Introduction
Osteonecrosis of the Femoral Head (ONFH) is a
prevalent clinical condition characterized by the degeneration of
subchondral bone, leading to severe hip pain and substantial
functional impairment (1,2). The disease is multifactorial, with the
primary cause being disruption of blood supply to the femoral head,
resulting in the death of bone cells, subsequent collapse of the
femoral head, and degenerative changes in the hip joint. While
corticosteroid use, alcohol consumption, sickle cell disease, and
trauma are well-established risk factors for ONFH, the precise
mechanisms underlying its pathogenesis remain unclear (3,4).
Treatment options for early-stage ONFH range from surgical
procedures aimed at preserving hip function to pharmacological
therapies (5,6). However, most patients ultimately
require total hip arthroplasty, which remains the most effective
treatment (7,8). Moreover, many young patients face the
need for revision surgeries, exacerbating the socioeconomic burden.
Consequently, there is an urgent need to identify novel biomarkers
and therapeutic targets, which are crucial for early diagnosis and
the development of new treatment strategies for ONFH.
Recent research has increasingly focused on the role
of biomolecules in ONFH pathogenesis. For instance, Yu et al
demonstrated the significant influence of immune cell infiltration
on ONFH onset and progression, highlighting a close correlation
with specific immune cell subpopulations (9). Fang et al identified that
immune and m6A regulatory factors play pivotal roles in mediating
the inflammatory response in ONFH, particularly involving
Th2-dominated immune cells and inflammatory mediators (10). However, the detailed pathological
processes remain incompletely understood. Additionally, gene chips
and high-throughput sequencing technologies have proven valuable
for identifying pathogenic genes, providing reliable and actionable
insights (11). Thus, investigating
novel molecular markers and immune cell infiltration in ONFH is
critical for elucidating its underlying pathogenic mechanisms.
This study integrated microarray datasets from the
Gene Expression Omnibus (GEO) database with cartilage samples from
patients with ONFH and supplemented these with our own microarray
data derived from subchondral bone samples of ONFH individuals.
Using machine learning techniques, this study identified key genes,
validated them via ROC analysis, and compared immune cell
infiltration across 22 immune cell subtypes in ONFH and healthy
controls using the CIBERSORT algorithm. This study further
identified key molecular pathways and functional networks within
ONFH tissues, employing bioinformatics analysis to elucidate the
roles of differentially expressed mRNAs. Ultimately, critical
feature genes were identified as molecular markers for both ONFH
diagnosis and immune infiltration.
Materials and methods
Obtaining and handling data
The GEO database (http://www.ncbi.nlm.nih.gov/geo/) was utilized to
download experimental group files for GSE74089 and GSE123568. The
dataset, consisting of 14 normal synovial tissue samples (labeled
as ‘normal’) and 34 ONFH synovial tissue samples (labeled as
‘ONFH’), was processed using the limma package in R software
(version 3.58.1) for data correction and normalization. The samples
were divided into two groups: the ONFH group (designated as
‘Treatment’) and the normal group (designated as ‘Control’).
Additionally, GSE178557, which included four samples each of ONFH
and normal synovial tissue, was used as an independent validation
dataset for external verification.
Identification of DEGs and
characteristic genes
To identify differentially expressed genes (DEGs), a
multi-step approach was employed. The limma package in R (version
3.58.1) was used to filter DEGs from all datasets, applying the
criteria of logFC >1 and an adjusted P-value <0.05. Following
DEG screening, feature genes were selected using Lasso regression,
with the glmnet (version 4.1-7) and e1071 (version 1.7-14)
packages. Additionally, the Support Vector Machine (SVM) algorithm,
implemented via the e1071 package (version 1.7-14), was applied to
DEGs to identify another set of feature genes. Finally, Random
Forest (RF) analysis was conducted on the DEGs using ggplot2
software, and genes with a gene importance score greater than 2
(rfGenes >2) were selected. By intersecting the characteristic
genes identified by these three methods, a final subset of
significant genes was obtained. One of these genes was chosen for
further investigation as a potential marker gene.
Evaluate the accuracy of
characteristic genes
A violin plot was generated using the ggpubr package
(version 0.6.0) to visualize the expression differences of the
selected feature genes between the ONFH (treatment) and normal
(control) groups.
Statistical significance was determined with
P<0.05. The discriminative ability of the identified feature
genes in distinguishing normal from ONFH samples was assessed
through Receiver Operating Characteristic (ROC) curve analysis
using the pROC package. The accuracy of the ROC curve was measured
by calculating the area under the curve (AUC), with an AUC value
greater than 0.7 considered indicative of satisfactory performance.
Finally, ROC curve analysis and differential expression analysis
were performed on the validation group samples to confirm the
validity of the experimental group findings.
Analysis of the correlation of
signature genes
After identifying the disease-related signature gene
through the established screening criteria, samples were classified
into high and low expression groups based on the signature gene
expression levels using the limma package (version: 3.58.1) in R.
The filtering criteria were set as logFC >1 and adj.P.Val
<0.05. DEGs associated with the signature gene were identified
through correlation analysis, with genes exhibiting a strong
correlation selected for further analysis. Data visualization was
performed using heatmaps and volcano plots, while gene correlations
were illustrated through correlation bubbles generated using the
corrplot package.
Functional enrichment analysis
Kyoto Encyclopedia of Genes and Genomes (KEGG)
pathway enrichment analysis was performed to identify significantly
enriched pathways (https://www.kegg.jp/). To explore the functional
enrichment of DEGs linked to the signature gene, Gene Ontology (GO)
enrichment analyses were performed, focusing on biological
processes (BPs), cellular components (CCs), and molecular functions
(MFs). Gene Set Enrichment Analysis (GSEA) was performed to assess
the enrichment and scoring values for the high and low expression
groups of the signature gene, providing insights into their
significance in relevant biological processes and functions.
Immune cell infiltration and
correlation analysis
R software was employed to analyze immune molecular
functions in ONFH and normal group samples, identifying significant
differences between the two groups. Immune cell infiltration was
assessed using the CIBERSORT tool, which applied an inverse
convolution technique to estimate the proportions of 22 lymphocyte
subpopulations across tissue samples (12). Following immune cell infiltration
analysis, the relationship between immune infiltrating cells and
gene expression was investigated using the Spearman correlation
algorithm with a significance threshold of P<0.05. This
correlation analysis aimed to determine the associations between
characteristic genes and immune cell infiltration, with the results
presented in graphical format.
Investigation of the signature gene's
RNA network
To predict the regulatory relationships between
miRNAs and the signature gene, TargetScan (https://www.targetscan.org/) (13), miRanda (http://mirtoolsgallery.tech/mirtoolsgallery/)
(14), and miRDB (https://mirdb.org/) (15) were used. In TargetScan, a Context
score percentile cutoff of ≥90 was applied, and in miRDB, the
Target Score cutoff was set at ≥90. The Director package (version:
3.1.9) was downloaded from the miRanda database to obtain the top
1% ranked genes. These results were consolidated for further
analysis. Additionally, the spongeScan database was utilized to
identify long non-coding RNAs (lncRNAs) that may compete with
miRNAs for binding via microRNA response elements (MREs).
Finally, the ceRNA regulatory network, consisting of the signature
gene, miRNAs, and lncRNAs, was visualized using Cytoscape
v3.10.1.
Isolation and cultivation of
chondrocytes
Synovial cartilage samples were collected from
patients diagnosed with traumatic femoral neck fractures and severe
femoral head necrosis who underwent joint replacement surgery at
the First Affiliated Hospital of Guangxi Medical University between
January 2021 and December 2023. The study included a control group
of 6 patients with femoral neck fractures (5 males and 1 female)
and an experimental group of 6 patients with severe femoral head
necrosis (4 males and 2 females). All procedures were approved by
the Medical Ethics Committee of the First Affiliated Hospital of
Guangxi Medical University (Ethical approval no. 2021-E67-01).
Immediately after surgery, femoral head cartilage
samples were stored at -80˚C for subsequent processing.
Approximately 100 g of femoral head cartilage was excised from each
patient, cut into small pieces, and subjected to enzymatic
digestion using trypsin and type I collagenase. The released
chondrocytes were harvested and cultured in high-glucose Dulbecco's
Modified Eagle Medium (DMEM) supplemented with 10% fetal bovine
serum and antibiotics. The cells were maintained at 37˚C in a
humidified incubator with 5% CO2. Chondrocytes from
passage 3 were used for all subsequent experiments.
RNA extraction, cDNA synthesis, and
quantitative PCR
Synovial cartilage samples were obtained from
patients with ONFH and normal femoral head cartilage. Total RNA was
extracted from approximately 50 g of cartilage tissue using the
RNAeasy™ Plus Animal RNA Isolation Kit (Beyotime
Biotechnology), following the manufacturer's instructions. A total
of 1 µg of RNA was reverse-transcribed into complementary DNA
(cDNA) using the PrimeScript™ RT Reagent Kit with gDNA Eraser
(Takara).
Quantitative real-time PCR (qRT-PCR) was performed
to assess ALDH5A1 mRNA expression levels in both ONFH and normal
femoral head cartilage. Gene-specific primers were designed based
on GenBank sequences. The qRT-PCR reaction consisted of 0.5 µl of
each primer, 1.5 µl of distilled water, 2.5 µl of cDNA, and 5 µl of
SYBR Premix Ex Taq mix (Thermo Fisher Scientific, Inc.), making up
a final volume of 10 µl. GAPDH was used as the internal control,
and relative expression levels of ALDH5A1 were calculated using the
2-ΔΔCt method. All experiments
were conducted in triplicate.
Primer sequences were as follows: ALDH5A1, forward,
5'-ATCACCCCGTGGAATTTCCC-3'; reverse, 5'-TTCACCACGACAGTACAGCC-3'.
GAPDH, forward, 5'-CACCCACTCCTCCACCTTTGAC-3'; reverse,
5'-GTCCACCACCCTGTTGCTGTAG-3'.
