Introduction
Bacterial infection of the central nervous system is
a common and serious threat to human life, and requires timely and
effective antibiotic treatment (1). For this to occur, a rapid and
accurate detection and identification of the cerebrospinal fluid
(CSF) pathogens is necessary. The diagnosis of intracranial
infection mainly relies on CSF bacterial culture, which is
considered to be the gold standard due to its high specificity
(2,3). However, this is a time consuming
procedure, and specific bacteria are difficult to cultivate. In
addition, given the influence of antibiotics, among other factors,
the positive rate of CSF culture is minimal, i.e. ~10% in the
hospital and 10–20% in the majority of studies (3,4).
Therefore, the current methods of etiological examination are
inadequate when compared with the advancements in clinical
treatment (1). Microarray
technology has the advantages of easy operation, rapid detection
and the ability to simultaneously detect a large number of specific
molecules. In the present study, several types of typical
intracranial infection-causing pathogens and their common
resistance genes were selected based on their specific DNA
sequence. A small microarray was designed and prepared to study the
application value of microarray technology in intracranial
infection.
Materials and methods
Types of bacteria and drug resistance
genes
Based on the common intracranial infection-causing
bacteria indicated by the Neurosurgery Department of The Affiliated
Hospital of Nantong University (Nantong, China) and The Affiliated
Haian People’s Hospital of Nantong University (Haian, China), the
following Gram-positive and Gram-negative cocci and bacilli were
used: Staphylococcus aureus, Klebsiella pneumoniae, Escherichia
coli and Streptococcus pneumoniae. In addition, the six
most common drug resistance genes of these four bacteria (mecA,
OXA-23, SHV, CTX-M, TEM and PBP1a) were tested. This study was
conducted in accordance with the Declaration of Helsinki and with
approval from the Ethics Committee of the Affiliated Haian People’s
Hospital of Nantong University (Haian, China). Written informed
consent was obtained from all participants.
Clinical specimen collection
Based on the detected stains, 30 CSF samples from
patients with clinically diagnosed intracranial infection were
collected between January 2010 and August 2011 at the Neurosurgery
Departments of the Affiliated Hospital of Nantong University
(Nantong, China) and the Affiliated Haian People’s Hospital of
Nantong University. Among the samples, bacterial culture revealed
that 12 cases were positive (five for S. aureus, three for
K. pneumoniae, two for E. coli and two for S.
pneumoniae) and 18 cases were negative (the negative CSF
samples within the same period were generated numbers and randomly
selected). The clinical diagnosis of intracranial bacterial
infection was based on the Harrison standard (5), combined with the following
neurosurgical characteristics: i) Risk factors for intracranial
infection, including CSF leak, open brain injury, surgery for an
extended period (>4 h), >2 surgeries and external drainage of
CSF; ii) clinical manifestations of fever, headache, vomiting or
meningeal irritation; and iii) white blood cell count
>1.18×109cells/l, glucose <1.9 mmol/l and protein
>2.2g/l, as assessed with CSF testing.
CSF bacterial culture
Loop-picked turbid CSF was inoculated on blood agar
and chocolate agar plates at 35°C under 5% CO2 for 24
h.
Bacterial DNA extraction from CSF
Each CSF sample (~2 ml) was collected and
centrifuged at 8,000 × g for 10 min. The supernatant was then
discarded and the sediment was suspended in sterile saline, prior
to being subjected to centrifugation at 8,000 × g for 10 min.
Having discarded the supernatant, a DNeasy® Blood &
Tissue kit (Qiagen GmbH, Hilden, Germany) was used. DNA lysate
(~180 μl) was added to the sediment, the mixture was placed in a
37°C water bath for 30 min and ~25 μl proteinase K being added. The
mixture was subsequently placed in a 56°C water bath for 30 min and
200 μl ethanol was added using spin columns to extract the sample
DNA.
Primer design and synthesis of
probes
The specific DNA sequences were screened for four
types of bacteria and six resistance genes from GenBank (http://www.ncbi.nlm.nih.gov/genbank/),
using the software Primer Premier 5.0 (Premier Biosoft, Palo Alto,
CA, USA) to design 10 pairs of PCR primers and probes (Table I). The 16S rDNA gene codes for
prokaryotic ribosomal small subunit rRNA (16S rRNA) and is the most
common and useful ‘molecular clock’ in bacterial taxonomic studies.
