Introduction
Osteoporosis (OP) is the most common type of
skeletal disease, regardless of sex (1). As a progressive skeletal disease, OP is
characterized by low bone mass and damaged bone structure, which
can result in rarefaction of bone and an increased risk of fracture
(2). In patients with fractures
caused by OP, hip fractures are commonly observed (1). Previous evidence indicated that
OP-induced factures are typically associated with higher fatality
rates and can limit the activity of the patient, thereby increasing
medical burden in China (1).
According to the latest global research, it has been reported that
menopausal women have the highest risk of OP (1). As age increases, the incidence rate of
hip fracture in women aged between 55 and 59 is 34%, which is
almost 5-fold greater (6.6%) in those aged >85 (3). Bone metabolism relies on the regulation
between the osteoblasts and osteoclasts. In patients with OP,
osteoclasts, which originate from the mononuclear macrophages of
bone marrow hematopoietic stem cells, can enhance the bone
absorption (4). Thus, research on
the mononuclear cells is of great significance in enhancing the
understanding on the mechanism and improving the treatment of
OP.
Long non-coding (lnc)RNAs have a transcriptional
length of >200 nucleotides and can regulate gene expression on
various levels, including epigenetic, transcriptional and
post-transcriptional regulation, without encoding any proteins
(5–12). LncRNAs have become an area of
interest in research, and it has been revealed that lncRNAs not
only exist in the blood of patients with tumors, but are stably
present, according to some reports, in the plasma or serum of some
human non-neoplastic diseases (6).
Hence, specific lncRNAs have been widely used in the diagnosis and
treatment of various diseases, and may have potential clinical
application value (13,14). This value provides a new direction in
studying the etiology, pathogenesis and prognosis of associated
diseases. In recent years, circulating lncRNAs have been identified
to serve an important role in diseases, including cardiovascular
disease (15), refractory asthma
(16), acute kidney injury (13), diabetes mellitus, major depressive
disorder (17) and Hashimoto's
thyroiditis (18). Nevertheless, a
few studies have focused on the effects of circulating lncRNAs in
OP.
In bone tissues, the activity of osteoblasts and
osteoclasts is precisely coordinated (9,19).
During bone remodeling, osteoclasts uptake the bone and then
recruit bone marrow mesenchymal stem cells (BM-MSCs) for subsequent
differentiation and bone formation (19). Under certain conditions, BM-MSCs
possess the potential to differentiate into different types of
cells, including osteoblasts, chondrocytes, adipocytes, muscle
cells and tendon cells (20), which
can provide a cell source for bone growth and bone repair.
Osteoblasts are the primary functional cells in bone formation
(20). They are distributed on the
surface of bone, which can produce the organic components of the
extracellular matrix as well as regulate the synthesis, secretion
and mineralization of bone matrix (19,20).
Notably, osteoblasts can be identified using the following indexes:
Alkaline phosphatase (ALP), bridge proteins, bone morphogenetic
proteins, and osteocalcin and bone sialoproteins (19,20).
Furthermore, osteoclasts are distributed in the bone resorption
lacuna, and are responsible for bone resorption (19,20).
There is a large number of experimental evidence
suggesting that BM-MSCs are associated with OP. The present study
aimed to assess the biological function of key long non-coding
(lnc)RNAs in the occurrence and development of osteoporosis, and to
further investigate its underlying molecular mechanism.
Materials and methods
Data collection
Data retrieval was performed with the Gene
Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/geo/index.cgi) using
‘osteoporosis’ and ‘HG-133A’ as key words. A total of 10 datasets
were identified. Since detection with the probe in HG-133A could
identify various lncRNAs, the HG-133A platform was selected as the
alternative platform in the present study. In the present study, a
total of three datasets, GSE56815, GSE7158 and GSE2208, were
acquired, in which GSE2208 was eliminated as this did not include
the original CEL files. In the GSE56815 dataset, there were 40
microarrays in monocytes of normal bone and 40 of OP, whereas in
the GSE7158 dataset there were 14 of normal bone and 14 of OP.
Inclusion and exclusion criteria of OP was based on a previous
guideline (21). Serum and blood
cells of 40 cases were collected (14 males, 26 females; age range,
43–72 years) from patients admitted to The First Affiliated
Hospital of Nanjing Medical University (Nanjing, Jiangsu, China)
between July 2017 and February 2018. Written informed consent was
obtained from all the participants. One tube of venous blood of all
patients enrolled was collected immediately with an
anticoagulant-free red blood collection tube following admission.
The present study was approved by the Ethical Committee of Nanjing
Medical University and patient consent was obtained prior to
enrollment. The tube was placed in the refrigerator at 4°C for 30
min, centrifuged at 2,500 × g for 15 min at room temperature, and
then the upper serum was collected and placed in refrigerator at
20°C.
RankProd analysis of differential
lncRNAs
To comprehensively analyze the differentially
expressed genes in the two datasets, analysis was performed with
RankProd R package (22). Firstly,
datasets were merged in accordance of the Combat method with the
Slicomerging R package to eliminate the difference between the two
groups (23). Proportion of false
positives (pfp) of differentially expressed genes were identified
using 1,000 permutations, and the list of upregulated or
downregulated probes was determined on the basis of pfp
(pfp<0.01) and fold change (FC) value (FC>1, upregulated;
FC<1, downregulated).
Isolation, culture and differentiation
of BM-MSCs
Primary BM-MSCs were collected from 10 3-week-old
female Sprague-Dawley rats (weight range, 50–60 g) and the bone
marrow was flushed from the femur and tibia of rats using a 5-ml
syringe. All rats were housed in a temperature controlled room
(21±2°C), under a 12 h light/dark cycle, with free access to water
and food. BM-MSCs were cultured in Dulbecco's modified Eagle's
medium (DMEM; Gibco; Thermo Fisher Scientific, Inc., Waltham, MA,
USA) supplemented with 10% fetal bovine serum (FBS; Gibco; Thermo
Fisher Scientific, Inc.), 1% L-glutamine (Gibco; Thermo Fisher
Scientific, Inc.), 1% penicillin and 1% HEPES (Gibco; Thermo Fisher
Scientific, Inc.) in an atmosphere containing 5% CO2 at
37°C. Medium was changed every 2 days to remove unattached cells.
