Introduction
Abnormal scar formation in wound healing causes
aesthetic issues, particularly for female patients. Abnormal scars,
including keloid and hypertrophic scars, are characterized by
abnormal fibroproliferation and increased extracellular matrix
production (1,2). In previous years, there have been a
number of studies investigating the relationship between p53
gene polymorphisms and breast cancer (3,4).
Abnormal scars, particularly keloid scars, are considered to have
specific characteristics that are similar to those of benign dermal
fibroproliferative tumors. The p53 mutations in dermal
fibroblasts are considered to contribute to the formation of
keloids (5,6). In addition, the relationship between
p53 polymorphisms and the susceptibility to form
hypertrophic scars is an area of interest for a number of
researchers (7–9). However, the majority of studies have
focused on the differences between patients with abnormal scars and
normal individuals. The aim of the present study was to investigate
the relationship between the polymorphisms of p53 codon-72
and the occurrence of hypertrophic scars for patients receiving a
caesarean section (CS).
Materials and methods
Patients
In total, 260 female patients with an average age of
29.4 years (ranging between 20 and 39 years) and of the same
ethnicity were selected from the Nanfang Hospital of Southern
Medical University (Guangzhou, China) for the study. The patients
did not have recorded histories of pathological scar formation or
benign or malignant tumors. Blood samples were collected one week
following the CS. Informed consent was obtained from each patient
prior to enrollment in the study. All specimens were obtained
following informed consent and procedures were conducted in
accordance with the Ethical Standards of the Declaration of
Helsinki (10). Written permission
was provided by the hospital and the local ethics committee of
Nanfang Hospital of Southern Medical University for the study.