The constant region of 16S rRNA is a common feature of all bacteria
(6). Therefore, based on the
constant region of the 16S gene found in all strains of bacteria,
primers and a probe were designed as a positive reference. All
primers and probes were synthesized by Shanghai Invitrogen
Biotechnology Co., Ltd. (Shanghai, China).
 | Table ISequences of multiplex polymerase
chain reaction primers and microarray hybridization probes. |
Table I
Sequences of multiplex polymerase
chain reaction primers and microarray hybridization probes.
| Name of bacteria and
resistance genes | Primer and probe
sequence (5′-3′) | Product size, bp |
|---|
| Staphylococcus
aureus | Sense primer:
TAAAGCGATTGATGGTGATACG | 238 |
| Antisense primer:
AGCCAAGCCTTGACGAACTA | |
| Probe:
AGCGAGCATACGGCAATACTCGTTGACTGCCTCTTCGCTGT | |
| Klebsiella
pneumoniae | Sense primer:
GCCTTGACCGCTGGGAAAC | 319 |
| Antisense primer:
GGCGTATCCCGCAGATAAAT | |
| Probe:
CAACGCACTGACCATACCTACTTTGTTATTCGGGCCAAGC | |
| Escherichia
coli | Sense primer:
CATGCGGTTCAGCCACGGTT | 471 |
| Antisense primer:
GCGCCAGTATTCCGCACCAA | |
| Probe:
CGAATCAGTCTTGCTCATCGTCGCTATCTGGCTGACTGCTT | |
| Pneumococcal | Sense primer:
CATTGTCTTAGGCGGAG | 679 |
| Antisense primer:
ATTGGTGTATTGACTGC | |
| Probe:
CGTTGCCGAGTTTCCATGTAGGTCTTTACCATAGTAGTTTTG | |
| 16S | Sense primer:
AGGAGGTGATCCAACCGCA | 370 |
| Antisense primer:
AACTGGAGGAAGGTGGGGAT | |
| Probe:
AGCTCACCATGTACGAACTGGGTGAATACGTTCCCGGGCCTTGT | |
| mecA | Sense primer:
GGCTATCGTGTCACAATCGTTGACG | 170 |
| Antisense primer:
GGGTGGATAGCAGTACCTGAGCCA | |
| Probe:
CGTATCGACTGCATCAATCCAGATGGCAAAGATATTCAACTAACT | |
| OXA-23 | Sense primer:
ATGGAAGGGCGAGAAAAGG | 127 |
| Antisense primer:
TTGCATGAGATCAAGACCGATA | |
| Probe:
AGTGGATCTTGTACGTGGACCGCAAGTTCCTGATAGACTGGGACTGCC | |
| SHV | Sense primer:
GCCTTGACCGCTGGGAAAC | 319 |
| Antisense primer:
GGCGTATCCCGCAGATAAAT | |
| Probe:
CGAATCAGTCTTGCTCATCGTGTCGCCCTGCTTGGCCCGGATAAC | |
| CTX-M | Sense primer:
CGGGAGGCAGACTGGGTGT | 381 |
| Antisense primer:
TCGGCTCGGTACGGTCGA | |
| Probe:
CCTGACTGCAATAGATCCTGACGGCCATCACTTTACTGGTGCTGC | |
| TEM | Sense primer:
GTCGCCGCATACACTATTCTCA | 258 |
| Antisense primer:
CGCTCGTCGTTTGGTATGG | |
| Probe:
GTCAGCGAGAACATGTGTACGCGGTTAGCTCCTTCGGTCCTCCG | |
| PBP1a | Sense primer:
AGTATTCACTACTCAAATGC | 557 |
| Antisense primer:
GCTACAAATTGAGAGGTGTT | |
| Probe:
CACTGAACAGCTGACATACG GGCAGCGTAAGCAGCAGCCATCT | |
Multiplex polymerase chain reaction
(PCR)
The experiments were divided into two groups
(Table I). The first group
underwent microbiological testing, while the second group underwent
resistance gene testing with water as a negative control. The
reaction system contained the following: 1.5 μl buffer (10X), 0.2
μl deoxyribonucleotide triphosphate (10 mmol/l), 1.0 μl DNA (20
ng/μl), 0.2 μl Taq enzyme, 0.2 μl primers ×2 (20 μmol/l),
0.6 μl MgCl2 (25 mmol/l) and 11.1 μl H2O. The
total volume was 15 μl (Tm, 56°C; 30 cycles). The PCR
products were analyzed using agarose gel electrophoresis (2%
agarose; voltage, 150 V; running time, 15 min), and the bands were
observed.