Following 5 days of incubation, the primary cells were trypsinized
and passaged. Generation 3–5 of BM-MSCs were used in the present
study. Notably, 10% FBS, 10 nmol/l dexamethasone (Sigma-Aldrich;
Merck KGaA, Darmstadt, Germany), 10 mmol/l β-glycerophosphate
(Sigma-Aldrich; Merck KGaA), 50 µg/ml ascorbic acid (Sigma-Aldrich;
Merck KGaA), 1% L-glucose, 1% penicillin-streptomycin and 1% HEPES
were added to a high-glucose DMEM (DMEM-HG; Gibco; Thermo Fisher
Scientific, Inc.) in order to make up the osteogenic medium and
induce osteogenic differentiation of MSCs.
Cell surface markers of MSCs
MSC surface markers of cells isolated from rats were
detected using flow cytometry with the following antibodies:
Fluorescein isothiocyanate (FITC) hamster anti-rat cluster of
differentiation (CD)29 (1:100; cat. no. 564131), FITC mouse
anti-rat CD44H (1:100; cat. no. 550974), FITC mouse anti-rat CD45
(1:100; cat. no. 561587), FITC mouse anti-rat CD90 (1:100; cat no.
554894). All antibodies were purchased from BD Biosciences
(Franklin Lakes, NJ, USA) and were incubated at 20°C for 1 h. FITC
Goat anti-Mouse Immunoglobulin G (H+L) Cross-Adsorbed Secondary
Antibodies was purchased from Thermo Fisher Scientific, Inc.
(1:1,000; cat. no. 31541) and incubated at 20°C for 30 min. Human
TruStain FcX™ (BioLegend, Inc., San Diego, CA, USA; cat. no.
422301) was used for blocking at room temperature for 5 min. Flow
cytometer (FACSCalibur; BD Bioscience) was used for analysis. Data
were obtained and analyzed using CellQuest professional software
(Version 3.3; Becton, Dickinson and Company; Franklin Lakes, NJ,
USA). Generation three BM-MSCs were suspended in PBS, incubated
with 0.5 µg/ml of each antibody. Unstained MSCs served as
controls.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
Total RNA was extracted from BM-MSCs using TRIzol
reagent (Invitrogen; Thermo Fisher Scientific, Inc.) according to
the manufacturer's instructions. Total RNA concentration was
measured using a NanoDrop 2000 (Thermo Fisher Scientific, Inc.) and
1 µg of total RNA was reverse transcribed into cDNA using a reverse
transcription kit (Takara Bio, Inc., Otsu, Japan). RT-qPCR was
performed on an ABI7900 Fast Real-Time System (Applied Biosystems;
Thermo Fisher Scientific, Inc.) using the SYBR Premix Ex Taq kit
(Takara Bio, Inc.). The thermocycling conditions were as follows:
pre-denaturation at 95°C for 5 min, denaturation at 95°C for 30
sec, annealing at 60°C for 45 sec, extension at 72°C for 3 min,
with 35 cycles and extension at 72°C for 5 min. PCR products were
then stored at 4°C. The primers utilized in PCR were as follows:
XIST forward, 5′-AGGCTGGCTGGAATAAAGG-3′ and reverse,
5′-TATGAAAAGGGAGGCGTGGT-3′; ALP forward, 5′-GGGACTGGTACTCGGACAAT-3′
and reverse, 5′-GGCCTTCTCATCCAGTTCAT-3′; Bglap forward,
5′-CATGAGGACCCTCTCTCTGC-3′ and reverse, 5′-TGGACATGAAGGCTTTGTCA-3′;
Runx2 forward, 5′-GCACCCAGCCCATAATAGA-3′ and reverse,
5′-TTGGAGCAAGGAGAACCC-3′; GAPDH forward,
5′-GGCACAGTCAAGGCTGAGAATG-3′ and reverse,
5′-ATGGTGGTGAAGACGCCAGTA-3′. The 2−ΔΔCq method was used.
GAPDH was used as a standard control for data analysis (24).
Western blot analysis
The proteins were extracted from BM-BMSCs.
Subsequently, the protein concentration was detected using the BCA
method (Pierce; Thermo Fisher Scientific, Inc.) and adjusted using
the radioimmunoprecipitation assay (Beyotime Institute of
Biotechnology, Shanghai, China). A total of 30 µg protein were
loaded per lane for electrophoresis. Following SDS-PAGE (5%
concentrated gel and 10% separation gel) the sample was transferred
to a polyvinylidene fluoride (PVDF) membrane (EMD Millipore,
Billerica, MA, USA), which was then soaked in 5% skimmed milk at
room temperature for 1 h. The PVDF membrane was subsequently cut
and respectively incubated with the following antibodies at 4°C
overnight: Runx2 (1:500; cat. no. ab76956); rabbit polyclonal ALP
(1:500; cat. no. ab83259) and rabbit polyclonal GAPDH (1:500; cat.
no. ab37168; all, Abcam, Cambridge, UK) The following day, the
bands were washed three times with PBS (for 10 min each time).
Subsequently, the bands were incubated with the following secondary
antibodies for 1 h at 20°C: Horseradish peroxidase conjugated goat
anti-rabbit Immunoglobulin G (1:2,000; cat. no. ab6721; Abcam).
Samples were then washed three times with PBS (for 10 min each
time). Lastly, ECL solution (Merck KGaA) was applied to bands in
order to detect the protein expression levels. Protein bands were
analyzed using Image J software (Version 1.38; National Institutes
of Health, Bethesda, MD, USA). GAPDH served as the internal
control.
Cell transfection
BM-MSCs were resuspended in antibiotic-free DMEM and
re-seeded into 6-well plates at a density of 3×105
cells/well. BM-MSCs were incubated for 18–24 h and transfected
using a Lipofectamine 2000 kit (Sigma-Aldrich; Merck KGaA).
Liposomes, si-NC, si- X-inactive specific transcript (XIST),
pcDNA-NC and pcDNA-XIST were all purchased from Shanghai Genechem
(Shanghai, China). Detailed cell transfection was performed
according to the instructions. siRNA at the concentration of 50 nM
was added to each well and then incubated for 48 h. The sequences
of the three si-XISTs were as follows: si-XIST 1#,
5′-GGCCTGTTATGTGTGTGATTATATT-3′; si-XIST 2#,
5′-GCCAACTGTCTGCTTAAGAAA-3′; and si-XIST 3#,
5′-GCTGCTAGTTTCCCAATGATA-3′.
