Introduction
Pelvic inflammatory disease (PID) is a common
disorder induced by ascending infection of the upper female genital
tract by, for example, Neisseria gonorrhoeae, Chlamydia
trachomatis, genital mycoplasmas, gram-negative bacteria or
gram-positive bacteria (1–4), the symptoms of which include
endometritis, salpingitis and peritonitis (5). This disease is considered to be a great
threaten to women as it can lead to a poor outcome with severe
sequelae, such as tubal factor infertility, ectopic pregnancy and
chronic pelvic pain. The diagnosis of PID is dependent upon
evaluation of the patient's history, physical examination,
laboratory studies and imaging (5).
However, the sensitivity and specificity of clinical diagnosis have
been reported to be only 87 and 50%, respectively (6). Although laparoscopy is the gold
standard for diagnosis, its routine use is limited by poor
compliance (7). Therefore, a novel
and non-invasive method is necessary for clinical diagnosis of
PID.
Metabolomics, a key technology for the evaluation of
system biology, focuses on the comprehensive qualification and
quantification of all endogenous metabolites of low molecular mass
in vivo, and thus reveals the pathological and physiological
changes of the whole biosystem from the aspect of metabolism
(8,9). Metabolomics has been carried out to
identify disease-related metabolites as biomarkers that contribute
to the non-invasive diagnosis of diseases. A previous study has
found that 3-hydroxybutyrate, N-acetyl-glycine,
3-hydroxy-2-methyl-butanoic acid and nonanedioic acid in the plasma
can serve as biomarkers for the diagnosis of breast cancer
(10). The levels of sarcosine in
urine can contribute to the diagnosis of prostate cancer (11), and desmosterol in plasma has been
identified as a new specific biomarker of Alzheimer's disease
(12). However, to the best of our
knowledge, no metabolomic study has yet been carried out to search
for novel metabolite biomarkers of PID.
Gas chromatography coupled with mass spectrometry
(GC/MS) is one of the core analytical methods for metabolomic study
and has been used as a platform in non-targeted metabolomic
analysis (13,14). Through GC-MS analysis, the chemical
structure of metabolites can be inferred by comparing the mass
spectrometric fragmentation pattern with mass spectral libraries,
which facilitates the identification of biomarkers. Silylation is a
common method of derivatization, the application of which in GC-MS
analysis can enable the detection of more metabolites, including
carbohydrates, amino acids and nucleosides (15).
Following renal concentration, urine amplifies the
circulating levels of metabolites and thus exhibits metabolomic
alterations more clearly. In addition, urine collection is
non-invasive. Hence, urine is of interest in metabolomic research.
In the majority of medical research, animal experiments are usually
carried out first to estimate the importance of conducting clinical
experiments and offer valuable information. Therefore, on the basis
of silylation and GC-MS analysis, the present urine non-targeted
metabolomic study was carried out in a rat model of PID to
illustrate the metabolic alterations associated with PID, and
identify the potential biomarkers of PID accordingly.
Materials and methods
Reagents and materials
Ultrapure water was produced using a Milli-Q Plus
Water Purification System (EMD Millipore, Milford, MA, USA).
Pentobarbital was purchased from Xiya Reagent Co., Ltd. (Chengdu,
China). Progesterone injection was obtained from Zhejiang Xianju
Pharmaceutical Co., Ltd. (Taizhou, China).
N,O-Bis(trimethylsilyl)-trifluoro-acetamide-trimethylchlorosilane
(BSTFA-TMCS) (99:1, v/v), urease, margaric acid, tropic acid,
methoxylamine hydrochloride, pyridine and metabolite standards were
purchased from Sigma-Aldrich (St. Louis, MO, USA). Absorbable
gelatin sponge was purchased from Jinling Pharmaceutical Co., Ltd.
(Nanjing, China).
PID model construction and sample
collection
The experimental procedures were approved by the
Animal Care and Use Committee of Central South University
(Changsha, China). The harm to animals was minimized with the
greatest efforts. Sixteen female specific pathogen-free Sprague
Dawley (SD) rats, aged 9 weeks and weighing 210–230 g, were
purchased from Silaikejingda Experimental Animal Co., Ltd.
(Changsha, China). They were randomly divided into two groups,
namely the control group and the PID group. Standard laboratory
chow was available ad libitum through the whole experiment.
All the rats were acclimated for a week and then injected
subcutaneously with 10 mg progesterone 7 days prior to
experimentation. PID modeling was conducted with reference to a
previous method (16,17). Ureaplasma urealyticum
(t-strain mycoplasma) and a pathogenic Escherichia coli
strain (No. 0573) were obtained from Clinical Laboratory Department
of the Maternal and Child Health Hospital of Hunan Province
(Changsha, China). An absorbable gelatin sponge, with a volume of
0.125 ml, was immerged into a microbe-mixing solution with a U.
urealyticum concentration of 1×108 ccu/ml and E.
coli concentration of 1×108 cfu/ml. Then, the cervix of
each rat in the PID group was implanted with the microbe-containing
gelatin sponge, and the rat was positioned upside down for 3 min.
The cervixes of control group rats were implanted with gelatin
sponges immerged in saline. This infection procedure was conducted
once every 2 days and repeated four times. Surface disinfection was
carried out with 70% alcohol following each infection. On the 21st
day from the first infection, a 24 h urine sample of each rat was
collected and stored at −80°C. Then, vaginal swab samples from rats
were obtained to detect U. urealyticum, and E. coli
using a mycoplasma detection kit and TDR-300B automatic microbial
analysis system (Tiandiren Biotech Co., Ltd., Changsha, China),
respectively. After that, rats were anesthetized with pentobarbital
(dose, 30 mg/kg). Plasma and the right uterus and fallopian tube
were collected and stored at −80°C, and the left uterus and
fallopian tube were used for histological examination. Then, all
the rats were sacrificed via cervical dislocation.