Protein extraction and western
blotting
Total protein was extracted from femoral head
cartilage using RIPA buffer containing PMSF (BOSTER), phosphatase
inhibitors (CWBIO), and protease inhibitors (MCE). Protein
concentration was determined using the BCA assay (Beyotime
Biotechnology) following the manufacturer's instructions.
Protein samples were mixed with loading buffer and
heated at 100˚C for 10 min. Proteins were separated by SDS-PAGE
(10% polyacrylamide gel) and transferred to PVDF membranes.
Membranes were blocked with 5% skim milk for 2 h at room
temperature, followed by overnight incubation at -4˚C with primary
antibodies against ALDH5A1 and GAPDH (Proteintech) at a 1:5,000
dilution. After washing with PBST, membranes were incubated for 1 h
at room temperature with a rabbit anti-rat IgG secondary antibody.
Protein signals were detected using an Odyssey infrared imaging
system, and chemiluminescence was visualized using enhanced
chemiluminescence (Beyotime Biotechnology).
Immunofluorescence staining
Chondrocytes were cultured overnight on sterile
glass coverslips at 37˚C with 5% CO2. Cells were fixed
with 4% paraformaldehyde for 15 min, permeabilized with
permeabilization solution (Beyotime Biotechnology) for 10 min, and
blocked with immunofluorescence blocking buffer (Beyotime
Biotechnology) for 1 h at room temperature. After washing, cells
were incubated overnight at 4˚C with anti-ALDH5A1 primary antibody
(Proteintech, China), followed by 2-h incubation with
FITC-conjugated goat anti-rabbit IgG secondary antibody (Epizyme
Biotech, China) in the dark. Nuclei were stained with DAPI (1
µg/ml) for 10 min. Images were captured using a Revolve2 inverted
fluorescence microscope (Echo), and fluorescence intensity was
quantified using ImageJ software.
Statistical analyses
Bioinformatics analysis was performed using R (x64
4.3.1). For the in vitro experiment, data were analyzed with
GraphPad Prism 8.0.2 and ImageJ and are presented as mean ±
standard error. Statistical differences between groups were
evaluated using an unpaired Student's t-test, with P<0.05
considered statistically significant.
Statement
This study was approved by the Medical Ethics
Committee of the First Affiliated Hospital of Guangxi Medical
University (approval number: 2021-E67-01) in Guangxi, China. The
handling of clinical tissue samples adhered to the ethical
standards outlined in the Declaration of Helsinki.
Results
Identification of DEGs and signature
genes
The limma package (version 3.58.1, Department of
Mathematics and Statistics, University of Melbourne, Australia) in
R software was employed to apply adjusted P-values across all
datasets. Using filtering criteria of adjusted P-value <0.05 and
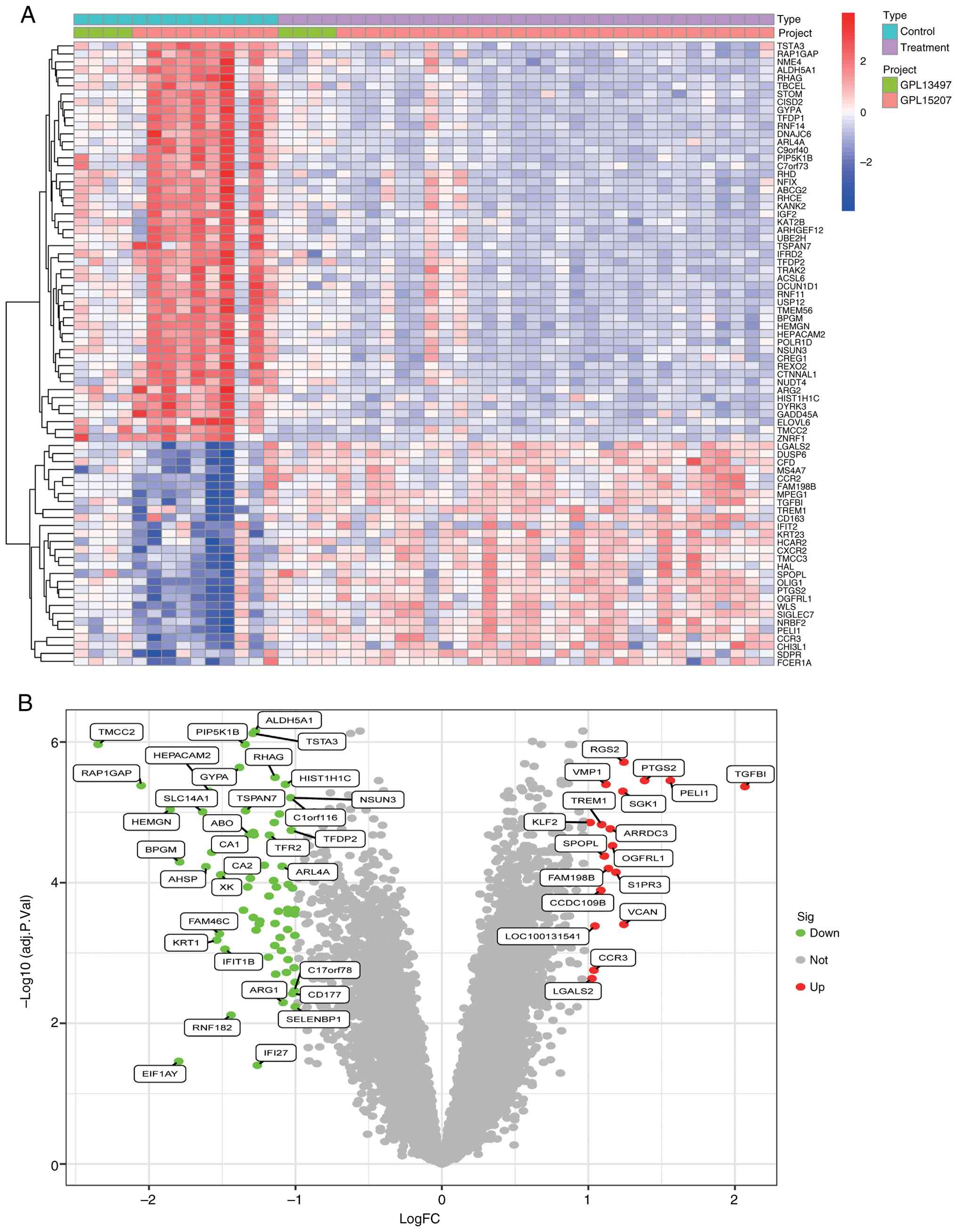
logFC >1, 51 significant DEGs were identified, including 33
downregulated and 18 upregulated genes. The heatmap (Fig. 1A) visualizes the expression patterns
of these DEGs, while the volcano plot (Fig. 1B) illustrates their differential
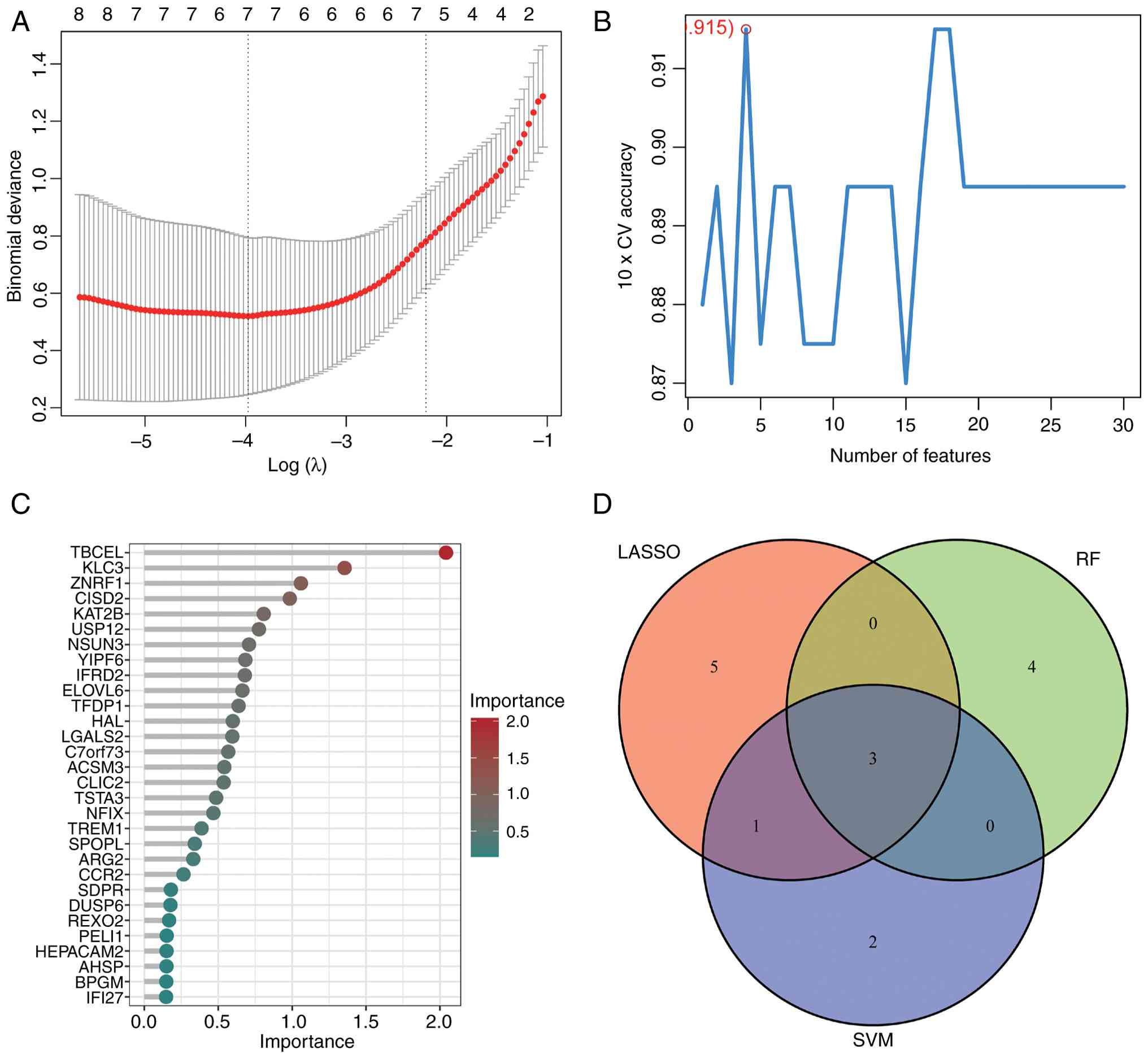
expression levels. Furthermore, 51 DEGs were cross-validated using
Lasso regression, SVM, and RF algorithms in R (Fig. 2A-C), yielding a Venn diagram
(Fig. 2D) that identified three key
characteristic genes: ALDH5A1, TBCEL, and KAT2B (Table I). Based on prior research, ALDH5A1
has been linked to mitochondrial dysfunction and anti-inflammatory
effects (16) and was selected as a
marker gene for further investigation. Basic information for
ALDH5A1 was retrieved from the GeneCards database (https://www.genecards.org/): Protein ID: ALDH5A1
(UniProtKB/SwissProt: P51649); Primary intracellular localization:
mitochondria; Protein size: 535 amino acids; Molecular weight:
57,215 daltons.
