Introduction
Esophageal cancer (ESCA) is one of the most
frequently diagnosed tumors of the digestive system worldwide,
ranking seventh in incidence and sixth in mortality (1). Esophageal squamous cell carcinoma
(ESCC), the main histological subtype of ESCA, is prevalent in
developing countries such as the Middle East and East Asia,
particularly in China (2,3). Although numerous clinical trials have
greatly improved the therapeutic efficacy of ESCC, the 5-year
survival rate of patients with ESCC is ~20% (4). In addition, the lack of effective
diagnostic and therapeutic biomarkers results in poor prognosis for
patients with ESCC (5). Therefore,
it is imperative to identify novel molecular markers to improve the
prognosis and therapeutic efficacy in patients with ESCC.
Peroxiredoxins (PRDXs) constitute a family of
peroxidase enzymes (PRDX1-6) characterized by a catalytic cysteine
residue embedded within the evolutionarily conserved motif PXXXTXXC
(6-8).
From a functional perspective, the six mammalian peroxiredoxins
(PRDX1-6) exhibit distinct characteristics in terms of subcellular
localization, susceptibility to hyperoxidation, substrate
specificity, and the presence or absence of an additional resolving
cysteine residue (9,10). Several studies have demonstrated
that PRDX4, a major factor of redox balance, shields cells from
reactive oxygen species (ROS)-induced oxidative damage by
diminishing peroxides, such as hydrogen peroxide
(H2O2) (11,12).
Elevated ROS levels have been implicated in the pathological and
physiological processes of a broad range of diseases, such as
cancer and neurodegenerative disorders (13,14).
In addition, tumor cells often experience oxidative stress due to
increased ROS levels (15), which
in turn results in damaged biomacromolecules (lipids, proteins, and
DNA), thus triggering strong cell toxicity (16). Wang et al (17) found that PRDX4 knockdown elicited
ROS overproduction in hepatocellular carcinoma (17). PRDX4 has been widely reported to
participate in the development and progression of various tumors,
such as breast cancer (18),
hepatoblastoma (19), renal cell
carcinoma (20), uterine corpus
endometrial carcinoma (21),
pancreatic ductal adenocarcinoma (22), and gastric cancer (23). Moreover, PRDX4 has been developed as
a biomarker and a novel therapeutic target for renal papillary cell
carcinoma and pancreatic cancer (24). These studies suggest that targeting
PRDX4 may be an attractive therapeutic strategy for a variety of
tumors. However, whether PRDX4-mediated tumor-promoting effects
make it an attractive target for patients with ESCC remains to be
investigated.
Ferroptosis, firstly discovered in 2012 is a type of
iron-dependent regulated cell death caused by excessive lipid
peroxidation (25), and is
different from other programmed cell death processes, including
apoptosis, necrosis, and pyroptosis (26). Numerous antioxidants and other
related factors, such as glutathione peroxidase 4 (GPX4),
ferroptosis suppressor protein 1, and dihydroorotate dehydrogenase,
have been shown to inhibit ferroptosis (27-29).
Several studies have revealed the versatile functions of
ferroptosis in numerous tumors, including cell growth, metastasis,
immune surveillance, and therapeutic response (30-33).
Recent studies have focused on the relationship between PRDX
proteins and ferroptosis. For example, genetic ablation or
knockdown of PRDX6 was shown to sensitize lung endothelial cells to
erastin-induced ferroptosis (34).
In addition, loss of PRDX6 downregulated selenoprotein expression
and induced ferroptosis through GPX4 suppression (35). A previous study revealed that PRDX4
knockdown markedly promotes ROS production and induces ferroptosis
in hepatic fibrosis (36). These
studies indicate a strong association between PRDX and ferroptosis
and suggest that targeting ferroptosis may be an emerging novel
therapeutic strategy for patients with ESCC. However, whether PRDX4
participates in the regulation of ferroptosis and its precise
molecular mechanisms in ESCC remain unclear.
The aim of the present study was to investigate the
role of PRDX4 in ESCC and its underlying mechanism. PRDX4
expression was analyzed in clinical ESCC samples and cell lines,
and its association with clinicopathological features was assessed.
PRDX4 knockdown and overexpression models were used to evaluate its
effects on cell proliferation, invasion, and ferroptosis. The
involvement of the PI3K/AKT signaling pathway was examined to
elucidate the molecular mechanism.
Materials and methods
Bioinformatics analysis
PRDX4 expression and pan-cancer prognostic values
were analyzed using the Sangerbox 3.0 online database (http://vip.sangerbox.com/home.html). PRDX4
expression in ESCA and its association with patient survival was
investigated using the UALCAN online database (http://ualcan.path.uab.edu/). The effect of PRDX4 on
the overall survival of patients with ESCA was analyzed using the
online databases, UALCAN and GEPIA (http://gepia.cancer-pku.cn/). GSE111011 was downloaded
from the GEO database (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE111011)
for PRDX4 expression assay.
ESCC samples, cell lines, and cell
cultures
A total of 65 ESCC samples along with paired normal
samples were procured from the First Affiliated Hospital of
Zhengzhou University (Zhengzhou, China). Patients with ESCC who had
not received chemotherapy or radiotherapy were selected and
provided written informed consent. Among the aforementioned cases,
there were 39 males and 26 females, with a median age of 61 years
(age range, 25-83 years). The experiments were approved (approval
no. ZZUIRB 2023-239) by the Research and Ethics Committee of
Zhengzhou University (Zhengzhou, China). The human ESCC cell line
KYSE30 was purchased from Procell Life Science &Technology Co.,
Ltd. Normal esophageal epithelial cell line Het-1A and human ESCC
cell lines KYSE520, KYSE70, KYSE450, and KYSE270, which were
authenticated by STR profiling, were purchased from Qingqi Shanghai
Biotechnology Development Co., Ltd. ESCC cells were cultured in
RPMI-1640 medium (Procell Life Science & Technology Co., Ltd.)
supplemented with 10% fetal bovine serum (FBS; Suzhou ShuangRu
Biotech Co., Ltd.), as well as penicillin (100 U/ml) and
streptomycin (0.1 mg/ml) solution (Solarbio Life Sciences), and
were incubated at 37˚C with 5% CO2.
Transfection of siRNA and plasmid
ESCC cells (KYSE30 and KYSE270) were plated in
6-well plates at 2x105 cells per well and cultured in a
CO2 incubator for 24 h. When the cell confluency reached
~80%, 1.5 µg empty pcDNA3.1 vector and 1.5 µg PRDX4 overexpression
vector, pcDNA3.1-PRDX4, were used to transfect into KYSE30 cells
using Lipo8000™ Transfection Reagent (cat. no. C0533;
Beyotime Institute of Biotechnology) according to manufacturer's
protocol for 48 h at 37˚C. The old medium was replaced with fresh
medium 6 h after transfection. The transfected cells were then
incubated for an additional 48 h at 37˚C before being harvested for
further analyses. In addition, a control siRNA (50 nM; cat. no.
sc-37007; Santa Cruz Biotechnology, Inc.) and PRDX4 siRNA (50 nM;
cat. no. sc-40835; Santa Cruz Biotechnology, Inc.) were transfected
into KYSE270 cells using Lipo8000™ Transfection Reagent
according to the manufacturer's instructions for 48 h at 37˚C. The
old medium was replaced with fresh medium 6 h after transfection.
The transfected cells were then incubated for an additional 48 h at
37˚C before being harvested for further analyses. Both the control
siRNA and PRDX4 siRNA were commercially purchased from Santa Cruz
Biotechnology, Inc. as validated, ready-to-use reagents with
demonstrated high knockdown efficiency; however, the manufacturer's
product documentation did not disclose the specific siRNA
sequences.
Cell Counting Kit-8 (CCK-8) assay
A CCK-8 kit (cat. C0038; Beyotime, China) was used
to assess cell proliferation. In brief, KYSE30 and KYSE270 cells
with different transfection efficiencies (3,000 cells per well)
were plated in a 96-well plate. Cell viability was measured at 24,
48, 72 and 96 h post-seeding by adding 10 µl of CCK-8 reagent to
each well, followed by incubation for 2 h at 37˚C. The absorbance
value at 450 nm was determined at 24, 48, 72 and 96 h using a
microplate reader.
Colony formation assay
A colony formation assay was performed to evaluate
the colony formation ability of KYSE30 and KYSE270 cells. Briefly,
transfected ESCC cells were plated in 6-well plates at a density of
1x103 cells per well. The culture medium was replaced
with fresh medium, and the cells were cultured for 9-12 days.
Finally, the cells were fixed with 4% paraformaldehyde (cat. no.
P1110; Solarbio Life Sciences) for 20 min at room temperature and
stained with 0.1% crystal violet (cat. no. IC0600; Solarbio Life
Sciences) for 30 min at room temperature. Colonies containing
>50 cells were counted using ImageJ 1.8.0 software (NIH).
EdU staining
EdU staining was performed to detect the
proliferative abilities of the ESCC cells. Briefly, transfected
KYSE30 and KYSE270 cells were plated in 48-well plates at
5x103 cells per well for 48 h. EdU (1:1,000 dilution;
cat. no. C10310-1; Guangzhou RiboBio Co., Ltd.) was added to each
well and incubated for 2 h at 37˚C. The cells were then fixed with
4% paraformaldehyde for 30 min at room temperature. Subsequently,
glycine solution was added to each well for 5 min, and the
penetrant reagent was added to each well for another 10 min. Next,
1X Apollo® staining solution (included in kit cat. no.
C10310-1; Guangzhou RiboBio Co., Ltd.) was added to each well for
30 min at room temperature. Finally, ESCC cells were treated with
Hoechst (1:100 dilution; included in kit cat. no. C10310-1;
Guangzhou RiboBio Co., Ltd.) for 30 min at room temperature.