Alkaline phosphatase (ALP)
staining
ALP staining was performed using the ALP detection
kit (Sigma Aldrich; Merck KGaA) according to the manufacturer's
instructions. MSCs were seeded in 24-well plates at a density of
7×10 4 cells/well. Differentiation into osteoblasts was induced by
osteogenic induction medium at a cell density of 60%. Following 10
days of induction, the cells were fixed with 4% formaldehyde and 5%
citrate in acetone at room temperature for 30 sec. The fixed cells
were further washed with PBS and incubated with 0.2% naphthol AS-BI
and 0.2% diazonium salt for 15 min at room temperature. Once the
working solution was discarded, the cells were washed with PBS
again. Images were captured using a light microscope at a
magnification of ×4.
Alizarin Red S (ARS) staining
The morphology of BM-BMSCs was observed with a
microscope. Once the medium was discarded, the cells were washed
twice with PBS and then fixed with 1 ml 4% paraformaldehyde at 20°C
for 20 min. Cells were then washed twice with PBS. Following this,
cells were stained with 1 ml ARS at room temperature for 10 min.
The cell samples were washed twice with PBS for 10 min each and
images were captured with an optical microscope (magnification,
×4).
Statistical analysis
Statistical analyses were performed with SPSS
software (v22.0; IBM Corp., Armonk, NY, USA). In addition, GraphPad
Prism software 5.0 (GraphPad Software, Inc., La Jolla, CA, USA) was
used for image editing. Comparisons between multiple groups were
performed using one-way analysis of variance followed by a post hoc
test (Fisher's Least Significant Difference). P<0.05 was
considered to indicate a statistically significant difference.
Results
Alterations in the expression levels
of 6 lncRNAs in patients with OP
Data were merged using the combat method and the
Slicomerging R package, and the differentially expressed genes were
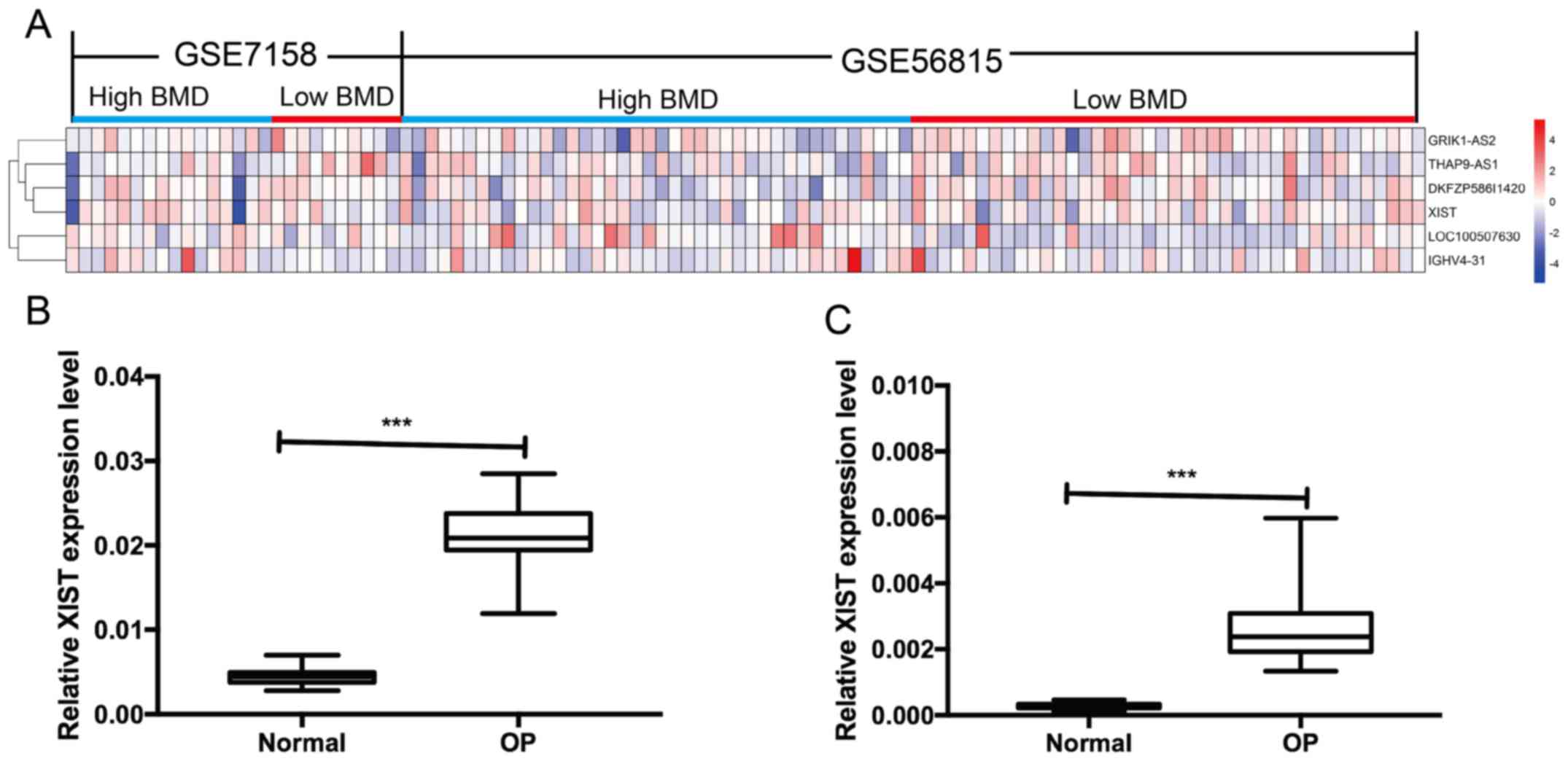
calculated in accordance with the RankProd method. The results
indicated that there were 1,060 downregulated genes and 765
upregulated genes, in which there were 4 upregulated lncRNAs and 2
downregulated lncRNAs (Fig. 1A and
Table I), including LOC100507630,
IGHV4-31, GRIK1-AS2, DKFZP586I1420, THAP9-AS1 and XIST. XIST is an
important lncRNA that has been previously reported in a variety of
diseases, but not in OP (25,26).
