|
1
|
Sarvananthan N, Surendran M, Roberts EO,
et al: The prevalence of nystagmus: the Leicestershire nystagmus
survey. Invest Ophthalmol Vis Sci. 50:5201–5206. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Cabot A, Rozet JM, Gerber S, et al: A gene
for X-linked idiopathic congenital nystagmus (NYS1) maps to
chromosome Xp11.4–p11.3. Am J Hum Genet. 64:1141–1146.
1999.PubMed/NCBI
|
|
3
|
Schiaffino MV, Bassi MT, Galli L, et al:
Analysis of the OA1 gene reveals mutations in only one-third of
patients with X-linked ocular albinism. Hum Mol Genet. 4:2319–2325.
1995. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Bassi MT, Schiaffino MV, Renieri A, et al:
Cloning of the gene for ocular albinism type 1 from the distal
short arm of the X chromosome. Nat Genet. 10:13–19. 1995.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Tarpey P, Thomas S, Sarvananthan N, et al:
Mutations in FRMD7, a newly identified member of the FERM family,
cause X-linked idiopathic congenital nystagmus. Nat Genet.
38:1242–1244. 2006. View
Article : Google Scholar : PubMed/NCBI
|
|
6
|
Schorderet DF, Tiab L, Gaillard MC, et al:
Novel mutations in FRMD7 in X-linked congenital nystagmus. Mutation
in brief #963. Online. Hum Mutat. 28:5252007.PubMed/NCBI
|
|
7
|
Zhang B, Liu Z, Zhao G, et al: Novel
mutations of the FRMD7 gene in X-linked congenital motor nystagmus.
Mol Vis. 13:1674–1679. 2007.PubMed/NCBI
|
|
8
|
Shiels A, Bennett TM, Prince JB and
Tychsen L: X-linked idiopathic infantile nystagmus associated with
a missense mutation in FRMD7. Mol Vis. 13:2233–2241.
2007.PubMed/NCBI
|
|
9
|
Thomas S, Proudlock FA, Sarvananthan N, et
al: Phenotypical characteristics of idiopathic infantile nystagmus
with and without mutations in FRMD7. Brain. 131:1259–1267. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Betts-Henderson J, Bartesaghi S, Crosier
M, et al: The nystagmus-associated FRMD7 gene regulates neuronal
outgrowth and development. Hum Mol Genet. 19:342–351. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Wang ET, Sandberg R, Luo S, et al:
Alternative isoform regulation in human tissue transcriptomes.
Nature. 456:470–476. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
de la Grange P, Gratadou L, Delord M,
Dutertre M and Auboeuf D: Splicing factor and exon profiling across
human tissues. Nucleic Acids Res. 38:2825–2838. 2010.PubMed/NCBI
|
|
13
|
Li Q, Lee JA and Black DL: Neuronal
regulation of alternative pre-mRNA splicing. Nat Rev Neurosci.