Histological examination
The left uterus and fallopian tube of each rat were
fixed in neutral-buffered formalin (10%), embedded in paraffin, cut
into 2-µm sections, and stained with hematoxylin and eosin
(H&E). Three different fields for each tissue sample were
examined using light microscopy (DM500l; Leica Microsystems GmbH,
Wetzlar, Germany) at a magnification of ×100 by a blinded observer.
The inflammation of the uterus and fallopian tube of each animal
was semi-scored by the observer's evaluation of the degree of
inflammatory cell infiltration (graded from 0 to 3) for each of the
three fields.
C-reactive protein (CRP), interleukin
(IL)-1β and IL-6 determinations
The concentration of CRP in plasma was measured
using enzyme-linked immunosorbent assay (ELISA) kits (RayBiotech,
Norcross, GA, USA). The right upper genital tract (including uterus
and fallopian tube) of each rat was weighed, followed by the
addition of physiological saline at the ratio of 5:1 (v/w) and
homogenization. The total protein of the homogenate was measured
using a bicinchoninic acid assay kit (Beyotime Institute of
Biotechnology, Shanghai, China). The concentrations of IL-1β and
IL-6 in the homogenate were determined using the ELISA kits and
presented in units of µg/g protein of homogenate. These procedures
were performed according to the protocols of the manufacturers.
Urine sample preparation
Thawed urine samples were diluted to give a
creatinine concentration of 2.5 mmol/l as determined using a JEOL
JCA-BM1650 clinical biochemistry analyzer (JEOL Ltd., Tokyo,
Japan). A 30-unit quantity of urease was added to 100 µl urine and
the mixture was incubated for 30 min at 37°C. Then, 50 µl of a
mixed solution of margaric acid and tropic acid, each of
concentration 0.5 mg/ml, was added as an internal standard. An
aliquot of 800 µl ethanol was mixed with the sample, and then the
mixture was centrifuged at 12,000 × g for 15 min. The supernatant
was transferred to a new tube and stored at −20°C for 10 min. After
drying the supernatant in a vacuum dryer at room temperature, 100
µl methoxylamine hydrochloride (20 mg/ml in pyridine) was added
with mixing, and the resultant mixture was incubated at 37°C for 2
h. Then, 100 µl BSTFA-TMCS was added and the mixture was heated to
70°C for 1 h. A quality control (QC) sample was prepared by mixing
urine from each rat and handling according to the above
protocol.
GC-MS analysis
GC-MS analysis was undertaken on an Agilent 7890 gas
chromatography system equipped with an Agilent 5975C mass analyzer
(Agilent Technologies, Inc., Palo Alto, CA, USA). Separation was
conducted on an Agilent DB-5MS capillary column (30 m × 0.25 mm
internal diameter × 0.25 µm film thickness). Each 1 µl aliquot of
the derivatized solution or reference standard was injected in the
splitless mode and helium was used as the carrier gas with flow
rate of 1.5 ml/min. The temperatures of the inlet, transfer line
and ion source were maintained at 250, 300 and 230°C, respectively.
The GC temperature programming was set to 4 min isothermal heating
at 60°C, followed by the first ramp at 6°C/min to 150°C and holding
for 8 min, second ramp at 8°C/min to 280°C and holding for 10 min,
and third ramp at 12°C/min to 320°C. Data acquisition was achieved
using MS in the electron impact mode at 70 eV and in the full-scan
monitoring mode with a mass to charge (m/z) ratio from 35 to 750.
The chromatogram acquisition and detection of mass spectral peaks
were performed using G1701EA GC/MD ChemStation software (Agilent
Technologies, Inc.).
Data analysis and potential biomarker
identification
The peak areas of metabolites in each sample were
normalized to the peak area of creatinine. Internal standards were
used to calibrate the retention time of all metabolites and monitor
sampling and instrumental condition. Then, a data set of all
samples, consisting of the retention time and the normalized peak
area of metabolites, was generated and imported into SIMCA-P
software (version 11.5; Umetrics; MKS Instruments, Umeå, Sweden)
for multivariate analysis. The data were processed by unit variance
scaling and were mean-centered, followed by multivariate analysis
including principal component analysis (PCA) and partial least
squares discriminant analysis (PLS-DA) between the PID group and
control group. The unsupervised statistical analysis PCA was used
to describe associations and patterns among a set of variables; R2X
and Q2 are two measures of PCA model quality. Metabolites relevant
for group discrimination were selected by the values of variable
importance in the projections (VIP) >1 constructed from PLS-DA
analysis. The Student's t-test was employed for the metabolites
with VIP>1, and the differentiating metabolites were selected
when P<0.05 with the Benjamini and Hochberg procedure to control
the false discovery rate (FDR) (18). The mass fragmentation patterns of
these potential biomarkers were compared with the NIST/EPA/NIH Mass
Spectral Library 2011 (version NIST11; Gaithersburg, MD, USA) to
predict their chemical structures. The further identification of
these differentiating metabolites was performed by comparing their
mass spectra and chromatographic retention times to those of
standards.
Statistical analysis
Data are presented as mean ± standard deviation
(SD). Statistical analysis was performed by unpaired Student's
t-test using SPSS for Windows 16.0 (SPSS, Inc., Chicago, IL, USA).
P<0.05 was considered to indicate a statistically significant
result.
