Introduction
Genital human papillomavirus (HPV) infectious types
HPV16/18 are very common in numerous populations. However, more
than 90% of genital HPV infections regress without intervention
within a short period of time. The remaining 5–10% of HPV-infected
patients develop long-term infections, and such persistent
infections are associated with an increased risk of developing
cervical cancer (1). This suggests
that, apart from HPV, other factors or co-factors are involved in
this process (2). To date, little
is known regarding the cellular factors that may facilitate
cervical carcinogenesis. In searching for factors responsible for
susceptibility to persistent HPV infection, our study focused on
insulin-like growth factor 1 (IGF1) protein. IGF1 is one of the
cellular factors involved in a variety of cellular processes, such
as cell proliferation, differentiation and apoptosis.
IGF1 exerts its effect mostly by binding the
specific receptor, IGF1 receptor (IGF1R), which causes
autophosphorylation of its β subunit and, therefore, initiates a
cascade of downstream kinase activation (3). IGF1R mediates IGF1 actions on all
cell types, and these actions reveal diversity in terms of tissue
specificity and developmental stage (4). The IGF1 protein is expressed mostly
in the liver; however, tissue-specific expression is also evident.
Thus, IGF1 exhibits both endocrine and auto/paracrine activity,
respectively. The circulating fraction of IGF1 is transported bound
to IGF1 binding proteins (IGF1BPs) types 1–6. The main transporter
is IGF1BP3, which modulates the bioactivity of IGF1 by limiting its
access to IGF1R (5).
The IGF1 gene is located on chromosome 12 and
extends over 85 kb. The gene comprises six exons separated by long
introns. Transcription starts from one of the two possible
promoters (P1 and P2) located in exons 1 and 2, respectively. Exons
1 and 2 are differently spliced to exon 3, producing alternative
class 1 and class 2 transcripts. The biochemical mechanisms
controlling the usage of IGF1 promoters are unresolved. Both P1 and
P2 regulate transcription over scattered start sites, and it is
suggested that heterogeneous transcription initiation compensates
for the lack of typical proximal promoter elements, such as TATA-
or CCAAT-boxes (6). In both
IGF1 gene promoters, multiple specificity protein 1 (SP1)
binding elements are present, which indicates that SP1 may be an
important IGF1 transcription regulator. SP1 is a sequence-specific
ubiquitous transcription factor that binds with high affinity to
GC-boxes, and to GT- and CT-boxes with significantly lower affinity
(7).
By contrast, exons 5 and 6 demonstrate alternative
splicing patterns, which give rise to six IGF1 precursors: classes
1A and 2A contain exons 3–4 and 6 of the transcript and form the
IGF1Ea isoform with C-terminal Ea extension peptide. Classes 1B and
2B contain exons 3–5 (IGF1Eb isoform), and class 1C and 2C isoform
(IGF1Ec) arises from an internal splice site within exon 5, which
links 49 nucleotides of exon 5 to exon 6.
All of these propeptides undergo subsequent
proteolytic processes and eventually result in one mature 70-kDa
IGF1 protein encoded by exons 3 and 4, which is either released
into the bloodstream or into the extracellular matrix (8). The physiological role of propeptides
and alternative E peptides generated from isoforms IGF1Ea, IGF1Eb
and IGF1Ec remains unclear. It has been proposed that different
IGF1 propeptides have distinct biological effects and may act
through various signaling pathways (9). Little is known about the regulation
of IGF1 pre-mRNA splicing. One of the putative factors implicated
in this process is FOX2 (RNA binding motif protein 9; RMB9), a
splicing regulator that binds exclusively to the UGCAUG and AGCAUG
sites downstream or upstream of the alternatively spliced exons.
Notably, FOX2 acts both as an enhancer and silencer of splicing,
depending on the location of the binding site (10–12).
It has previously been observed that up-regulation
of the circulating IGF1 level is correlated with several types of
cancer, including prostate, breast, colorectal and lung cancer,
which suggests a role for IGF1 as a risk factor for cancer
(4). Numerous studies have focused
mostly on the measurement of circular IGF1, while its role as a
tissue growth factor has been underestimated. Little is known about
IGF1 tissue expression in cervical cancer cells, and the available
data are inconsistent (13,14).
The turning point in studies on IGF1 action was the
detection of the IGF1Eb isoform in the nucleus of human cells by
Tan et al (15). Also, our
previous immunohistochemical studies indicated the presence of IGF1
in the nuclei of a reproductive layer of paraepidermal epithelium
in intraepithelial neoplasia (low-grade squamous intraepithelial
neoplasia; L-SIL, and high-grade squamous intraepithelial
neoplasia; H-SIL) and in cervical cancer cells. The nuclear
localization of IGF1 suggests that it may also have intranuclear
activity (13).
The aim of our study was to evaluate the expression
of total IGF1 and IGF1 isoforms (IGF1Ea, IGF1Eb and IGF1Ec) in
HPV-positive cervix samples in women with pre-cancerous (L-SIL and
H-SIL) and cancer lesions in relation to control HPV-negative
cervix. In addition, we analyzed the usage of IGF1 gene
promoters P1 and P2 in study cells as well as the transcription
factor SP1, splicing factor FOX2 and IGF1R expression levels.
Materials and methods
Research material
The study was performed in a group of 109 women aged
23 to 61 years (median 49). There were no significant differences
in the mean age of women who underwent surgery for squamous cell
cervical carcinoma compared to the control women. However, the
women with H-SIL a statistically lower mean age compared to the
women with squamous cell cervical carcinoma and the controls
(p<0.05).