Fluorescence images were acquired using an inverted fluorescence
microscope, and at least five random fields of view were captured
per well. EdU-positive cells were quantified using ImageJ 1.8.0
software [National Institutes of Health (NIH)].
Cell migration
A 24-well Transwell chamber (8-µm pores; Corning,
Inc.) was used to examine the cell migration ability. Transfected
KYSE30 and KYSE270 cells (5x103) cultured in 200 µl of
FBS-free medium were seeded into a Matrigel-uncoated upper chamber
(Transwell chamber, Corning, Inc.), and the lower chamber was
filled with 600 µl of medium containing 20% FBS. Cells were
maintained at 37˚C in a 5% CO2 incubator for 48 h, fixed
using 4% paraformaldehyde for 30 min at room temperature, stained
with 0.1% crystal violet for 30 min at room temperature, and washed
with PBS. Images were captured using an optical microscope.
Cell invasion
Cell invasion was investigated using a 24-well
Transwell chamber with Matrigel (BD Biosciences, Inc.), and
Matrigel-coated inserts were thawed and rehydrated with serum-free
medium for 1 h at 37˚C prior to cell seeding. Transfected KYSE30
and KYSE270 cells (5x103) in serum-free medium were
added to the top chamber of each well, and 600 µl complete medium
supplemented with 20% FBS was applied to the lower chamber. The
cells were incubated at 37˚C in a 5% CO2 incubator for
48 h. Subsequently, the invasive cells were fixed using methanol
for 30 min at room temperature, and stained with crystal violet for
30 min at room temperature. The stained invasive cells were counted
using an optical microscope.
Analysis of malondialdehyde (MDA),
lipid peroxidation (LPO) and glutathione (GSH) levels
KYSE30 and KYSE270 cells (1x106) were
harvested 48 h after transfection and seeded into 6-well plates.
The cells were then treated with erastin (5 µM), ferrostatin-1
(Fer-1; 10 µM), 740 Y-P (20 µM) and LY294002 (20 µM) (all from
TargetMol Chemicals Corporation) for 24 h at 37˚C. Lipid
peroxidation indicators were measured according to the
manufacturer's protocol. MDA levels were detected using an MDA
Content Detection kit (cat. no. S0131S; Beyotime Institute of
Biotechnology), LPO levels were determined using an LPO Content
Assay kit (cat. no. BC5245; Solarbio Life Sciences), and the GSH
levels were measured using a GSH Detection kit (cat. no. BC1175;
Solarbio Life Sciences).
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA was extracted using TRIzol reagent
(Solarbio Life Sciences), and transcribed into cDNA using the
Prime-Script RT kit (Takara Bio, Inc.). cDNA was then added to the
reaction system for PCR amplification using the SYBR Green PCR
Master Mix (Takara Bio, Inc.). Subsequently, RT-qPCR was performed
in 384-well plates using the real-time PCR system,
LightCycler® 480 (Roche Diagnostics) according to the
manufacturer's protocol. The following primers were used: PRDX4
(NM_006406, product size: 171 bp) forward,
5'-CGAAGATTTCCAAGCCAGCG-3' and reverse, 5'-CAAGTCTGTCGCCAAAAGCG-3';
β-actin (NM_001101, produce size: 192 bp) forward:
5'-AACTGGGACGACATGGAGAAAA-3' and reverse,
5'-GGATAGCACAGCCTGGATAGCA-3'. β-actin was used as the control.
Finally, the relative level of PRDX4 was analyzed from three
independent experiments using the
2-∆∆Cq method (37).
Western blotting
Total protein was extracted using RIPA lysis buffer
(Solarbio Life Sciences). Protein concentration was determined
using a BCA Protein Assay kit (Solarbio Life Sciences). Total
proteins (50 µg) from each group were separated using 10% SDS-PAGE
and transferred onto PVDF membranes (MilliporeSigma). After
blocking with 5% skimmed milk, PVDF membranes were incubated with
primary antibodies at 4˚C overnight. The primary antibodies were as
follows: Anti-PRDX4 (1:2,000 dilution; cat. no. 10703-1-AP),
anti-E-cadherin (1:20,000 dilution; cat. no. 20874-1-AP),
anti-N-cadherin (1:2,000 dilution; cat. no. 22018-1-AP),
anti-Vimentin (1:20,000 dilution; cat. no. 10366-1-AP), and
anti-GPX4 (1:1,000 dilution; cat. no. 30388-1-AP; all from
Proteintech Group, Inc.), anti-solute carrier family 7 member 11
(SLC7A11; 1:1,000 dilution; cat. no. DF12509; Affinity Biosciences,
Ltd.), anti-prostaglandin-endoperoxide synthase 2 (PTGS2; 1:1,000
dilution; cat. no. 27308-1-AP; Proteintech Group, Inc.),
anti-p-PI3K (1:1,000 dilution; cat. no. AP0427; ABclonal Biotech
Co., Ltd.), anti-PI3K (1:1,000 dilution; cat. no. A19742; ABclonal
Biotech Co., Ltd.), anti-p-AKT (1:2,000 dilution; cat. no.
28731-1-AP; Proteintech Group, Inc.), anti-AKT (1:2,000 dilution;
cat. no. 10176-2-AP; Proteintech Group, Inc.) and anti-β-actin
(1:2,000 dilution; cat. no. 20536-1-AP; Proteintech Group, Inc.).
After washing with Tris-buffered saline (TBS) with 0.05% Tween 20,
the PVDF membranes were incubated with horseradish peroxidase
(HRP)-labeled secondary antibody (1:10,000 dilution; cat. no.
RGAR001; Proteintech Group, Inc.) for 2 h at 37˚C, and then rinsed
with TBST. Subsequently, the PVDF membranes were incubated with
chemiluminescent HRP substrate (Affinity Biosciences, Ltd.).
Finally, the relative protein levels were analyzed using the ImageJ
1.8.0 software (NIH).
Immunohistochemistry (IHC)
The tissue sections were fixed with 10% formalin for
24 h at room temperature, embedded in paraffin, and continuously
cut into 4-6 µm sections. After dewaxing, rehydration, and antigen
retrieval in citrate buffer (pH 6.0) using microwave heating and
blocking with 10% goat serum (cat. no. C0265; Beyotime Insitute of
Biotechnology) for 1 h at room temperature, tissue sections were
incubated with anti-PRDX4 (1:2,000 dilution; cat. no. 10703-1-AP;
Proteintech Group, Inc.) overnight at 4˚C. After washing, the
corresponding HRP-labeled secondary antibody (1:5,000; cat. no.
RGAR001; Proteintech Group, Inc.) were added to the tissue slides.
The DAB reagent was used to generate the staining signals. The
images were obtained using an optical microscope.
Statistical analysis
All experimental data are expressed as the mean ±
SD, and were analyzed using GraphPad Prism software (version 8.0)
for statistical assay. All experiments were independently repeated
at least three times with distinct biological samples. Prior to
parametric analyses, a normality distribution test was conducted to
assess the data. Student's t-test or Mann-Whitney U test were
utilized to compare two groups, and Differences among three or more
groups were assessed by one-way ANOVA followed by Tukey's HSD post
hoc test. P<0.05 was considered to indicate a statistically
significant difference.
Results
PRDX4 is highly expressed in ESCA
PRDX4 expression was investigated in ESCA using TCGA
database. PRDX4 was highly expressed in all the investigated tumor
types, with low expression in pancreatic adenocarcinoma (PAAD) and
kidney chromophobe (KICH) tumors (Fig.
1A). Data from the UALCAN database revealed that the expression
of PRDX4 in ESCA samples was significantly higher than that in
normal samples (Fig. 1B), which was
also validated by GEO dataset GSE111011 (Fig. 1C). In addition, RT-qPCR revealed
that the mRNA levels of PRDX4 in 65 ESCC samples were significantly
higher than those in the paired normal samples (Fig. 1D). Furthermore, the expression of
PRDX4 was examined in eight ESCC samples and paired normal samples
by western blotting and it was found that the expression of PRDX4
in all ESCC samples was higher than that in paired normal samples
(Fig. 1E and F), which was also verified by IHC assay
(Fig. 1G). Further investigation
revealed that PRDX4 expression in a panel of ESCC cell lines was
significantly higher than that in the normal esophageal epithelial
cell line Het-1A (Fig. 1H-J). These
findings indicated that PRDX4 was highly expressed in ESCC tissues
and cells.