Microarray preparation
The PCR products were diluted to 50 mmol/l with
spotting solution and added to 384-well plates (Corning Life
Sciences, Tewksbury, MA, USA) at 10 μl per well. Under 60% humidity
and 25°C, contact spotting (Omni Grid™ 100 microarray spotter;
GeneMachine, USA) was performed to load the probe point to the
optical level of the aldehyde modification chip (Boao Biology Co.,
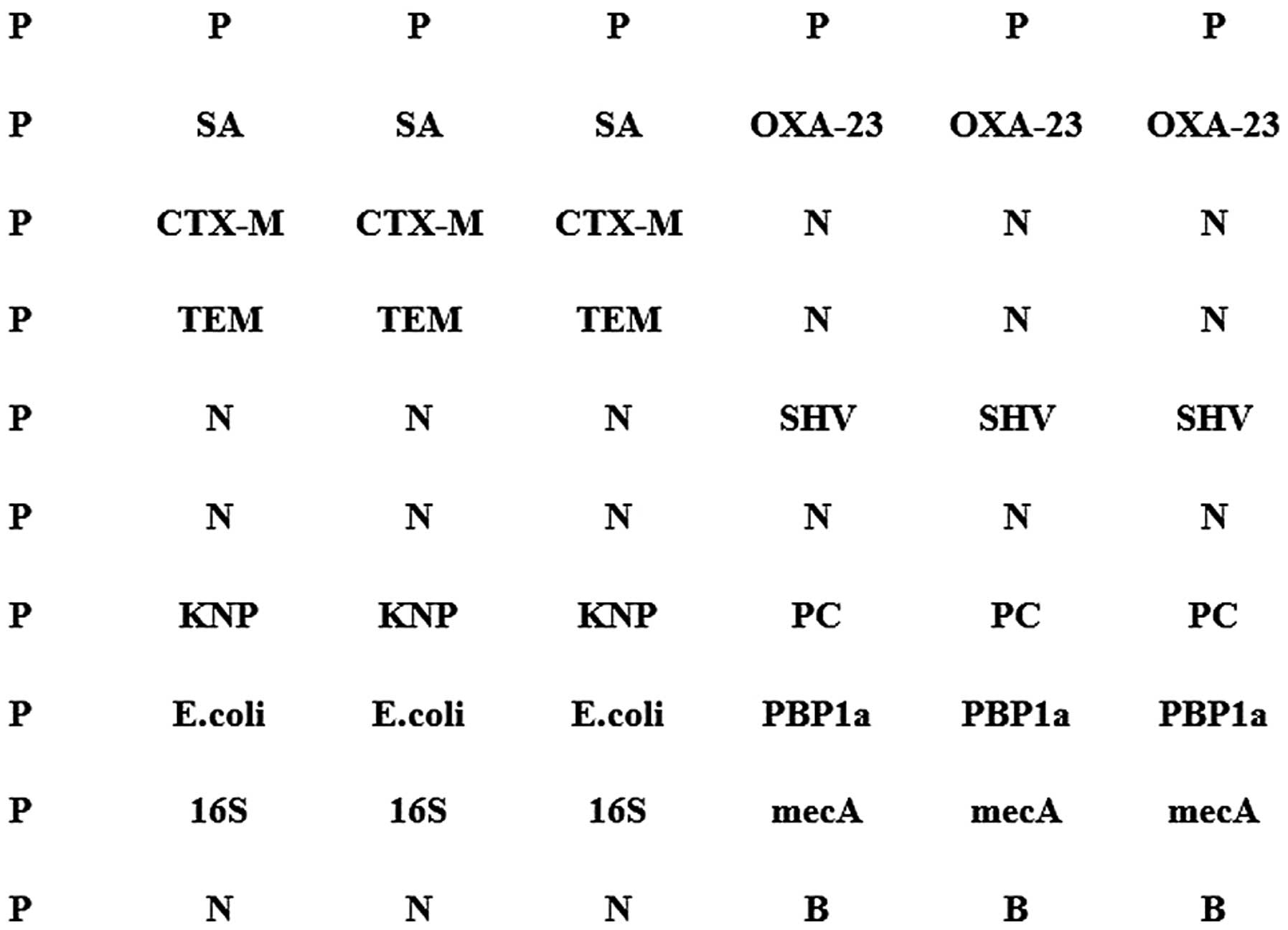
Ltd., Beijing, China). The matrix measured 10×7, and all points
were randomly arranged. Each probe set was repeated in triplicate
(Fig. 1). The microarray was
supplied by Shanghai Biochip Co., Ltd. (Shanghai, China) and placed
in the oven during storage.
Microarray hybridization and result
interpretation
The PCR products were fluorescently labeled under
the following conditions: 96°C for 3 min, followed by 66°C for 30
sec, 72°C for 20 sec for 35 cycles and extension at 72°C for 5 min.