Results
PID model construction
The vaginal swab samples of rats from the PID group
tested positive for both U. urealyticum and E. coli, whereas the
control rats tested negative for the two pathogens. These results
indicated that infection in the PID group rats was successfully
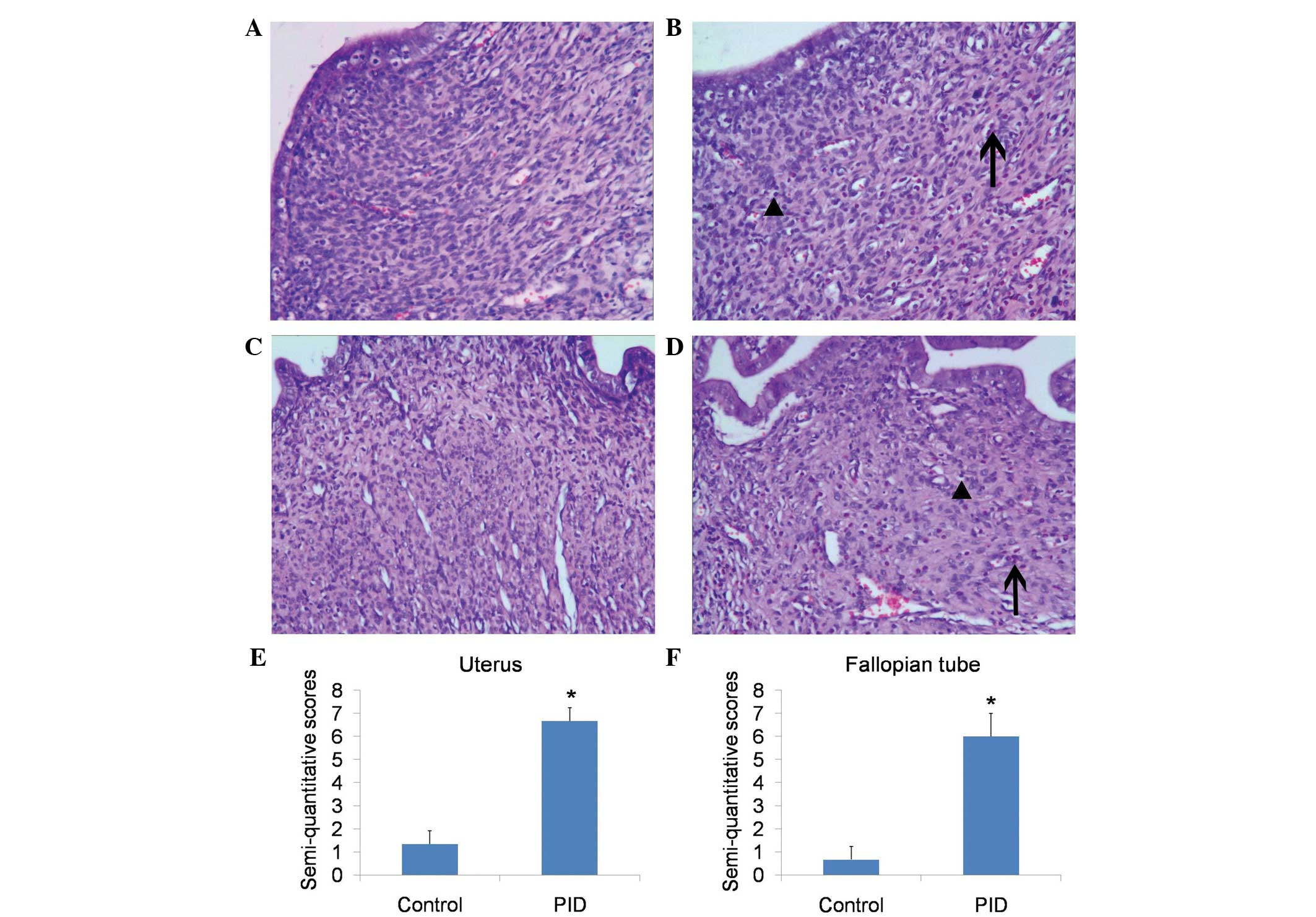
achieved. Histological examination revealed that, in comparison
with control rats, upper genital tract infection with these
pathogens caused significant inflammation, of the uterus and the
fallopian tubes (Fig. 1A–D),
including mass neutrophil and lymphocyte infiltration.
Semi-quantitative scoring of the inflammation of the uterus
(Fig. 1E) and fallopian tubes
(Fig. 1F) showed a significant
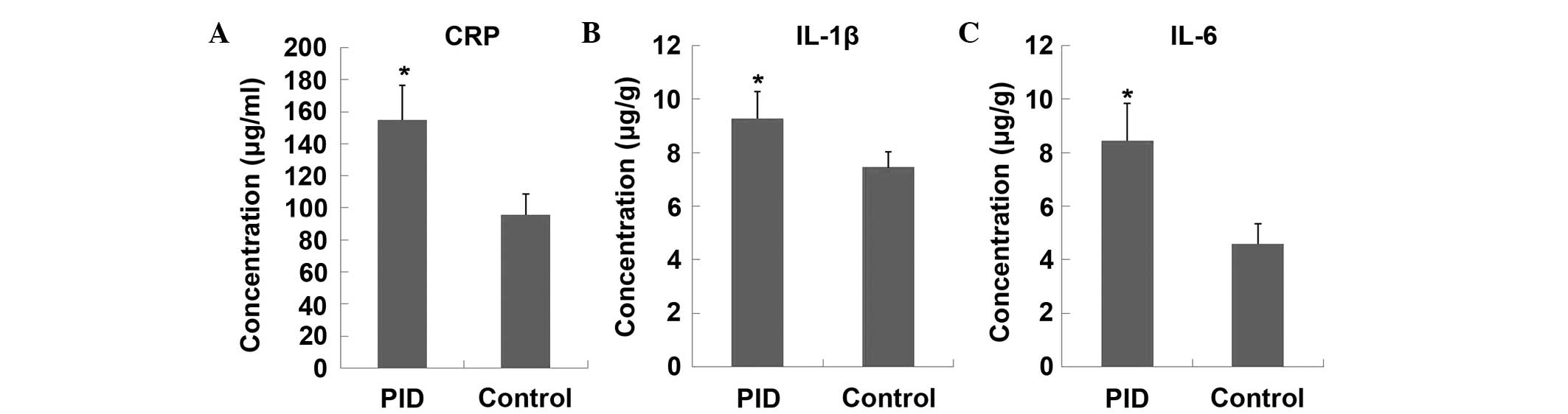
difference between the PID and control groups. The CRP levels in
plasma, and expression levels of IL-1β and IL-6 in the upper
genital tract (including the uterus and fallopian tube) were
significantly higher in PID model rats, as compared with the
control animals (Fig. 2), which
indicated that an inflammatory response occurred in upper genital
tract.
