Introduction
Osteosarcoma (OS) is the most prevalent malignant
bone tumour in children and adolescents, and it generally occurs in
the metaphysis of long bones (1,2).
Furthermore, OS is characterized by high relapse, and ~80% of OS
patients eventually develop metastatic disease (1,3). Despite
the combination of surgery, radiotherapy and chemotherapy, the
5-year survival of advanced OS patients is still poor (1,2).
Therefore, it is urgently required to explore the potential
molecular mechanism underlying OS progression and to identify
effective molecular targets for the treatment of OS.
Long non-coding RNAs (lncRNAs) are a type of
regulatory transcript >200 nucleotides in length that mainly
function through sponging target mRNAs, microRNAs (miRNAs or miRs)
or proteins (4,5). miRNAs are a class of regulatory
transcripts containing 22–25 nucleotides and are important gene
expression regulators through their interaction with the
3′-untranslated region of their target mRNAs, resulting in mRNA
degradation or translation repression (6,7). By
regulating the expression of their targets, both lncRNAs and miRNAs
have been demonstrated to serve key roles in the development and
progression of various human cancers including OS (7–9). For
instance, miR-33b inhibits the proliferation and migration of
osteosarcoma cells via targeting hypoxia-inducible factor 1-α
(7). The lncRNA metastasis
associated lung adenocarcinoma transcript 1 promotes the
proliferation and metastasis of osteosarcoma cells via
downregulation of miR-509 and, thus, activation of the Rac1/c-Jun
N-terminal kinasepathway (8).
The lncRNA X inactive-specific transcript (XIST) is
expressed exclusively from the X inactivation centre of the
inactive X chromosome and is essential for the initiation and
spread of X-inactivation (10).
Previous studies have reported that XIST is frequently upregulated
in human cancers, and it functions as an oncogene (11,12). For
instance, XIST promotes cell growth and metastasis through
regulating the miR-139-5p mediated Wnt/β-catenin signalling pathway
in bladder cancer (13). Li et
al previously reported that XIST promotes transforming growth
factor (TGF)-β-induced epithelial-mesenchymal transition by
regulating the miR-367/141-zinc finger E-box binding homeobox 2
axis in non-small-cell lung cancer (14). Recently, XIST has been demonstrated
to be upregulated in OS tissues and cell lines, and its
upregulation is significantly associated with advanced tumour size,
advanced clinical stage, distant metastasis and shorter survival
time of OS patients (15). In
addition, XIST expression has been demonstrated to be an
independent prognostic factor for the survival of patients with OS
(15). Furthermore, several miRNAs
have been identified to be sponged by XIST in OS cells, including
miR-320b (16), miR-193a-3p
(17), miR-195-5p (18) and miR-21-5p (19). For example, Lv et al
demonstrated that XIST promoted OS progression by targeting RAP2B
via miR-320b (16). Yang et
al reported that XIST enhanced OS growth by targeting YAP via
miR-195-5p (18). However, whether
XIST also interacts with other miRNAs in OS cells remains
unclear.
The aim of the present study was to explore the
molecular mechanism by which XIST promotes OS growth and metastasis
in vitro.
Materials and methods
Clinical tissues
The present study was approved by the Medical Ethics
Committee of First Affiliated Hospital of Hunan College of
Traditional Chinese Medicine (Zhuzhou, China). OS tissues and
matched adjacent non-tumour tissues were collected from 35 patients
with primary OS at the Department of Orthopaedics, First Affiliated
Hospital of Hunan College of Traditional Chinese Medicine between
April 2014 and March 2017. Written informed consent was obtained
from all participating patients. These patients included 15 females
and 20 males, aged from 11–33 years with a mean age of 18.5 years.
The exclusion criterion was that no patient received radiotherapy
or chemotherapy prior to surgical resection. These tissues were
immediately frozen and stored in liquid nitrogen prior to use.
Cell culture
Human OS cell lines (Saos-2, U2OS, HOS and MG63) and
normal human osteoblast cell line HFOB1.19 were purchased from Cell
Bank of Chinese Academy of Sciences (Shanghai, China). These cells
were cultured in Dulbecco's modified Eagle's medium (DMEM; Thermo
Fisher Scientific, Inc., Waltham, MA, USA) supplemented with 10%
foetal bovine serum (FBS; Thermo Fisher Scientific, Inc.). Cells
were incubated at 37°C in a humidified atmosphere with 5%
CO2.
Cell transfection
For XIST and miR-137 function analysis, Saos-2 and
U2OS cells were transiently transfected with 100 µM of negative
control (NC) small interfering (si)RNA (4390843; Thermo Fisher
Scientific, Inc.), 100 µM of XIST siRNA (4390771; Thermo Fisher
Scientific, Inc.), 4 ug pcDNA3.1 vector (V79020; Thermo Fisher
Scientific, Inc.), 4 ug of pcDNA-XIST expression plasmid (P00238;
Amspring, Changsha, China), 100 µM of miR-137 inhibitor (4464084;
Thermo Fisher Scientific, Inc.) or 100 µM of NC inhibitor (4464076;
Thermo Fisher Scientific, Inc.) using Lipofectamine 2000 (Thermo
Fisher Scientific, Inc.) according to the manufacturer's
instructions. Following 48 h of transfection, the expression levels
of XIST or miR-137 were examined to confirm the transfection
efficiency.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR) analysis
Total RNA was extracted from tissues and cells using
TRIzol reagent (Thermo Fisher Scientific, Inc.). For XIST
expression detection, RT-qPCR was conducted using the OneStep
RT-PCR kit (Qiagen, Inc., Valencia, CA, USA) according to the
manufacturer's instructions. GAPDH was used as an internal
reference. For miR-137 expression detection, RT-qPCR was conducted
using the Mir-X™ miRNA qRT-PCR SYBR® kit (Clontech
Laboratories, Inc., Mountainview, CA, USA) according to the
manufacturer's instructions. U6 was used as an internal reference.