The products were then placed in the dark at 4°C. The
fluorescently-labeled products were subjected to DNA hybridization
(Thermo Hybaid Maxi 14 Hybridization Oven; Thermo Hybaid, Ulm,
Germany) at 48°C for 2 h. The hybridized chip was then scanned
(GenePix® 4000B confocal laser scanner; Molecular
Devices, LLC, Sunnyvale, CA, USA), and GenePix® Pro 6.0
Acquisition and Analysis Microarray Software (Molecular Devices,
LLC) was used to assess the fluorescence signal intensity value of
each probe set. The low-signal locus was removed, and values higher
than the cutoff value [signal-to-noise ratio, 3.0] were deemed as a
valid signal.
Results
Multiplex PCR
Ten pairs of primers were designed to amplify the
corresponding target bacteria and resistance gene sequences. The
PCR products were then subjected to agarose gel electrophoresis,
showing clear bands of the appropriate size (Fig. 2). The results demonstrated the
specification and effectiveness of the designed primer
sequences.
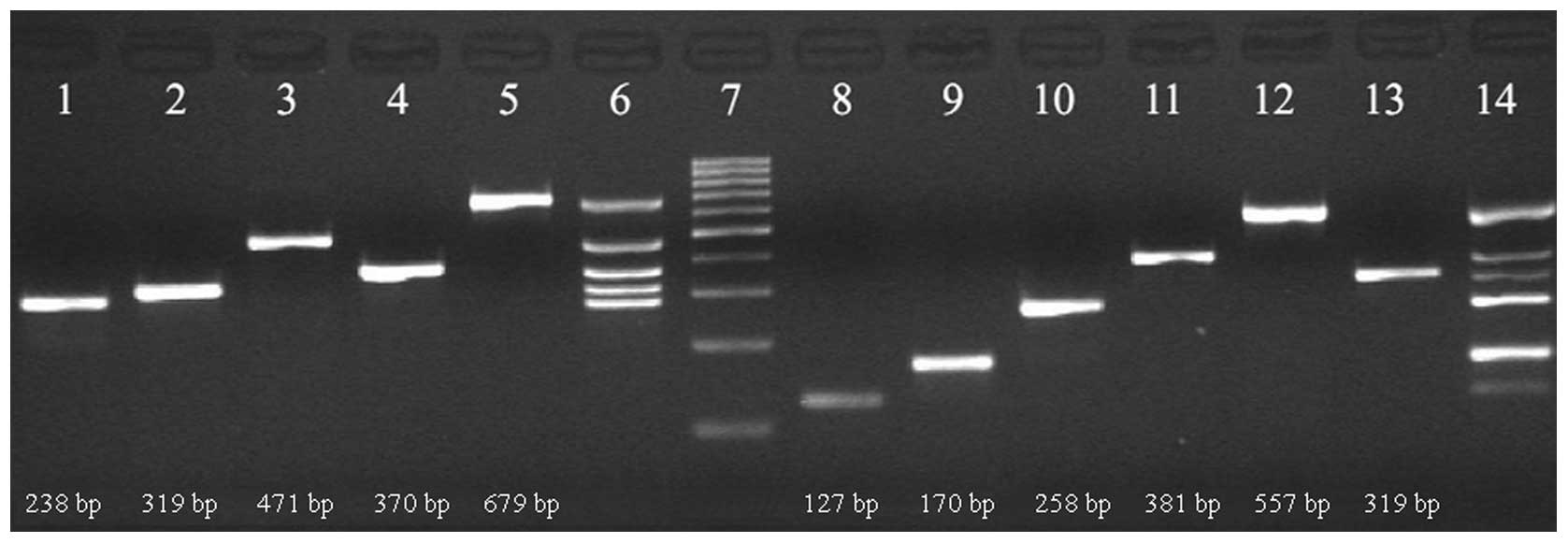 | Figure 2Detection of bacterial strains and
resistance genes using electrophoresis of the PCR products. Lane 1,
Staphylococcus aureus; 2, Klebsiella pneumoniae; 3,
Escherichia coli; 4, 16S; 5, Streptococcus
pneumoniae; 6, multiplex PCR of bacteria; 7, Marker; 8, OXA-23;
9, mecA; 10, TEM; 11, CTX-M; 12, PBP1a; 13, SHV; 14, multiplex PCR
of resistance genes. The marker strips from the bottom to the top
were: 100, 200, 300, 400, 500, 600, 700, 800, 900 and 1,000 bp.
