Introduction
Breast cancer is a common clinical condition with
increasing morbidity and high mortality rates (1). Previous studies have suggested that the
development of breast cancer is associated with inactivated tumor
suppressor genes, dysfunctional signaling transduction pathways and
other malfunctioning molecular signaling processes (2–4). With
the development of chemotherapy, hormonal therapy, immunotherapy,
gene therapy and other treatment technologies, the long-term
survival of breast cancer patients has become possible. Breast
cancer patients continue to succumb to the disease due to tumor
metastasis, drug resistance and other reasons including hemorrhage,
infection and recurrence (5).
Therefore, finding novel biomarkers for use in the early diagnosis
and treatment of breast cancer has become increasingly studied.
A high-throughput sequencing study revealed the
existence of types of non-coding RNA in gene transcripts, including
small non-coding RNA and long non-coding (lnc)RNA (6). LncRNA has a length of >200
nucleotides and a similar structure to mRNA, including a 5 cap and
poly-A tail. However, lncRNA is unable to encode proteins due to
the absence of an open reading frame (7). Increasing numbers of studies have
indicated that lncRNA may play an important role in various
cellular processes, including cell differentiation, proliferation
and apoptosis (8,9). Notably, a previous study revealed a
novel function of lncRNA in the development of prostate cancer
(10). In addition, long non-coding
homeobox antisense intergenic RNA is reported to be a highly
valuable clinical biomarker for the prognosis and early diagnosis
of ovarian cancer (11).
The primary functions of lncRNA include genomic
imprinting, chromatin remodeling, cell cycle regulation and
transcriptional regulation of adjacent mRNA (13). The γ-synuclein gene (SNCG), also
known as breast cancer-specific protein 1, encodes a synaptic
protein that was first identified in breast cancer. The expression
of SNCG has been closely associated with tumor invasion and
metastasis in a number of cancer types, including breast, stomach
and liver cancer, as well as the clinical staging of cancer and the
extent of lymph node metastasis (14–16).
Activation of SNCG expression, induced by demethylation, has been
observed to stimulate cell invasion and metastasis by activating
the mitogen-activated protein kinase signaling pathway and the
phosphorylation of the activator protein-1 transcription factor
(17). In addition, SNCG is able to
induce resistance to chemotherapy by inhibiting the c-Jun
N-terminal kinase signaling pathway.
The present study examined the expression levels of
lncRNA-AK058003 in breast cancer tissues and the effects on cell
proliferation, invasion and migration. In addition, the potential
of lncRNA-AK058003 as a transcriptional regulator of SNCG was
investigated, as well as the underlying mechanism of action.
Materials and methods
Tissue samples
Breast cancer and adjacent tissue samples were
collected from 30 patients (age range, 36–68 years; mean age, 42.5
years; median age, 45) who had received a clinical and pathological
diagnosis of breast cancer between December 2012 and December 2013.
The distance between the tumor tissue edge and the adjacent tissue
was >5 cm. Tissue samples were rinsed in saline and immediately
frozen with liquid nitrogen for storage at −80°C. The pathological
stage of the breast cancer tissues and the extent of lymph node
metastasis were diagnosed by two pathologists. The tissue samples
were divided into well-, moderately- and poorly-differentiated
groups according to the degree of differentiation, and were further
classified into lymph node metastasis (N1) or lymph node
non-metastasis (N0) groups depending on the extent of metastasis.
The study was approved by the Ethics Committee of Shandong
University (Jinan, China) and informed consent was obtained from
all the participants or their families.
Reagents
Total RNA extraction reagent TRIzol®, Dulbeccos
modified Eagles medium (DMEM) and Lipofectamine® 2000 were
purchased from Invitrogen Life Technologies (Carlsbad, CA, USA).
MCF-7 breast cancer cells were purchased from the Cell Bank of the
Chinese Academy of Sciences (Shanghai, China). Fetal bovine serum
was purchased from Gibco Life Technologies (Grand Island, NY, USA)
and Transwell® chambers were purchased from Corning, Inc.
(Tewksbury, MA, USA). A rabbit anti-human SNCG polyclonal antibody
was purchased from Abcam (Boston, MA, USA) and a Reverse
Transcription System kit was purchased from Chengdu Bo Ruike
Biotechnology Ltd. (Chengdu, China). The SYBR® Green quantitative
polymerase chain reaction (qPCR) reagent was obtained from Kapa
Biosystems, Inc. (Wilmington, MA, USA).