8:819–831. 2007. View
Article : Google Scholar
|
|
14
|
Venables JP: Alternative splicing in the
testes. Curr Opin Genet Dev. 12:615–619. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
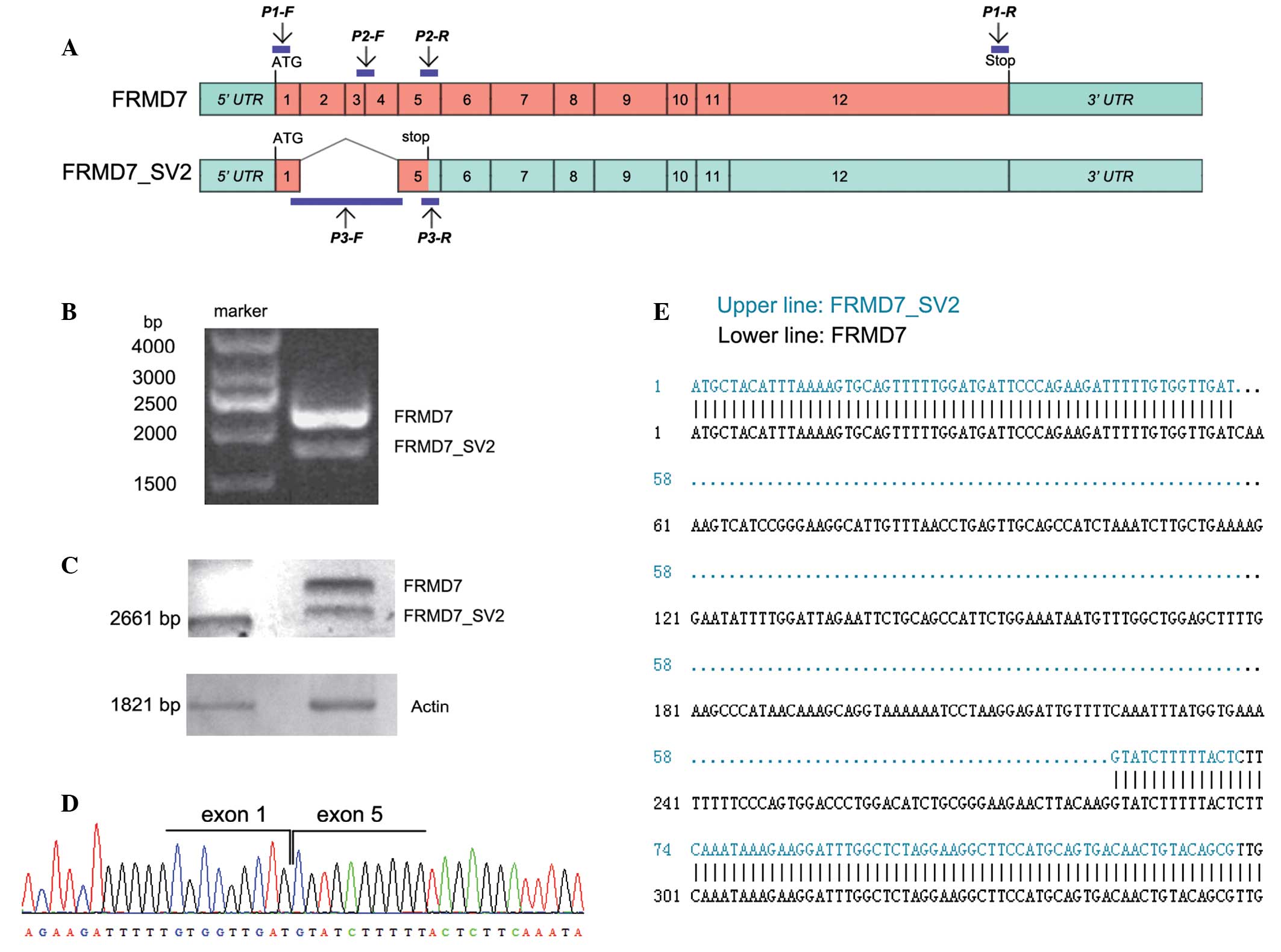