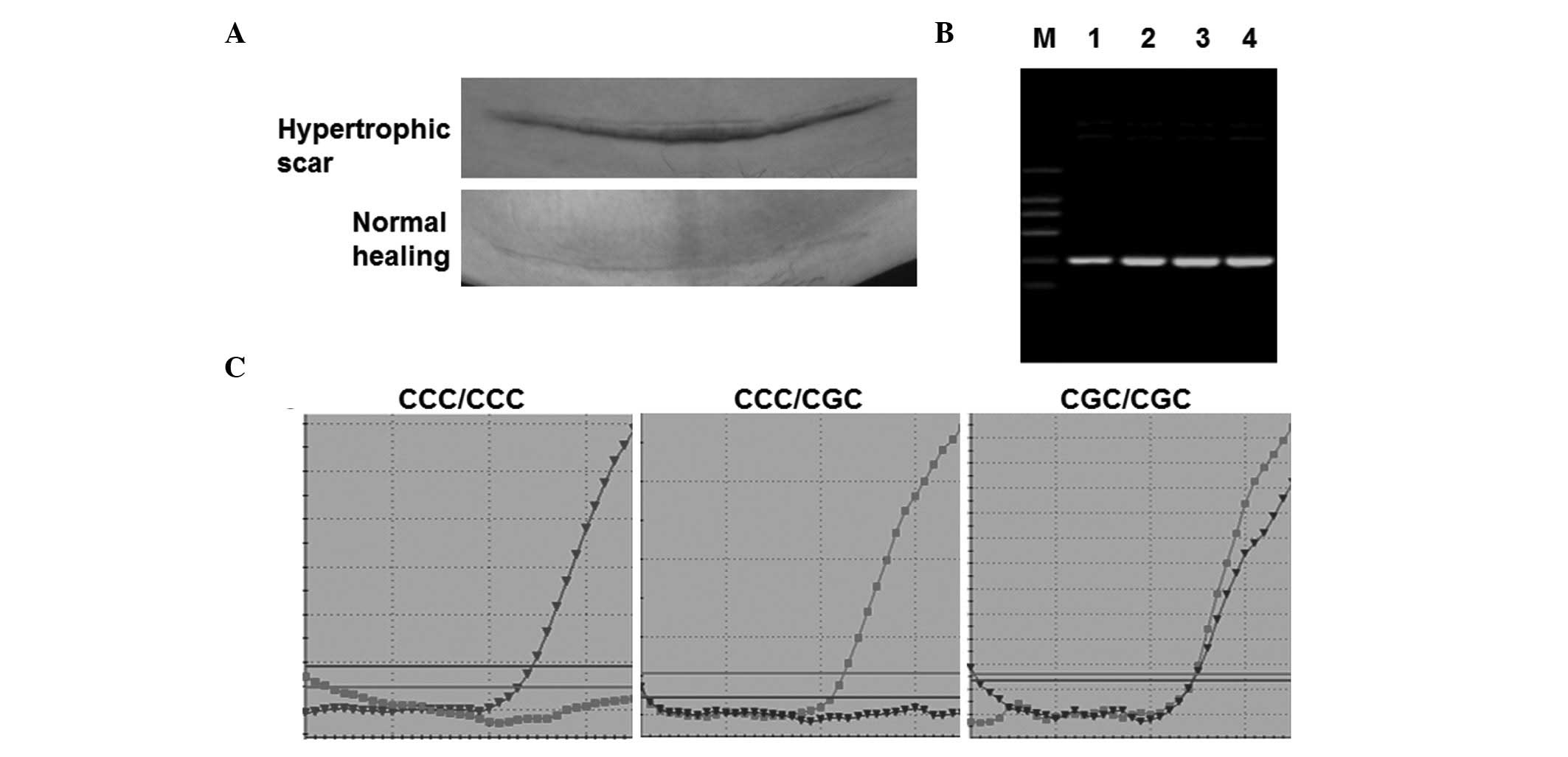
Normal healing, hypertrophic scars (Fig. 1A) and keloid scars were defined
according to the following clinical features. Keloid scars became
irritated two or three months following surgery and the scars were
recognized as round and smooth. Keloids usually extended beyond the
area of the original injury and showed no tendency to regress in
the 12 months. No keloid scars were identified in the patients.
Hypertrophic scars generally appeared within 4 weeks following
surgery. Initially hypertrophic scar lesions were often
erythematous and brownish-red but then became pale. However,
hypertrophic scars never extended beyond the area of the original
injury and regressed spontaneously when re-examined during the
12–18 month follow-up investigations. Normal healing scars
regressed spontaneously within 3 months.
Fluorescence quantitative polymerase
chain reaction (qPCR)
Genomic DNA was extracted from 2 ml blood samples
collected from the peripheral veins of each patient. A pair of
molecular beacon probes were designed and labeled with fluorescent
dyes to detect the single nucleotide polymorphism (SNP) of
p53 codon-72. The 5′ end of the CCC and CGC probes were
labeled with carboxyfluorescein (Fam) and hexachlorofluorescein
(Hex), respectively (Sangon Biotech Co., Ltd., Shanghai, China),
the 3′ end was conjugated with a quenching complex [Black Hole
Quencher-1 (BHQ-1); Biosearch Technologies, Inc., Novato, CA, USA].
The sequences of the probes were as follows: MBC,
5′-Fam-ATGCAGCCCCcCGTGGCCCCTGCAT-BHQ-1; and MBG,
5′-Hex-ATGCAGCCCCgCGTGGCCCCTGCAT -BHQ-1. Two primers, designed
using Primer 5 software (Premier Biosoft, Palo Alto, CA, USA) and
synthesized by Shanghai Genechem Co., Ltd. (Shanghai, China), were
used to amplify the p53 380-bp fragment in exon 4. The
primer sequences were as follows: Forward, TPF,
5′-GACCTGGTCCTCTGACTGCT-3′; and reverse, TPR,
5′-GATACGGCCAGGCATTGAAG-3′.