Cell cultures
MCF-7 cells were cultured in DMEM, supplemented with
10% fetal bovine serum, 100 U/ml penicillin and 100 µg/ml
streptomycin, in an incubator at 37°C with 5% CO2. One
day prior to transfection, log-phase MCF-7 cells were seeded into
24-well plates (2×105/well) and classified into control,
small interfering (si)RNA-AK058003 interference, scrambled siRNA
negative control (NC) and mock groups. Transfection was performed
on the following day when the cells had reached 70% confluence.
Lipofectamine® 2000 (1 µl) and 1.5 µl siRNA solution (20 pmol/µl;
RiboBio Co., Ltd, Guangzhou, China) were added separately to two
Eppendorf tubes containing 50 µl Opti-MEM® I (Thermo Fisher
Scientific, Waltham, MA, USA). After standing for 5 min, the two
solutions were combined and left to stand for a further 20 min at
room temperature. The mixture was subsequently transferred to
culture plates to incubate for 6 h. The medium was then replaced
with fresh, high-glucose DMEM, supplemented with 10% fetal bovine
serum, prior to continued incubation. After 48 and 72 h of
incubation, cells were collected for the determination of gene and
protein expression levels.
Bioinformatics
SNCG is strongly associated with breast cancer and
thus coding genes upstream and downstream of SNCG was searched to
check for the existence of lncRNA Information regarding the genomic
location of AK058003 (8.4 kb upstream of SNCG on the same
chromosome 10) was obtained by searching the UCSC database
(http://genome.ucsc.edu/). The details of coding
genes that were within 50 kb upstream and downstream of AK058003
were obtained from GenBank (http://www.ncbi.nlm.nih.gov/genbank/). No other genes
were analyzed in the present study.
qPCR
Total RNA of the breast cancer tissues was extracted
using TRIzol®, according to the manufacturer's instructions. A 1-µg
sample of the total RNA was used for the reverse transcription
reaction and 1-µl was used for qPCR. The relative expression levels
were analyzed using the 2−ΔΔCt method. All experiments
were performed in triplicate and the averages were calculated, with
GAPDH used as the internal reference. The primers used were as
follows: AK058003 forward, 5-CAGATGGCTGAGGTGGAAGG-3 and reverse,
5-GACAAGGTCTCGCTCTTTTGCT-3; SNCG forward, 5-CACCCTCTGGTCCTTCTG-3
and reverse, 5-AGGAGTGGGCTCAAGTTT-3′; GAPDH forward,
5′-AGGTGAAGGTCGGAGTCAAC-3′ and reverse, 5′-CGCTCCTGGAAGATGGTGAT-3′.
The conditions for PCR were as follows: 95°C for 1 min, 95°C for 15
sec, 60°C for 30 sec, 72°C for 15 sec, for a total of 40 cycles,
followed by 72°C for 5 min.
Western blot analysis
Following liposome transfection for 72 h, cells from
the control, interference, mock and NC groups were washed twice
with cold phosphate-buffered saline, and radioimmunoprecipitation
assay lysis buffer was added for total protein extraction. Sodium
dodecyl sulfate-polyacrylamide gel electrophoresis was performed to
separate the proteins, which were subsequently transferred onto a
polyvinylidene difluoride membrane. The membrane was incubated with
appropriate concentrations of primary antibodies (SNCG, 1:1,000;
GAPDH, 1:2,000; Bioworld Technology Inc., St. Louis Park, MN, USA)
overnight at 4°C. After washing the membrane with
phosphate-buffered saline with Tween-20 three times for 15 min, the
membrane was incubated with horseradish peroxidase-conjugated
secondary antibodies (goat anti-mouse, 1:3,000, #BS11502; goat
anti-rabbit, 1:1,000, #BS12478; all from Bioworld Technology Inc.)
for 1 h at room temperature. Following three washes with
phosphate-buffered saline with Tween-20 for 15 min, the membrane
was visualized using an enhanced chemiluminescence detection kit
(Sigma-Aldrich, St. Louis, MO, USA) for imaging. Image
Lab™ software (Bio-Rad, Hercules, CA, USA) was used to
acquire and analyze the imaging signals. The relative content of
the SNCG protein was expressed as the SNCG/GAPDH ratio.