Among the patients, 87 were treated for L-SIL, H-SIL
and squamous cell cervical carcinoma between 2007–2009 at the
Department of Gynecology and Obstetrics, Medical University of
Lublin, Poland. The study was approved by the Ethics Committee of
the Medical University of Lublin. Cervical cells were collected
from the study subjects via cervical scrape. For cancer and
neoplasmatic cell localization, all specimens initially underwent
H&E staining followed by pathological review. Cervical sections
comprised of at least 70% cancer cells were used as cancer samples.
The tissue samples were frozen immediately in liquid nitrogen and
stored at −80°C until further use.
The study group consisted of postoperative tissues
from patients diagnosed with L-SIL (n=28), H-SIL (n=30) and
squamous cell cervical carcinoma (n=29). The control group (n=22)
consisted of normal cervical tissue specimens obtained from
patients undergoing hysterectomy due to uterine leiomyomas.
Histopathological criteria formulated by the World
Health Organization (WHO) were used to establish the diagnosis of
squamous cell cervical carcinoma (16). With respect to the
dedifferentiation of neoplastic cells, all patients were classified
as G2 with moderately differentiated squamous cell carcinomas (WHO
grading G1–G3). According to FIGO clinical staging, among the
squamous cell cervical carcinoma tissues, 15 patients were at stage
I and 14 at stage IIA (17).
HPV identification
Genomic DNA was isolated from the study tissues
using the QIAamp DNA Midi kit (Qiagen) according to the
manufacturer’s protocol. The purity and concentration of genomic
DNA was analyzed spectrophotometrically. HPV infection was
identified by PCR amplification of the HPV gene sequence using
primers MY09 and MY11 complementary to the genome sequence of at
least 33 types of HPV viruses as described previously (18). The reaction mixture contained 15
ng/μl DNA and the following reagents: 0.5 units Taq DNA
polymerase (Fermentas), PCR buffer and magnesium chloride
(Fermentas) at the final concentration of 1.5 mM, primers at the
final concentrations of 0.25 mM each and dNTPs (Promega) at the
final concentration of 0.2 mM. PCR products were run in 1.2%
agarose gel electrophoresis with ethidium bromide and visualized
under UV light. The product length was assessed according to
MassRuler marker (Fermentas). PCR products were randomly eluted
from agarose gel using the QIAquick Gel Extraction kit (Qiagen) and
sequenced. The sequencing results were computationally analyzed
using the BLAST database (http://blast.ncbi.nlm.nih.gov).
RNA extraction and cDNA synthesis
Total RNA was extracted using the RNeasy-Fibrous
Tissue mini-kit (Qiagen) according to the manufacturer’s protocol.
RNA purification included double DNase treatment. RNA
quantification was performed spectrophotometrically, and the
quality was assessed by agarose-formaldehyde gel
electrophoresis.
For each sample, after gDNA removal, 1 μg RNA was
converted into first-strand cDNA in a reverse transcription
reaction using the QuantiTect Reverse Transcription kit (Qiagen)
following the manufacturer’s instructions.
Primers
The primers used for real-time PCR are presented in
Table I.
 | Table I.Primers used in real-time PCR and PCR
product sizes. |
Table I.
Primers used in real-time PCR and PCR
product sizes.
| Gene/isoform | Primer
sequence | Fragment length
(bp) | Annealing
temperature (°C) |
|---|
| IGF1 |
GCTCTTCAGTTCGTGTGTGG | 171 | 60 |
|
TGACTTGGCAGGCTTGAGG | | |
| IGF1Ea |
TCGTGGATGAGTGCTGCTTCCG | 144 | 60 |
|
TCAAATGTACTTCCTTCTGGGTCTTG | | |
| IGF1Eb |
ATCTACCAACAAGAACACG | 140 | 56 |
|
TACTTCCAATCTCCCTCC | | |
| IGF1Ec |
ACCAACAAGAACACGAAGTC | 126 | 58 |
|
CATGTCACTCTTCACTCCTC | | |
| Class 1 IGF1
(P1) |
CAGCAGTCTTCCAACCCA | 102 | 61 |
|
CACAGCGCCAGGTAGAAGAGATGC | | |
| Class 2 IGF1
(P2) |
CACCTACAGTGAAGATGCACACC | 101 | 67 |
|
CGTCTCCGGTCCAGCCGTGGC | | |
| FOX2 |
CGAGAATAGTGCTGATG | 103 | 55 |
|
GGTCTTACACGTGCTGTAG | | |
| SP1 |
CCCAACCCCAAGCCGGTC | 85 | 65 |
|
CCCCCGAGCCCCTTCC | | |
| IGF1R |
GGGAATGGAGTGCTGTATG | 258 | 60 |
|
CACAGAAGCTTCGTTGAGAA | | |
| RPLP0 |
CCTCATATCCGGGGGAATGTG | 95 | 56 |
|
GCAGCAGCTGGCACCTTATTG | | |
| GAPDH |
AAGGTCGGAGTCAACGGATTT | 60 | 58 |
|
ACCAGAGTTAAAAGCAGCCCTG | | |
Primers specific to the human IGF1 gene were
designed according to sequence number NT_019546.15. Primers to
quantify total IGF1 were complementary to constitutive exons 3 and
4 (13). In order to distinguish
IGF1 mRNA isoforms, primers were complementary to different
exons. To amplify isoform IGF1Ea, the primers were complementry to
sequences in exons 4 and 6; for isoform IGF1Eb, exons 5a and 5b;
for IGF1Ec, exons 5a and 6. To specifically indicate transcripts
class I derived from the P1 promoter, primers were complementary to
exons 1 and 3, and for class II, exons 2/3 and 3.