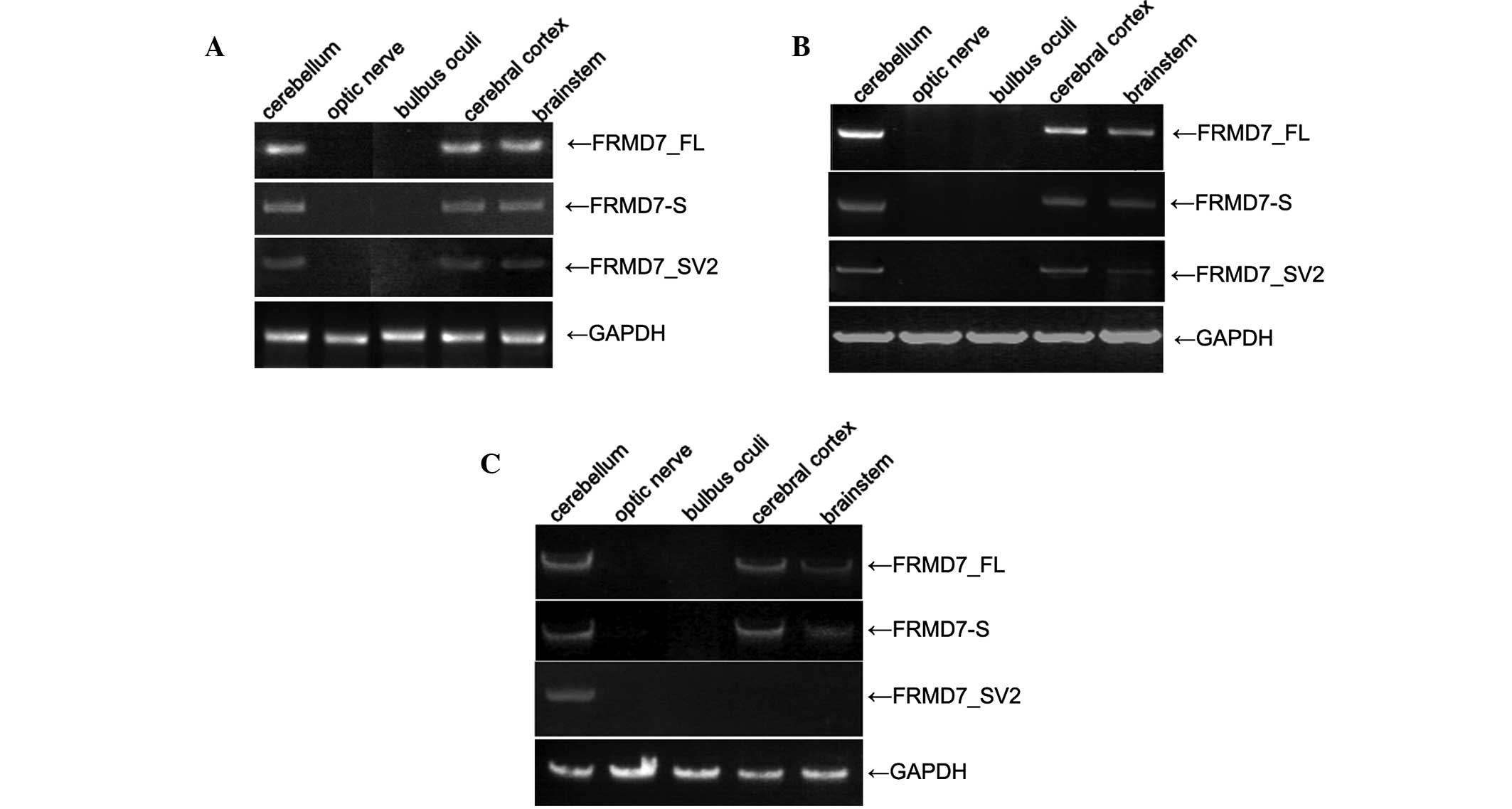
Li Y, Pu J, Liu Z, et al: Identification
of a novel FRMD7 splice variant and functional analysis of two
FRMD7 transcripts during human NT2 cell differentiation. Mol Vis.