Therefore, XIST was chosen as a candidate lncRNA to explore its
associated mechanism in OP. The expression of lncRNA XIST in serum
and peripheral blood monocytes was detected by RT-qPCR in patients
with OP and normal subjects. The results indicated that the
expression level of lncRNA XIST in the serum and peripheral blood
monocytes of patients with OP was significantly increased compared
with that in normal subjects (Fig. 1B
and C).
 | Table I.Significantly dysregulated probes
identified by RankProd in osteoporosis. |
Table I.
Significantly dysregulated probes
identified by RankProd in osteoporosis.
| ID | Symbol |
FC_class1.class2 | pfp | P-value |
|---|
| 207476_at | LOC100507630 | 0.886603422 | 0 | <0.001 |
| 217281_x_at | IGHV4-31 | 0.823655383 |
1.00×10−04 | <0.001 |
| 210818_s_at | GRIK1-AS2 | 1.091941472 | 0 | <0.001 |
| 213546_at | DKFZP586I1420 | 1.107174491 |
2.00×10−04 | <0.001 |
| 215009_s_at | THAP9-AS1 | 1.153535587 |
4.00×10−04 | <0.001 |
| 221728_x_at | XIST | 1.045369015 | 0.0029 |
1.00×10−04 |
BM-MSCs purity
Primary BM-MSCs were isolated from rat bone marrow
and passaged successfully. BM-MSCs adhered to plastic cultured
dishes and exhibited a typical spindle-shaped morphology (Fig. 2A). Following 14 days of osteogenic
induction, a large number of mineralized nodules were observed, as
demonstrated by ARS staining (Fig.
2B). The immunophenotype was identified by flow cytometry. As
demonstrated in Fig. 2C, the results
indicated that the isolated cells were positive for
mesenchymal-associated markers CD29 (99.82%), CD44 (99.73%) and
CD90 (99.68%). However, few cells were positive for CD45 (0.01%). A
large number of calcified nodules were identified by ARS staining
following 14 days of culture in osteogenic medium. However, this
was not observed in the control group, which demonstrated the
ability of the MSCs to differentiate into osteoblasts. In order to
further verify that BM-MSCs could be differentiated into
osteoblasts, the osteogenic marker genes ALP, runt related
transcription factor 2 (Runx2) and bone γ-carboxyglutamic