The reaction conditions were as follows: 95°C for 3 min, followed
by 40 cycles of 95°C for 30 sec and 60°C for 30 sec. The relative
expression was analysed using the 2−ΔΔCq method
(20). Primers used were as follows:
XIST, forward 5′-ACGCTGCATGTGTCCTTAG-3′ and reverse
5′-GAGCCTCTTATAGCTGTTTG-3′; GAPDH, forward
5′-GGAGCGAGATCCCTCCAAAAT-3′ and reverse
5′-GGCTGTTGTCATACTTCTCATGG-3′; miR-137, forward
5′-GCAGCAAGAGTTCTGGTGGC-3′ and reverse 5′-TGGAACCAGTGCGAAAACAC-3′;
and U6, forward 5′-CTCGCTTCGGCAGCACA-3′ and reverse
5′-AACGCTTCACGAATTTGCGT-3′.
Western blotting
Saos-2 and U2OS cells were lysed in cold
radioimmunoprecipitation assay buffer (Thermo Fisher Scientific,
Inc.). The protein concentration was examined using a Bicinchoninic
Acid Protein Assay kit (Thermo Fisher Scientific, Inc.). Proteins
(50 µg per lane) were separated by 10% SDS-PAGE and then
transferred to polyvinylidene difluoride membranes (Thermo Fisher
Scientific, Inc.). The membranes were blocked in 5% non-fat milk in
PBS (Thermo Fisher Scientific, Inc.) containing 0.1% Tween-20
(Thermo Fisher Scientific, Inc.) at room temperature for 3 h.
Following that, the membranes were incubated with rabbit
anti-matrix metalloproteinase (MMP)2 (1:1,000; ab92536; Abcam,
Cambridge, MA, USA), rabbit anti-MMP9 (1:500; ab76003; Abcam) or
rabbit anti-GAPDH (1:1,000; ab9485; Abcam) primary antibodies at
4°C overnight. Then, the membranes were incubated with a
horseradish peroxidase-conjugated goat anti-rabbit secondary
antibody (1:5,000; ab205718; Abcam) at room temperature for 30 min.
The protein bands were detected using an Enhanced Chemiluminescence
Western Blotting kit (Thermo Fisher Scientific, Inc.) and
quantified using Image Lab analysis software version 3.1 (Bio-Rad
Laboratories, Inc., Hercules, CA, USA).
Cell counting kit (CCK)-8 assay
Saos-2 and U2OS cells (5,000 cells/well) were seeded
into 96-well plates. Following incubation at 37°C for 0, 24, 48 or
72 h, 10 ml CCK-8 reagent (Beyotime Institute of Biotechnology,
Haimen, China) was added to each well. The cells were then
incubated at 37°C for 2 h. The absorbance was examined at a
wavelength of 450 nm using an ELx808 absorbance reader (BioTek
Instruments, Inc., Winooski, VT, USA).
Cell invasion assay
The Saos-2 and U2OS cell suspensions
(1×105 cells per well) were added to the upper chamber
of a Transwell plate that was pre-coated with Matrigel (Chemicon;
EMD Millipore, Billerica, MA, USA), and 300 µl DMEM containing 10%
FBS was added to the lower chamber. Cells were then incubated at
37°C for 24 h. Following that, the cells on the interior of the
inserts were removed using a cotton-tipped swab, while the cells on
the lower surface of the membrane were stained with gentian violet
(Sigma-Aldrich; Merck KGaA, Darmstadt, Germany) at room temperature
for 30 min. The invading cells were counted under a light
microscope (magnification, ×200; Olympus Corp., Tokyo, Japan).
Bioinformatics analysis and Luciferase
reporter gene assay
The target miRNAs of XIST were predicted using
RNAhybrid 2.12 (http://bibiserv.techfak.uni-bielefeld.de/rnahybrid/).
The fragment from XIST containing the wild-type (WT) or mutant type
(MT) miR-137 binding sites was cloned into pmirGLO Dual-luciferase
Target Vector (Promega Corporation, Madison, WI, USA), generating
WT or MT XIST plasmids. Saos-2 and U2OS cells were co-transfected
with miR-137 mimic (4464066; Thermo Fisher Scientific, Inc.) or
miRNA-NC (4464058; Thermo Fisher Scientific, Inc.), and WT (or MT)
XIST plasmids using Lipofectamine 2000. Following 48 h of
transfection, luciferase reporter gene assay was performed using
the Dual-Luciferase Reporter Assay System (Promega Corporation).
Firefly luciferase activity was normalized to Renilla
luciferase activity.
Statistical analysis
Data are presented as the mean ± standard deviation
of at least three repeated experiments. SPSS version 20.0 software
(IBM Corp., Armonk, NY, USA) was used to conduct statistical
analyses. Differences were examined using Student's t-test for
two-group comparison. Differences were examined using analysis of
variance for comparison of more than two groups followed by
Turkey's post hoc test. Pearson correlation analysis was performed
to examine the correlation between the XIST and miR-137 expression
in OS tissues. P<0.05 was considered to indicate a statistically
significant difference.