Other real-time PCR primers were used to quantify
SP1. Primers for IGF1R and the reference genes GAPDH and RPLP0 were
as previously described (14,19).
Primer specificity was assessed by a PCR reaction.
The PCR reaction was carried out in a final volume of 10 μl, and
the mixture contained 15 ng/μl cDNA and the following reagents: 0.5
units Taq DNA polymerase, PCR buffer and magnesium chloride
at the final concentration of 1.5 mM, primers at the final
concentrations of 0.25 mM each, and dNTPs at the final
concentration of 0.2 mM. The reaction was run under the following
conditions: initial denaturation at 90°C for 10 min, followed by 45
cycles of denaturation at 95°C for 10 sec, annealing (temperature
dependant on primer sequences; Table
I) and elongation at 72°C for 20 sec. Each primer pair
generated a single product observed as a single band of appropriate
size (Table I) in 2% agarose gel
electrophoresis. PCR products were eluted from agarose gel using
the QIAquick Gel Extraction kit, and were sequenced to confirm the
expected product presence. The sequencing results were
computationally analyzed using the BLAST database (http://blast.ncbi.nlm.nih.gov).
Real-time PCR
Quantitative real-time PCR analysis was performed in
an automated fluorometer (Rotor-Gene 6000; Corbett Research) using
the SYBR Green PCR master mix (Applied Biosystems, UK). GAPDH and
RPLP0 were used as reference genes, and the selection was based on
data from a previously described validation study (19). The PCR reaction was carried out at
a final volume of 10 μl containing 2 μl of cDNA, primers at a
concentration of 1 μM each and SYBR Green PCR master mix. The
samples were run in triplicate and placed in a 72-well plate. The
reaction was performed under the following conditions: initial
denaturation at 90°C for 10 min, followed by 45 cycles of
denaturation at 95°C for 10 sec, annealing (temperature dependant
on primer sequences; Table I) and
elongation at 72°C for 20 sec. PCR products were quantified
according to the standard curves produced by amplification of the
