CCP antibodies (r=-0.58, P<0.001) in RA
patients (Fig. 1).
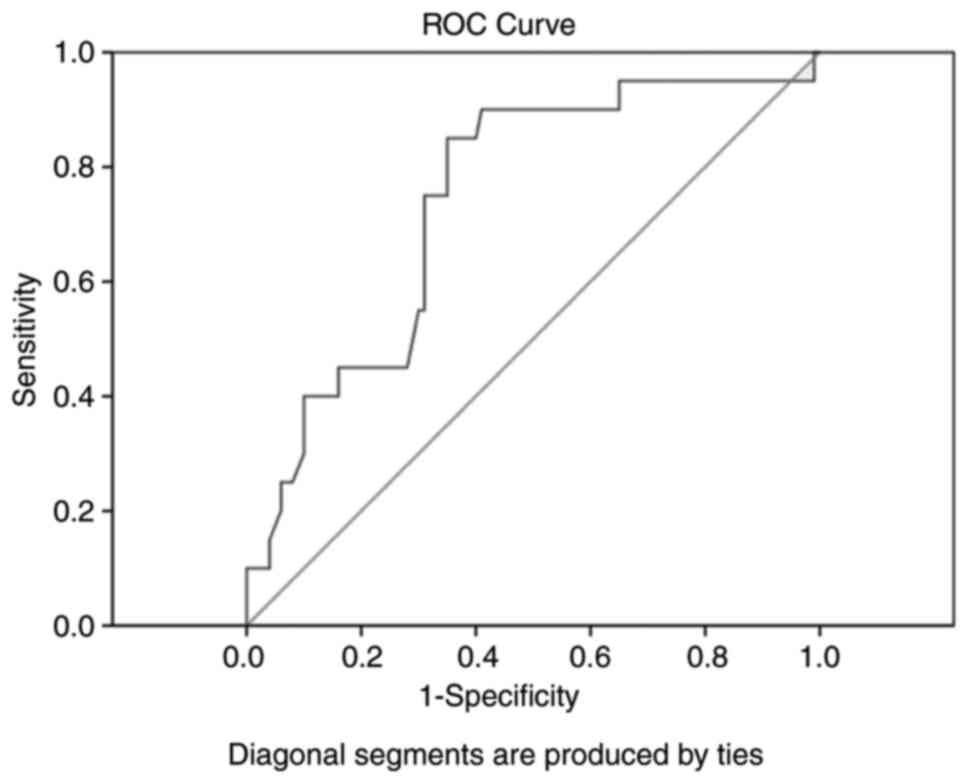
ROC curve analysis revealed high diagnostic value
for lncRNA GAS5 with respect to the diagnosis of RA, with 89%
sensitivity, 70% specificity, 0.75 area under the curve and 76%
accuracy (Fig. 2). Multivariate
regression analysis was used on ESR, CRP, anti-CCP and lncRNA GAS5
levels to determine which parameters independently predicted RA.
The results showed that ESR [odds ratio (OR)=2.89, P=0.03], CRP
(OR=4.09, P=0.01) and anti-CCP antibodies (OR=1.41, P=0.04) were
independent predictors for RA (Table
IV).
 | Table IVMultivariate regression analysis of
laboratory parameters for prediction of rheumatoid arthritis. |
Table IV
Multivariate regression analysis of
laboratory parameters for prediction of rheumatoid arthritis.
| Variable | Odd ratio (95%
confidence interval) | P-value |
|---|
| Erythrocyte
sedimentation rate, mm/h | 2.89
(1.09-7.64) | 0.03a |
| C-reactive protein,
mg/dl | 4.09
(1.35-12.38) | 0.01a |
| Anti-cyclic
citrullinated peptide antibodies, U/ml | 1.41
(0.65-2.91) | 0.04a |
| Long non-coding RNA
growth arrest specific 5 | 0.49
(0.18-1.37) | 0.17 |
Table V shows the
genotype and allele frequencies of rs2067079, rs6790 and rs17359906
SNPs of the lncRNA GAS5 gene in patients with RA and normal
controls. The TT genotype of rs2067079 SNP was significantly
related to a decreased susceptibility of RA [TT vs. C: OR=2.358;
95% confidence interval (CI), 1.114-5.131; P=0.045]. Moreover,
there was a reduced risk of rs2067079 SNP with a recessive pattern
(TT vs. TC + CC: OR=2.374; 95% CI, 1.091-5.123; P=0.037). The A
allele of the rs6790 SNP was present at a higher frequency in the
patients compared with the controls, but the difference was not
statistically significant (P=0.06). The rs6790 SNP was associated
with the risk of RA in the recessive model (AA vs. GA + GG:
OR=2.55; 95% CI, 1.39–5.32; P=0.02). Regarding rs17359906 SNP, no
significant difference was noted between patients and controls with
regard to genotype or allele.