Assessment of the repeatability and
stability of the analytical method
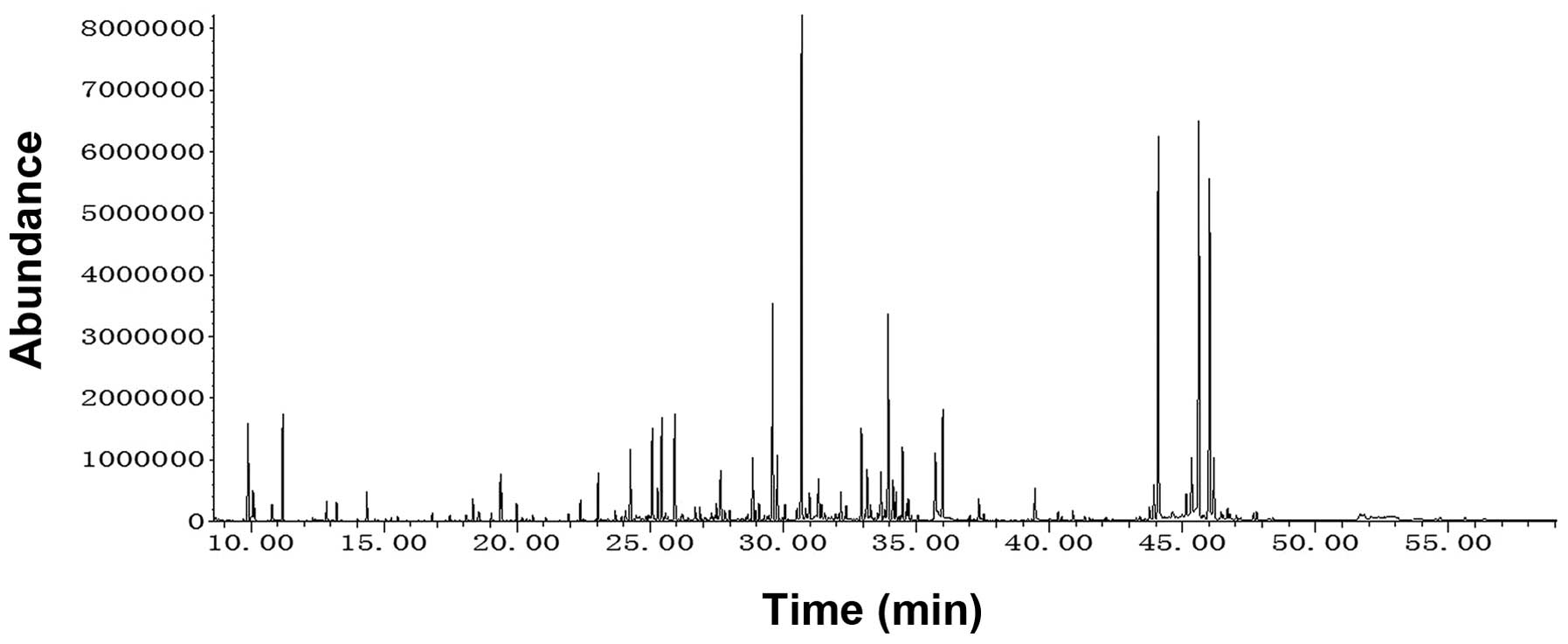
A representative chromatogram for the GC-MS coupled
with derivatization method is shown in Fig. 3. Repeatability and stability of the
analytical method are essential for obtaining valid metabolomic
data. In the present study, a stability test was performed by
analyzing 6 injections of a QC sample, and the relative standard
deviations (RSDs) of eight common ions were <0.27% for retention
times and <5.35% for peak areas. The repeatability was evaluated
by the analysis of 6 QC samples, with the RSDs of peak area
<11.28%. These results validated the analytical method for
metabolite determination.
Multivariate statistical analysis
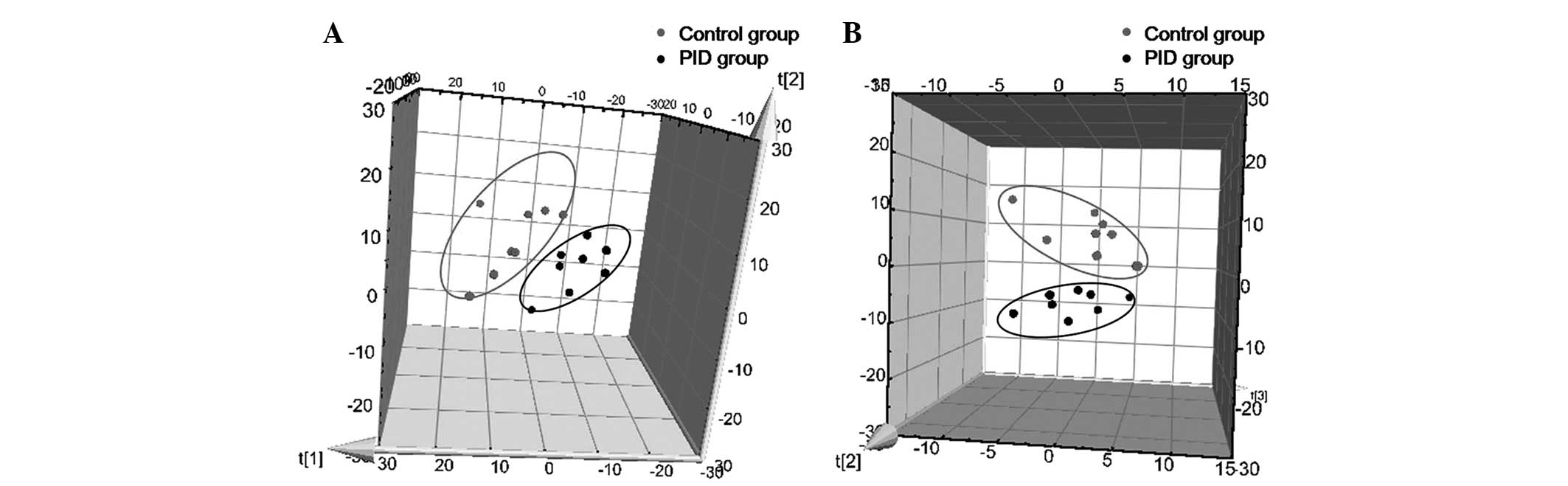
A three-component PCA model was selected to reveal
the general metabolic differences between groups, with R2X=0.751
and Q2=0.412. As presented in Fig.