serial dilutions, and the expected product was verified according
to the melting point.
Statistical analysis
The measured parameters were statistically evaluated
as quotient scales by means of the median, lower and upper quartile
with range of trait variability. Because of the skewed distribution
of the analyzed parameters evaluated by the Shapiro-Wilk W test or
non-homogeneous variance calculated by the Fischer F test,
non-parametric tests were used to analyze differences between
subgroups. The Kruskal-Wallis test and multiple post hoc
analysis were used for the comparison of more than two groups.
Spearman’s rank correlation coefficient (R) was used for the
assessment of the relationship between subgroups. Levels of
significance with 5% error (p<0.05) were chosen to calculate the
statistical significance of differences or correlations. The
obtained results are presented in the tables and figures. All
calculations were carried out using the Statistica 7.1 software
package (StatSoft, Poland).
Results
The research material underwent HPV identification.
For the study, 109 samples were selected and divided into
HPV-negative controls and HPV-positive L-SIL, H-SIL and cancer
cells.
Using the real-time PCR technique, all possible
IGF1 mRNA isoforms were identified in the pre-cancer, cancer
and control epithelial cells of the cervix samples.
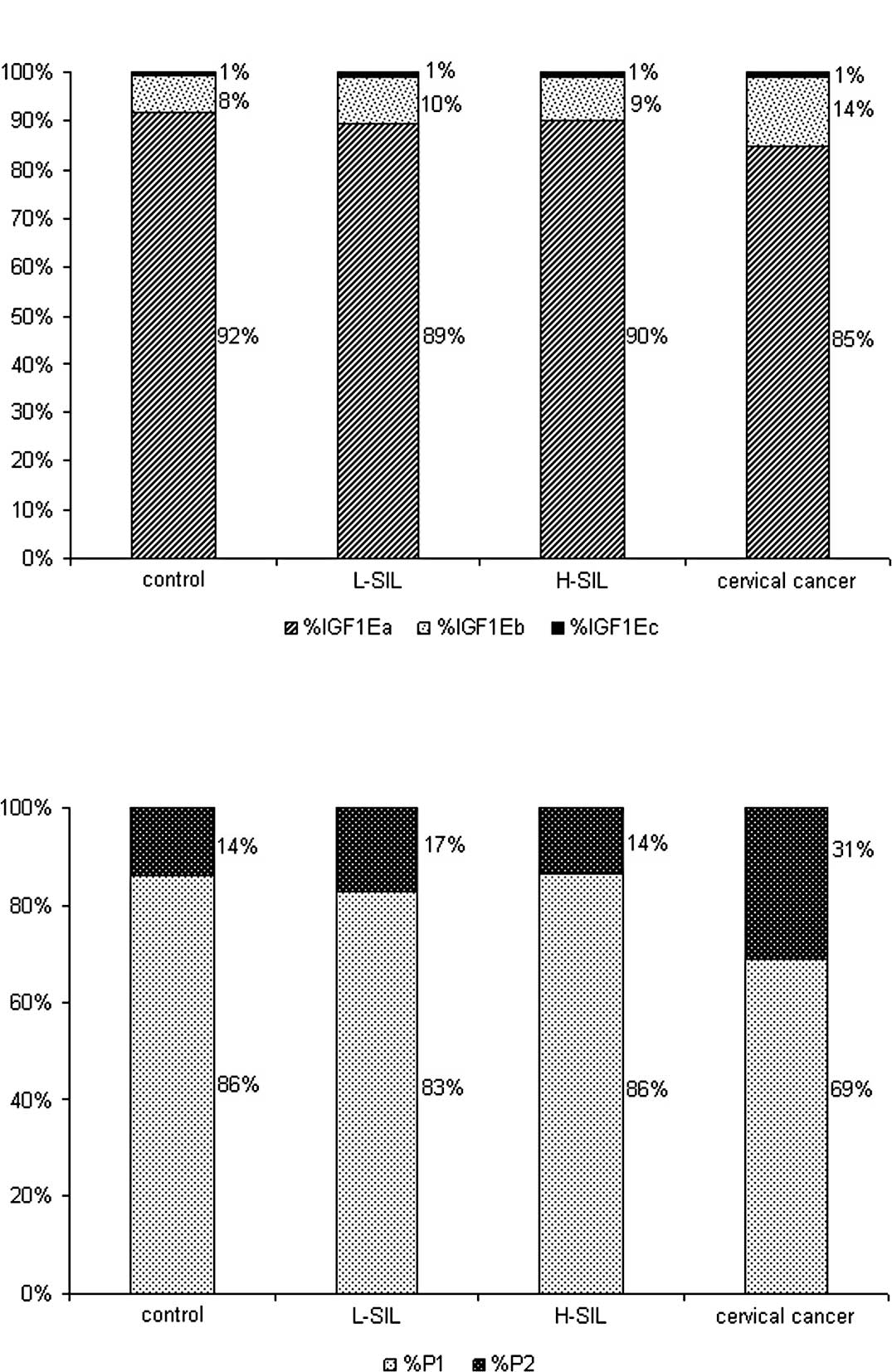
The predominant splicing isoform in all study cells
was IGF1Ea (85–92%). The expression of the IGF1Eb isoform, which
was differential and dependent on the stage of carcinogenesis,
ranged between 8% in the control tissue and 14% in the cervical
cancer. The expression of the IGF1Ec isoform appeared to be at the
lowest level and was consistently at ∼1% in all the studied tissues
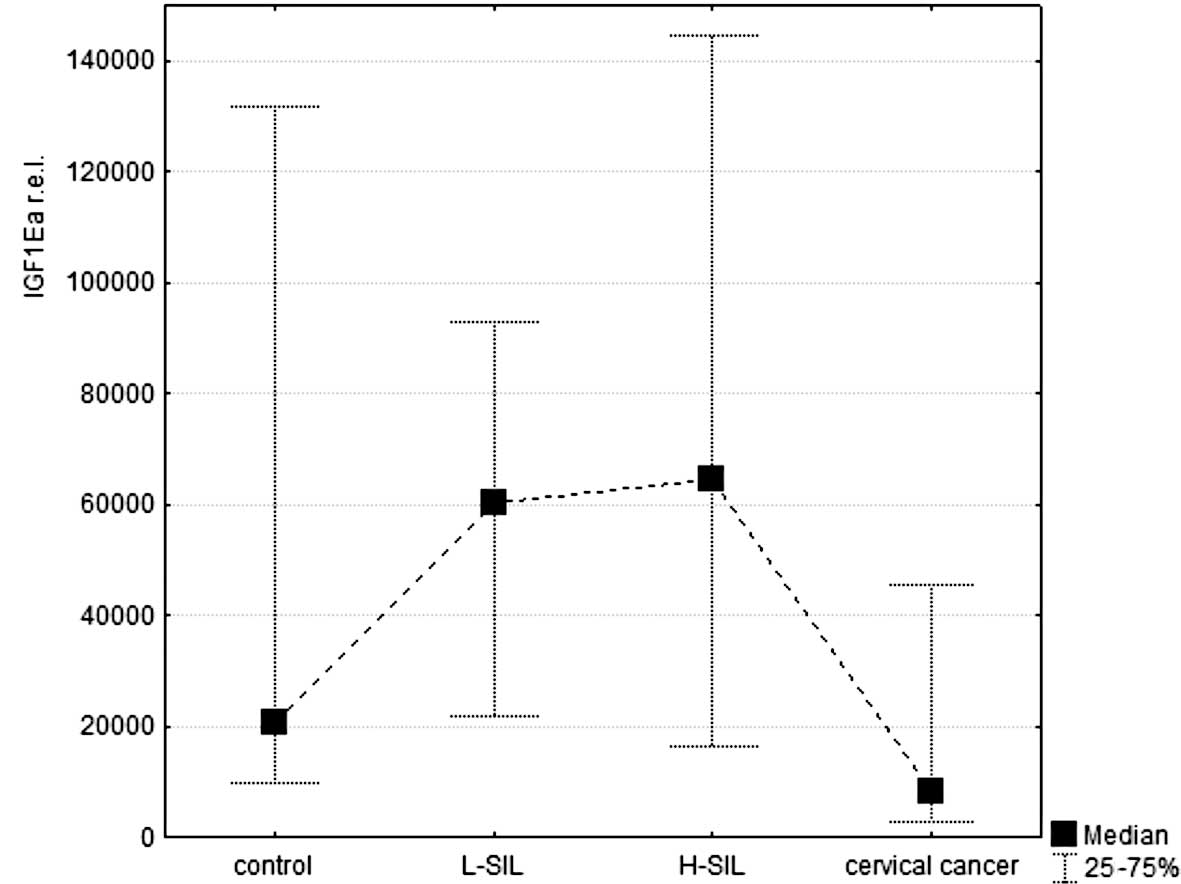
(Fig. 1A). Detailed analysis of
the mRNA relative expression levels of each splicing isoform
(normalized according to reporter genes) showed up-regulation of
all isoforms in L-SIL and H-SIL, compared to the control and the
cervical cancer tissue (Fig.