 | Table VGenotype and allele frequencies of
SNPs of lncRNA GAS5 in the different study groups. |
Table V
Genotype and allele frequencies of
SNPs of lncRNA GAS5 in the different study groups.
| A, rs2067079
C>T |
|---|
| Genotype | Controls, n=150
(%) | Patients, n=200,
(%) | P-value | OR (95% CI) |
|---|
|
TT | 6(4) | 3 (1.5) | 0.045a | 2.358
(1.114-5.131) |
|
CT | 36(24) | 50(25) | 0.777 | 0.951
(0.732-1.261) |
|
CC | 108(72) | 147 (73.5) | Reference | |
|
C
allele | 252(84) | 345 (86.2) | 0.396 | 0.942
(0.763-1.148) |
|
T
allele | 48(16) | 55 (13.8) | Reference | |
| Dominant model | | | | 1.042
(0.804-1.343) |
|
CC | 108(72) | 147 (73.5) | 0.8 | |
|
CT + TT | 42(28) | 53 (26.5) | Reference | |
| Recessive
model | | | | 2.374
(1.091-5.123) |
|
TT | 6(4) | 3 (1.5) | 0.037a | |
|
CT + CC | 144(96) | 197 (98.5) | Reference | |
| B, rs6790
G>A |
| Genotype | Controls, n=150
(%) | Patients, n=200,
(%) | P-value | OR (95% CI) |
|
AA | 12(8) | 40(20) | 0.09 | 2.78
(1.265.88) |
|
GA | 66(44) | 80(40) | 0.56 | 1.06
(0.64-1.78) |
|
GG | 72(48) | 80(40) | Reference | |
|
G
allele | 210(70) | 240(60) | 0.06 | 1.49
(1.04-2.12) |
|
A
allele | 90(30) | 160(40) | Reference | |
| Dominant model | | | | 1.33
(0.82-2.15) |
|
GG | 72(48) | 80(40) | 0.49 | |
|
GA + AA | 78(52) | 120(60) | Reference | |
| Recessive
model | | | | 2.55
(1.39-5.32) |
|
AA | 12(8) | 40(20) | 0.02a | |
|
GA + GG | 138(92) | 160(80) | Reference | |
| C, rs17359906
G>A |
| Genotype | Controls, n=150
(%) | Patients, n=200,
(%) | P-value | OR (95% CI) |
|
AA | 16 (10.6) | 18(9) | 0.61 | 1.098
(0.761-1.574) |
|
GA | 70 (46.7) | 89 (44.5) | 0.2 | 1.269
(0.864-1.839) |
|
GG | 64 (42.7) | 93 (46.5) | Reference | |
|
G
allele | 198(66) | 275 (68.8) | 0.07 | 1.149
(0.979-1.349) |
|
A
allele | 102(34) | 125 (31.2) | Reference | |
| Dominant model | | | | 0.839
(0.679-1.044) |
|
GG | 64 (42.7) | 93 (46.5) | 0.13 | |
|
GA + AA | 86 (57.3) | 107 (53.5) | Reference | |
| Recessive
model | | | | 0.848
(0.588-1.209) |
|
AA | 16 (10.6) | 18(9) | 0.36 | |
|
GA + GG | 134 (89.4) | 182(91) | Reference | |
The association between lncRNA GAS5 levels and their
corresponding genotype frequencies in the three SNPs were next
analyzed. No significant differences were noted in the levels of
lncRNA GAS5 amongst the different genotypes in patients with RA
(Table VI).