4A, the PID group was clearly separated from the control group.
Subsequently, a PLS-DA was employed to further elucidate the
metabolic differentiation associated with PID. A clear separation
is shown in Fig. 4B, with three
predictive components and three orthogonal components (R2X=0.652,
R2Y=0.934, Q2=0.720). These results indicated that the metabolic
profile of rats was evidently disturbed by PID.
Potential biomarker
identification
Eighteen differentiating metabolites were selected
as potential biomarkers with VIP>1 and P<0.05, followed by
the identification of chemical structures using commercial
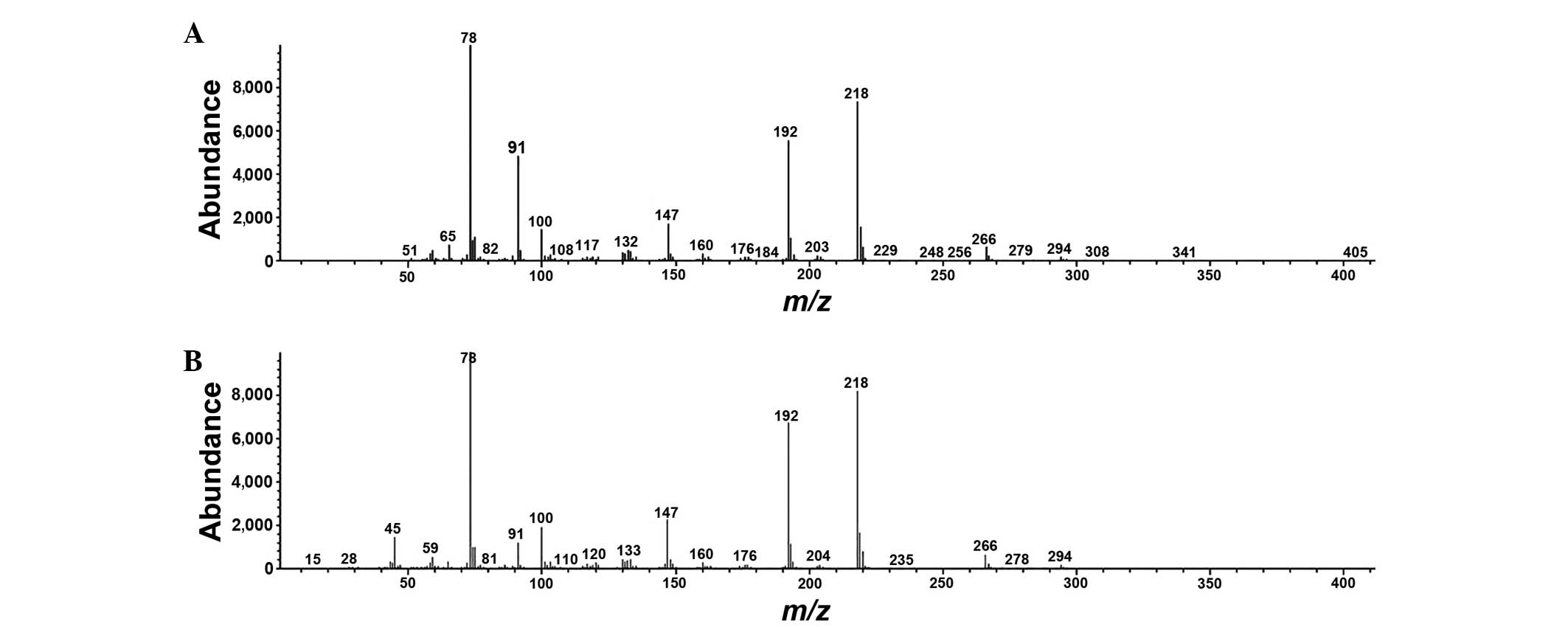
standards. Typical mass spectra of phenylalanine in urine and
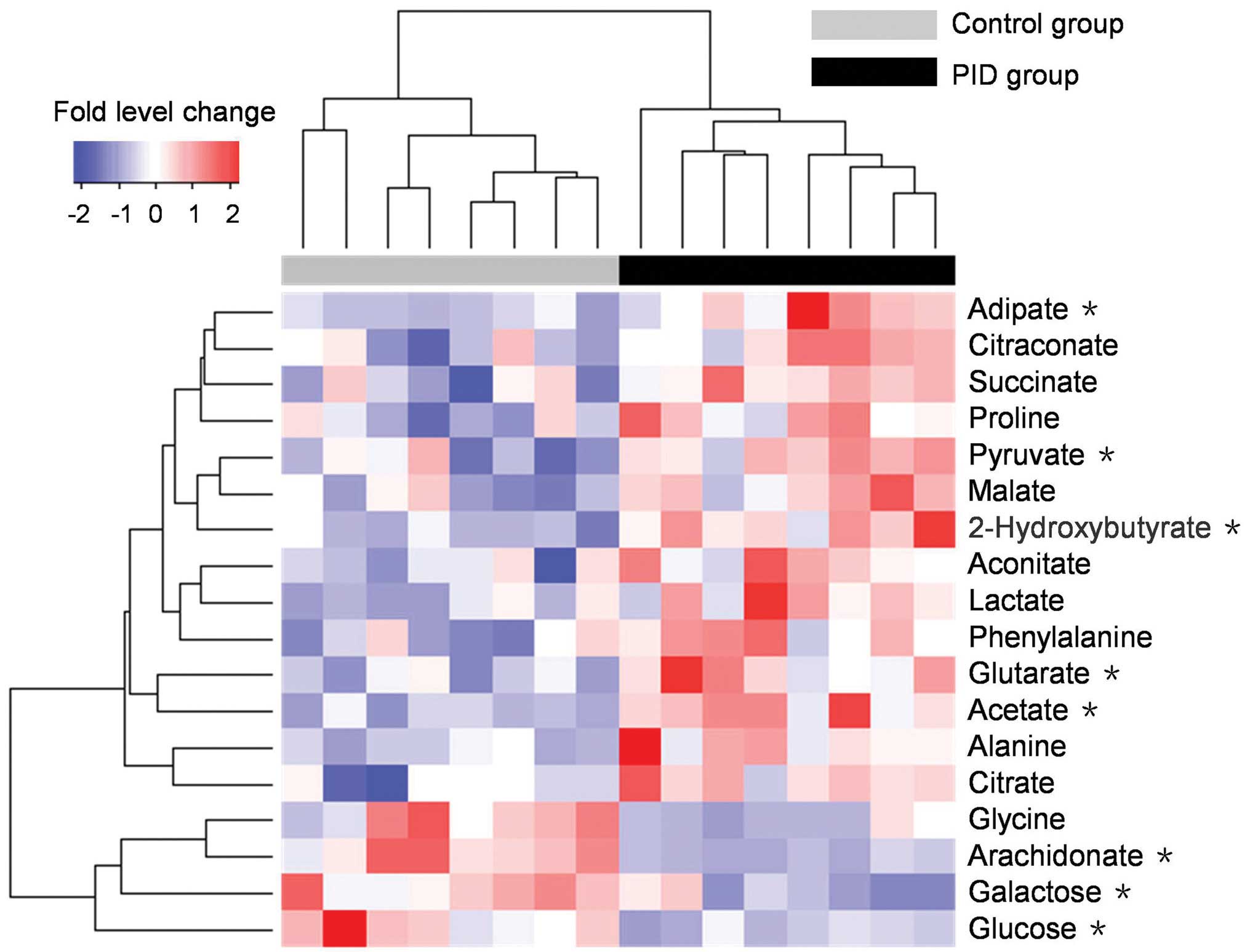
phenylalanine standard are presented in Fig. 5. A heat map of the potential
biomarkers is presented in Fig. 6,
which visualizes the fold level changes between the rats in the PID
and control groups. These differentiating metabolites are four
amino acids, three fatty acids, nine organic acids and two sugars.
The levels of 14 metabolites were notably increased in the urine of
the rats with PID compared with their levels in control rats, but
the levels of four metabolites, namely glycine, glucose, galactose
and arachidonate, were decreased.
Discussion
To the best of our knowledge, no previous study has
delineated the metabolomic changes that are associated with PID.
The aim of the present study was to examine the PID-associated
disturbance of metabolism in rats with the further aim of
identifying potential metabolite biomarkers for use in the
diagnosis of this disease.
Toll-like receptors (TLRs) play a key role in
evoking the inflammatory response to microbial infection in the
genital tract (19). U.
urealyticum (20) can be
recognized by TLR 2, and pathogenic E. coli (21) causes an inflammatory response through
TLR 4. Furthermore, both U. urealyticum and E. coli
are common pathogens of PID (22).
Therefore, to activate multiple TLRs and induce a quick and
enhanced inflammatory response in the genital tract, a solution
containing a mixture of U. urealyticum and E. coli
was used to construct a PID rat model, as in previous studies
(16,17). Following infection, mass neutrophil
and lymphocyte infiltrations indicated inflammation in the uterus
and fallopian tube (Fig. 1). The
elevation of CRP levels in plasma has been used to differentiate
inflammatory from non-inflammatory pelvic pathology in vivo
(23). CRP examination has been
included in the criteria for the diagnosis of PID proposed in the