2A–C). Differences between the relative expression of each
isoform A–C in H-SIL and cervical cancer were statistically
significant (p<0.05). Additionally, for IGF1Eb, this difference
was also statistically significant when compared to L-SIL and
cervical cancer.
Promoter usage in the IGF1 gene was
determined by identification of either exon 1 or exon 2 in the IGF1
transcripts. IGF1 transcription in the studied cells was
mostly controlled by the P1 promoter localized in exon 1 (69–86% of
class I transcripts). However, exon 2 was detected in 14–31% of the
transcripts, which indicates considerable P2 promoter activity
(class 2 transcripts). The greatest contribution of the P2 promoter
in total IGF1 expression was observed in the cervical cancer
samples (31%) (Fig. 1B). Detailed
analysis of the relative expression levels showed no significant
differences in promoter usage between the cancer, L-SIL, H-SIL and
control tissues (Fig. 2D and
E).
The mechanism of IGF1 pre-mRNA alternative
splicing and promoter usage regulation remains unknown. However,
the analysis of alternative splicing factor FOX2 expression levels
(unpublished data) showed a statistically significant correlation
between the FOX2 mRNA expression level and an increase in
the IGF1Eb isoform (Table II).
 | Table II.Statistical analysis of the
expression of IGF1 mRNA isoforms, promoter activity and total IGF1
expression correlated with mRNA expression levels of FOX2, SP1 and
IGF1R, respectively, at various stages of carcinogenesis. |
Table II.
Statistical analysis of the
expression of IGF1 mRNA isoforms, promoter activity and total IGF1
expression correlated with mRNA expression levels of FOX2, SP1 and
IGF1R, respectively, at various stages of carcinogenesis.
| Control | L-SIL | H-SIL | Cervical
cancer |
|---|
| IGF1Ea and
FOX2 | 0.36 | 0.69b | 0.62b | 0.59a |
| IGF1Eb and
FOX2 | 0.21 | 0.64b | 0.68b | 0.63a |
| IGF1Ec and
FOX2 | 0.25 | 0.61b | 0.64b | 0.52 |
| P1 and SP1 | 0.92b | 0.57b | 0.55b | 0.15 |
| P2 and SP1 | 0.77b | 0.44a | 0.34a | 0.25 |
| Total IGF1 and
IGF1R | 0.52 | 0.40 | −0.21 | 0.45 |
SP1 transcription factor, the activator for the P1
and P2 promoter, appeared to be up-regulated in dysplasia
(unpublished data), which was correlated with the increased IGF1
level in these tissue samples (Table
II). No correlation between SP1 and promoter usage was detected
in the cancer tissue.
The expression level of IGF1-specific receptor IGF1R
mRNA was also quantified, and was observed to be doubled in the
cancer tissue compared to the controls, while slight up-regulation
of IGF1R mRNA was observed in neoplasia, L-SIL and H-SIL tissues
(unpublished data). No statistically significant correlation was
found between tissue IGF1 and IGF1R expression levels (Table II).
Discussion
IGF1 is a circulating peptide hormone and a locally
acting growth factor with endocrine, paracrine and autocrine
functions (3). Recently, a number
of studies have focused on the identification of different IGF1
isoforms and the evaluation of their functional significance
(9,20–26).
The physiological differences between the functions
of the local and circulating fraction of IGF1 have not been
completely established. However, it has been proposed that
different IGF1 isoforms (IGF1Ea, IGF1Eb and IGF1Ec) have various
functions in muscle repair and growth (20). After injury to skeletal muscle, the
IGF1Eb mRNA splice variant is initially up-regulated, followed by
up-regulation of the IGF1Ea splice variant at later time points.
Up-regulation of IGF1Eb mRNA correlates with myoblast
proliferation, whereas up-regulation of IGF1Ea mRNA is linked to
differentiation into mature myofibers (21,22).
Other studies have indicated that the locally acting IGF1 isoform
also has neuroprotective effects and protects cardiomyocytes from
hypertrophic and oxidative stress (9,23,24).
Ohtsuki et al, using the PCR method, demonstrated that
IGF1Ea mRNA was more abundant than IGF1Eb mRNA in a number of
organs (uterus, ovary, liver and kidney) in the mouse, and was both
organ specifically and temporally regulated (25). Additionally, it has been suggested
that E-peptides modulate the cell entry of mature IGF1 (26).
In our study, we detected all three IGF1 isoforms in
the HPV-positive L-SIL, H-SIL and cancer tissues as well as in the
HPV-negative control cervical epithelium. However, we observed
increased mRNA expression of all IGF1 isoforms in L-SIL and H-SIL
compared to normal cervix and the cervix of women with cervical
cancer (Fig. 2A–C). Notably, the
percentage contribution of isoform IGF1Eb to total IGF1 expression
was higher in cervical cancer compared to that in other stages of
carcinogenesis (Fig. 1A). We
suggest that this shift in the balance between IGF1 isoforms
towards mitogenic IGF1Eb in cervical cancer cells may be
significant in cancer progression. It has previously been suggested
that E-peptides may have functions distinct from mature IGF1.
Siegfried et al demonstrated that E-peptide derived from the
IGF1Eb isoform exhibits mitogenic activity (25). Later, it was shown that E-peptides
increase IGF1 uptake into cells and enhance mature IGF1 activity
(26). Therefore, we propose that
a shift in the ratio of IGF1Ea to IGF1Eb isoforms towards IGF1Eb
may also serve as a biomarker for the prognosis of cervical cancer
development.