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT)
assay
Cells were seeded into 96-well plates at a density
of 3,000 cells/well. Next, 20 µl MTT (5 mg/ml; Sigma-Aldrich) was
added to each well prior to incubation for 4 h at 37°C, which was
followed by the addition of 150 µl dimethyl sulfoxide. After 4 h of
incubation, the absorbance of each well was measured at 490 nm
(Multiskan™ GO reader; Thermo Fisher Scientific). A cell
proliferation curve was plotted using optical density values.
Transwell® chamber assay
Matrigel (BD Biosciences, Franklin Lakes, NJ, USA)
was thawed at −4°C overnight and diluted with serum-free RPMI-1640
medium (dilution, 1:2). A 50-µl sample of this mixture was
deposited evenly into a 24-well chamber (8 µm), which was incubated
for 60 min at 37°C. Upon solidifying, 2×105 MCF-7 cells
from each group were seeded into the upper chamber containing 300
µl serum-free RPMI-1640 medium. In addition, 500 µl RPMI-1640
medium supplemented with 10% fetal bovine serum was added to the
lower chamber. The upper chamber was removed after 24 h and the
cells in the upper chamber were discarded. The membrane was fixed
with 4% formaldehyde for 10 min and stained using the Giemsa method
for microscopic observation of five random fields (magnification,
×200; Olympus IX83; Olympus Corporation, Tokyo, Japan), from which
the number of cells that had migrated was calculated. All
procedures were conducted on ice using pipetting tips cooled to
4°C.
Statistical analysis
Data were analyzed using SPSS 16.0 software (SPSS,
Inc., Chicago, IL, USA). Measurement data are expressed as the mean
± standard deviation. The t-test was used for comparisons between
groups, where P<0.05 was considered to indicate a statistically
significant difference.
Results
LncRNA-AK058003 and SNCG mRNA
expression levels are enhanced in breast cancer tissue
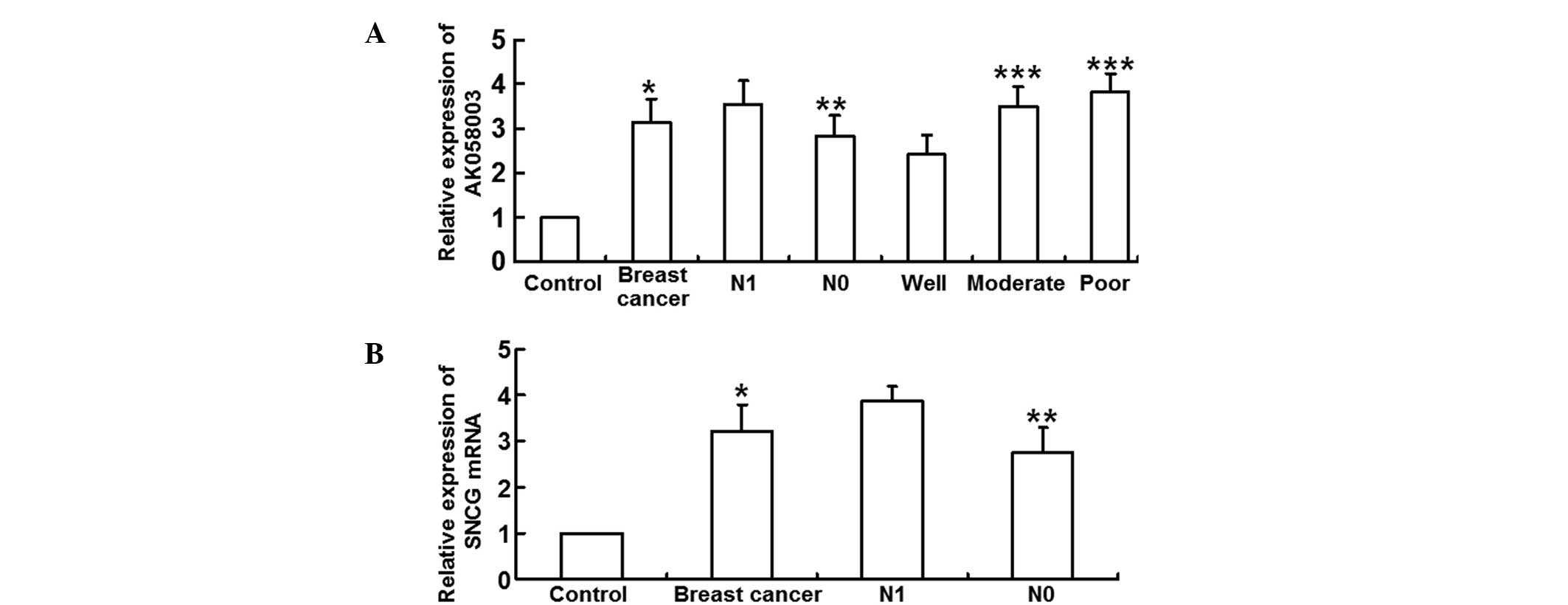
qPCR was employed to measure the expression levels
of lncRNA-AK058003 and SNCG mRNA. The results revealed that
relative lncRNA-AK058003 expression levels were significantly
higher in the breast cancer tissues (3.14±1.05 times) when compared
with the adjacent tissues (P<0.05). In addition, lncRNA-AK058003
expression levels in the N1 group (3.55±1.04) were significantly
higher when compared with the N0 group (2.82±0.96; P<0.05).