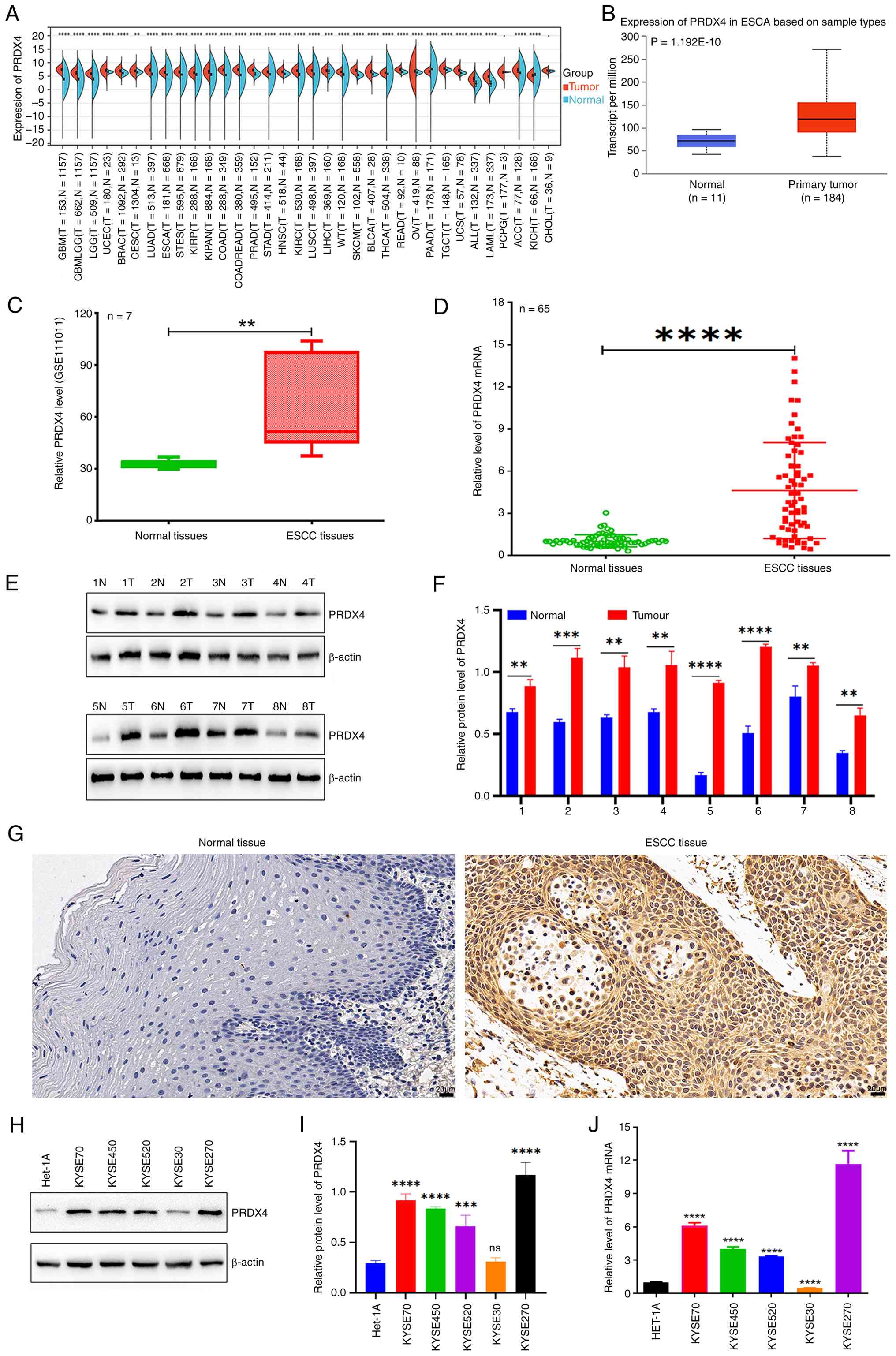 | Figure 1PRDX4 exhibits high expression in
ESCC tissues and cells. (A) Sangerbox 3.0 online software assay for
PRDX4 expression in pan-cancer. (B) UALCAN database investigation
for PRDX4 expression in ESCA samples and normal esophageal
epithelial tissues. (C) GEO dataset GSE111011 was used to
investigate the expression of PRDX4 in ESCC samples and paired
normal samples. (D) RT-qPCR assay was used to assess the expression
of PRDX4 in 65 ESCC samples and paired normal samples. (E) Western
blot analysis of the protein expression of PRDX4 in eight ESCC
samples and paired normal samples. (F) The relative protein levels
of PRDX4 in eight ESCC samples and paired normal samples. (G) IHC
detection of PRDX4 expression in normal tissues and ESCC tissues.
Scale bar, 20 µm. (H) Western blot analysis of PRDX4 protein
expression in ESCC cell lines (KYSE70, KYSE450, KYSE520, KYSE30 and
KYSE270) and normal esophageal epithelial cell line Het-1A. (I) The
relative protein levels of PRDX4 in ESCC cell lines and Het-1A
cells. (J) RT-qPCR assay of PRDX4 mRNA expression in the
aforementioned ESCC cell lines and Het-1A cells.
**P<0.01, ***P<0.001 and
****P<0.0001, indicate statistical significance.
PRDX4, peroxiredoxin 4; ESCC, esophageal squamous cell carcinoma;
ESCA, esophageal carcinoma; RT-qPCR, reverse
transcription-quantitative polymerase chain reaction; GEO, Gene
Expression Omnibus; IHC, immunohistochemistry; ns, not
significant. |
High PRDX4 expression as a potential
prognostic marker in ESCC
To further dissect the clinical value of PRDX4 in
ESCC, TCGA database was used to investigate the expression of PRDX4
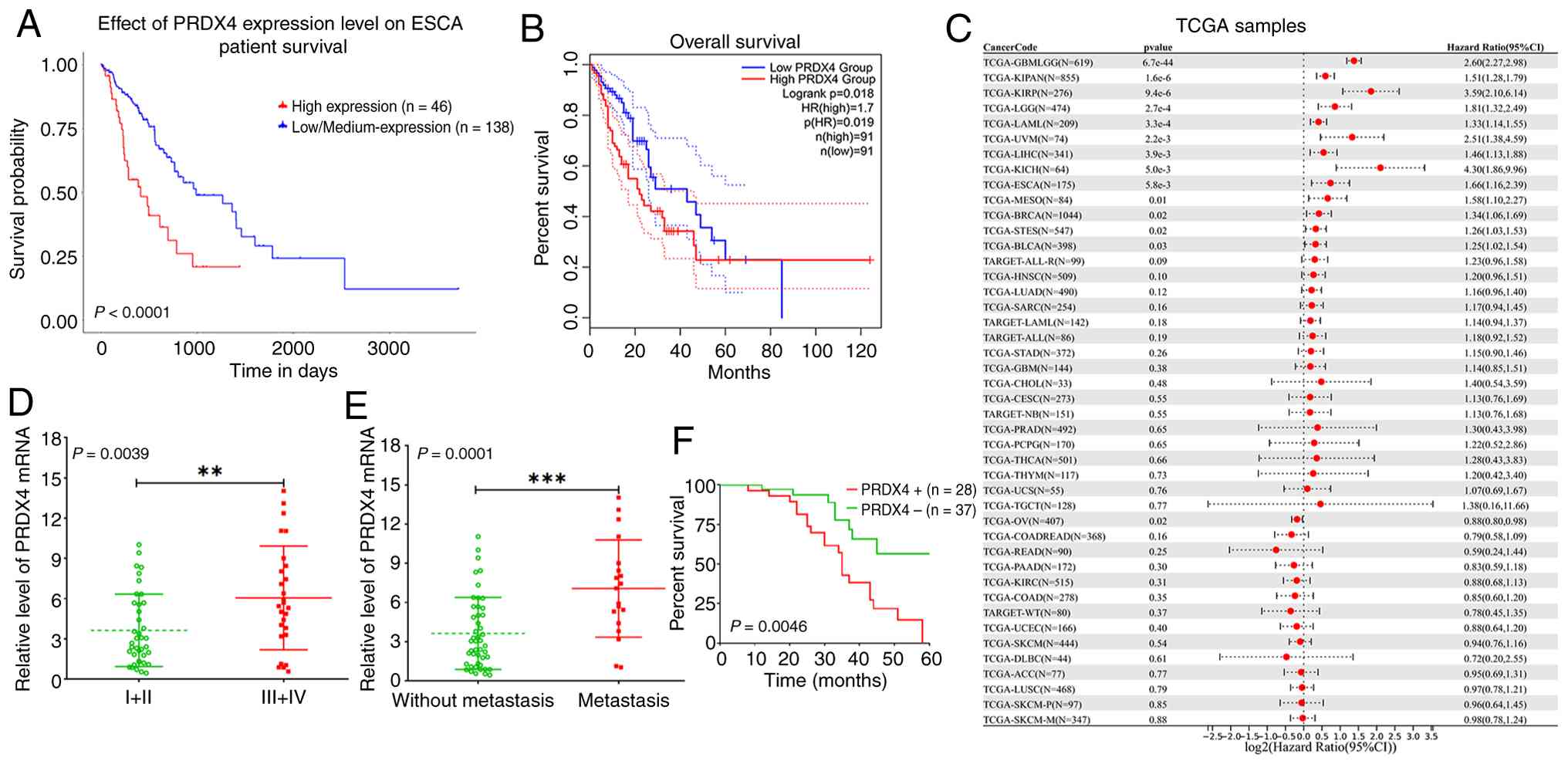
on the survival rate of patients with ESCA. Patients with high
PRDX4 expression had lower survival rates than those with low PRDX4
expression (Fig. 2A and B). Moreover, high PRDX4 expression
predicted a poor prognosis in patients with ESCA (Fig. 2C). Further investigation revealed
that high PRDX4 expression was closely associated with TNM staging
and lymph node metastasis (Fig. 2D
and E). Notably, patients with ESCC
with high PRDX4 expression had lower survival rates than those with
low PRDX4 expression (Fig. 2F).
These findings indicated that PRDX4 is involved in ESCC progression
and may serve as a novel prognostic marker for patients with this
disease.