 | Table IAlgorithms for genetic screening
results. |
Table I
Algorithms for genetic screening
results.
| Algorithm | Genes |
|---|
| LASSO | ALDH5A1, TSTA3,
TMCC2, TBCEL, ARG2, KAT2B, HIST1H1C, TREM1, LTF |
| SVM | ALDH5A1, KAT2B,
ARG2, TBCEL, YPEL4, FHDC1 |
| RF | ALDH5A1, TBCEL,
KLC3, ZNRF1, CISD2, KAT2B, USP12 |
Accuracy analysis of characteristic
genes
To evaluate the discriminative ability of the three
identified feature genes, ROC analysis was performed for ALDH5A1,
TBCEL, and KAT2B (Fig. S1). The
results indicated that all three genes possessed robust diagnostic
potential; however, ALDH5A1 exhibited the superior diagnostic
accuracy with an AUC of 0.968, compared to 0.931 for TBCEL and
0.899 for KAT2B. Consequently, ALDH5A1 was selected as the optimal
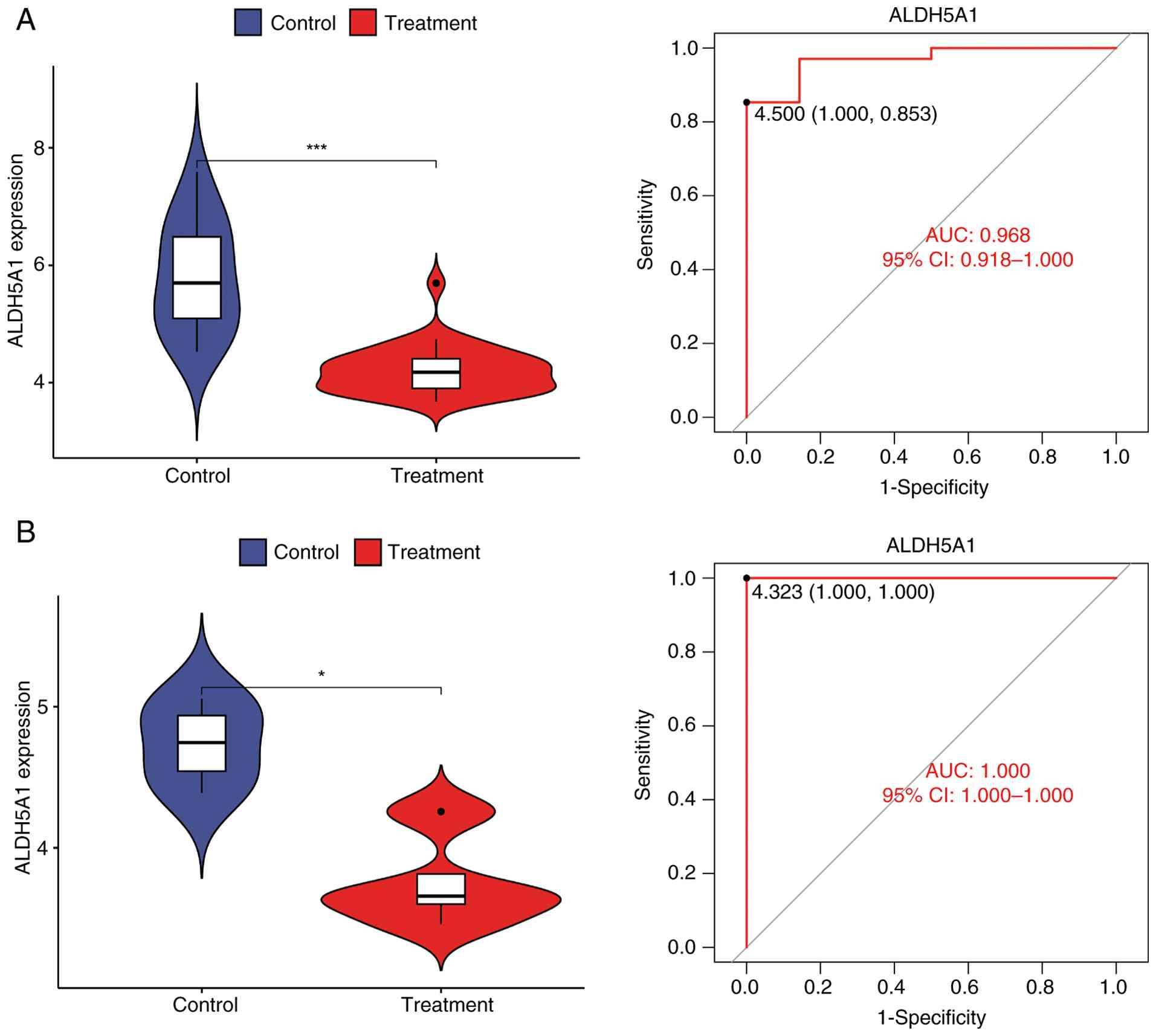
biomarker for further analysis. The violin plot generated using the
ggpubr software (Fig. 3A) indicated
that the expression level of ALDH5A1 in the ONFH group was
significantly lower than in the normal group (P<0.05).
Validation of these findings in the validation group was confirmed
through the violin plot (Fig. 3B)
and corresponding ROC curve, further supporting the reliability and
accuracy of the ALDH5A1 marker identified in the experimental group
samples.
Correlation analysis of characteristic
genes
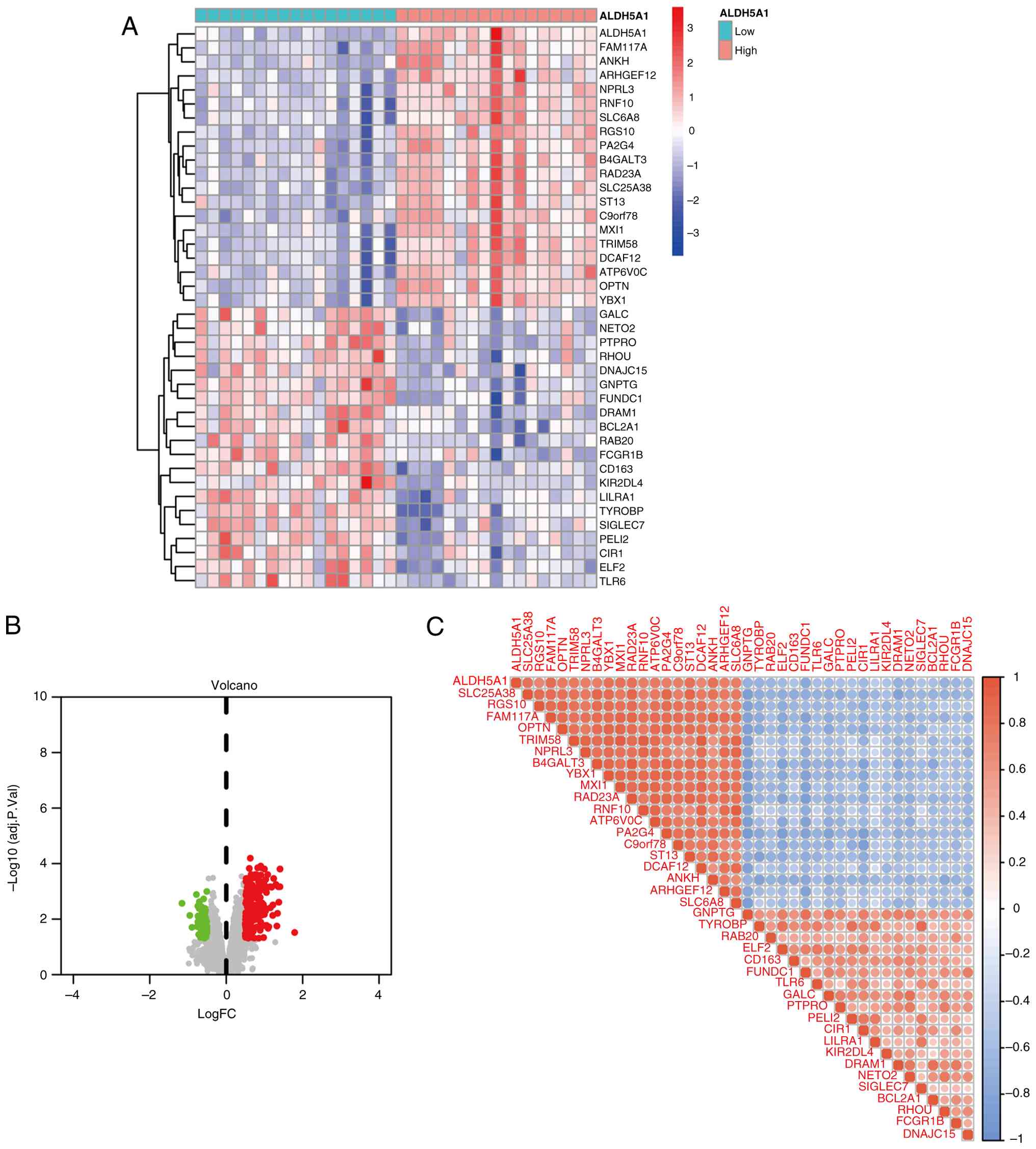
In the correlation study of ALDH5A1, as shown in the
volcano map and differential heatmap (Fig. 4A and B), a correlation analysis filter
identified 39 DEGs associated with ALDH5A1. Among them, 20 were
upregulated and 19 were downregulated. The correlation and
expression pattern of these ALDH5A1-related DEGs are depicted in
the bubble plot (Fig. 4C).