USA (24). In the PID model in the
present study, the elevated CRP levels in plasma (Fig. 2A) illustrated the inflammatory
response. The pro-inflammatory cytokine IL-1β is a key factor in
the pathological development of PID (25), and the cytokine IL-6 is involved in
the processes of chronization and suppression of the synthesis of
IL-1β in the second phase of the immune response (26). Both IL-1β and IL-6 exhibit
significant elevation in patients with PID (27). Hence, as shown in Fig. 2B and C, the elevated expression of
IL-1β and IL-6 in the uterus and fallopian tube validated the local
inflammatory response in the rat PID model.
The biochemical reactions associated with the
differentiating metabolites can be determined using the Kyoto
Encyclopedia of Genes and Genomes (KEGG; http://www.genome.jp/kegg/) and the Human Metabolome
Database (HMDB, http://www.hmdb.ca/). The present
study illustrated that the comprehensive metabolic profile changes
associated with PID in rats are associated with sugar metabolism,
the citric acid cycle (TCA cycle), amino acid metabolism and fat
acid metabolism. The concurrence of reductions in the levels of
sugars (glucose and galactose) and an increase in pyruvate levels
indicated enhanced glycolysis in PID rats. In addition, the
upregulation of citrate suppresses citrate synthase activity,
followed by increased lactate production from pyruvate. The
elevated levels of intermediate products in the TCA cycle,
including citrate, aconitate, succinate and malate, suggest
possible defects in the mitochondrial respiratory system, and the
concentration changes of alanine, glycine, phenylalanine and
proline indicate alterations of amino acid metabolism. The increase
of glutarate and adipate in PID rats illustrates an abnormality of
fat acid metabolism that is also present in arthritis (28,29).
Leukotriene B4 (LTB4) is able to promote leucocyte recruitment
(30), and increased prostaglandin
E2 (PGE2) levels lead to edema and a hyperalgesic response
(31). The overproduction of both in
women with PID (32) may be involved
in modulation of the inflammatory response. As arachidonate is the
primary precursor of LTB4 and PGE2, high levels of production of
LTB4 and PGE2 will consume considerable arachidonate. Consistently,
a reduction in the levels of arachidonate was observed in the rats
with PID in the present study. The elevation of citraconate also
occurs in experimental arthritis (29), indicating a modulation of branched
pentadioic acid metabolism relating to an inflammatory response.