PRDX4 knockdown suppresses cell
proliferation in ESCC cells
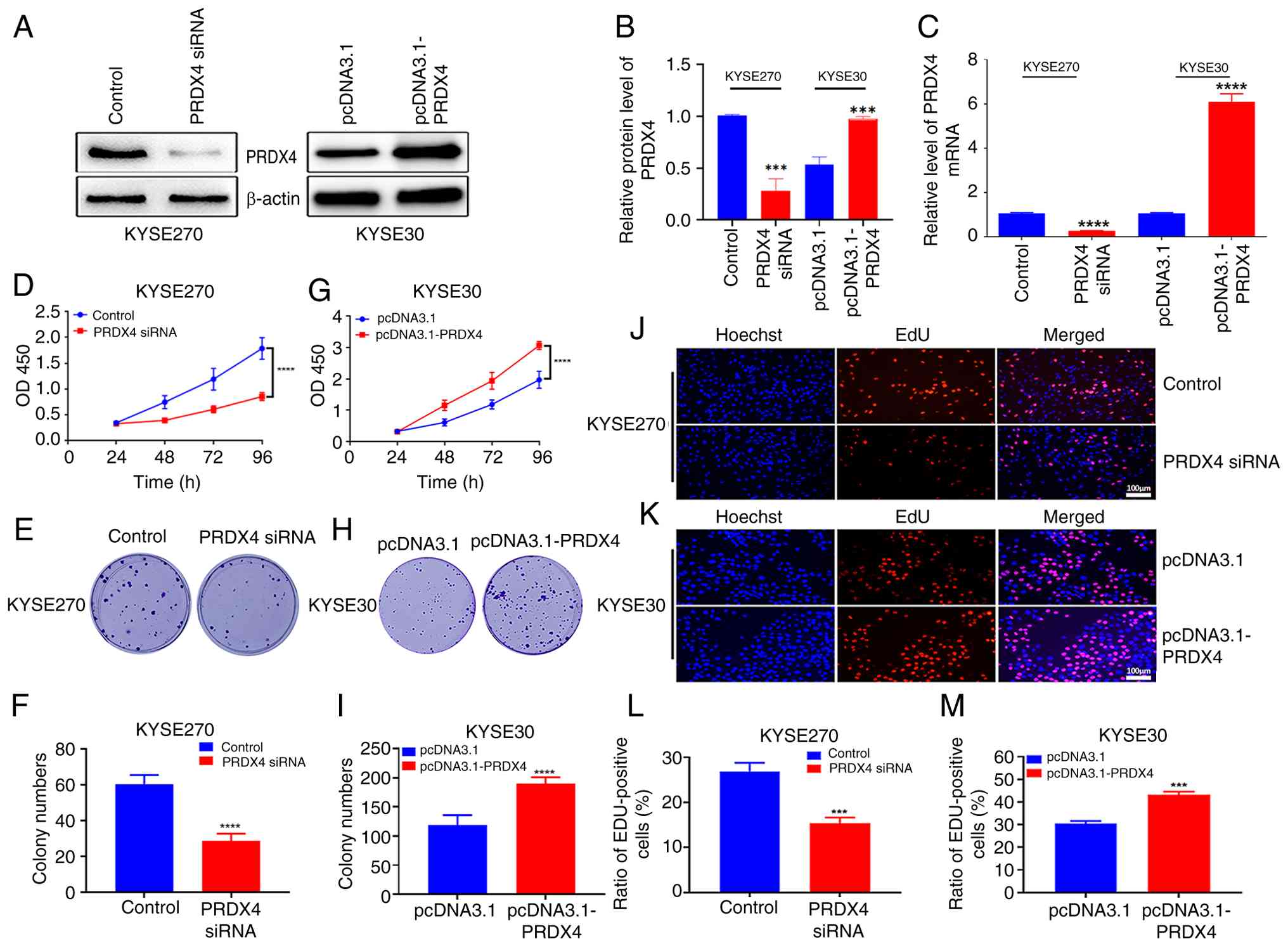
To further explore the biological role of PRDX4 in
ESCC, ESCC cells were transfected with PRDX4 siRNA and the
overexpression vector pcDNA3.1-PRDX4. PRDX4 siRNA significantly
downregulated PRDX4 expression in KYSE520 cells with high PRDX4
expression (Fig. 3A and B), whereas pcDNA3.1-PRDX4 markedly
upregulated PRDX4 expression in KYSE30 cells with low PRDX4
expression (Fig. 3A and B), which was further validated using
RT-qPCR (Fig. 3C). Further
investigation indicated that PRDX4 depletion inhibited cell
proliferation and colony formation (Fig. 3D-F), whereas PRDX4 overexpression
had the opposite effects (Fig.
3G-I). Furthermore, EdU staining revealed that PRDX4
downregulation markedly reduced the number of EdU-positive cells,
whereas PRDX4 overexpression markedly increased the number of
EdU-positive cells (Fig. 3J-M).
These findings indicated that PRDX4 functions as an oncogene in
ESCC cells.
PRDX4 depletion inhibits cell
migration and invasion in ESCC cells
In the present study, the role of PRDX4 in ESCC cell
migration and invasion was explored. The results revealed that
PRDX4 downregulation markedly suppressed ESCC cell migration and
invasion (Fig. 4A-C). Western blot
analysis demonstrated that PRDX4 knockdown markedly promoted the
expression of E-cadherin, but reduced the expression of N-cadherin
and vimentin (Fig. 4D and E). Conversely, PRDX4 overexpression
significantly increased the migration and invasion abilities of
ESCC cells (Fig. 4F-H), reduced
E-cadherin expression, and increased N-cadherin and vimentin
expression (Fig. 4I and J). These findings indicated that PRDX4 is
an important regulator of ESCC cell invasion.
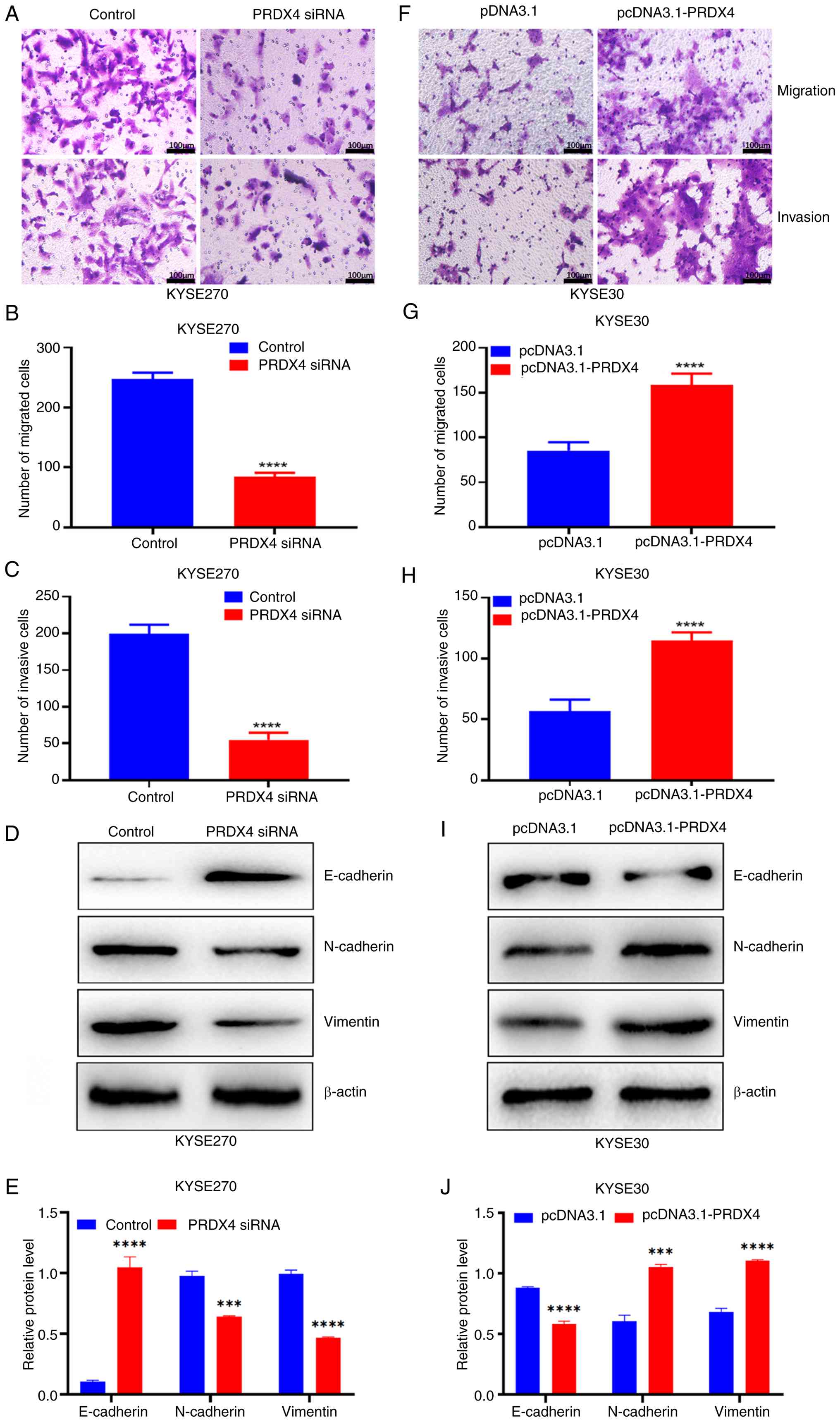 | Figure 4PRDX4 downregulation suppresses cell
migration and invasion in ESCC cells. (A) PRDX4 knockdown
suppresses cell migration and invasion in KYSE270 cells after
transfection with PRDX4 siRNA. Scale bar, 100 µm. (B) Statistical
analysis of the number of migratory cells in KYSE270 cells
transfected with PRDX4 siRNA. (C) Statistical analysis of the
number of invasive cells in KYSE270 cells transfected with PRDX4
siRNA. (D) Western blot analysis of the expression levels of
E-cadherin, N-cadherin and vimentin in KYSE270 cells transfected
with PRDX4 siRNA. (E) The relative protein levels of E-cadherin,
N-cadherin and vimentin in KYSE270 cells transfected with PRDX4
siRNA. (F) PRDX4 overexpression suppresses cell migration and
invasion in KYSE30 cells after transfection with pcDNA3.1-PRDX4.
Scale bar, 100 µm. (G) Statistical analysis of the number of
migratory cells in KYSE30 cells transfected with pcDNA3.1-PRDX4.