Functional enrichment analysis
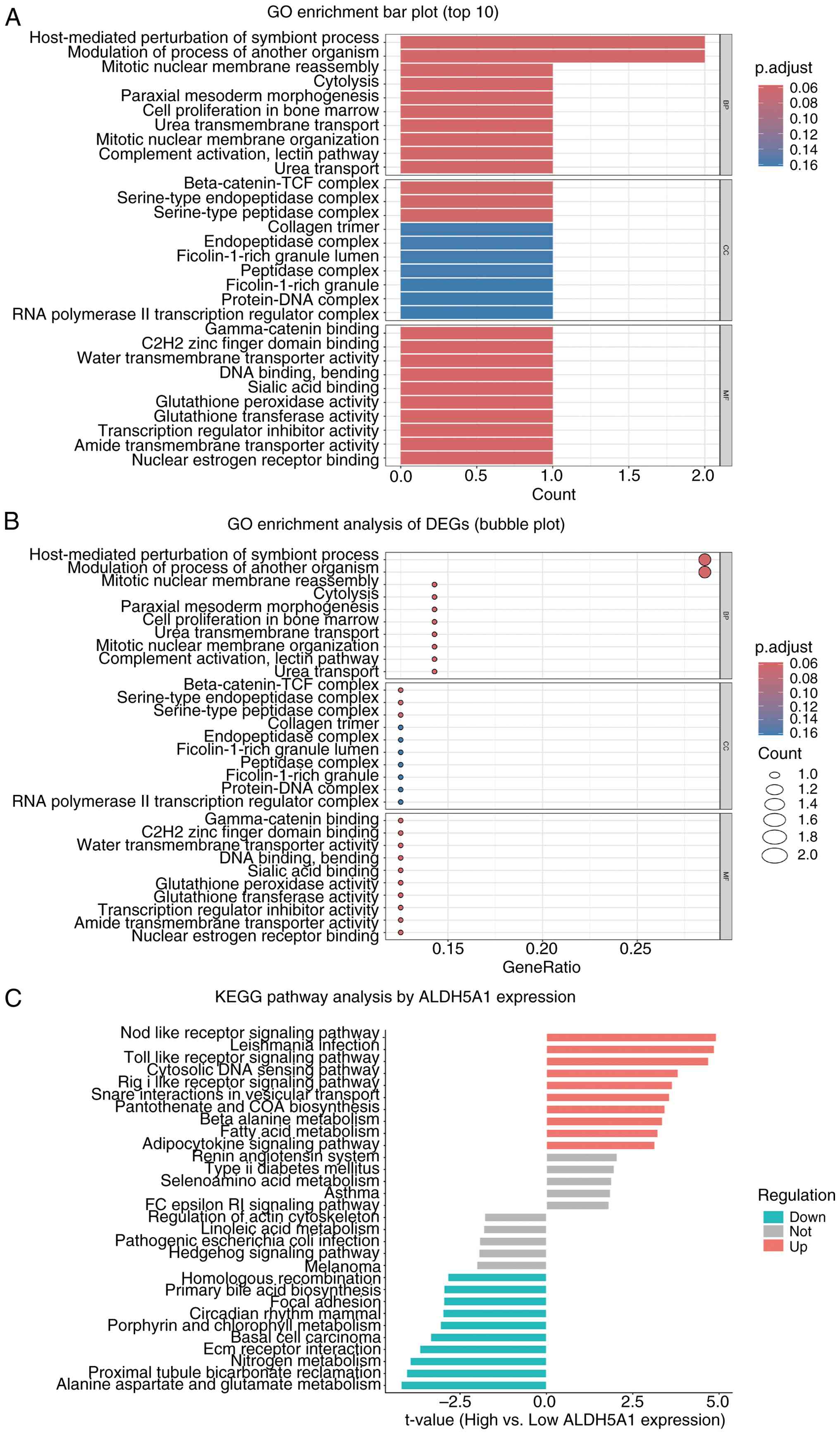
The results of the GO enrichment analysis are
detailed in Fig. 5. The bar plot
(Fig. 5A) highlights the top
significant terms, showing that ALDH5A1-related DEGs are primarily
enriched in biological processes such as anion transmembrane
transport and regulation of autophagy. The bubble plot (Fig. 5B) complements this, displaying
GeneRatio, adjusted P-values, and gene counts for these significant
terms. In terms of CC, DEGs are predominantly localized in
lysosomes and azurophilic granules. Regarding MF, DEGs are strongly
associated with transmembrane transporter activity. These results
suggest that ALDH5A1 may influence ONFH pathophysiology by
participating in signal transduction pathways that regulate
autophagy. According to Gene Set Variation Analysis (GSVA),
upregulated genes among the DEGs related to ALDH5A1 are primarily
enriched in pathways such as nitric oxide receptor signaling,
Leishmania infection, TOLL-like receptor signaling, cytosolic DNA
sensing, RIG-like receptor signaling, and vesicle transport.
Conversely, downregulated genes are enriched in pathways associated
with proximal tubular bicarbonate reclamation, nitrogen metabolism,
ECM-receptor interactions, basal cell carcinoma, porphyrin and
chlorophyll metabolism, and the mammalian circadian rhythm
(Fig. 5C).
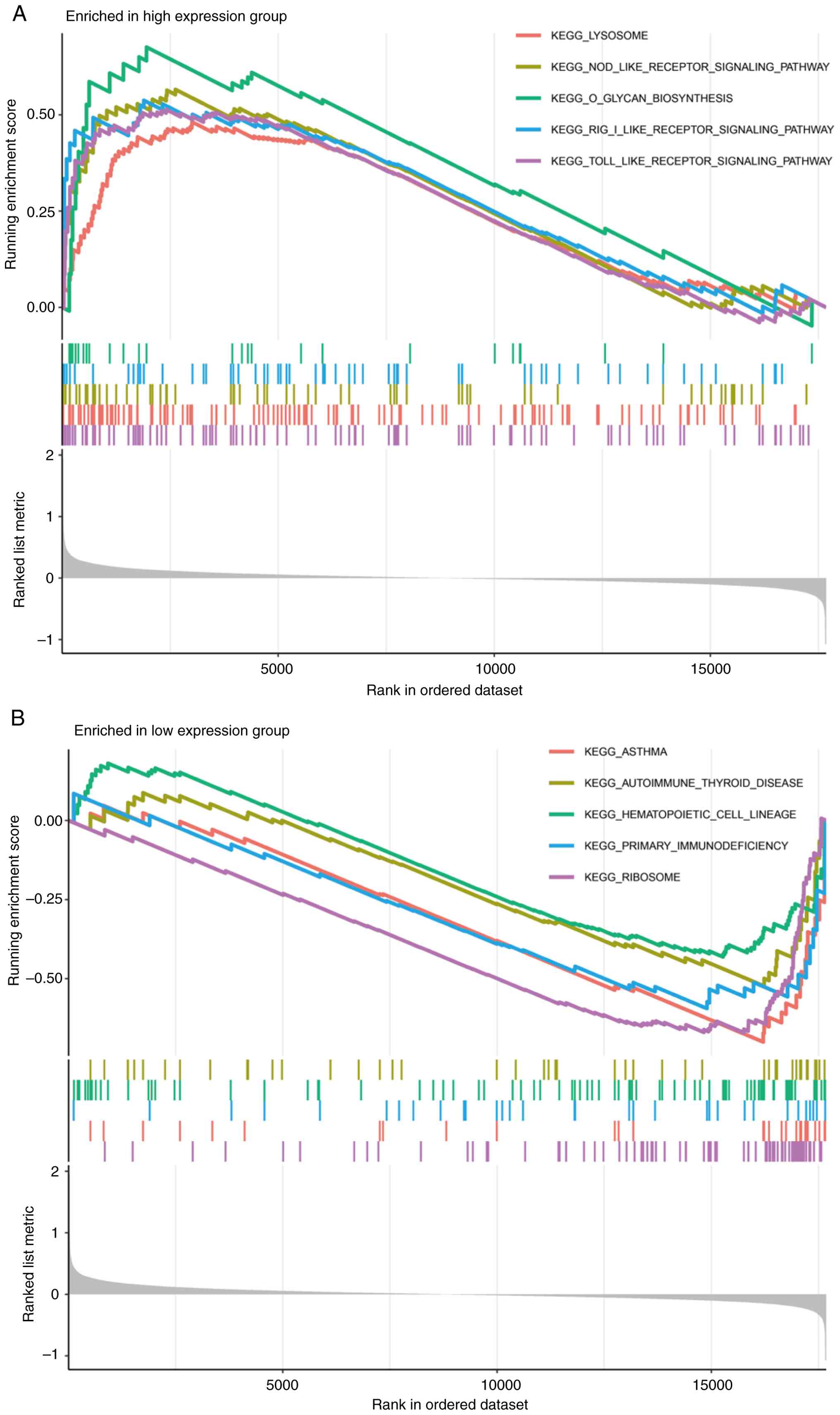
Gene set enrichment analysis
By classifying the experimental samples into high
and low expression groups of ALDH5A1, this study further explored
its biological significance in ONFH through GSEA. The high
expression group was significantly enriched in five activities and
pathways related to lysosomal function and receptor signaling
pathways (Fig. 6A). In contrast,
the low expression group exhibited enrichment in five biological
signaling pathways, including primary immune cell efficiency,
ribosome activity, hematopoietic cell function, asthma, and
autoimmune thyroid disease (Fig.