LncRNA-AK058003 expression in the well-differentiated group
(2.42±0.89) was significantly lower than that in the moderate
(3.51±0.86) and poor differentiation groups (3.82±0.82; P<0.05),
with the moderate differentiation group exhibiting no statistically
significant difference when compared with the poor differentiation
group (P>0.05; Fig. 1A). In
addition, the relative SNCG mRNA expression levels increased
significantly in the breast cancer tissues (3.22±0.56 times) when
compared with the adjacent tissues (P<0.05). Relative SNCG mRNA
expression levels were higher in the N1 group (3.87±0.37 times)
when compared with the N0 group (P>0.05; Fig. 1B). These observations indicated that
lncRNA-AK058003 and SNCG mRNA expression levels were elevated in
breast cancer tissues.
Expression of lncRNA-AK058003 is
inhibited by AK058003 siRNA
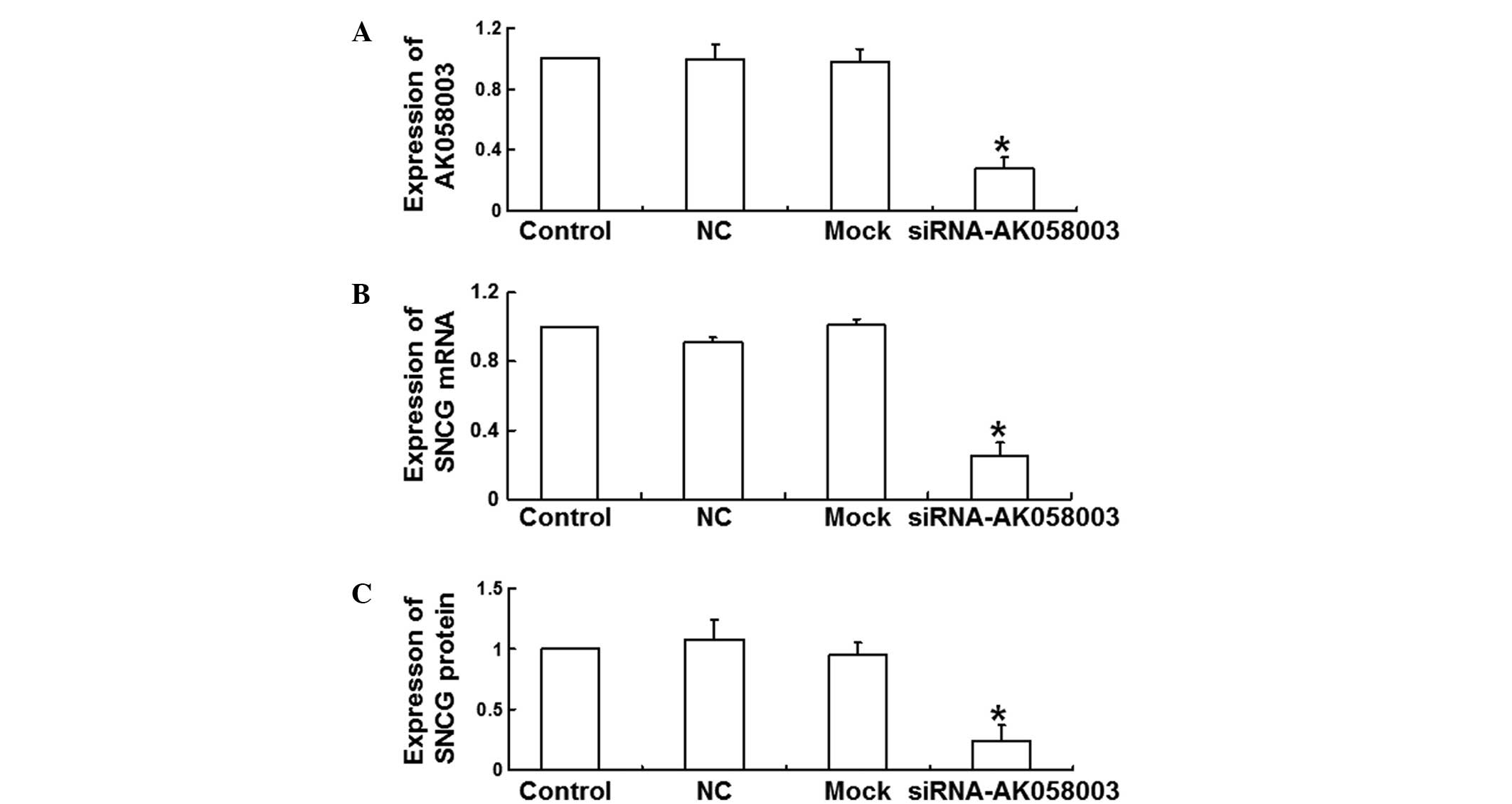
LncRNA-AK058003 expression levels were determined 48
h after siRNA transfection in MCF-7 cells in order to analyze the
effect of AK058003 siRNA on the expression of lncRNA-AK058003. The
results revealed that siRNA transfection significantly reduced the
expression levels of lncRNA-AK058003 when compared with the NC and
mock groups (P<0.05; Fig. 2A).
These data demonstrated that the expression of lncRNA-AK058003 was
inhibited by AK058003 siRNA.
SNCG mRNA and protein expression
levels are reduced by AK058003 siRNA
qPCR and western blot analysis were performed to
investigate the effects of AK058003 siRNA on the mRNA and protein
expression levels of SNCG. The results of qPCR revealed that the
relative SNCG mRNA expression levels decreased significantly at 48
h after transfection with AK058003 siRNA (0.25±0.08 times of
control; P<0.05; Fig. 2B).
Consistent with this result, relative SNCG protein expression
levels also decreased significantly (0.24±0.13 times of control;
P<0.05; Fig. 2C), indicating that
SNCG mRNA and protein expression levels were reduced by AK058003
siRNA.
MCF-7 cell proliferation is inhibited
by AK058003 siRNA
An MTT assay was performed to assess the effects of
AK058003 siRNA on MCF-7 cell proliferation. Cell proliferation in
the interference group was significantly reduced when compared with
the control, NC and mock groups at 24, 48 and 72 h following the
AK058003 siRNA treatment (P<0.05; Fig. 3), indicating that MCF-7 cell
proliferation was inhibited by AK058003 siRNA.