Results
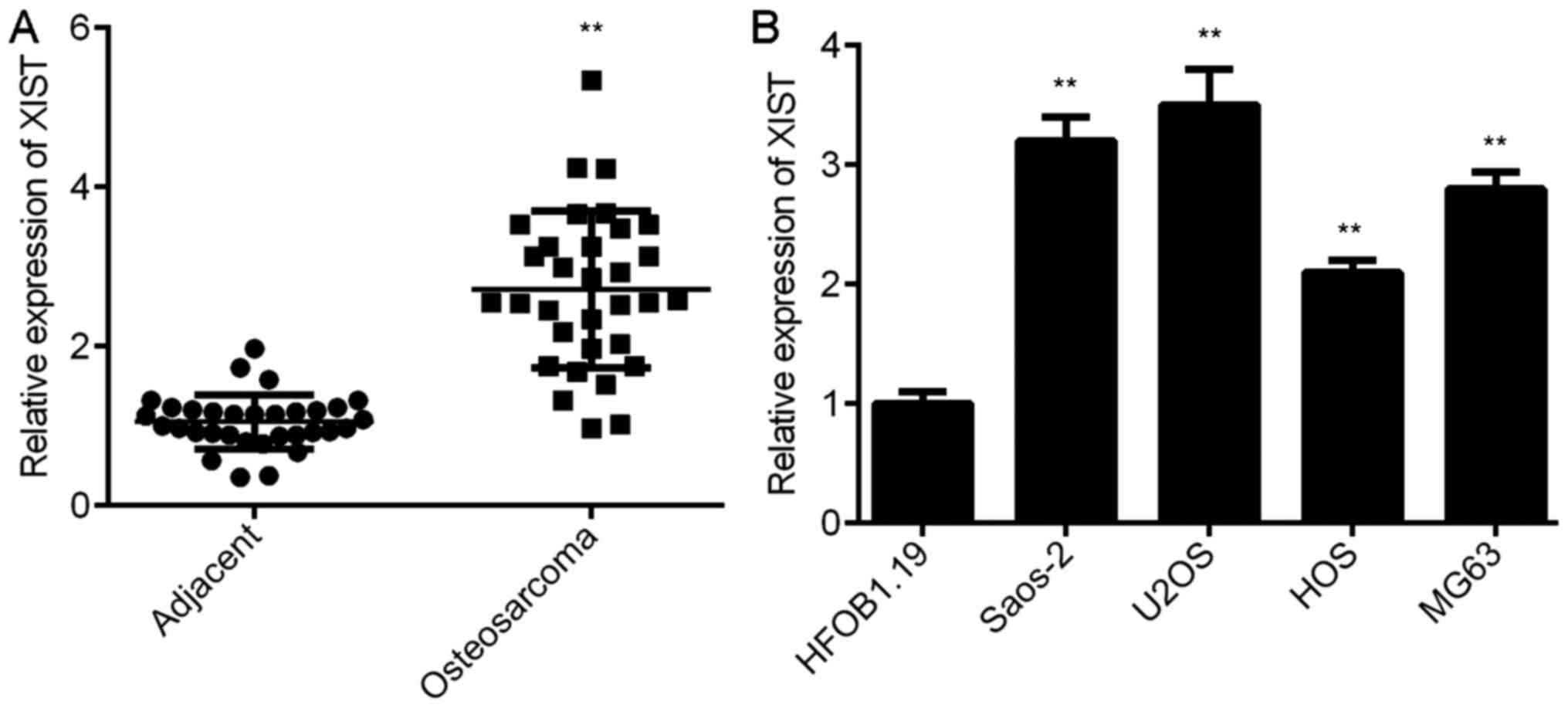
XIST is upregulated in OS
To study the role of XIST in OS, the expression
levels of XIST were initially examined in OS tissues and cell
lines. RT-qPCR data demonstrated that XIST was significantly
upregulated in OS tissues compared with that of matched adjacent
non-tumour tissues (Fig. 1A).
Similar findings demonstrated that the expression of XIST was
increased in OS cell lines (MG63, Saos-2, U2OS and HOS) compared
with that of normal human osteoblast cell line HFOB1.19 (Fig. 1B). Therefore, XIST is upregulated in
OS.
XIST negatively regulates the
expression of miR-137 in OS cells by sponging
LncRNAs have been demonstrated to function by
interacting directly with miRNAs (4,5). Thus,
bioinformatics analysis was performed to identify the target miRNAs
of XIST. Putative binding sites were identified between XIST and
miR-137 (Fig. 2A). To validate this
prediction, a luciferase reporter plasmid was constructed
containing the WT or MT miR-137 binding sites in XIST (Fig. 2A) and luciferase reporter gene assay
was performed using Saos-2 and U2OS cells. The data demonstrated
that transfection with the miR-137 mimic caused a significant
reduction in the luciferase activity of WT XIST but did not
significantly affect the luciferase activity of MT XIST in Saos-2
and U2OS cells (Fig. 2B and C).
These data indicate that XIST can directly sponge miR-137 in OS
cells.