6B). These findings highlight the potential role of ALDH5A1 in
regulating signal transduction and autophagy, highlighting its
critical involvement in the pathogenesis of ONFH.
Correlation and immune cell
infiltration analysis
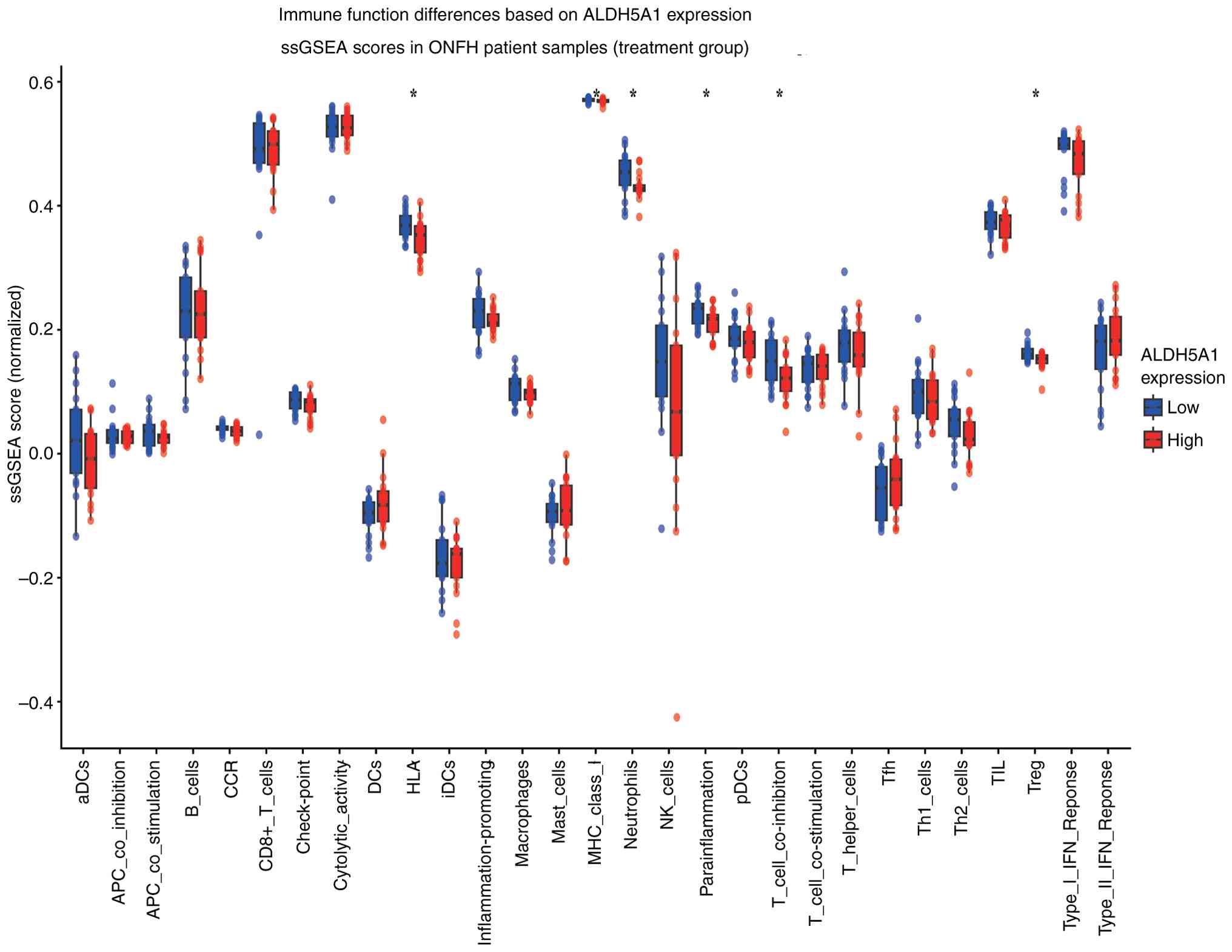
Immune-related functional analysis revealed
significant differences between the two ALDH5A1 expression groups
in terms of HLA and MHC class I molecular functions, neutrophil and
T cell co-inhibition, and regulatory T cell functions (Fig. 7). These functions were particularly
active in the low ALDH5A1 expression group. Immune cell
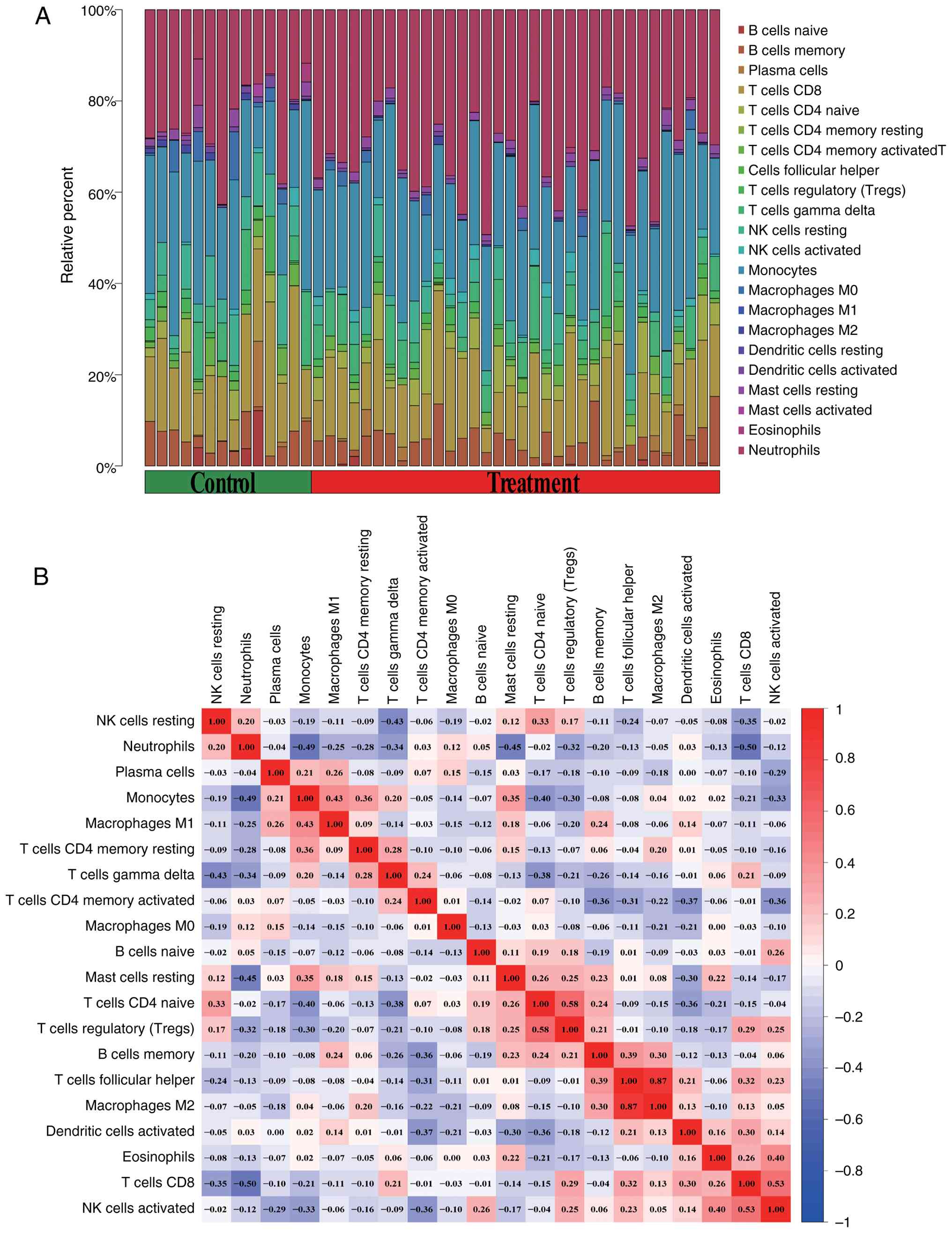
infiltration analysis, depicted in the bubble plot (Fig. 8A), shows the distribution of 22
immune cell types across the normal and ONFH groups. The volcano
plot (Fig. 8B) illustrates a strong
negative correlation between activated CD4+ memory T
cells and memory B cells, T cells, follicular helper cells,
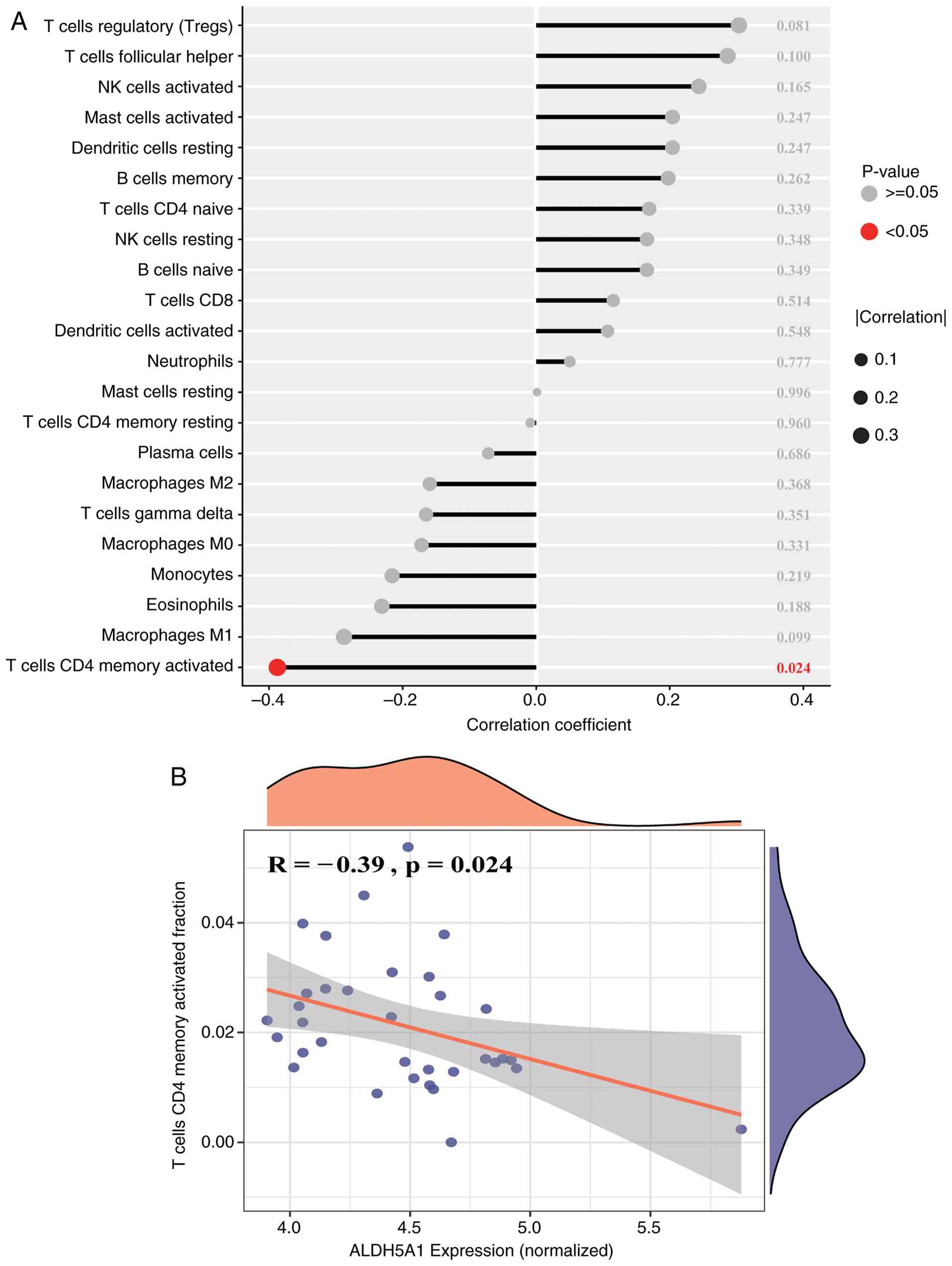
dendritic cells, and activated NK cells. Additionally, the lollipop
plot (Fig. 9A) demonstrates a
significant association (P<0.05) between immune cells and
ALDH5A1, with the most notable finding being a negative correlation
between ALDH5A1 and activated CD4+ memory T cells
(Fig. 9B). These results highlight
the complex interactions between immune cells that contribute to
ONFH progression and suggest that ALDH5A1 plays a key regulatory
role in this immune network.