For PCR, 0.5 μl of each primer, 1 μl genomic DNA,
0.2 μl Fam-C primer and 0.2 μl Hex-G primer were mixed and then
diluted with DNase- and RNase-free water to provide a total
reaction volume of 20 μl. The reaction was performed with a
Stratagene Mx3000P qPCR system (Agilent Technologies, Santa Clara,
CA, USA). The amplified gene products were sent to Sangon Biotech
Co., Ltd. (Shanghai, China) for sequencing. PCR products were
detected by electrophoresis.
Nested PCR
Nested PCR was preformed to obtain the p53 380-bp
fragment in exon 4. Briefly, DNA extracted from peripheral blood
was amplified by the first set of primers. The product obtained was
then used as the template for a second amplification with primers
of TPF and TPR. Finally, the PCR products from the second
amplication were analyzed by agarose gel electrophoresis.
Follow-up investigations
Follow-up investigations were performed for a period
of 12–18 months. During this period, patients were required to
return to the hospital at any time when a fibroproliferative scar
appeared to grow beyond the confines of the original wound.
Patients were also required to report to the hospital if symptoms,
including skin erythema, pruritus or pain, developed at the site of
surgical incision.
Statistical analysis
The relationship between the p53 gene
polymorphisms and the occurrence of hypertrophic scars was
evaluated by a χ2 test and a Student’s t-test. Moreover,
the significant association between p53 gene polymorphisms
and hypertrophic scars was also evaluated by a corrected
χ2 test. SPSS 13.0 (SPSS, Inc., Chicago, IL, USA) was
used to carry out statistical analysis. P<0.05 was considered to
indicate a statistically significant difference.
Results
Clinical characteristics of the
patients
Peripheral venous blood samples were collected from
260 patients in the study. A total of 249 patients underwent the
lower abdominal CS (Fig. 1A) via a
transverse incision, while the remaining 11 patients underwent the
traditional longitudinal CS incision.
The products of nested PCR were detected following
an electrophoresis reaction. As shown in Fig. 1B, only a 380-bp DNA band was
detected in all peripheral blood samples. No non-specific
amplification was observed in those groups, indicating that PCR was
specific.
Fluorescence qPCR
To determine the gene p53 codon-72
polymorphisms, fluorescence qPCR was performed. As shown in
Fig. 1C, samples with only MBC
probe-positive signals were considered as CCC/CCC homozygous and
those with only MBG probe-positive signals were considered to be
CGC/CGC homozygous. Samples with MBC and MBG probe-positive signals
were considered to be CCC/CGC heterozygous. These results indicate
that the three possible genotypes were successfully detected by
fluorescence qPCR.
All patients received a 12–18 month follow-up
investigation. During this period, hypertrophic scars developed in
22 patients, with an incidence rate of 8.47%. As shown in Table I, among the patients with the
CCC/CCC genotype, nine patients had hypertrophic scars and 46
patients showed normal healing, which is a ratio of 0.19. However,
the follow-up investigations indicated that the presence of a
homozygous or heterozygous C-to-G alteration at the codon-72 site
in gene p53 resulted in 13 patients with hypertrophic scars
and 192 patients with normal healing, which is a ratio of 0.07
(Table I). Therefore, patients
with the CCC/CCC genotype had a higher risk of developing
hypertrophic scars compared with that of patients with CCC/CGC or
CGC/CGC genotypes. The differences between these two groups were
statistically significant (P<0.05). The positive predictive
value (PPV) and negative predictive value were 9/55 = 163.6/1,000
and 192/205 = 936.6/1,000, respectively. These results indicate
that patients with the CCC/CCC genotype had a higher risk of
developing hypertrophic scars compared with that of patients with
CCC/CGC or CGC/CGC genotypes.
 | Table IComparison between the C/C and C/G or
G/G genotypes. |
Table I
Comparison between the C/C and C/G or
G/G genotypes.