17:2986–2996. 2011.PubMed/NCBI
|
|
16
|
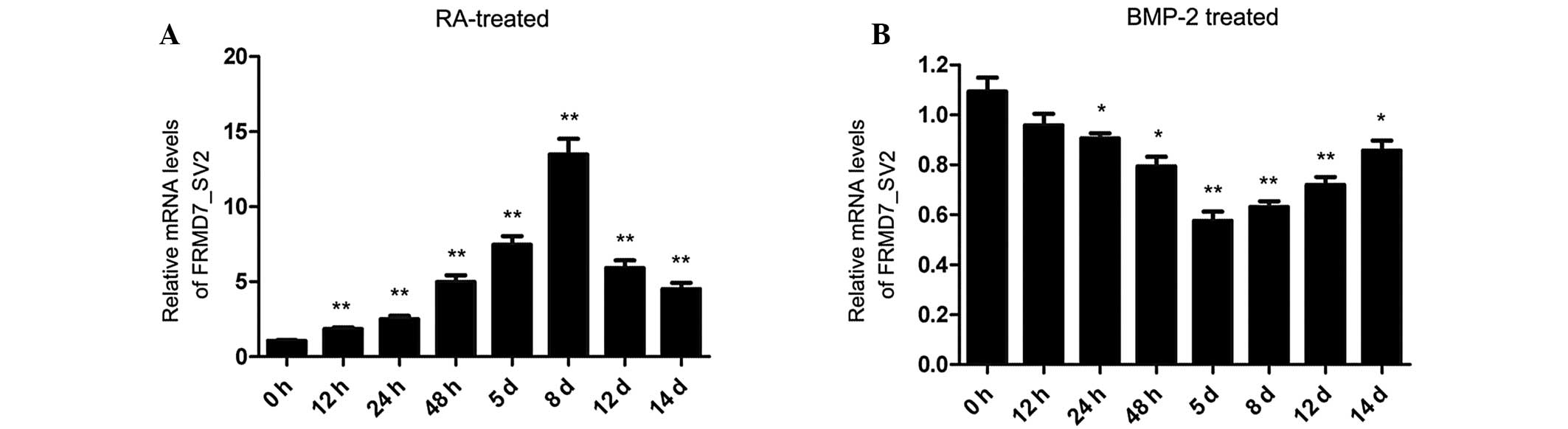
Andrews PW: Retinoic acid induces neuronal
differentiation of a cloned human embryonal carcinoma cell line in
vitro. Dev Biol. 103:285–293. 1984. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Pleasure SJ, Page C and Lee VM: Pure,
postmitotic, polarized human neurons derived from NTera 2 cells
provide a system for expressing exogenous proteins in terminally
differentiated neurons. J Neurosci. 12:1802–1815. 1992.
|
|
18
|
Chadalavada RS, Houldsworth J, Olshen AB,
et al: Transcriptional program of bone morphogenetic
protein-2-induced epithelial and smooth muscle differentiation of
pluripotent human embryonal carcinoma cells. Funct Integr Genomics.
5:59–69. 2005. View Article : Google Scholar
|
|
19
|
Caricasole A, Ward-van Oostwaard D,
Zeinstra L, et al: Bone morphogenetic proteins (BMPs) induce
epithelial differentiation of NT2D1 human embryonal carcinoma
cells. Int J Dev Biol. 44:443–450. 2000.PubMed/NCBI
|
|
20
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
|
|
21
|
Johnson JM, Castle J, Garrett-Engele P, et
al: Genome-wide survey of human alternative pre-mRNA splicing with
exon junction microarrays. Science. 302:2141–2144. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Dredge BK, Polydorides AD and Darnell RB:
The splice of life: alternative splicing and neurological disease.
Nat Rev Neurosci. 2:43–50. 2001. View
Article : Google Scholar : PubMed/NCBI
|
|
23
|
Stamm S, Ben-Ari S, Rafalska I, et al:
Function of alternative splicing. Gene. 344:1–20. 2005. View Article : Google Scholar
|
|
24
|
Yeo G, Holste D, Kreiman G and Burge CB:
Variation in alternative splicing across human tissues. Genome
Biol. 5:R742004. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Pu J, Li Y, Liu Z, et al: Expression and
localization of FRMD7 in human fetal brain, and a role for F-actin.
Mol Vis. 17:591–597. 2011.PubMed/NCBI
|
|
26
|
Martin JH, Cooper SE, Hacking A and Ghez
C: Differential effects of deep cerebellar nuclei inactivation on
reaching and adaptive control. J Neurophysiol. 83:1886–1899.
2000.PubMed/NCBI
|
|
27
|
Dieterich M and Brandt T: Functional brain
imaging of peripheral and central vestibular disorders. Brain.
131:2538–2552. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Harris CM, Walker J, Shawkat F, et al: Eye
movements in a familial vestibulocerebellar disorder.
Neuropediatrics. 24:117–122. 1993. View Article : Google Scholar : PubMed/NCBI
|