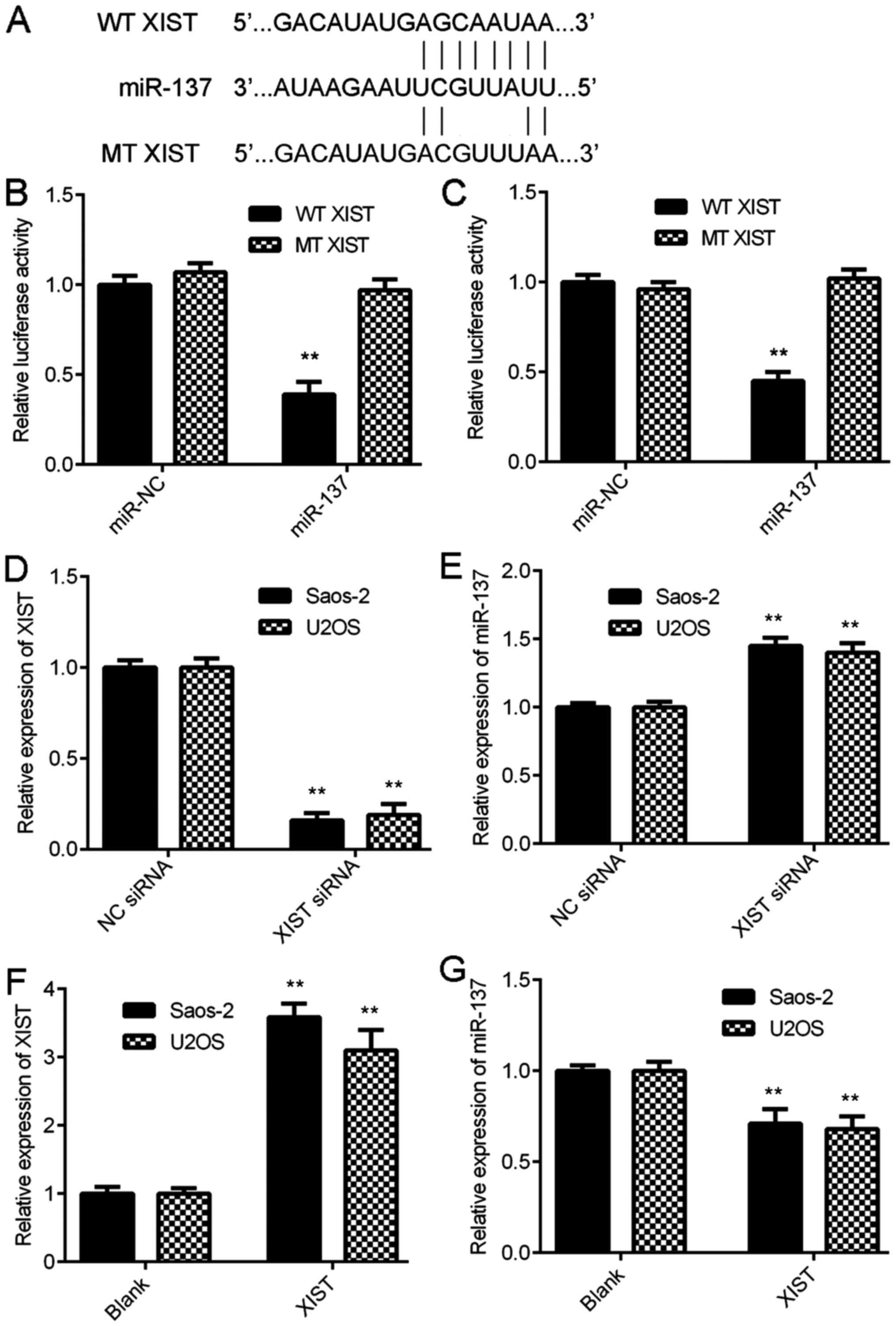 | Figure 2.XIST negatively regulates the
expression of miR-137 in OS cells by sponging. (A) The luciferase
reporter plasmid containing the WT or MT miR-137 binding sites in
XIST were constructed. (B) Saos-2 and (C) U2OS cells were used for
the luciferase reporter gene assay, and transfection with miR-137
mimic significantly inhibited the luciferase activity of WT XIST in
OS cells but did not significantly affect the luciferase activity
of MT XIST. **P<0.01 vs. miR-NC. (D) Saos-2 and (E) U2OS cells
were transfected with XIST siRNA or NC siRNA, and RT-qPCR was
conducted to examine the expression of XIST and miR-137.
**P<0.01 vs. NC siRNA. (F) Saos-2 and (G) U2OS were transfected
with a XIST plasmid or blank vector, and RT-qPCR was conducted to
examine the expression of XIST and miR-137. **P<0.01 vs. Blank.
XIST, X inactive-specific transcript; miR, microRNA; OS,
osteosarcoma; WT, wild-type; MT, mutant type; NC, negative control;
siRNA, small interfering RNA; RT-qPCR, reverse
transcription-quantitative polymerase chain reaction. |
The effect of XIST on the miR-137 expression was
further studied in OS cells. XIST siRNA or XIST plasmid was used to
transfect Saos-2 and U2OS cells. As presented in Fig. 2D, transfection with XIST siRNA
significantly inhibited its expression compared with the NC siRNA
group. Further experiments demonstrated that inhibition of XIST
expression caused a significant increase in miR-137 expression in
Saos-2 and U2OS cells (Fig. 2E). In
contrast, transfection with XIST plasmid significantly upregulated
its expression compared with the blank group (Fig. 2F). Furthermore, overexpression of
XIST significantly reduced the expression of miR-137 in Saos-2 and
U2OS cells (Fig. 2G). These data
suggest that XIST can negatively regulate the miR-137 expression in
OS cells by directly sponging it.