(H) Statistical analysis of the number of invasive cells in KYSE30
cells transfected with pcDNA3.1-PRDX4. (I) Western blot analysis of
the expression levels of E-cadherin, N-cadherin and vimentin in
KYSE30 cells transfected with pcDNA3.1-PRDX4. (J) The relative
protein levels of E-cadherin, N-cadherin and vimentin in KYSE30
cells transfected with pcDNA3.1-PRDX4. ***P<0.001 and
****P<0.0001, indicate statistical significance.
PRDX4, peroxiredoxin 4; ESCC, esophageal squamous cell carcinoma;
siRNA, small interfering RNA. |
PRDX4 knockdown promotes ferroptosis
in ESCC cells
Considering the important role of PRDX4 in oxidative
stress, it was investigated whether PRDX4 affected ferroptosis in
ESCC cells. It was revealed that PRDX4 depletion increased the
levels of MDA and LPO but reduced the levels of GSH in ESCC cells
(Fig. 5A). Furthermore, western
blotting revealed that PRDX4 knockdown reduced the expression of
ferroptosis inhibitory factors, such as GPX4 and SLC7A11, but
increased the levels of the ferroptosis-promoting factor, PTGS2
(Fig. 5B and C). By contrast, PRDX4 overexpression
significantly reduced the levels of MDA and LPO, but increased the
levels of GSH in ESCC cells (Fig.
5D), accompanied by enhanced GPX4 and SLC7A11 levels and
decreased PTGS2 levels (Fig. 5E and
F). Notably, Fer-1 partly reversed
the changes in the levels of MDA, LPO, and GSH, as well as the
expression of GPX4, SLC7A11, and PTGS2 proteins induced by PRDX4
siRNA in KYSE270 cells (Fig. 5G-I).
Similarly, erastin effectively induced ferroptosis, which was
suppressed by PRDX4 overexpression in KYSE30 cells, coupled with
alterations in the levels of key ferroptosis-related proteins, such
as GPX4, SLC7A11, and PTGS2 (Fig.
5J-L). These findings indicated that PRDX4 may participate in
the regulation of ferroptosis in ESCC cells.
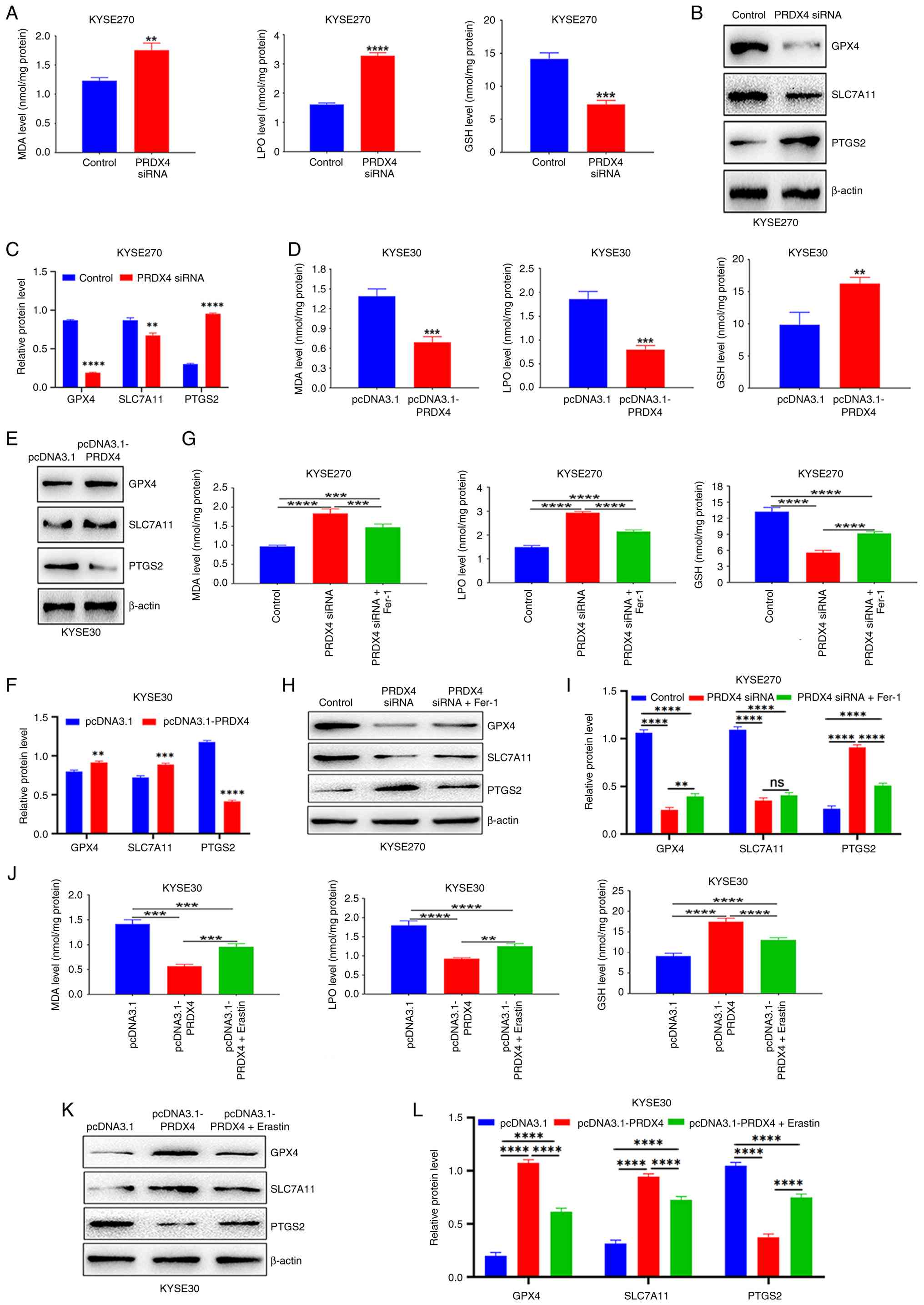 | Figure 5PRDX4 is an important regulator of
ferroptosis in ESCC cells. (A) Determination of MDA, LPO and GSH
contents in KYSE270 cells after transfection with PRDX4 siRNA. (B)
Western blot analysis of the protein levels of GPX4, SLC7A11 and
PTGS2 in KYSE270 cells transfected with PRDX4 siRNA. (C) The
relative protein levels of GPX4, SLC7A11 and PTGS2 in KYSE270 cells
transfected with PRDX4 siRNA. (D) Determination of MDA, LPO and GSH
contents in KYSE30 cells after transfection with pcDNA3.1-PRDX4.
(E) Western blot analysis of the protein levels of GPX4, SLC7A11
and PTGS2 in KYSE30 cells transfected with pcDNA3.1-PRDX4. (F) The
relative protein levels of GPX4, SLC7A11 and PTGS2 in KYSE30 cells
transfected with pcDNA3.1-PRDX4. (G) Detection of the levels of
MDA, LPO and GSH in the control group, PRDX4 siRNA group and PRDX4
siRNA plus Fer-1 group in KYSE270 cells. (H) Western blot analysis
of the protein expression levels of GPX4, SLC7A11 and PTGS2 in the
control group, PRDX4 siRNA group and PRDX4 siRNA plus Fer-1 group
in KYSE270 cells. (I) The relative protein levels of GPX4, SLC7A11
and PTGS2 in the control group, PRDX4 siRNA group and PRDX4 siRNA
plus Fer-1 group in KYSE270 cells. (J) Detection of the levels of
MDA, LPO and GSH in the pcDNA3.1 group, pcDNA3.1-PRDX4 group and
pcDNA3.1-PRDX4 plus erastin group in KYSE30 cells. (K) Western blot
analysis of the protein expression levels of GPX4, SLC7A11 and
PTGS2 in the pcDNA3.1 group, pcDNA3.1-PRDX4 group and
pcDNA3.1-PRDX4 plus erastin group in KYSE30 cells. (L) The relative
protein levels of GPX4, SLC7A11 and PTGS2 in the pcDNA3.1 group,
pcDNA3.1-PRDX4 group and pcDNA3.1-PRDX4 plus erastin group in
KYSE30 cells. **P<0.01, ***P<0.001 and
****P<0.0001, indicate statistical significance.
PRDX4, peroxiredoxin 4; ESCC, esophageal squamous cell carcinoma;
siRNA, small interfering RNA; MDA, malondialdehyde; LPO, lipid
peroxidation; GSH, glutathione; GPX4, glutathione peroxidase 4;
SLC7A11, solute carrier family 7 member 11; PTGS2,
prostaglandin-endoperoxide synthase 2; Fer-1, ferrostatin-1; ns,
not significant. |
PRDX4 suppresses ferroptosis by
regulating the PI3K/AKT signaling pathway
To further elucidate the potential molecular
mechanisms of PRDX4 in the regulation of ferroptosis, the effects
of PRDX4 on the PI3K/AKT signaling pathway in ESCC cells were
explored. It was found that the increased levels of MDA and LPO and
decreased levels of GSH, upon PRDX4 depletion were partly reversed
by the PI3K activator, 740 Y-P (Fig.
6A). Notably, this activator significantly reduced the levels
of MDA and LPO and enhanced the levels of GSH (Fig. 6A). Further investigation revealed
that PRDX4 depletion reduced the expression of GPX4, p-PI3K, and
p-AKT, which was partially recovered after 740 Y-P treatment
(Fig. 6B and C). Furthermore, the decrease in the levels
of MDA and LPO and the increase in the levels of GSH induced by
PRDX4 overexpression were significantly reversed after treatment
with the PI3K inhibitor, LY294002 (Fig.
6D). Notably, LY294002 significantly increased the levels of
MDA and LPO, but reduced the levels of GSH (Fig. 6D). In addition, western blotting
demonstrated that PRDX4 promoted the expression of GPX4, p-PI3K,
and p-AKT, and this effect was reversed when PRDX4 was
overexpressed in combination with LY294002 (Fig. 6E and F). These findings indicated that PRDX4
suppresses ferroptosis by activating the PI3K/AKT signaling pathway
in ESCC cells.