 | Table VIAssociation of lncRNA GAS5 levels
with their respective genotype in the patients with rheumatoid
arthritis. |
Table VI
Association of lncRNA GAS5 levels
with their respective genotype in the patients with rheumatoid
arthritis.
| Variable | Genotype | n | lncRNA GAS5 | P-value |
|---|
| rs2067079
C>T | CC | 147 | 0.65 | 0.405 |
| | CT | 50 | 0.58 | |
| | TT | 3 | 0.90 | |
| rs6790 G>A | GG | 80 | 0.89 | 0.551 |
| | GA | 80 | 0.56 | |
| | AA | 40 | 0.61 | |
| rs17359906
G>A | GG | 93 | 0.68 | 0.519 |
| | GA | 89 | 0.81 | |
| | AA | 18 | 0.51 | |
Discussion
RA is an aggressive eradicative disorder
characterized by joint destruction. The pathophysiology of RA is
complicated and includes individual (genetic predispositions) and
environmental (bacterial or viral infections) factors (18).
Recent studies have reported that lncRNAs are
associated with numerous biological processes, particularly with
immune cell function and the genetic pathogenesis of immune-related
disorders (4,19).
Furthermore, lncRNAs serve diverse crucial roles,
which involve suppressing the transcriptional machinery,
influencing mRNA stabilization, regulating DNA modulation,
promoting RNA degradation and inducing immune cell apoptosis, in
several physiological processes (20). A previous study has reported that
the expression of ~85 lncRNAs in CD14+ monocytes in
patients with RA is significantly regulated by TNF-α and IL-6
receptor inhibitors. These results suggest that lncRNAs are
involved in RA pathophysiology (21).
LncRNA GAS5, located at chromosome 1q25, which was
initially segregated from mice NIH 3T3 cells, is a small nucleolar
RNA host gene (22,23). LncRNA GAS5 levels are decreased in
CD4+ T and B cells in patients with RA and SLE compared
with control healthy individuals (24). The expression of lncRNA GAS5
increases during growth arrest induced by serum starvation or
suppression of mammalian target of rapamycin (24).
Furthermore, GAS5 can stimulate cell cycle arrest at
the G0/G1 phase by suppressing the transcription of genes, such as
glucocorticoid receptors and androgen, indicating that GAS5
participates in the pathophysiology of RA and SLE by inducing cell
cycle arrest. GAS5 also serves as a microRNA (miR)sponge, such as
for miR-21. As miR-21 is generally implicated in enhancing Th17
cell differentiation (25,26) and participates in the pathogenesis
of several autoimmune disorders, the decreased expression of GAS5
may be involved in the pathophysiology RA and SLE via uncontrolled
miR-21 function in Th17 cells (27). In the present study, the serum
levels of lncRNA GAS5 were investigated in patients with RA and
healthy controls. Considering the criteria for RA diagnosis, ESR,
CRP levels, and anti-CCP antibody levels were assessed and the
results showed that the mean values of these parameters
significantly increased in the patients compared with the controls
(P<0.0001). The mean levels of lncRNA GAS5 significantly
decreased in the patients with RA compared with the controls
(P<0.0001).
These findings are consistent with those of Mayama
et al (10), who
investigated the expression of lncRNA GAS5 in various autoimmune
diseases. They revealed that GAS5 expression significantly
decreased in the CD4+ T and B cells of patients with RA
compared with healthy controls.
Wu et al (28) also reported that GAS5 expression
significantly decreased in the plasma of patients with SLE compared
with healthy controls. Furthermore, Li et al (29) examined three lncRNAs (Linc0597,
lnc-DC and GAS5) in patients with SLE and found that the levels of
these lncRNAs was significantly decreased in patients with SLE
compared with healthy controls.
Several studies have proposed that numerous
lncRNA-related SNPs are associated with the pathophysiology of
autoimmune disorders. The GAS5 gene exists in the disease
susceptibility region in the mice BXSB strain and causes
glomerulonephritis with SLE. A locus on chromosome 9p21, including
lncRNA ANRIL, is significantly associated with predisposition to
type II diabetes. Furthermore, several related SNPs alter the
transcription and processing of ANRIL transcripts in the
pathogenesis of human diseases (30,31).