| p53
polymorphisms | | | |
|---|
|
| | | |
|---|
| Groups | CCC/CCC | CCC/CGC or
CGC/CGC | Sum | χ2 | P-value |
|---|
| Normal healing | 46 | 192 | 238 | | |
| Hypertrophic
scar | 9 | 13 | 22 | | |
| Total | 55 | 205 | 260 | 4.404 | 0.036a |
Discussion
Possible mechanisms underlying the pathogenesis of
hypertrophic scars include excessive inflammation, excessive
angiogenesis, abnormal growth factor levels and delayed apoptosis
of fibrotic myofibroblasts due to alterations to the p53
gene (11–14). In p53-null mice, it was found that
scar hypertrophy and cellular density significantly increased due
to the downregulation of cellular apoptosis (15). p53 mutations in hypertrophic scars
and keloid fibroblasts were detected from cultured cells (16). The p53 codon-72 (Arg 72
Pro), which is associated with various types of cancers, is the
most extensively studied SNP in the p53 gene. Specific
studies have reported that the p53 codon-72 CGT/CCT SNP is
also associated with abnormal scar susceptibility; however, this
remains controversial. Wang et al (7) analyzed codon 72 of p53 in 54
patients with keloids and 30 patients with hypertrophic scars using
restriction fragment length polymorphism. The study observed that
the frequencies of the Pro- and Arg-encoding alleles in the
hypertrophic scar patients deviated significantly from those in the
normal controls. However, there was no significant difference
between the keloid patients and healthy individuals. Zhuo et
al (17) analyzed 45 patients
with keloids by PCR-reverse dot blotting and identified that
patients with the Pro/Pro genotype had a higher risk of forming a
keloid scar than patients with Pro/Arg and Arg/Arg genotypes.
However, Yan et al (9)
hypothesized that there were no significant differences in the
distribution of the p53 codon-72 polymorphism between keloid
patients and healthy controls, but that the Arg/Arg genotype may
affect the formation of keloids in the shoulder and back.
A study has shown that the p53 codon-72
CCC/CCC genotype may result in keloid susceptibility in the
Guangdong district (17).
Furthermore, it has been reported that the Pro 72 and Arg 72
variants differ in functional activities (18). In addition, p53 codon-72
polymorphisms may change gene expression, resulting in changes to
p53 function, including activating gene transcription, inducing
apoptosis and inhibiting cell transformation (7). The Pro 72 variant was shown to
activate transcription effectively and upregulate the downstream
genes. However, the Arg 72 variant was not only a suppressor of
cellular transformation, but also induced apoptosis more
effectively than the Pro 72 variant.
In the present study, the PPV of abnormal scars
following CS in patients with the p53 codon-72 CCC/CCC
genotype was 163.6/1,000. Patients with the p53 codon-72
CCC/CCC genotype were the susceptible population, whose relative
risk of hypertrophic scars was 2.8896-fold higher than that of the
other individuals. The results of the present study indicate that
patients with the CCC/CCC genotype had a higher risk of developing
hypertrophic scars compared with that of patients with CCC/CGC or
CGC/CGC genotypes.
Acknowledgements
The study was supported by a grant from the National
Natural Science Foundation of China (no. 81071589).
References
|
1
|
Ceelen W, Pattyn P and Mareel M: Surgery,
wound healing, and metastasis: Recent insights and clinical
implications. Crit Rev Oncol Hematol. 89:16–26. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Greaves NS, Ashcroft KJ, Baguneid M and
Bayat A: Current understanding of molecular and cellular mechanisms
in fibroplasia and angiogenesis during acute wound healing. J
Dermatol Sci. 72:206–217. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Kazemi M, Salehi Z and Chakosari RJ: TP53
codon 72 polymorphism and breast cancer in northern Iran. Oncol
Res. 18:25–30. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Kara N, Karakus N, Ulusoy AN, Ozaslan C,
Gungor B and Bagci H: P53 codon 72 and HER2 codon 655
polymorphisms in Turkish breast cancer patients. DNA Cell Biol.