ceRNA network for detecting
characteristic genes
LncRNAs have gained increasing attention in genetic
research due to their substantial effects on various biological
processes, including cellular regulation. Using three
databases-TargetScan, miRanda, and miRDB-this study analyzed the
regulatory interactions between ALDH5A1 and miRNAs. Our findings
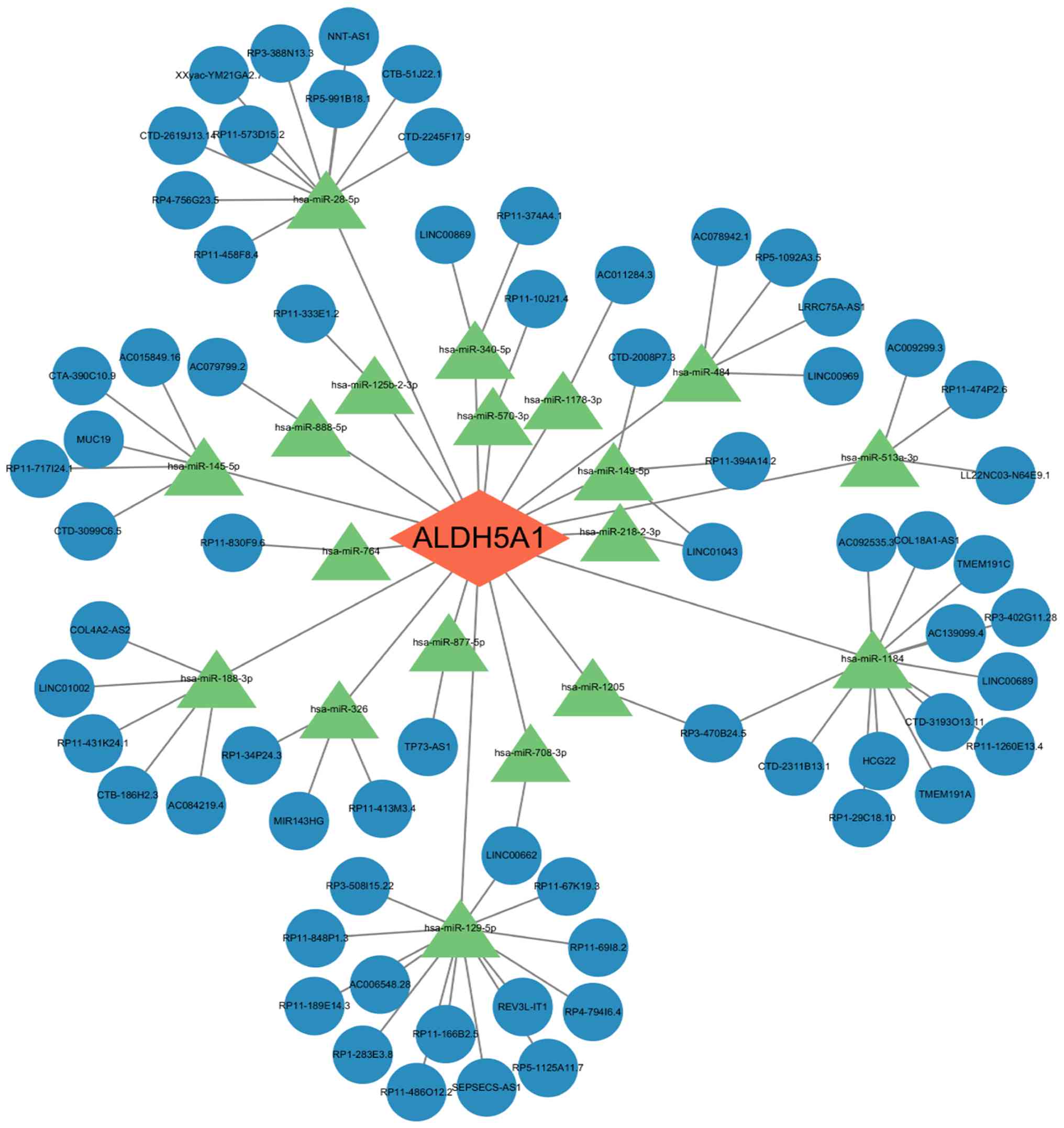
revealed 18 miRNAs that regulate ALDH5A1. Additionally, 68 lncRNAs
that compete with these miRNAs for binding to MREs were identified
(Fig. 10). These results suggest
that ALDH5A1 regulates both its own expression and that of other
molecular targets through the ceRNA network, which may play a
pivotal role in the pathogenesis of avascular necrosis of the
femoral head.
Bioinformatics and experimental
analysis of ALDH5A1 expression
Bioinformatics analysis revealed a significant
decrease in ALDH5A1 expression in chondrocytes from the ONFH group
compared to the normal group (P<0.05). This was corroborated by
qRT-PCR, which showed a marked reduction in ALDH5A1 mRNA levels in
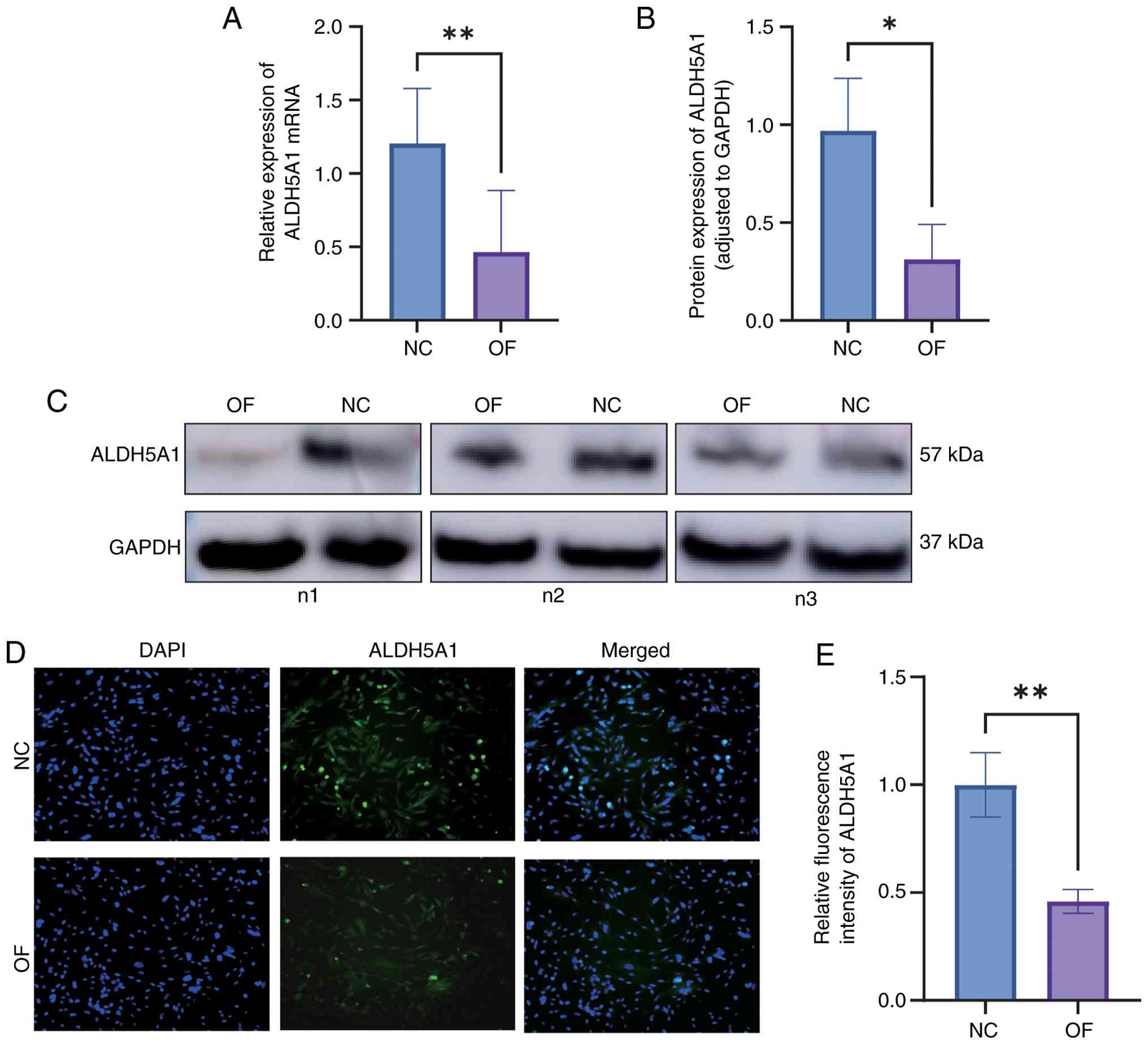
the ONFH group (Fig. 11A). Western
blot analysis further confirmed that ALDH5A1 protein expression was
significantly lower in the ONFH group than in the normal group
(P<0.05) (Fig. 11B and C). Immunofluorescence staining also
demonstrated reduced ALDH5A1 fluorescence intensity in the ONFH
group relative to the normal group (Fig. 11D and E). The relative fluorescence intensity of
ALDH5A1 in the ONFH group was significantly lower (P<0.01),
providing additional support for the decreased ALDH5A1 expression
in ONFH chondrocytes. These findings from bioinformatics, qRT-PCR,
Western blot, and immunofluorescence analyses collectively confirm
that ALDH5A1 expression is significantly reduced in ONFH samples
compared to normal controls.