PCR, polymerase chain reaction. |
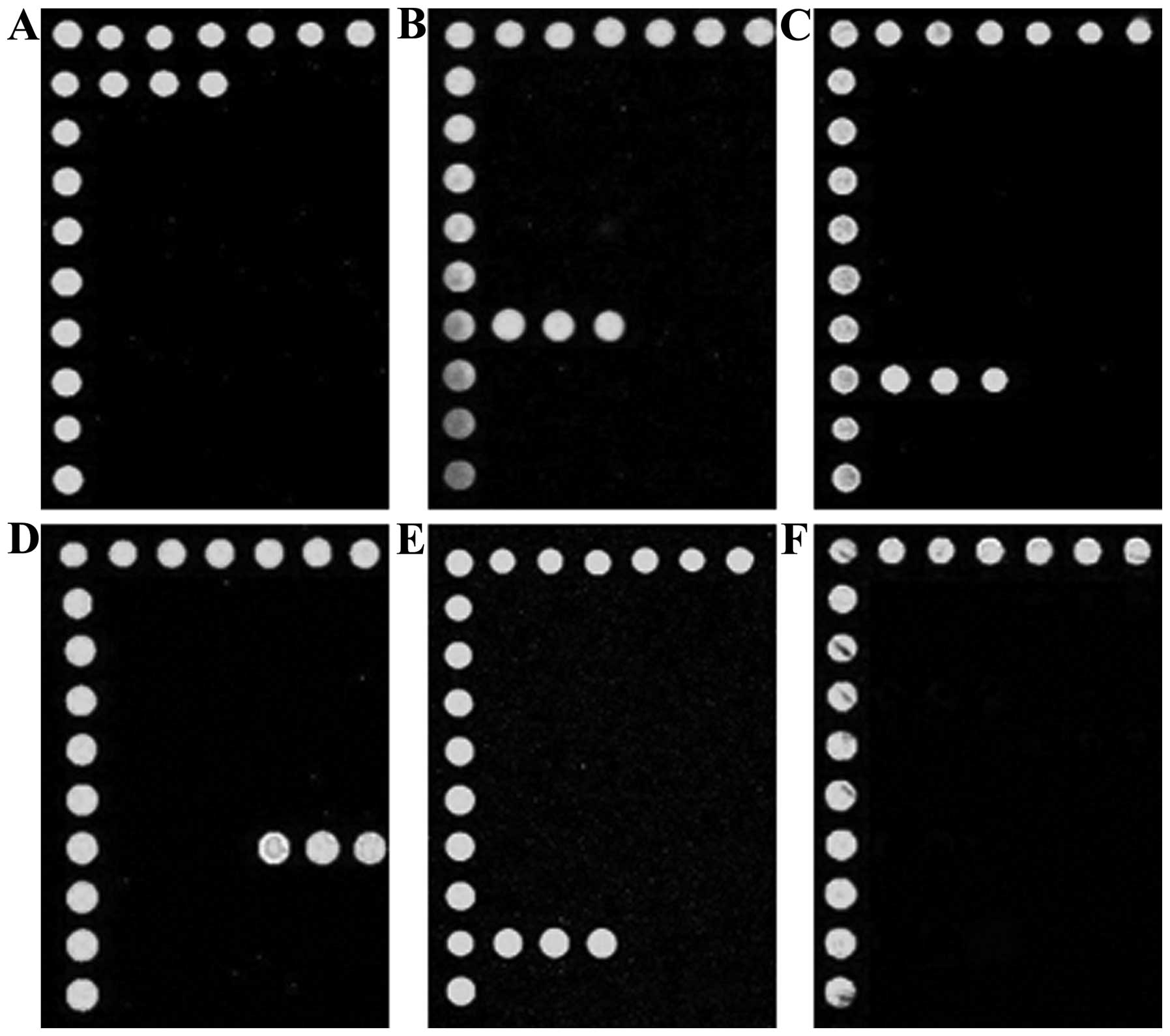
Microarray hybridization
Following hybridization with the multiplex PCR
products and specific probes, the microarray showed green
fluorescence at the corresponding sites (Fig. 3), whereas the negative control
showed no fluorescence. As a result, the bacterial species and
resistance genes were identified.