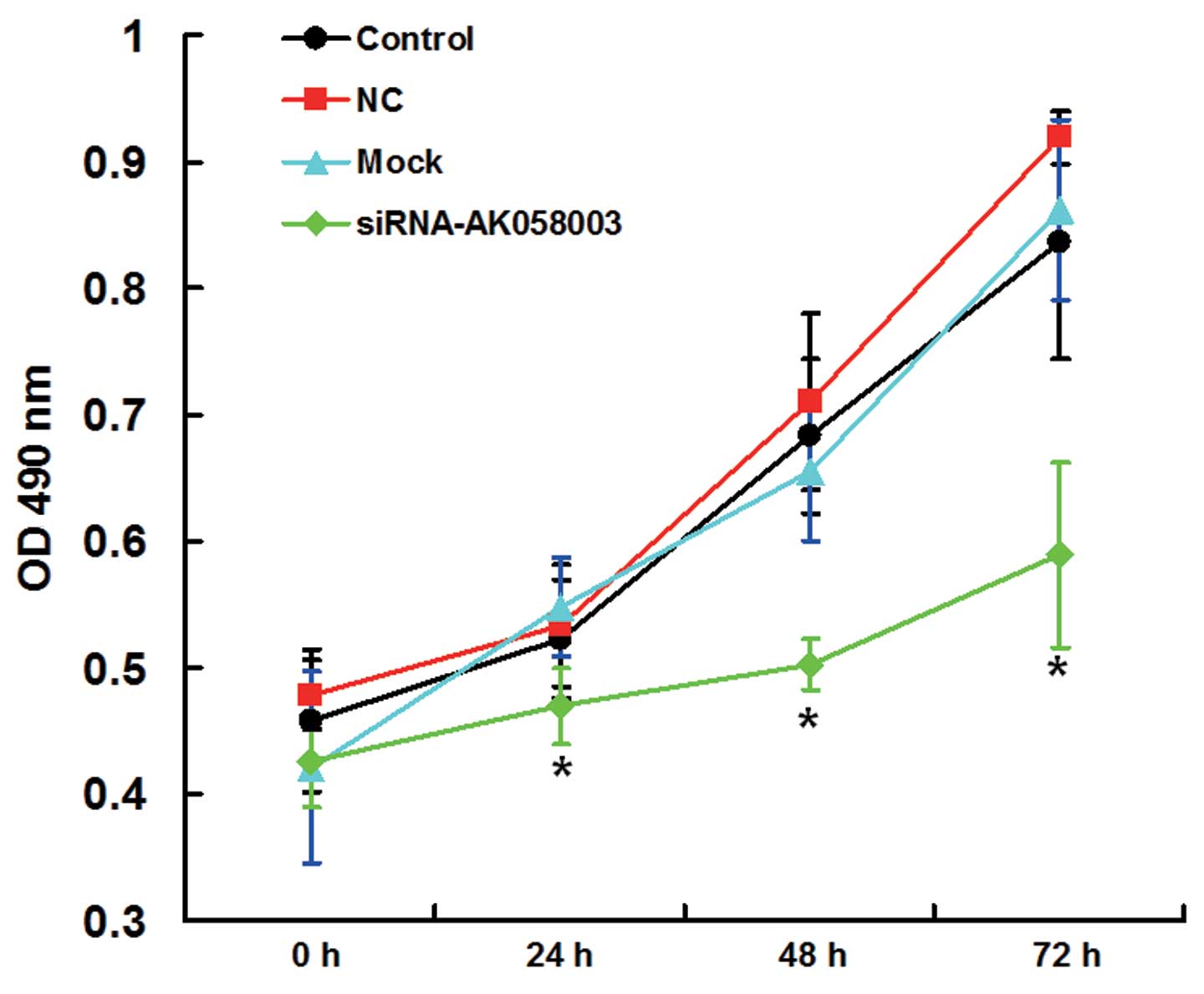 | Figure 3.Proliferation curves of MCF-7 cells in
the control, mock, NC and siRNA-AK058003 groups, as determined
using a 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium
bromide assay and measuring the OD at 490 nm. Data are expressed as
the mean ± standard deviation. *P<0.05, vs. control, NC or mock
groups. NC, negative control group; OD, optical density; siRNA,
small interfering RNA. |
MCF-7 cell migration and invasion are
reduced by AK058003 siRNA
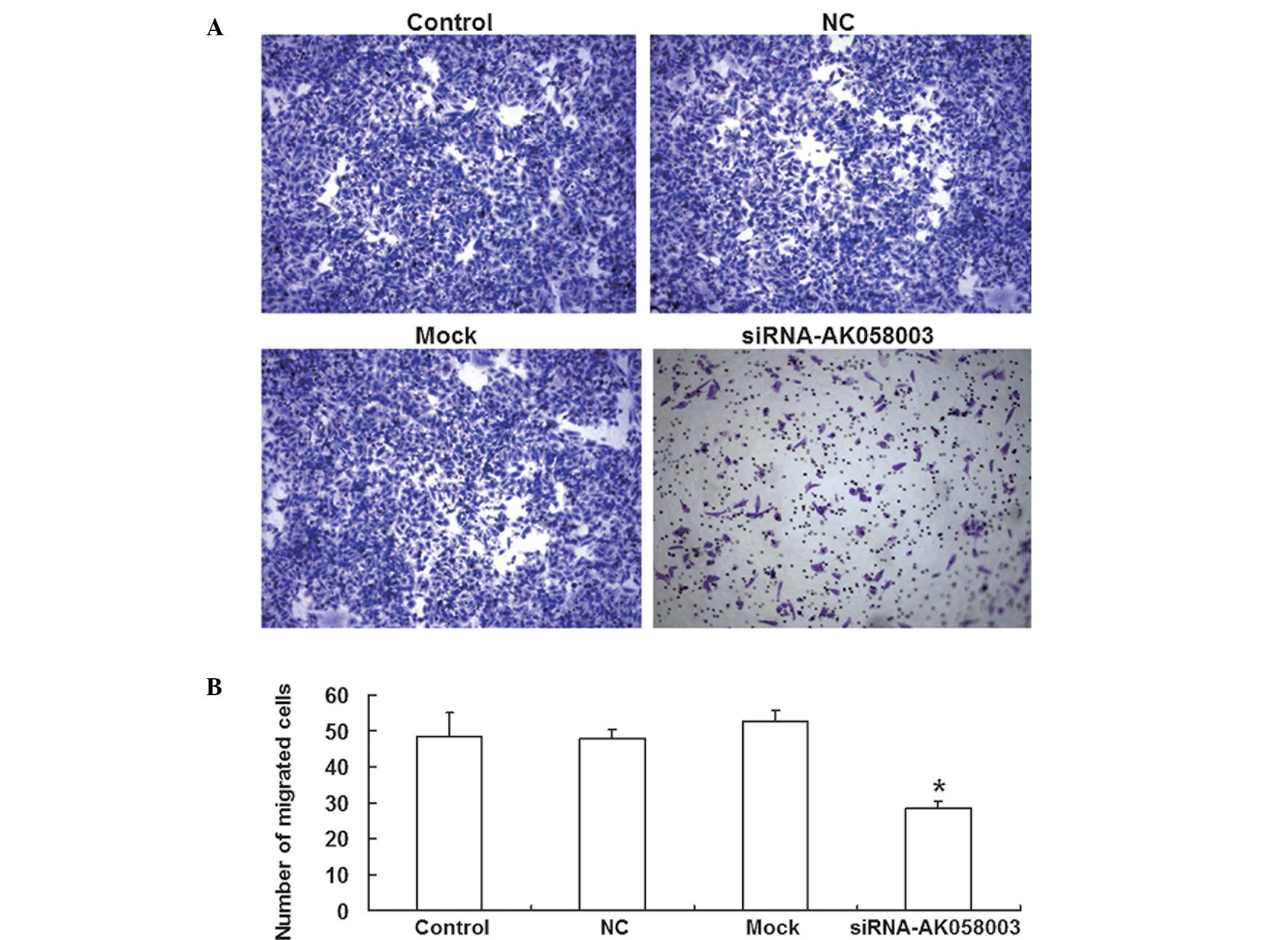
A Transwell® chamber assay was employed to assess
the effects of AK058003 siRNA on MCF-7 cell migration. The number
of MCF-7 cells that passed through the membrane was significantly
reduced in the AK058003 siRNA group (28.6±1.75; P<0.05) when
compared with the control (48.4±6.49), NC (47.8±2.65) and mock
groups (52.4±3.16; Fig. 4). These
results indicated that MCF-7 cell migration and invasion were
reduced by AK058003 siRNA.
Discussion
Mammalian genomes encode hundreds of lncRNAs with
numerous mechanisms of action. Notably, lncRNAs play an important
role in dosage compensation, epigenetics, the cell cycle and
differentiation (12). The primary
functions of lncRNA include genomic imprinting, chromatin
remodeling, mRNA transcriptional regulation and cell cycle
regulation (13). The present study
determined the expression levels of lncRNA-AK058003 in breast
cancer tissue and the potential correlation with the expression of
SNCG, which regulates MCF-7 breast cancer cells. LncRNA-AK058003
expression levels were found to be significantly increased in the
breast cancer tissue, and particularly in cases that had lymph node
metastasis. The well-differentiated group exhibited significantly
lower lncRNA-AK058003 expression levels when compared with the
moderate and poor differentiation groups. Thus, the level of
lncRNA-AK058003 expression was demonstrated to be closely
associated with the extent of lymph node metastasis and cell
differentiation. In addition, SNCG expression levels were
significantly increased in the breast cancer tissue and correlated
positively with lymph node metastasis. Downregulation of
lncRNA-AK058003 using siRNA in breast cancer MCF-7 cells
significantly reduced the level of SNCG expression. Furthermore,
MTT and Transwell® assays demonstrated that the proliferation,
invasion and migration abilities of the MCF-7 cells were
significantly reduced by the downregulation of lncRNA-AK058003
expression. These findings demonstrated that lncRNA-AK058003
regulated SNCG expression, possibly by affecting the SNCG
methylation levels. However, the exact mechanism underlying this
effect requires further study.