In the present study, the association between
rs2067079, rs6790 and rs17359906 SNPs of the lncRNA GAS5 gene with
RA risk in the Egyptian population was assessed. The TT genotype of
rs2067079 SNP was significantly associated with a decreased risk of
RA. Moreover, there was a reduced risk of RA with the rs2067079 SNP
with a recessive pattern.
Li et al (29) performed a study to elucidate the
possible association between the rs2067079 C>T SNP of lncRNA
GAS5 and SLE risk, but found no significant association.
Eftekharian et al (32)
reported an association between rs2067079 SNP of lncRNA GAS5 and
multiple sclerosis risk. They found a significantly higher
percentage of the T allele of rs2067079 SNP in patients with MS
than in healthy controls, and showed that this variant was
associated with MS risk in a co-dominant and recessive patterns
(32).
Several studies have revealed that certain SNPs in
the promoter region of GAS5 can regulate its expression and are
associated with the susceptibility to carcinomas. For example,
rs6790 SNP is a biomarker for adverse effects resulting from
chemoradiotherapy in patients with nasopharyngeal carcinoma
(33). CHIP-seq data revealed that
rs6790 SNP is located in an active promoter region or enhancer
region, and silico analysis detected that this variant has a
considerable feature of expression quantitative trait locus. Both
CHIP-seq data and silico analysis revealed the involvement
of rs6790 SNP in the variations of GAS5 expression. In addition,
rs6790 was revealed to exert repressor effects of GAS5 on the
transcriptional activity of glucocorticoid receptor in autoimmune
disorders (32).
The A allele of rs6790 SNP existed at a higher
frequency in the patients compared with the controls, but it was
not statistically significant. rs6790 was associated with the risk
of RA in the recessive model. Wu et al (34) reported that the genotype frequencies
of the rs6790 SNP of lncRNA GAS5 gene was significantly associated
with the risk of RA, but they did not reveal the relation of this
SNP with RA risk with dominant and recessive patterns. Thus,
further studies are required to explain the specific role of lncRNA
GAS5 and its polymorphisms in RA development.
In conclusion serum lncRNA GAS5 levels were
significantly decreased in patients with RA, which indicates that
it is a candidate biomarker for the early diagnosis of RA,
highlighting its potential for use in the early detection of RA for
improved treatment outcomes. The TT genotype of rs2067079 SNP of
lncRNA GAS5 was significantly associated with a decreased risk of
RA. Furthermore, there was a reduced risk of rs2067079 SNP with a
recessive pattern, whereas rs6790 SNP was associated with RA risk
in the recessive model.
Acknowledgements
Not applicable.
Funding
This research did not receive any specific grant from funding
agencies, it was entirely sponsored by Fayoum University under the
supervision of its research committee.
Availability of data and material
The datasets used and/or analyzed during the present
study are available from the corresponding author on reasonable
request.
Authors' contributions
AME, HSES, NKA and SNG performed the experiments and
participated in data analysis. SS recruited the participants and
participated in data analysis. All authors participated in
designing the study, performing statistical analyses and writing
the manuscript. All authors have read and approved the final
manuscript. AME, HSES, NKA, SNG and SS confirm the authenticity of
all the raw data.
Ethics approval and consent to
participate
Written informed consent was obtained from all
participants, and this study was approved by the Ethics Committee
of the Faculty of Medicine, Fayoum University. The study procedures
were performed in accordance with the ethical code of the World
Medical Association Declaration of Helsinki for human
experiments.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Lee DM and Weinblatt ME: Rheumatoid
arthritis. Lancet. 358:903–911. 2001.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Solus JF, Chung CP, Oeser A, Li C, Rho YH,
Bradley KM, Kawai VK, Smith JR and Stein CM: Genetics of serum
concentration of IL-6 and TNFα in systemic lupus erythematosus and
rheumatoid arthritis: A candidate gene analysis. Clin Rheumatol.
34:1375–1382. 2015.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Yamanaka H: TNF as a target of
inflammation in rheumatoid arthritis. Endocr Metab Immune Disord
Drug Targets. 15:129–134. 2015.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Wu GC: Immunoregulation function of long
noncoding RNA in rheumatic diseases. Chin J Dis Control Prev.