Microarray comparison with CSF culture
results
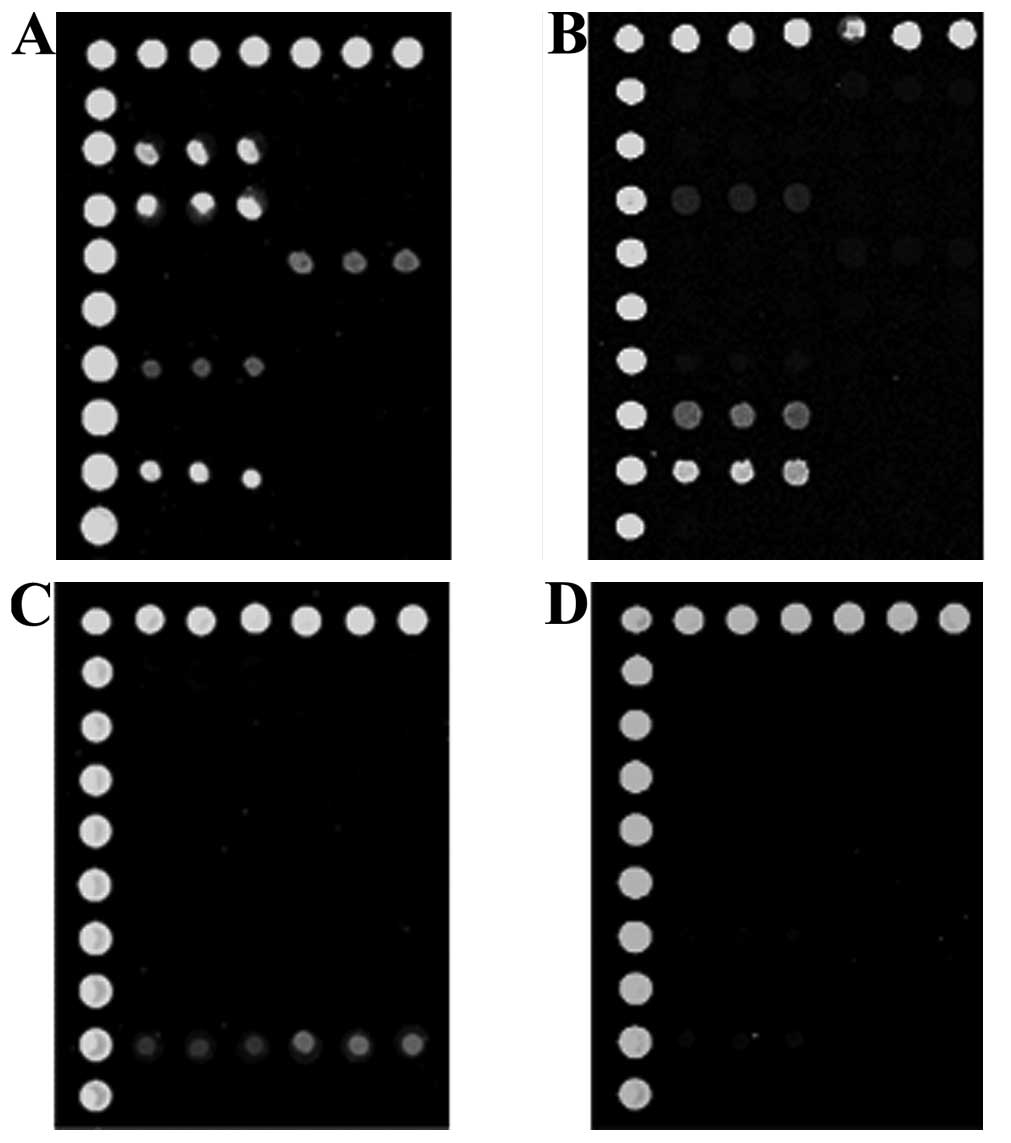
The microarray procedure took one day, whereas
bacterial culture and sensitivity testing took 4–5 days. A total of
12 CSF samples with positive bacterial cultures were identified as
being positive for bacterial strains and resistance genes (Fig. 4A) using microarray, including five
that were positive for S. aureus, three for K.
pneumonia, two for E. coli and two for S.
pneumoniae, consistent with the bacterial culture results.
Among the 18 specimens that had negative bacterial culture results,
bacteria and drug resistance genes were identified in a number of
samples (Fig. 4B), including one
sample positive for S. aureus and two for E. coli.
The 16S gene without bacteria was detected in eight cases. However,
the majority of these 8 cases (six) were positive for resistance
genes (Fig. 4C). Samples from
seven patients were without detectable 16S or drug resistance genes
(Fig. 4D).
Discussion
Although CSF bacterial culture is an important
technique for the diagnosis of intracranial bacterial infection,
the positive rate is too low and the process is time consuming
(2,7). These disadvantages hinder the
application of CSF bacterial culture in clinical treatment;
therefore, various testing methods have been proposed as a
supplement or substitute for bacterial culture, for example PCR
(2,3,8) and
immunological analyses (9).