Furthermore, up-regulation of the IGF1Eb isoform in
cervical cancer was found to correspond to simultaneous expression
of HPV viral E6 and E7 proteins during the late stages of neoplasia
(1). HPV viral early proteins are
involved in a variety of intracellular processes, including
splicing. It has previously been demonstrated by Bell et al
that HPV E4 protein interacts with SR-specific kinase SRPK1, which
consequently impairs spliceosome assembly (27). In addition, Bodaghi et al
showed that HPV16 proteins E2 and E6 are RNA-binding proteins and
may affect splicing both by binding directly to pre-mRNA and by
interacting with splicing factors (28). It was previously described that HPV
proteins also regulate gene expression at the transcriptional level
and affect splicing by the up-regulation of splicing factors. In
fact, the HPV E2 protein acts as a transcription factor, not only
in terms of viral gene regulation, but also by altering the
expression of multiple cellular genes, including splicing factors
SR (arginine/serine-rich splicing factor protein family), splicing
factor 2/alternative splicing factor (SF/ASF) and heterogeneous
nuclear ribonucleoprotein A1 (hnRNPA1) (29,30).
A gradual increase in the expression level of the
FOX2 splicing factor in cervical epithelium upon cervical
carcinogenesis has been observed. These results are consistent with
data obtained by Venables et al for breast cancer, but
differ from those found for ovarian cancer (31). It has been suggested that
downstream sites demonstrate the character of the splicing
enhancer, while upstream sites function as silencers (12). In the IGF1 gene, the FOX2
putative binding sites are the upstream exon 5 and downstream exon
6. Our results indicate that the up-regulation of FOX2 may prevent
the exclusion of exon 5 and at the same time enhance the exclusion
of exon 6, which results in the up-regulation of the IGF1Eb isoform
and a reduction of IGF1Ea in dysplasia and cervical cancer tissue.
This putative mechanism resulted in correlations between IGF1 mRNA
splicing isoforms and FOX2 expression levels (Table II).
In this study, we found that the up-regulation of
total IGF1 expression, which is mostly controlled by P1, was
increased in both stages of neoplasia (L-SIL and H-SIL) (Fig. 2D and E). The mechanism of
preferential transcription initiation from the P1 promoter as well
as increased P2 usage in cancer (Fig.
1B) remains unresolved. However, HPV proteins, which are
overexpressed in neoplasia (e.g., E2) (1), may be involved in the regulation of
IGF1 promoter activity. Furthermore, we demonstrated the
up-regulation of SP1 transcriptional factor in HPV-positive tissues
(unpublished data). Of note, SP1 binding sites have been found in
both the P1 and P2 IGF1 promoters. Thus, we suggest that the
correlation of SP1 expression with P1 and P2 IGF1 promoter
activity in neoplasia is related to the up-regulation of
IGF1 expression caused by SP1 action. It has not only been
suggested that SP1 activates IGF1, but that the SP1 binding site is
also a target of IGF1 action in other genes activated in cancer
(32), which may be an additional
explanation for the considerably strong correlation between the
expression levels of these factors (Table II). In addition, the SP1-binding
site is also situated in the HPV E6/E7 promoter region, and is
probably the only one recognized by cellular transcription factors
essential for the function of early transcription initiation
(33,34). Thus, we suggest that the
up-regulation of SP1 during the early stage of dysplasia may
provoke expression of the HPV oncogenic proteins E6 and E7, and may
therefore serve as a prognostic factor for cancer development.
Additionally, recent studies on SP1 expression upon HPV infection
showed concurrent expression of SP1 with L1 proteins in
differentiating keratinocytes in mice (35).
Measurement of the expression level of the
IGF1-specific receptor IGF1R revealed a significant up-regulation
in cervical cancer tissue (unpublished data). This result is
consistent with previously described data obtained in mice, where
the overexpression of IGF1R facilitated tumor development (36,37).
Moreover, increased expression of IGF1R has been linked to several
cancer types in humans (38,39).
It has been proposed that the up-regulation of IGF1R in cancer
cells may be due to loss of transcriptional control by major
tumor-suppressor genes (40).
Additionally, immunohistochemistry with an anti-IGF1R antibody
revealed that the expression levels of IGF1R in H-SIL and invasive
squamous cell carcinoma were higher than the level in normal
cervical epithelial cells. Kuramoto et al showed that the
distribution of IGF1R overexpression in L-SIL and H-SIL was
co-localized with that of HPV E6 oncoprotein expression (41). We did not find any significant
correlation between IGF1R and the local expression of total IGF1
(Table II). However, this result
can be explained by the fact that our study focus was on locally
expressed IGF1, while IGF1R is activated not only by the local, but
also by the circular IGF1 fraction (5).
Taken together, our results demonstrate the
differential expression of IGF1 mRNA splicing isoforms in
HPV-positive pre-cancerous and cervical cancer cells compared to
HPV-negative control tissue. Both IGF1 promoters appeared to
be active in normal and tumorous cervical cells; however, the
balance between them was unequal. Moreover, we demonstrated a
strong correlation in the expression of IGF1 isoforms with cellular
factors FOX2 and SP1.
Thus, we suggest that the role of IGF1 in the
development and progression of cervical cancer appears to be more
complex than previously described. In particular, due to the
up-regulation of the IGF1Eb isoform with mitogenic properties, its
significance in cancerogenesis may be more pronounced. Although the
impact of HPV infection on the course of IGF1 pre-mRNA splicing as
well as on IGF1 promoter activation has not yet been resolved,
numerous research results have demonstrated that HPV proteins may
impair this process. Furthermore, HPV proteins affect splicing both
directly, as spliceosome components, and indirectly, via the
activation of splicing factors.