acid-containing protein (Bglap) were detected in cells cultured for
1, 3, 7 and 14 days. On the 3rd and 7th day, the expression levels
of the marker genes mentioned above were significantly increased
(Fig. 2D). By contrast, the
expression level of XIST in the cells was significantly decreased
in a time-dependent manner (Fig.
2E).
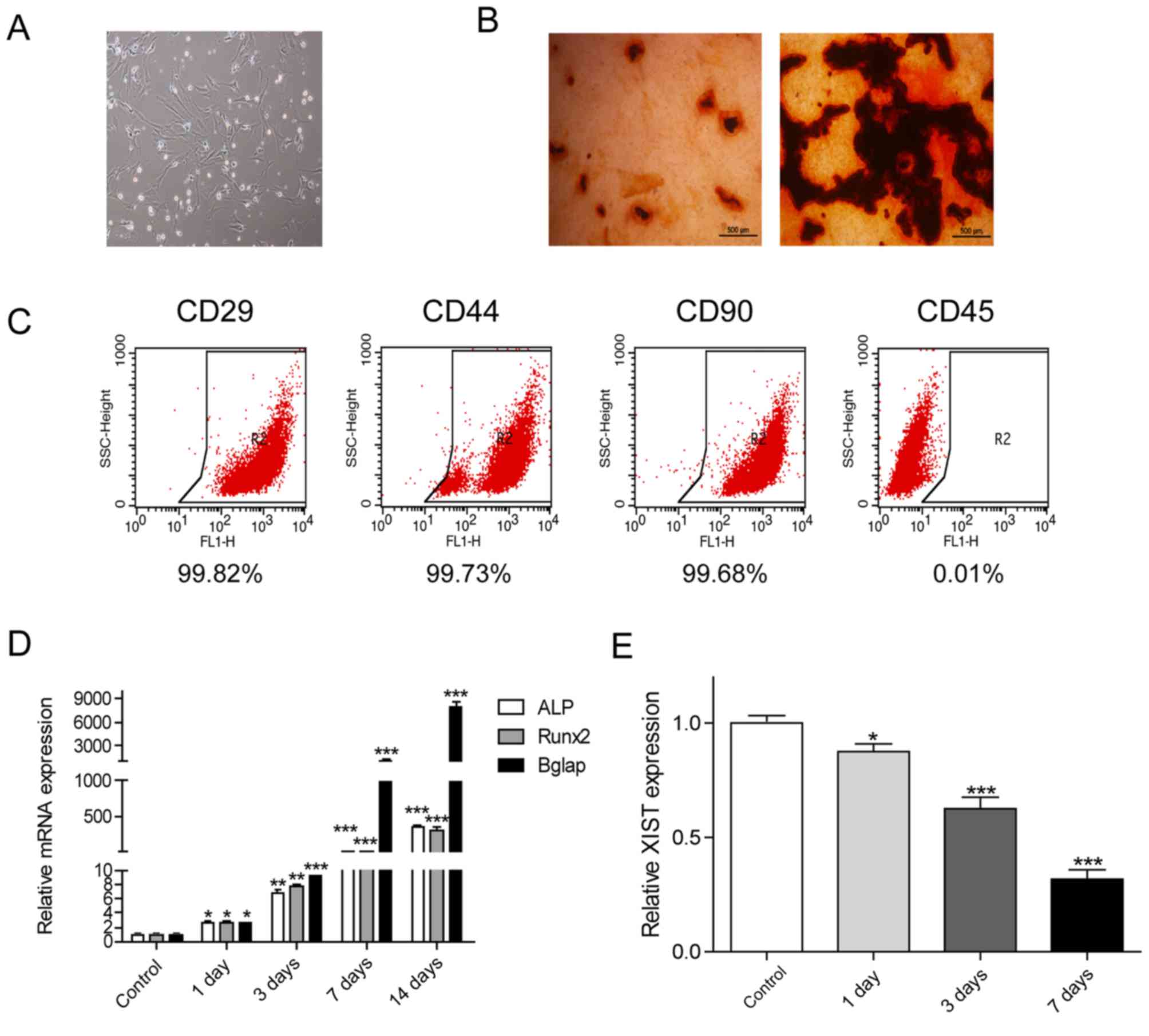 | Figure 2.Phenotypic identification of bone
marrow mesenchymal stem cells. (A) The shape of MSCs on the 4th
day. A typical long fusiform shape was observed in the MSCs
(magnification, ×40). (B) Following cultured in osteogenic
induction medium for 14 days, MSCs exhibited more mineralized
nodules according to Alizarin Red S staining, whereas the control
group did not. (C) Identification of MSC-specific surface antigens,
including positive identification of CD29, CD44 and CD90, and
negative identification of CD45 using flow cytometry. (D) The
expression levels of osteoblast marker genes ALP, Runx2 and Bglap
on different days of induction were significantly increased most
obviously on the 3rd and 7th day of induction. (E) The expression
of XIST decreased in a time-dependent manner. *P<0.05,
**P<0.01 and ***P<0.001 vs. the control group. MSCs,
mesenchymal stem cells; XIST, X-inactive specific transcript; ALP,
alkaline phosphatase; Runx2, runt related transcription factor 2;
Bglap, bone γ-carboxyglutamic acid-containing protein; CD, cluster
of differentiation. |
XIST inhibits osteogenic
differentiation of BM-MSCs
To investigate the effect of XIST on the expression
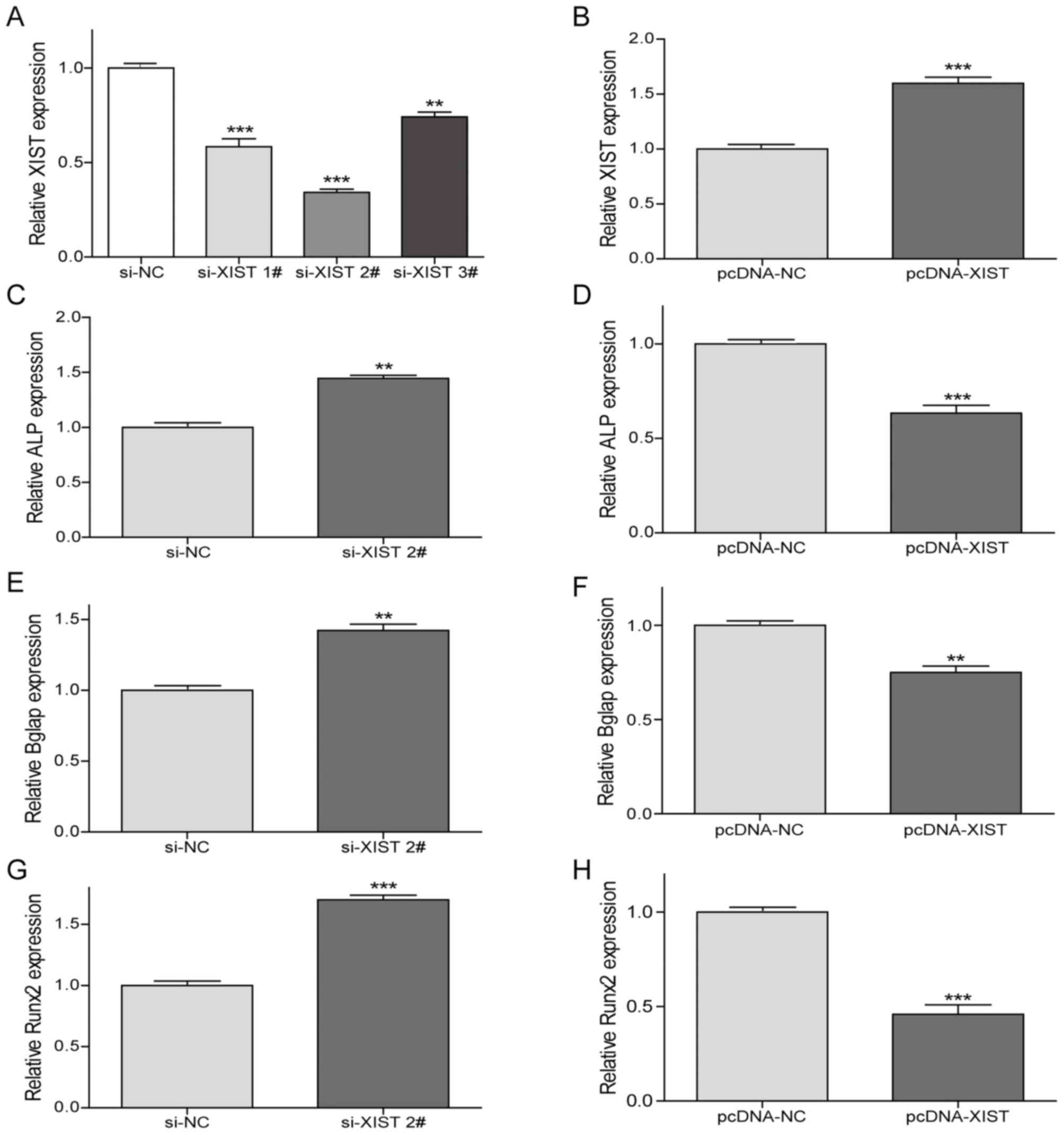
of the osteogenic marker genes, RT-qPCR was performed (Fig. 3). Following si-XIST transfection, the
expression level of XIST in BM-MSCs was significantly reduced, and
the most significant decrease was induced by si-XIST 2# (Fig. 3A). Subsequently, si-XIST2 was
selected for the remaining interference experiments. Notably, XIST
expression levels were significantly increased with pcDNA-XIST
transfected in BM-MSCs, which confirmed the transfection efficiency
(Fig. 3B).