20:1165–1171. 2016.
|
|
5
|
Messemaker TC, Frank-Bertoncelj M, Marques
RB, Adriaans A, Bakker AM, Daha N, Gay S, Huizinga TW, Toes REM,
Mikkers HMM and Kurreeman F: A novel long non-coding RNA in the
rheumatoid arthritis risk locus TRAF1-C5 influences C5 mRNA levels.
Genes Immun. 17:85–92. 2016.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Zhang F, Wu L, Qian J, Qu B, Xia S, La T,
Wu Y, Ma J, Zeng J, Guo Q, et al: Identification of the long
noncoding RNA NEAT1 as a novel inflammatory regulator acting
through MAPK pathway in human lupus. J Autoimmun. 75:96–104.
2016.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Haywood MEK, Rose SJ, Horswell S, Lees MJ,
Fu G, Walport MJ and Morley BJ: Overlapping BXSB congenic
intervals, in combination with microarray gene expression, reveal
novel lupus candidate genes. Genes Immun. 7:250–263.
2006.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Wu Y, Zhang F, Ma J, Zhang X, Wu L, Qu B,
Xia S, Chen S, Tang Y and Shen N: Association of large intergenic
noncoding RNA expression with disease activity and organ damage in
systemic lupus erythematosus. Arthritis Res Ther.
17(131)2015.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Kino T, Hurt DE, Ichijo T, Nader N and
Chrousos GP: Noncoding RNA gas5 is a growth arrest- and
starvation-associated repressor of the glucocorticoid receptor. Sci
Signal. 3(ra8)2010.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Mayama T, Marr AK and Kino T: Differential
expression of glucocorticoid receptor Noncoding RNA repressor gas5
in autoimmune and inflammatory diseases. Horm Metab Res.
48:550–557. 2016.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Okada Y, Wu D, Trynka G, Raj T, Terao C,
Ikari K, Kochi Y, Ohmura K, Suzuki A, Yoshida S, et al: Genetics of
rheumatoid arthritis contributes to biology and drug discovery.
Nature. 506:376–381. 2014.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Kumar V, Wijmenga C and Withoff S: From
genome-wide association studies to disease mechanisms: Celiac
disease as a model for autoimmune diseases. Semin Immunopathol.
34:567–580. 2012.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Aletaha D, Neogi T, Silman AJ, Funovits J,
Felson DT, Bingham III CO, Birnbaum NS, Burmester GR, Bykerk VP,
Cohen MD, et al: 2010 Rheumatoid arthritis classification criteria:
An American college of rheumatology/European league against
rheumatism collaborative initiative. Ann Rheum Dis. 69:1580–1588.
2010.PubMed/NCBI View Article : Google Scholar
|
|
14
|
World Medical Association (WMA): WMA
Declaration of Helsinki - Ethical Principles for Medical Research
Involving Human Subjects. WMA, Helsinki, 1964. https://www.wma.net/policies-post/wma-declaration-of-helsinki-ethical-principles-for-medical-research-involving-human-subjects/.
Latest Updated July 9, 2018.
|
|
15
|
Sinton JR: Clinical value of some methods
of estimating erythrocyte sedimentation rate. Br Med J. 1:391–393.
1948.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Salazar LA, Hirata MH, Cavalli SA, Machado
MO and Hirata RD: Optimized procedure for DNA isolation from fresh
and cryopreserved clotted human blood useful in clinical molecular
testing. Clin Chem. 44:1748–1750. 1988.PubMed/NCBI
|
|
17
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
18
|
McInnes IB and Schett G: The pathogenesis
of rheumatoid arthritis. N Engl J Med. 365:2205–2219.