Further investigation towards a better understanding
of the mechanisms of IGF1 expression in HPV-infected cells will be
helpful to develop highly sensitive and multifactorial tools for
the prevention, diagnosis and treatment of cervical cancer.
Acknowledgements
This study was supported by the
ministry of Higher Education in Poland, grant no. N N401219824. The
authors acknowledge Dr Agata Smolen and Professor Maria Kaczmarek
for their assistance with the statistical analysis.
References
|
1.
|
Doorbar J: Molecular biology of human
papillomavirus infection and cervical cancer. Clin Sci.
110:525–541. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
2.
|
Madkan VK, Cook-Norris RH, Steadman MC,
Arora A, Mendoza N and Tyring SK: The oncogenic potential of human
papillomaviruses: a review on the role of host genetics and
environmental cofactors. Br J Dermatol. 157:228–241. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
3.
|
Samani AA, Yakar S, LeRoith D and Brodt P:
The role of the IGF system in cancer growth and metastasis:
overview and recent insights. Endocr Rev. 28:20–47. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
4.
|
Cohen P: Overview of the IGF-I system.
Horm Res. 65(Suppl 1): 3–8. 2006. View Article : Google Scholar
|
|
5.
|
Pollak M: Insulin and insulin-like growth
factor signalling in neoplasia. Nat Rev Cancer. 8:915–928. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
6.
|
Mittanck DW, Kim SW and Rotwein P:
Essential promoter elements are located within the 5′ untranslated
region of human insulin-like growth factor-I exon I. Mol Cell
Endocrinol. 126:153–163. 1997.
|
|
7.
|
Wierstra I: Sp1: emerging roles – beyond
constitutive activation of TATA-less housekeeping genes. Biochem
Biophys Res Commun. 372:1–13. 2008.
|
|
8.
|
LeRoith D and Roberts CT Jr: The
insulin-like growth factor system and cancer. Cancer Lett.
195:127–137. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
9.
|
Musarò A, Dobrowolny G and Rosenthal N:
The neuroprotective effects of a locally acting IGF-1 isoform. Exp
Gerontol. 42:76–80. 2007.PubMed/NCBI
|
|
10.
|
Gee SL, Aoyagi K, Lersch R, Hou V, Wu M
and Conboy JG: Alternative splicing of protein 4.1R exon 16:
ordered excision of flanking introns ensures proper splice site
choice. Blood. 95:692–699. 2000.PubMed/NCBI
|
|
11.
|
Baraniak AP, Chen JR and Garcia-Blanco A:
Fox-2 mediates epithelial cell-specific fibroblast growth factor
receptor 2 exon choice. Mol Cell Biol. 26:1209–1222. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
12.
|
Zhang C, Zhang Z, Castle J, Sun S, Jahnson
J, Krainer AR and Zhang MQ: Defining the regulatory network of the
tissue-specific splicing factors FOX-1 and FOX-2. Genes Dev.
22:2550–2563. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
13.
|
Jozefiak A, Pacholska-Bogalska J,
Myga-Nowak M, Kedzia W, Kwasniewska A, Luczak M, Kedzia H and
Gozdzicka-Jozefiak A: Serum and tissue levels of insulin-like
growth factor-I in women with dysplasia and HPV-positive cervical
cancer. Mol Med Rep. 1:231–237. 2008.PubMed/NCBI
|
|
14.
|
Serrano ML, Sánchez-Gómez M and Bravo MM:
Insulin-like growth factor system gene expression in cervical
scrapes from women with squamous intraepithelial lesions and
cervical cancer. Growth Horm IGF Res. 17:492–499. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
15.
|
Tan DS, Cook A and Chew SL: Nucleolar
localization of an isoform of the IGF-I precursor. BMC Cell Biol.
3:172002. View Article : Google Scholar : PubMed/NCBI
|
|
16.
|
Tavassoli FA and Deville P; World Health
Organization Classification of Tumors: Pathology, Genetics. Tumours
of the Breast and Female Genital Organs. IRAC Press; Lyon: 2003
|
|
17.
|
Kosary CL: FIGO stage, histology,
histologic grade, age and race as prognostic factors in determining
survival for cancers of the female gynecological system: an
analysis of 1973–87 SEER cases of cancers of the endometrium,
cervix, ovary, vulva, and vagina. Semin Surg Oncol. 10:31–46.
1994.PubMed/NCBI
|
|
18.
|
Manos MM, Ting Y, Wright DK, Lewis AI,
Broker TR and Wolinsky SM: The use of polymerase chain reaction
amplification for the detection of genital human papillomaviruses.
Cancer Cells. 7:209–214. 1989.
|
|
19.
|
Daud II and Scott ME: Validation of
reference genes in cervical cell samples from human
papillomavirus-infected and -uninfected women for quantitative
reverse transcription-PCR assays. Clin Vaccine Immunol.
15:1369–1373. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
20.
|
Philippou A, Papageorgiou E, Bogdanis G,
Halapas A, Sourla A, Maridaki M, Pissimissis N and Koutsilieris M:
Expression of IGF-1 isoforms after exercise-induced muscle damage
in humans: characterization of the MGF E peptide actions in vitro.
In Vivo. 23:567–575. 2009.PubMed/NCBI
|
|
21.
|
Matheny RW, Nindl BC and Adamo MI:
Mechano-growth factor: a putative product of IGF 1 gene expression
involved in tissue repair and regeneration. Endocrinology.