In summary, lncRNA-AK058003 expression levels were
demonstrated to be significantly increased in breast cancer
tissues. Furthermore, lncRNA-AK058003 expression was shown to
promote breast cancer proliferation, invasion and metastasis by
regulating SNCG expression, which may prove useful for the early
diagnosis and prognosis of breast cancer patients.
Acknowledgements
The authors thank Dr Yongzheng Min from the
Department of Urology at the General Hospital of Jinan Military
Region (Jinan, China) for valuable suggestions and instructions
during this study.
References
|
1
|
Depla AL, Scharloo-Karels CH, de Jong MA,
et al: Treatment and prognostic factors of radiation-associated
angiosarcoma (RAAS) after primary breast cancer: a systematic
review. Eur J Cancer. 50:1779–1788. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Harbeck N, Beckmann MW, Rody A, et al:
HER2 dimerization inhibitor pertuzumab - mode of action and
clinical data in breast cancer. Breast Care (Basel). 8:49–55. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Koul HK, Pal M and Koul S: Role of p38 MAP
kinase signal transduction in solid tumors. Genes Cancer.
4:342–359. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Plaza-Menacho I, Mologni L and McDonald
NQ: Mechanisms of RET signaling in cancer: current and future
implications for targeted therapy. Cell Signal. 26:1743–1752. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Lianos GD, Vlachos K, Zoras O, Katsios C,
Cho WC and Roukos DH: Potential of antibody-drug conjugates and
novel therapeutics in breast cancer management. Onco Targets Ther.
7:491–500. 2014.PubMed/NCBI
|
|
6
|
Peschansky VJ and Wahlestedt C: Non-coding
RNAs as direct and indirect modulators of epigenetic regulation.
Epigenetics. 9:3–12. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Kornfeld JW and Brüning JC: Regulation of
metabolism by long, non-coding RNAs. Front Genet. 5:572014.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Negishi M, Wongpalee SP, Sarkar S, et al:
A new lncRNA, APTR, associates with and represses the CDKN1A/p21
promoter by recruiting polycomb proteins. PLoS One. 9:e952162014.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Wang P, Xue Y, Han Y, et al: The
STAT3-binding long noncoding RNA lnc-DC controls human dendritic
cell differentiation. Science. 344:310–313. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Ye ZQ, Wang T and Song W: Long noncoding
RNAs in prostate cancer. Zhonghua Nan Ke Xue. 20:963–968. 2014.[(In
Chinese)]. PubMed/NCBI
|
|
11
|
Huang L, Liao LM, Liu AW, Wu JB, Cheng XL,
Lin JX and Zheng M: Overexpression of long noncoding RNA HOTAIR
predicts a poor prognosis in patients with cervical cancer. Arch
Gynecol Obstet. 290:717–723. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Rossi MN and Antonangeli F: LncRNAs: New
players in apoptosis control. Int J Cell Biol. 2014:4738572014.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Hrdlickova B, de Almeida RC, Borek Z and
Withoff S: Genetic variation in the non-coding genome: Involvement
of micro-RNAs and long non-coding RNAs in disease. Biochim Biophys
Acta. 1842:1910–1922. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Singh VK and Jia Z: Targeting synuclein-γ
to counteract drug resistance in cancer. Expert Opin Ther Targets.
12:59–68. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Chen J, Huang S, Wu KJ, et al: The
correlation of synuclein-γ and matrix metalloproteinase 9 in breast
cancer. Zhonghua Wai Ke Za Zhi. 51:641–644. 2013.[(In Chinese)].
PubMed/NCBI
|
|
16
|
Luo JH, Zhou J and Gao Y: Correlation
between periostin and SNCG and esophageal cancer invasion,
infiltration and apoptosis. Asian Pac J Trop Med. 6:516–519. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Lu A, Zhang F, Gupta A and Liu J: Blockade
of AP1 transactivation abrogates the abnormal expression of breast
cancer-specific gene 1 in breast cancer cells. J Biol Chem.
277:31364–31372. 2002. View Article : Google Scholar : PubMed/NCBI
|