Discussion
ONFH is a complex, progressive disease primarily
characterized by microcirculatory disorders and necrosis of bone
marrow mesenchymal stem cells. Despite extensive research, the
mechanisms underlying ONFH remain unclear, and the absence of
reliable diagnostic markers often results in delayed treatment,
eventually necessitating joint replacement (17). This study explores mRNA expression
profiles in the peripheral blood serum of patients with ONFH and
healthy controls, incorporating data from hip joint cartilage.
Differential analysis and machine learning algorithms were employed
to identify potential biomarkers, while the CIBERSORT algorithm was
used to investigate correlations between immune cell populations
and ONFH. Additionally, mRNA-miRNA-lncRNA relationships linked to
key genes were predicted. The expression of hub genes was validated
through qRT-PCR and Western blot, with results consistent with our
bioinformatics analysis.
A bioinformatic analysis identified DEGs associated
with ONFH. A total of 51 relevant genes were screened, including 18
upregulated and 33 downregulated genes, which may serve as
significant biomarkers for the pathogenesis and progression of
ONFH. Through methods such as LASSO regression, the SVM algorithm,
and RF analysis, ALDH5A1, TBCEL, and KAT2B were identified as key
genes. ALDH5A1 encodes succinate semialdehyde dehydrogenase, an
enzyme involved in mitochondrial glutamate metabolism, regulating
energy metabolism and maintaining mitochondrial integrity.
Depletion of ALDH5A1 can result in mitochondrial dysfunction,
impaired γ-aminobutyric acid (GABA) metabolism (18), and is associated with reduced ATP
production, increased oxidative stress, and compromised
mitochondrial dynamics, leading to cell apoptosis and inflammation
(19). Following ischemic necrosis
of the femoral head, an oxidative stress microenvironment develops
in the necrotic bone area, driven by excessive reactive oxygen
species (ROS) and apoptosis-inducing factors released from damaged
mitochondria (20). While low
levels of ROS participate in essential cellular signaling,
excessive ROS production induces oxidative stress, disrupting
cellular homeostasis and impairing function (21-23).
ROS are also critical in activating autophagy and osteoblast
apoptosis (24,25). Moreover, glucocorticoids, a major
cause of non-traumatic ONFH, promote ROS production in osteoblasts,
activating the JNK/c-Jun signaling pathway, which triggers
autophagy and apoptosis (26). ROS
inhibitors can reduce ROS levels, inhibit the JNK/c-Jun pathway,
and reverse glucocorticoid-induced osteoblast apoptosis and
autophagy (27). Furthermore,
glucocorticoids impair cell angiogenesis, disrupt mitochondrial
function, and hinder the blood supply to the femoral head,
ultimately leading to bone necrosis (28).
Differential analysis of ALDH5A1 gene expression
between the two groups revealed that its expression was
significantly lower in the ONFH group compared to the normal group,
further supporting our hypothesis. This suggests that ALDH5A1 may
play a pivotal role in ONFH. To explore its potential function, the
samples were divided into high and low ALDH5A1 expression groups,
and enrichment analysis was performed on the DEGs. The GO results
showed that the enriched genes were primarily involved in cellular
autophagy, anion transmembrane transport, and proteasome-mediated
ubiquitin-dependent protein catabolism. GSVA indicated that
upregulated genes were enriched in pathways such as nitric oxide
receptor signaling, Leishmania infection, TOLL-like receptor
signaling, RIG-like receptor signaling, and vesicle
transport-related interactions. These findings strongly implicate
ALDH5A1-related DEGs in pathways like autophagy and lysosomal
function, forming the basis for our mechanistic hypothesis.
Although direct experimental validation of ALDH5A1's role in these
specific pathways was not conducted in this study, the functional
parallels with other members of the ALDH family provide substantial
support for our hypothesis. For instance, ALDH2, a
well-characterized member of the family, protects against bone loss
by regulating mitochondrial ROS and mitophagy, and modulates cell
death pathways, including autophagy and ferroptosis, through
detoxification of lipid peroxidation products. Additionally, other
ALDH family members, such as ALDH1a3, sensitize cells to
ferroptosis via autophagy-dependent ferritinophagy, while ALDH7A1
and ALDH3A1 act as inhibitors of ferroptosis (29,30).
Although research on ALDH5A1 in these contexts is limited, the
functional conservation observed across the ALDH family strengthens
the plausibility of our findings. It is hypothesized that the
downregulation of ALDH5A1 contributes to ONFH pathogenesis through
mechanisms akin to its family members, such as impaired
mitochondrial integrity, failure to detoxify ROS, and subsequent
dysregulation of protective autophagy and ferroptosis pathways
(31,32). This positions ALDH5A1 not only as a
potential biomarker but also as a novel mechanistic component in
ONFH pathology.
Our analysis also highlighted significant
differences in immune cell functions, including HLA and T cell
co-inhibition, between the high and low ALDH5A1 expression groups
(9). Notably, ALDH5A1 expression
was significantly negatively correlated with activated
CD4+ memory T cells (33), suggesting a potential link between
ALDH5A1's metabolic role and the immune microenvironment in ONFH,
which warrants further investigation (34). Finally, a predictive
mRNA-miRNA-lncRNA ceRNA network was constructed, revealing complex
upstream regulatory mechanisms for ALDH5A1 (Fig. 10). While this network is purely
predictive, it provides a foundation for future studies on the
regulation of ALDH5A1 in ONFH.
The identification of ALDH5A1 as a robust biomarker
paves the way for several potential clinical applications. In
diagnostic settings, ALDH5A1 expression, possibly measured in
synovial fluid aspirates or synovial biopsies, could serve as an
objective molecular marker, complementing standard imaging
techniques. This could be particularly useful for diagnosing
early-stage ONFH, even before significant radiographic changes are
evident. Additionally, ALDH5A1 may prove valuable as a prognostic
tool. It is hypothesized that its expression level at diagnosis
could correlate with disease severity or progression. Specifically,
markedly low ALDH5A1 levels may indicate a higher risk or faster
progression to femoral head collapse, aiding in patient
stratification for either joint-preserving procedures or total hip
arthroplasty. While our study focused on tissue samples, future
research should explore whether the ALDH5A1 protein or its
downstream metabolites can be detected as a non-invasive blood
biomarker, representing a significant advancement in ONFH
management.
However, this study has several limitations.
Firstly, a key limitation is the mismatch in tissue types between
our discovery and validation cohorts. Our bioinformatics analysis
was based on publicly available synovial tissue datasets, while
experimental validation was conducted on articular cartilage. These
tissues are histologically distinct. Our rationale is based on the
premise that ONFH evolves into a ‘whole-joint’ disease, where
secondary synovitis and cartilage degradation are linked by a
shared microenvironment. While our data supports this hypothesis,
this constitutes indirect validation. Future studies should confirm
these findings in matched synovial, cartilage, and subchondral bone
samples from the same patient cohort. Secondly, the sample size for
our in vitro experimental validation was relatively small
(n=6 per group), limiting the statistical power and reliability of
our conclusions. Larger patient cohorts will be necessary to
confirm our findings. Thirdly, the validated molecules have not yet
been confirmed in vivo. Additional research is needed to
validate the role of these molecules in ONFH in vivo.
In conclusion, this study used machine learning
methods to identify ALDH5A1 as a key gene associated with ONFH,
suggesting its potential as a prognostic marker. This finding was
validated through ROC curves and in vitro experiments.
Furthermore, significant correlations between key genes and immune
cells were observed, and the ceRNA network relationships between
miRNAs, lncRNAs, and hub genes were predicted. Our findings
contribute to a deeper understanding of ONFH pathogenesis and
provide potential molecular markers for the prognosis and treatment
of the disease.
Supplementary Material
ROC curve analysis of key genes for
diagnostic performance. (A) ROC curve of ALDH5A1, showing its
ability to distinguish cases from controls (AUC and 95% CI are
indicated). (B) ROC curve of TBCEL, showing its ability to
distinguish cases from controls (AUC and 95% CI are indicated). (C)
ROC curve of KAT2B, showing its ability to distinguish cases from
controls (AUC and 95% CI are indicated).
Acknowledgements
Not applicable.