2-Hydroxybutyrate is a by-product of ophthalmate synthesis, a high
level of which indicates glutathione depletion (33). The increased levels 2-hydroxybutyrate
observed in rats with PID in this study may suggest a shortage of
glutathione and oxidative stress in the genital tract, which is
also observed in endometriosis (34). Acetic acid is a substance existing as
an antibacterial agent in the vaginal lubrication fluid of humans
(35). Notably, elevated pyruvate
levels may have promoted acetate production in the present study,
as pyruvate can be metabolized by pyruvate dehydrogenase to produce
acetate (36). This elevation of
acetate levels may be considered as a physiological response to
defend against the following infection.
In conclusion, a PID model was constructed by
multi-pathogenic infection of the upper genital tract in rats.
Infiltration of inflammatory cells and elevated expression levels
of cytokines in the uterus and fallopian tube validated the disease
model. The results reveal PID-associated metabolomic changes in the
rat PID model. Eighteen differentiating metabolites were found,
which are associated with sugar metabolism, the TCA cycle, amino
acid metabolism and fatty acid metabolism. These metabolites could
be potential biomarkers of PID. This study also offers a new
approach to evaluate the effect of anti-PID drugs in pre-clinical
or clinical trials.
Acknowledgements
The authors thank the Clinical Laboratory Department
and Department of Pathology in the Maternal and Child Health
Hospital of Hunan Province for experimental support. The present
study was supported by the National Natural Science Foundation of
China (grant no. 81501218), China Postdoctoral Science Foundation
(grant no. 2015M582330), Science and Technology Project of Hunan
Province (grant no. 2015RS4054) and Administration of Traditional
Chinese Medicine of Hunan Province (grant no. 201402).
Glossary
Abbreviations
Abbreviations:
|
GC/MS
|
gas chromatography-mass
spectrometry
|
|
PID
|
pelvic inflammatory disease
|
|
PCA
|
principal component analysis
|
|
PLS-DS
|
partial least squares-discriminant
analysis
|
|
TCA cycle
|
citric acid cycle
|
|
BSTFA-TMCS
|
N,O-bis(trimethylsilyl)-trifluoroacetamide-trimethylchlorosilane
|
|
CRP
|
C-reactive protein
|
|
IL
|
interleukin
|
|
QC
|
quality control
|
|
VIP
|
variable importance in the
projections
|
|
TLR
|
toll-like receptor
|
|
LTB4
|
leukotriene B4
|
|
PGE2
|
prostaglandin E2
|
References
|
1
|
World Health Organization: Global
incidence and prevalence of selected curable sexually transmitted
infections: 2008. Reprod Health Matters. 20:207–209. 2012.
View Article : Google Scholar
|
|
2
|
Quentin R and Verdon R: Microbiologic
basis of diagnosis and treatment of pelvic inflammatory disease. J
Gynecol Obstet Biol Reprod (Paris). 41:850–863. 2012.(In French).
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Saini S, Gupta N Aparna, Batra G and Arora
DR: Role of anaerobes in acute pelvic inflammatory disease. Indian
J Med Microbiol. 21:189–192. 2003.PubMed/NCBI
|
|
4
|
Zhang DZ, Wen JY, Zhou WC and Wu XY:
Pathogenic bacteria distribution and drug resistance isolated from
women with pelvic inflammatory disease. Zhonghua Yi Yuan Gan Ran
Xue Za Zhi. 19:1747–1750. 2009.(In Chinese).
|
|
5
|
Soper DE: Pelvic inflammatory disease.
Obstet Gynecol. 116:419–428. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Blenning CE, Muench J, Judkins DZ and
Roberts KT: Clinical inquiries. Which tests are most useful for
diagnosing PID? J Fam Pract. 56:216–220. 2007.PubMed/NCBI
|
|
7
|
Jaiyeoba O and Soper DE: A practical
approach to the diagnosis of pelvic inflammatory disease. Infect
Dis Obstet Gynecol. 2011:7530372011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Nicholson JK, Lindon JC and Holmes E:
‘Metabonomics’: Understanding the metabolic responses of living
systems to pathophysiological stimuli via multivariate statistical
analysis of biological NMR spectroscopic data. Xenobiotica.
29:1181–1189. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Coen M, Holmes E, Lindon JC and Nicholson
JK: NMR-based metabolic profiling and metabonomic approaches to
problems in molecular toxicology. Chem Res Toxicol. 21:9–27. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Asiago VM, Alvarado LZ, Shanaiah N, Gowda
GA, Owusu-Sarfo K, Ballas RA and Raftery D: Early detection of
recurrent breast cancer using metabolite profiling. Cancer Res.
70:8309–8318. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Sreekumar A, Poisson LM, Rajendiran TM,
Khan AP, Cao Q, Yu J, Laxman B, Mehra R, Lonigro RJ, Li Y, et al:
Metabolomic profiles delineate potential role for sarcosine in
prostate cancer progression. Nature. 457:910–914. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Sato Y, Suzuki I, Nakamura T, Bernier F,
Aoshima K and Oda Y: Identification of a new plasma biomarker of
Alzheimer's disease using metabolomics technology. J Lipid Res.