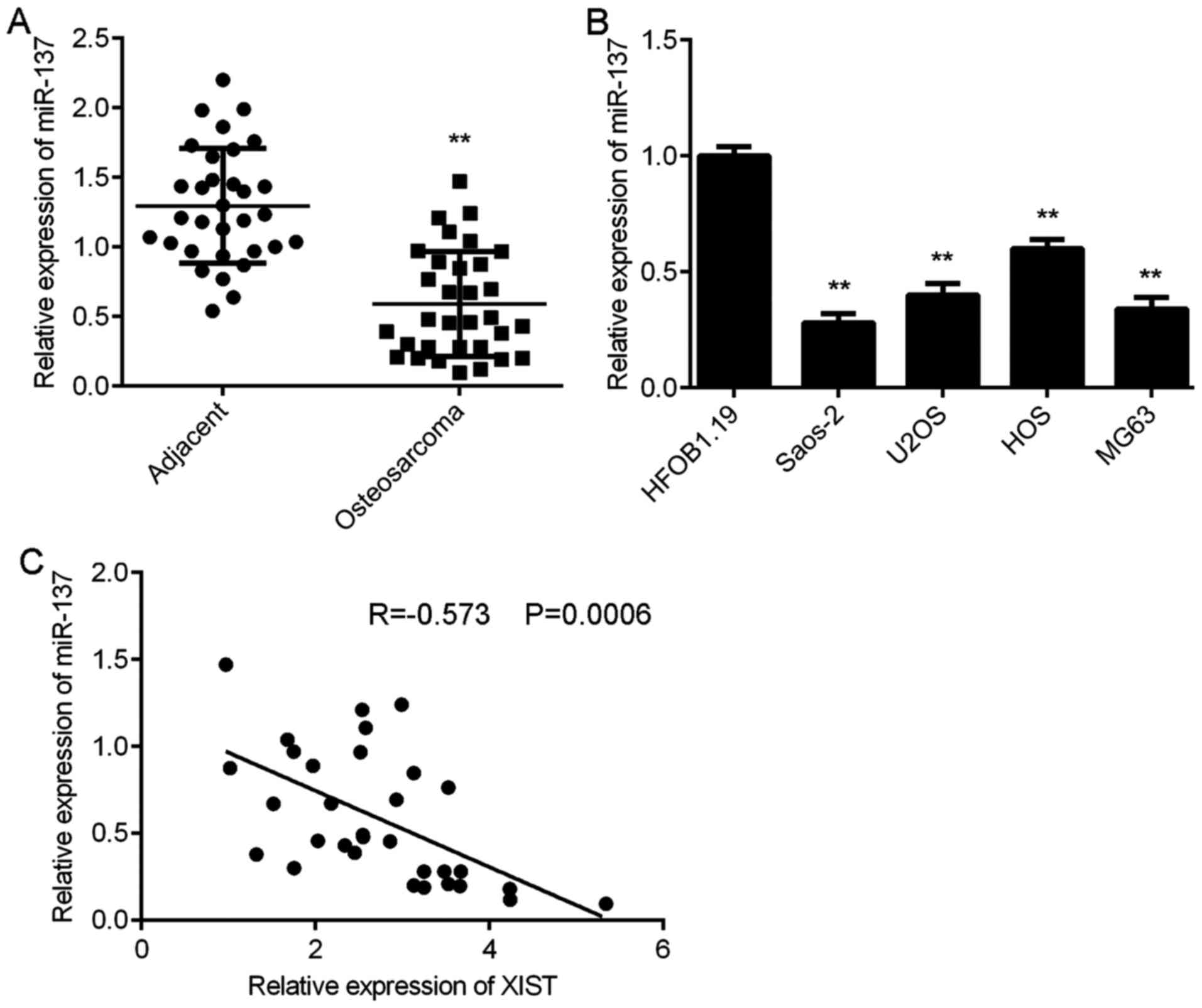
miR-137 is downregulated in OS
The expression of miR-137 in OS was further
examined. RT-qPCR data demonstrated that the expression levels of
miR-137 were significantly reduced in OS tissues compared with
those of adjacent non-tumour tissues (Fig. 3A). Consistently, miR-137 was also
downregulated in OS cell lines compared with that of normal human
osteoblasts cell line HFOB1.19 (Fig.
3B). Notably, an inverse correlation was identified between the
XIST and miR-137 expression in OS tissues (Fig. 3C). These findings suggest that the
reduced expression of miR-137 may be due to the increased
expression of XIST in OS tissues.
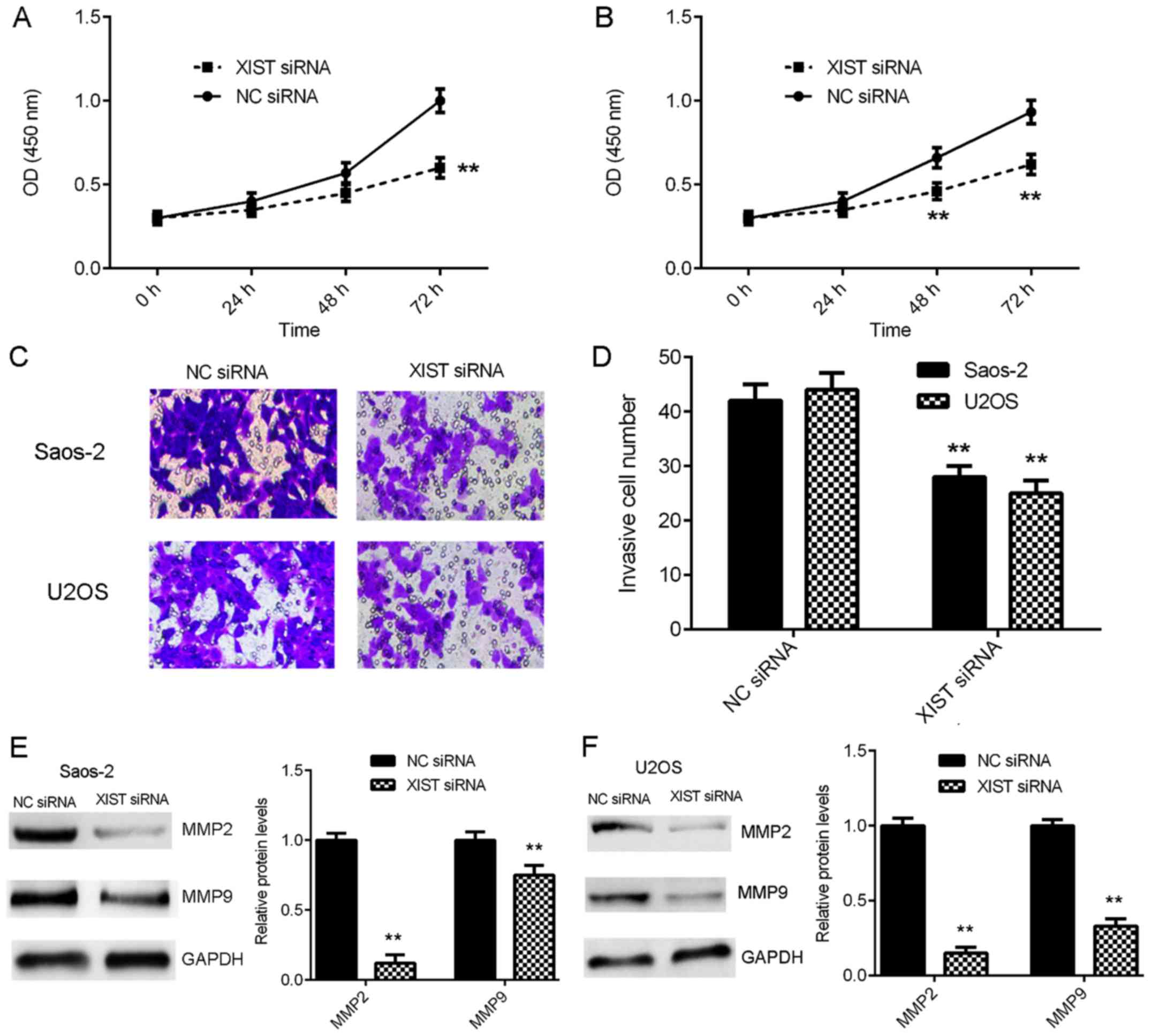
Inhibition of XIST expression
suppresses the proliferation and invasion of OS cells
The effects of XIST downregulation on OS cell
proliferation and invasion were evaluated. CCK-8 assay data