151:3865–3875. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
22.
|
Barton ER: The ABC of IGF 1 isoforms:
impact on muscle hypertrophy and implication for repair. Appl
Physiol Nutr Metab. 31:791–797. 2006. View
Article : Google Scholar : PubMed/NCBI
|
|
23.
|
Górecki DC, Beręsewicz M and Zabłocka B:
Neuroprotective effects of short peptides derived from the
insulin-like growth factor 1. Neurochem Int. 51:451–458.
2007.PubMed/NCBI
|
|
24.
|
Vinciquerra M, Santini MP, Claycomb WC,
Ladurner AG and Rosenthal N: Local-IGF-1 isoform protects
cardiomyocytes from hypertrophic and oxidative stresses via Sirt1
activity. Aging (Albany NY). 43–62. 2010.PubMed/NCBI
|
|
25.
|
Ohtsuki T, Otsuki M, Murakami Y, Maekowa
T, Yamamoto T, Kasaka K, Takenchi S and Takahashi S: Organ-specific
and age-dependent expression of insulin-like growth factor-I
(IGF-I) mRNA variants: IGF-IA and IB mRNAs in the mouse. Zoolog
Sci. 22:1011–1021. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
26.
|
Pfeffer LA, Brisson BK, Lei H and Barton
ER: The insulin-like growth factor (IGF)-I E-peptides modulate cell
entry of the mature IGF-I protein. Mol Biol Cell. 20:3810–3817.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
27.
|
Bell I, Martin A and Roberts S: The E1^E4
protein of human papillomavirus interacts with the
serine-arginine-specific protein kinase SRPK1. J Virol.
81:5437–5448. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
28.
|
Bodaghi S, Jia R and Zheng ZM: Human
papillomavirus type 16 E2 and E6 are RNA-binding proteins and
inhibit in vitro splicing of pre-mRNAs with suboptimal splice
sites. Virology. 386:32–43. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
29.
|
Cheunim T, Zhang J, Milligan SG,
McPhillips MG and Graham SV: The alternative splicing factor hnRNP
A1 is up-regulated during virus-infected epithelial cell
differentiation and binds the human papillomavirus type 16 late
regulatory element. Virus Res. 131:189–198. 2008. View Article : Google Scholar
|
|
30.
|
Mole S, McFarlane M, Chuen-Im T, Milligan
SG, Millan D and Graham SV: RNA splicing factors regulated by HPV16
during cervical tumour progression. J Pathol. 219:383–391. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
31.
|
Venables JP, Klinck R, Koh C, et al:
Cancer-associated regulation of alternative splicing. Nat Struct
Mol Biol. 16:670–676. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
32.
|
Li T, Chen YH, Liu TJ, Jia J, Hampson S,
Shan YX, Kibler D and Wang PH: Using DNA microarray to identify Sp1
as a transcriptional regulatory element of insulin-like growth
factor 1 in cardiac muscle cells. Circ Res. 93:1202–1209. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
33.
|
Gloss B and Bernard HU: The E6/E7 promoter
of human papillomavirus type 16 is activated in the absence of E2
proteins by a sequence-aberrant Sp1 distal element. J Virol.
64:5577–5584. 1990.PubMed/NCBI
|
|
34.
|
McPhillips MG, Veerapraditsin T, Cumming
SA, Karali D, Milligan SG, Boner W, Morgan IM and Graham SV:
SF2/ASF binds the human papillomavirus type 16 late RNA control
element and is regulated during differentiation of virus-infected
epithelial cells. J Virol. 78:10598–10605. 2004. View Article : Google Scholar
|
|
35.
|
Li B, Wang X, Zhou F, Saunders NA, Frazer
IH and Zhao KN: Up-regulated expression of Sp1 protein coincident
with a viral protein in human and mouse differentiating
keratinocytes may act as a cell differentiation marker.
Differentiation. 76:1068–1080. 2008. View Article : Google Scholar
|
|
36.
|
Carboni JM, Lee AV, Hadsell DL, et al:
Tumor development by transgenic expression of a constitutively
active insulin-like growth factor I receptor. Cancer Res.
65:3781–3787. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
37.
|
Linnerth NM, Siwicky MD, Campbell CI,
Watson KL, Petrik JJ, Whitsett JA and Moorehead RA: Type I
insulin-like growth factor receptor induces pulmonary
tumorigenesis. Neoplasia. 11:672–682. 2009.PubMed/NCBI
|
|
38.
|
Chong YM, Colston K, Jiang WG, Sharma AK
and Mokbel K: The relationship between the insulin-like growth
factor-1 system and the oestrogen metabolising enzymes in breast
cancer tissue and its adjacent non-cancerous tissue. Breast Cancer
Res Treat. 99:275–288. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
39.
|
Weber MM, Fottner C, Liu SB, Jung MC,
Engelhardt D and Baretton GB: Overexpression of the insulin-like
growth factor I receptor in human colon carcinomas. Cancer.
95:2086–2095. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
40.
|
Werner H, Shalita-Chesner M, Abramovitch
S, Idelman G, Shaharabani-Gargir L and Glaser T: Regulation of the
insulin-like growth factor-I receptor gene by oncogenes and
antioncogenes: implications in human cancer. Mol Genet Metab.
71:315–320. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
41.
|
Kuramoto H, Hongo A, Liu Y, Ojima Y,
Nahamura K, Seki N, Kodama J and Hiramatsu Y: Immunohistochemical
evaluation of insulin-like growth factor 1, receptor status in
cervical cancer specimens. Acta Medica Okayama. 62:251–259.
2008.PubMed/NCBI
|