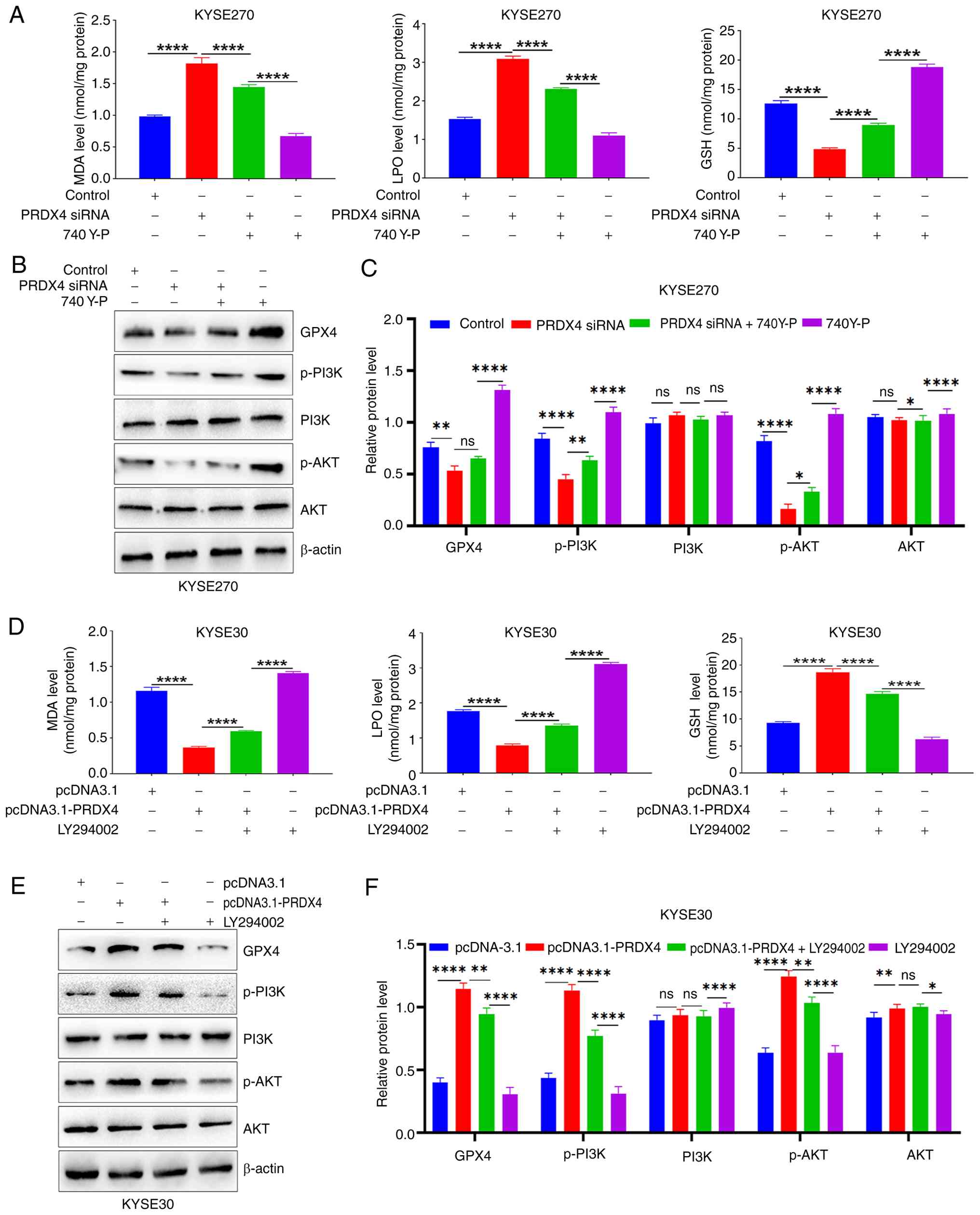 | Figure 6PRDX4 suppresses ferroptosis of ESCC
cells by activating the PI3K/AKT signaling pathway. (A)
Determination of MDA, LPO and GSH contents in the absence or
presence of the PI3K activator 740 Y-P after PRDX4 knockdown in
KYSE270 cells. (B) Western blot analysis of the protein expression
levels of GPX4, p-PI3K, PI3K, p-AKT and AKT in the absence or
presence of the PI3K activator 740 Y-P after PRDX4 knockdown in
KYSE270 cells. (C) The relative protein levels of GPX4, p-PI3K,
PI3K, p-AKT and AKT in the absence or presence of the PI3K
activator 740 Y-P after PRDX4 knockdown in KYSE270 cells. (D)
Detection of MDA, LPO and GSH contents in the absence or presence
of the PI3K inhibitor LY294002 after PRDX4 overexpression in KYSE30
cells. (E) Western blot analysis of the protein expression levels
of GPX4, p-PI3K, PI3K, p-AKT and AKT in the absence or presence of
the PI3K inhibitor LY294002 after PRDX4 overexpression in KYSE30
cells. (F) The relative protein levels of GPX4, p-PI3K, PI3K, p-AKT
and AKT in the absence or presence of the PI3K inhibitor LY294002
after PRDX4 overexpression in KYSE30 cells. *P<0.05,
**P<0.01, and ****P<0.0001, indicate
statistical significance. PRDX4, peroxiredoxin 4; ESCC, esophageal
squamous cell carcinoma; PI3K, phosphoinositide 3-kinase; AKT,
protein kinase B; MDA, malondialdehyde; LPO, lipid peroxidation;
GSH, glutathione; GPX4, glutathione peroxidase 4; p-PI3K,
phosphorylated PI3K; p-AKT, phosphorylated AKT; ns, not
significant. |
Discussion
The relationship between redox homeostasis and tumor
development remains a hot topic of research. Therefore, targeting
the genes related to redox homeostasis may provide therapeutic
opportunities for various tumors. PRDX4 is an important antioxidant
enzyme that widely participates in tumor progression and may be a
potential prognostic biomarker for a variety of tumors (23,38,39),
however, its role and molecular mechanism in the development and
progression of ESCC remains unknown. The present study verified the
high expression of PRDX4 in ESCC samples and cells, which was
strongly linked to TNM staging and lymph node metastasis, and
explained the dismal prognosis of patients with ESCA. Further
investigations revealed that PRDX4 depletion suppressed ESCC cell
proliferation, whereas PRDX4 overexpression had the opposite
effect. Notably, PRDX4 knockdown induced ferroptosis by inhibiting
the PI3K/AKT signaling pathway in ESCC cells. The findings
indicated that targeting PRDX4 is a promising therapeutic approach
for patients with ESCC.
Several studies have demonstrated that PRDX4 is
frequently overexpressed in various tumor types. Kim et al
(40) confirmed that PRDX4 was
highly expressed in human glioblastoma cells and mouse models.
Ummanni et al (39) found
that PRDX3 and PRDX4 were overexpressed in prostate cancer samples,
as was determined by reverse-phase protein arrays, and were
negatively associated with TMPRSS2-ERG gene fusion (39). The iTRAQ technique revealed that
PRDX4 is a metastasis-associated protein in oral squamous cell
carcinoma (41). Notably, PRDX4 may
be a novel prognostic marker for various tumors, including renal
papillary cell carcinoma (24),
colon adenocarcinoma (42), head
and neck squamous cell carcinoma (43), and gastric cancer (23). These studies highlighted the
clinical value of PRDX4 in most tumor types. In the present study,
using bioinformatics analysis, it was found that PRDX4 is widely
overexpressed in pan-cancer and ESCA, which was further validated
in ESCC tissues and cells. Notably, high PRDX4 expression was
strongly associated with TNM staging and lymph node metastasis in
patients with ESCC. Patients with high PRDX4 expression exhibited
shorter survival rates than those with low expression levels.
Therefore, PRDX4 may be an important prognostic factor for patients
with ESCA. The findings indicated that PRDX4 participates in the
development and progression of ESCC and may be a potential
prognostic factor for patients with ESCC.
Previous studies have highlighted the roles of PRDX4
in the regulation of cell proliferation, apoptosis, metastasis, and
radiation resistance. Wang et al (17) found that PRDX4 suppressed anoikis
and promoted growth and metastasis via the β-catenin/ID2 signaling
pathway in hepatocellular carcinoma. In another study, PRDX4
depletion suppressed cell growth and radiation resistance, coupled
with increased ROS levels, apoptosis, and DNA damage in
glioblastoma cells (40). Our data
also demonstrated that PRDX4 depletion suppressed cell
proliferation and colony formation and reduced the number of
EdU-positive cells, whereas PRDX4 overexpression promoted cell
proliferation and colony formation and increased the number of
EdU-positive cells in ESCC cells. In addition, the inhibition of
cell migration and invasion, along with an increase of E-cadherin
and a decrease in N-cadherin and vimentin expression, were observed
upon PRDX4 knockdown; however, the opposite effects were observed
following PRDX4 overexpression. These findings indicated that PRDX4
is a promising therapeutic target for patients with ESCC.