indicated that the proliferation of Saos-2 and U2OS cells was
significantly downregulated following knockdown of XIST compared
with that of the NC siRNA group (Fig. 4A
and B). Transwell assay data demonstrated that the invasion of
Saos-2 and U2OS cells was also significantly reduced in the XIST
siRNA group compared with the NC siRNA group (Fig. 4C and D). Furthermore, the protein
levels of MMP2 and MMP9, which are not direct target genes of XIST
and miR-137, but are key factors associated with tumour cell
invasion, were also significantly decreased following inhibition of
XIST expression (Fig. 4E-F). These
findings suggest that XIST may serve a promoting role in OS growth
and metastasis.
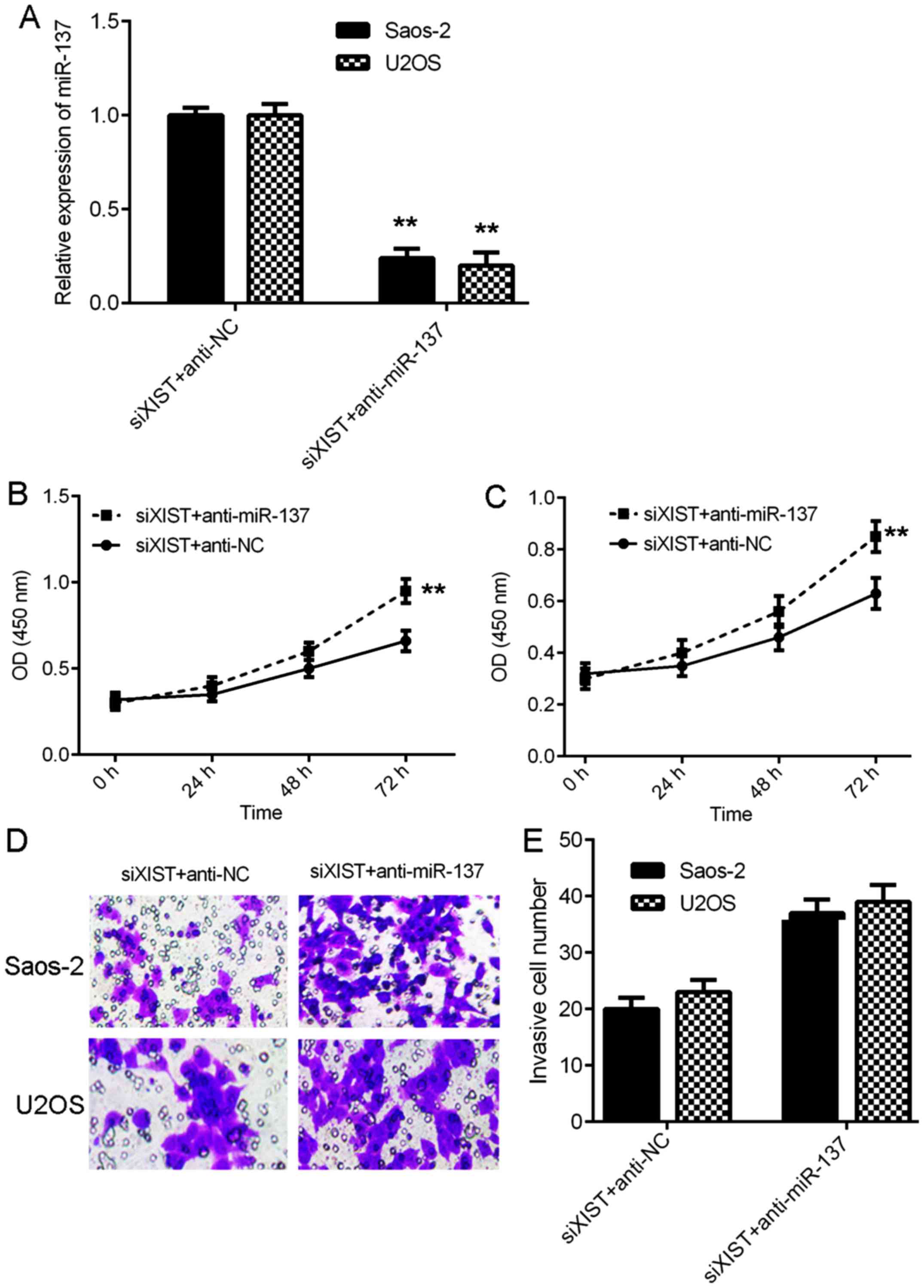
miR-137 is associated with
XIST-mediated OS cell proliferation and invasion
The above findings suggest that miR-137 may be
associated with XIST-mediated OS cell proliferation and invasion.