The expression levels of ALP, Bglap and Runx2 genes
were significantly increased by si-XIST 2# (Fig. 3C, E and G). Conversely, the
expression levels of the above osteoblast-associated genes were
significantly reduced following the overexpression of XIST
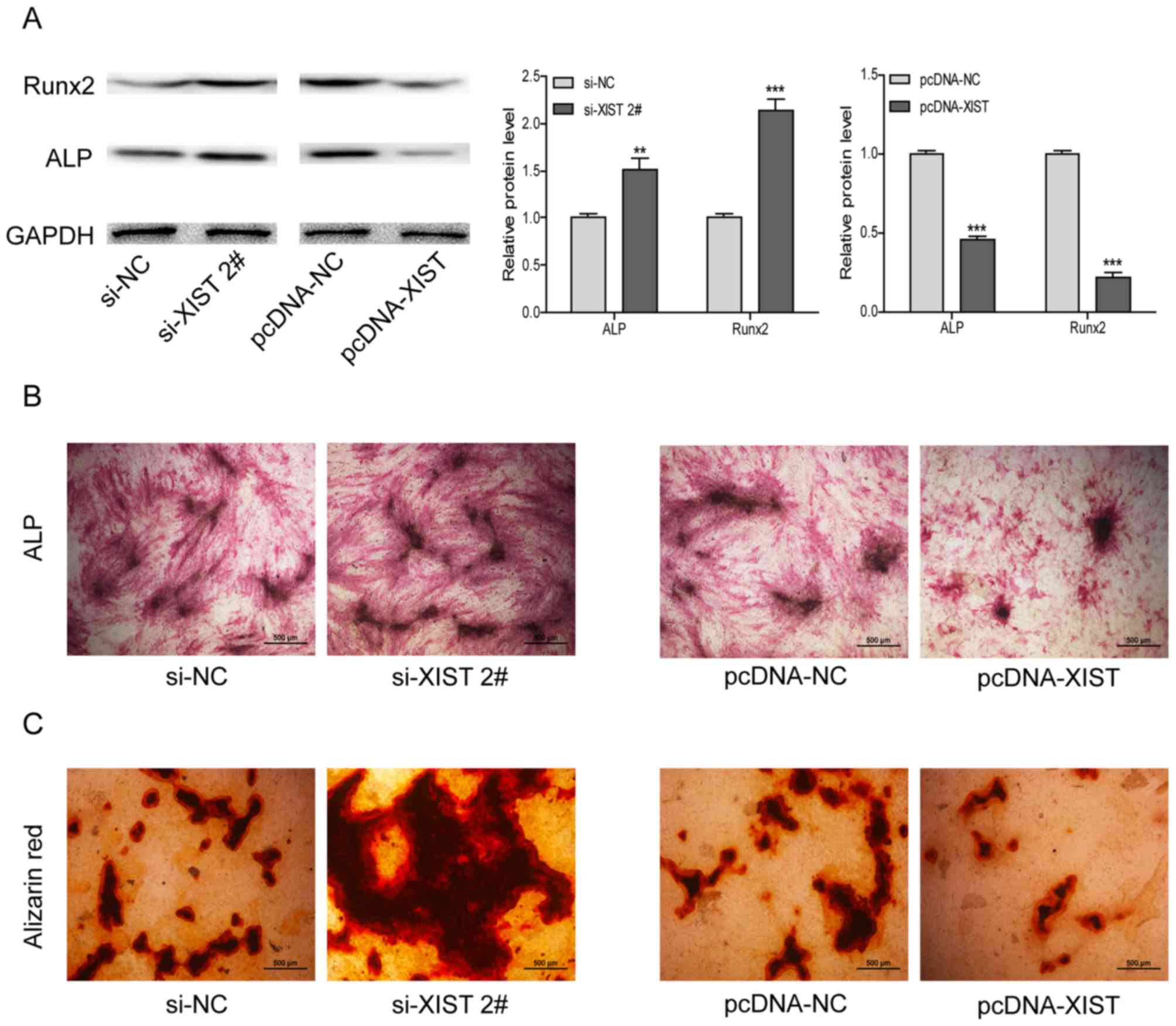
(Fig. 3D, F and H). Correspondingly,
the expression trends of ALP and Runx2 proteins were consistent
with their relative RNAs (Fig. 4A).
Furthermore, ALP staining was enhanced following XIST interference
and reduced with XIST overexpression (Fig. 4B).
Following interfering with XIST, the number of
mineralized nodules was increased, as indicated with the increased
ARS staining. Conversely, smaller and fewer nodules were observed
when XIST was overexpressed (Fig.
4C). These results suggested that overexpression of XIST could
inhibit osteogenic differentiation of BM-MSCs.
Discussion
In a normal cycle of bone metabolism, bone
absorption and formation are in a dynamic balance. Bone absorption
of osteoclasts and bone formation of osteoblasts are coupled for
the continuous reconstruction of bones. However, once the coupled
balance is broken, it can result in an increase in bone absorption
of osteoclasts, or a decrease in bone formation of osteoblasts,
thereby leading to the former process being greater than the
latter, and giving rise to OP (27–29).
XIST, a lncRNA indispensable for silencing the
transcription of chromosome X in mammals, is critical for the
deactivation of chromosome X (25).
In recent years, more studies have demonstrated that XIST is
correlated with the development and progression of tumors (30–32). In
glioblastoma, XIST knockdown can inhibit the progression of tumors
by reducing cell proliferation, invasion and migration and inducing
cell apoptosis (30). In patients
with gastric cancer it has been reported that the high expression
of XIST is associated with an increase in tumor volume, lymphatic
metastasis, distant metastasis and tumor, node, metastasis staging
(33,34). Furthermore, lncRNA maternally
expressed 3 and differentiation antagonizing non-protein coding RNA
are also critical for bones; however, no study has reported on the
role of XIST (35,36).
Bone is a dynamic tissue that is in continuously
modified (19). It is primarily
composed of two types of cells, osteoblasts and osteoclasts; the
former promote bone formation, while the latter promote bone
resorption (19). These two types of
bone cells are at equilibrium in normal bone. However, once this
balance is disrupted, a variety of bone metabolic diseases can be
triggered (37). Bglap and Runx2,
two important markers of bone formation, are tightly bound to bone
hydroxyapatite and calcium and are indirectly involved in
osteoblast activation during osteogenesis (19). ALP is also an important marker of
early osteogenic differentiation of BM-MSCs, which serves a key
role in the in vitro calcification of BM-MSCs (38–40).
Therefore, early osteogenic differentiation of BM-MSCs can be
observed through the detection of ALP synthesis (41). Calcium nodules are formed when
mineralization occurs in BM-MSCs. The formation of calcium nodules
is a sign of maturation of cells (19). The degree of osteogenic
differentiation of BM-MSCs can be analyzed by detecting the
formation of calcium nodules in BM-MSCs (42). In the present study, RankProd
analysis with the retrieval of databases demonstrated that XIST was
highly expressed in patients with OP. It was indicated that
overexpression of XIST significantly decreased the gene and protein
expression levels of ALP, Bglap and Runx2, whereas ALP staining and
calcification in ARS staining were reduced. These results indicated
that XIST inhibited osteogenic differentiation of BM-MSCs, which
was consistent with the data analysis.