53:567–576. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Fiehn O, Kopka J, Dormann P, Altmann T,
Trethewey RN and Willmitzer L: Metabolite profiling for plant
functional genomics. Nat Biotechnol. 18:1157–1161. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wiklund S, Johansson E, Sjöström L,
Mellerowicz EJ, Edlund U, Shockcor JP, Gottfries J, Moritz T and
Trygg J: Visualization of GC/TOF-MS-based metabolomics data for
identification of biochemically interesting compounds using OPLS
class models. Anal Chem. 80:115–122. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Hong Z, Lin Z, Liu Y, Tan G, Lou Z, Zhu Z,
Chai Y, Fan G, Zhang J and Zhang L: Innovative microwave-assisted
oximation and silylation procedures for metabolomic analysis of
plasma samples using gas chromatography-mass spectrometry. J
Chromatogr A. 1254:14–22. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Cai XF, Chen LF, Wang ZL and Gu YE: Effect
of acupoint injection by Astragalus injection on local SIgA
and pathomorphology changes in rats with chronic pelvic
inflammatory disease. Zhongguo Zhong Yao Za Zhi. 31:1361–1364.
2006.(In Chinese). PubMed/NCBI
|
|
17
|
Li X, Guo J, Shi Z and Nie J: Effect of
Fuke Qianjin tablets on inflammatory cytokines in blood serum in
rats with chronic pelvic inflammatory disease. Zhongguo Shi Yan
Fang Ji Xue Za Zhi. 19:226–228. 2013.(In Chinese).
|
|
18
|
Benjamini Y and Hochberg Y: Controlling
the false discovery rate: A practical and powerful approach to
multiple testing. J R Stat Soc Series B Stat Methodol. 57:289–300.
1995.
|
|
19
|
Sonnex C: Toll-like receptors and genital
tract infection. Int J STD AIDS. 21:153–157. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
He J, You X, Zeng Y, Yu M, Zuo L and Wu Y:
Mycoplasma genitalium-derived lipid-associated membrane proteins
activate NF-kappaB through toll-like receptors 1, 2 and 6 and CD14
in a MyD88-dependent pathway. Clin Vaccine Immunol. 16:1750–1757.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Sheldon IM, Rycroft AN, Dogan B, Craven M,
Bromfield JJ, Chandler A, Roberts MH, Price SB, Gilbert RO and
Simpson KW: Specific strains of Escherichia coli are
pathogenic for the endometrium of cattle and cause pelvic
inflammatory disease in cattle and mice. PLoS One. 5:e91922010.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zhou B, Cong L and Sha Y: Pathogens of
transmitted disease in the pathogenesis of acute pelvic
inflammatory disease. Zhonghua Fu Chan Ke Za Zhi. 36:539–541.
2001.(In Chinese). PubMed/NCBI
|
|
23
|
Angerman NS, Evans MI, Moravec WD,
Schumacher GF and Hajj SN: C-reactive protein in the evaluation of
antibiotic therapy for pelvic infection. J Reprod Med. 25:63–66.
1980.PubMed/NCBI
|
|
24
|
Workowski KA and Berman S: Centers for
Disease Control and Prevention (CDC): Sexually transmitted diseases
treatment guidelines, 2010. MMWR Recomm Rep. 59:1–110.
2010.PubMed/NCBI
|
|
25
|
Cheng W, Shivshankar P, Li Z, Chen L, Yeh
IT and Zhong G: Caspase-1 contributes to Chlamydia
trachomatis-induced upper urogenital tract inflammatory
pathologies without affecting the course of infection. Infect
Immun. 76:515–522. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Trunov A, Obukhova O, Gorbenko O, Shvayk A
and Trunova L: Cytokines, estradiol and progesterone in the plasma
of women of reproductive age with pelvic inflammatory disease in
remission. Adv Biosci Biotechnol. 4:7272013. View Article : Google Scholar
|
|
27
|
Lee SA, Tsai HT, Ou HC, Han CP, Tee YT,
Chen YC, Wu MT, Chou MC, Wang PH and Yang SF: Plasma
interleukin-1beta, −6, −8 and tumor necrosis factor-alpha as highly
informative markers of pelvic inflammatory disease. Clin Chem Lab
Med. 46:997–1003. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Weljie AM, Dowlatabadi R, Miller BJ, Vogel
HJ and Jirik FR: An inflammatory arthritis-associated metabolite
biomarker pattern revealed by 1H NMR spectroscopy. J Proteome Res.
6:3456–3464. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Yue R, Zhao L, Hu Y, Jiang P, Wang S,
Xiang L, Liu W, Zhang W and Liu R: Rapid-resolution liquid
chromatography TOF-MS for urine metabolomic analysis of
collagen-induced arthritis in rats and its applications. J
Ethnopharmacol. 145:465–475. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Hotter G, Closa D, Prats N, Pi F, Gelpí E
and Roselló-Catafau J: Free radical enhancement promotes leucocyte
recruitment through a PAF and LTB4 dependent mechanism. Free Radic
Biol Med. 22:947–954. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Seibert K, Zhang Y, Leahy K, Hauser S,
Masferrer J, Perkins W, Lee L and Isakson P: Pharmacological and
biochemical demonstration of the role of cyclooxygenase 2 in
inflammation and pain. Proc Natl Acad Sci USA. 91:12013–12017.
1994. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Heinonen PK, Aine R and Seppälä E:
Peritoneal fluid leukotriene B4 and prostaglandin E2 in acute
salpingitis. Gynecol Obstet Invest. 29:292–295. 1990. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Soga T, Baran R, Suematsu M, Ueno Y, Ikeda
S, Sakurakawa T, Kakazu Y, Ishikawa T, Robert M, Nishioka T and
Tomita M: Differential metabolomics reveals ophthalmic acid as an
oxidative stress biomarker indicating hepatic glutathione
consumption. J Biol Chem. 281:16768–16776. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Jana SK, Dutta M, Joshi M, Srivastava S,
Chakravarty B and Chaudhury K: 1H NMR based targeted
metabolite profiling for understanding the complex relationship
connecting oxidative stress with endometriosis. Biomed Res Int.
2013:3290582013. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Valore EV, Park CH, Igreti SL and Ganz T:
Antimicrobial components of vaginal fluid. Am J Obstet Gynecol.
187:561–568. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Krebs HA and Johnson WA: Metabolism of
ketonic acids in animal tissues. Biochem J. 31:645–660. 1937.
View Article : Google Scholar : PubMed/NCBI
|