To test this hypothesis, XIST siRNA transfected cells were then
transfected with an miR-137 inhibitor to downregulate the
expression of miR-137 in OS cells. RT-qPCR data demonstrated that
miR-137 was significantly downregulated in the siXIST+anti-miR-137
group compared with the siXIST+anti-NC group (Fig. 5A). Furthermore, CCK-8 assay data
demonstrated that the proliferation of Saos-2 and U2OS cells was
significantly increased in the siXIST+anti-miR-137 group compared
with the siXIST+anti-NC group (Fig. 5B
and C). Similarly, Transwell assay data demonstrated that the
invasion of Saos-2 and U2OS cells was also upregulated in the
siXIST+anti-miR-137 group compared with the siXIST+anti-NC group
(Fig. 5D-E). Therefore, the present
findings suggest that XIST serves a promoting role in the
regulation of OS cell proliferation and invasion by directly
sponging miR-137.
Discussion
XIST has been implicated in OS, but the underlying
molecular mechanism remains largely unclear. In the present study
it was reported that XIST was significantly upregulated in OS
tissues and cell lines compared with adjacent non-tumour tissues
and normal human osteoblast cells. Bioinformatics analysis and
luciferase reporter gene assay data demonstrated that XIST could
directly target miR-137, and the expression of miR-137 was
negatively regulated by XIST in Saos-2 and U2OS cells. Furthermore,
miR-137 was markedly downregulated in OS tissues and cell lines,
and an inverse association was observed between XIST and miR-137
expression in OS tissues. Knockdown of XIST caused a significant
reduction in cell proliferation and invasion and suppressed MMP2
and MMP9 protein levels in Saos-2 and U2OS cells. Furthermore,
inhibition of miR-137 expression abolished the effects of XIST
downregulation on the proliferation and invasion of OS cells.
Recently, the increased expression of XIST has been
observed in different human cancers, and previous studies have
demonstrated that XIST serves an oncogenic role (21,22). For
instance, Sun et al reported that XIST was significantly
upregulated in pancreatic cancer, and it exerts oncogenic functions
via miR-34a-5p (21). Jiang et
al demonstrated that XIST was significantly upregulated in
NSCLC, and inhibition of XIST expression effectively suppressed
NSCLC cell viability and invasion by regulating the
miR-137/paxillin axis (22). In the
present study, it was demonstrated that the expression levels of
XIST were significantly upregulated in OS tissues and cells
compared with adjacent non-tumour tissues and normal human
osteoblast cell line HFOB1.19. These findings were consistent with
several previous studies (15,16). It
was also demonstrated that inhibition of XIST expression by siRNA
significantly reduced the proliferation and invasion of Saos-2 and
U2OS cells. Similarly, Yang et al (18) also demonstrated that knockdown of
XIST inhibited OS cell proliferation and invasion in vitro
and tumour growth in vivo.
Previous studies have demonstrated that lncRNAs
including XIST can negatively regulate the expression of miRNAs
through sponging (23,24). For instance, Zhang et al
demonstrated that XIST promoted gastric cancer progression through
TGF-β1 via targeting miR-185 (23).
Cheng et al reported that XIST suppressed nasopharyngeal
carcinoma progression by activating miR-491-5p (24). In OS, several miRNAs have been
identified as the targets of XIST, such as miR-195-5p (18), miR-21-5p (19) and miR-320b (16). In the present study, bioinformatics
analysis data demonstrated that miR-137 was a potential target of
XIST, which was confirmed by luciferase reporter gene assay. The
expression of miR-137 was negatively regulated by XIST in OS cells.
Furthermore, it was observed that miR-137 was significantly
downregulated in OS tissues and cell lines, and the miR-137 levels
were inversely correlated to the XIST levels in OS tissues. These
findings suggest that the upregulation of XIST may contribute to
the downregulation of miR-137 in OS tissues.
A previous study has demonstrated that miR-137
functions as a tumour suppressor in OS via targeting EZH2 (25). In addition, Li et al also
reported that miR-137 is downregulated in OS tissues and cell
lines, consistent with the present findings, and can inhibit OS
cell proliferation and migration through directly targeting FXYD6
(26). However, the molecular
mechanism of miR-137 underlying OS progression remains largely
unclear. In the present study, it was identified that inhibition of
miR-137 reversed the suppressive effects of XIST downregulation on
proliferation and invasion of Saos-2 and U2OS cells. These findings
suggest that miR-137 functions as a downstream effector of XIST in
OS cells.
To the best of our knowledge, this is the first
study to report that XIST has promoting effects on proliferation
and invasion of OS cells by sponging miR-137. Therefore, these
finding suggest suggest that XIST may become a potential
therapeutic target for the treatment of OS.
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
All data generated or analysed during this study are
included in this published article.
Authors' contributions
HL collected clinical tissues, examined the
expression in tissues, and wrote the manuscript. SH and HY designed
the study and revised the manuscript. JC, BX and LL performed the
in vitro experiments.
Ethics approval and consent to
participate
This study was approved by the Medical Ethics
Committee of First Affiliated Hospital of Hunan College of
Traditional Chinese Medicine (Zhuzhou, China) and written informed
consents was obtained from all patients.