The present study had some limitations. Firstly, the
present study only demonstrated the association between XIST and
BM-MSCs; however, the detailed molecular mechanism has not been
fully elucidated. Further research should be performed to explore
how XIST influences the differentiation of BM-MSCs and what types
of molecules or signaling pathways takes part in this effect.
Notably, si-XIST and pcDNA-XIST transgenic mice will be used in
future work to investigate the development of OP, and to analyze
the expression of XIST and osteoblast identification indexes.
In conclusion, the present findings suggest that
XIST is highly expressed in patients with OP, and XIST is able to
inhibit osteogenic differentiation of BM-MSCs.
Acknowledgements
Not applicable.
Funding
The current study was supported by the National
Natural Science Foundation of China (grant no. 81802149) and
Science and Technology Development Fund of NJMU (grant no.
2017NJMU130).
Availability of data and materials
All data generated or analyzed during the present
study are included in this published article.
Authors' contributions
XCh, SZ and BL designed the study and performed the
experiments. XCa, LY and WW established the animal models. DG, JY
and ZY collected the data. XCh and CJ analyzed the data. XCh and LY
prepared the manuscript. All authors read and approved the final
manuscript.
Ethics approval and consent to
participate
This study was approved by the Animal Ethics
Committee of Nanjing Medical University Animal Center. This study
was also approved by the Medical Ethics Committee of Nanjing First
Hospital (Nanjing, China) and patient consent was obtained.
Patient consent for publication
Written informed consent was obtained from all the
participants.
Competing interests
The authors declare no competing interests.
References
|
1
|
Cosman F, de Beur SJ, LeBoff MS, Lewiecki
EM, Tanner B, Randall S and Lindsay R: Erratum to: Clinician's
guide to prevention and treatment of osteoporosis. Osteoporos Int.
26:2045–2047. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Ge DW, Wang WW, Chen HT, Yang L and Cao
XJ: Functions of microRNAs in osteoporosis. Eur Rev Med Pharmacol
Sci. 21:4784–4789. 2017.PubMed/NCBI
|
|
3
|
Pfeilschifter J, Cooper C, Watts NB,
Flahive J, Saag KG, Adachi JD, Boonen S, Chapurlat R, Compston JE,
Díez-Pérez A, et al: Regional and age-related variations in the
proportions of hip fractures and major fractures among
postmenopausal women: The Global Longitudinal Study of Osteoporosis
in Women. Osteoporos Int. 23:2179–2188. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Wang WW, Yang L, Wu J, Gao C, Zhu YX,
Zhang D and Zhang HX: The function of miR-218 and miR-618 in
postmenopausal osteoporosis. Eur Rev Med Pharmacol Sci.
21:5534–5541. 2017.PubMed/NCBI
|
|
5
|
Court F, Baniol M, Hagege H, Petit JS,
Lelay-Taha MN, Carbonell F, Weber M, Cathala G and Forne T:
Long-range chromatin interactions at the mouse Igf2/H19 locus
reveal a novel paternally expressed long non-coding RNA. Nucleic
Acids Res. 39:5893–5906. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Garmire LX, Garmire DG, Huang W, Yao J,
Glass CK and Subramaniam S: A global clustering algorithm to
identify long intergenic non-coding RNA-with applications in mouse
macrophages. PLoS One. 6:e240512011. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Gibb EA, Vucic EA, Enfield KS, Stewart GL,
Lonergan KM, Kennett JY, Becker-Santos DD, MacAulay CE, Lam S,
Brown CJ and Lam WL: Human cancer long non-coding RNA
transcriptomes. PLoS One. 6:e259152011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Korostowski L, Raval A, Breuer G and Engel
N: Enhancer-driven chromatin interactions during development
promote escape from silencing by a long non-coding RNA. Epigenetics
Chromatin. 4:212011. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Muers M: RNA: Genome-wide views of long
non-coding RNAs. Nat Rev Genet. 12:7422011. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Saxena A and Carninci P: Long non-coding
RNA modifies chromatin: Epigenetic silencing by long non-coding
RNAs. Bioessays. 33:830–839. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Schorderet P and Duboule D: Structural and
functional differences in the long non-coding RNA hotair in mouse
and human. PLoS Genet. 7:e10020712011. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Yang Z, Zhou L, Wu LM, Lai MC, Xie HY,
Zhang F and Zheng SS: Overexpression of long non-coding RNA HOTAIR
predicts tumor recurrence in hepatocellular carcinoma patients
following liver transplantation. Ann Surg Oncol. 18:1243–1250.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Lorenzen JM, Schauerte C, Kielstein JT,
Hubner A, Martino F, Fiedler J, Gupta SK, Faulhaber-Walter R,
Kumarswamy R, Hafer C, et al: Circulating long noncoding RNATapSaki
is a predictor of mortality in critically ill patients with acute
kidney injury. Clin Chem. 61:191–201. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Kumarswamy R, Bauters C, Volkmann I, Maury
F, Fetisch J, Holzmann A, Lemesle G, de Groote P, Pinet F and Thum
T: Circulating long noncoding RNA, LIPCAR, predicts survival in
patients with heart failure. Circ Res. 114:1569–1575. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Yang KC, Yamada KA, Patel AY, Topkara VK,
George I, Cheema FH, Ewald GA, Mann DL and Nerbonne JM: Deep RNA
sequencing reveals dynamic regulation of myocardial noncoding RNAs
in failing human heart and remodeling with mechanical circulatory
support. Circulation. 129:1009–1021. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Orsmark-Pietras C, James A, Konradsen JR,
Nordlund B, Söderhäll C, Pulkkinen V, Pedroletti C, Daham K,
Kupczyk M, Dahlen B, et al: Transcriptome analysis reveals
upregulation of bitter taste receptors in severe asthmatics. Eur
Respir J. 42:65–78. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Liu Z, Li X, Sun N, Xu Y, Meng Y, Yang C,