Multiplex PCR technology involves adding various specific primers
in the same PCR system to simultaneously detect multiple pathogens
or resistance genes (2,10). With regard to microarray
technology, a large number of probes must be fixed onto a support
to detect and analyze a variety of sample sequences (11). Since numerous pathogens cause
intracranial infection, the two technologies were combined in the
present study to detect four types of bacteria and six resistance
genes. All experiments were performed in one day, showing that the
process was an efficient means of genetic testing (12), facilitating the rapid detection of
pathogens causing intracranial infection.
Twelve cases of positive bacterial culture specimens
were identified to have the same strains based on the microarray
results. Among the 18 cases of culture-negative specimens, 11 were
shown to be positive following gene chip hybridization of the 16S
gene, demonstrating the presence of bacterial infection. This
indicated that microarray technology had a higher sensitivity than
CSF culture. Eight cases were shown to be positive for the 16S gene
without bacteria being identified. This may have been due to the
pathogens not belonging to one of the four species included in the
study design. The 15 positive specimens (12 positive in culture and
3 negative in culture but positive in microarray) showed only one
result, indicating that microarray technology had a high
specificity.
Among the 30 CSF samples that were diagnosed as
having intracranial bacterial infections, seven cases failed to
pass the gene chip detection for the presence of bacteria. There
were several possible reasons for this. The experiments were
conducted using a fluorescent-labeling method to perform the gene
hybridization and interpret the results. When the conventional
material for the fluorescent-labeling method was used, the
sensitivity was low (13). In
addition, the bacterial DNA content of specific samples was too
low. The required level of fluorescently-labeled DNA was not
achieved, even following several amplifications. Furthermore,
steric hindrance existed between the target molecule and probe, and
the hybridization probe molecules affected the quality of results
(14). Therefore, negative
microarray results were not able to be used as a reliable indicator
of a definitive negative clinical diagnosis. In addition,
microarray is not recommended for patients with a high positive
rate of bacterial culture, such as lung infection.
Antibiotic resistance is a common phenomenon,
particularly when antibiotics are frequently used (15). Microarray technology is capable of
rapidly detecting common resistance genes in the case of unknown
bacteria, in accordance with the various resistance genes supplying
direction for early clinical trials of drugs (16). TEM, CTX-M and OXA-23 are
hyperspectral β-lactamases that have been associated with drug
resistance to penicillin and cephalosporins (17). mecA is a type of
methicillin-resistant gene, and has been associated with drug
resistance to gentamicin, imipenem and cephalosporins (18). Furthermore, the KPC gene encodes
carbapenem, which has been associated with drug resistance to
imipenem (19). Results from the
experimental detection of resistance genes corroborated with the
susceptibility test results, showing the reliability of the
experimental detection of resistance genes. Of course, the
detection of resistance genes also has significant limitations, for
it may only help to avoid the use of partial tolerant antibiotics.
However, resistance gene detection is not able to guide the
selection of sensitive drugs and therefore is not able to
substitute for susceptibility testing. Nevertheless, in the case of
culture-negative CSF, the test result of resistance genes is the
only reference index available.
The small chip experiments of the current study
demonstrate that microarray technology has advantages in terms of
speed and sensitivity compared with traditional CSF bacterial
cultures. Based on the detection of resistance genes, microarray
technology also avoids the use of antibiotics as soon as possible.
However, specific disadvantages, such as high cost and low
sensitivity, exist. The present trends in microarray technology
have three main aspects (20,21):
i) High-density probe analysis; ii) microanalysis of the test
samples; and iii) microanalysis of the chip matrix area. These
aspects further enhance detection sensitivity and greatly reduce
the cost of testing. With the development of chip technology,
microarray technology shows potential for the diagnosis of
bacterial culture diseases with low positive rates, such as
intracranial infection.
Acknowledgements
This study was supported by the Social Development
Science and Technology Project of Nantong (grant no. S10935).
References
|
1
|
Lin AL and Safdieh JE: The evaluation and
management of bacterial meningitis: current practice and emerging
developments. Neurologist. 16:143–151. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Favaro M, Savini V, Favalli C and Fontana
C: A multi-target real-time PCR assay for rapid identification of
meningitis-associated microorganisms. Mol Biotechnol. 53:74–79.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Wu HM, Cordeiro SM, Harcourt BH, et al:
Accuracy of real-time PCR, Gram stain and culture for
Streptococcus pneumoniae, Neisseria meningitidis and
Haemophilus influenzae meningitis diagnosis. BMC Infect Dis.