Patient consent for publication
Written informed consents was obtained from all
patients.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Lindsey BA, Markel JE and Kleinerman ES:
Osteosarcoma overview. Rheumatol Ther. 4:25–43. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2015. CA Cancer J Clin. 65:5–29. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Maximov VV and Aqeilan RI: Genetic factors
conferring metastasis in osteosarcoma. Future Oncol. 12:1623–1644.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Wang S, Hui Y, Li X and Jia Q: Silencing
of lncRNA-CCDC26 restrains the growth and migration of glioma cells
in vitro and in vivo via targeting miR-203. Oncol Res.
26:1143–1154. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Li J, Zi Y, Wang W and Li Y: LncRNA MEG3
inhibits cell proliferation and metastasis in chronic myeloid
leukemia via targeting MiR-184. Oncol Res. 26:297–305. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Zhu K, He Y, Xia C, Yan J, Hou J, Kong D,
Yang Y and Zheng G: MicroRNA-15a inhibits proliferation and induces
apoptosis in CNE1 nasopharyngeal carcinoma cells. Oncol Res.
24:145–151. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Zhou Y, Yang C, Wang K, Liu X and Liu Q:
MicroRNA-33b inhibits the proliferation and migration of
osteosarcoma cells via targeting hypoxia-inducible factor-1alpha.
Oncol Res. 25:397–405. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Zhang Y, Dai Q, Zeng F and Liu H: MALAT1
promotes the proliferation and metastasis of osteosarcoma cells by
activating the Rac1/JNK pathway via targeting miR-509. Oncol Res.
Apr 27–2018.(Epub ahead of print).
|
|
9
|
Wang Y, Yang T, Zhang Z, Lu M, Zhao W,
Zeng X and Zhang W: Long non-coding RNA TUG1 promotes migration and
invasion by acting as a ceRNA of miR-335-5p in osteosarcoma cells.
Cancer Sci. 108:859–867. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Gilbert SL, Pehrson JR and Sharp PA: XIST
RNA associates with specific regions of the inactive X chromatin. J
Biol Chem. 275:36491–36494. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Chen DL, Chen LZ, Lu YX, Zhang DS, Zeng
ZL, Pan ZZ, Huang P, Wang FH, Li YH, Ju HQ and Xu RH: Long
noncoding RNA XIST expedites metastasis and modulates
epithelial-mesenchymal transition in colorectal cancer. Cell Death
Dis. 8:e30112017. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Xu Z, Xu J, Lu H, Lin B, Cai S, Guo J,
Zang F and Chen R: LARP1 is regulated by the XIST/miR-374a axis and
functions as an oncogene in non-small cell lung carcinoma. Oncol
Rep. 38:3659–3667. 2017.PubMed/NCBI
|
|
13
|
Hu Y, Deng C, Zhang H, Zhang J, Peng B and
Hu C: Long non-coding RNA XIST promotes cell growth and metastasis
through regulating miR-139-5p mediated Wnt/β-catenin signaling
pathway in bladder cancer. Oncotarget. 8:94554–94568.
2017.PubMed/NCBI
|
|
14
|
Li C, Wan L, Liu Z, Xu G, Wang S, Su Z,
Zhang Y, Zhang C, Liu X, Lei Z and Zhang HT: Long non-coding RNA
XIST promotes TGF-β-induced epithelial-mesenchymal transition by
regulating miR-367/141-ZEB2 axis in non-small-cell lung cancer.
Cancer Lett. 418:185–195. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Li GL, Wu YX, Li YM and Li J: High
expression of long non-coding RNA XIST in osteosarcoma is
associated with cell proliferation and poor prognosis. Eur Rev Med
Pharmacol Sci. 21:2829–2834. 2017.PubMed/NCBI
|
|
16
|
Lv GY, Miao J and Zhang XL: Long
non-coding RNA XIST promotes osteosarcoma progression by targeting
ras-related protein RAP2B via miR-320b. Oncol Res. 26:837–846.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Wu D, Nie X, Ma C, Liu X, Liang X, An Y,
Zhao B and Wu X: RSF1 functions as an oncogene in osteosarcoma and
is regulated by XIST/miR-193a-3p axis. Biomed Pharmacother.
95:207–214. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Yang C, Wu K, Wang S and Wei G: Long
non-coding RNA XIST promotes osteosarcoma progression by targeting
YAP via miR-195-5p. J Cell Biochem. 119:5646–5656. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Zhang R and Xia T: Long non-coding RNA
XIST regulates PDCD4 expression by interacting with miR-21-5p and
inhibits osteosarcoma cell growth and metastasis. Int J Oncol.
51:1460–1470. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Sun Z, Zhang B and Cui T: Long non-coding
RNA XIST exerts oncogenic functions in pancreatic cancer via
miR-34a-5p. Oncol Rep. 39:1591–1600. 2018.PubMed/NCBI
|
|
22
|
Jiang H, Zhang H, Hu X and Li W: Knockdown
of long non-coding RNA XIST inhibits cell viability and invasion by
regulating miR-137/PXN axis in non-small cell lung cancer. Int J
Biol Macromol. 111:623–631. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zhang Q, Chen B, Liu P and Yang J: XIST
promotes gastric cancer (GC) progression through TGF-beta1 via
targeting miR-185. J Cell Biochem. 119:2787–2796. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Cheng Q, Xu X, Jiang H, Xu L and Li Q:
Knockdown of long non-coding RNA XIST suppresses nasopharyngeal
carcinoma progression by activating miR-491-5p. J Cell Biochem.
119:3936–3944. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Feng Q, Wu Q, Liu X, Xiong Y and Li H:
MicroRNA-137 acts as a tumor suppressor in osteosarcoma by
targeting enhancer of zeste homolog 2. Exp Ther Med. 13:3167–3174.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Li ZM, Zhang HY, Wang YX and Wang WB:
MicroRNA-137 is downregulated in human osteosarcoma and regulates
cell proliferation and migration through targeting FXYD6. J Drug
Target. 24:102–110. 2016. View Article : Google Scholar : PubMed/NCBI
|