|
1
|
Qi P and Du X: The long non-coding RNAs, a
new cancer diagnostic and therapeutic gold mine. Mod Pathol.
26:155–165. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Amirkhah R, Schmitz U, Linnebacher M,
Wolkenhauer O and Farazmand A: MicroRNA-mRNA interactions in
colorectal cancer and their role in tumor progression. Genes
Chromosomes Cancer. 54:129–141. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Wang C, Hu DZ and Liu JZ: Identification
of critical TF-miRNA-mRNA regulation loops for colorectal cancer
metastasis. Genetics Mol Res. 14:5485–5495. 2015. View Article : Google Scholar
|
|
4
|
Zhang C, Wang C, Jia Z, Tong W, Liu D, He
C, Huang X and Xu W: Differentially expressed mRNAs, lncRNAs, and
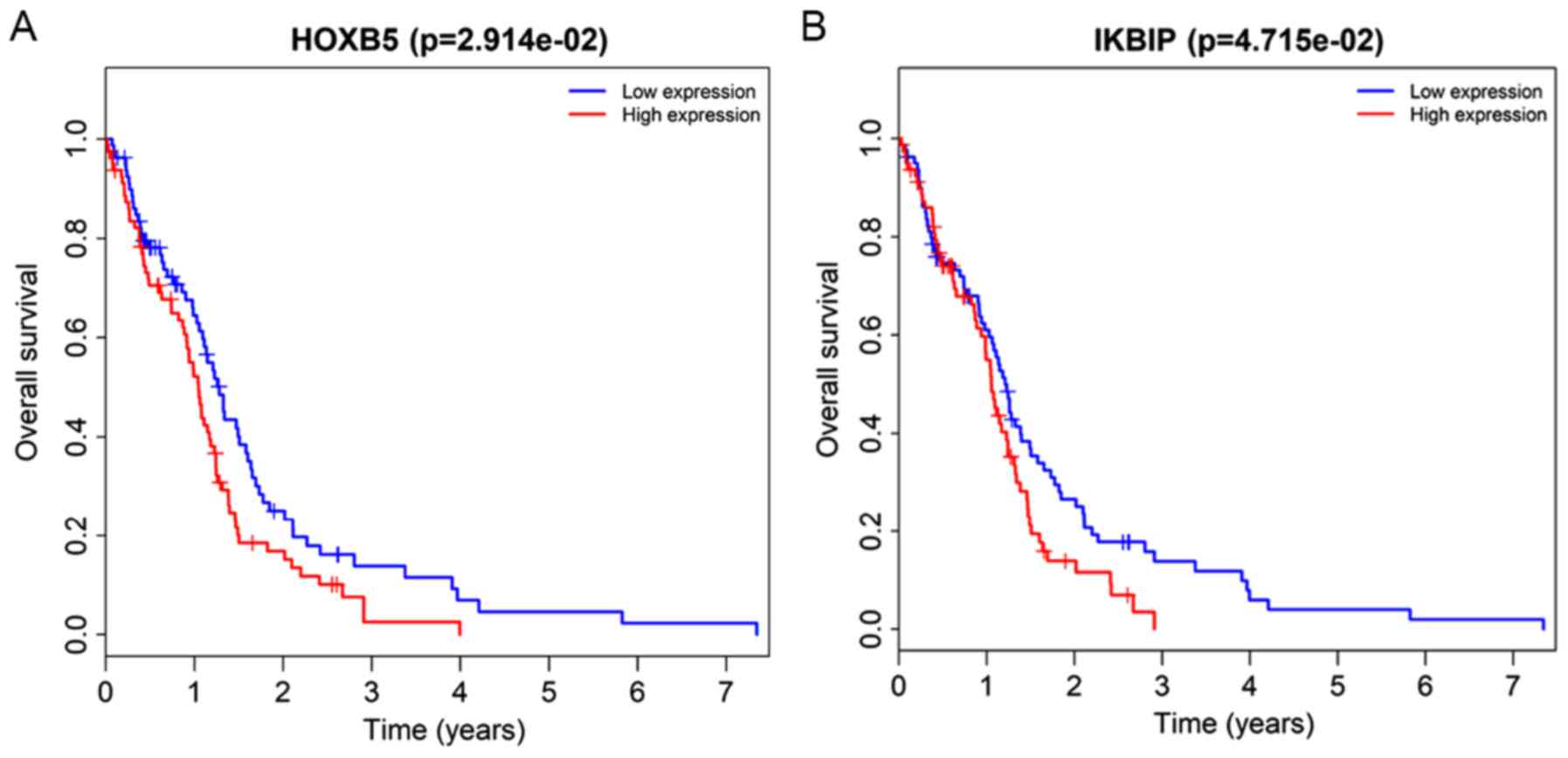
miRNAs with associated co-expression and ceRNA networks in
ankylosin spondylitis. Oncotarget. 8:113543–113557. 2017.PubMed/NCBI
|
|
5
|
Salmena L, Poliseno L, Tay Y, Kats L and
Pandolfi PP: A ceRNA hypothesis: The Rosetta stone of a hidden RNA
language? Cell. 146:353–358. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Liu K, Guo L, Guo Y, Zhou B, Li T, Yang H,
Yin R and Xi T: AEG-1 3-untranslated region functions as a ceRNA in
inducing epithelial-mesenchymal transition of human non-small cell
lung cancer by regulating miR-30a activity. Eur J Cell Biol.