13:262013. View Article : Google Scholar
|
|
4
|
Srinivasan L, Pisapia JM, Shah SS, Halpern
CH and Harris MC: Can broad-range 16S ribosomal ribonucleic acid
gene polymerase chain reactions improve the diagnosis of bacterial
meningitis? A systematic review and meta-analysis. Ann Emerg Med.
60:609–620. 2012. View Article : Google Scholar
|
|
5
|
Harrison PJ: Bacterial meningitis. Lancet.
2:8481979. View Article : Google Scholar
|
|
6
|
Santoni D and Romano-Spica V: A gzip-based
algorithm to identify bacterial families by 16S rRNA. Lett Appl
Microbiol. 42:312–314. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Etyang AO, Amayo EO, Bhatt SM, Wamola IA
and Maritim MC: Comparison of bedside inoculation of culture media
with conventional cerebrospinal fluid culture method in patients
with bacterial meningitis. East Afr Med J. 86:476–479. 2009.
|
|
8
|
Sanou M, Palenfo D, Bisseye C, et al:
Molecular diagnosis of bacterial meningitis in Burkina Faso. Med
Sante Trop. 23:93–99. 2013.(In French).
|
|
9
|
Ouédraogo SM, Yaméogo TM, Kyelem CG, et
al: Acute bacterial meningitis with soluble antigen detected by
latex particle agglutination tests at the Sourô-Sanou University
Hospital of Bobo-Dioulasso (Burkina Faso). Med Sante Trop.
22:412–416. 2012.(In French).
|
|
10
|
Gupta S, Bandyopadhyay D, Paine SK, et al:
Rapid identification of mycobacterium species with the aid of
multiplex polymerase chain reaction (PCR) from clinical isolates.
Open Microbiol J. 4:93–97. 2010.PubMed/NCBI
|
|
11
|
Palka-Santini M, Pützfeld S, Cleven BE,
Krönke M and Krut O: Rapid identification, virulence analysis and
resistance profiling of Staphylococcus aureus by gene
segment-based DNA microarrays: application to blood culture
post-processing. J Microbiol Methods. 68:468–477. 2007.PubMed/NCBI
|
|
12
|
Shi J, Wu Y, Cai M and Shang S: Rapid
diagnosis of herpetic encephalitis in children by PCR-microarray
technology for simultaneous detection of seven human herpes
viruses. Eur J Pediatr. 169:421–425. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Mathias PC, Jones SI, Wu HY, et al:
Improved sensitivity of DNA microarrays using photonic crystal
enhanced fluorescence. Anal Chem. 82:6854–6861. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Southern EM: DNA microarrays. History and
overview. Methods Mol Biol. 170:1–15. 2001.PubMed/NCBI
|
|
15
|
Sengupta S, Chattopadhyay MK and Grossart
HP: The multifaceted roles of antibiotics and antibiotic resistance
in nature. Front Microbiol. 4:472013. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Bruant G, Maynard C, Bekal S, et al:
Development and validation of an oligonucleotide microarray for
detection of multiple virulence and antimicrobial resistance genes
in Escherichia coli. Appl Environ Microbiol. 72:3780–3784.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Chen H, Shu W, Chang X, Chen JA, Guo Y and
Tan Y: The profile of antibiotics resistance and integrons of
extended-spectrum beta-lactamase producing thermotolerant coliforms
isolated from the Yangtze River basin in Chongqing. Environ Pollut.
158:2459–2464. 2010. View Article : Google Scholar
|
|
18
|
Gardete S, de Lencastre H and Tomasz A: A
link in transcription between the native pbpB and the acquired mecA
gene in a strain of Staphylococcus aureus. Microbiology.
152:2549–2558. 2006. View Article : Google Scholar
|
|
19
|
Papp-Wallace KM, Bethel CR, Distler AM,
Kasuboski C, Taracila M and Bonomo RA: Inhibitor resistance in the
KPC-2 beta-lactamase, a preeminent property of this class A
beta-lactamase. Antimicrob Agents Chemother. 54:890–897. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Ahmad H, Sutherland A, Shin YS, et al: A
robotics platform for automated batch fabrication of high density,
microfluidics-based DNA microarrays, with applications to single
cell, multiplex assays of secreted proteins. Rev Sci Instrum.
82:0943012011. View Article : Google Scholar
|
|
21
|
Bumgarner R: Overview of DNA microarrays:
types, applications, and their future. Curr Protoc Mol Biol.
22.1.1–22.1.11. 2013.PubMed/NCBI
|