Ferroptosis is a cell-death pathway induced by the
accumulation of phospholipid peroxides (44). GPX4 is a critical enzyme that
protects cells from ferroptosis by reducing phospholipid peroxides
using GSH as the reductant. Thus, GPX4 suppression can induce
ferroptosis (45). SLC7A11, a key
factor involved in ferroptosis, can increase intracellular cystine
levels and GSH biosynthesis, thereby suppressing ferroptosis
(46,47). PTGS2, an enzyme involved in lipid
metabolism, promotes ferroptosis. To further explore the role of
PRDX4 in ferroptosis in ESCC cells, the levels of
ferroptosis-related markers were examined by western blotting. The
findings revealed that PRDX4 depletion significantly increased the
levels of MDA and LPO, but reduced the levels of GSH, whereas PRDX4
overexpression had the opposite effects. Stepwise investigation
demonstrated that PRDX4 knockdown reduced the expression of GPX4
and SLC7A11, but enhanced the expression of PTGS2. Notably, the
ferroptosis inhibitor Fer-1 partially reversed ferroptosis
triggered by PRDX4 knockdown, whereas the ferroptosis inducer
erastin partially promoted ferroptosis suppressed by PRDX4
overexpression in ESCC cells. Evidence suggests that PRDX4 is a
major factor controlling ferroptosis in ESCC cells. The PI3K/AKT
signaling pathway was shown to be critical for inhibiting
ferroptosis by affecting lipid metabolism (48). The PI3K/AKT pathway was demonstrated
to block ferroptosis by activating the NRF2/SLC7A11 signaling axis
during cerebral ischemia (49). In
the present study, it was found that the PI3K activator 740 Y-P
significantly reduced the levels of MDA and LPO and increased the
levels of GSH and GPX4 protein expression in ESCC cells, whereas
the PI3K inhibitor LY294002 exerted the opposite effects. Notably,
740 Y-P and LY294002 partially reversed the effects of PRDX4 siRNA
and PRDX4 overexpression. These findings indicated that PRDX4
suppresses ferroptosis by activating the PI3K/AKT signaling
pathway. However, future studies employing genome-wide CRISPR-Cas9
screening may systematically identify PRDX4-interacting genes and
novel regulatory networks involved in ferroptosis and tumor
suppression. In addition, more detailed mechanistic insights
require further identification of PRDX4-interacting proteins
through co-immunoprecipitation coupled with mass spectrometry,
which will help elucidate its functional role in ferroptosis, an
aspect that will be addressed in our future studies.
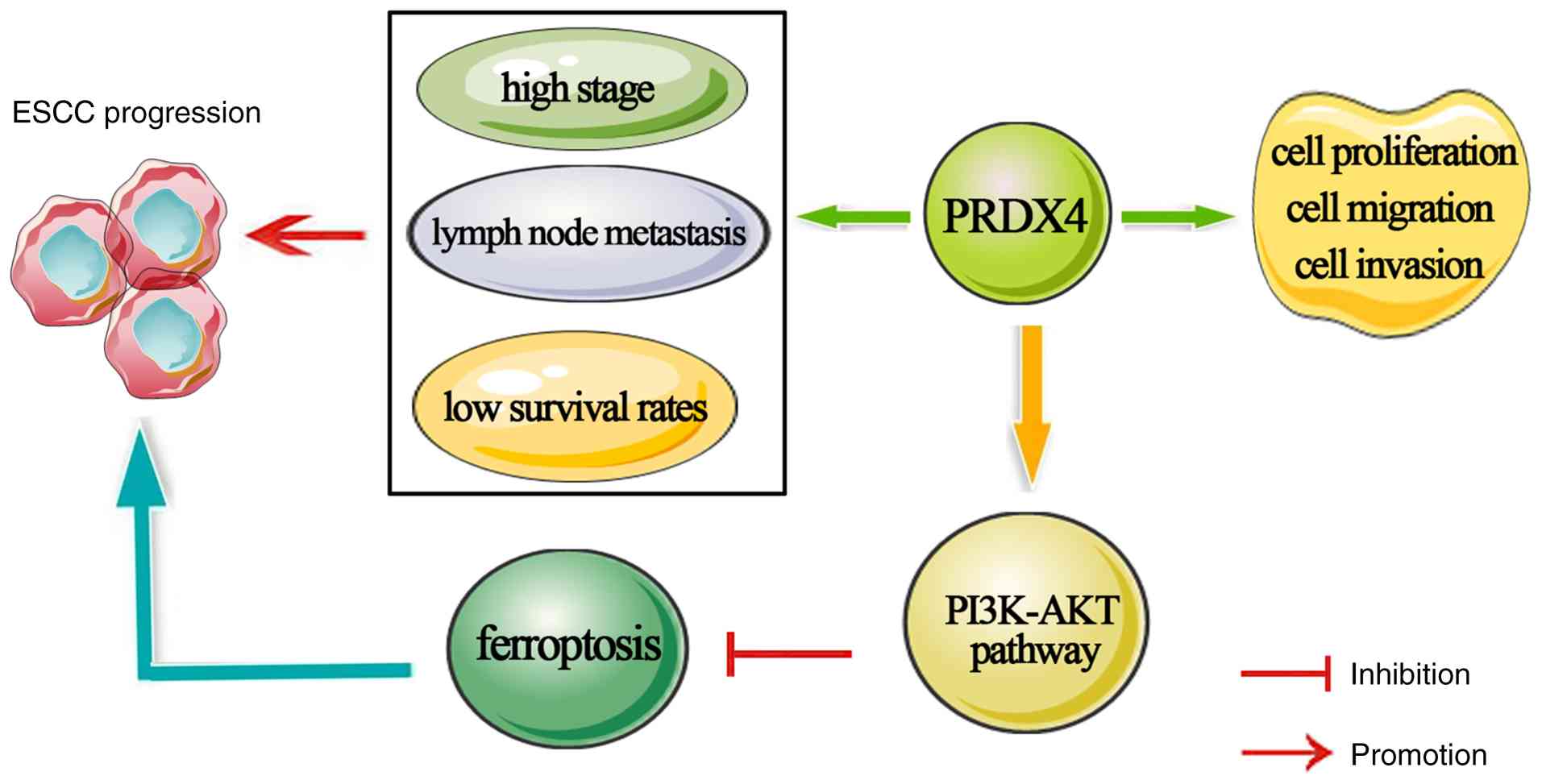
In conclusion, the present study revealed that PRDX4
is highly expressed in ESCC samples and cells. High PRDX4
expression was strongly linked to TNM stage and lymph node
metastasis in patients with ESCC and may be a potential prognostic
indicator in patients with ESCA. PRDX4 knockdown suppressed ESCC
cell proliferation and invasion by inactivating the PI3K/AKT
signaling pathway, which further triggered ferroptosis in these
cells (Fig. 7). These findings
suggest that targeting PRDX4 is a promising therapeutic strategy
for ESCC, thereby providing valuable insights into treatment
approaches for patients with ESCC.
Acknowledgements
Not applicable.
Funding
Funding: The present study was supported by the Key project of
Henan Provincial Science and Technology Research and Development
Joint Fund (grant no. 225200810011), and Henan Province's key
R&D and promotion projects (scientific and technological
research) projects (grant nos. 252102311046 and 222102310099).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
YLiu conceived and designed the current study. KL
and SX performed the majority of the experiments, and interpreted
the data. HL and TF performed data collection and interpretation.
YLi and RR conducted the measurement of lipid peroxidation
indicators, and interpreted the data. YX and SL contributed to the
statistical analysis of the data. YLiu and SX wrote the original
manuscript. KL and SX reviewed and revised the manuscript. KL and
YLiu confirm the authenticity of all the raw data. All authors read
and approved the final manuscript.
Ethical approval and consent to
participate
The present study was approved (approval no. ZZUIRB
2023-239) by the Research and Ethics Committee of Zhengzhou
University (Zhengzhou, China). Written informed consent was
obtained from all patients.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Bi B, Qiu M, Liu P, Wang Q, Wen Y, Li Y,
Li B, Li Y, He Y and Zhao J: Protein post-translational
modifications: A key factor in colorectal cancer resistance
mechanisms. Biochim Biophys Acta Gene Regul Mech.
1866(194977)2023.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Zeng H, Chen W, Zheng R, Zhang S, Ji JS,
Zou X, Xia C, Sun K, Yang Z, Li H, et al: Changing cancer survival
in China during 2003-15: A pooled analysis of 17 population-based
cancer registries. Lancet Glob Health. 6:e555–e567. 2018.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Siegel RL, Miller KD, Fuchs HE and Jemal
A: Cancer statistics, 2021. CA Cancer J Clin. 71:7–33.
2021.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Shi Y, Fang N, Li Y, Guo Z, Jiang W, He Y,
Ma Z and Chen Y: Circular RNA LPAR3 sponges microRNA-198 to
facilitate esophageal cancer migration, invasion, and metastasis.
Cancer Sci. 111:2824–2836. 2020.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Rhee SG and Kil IS: Multiple functions and
regulation of mammalian peroxiredoxins. Annu Rev Biochem.
86:749–775. 2017.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Rhee SG: Overview on peroxiredoxin. Mol
Cells. 39:1–5. 2016.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Fujii J, Ikeda Y, Kurahashi T and Homma T:
Physiological and pathological views of peroxiredoxin 4. Free Radic
Biol Med. 83:373–379. 2015.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Cox AG, Pearson AG, Pullar JM, Jönsson TJ,
Lowther WT, Winterbourn CC and Hampton MB: Mitochondrial
peroxiredoxin 3 is more resilient to hyperoxidation than
cytoplasmic peroxiredoxins. Biochem J. 421:51–58. 2009.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Perkins A, Nelson KJ, Parsonage D, Poole
LB and Karplus PA: Peroxiredoxins: Guardians against oxidative
stress and modulators of peroxide signaling. Trends Biochem Sci.
40:435–445. 2015.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Tavender TJ and Bulleid NJ: Peroxiredoxin
IV protects cells from oxidative stress by removing H2O2 produced
during disulphide formation. J Cell Sci. 123:2672–2679.
2010.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Zito E: PRDX4, an endoplasmic
reticulum-localized peroxiredoxin at the crossroads between
enzymatic oxidative protein folding and nonenzymatic protein
oxidation. Antioxid Redox Signal. 18:1666–1674. 2013.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Harris IS and DeNicola GM: The complex
interplay between antioxidants and ROS in cancer. Trends Cell Biol.