Funding
Funding: This research was supported by the Guangxi Natural
Science Foundation (grant no. 2023GXNSFAA026402), the Guangxi
Medical and Health Appropriate Technology Development and Extension
and Application Project (grant no. GZSY22-62), the Nanning Qingxiu
District Science and Plan Project (grant no. 2020018), the Guangxi
Science and Technology Base and Talent Special Project (grant no.
GuikeAD19254003), and the Health Department of Guangxi Zhuang
Autonomous Region Self-funded project (grant no. Z2013039).
Availability of data and materials
This study analyzed publicly available datasets,
which can be accessed through the Gene Expression Omnibus (GEO)
platform (https://www.ncbi.nlm.nih.gov/geo/). The data generated
in this study can be requested from the corresponding author.
Authors' contributions
TR, QQ, LP, XZ, JH, JM, YQ, ZL and JY participated
in conceptualizing the topic, designing the experimental study,
implementing and analyzing data. TR, QQ, LP, XZ, JH, JM, YQ, ZL and
JY acquired and interpreted graphs and figures, and drafted,
revised and critically reviewed the article. All authors have read
and approved the final manuscript, agreed to the journal for
submission, and have accepted responsibility for all aspects of the
work. TR and JY confirm the authenticity of all the raw data.
Ethics approval and consent to
participate
The study was approved by the Medical Ethics
Committee of the First Affiliated Hospital of Guangxi Medical
University (approval no. 2021-E67-01). Written informed consent was
obtained from the participants at the time of tissue collection for
the use of their discarded tissues in anonymized biomedical
research. The consent form explicitly included permission for the
use of any discarded pathological tissue for future research. The
Ethics Committee confirmed that re-consent for the use of tissue in
the present study was unnecessary, as the original consent was
broad enough to cover retrospective research using these anonymized
samples.
Patient consent for publication
Not applicable.
Competing interests
The authors have declared that they have no
competing interests.
References
|
1
|
Zhao DW, Zhang F, Wang BJ, Liu BY, Li L,
Kim SY, Goodman SB, Hernigou P, Cui Q, Lineaweaver WC, et al:
Guidelines for clinical diagnosis and treatment of osteonecrosis of
the femoral head in adults (2019 version). J Orthop Translat.
21:100–110. 2020.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Zhou W, Qu M, Lv YJ and Zhu JY: New
advances in stem cell therapy for osteonecrosis of the femoral
head. Curr Stem Cell Res Ther. 14:226–229. 2019.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Arbab D and König DP: Atraumatic femoral
head necrosis in adults. Dtsch Arztebl Int. 113:31–38.
2016.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Cohen-Rosenblum A and Cui QJ:
Osteonecrosis of the femoral head. Orthop Clin North Am.
50:139–149. 2019.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Migliorini F, Maffulli N, Baroncini A,
Eschweiler J, Tingart M and Betsch M: Prognostic factors in the
management of osteonecrosis of the femoral head: A systematic
review. Surgeon. 21:85–98. 2023.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Sadile F, Bernasconi A, Russo S and
Maffulli N: Core decompression versus other joint preserving
treatments for osteonecrosis of the femoral head: A meta-analysis.
Brit Med Bull. 118:33–49. 2016.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Cao HJ, Guan HF, Lai YX, Qin L and Wang
XL: Review of various treatment options and potential therapies for
osteonecrosis of the femoral head. J Orthop Translat. 4:57–70.
2015.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Mont MA, Salem HS, Piuzzi NS, Goodman SB
and Jones LC: Nontraumatic osteonecrosis of the femoral head: Where
do we stand today?: A 5-year update. J Bone Joint Surg Am.
102:1084–1099. 2020.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Yu R, Zhang J, Zhuo Y, Hong X, Ye J, Tang
S, Liu N and Zhang Y: ARG2, MAP4K5 and TSTA3 as diagnostic markers
of steroid-induced osteonecrosis of the femoral head and their
correlation with immune infiltration. Front Genet.
12(691465)2021.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Fang W, Peng P, Lin K, Xiao F, He W, He M
and Wei Q: m6A methylation modification and immune infiltration
analysis in osteonecrosis of the femoral head. J Orthop Surg Res.
19(183)2024.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Rodriques SG, Stickels RR, Goeva A, Martin
CA, Murray E, Vanderburg CR, Welch J, Chen LM, Chen F and Macosko
EZ: Slide-seq: A scalable technology for measuring genome-wide
expression at high spatial resolution. Science. 363:1463–1467.
2019.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Chen B, Khodadoust MS, Liu CL, Newman AM
and Alizadeh AA: Profiling tumor infiltrating immune cells with
CIBERSORT. Methods Mol Biol. 1711:243–259. 2018.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Shi Y, Yang F, Wei S and Xu G:
Identification of key genes affecting results of hyperthermia in
osteosarcoma based on integrative ChIP-Seq/TargetScan analysis. Med
Sci Monitor. 23:2042–1048. 2017.PubMed/NCBI View Article : Google Scholar
|
|
14
|
John B, Enright AJ, Aravin A, Tuschl T,
Sander C and Marks DS: Human MicroRNA targets. PLoS Biol.
2(e363)2004.
|
|
15
|
Chen YH and Wang XW: miRDB: An online
database for prediction of functional microRNA targets. Nucleic
Acids Res. 48:D127–D131. 2020.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Yang Q, Zhang P, Han L, Shi PS, Zhao ZF,
Cui DJ and Hong K: Mitochondrial-related genes PDK2, CHDH, and
ALDH5A1 served as a diagnostic signature and correlated with immune
cell infiltration in ulcerative colitis. Aging (Albany NY).
16:3803–3822. 2024.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Song Y, Du ZW, Yang QW, Ren M, Wang QY,
Wang A, Chen GY, Zhao HY, Yu T and Zhang GZ: Association of genes
variants in RANKL/RANK/OPG signaling pathway with the development
of osteonecrosis of the femoral head in Chinese population. Int J
Med Sci. 14:690–697. 2017.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Kim KJ, Pearl PL, Jensen K, Snead OC,
Malaspina P, Jakobs C and Gibson KM: Succinic semialdehyde
dehydrogenase: Biochemical-molecular-clinical disease mechanisms,
redox regulation, and functional significance. Antioxid Redox
Signal. 15:691–718. 2011.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Jackson B, Brocker C, Thompson DC, Black
W, Vasiliou K, Nebert DW and Vasiliou V: Update on the aldehyde
dehydrogenase gene (ALDH) superfamily. Hum Genomics. 5:283–303.
2011.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Brown MN, Gibson KM, Schmidt MA, Walters
DC, Arning E, Bottiglieri T and Roullet JB: Cellular and molecular
outcomes of glutamine supplementation in the brain of succinic
semialdehyde dehydrogenase-deficient mice. JIMD Rep. 56:58–69.
2020.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Hu XF, Wang L, Xiang G, Lei W and Feng YF:
Angiogenesis impairment by the NADPH oxidase-triggered oxidative
stress at the bone-implant interface: Critical mechanisms and
therapeutic targets for implant failure under hyperglycemic
conditions in diabetes. Acta Biomater. 73:470–487. 2018.PubMed/NCBI View Article : Google Scholar
|
|
22
|
McGarry T, Biniecka M, Veale DJ and Fearon
U: Hypoxia, oxidative stress and inflammation. Free Radic Biol Med.
125:15–24. 2018.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Wauquier F, Leotoing L, Coxam V, Guicheux
J and Wittrant Y: Oxidative stress in bone remodelling and disease.
Trends Mol Med. 15:468–477. 2009.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Shadel GS and Horvath TL: Mitochondrial
ROS signaling in organismal homeostasis. Cell. 163:560–569.
2015.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Volpe CMO, Villar-Delfino PH, dos Anjos
PMF and Nogueira-Machado JA: Cellular death, reactive oxygen
species (ROS) and diabetic complications. Cell Death Dis.
9(119)2018.PubMed/NCBI View Article : Google Scholar
|
|
26
|
De Meyer GRY, De Keulenaer GW and Martinet
W: Role of autophagy in heart failure associated with aging. Heart
Fail Rev. 15:423–430. 2010.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Assouline-Dayan Y, Chang C, Greenspan A,
Shoenfeld Y and Gershwin ME: Pathogenesis and natural history of
osteonecrosis. Semin Arthritis Rheu. 32:94–124. 2002.PubMed/NCBI
|
|
28
|
Peng PJ, Nie ZG, Sun F and Peng H:
Glucocorticoids induce femoral head necrosis in rats through the
ROS/JNK/c-Jun pathway. FEBS Open Bio. 11:312–321. 2021.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Yao X, Yu S, Jing X, Guo J, Sun K, Guo F
and Ye Y: PTEN inhibitor VO-OHpic attenuates GC-associated
endothelial progenitor cell dysfunction and osteonecrosis of the
femoral head via activating Nrf2 signaling and inhibiting
mitochondrial apoptosis pathway. Stem Cell Res Ther.
11(140)2020.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Shidoji Y, Hayashi K, Komura S, Ohishi N
and Yagi K: Loss of molecular interaction between cytochrome c and
cardiolipin due to lipid peroxidation. Biochem Biophys Res Commun.
264:343–347. 1999.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Li J, Cao F, Yin HL, Huang ZJ, Lin ZT, Mao
N, Sun B and Wang G: Ferroptosis: Past, present and future. Cell
Death Dis. 11(88)2020.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Su LJ, Zhang JH, Gomez H, Murugan R, Hong
X, Xu DX, Jiang F and Peng ZY: Reactive oxygen species-induced
lipid peroxidation in apoptosis, autophagy, and ferroptosis. Oxid
Med Cell Longev. 2019(5080843)2019.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Chen C, Zhao X, Luo Y, Li B, Li Q, Zhao C,
Huang Y and Kang P: Imbalanced T-cell subsets may facilitate the
occurrence of osteonecrosis of the femoral head. J Inflamm Res.
15:4159–4169. 2022.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Chen X, Li J, Kang R, Klionsky D and Tang
D: Ferroptosis: Machinery and regulation. Autophagy. 17:2054–2081.
2021.PubMed/NCBI View Article : Google Scholar
|