94:22–31. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Peng W, Si S, Zhang Q, Li C, Zhao F, Wang
F, Yu J and Ma R: Long non-coding RNA MEG3 functions as a competing
endogenous RNA to regulate gastric cancer progression. J Exp Clin
Cancer Res. 34:792015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Johnsson P, Ackley A, Vidarsdottir L, Lui
WO, Corcoran M, Grandér D and Morris KV: A pseudogene
long-noncoding-RNA network regulates PTEN transcription and
translation in human cells. Nat Struct Mol Biol. 20:440–446. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Fan Q and Liu B: Comprehensive analysis of
a long noncoding RNA-associated competing endogenous RNA network in
colorectal cancer. Onco Targets Ther. 11:2453–2466. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Du Z, Fei T, Verhaak RG, Su Z, Zhang Y,
Brown M, Chen Y and Liu XS: Integrative genomic analyses reveal
clinically relevant long noncoding RNAs in human cancer. Nat Struct
Mol Biol. 20:908–913. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Zhang W, Zhang J, Hoadley K, Kushwaha D,
Ramakrishnan V, Li S, Kang C, You Y, Jiang C, Song SW, et al:
MiR-181d: A predictive glioblastoma biomarker that downregulates
MGMT expression. Neuro Oncol. 14:712–719. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Robinson MD, McCarthy DJ and Smyth GK:
EdgeR: A bioconductor package for differential expression analysis
of digital gene expression data. Bioinformatics. 26:139–140. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Jeggari A, Marks DS and Larsson E:
MiRcode: A map of putative microRNA target sites in the long
non-coding transcriptome. Bioinformatics. 28:2062–2063. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Agarwal V, Bell GW, Nam JW and Bartel DP:
Predicting effective microRNA target sites in mammalian mRNAs.
Elife. 4:2015. View Article : Google Scholar
|
|
15
|
Chou CH, Chang NW, Shrestha S, Hsu SD, Lin
YL, Lee WH, Yang CD, Hong HC, Wei TY, Tu SJ, et al: MiRTarBase
2016: Updates to the experimentally validated miRNA-target
interactions database. Nucleic Acids Res. 44:D239–D247. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Wong N and Wang X: MiRDB: An online
resource for microRNA target prediction and functional annotations.
Nucleic Acids Res. 43:D146–D152. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Dennis G Jr, Sherman BT, Hosack DA, Yang
J, Gao W, Lane HC and Lempicki RA: DAVID: Database for annotation,
visualization, and integrated discovery. Genome Biol. 4:R602003.
View Article : Google Scholar
|
|
18
|
Yu G, Wang L-G, Han Y and He Q-Y:
ClusterProfiler: An R package for comparing biological themes among
gene clusters. OMICS. 16:284–287. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Roy S, Lahiri D, Maji T and Biswas J:
Recurrent glioblastoma: Where we stand. South Asian J Cancer.
4:163–173. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Ebert MS, Neilson JR and Sharp PA:
MicroRNA sponges: Competitive inhibitors of small RNAs in mammalian
cells. Nat Methods. 4:721–726. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Brown BD, Cantore A, Annoni A, Sergi LS,
Lombardo A, Della Valle P, DAngelo A and Naldini L: A
microRNA-regulated lentiviral vector mediates stable correction of
hemophilia B mice. Blood. 110:4144–4152. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zhang X, Zhang X, Hu S, Zheng M, Zhang J,
Zhao J, Zhang X, Yan B, Jia L, Zhao J, et al: Identification of
miRNA-7 by genome-wide analysis as a critical sensitizer for
TRAIL-induced apoptosis in glioblastoma cells. Nucleic Acids Res.
45:5930–5944. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Yu J, Cai X, He J, Zhao W, Wang Q and Liu
B: Microarray-based analysis of gene regulation by transcription
factors and microRNAs in glioma. Neurol Sci. 34:1283–1289. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Liang F, Yang M, Tong N, Fang J, Pan Y, Li
J and Zhang X: Identification of six key miRNAs associated with
breast cancer through screening large-scale microarray data. Oncol
Lett. 16:4159–4168. 2018.PubMed/NCBI
|
|
25
|
Li G, Huang M, Cai Y, Yang Y, Sun X and Ke
Y: Circ-U2AF1 promotes human glioma via derepressing
neuro-oncological ventral antigen 2 by sponging hsa-miR-7-5p. J
Cellular Physiol. 234:9144–9155. 2019. View Article : Google Scholar
|
|
26
|
Cao Y, Wang P, Ning S, Xiao W, Xiao B and
Li X: Identification of prognostic biomarkers in glioblastoma using
a long non-coding RNA-mediated, competitive endogenous RNA network.
Oncotarget. 5:41737–41747. 2016.
|
|
27
|
Lee JY, Kim JM, Jeong DS and Kim MH:
Transcriptional activation of EGFR by HOXB5 and its role in breast
cancer cell invasion. Biochem Biophys Res Commun. 18:2924–2930.
2018. View Article : Google Scholar
|
|
28
|
Xu H, Zhao H and Yu J: HOXB5 promotes
retinoblastoma cell migration and invasion via ERK1/2
pathway-mediated MMPs production. Am J Transl Res. 15:1703–1712.
2018.
|
|
29
|
Zhang B, Li N and Zhang H: Knockdown of
homeobox B5 (HOXB5) inhibits cell proliferation, migration, and
invasion in non-small cell lung cancer cells through inactivation
of the Wnt/β-catenin pathway. Oncol Res. 19:37–44. 2018. View Article : Google Scholar
|
|
30
|
Hofer-Warbinek R, Schmid JA, Mayer H,
Winsauer G, Orel L, Mueller B, Wiesner CH, Binder BR and de Martin
R: A highly conserved proapoptotic gene, IKIP, located next to the
APAF1 gene locus, is regulated by p53. Cell Death Differ.
11:1317–1325. 2004. View Article : Google Scholar : PubMed/NCBI
|