30:440–451. 2020.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Singh A, Kukreti R, Saso L and Kukreti S:
Oxidative stress: A key modulator in neurodegenerative diseases.
Molecules. 24(1583)2019.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Matés JM, Segura JA, Alonso FJ and Márquez
J: Oxidative stress in apoptosis and cancer: An update. Arch
Toxicol. 86:1649–1665. 2012.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Schieber M and Chandel NS: ROS function in
redox signaling and oxidative stress. Curr Biol. 24:R453–R462.
2014.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Wang W, Shen XB, Huang DB, Jia W, Liu WB
and He YF: Peroxiredoxin 4 suppresses anoikis and augments growth
and metastasis of hepatocellular carcinoma cells through the
β-catenin/ID2 pathway. Cell Oncol (Dordr). 42:769–781.
2019.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Jiang W, Wang M, Chen Q, Yu X, Liu G, He
X, Mei C and Ou C: Immune infiltration related PRDX4 facilitates
the malignant features and drug resistance of breast cancer. Sci
Rep. 15(27507)2025.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Liu Y, Han J, Shioya A, Zhang YX, Dung VA,
Oyama T, Guo X, Yang Q, Ito T and Yamada S: The immunohistochemical
combination of low SGLT2 expression and high PRDX4 expression
independently predicts shortened survival in patients undergoing
surgical resection for hepatoblastoma. Diagn Pathol.
20(2)2025.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Li H, Wang Z, Chen X, Li S and Zhang F:
Resveratrol downregulated PRDX4 expression to inhibit the
progression of renal cell carcinoma via Wnt/β-catenin pathway. Food
Sci Nutr. 13(e70352)2025.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Lei P, Yu L, Sun X, Hao J, Shi W, Sun H,
Guo X, Jia X, Liu T, Zhang DL, et al: Exploring the role of PRDX4
in the development of uterine corpus endometrial carcinoma. Med
Oncol. 41(48)2024.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Han J, Itoh T, Shioya A, Sakurai M, Oyama
T, Kumagai M, Takamura H, Okuro M, Mukai T, Kitakata H, et al: The
combination of the low immunohistochemical expression of
peroxiredoxin 4 and perilipin 2 predicts longer survival in
pancreatic ductal adenocarcinoma with peroxiredoxin 4 possibly
playing a main role. Histol Histopathol. 38:1415–1427.
2023.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Park SY, Lee YJ, Park J, Kim TH, Hong SC,
Jung EJ, Ju YT, Jeong CY, Park HJ, Ko GH, et al: PRDX4
overexpression is associated with poor prognosis in gastric cancer.
Oncol Lett. 19:3522–3530. 2020.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Kocatürk B: In silico analysis reveals
PRDX4 as a prognostic and oncogenic marker in renal papillary cell
carcinoma. Gene. 859(147201)2023.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Dixon SJ, Lemberg KM, Lamprecht MR, Skouta
R, Zaitsev EM, Gleason CE, Patel DN, Bauer AJ, Cantley AM, Yang WS,
et al: Ferroptosis: An iron-dependent form of nonapoptotic cell
death. Cell. 149:1060–1072. 2012.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Stockwell BR, Friedmann Angeli JP, Bayir
H, Bush AI, Conrad M, Dixon SJ, Fulda S, Gascón S, Hatzios SK,
Kagan VE, et al: Ferroptosis: A regulated cell death nexus linking
metabolism, redox biology, and disease. Cell. 171:273–285.
2017.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Yang WS, SriRamaratnam R, Welsch ME,
Shimada K, Skouta R, Viswanathan VS, Cheah JH, Clemons PA, Shamji
AF, Clish CB, et al: Regulation of ferroptotic cancer cell death by
GPX4. Cell. 156:317–331. 2014.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Bersuker K, Hendricks JM, Li Z, Magtanong
L, Ford B, Tang PH, Roberts MA, Tong B, Maimone TJ, Zoncu R, et al:
The CoQ oxidoreductase FSP1 acts parallel to GPX4 to inhibit
ferroptosis. Nature. 575:688–692. 2019.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Mao C, Liu X, Zhang Y, Lei G, Yan Y, Lee
H, Koppula P, Wu S, Zhuang L, Fang B, et al: DHODH-mediated
ferroptosis defence is a targetable vulnerability in cancer.
Nature. 593:586–590. 2021.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Wang SJ, Li D, Ou Y, Jiang L, Chen Y, Zhao
Y and Gu W: Acetylation is crucial for p53-mediated ferroptosis and
tumor suppression. Cell Rep. 17:366–373. 2016.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Jiang L, Kon N, Li T, Wang SJ, Su T,
Hibshoosh H, Baer R and Gu W: Ferroptosis as a p53-mediated
activity during tumour suppression. Nature. 520:57–62.
2015.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Zhao Y, Liu Z, Liu G, Zhang Y, Liu S, Gan
D, Chang W, Peng X, Sung ES, Gilbert K, et al: Neutrophils resist
ferroptosis and promote breast cancer metastasis through aconitate
decarboxylase 1. Cell Metab. 35:1688–1703.e10. 2023.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Dos Santos AF, Fazeli G, Xavier da Silva
TN and Friedmann Angeli JP: Ferroptosis: Mechanisms and
implications for cancer development and therapy response. Trends
Cell Biol. 33:1062–1076. 2023.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Torres-Velarde JM, Allen KN,
Salvador-Pascual A, Leija RG, Luong D, Moreno-Santillán DD,
Ensminger DC and Vázquez-Medina JP: Peroxiredoxin 6 suppresses
ferroptosis in lung endothelial cells. Free Radic Biol Med.
218:82–93. 2024.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Fujita H, Tanaka YK, Ogata S, Suzuki N,
Kuno S, Barayeu U, Akaike T, Ogra Y and Iwai K: PRDX6 augments
selenium utilization to limit iron toxicity and ferroptosis. Nat
Struct Mol Biol. 31:1277–1285. 2024.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Luo P, Liu D, Zhang Q, Yang F, Wong YK,
Xia F, Zhang J, Chen J, Tian Y, Yang C, et al: Celastrol induces
ferroptosis in activated HSCs to ameliorate hepatic fibrosis via
targeting peroxiredoxins and HO-1. Acta Pharm Sin B. 12:2300–2314.
2022.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Guo X, Noguchi H, Ishii N, Homma T, Hamada
T, Hiraki T, Zhang J, Matsuo K, Yokoyama S, Ishibashi H, et al: The
association of peroxiredoxin 4 with the initiation and progression
of hepatocellular carcinoma. Antioxid Redox Signal. 30:1271–1284.
2019.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Ummanni R, Barreto F, Venz S, Scharf C,
Barett C, Mannsperger HA, Brase JC, Kuner R, Schlomm T, Sauter G,
et al: Peroxiredoxins 3 and 4 are overexpressed in prostate cancer
tissue and affect the proliferation of prostate cancer cells in
vitro. J Proteome Res. 11:2452–2466. 2012.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Kim TH, Song J, Alcantara Llaguno SR,
Murnan E, Liyanarachchi S, Palanichamy K, Yi JY, Viapiano MS,
Nakano I, Yoon SO, et al: Suppression of peroxiredoxin 4 in
glioblastoma cells increases apoptosis and reduces tumor growth.
PLoS One. 7(e42818)2012.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Chang KP, Yu JS, Chien KY, Lee CW, Liang
Y, Liao CT, Yen TC, Lee LY, Huang LL, Liu SC, et al: Identification
of PRDX4 and P4HA2 as metastasis-associated proteins in oral cavity
squamous cell carcinoma by comparative tissue proteomics of
microdissected specimens using iTRAQ technology. J Proteome Res.
10:4935–4947. 2011.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Zhou H, Li L, Chen J, Hou S, Zhou T and
Xiong Y: Expression and prognostic value of PRDX family in colon
adenocarcinoma by integrating comprehensive analysis and in vitro
and in vivo validation. Front Oncol. 13(1136738)2023.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Cao R, Zhang W, Zhang H, Wang L, Chen X,
Ren X, Cheng B and Xia J: Comprehensive analysis of the PRDXs
family in head and neck squamous cell carcinoma. Front Oncol.
12(798483)2022.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Jiang X, Stockwell BR and Conrad M:
Ferroptosis: Mechanisms, biology and role in disease. Nat Rev Mol
Cell Biol. 22:266–282. 2021.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Stockwell BR: Ferroptosis turns 10:
Emerging mechanisms, physiological functions, and therapeutic
applications. Cell. 185:2401–2421. 2022.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Guo W, Zhao Y, Zhang Z, Tan N, Zhao F, Ge
C, Liang L, Jia D, Chen T, Yao M, et al: Disruption of xCT inhibits
cell growth via the ROS/autophagy pathway in hepatocellular
carcinoma. Cancer Lett. 312:55–61. 2011.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Huang Y, Dai Z, Barbacioru C and Sadée W:
Cystine-glutamate transporter SLC7A11 in cancer chemosensitivity
and chemoresistance. Cancer Res. 65:7446–7454. 2005.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Xu J, Li Y, Kang M, Chang C, Wei H, Zhang
C and Chen Y: Multiple forms of cell death: A focus on the PI3K/AKT
pathway. J Cell Physiol. 238:2026–2038. 2023.PubMed/NCBI View Article : Google Scholar
|
|
49
|
Fu C, Wu Y, Liu S, Luo C, Lu Y, Liu M,
Wang L, Zhang Y and Liu X: Rehmannioside A improves cognitive
impairment and alleviates ferroptosis via activating PI3K/AKT/Nrf2
and SLC7A11/GPX4 signaling pathway after ischemia. J
Ethnopharmacol. 289(115021)2022.PubMed/NCBI View Article : Google Scholar
|