29:387–392. 2010. View Article : Google Scholar
|
|
5
|
Saed GM, Ladin D, Olson J, Han X, Hou Z
and Fivenson D: Analysis of p53 gene mutations in keloids using
polymerase chain reaction-based single-strand conformational
polymorphism and DNA sequencing. Arch Dermatol. 134:963–967.
1998.PubMed/NCBI
|
|
6
|
Liu YB, Gao JH, Duan HJ and Liu XJ:
Investigation of p53 gene mutations in keloids using PCR-SSCP.
Zhonghua Zheng Xing Wai Ke Za Zhi. 19:258–260. 2003.(In
Chinese).
|
|
7
|
Wang CM, Hiko H and Nakazawa N:
Investigation of p53 polymorphism for genetic predisposition of
keloid and hypertrophic scar. Zhonghua Zheng Xing Wai Ke Za Zhi.
21:32–35. 2005.(In Chinese).
|
|
8
|
Zhuo Y, Gao JH, Luo SQ, Zeng WS, Hu ZQ, Lu
F and Zhao YZ: p53 gene codon 72 polymorphism and susceptibility to
keloid. Zhonghua Zheng Xing Wai Ke Za Zhi. 21:201–203. 2005.(In
Chinese).
|
|
9
|
Yan L, Lü XY, Wang CM, Cao R, Yin YH, Jia
CS and Zhuang Q: Association between p53 gene codon 72 polymorphism
and keloid in Chinese population. Zhonghua Zheng Xing Wai Ke Za
Zhi. 23:428–430. 2007.(In Chinese).
|
|
10
|
WMA Declaration of Helsinki - Ethical
Principles for Medical Research Involving Human Subjects. 59th WMA
General Assembly; Seoul, Republic of Korea. October 2008; Available
at http://www.wma.net/en/30publications/10policies/b3/uri.
Accessed February 7, 2014
|
|
11
|
Penn JW, Grobbelaar AO and Rolfe KJ: The
role of the TGF-β family in wound healing, burns and scarring: a
review. Int J Burns Trauma. 2:18–28. 2012.
|
|
12
|
van den Broek LJ, Kroeze KL, Waaijman T,
et al: Differential response of human adipose tissue-derived
mesenchymal stem cells, dermal fibroblasts and keratinocytes to
burn wound exudates: Potential role of skin-specific chemokine
CCL27. Tissue Eng Part A. 20:197–209. 2014.
|
|
13
|
Albrecht-Schgoer K, Schgoer W, Theurl M,
et al: Topical secretoneurin gene therapy accelerates diabetic
wound healing by interaction between heparan-sulfate proteoglycans
and basic FGF. Angiogenesis. 17:27–36. 2014. View Article : Google Scholar
|
|
14
|
Kong P, Xie X, Li F, Liu Y and Lu Y:
Placenta mesenchymal stem cell accelerates wound healing by
enhancing angiogenesis in diabetic Goto-Kakizaki (GK) rats. Biochem
Biophys Res Commun. 438:410–419. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Aarabi S, Bhatt KA, Shi Y, et al:
Mechanical load initiates hypertrophic scar formation through
decreased cellular apoptosis. FASEB J. 21:3250–3261. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
De Felice B, Garbi C, Santoriello M,
Santillo A and Wilson RR: Differential apoptosis markers in human
keloids and hypertrophic scars fibroblasts. Mol Cell Biochem.
327:191–201. 2009.PubMed/NCBI
|
|
17
|
Zhuo Y, Gao JH, Luo SQ, et al: Evaluation
of high-risk individuals for keloid and polymorphism of p53 gene
codon 72. Chin J Clin Rehabil. 9(22): 130–131. 2005.(In
chinese).
|
|
18
|
Kay C, Jeyendran RS and Coulam CB: p53
tumour suppressor gene polymorphism is associated with recurrent
implantation failure. Reprod Biomed Online. 13:492–496. 2006.
View Article : Google Scholar : PubMed/NCBI
|