2011.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Wu GC, Pan HF, Leng RX, Wang DG, Li XP, Li
XM and Ye DQ: Emerging role of long noncoding RNAs in autoimmune
diseases. Autoimmun Rev. 14:798–805. 2015.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Iyer MK, Niknafs YS, Malik R, Singhal U,
Sahu A, Hosono Y, Barrette TR, Prensner JR, Evans JR, Zhao S, et
al: The landscape of long noncoding RNAs in the human
transcriptome. Nat Genet. 47:199–208. 2015.PubMed/NCBI View
Article : Google Scholar
|
|
21
|
Müller N, Döring F, Klapper M, Neumann K,
Schulte DM, Türk K, Schröder JO, Zeuner RA, Freitag-Wolf S,
Schreiber S and Laudes M: Interleukin-6 and tumour necrosis
factor-α differentially regulate lincRNA transcripts in cells of
the innate immune system in vivo in human subjects with rheumatoid
arthritis. Cytokine. 68:65–68. 2014.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Zhang Z, Zhu Z, Watabe K, Zhang X, Bai C,
Xu M, Wu F and Mo YY: Negative regulation of lncRNA GAS5 by miR-21.
Cell Death Differ. 20:1558–1568. 2013.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Tu ZQ, Li RJ, Mei JZ and Li XH:
Down-regulation of long non-coding RNA GAS5 is associated with the
prognosis of hepatocellular carcinoma. Int J Clin Exp Pathol.
7:4303–4309. 2014.PubMed/NCBI
|
|
24
|
Pickard MR and Williams GT: Molecular and
cellular mechanisms of action of tumour suppressor GAS5 LncRNA.
Genes (Basel). 6:484–499. 2015.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Murugaiyan G, da Cunha AP, Ajay AK, Joller
N, Garo LP, Kumaradevan S, Yosef N, Vaidya VS and Weiner HL:
MicroRNA-21 promotes Th17 differentiation and mediates experimental
autoimmune encephalomyelitis. J Clin Invest. 125:1069–1080.
2015.PubMed/NCBI View
Article : Google Scholar
|
|
26
|
Dong L, Wang X, Tan J, Li H, Qian W, Chen
J, Chen Q, Wang J, Xu W, Tao C and Wang S: Decreased expression of
microRNA-21 correlates with the imbalance of Th17 and Treg cells in
patients with rheumatoid arthritis. J Cell Mol Med. 18:2213–2224.
2014.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Song J, Ahn C, Chun CH and Jin EJ: A long
non-coding RNA, GAS5, plays a critical role in the regulation of
miR-21 during osteoarthritis. J Orthop Res. 32:1628–1635.
2014.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Wu GC, Li J, Leng RX, Li XP, Li XM, Wang
DG, Pan HF and Ye DQ: Identification of long non-coding RNAs GAS5,
linc0597 and lnc-DC in plasma as novel biomarkers for systemic
lupus erythematosus. Oncotarget. 8:23650–23663. 2017.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Li J, Wu GC, Zhang TP, Yang XK, Chen SS,
Li LJ, Xu SZ, Lv TT, Leng RX, Pan HF and Ye DQ: Association of long
noncoding RNAs expression levels and their gene polymorphisms with
systemic lupus erythematosus. Sci Rep. 7(15119)2017.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Wallace C, Smyth DJ, Maisuria-Armer M,
Walker NM, Todd JA and Clayton DG: The imprinted DLK1-MEG3 gene
region on chromosome 14q32.2 alters susceptibility to type 1
diabetes. Nat Genet. 42:68–71. 2010.PubMed/NCBI View
Article : Google Scholar
|
|
31
|
Pasmant E, Sabbagh A, Vidaud M and Bièche
I: ANRIL, a long noncoding RNA, is an unexpected major hotspot in
GWAS. FASEB J. 25:444–448. 2011.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Eftekharian MM, Noroozi R, Komaki A,
Mazdeh M, Taheri M and Ghafouri-Fard S: GAS5 genomic variants and
risk of multiple sclerosis. Neurosci Lett. 701:54–57.
2019.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Li Q, Ma G, Sun SH, Xu Y and Wang B:
Polymorphism in the promoter region of lncRNA GAS5 is functionally
associated with the risk of gastric cancer. Clin Res Hepatol
Gastroenterol. 42:478–482. 2018.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Wu J, Zhang TP, Zhao YL, Li BZ, Leng RX,
Pan HF and Ye DQ: Decreased H19, GAS5, and linc0597 expression and
association analysis of related gene polymorphisms in rheumatoid
arthritis. Biomolecules. 10(55)2019.PubMed/NCBI View Article : Google Scholar
|