Wang Y and Zhang K: Microarray profiling and co-expression network
analysis of circulating lncRNAs and mRNAs associated with major
depressive disorder. PLoS One. 9:e933882014. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Peng H, Liu Y, Tian J, Ma J, Tang X, Rui
K, Tian X, Mao C, Lu L, Xu H, et al: The long noncoding RNA
IFNG-AS1 promotes T helper type 1 cells response in patients with
hashimoto's thyroiditis. Sci Rep. 5:177022015. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Mundy GR and Elefteriou F: Boning up on
ephrin signaling. Cell. 126:441–443. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Tang Y, Wu X, Lei W, Pang L, Wan C, Shi Z,
Zhao L, Nagy TR, Peng X, Hu J, et al: TGF-beta1-induced migration
of bone mesenchymal stem cells couples bone resorption with
formation. Nat Med. 15:757–765. 2009. View
Article : Google Scholar : PubMed/NCBI
|
|
21
|
Soen S, Fukunaga M, Sugimoto T, Sone T,
Fujiwara S, Endo N, Gorai I, Shiraki M, Hagino H, Hosoi T, et al:
Diagnostic criteria for primary osteoporosis: Year 2012 revision. J
Bone Miner Metab. 31:247–257. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Breitling R, Armengaud P, Amtmann A and
Herzyk P: Rank products: A simple, yet powerful, new method to
detect differentially regulated genes in replicated microarray
experiments. FEBS Lett. 573:83–92. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Taminau J, Meganck S, Lazar C, Steenhoff
D, Coletta A, Molter C, Duque R, de Schaetzen V, Weiss Solís DY,
Bersini H and Nowé A: Unlocking the potential of publicly available
microarray data using inSilicoDb and inSilicoMerging R/Bioconductor
packages. BMC Bioinformatics. 13:3352012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Yue M, Ogawa A, Yamada N, Charles Richard
JL, Barski A and Ogawa Y: Xist RNA repeat E is essential for ASH2L
recruitment to the inactive X and regulates histone modifications
and escape gene expression. PLoS Genet. 13:e10068902017. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Li GL, Wu YX, Li YM and Li J: High
expression of long non-coding RNA XIST in osteosarcoma is
associated with cell proliferation and poor prognosis. Eur Rev Med
Pharmacol Sci. 21:2829–2834. 2017.PubMed/NCBI
|
|
27
|
Cheung AM, Papaioannou A and Morin S:
Osteoporosis Canada Scientific Advisory Council: Postmenopausal
osteoporosis. N Engl J Med. 374:20962016.PubMed/NCBI
|
|
28
|
Tsai JN, Uihlein AV, Lee H, Kumbhani R,
Siwila-Sackman E, McKay EA, Burnett-Bowie SA, Neer RM and Leder BZ:
Teriparatide and denosumab, alone or combined, in women with
postmenopausal osteoporosis: The DATA study randomised trial.
Lancet. 382:50–56. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Rachner TD, Khosla S and Hofbauer LC:
Osteoporosis: Now and the future. Lancet. 377:1276–1287. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Yao Y, Ma J, Xue Y, Wang P, Li Z, Liu J,
Chen L, Xi Z, Teng H, Wang Z, et al: Knockdown of long non-coding
RNA XIST exerts tumor-suppressive functions in human glioblastoma
stem cells by up-regulating miR-152. Cancer Lett. 359:75–86. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Tantai J, Hu D, Yang Y and Geng J:
Combined identification of long non-coding RNA XIST and HIF1A-AS1
in serum as an effective screening for non-small cell lung cancer.
Int J Clin Exp Pathol. 8:7887–7895. 2015.PubMed/NCBI
|
|
32
|
Sirchia SM, Tabano S, Monti L, Recalcati
MP, Gariboldi M, Grati FR, Porta G, Finelli P, Radice P and Miozzo
M: Misbehaviour of XIST RNA in breast cancer cells. PLoS One.
4:e55592009. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Ma L, Zhou Y, Luo X, Gao H, Deng X and
Jiang Y: Long non-coding RNA XIST promotes cell growth and invasion
through regulating miR-497/MACC1 axis in gastric cancer.
Oncotarget. 8:4125–4135. 2017.PubMed/NCBI
|
|
34
|
Chen DL, Ju HQ, Lu YX, Chen LZ, Zeng ZL,
Zhang DS, Luo HY, Wang F, Qiu MZ, Wang DS, et al: Long non-coding
RNA XIST regulates gastric cancer progression by acting as a
molecular sponge of miR-101 to modulate EZH2 expression. J Exp Clin
Cancer Res. 35:1422016. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Tong X, Gu PC, Xu SZ and Lin XJ: Long
non-coding RNA-DANCR in human circulating monocytes: A potential
biomarker associated with postmenopausal osteoporosis. Biosci
Biotechnol Biochem. 79:732–737. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Wang Q, Li Y and Zhang Y, Ma L, Lin L,
Meng J, Jiang L, Wang L, Zhou P and Zhang Y: LncRNA MEG3 inhibited
osteogenic differentiation of bone marrow mesenchymal stem cells
from postmenopausal osteoporosis by targeting miR-133a-3p. Biomed
Pharmacother. 89:1178–1186. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Pagani F, Francucci CM and Moro L: Markers
of bone turnover: Biochemical and clinical perspectives. J
Endocrinol Invest. 28 (10 Suppl):S8–S13. 2005.
|
|
38
|
Johansen JS, Riis BJ, Delmas PD and
Christiansen C: Plasma BGP: An indicator of spontaneous bone loss
and of the effect of oestrogen treatment in postmenopausal women.
Eur J Clin Invest. 18:191–195. 1988. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Enomoto H, Furuichi T, Zanma A, Yamana K,
Yoshida C, Sumitani S, Yamamoto H, Enomoto-Iwamoto M, Iwamoto M and
Komori T: Runx2 deficiency in chondrocytes causes adipogenic
changes in vitro. J Cell Sci. 117:417–425. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Bai Y, Yin G, Huang Z, Liao X, Chen X, Yao
Y and Pu X: Localized delivery of growth factors for angiogenesis
and bone formation in tissue engineering. Int Immunopharmacol.
16:214–223. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Golub EE, Harrison G, Taylor AG, Camper S
and Shapiro IM: The role of alkaline phosphatase in cartilage
mineralization. Bone Miner. 17:273–278. 1992. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Giannoudis PV, Jones E and Einhorn TA:
Fracture healing and bone repair. Injury. 42:549–550, 2011.vv.
View Article : Google Scholar : PubMed/NCBI
|