Introduction
Microsatellites (MS), short tandem repeat sequences,
are composed of 1–6 base pairs (bps). Approximately 1 million
microsatellite loci, mostly as (CA)n, are dispersed in
the introns and exons of the human genome. Simple DNA repeats are
prone to expansion/contraction via the formation of secondary
structures during DNA synthesis. Such structures inhibit
replication forks and create opportunities for template-primer
slippage, making these repeats unstable (1). MS instability (MSI) is characterized by
alterations in length within MS, resulting from mutational
inactivation or epigenetic silencing of DNA mismatch repair genes
[e.g. MutS homolog 1 (MSH1), MSH2, MSH3 and mutL homolog 1]. These
mutations include coding region frameshift mutations caused by MSI,
which may drive oncogenesis by inactivating tumor-suppressor genes
or disrupting other non-coding regulatory sequences (2). Furthermore, as compared with the number
of repeats in the germline genome, an abnormal number of repeats,
in ≥30% of the microsatellite loci examined is defined as
microsatellite instability-high (MSI-H). MSI-H is known to occur in
~10% of sporadic colorectal cancers (CRCs) and 3% hereditary CRCs
(3).
CRC is the second and third most commonly diagnosed
cancer type among females and males worldwide, respectively
(4). CRC is a heterogeneous disease
and may be divided into certain molecular subtypes. Molecular
changes that occur in CRC may be categorized into three major
groups: i) Chromosomal instability, ii) MSI and iii) CpG island
methylation phenotype that silences gene function with aberrant
hypermethylation (5). The following
three major applications have been developed for the MSI phenotype:
i) Genetic evaluation of Lynch syndrome; ii) predicting the
response to chemotherapy drugs and iii) predicting the prognosis of
CRC patients. Thus, the MSI status has an important role in the
study of CRC.
Loss of heterozygosity (LOH) is another form of MS
alteration, which may be caused by mutation in one allele of a
gene. Early studies have demonstrated that 81% of patients with
sporadic breast cancer and 93% of patients with sporadic cancer (10
types) exhibit missense mutation LOH (6,7). It was
reported that a high frequency of LOH coincided with mutant P53
protein stabilization (8).
The human TP53 gene is located on chromosome 17p and
comprises 11 exons and 10 introns (9). The p53 protein is a phospho protein
consisting of 393 amino acids. Upon DNA damage, activation of p53
leads to cell cycle arrest, enabling the cells to repair the
damaged DNA. Exon mutations in the TP53 gene are the most commonly
observed genetic alterations in CRC with a prevalence of 50–70%
amongst CRC cases (10). Loss of
function of mutant p53 is a critical event in the progression of
CRC. Such mutations, which provide clues about the mechanisms of
genetic damage, tend to be differentially associated with other
cancer-associated genetic alterations and have prognostic and
clinical relevance (11).
Although the association of MSI and clinical
features of patients with CRC has been widely investigated,
differences between MSI and LOH alterations of a single MS in CRC
have not been previously addressed, to the best of our knowledge,
particularly the MS in the tumor suppressor gene TP53. Furthermore,
the associations between MS alterations (MSI and LOH) in TP53
introns and mutations in TP53 exons remain elusive. In the present
study, patients were stratified using different MS statuses,
including MSI, LOH and MS-stable (MSS) on TP53 intron and the
mutational profiles of exonsin the TP53 gene were evaluated.
Furthermore, their association with clinicopathological
characteristics in CRC was also explored. The present results
revealed that MSI alterations in TP53 introns are a valuable
predictive marker for the tumor-nodes-metastasis (TNM) stage of CRC
and LOH alteration may be a useful marker for the TP53 exon
mutation status.
Materials and methods
Patients
CRC samples were collected from the Clinical Data
and Biobank Resource of Beijing Friendship Hospital (a specimen
bank; Beijing, China) between November 2016 and November 2018. The
establishment of the specimen bank requires informed consent from
the patients, and therefore, ethical approval was obtained from the
ethics committee prior to the start of the study. According to the
TNM system classification of the American Joint Committee on Cancer
(12), specimens of CRC at stage
II/III were selected and the principal inclusion criteria were as
follows: Histologically proven papillary/tubular adenocarcinoma,
signet ring carcinoma and mucinous carcinoma of the colon or
rectum. Specimens were collected from fresh tumors and matched
normal tissues for the genetic analysis of MSI and LOH in a
specific MS (TP53ALU). A total of 512 specimens from 256 CRC
patients were stored at −80°C and analyzed. Relevant clinical data
were collected from the patients' medical charts. Vital status and
cause of death were obtained from medical records, tumor registry
correspondence or death certificate. The present study was approved
by the institutional review board of the Beijing Friendship
Hospital (Beijing, China).
Genomic DNA extraction
Genomic DNA was extracted from 256 pairs of CRC and
their matched normal tissues using a standard phenol-chloroform
extraction and ethanol precipitation method, as previously
described (13). DNA was quantified
using the absorbance ratio at 260/280 nm measured with a microplate
absorbance reader (Bio-Rad 680; Bio-Rad Laboratories, Inc.) and
then further analyzed by agarose gel electrophoresis. DNA samples
were diluted to a concentration of 50 ng/µl and stored at
−80°C.
MS analysis
An MS locus in intron 1 of the tumor suppressor gene
TP53 (referred to as TP53ALU) with the repeat unit
(AAAAT)8 was assessed. The sequences of the primers
designed were as follows: 5′-GGCAATAAGAGCTGAGACTCC-3′ (sense) and
5′-GACAAAACATCCCCTACCAAA-3′ (anti-sense). The forward primer of the
locus was labeled at the 5′ end with a fluorescent marker (labeled
using 6-carboxyfluorescein) for later use in short tandem repeat
(STR) scanning. The PCR amplification system contained 2 µl 10X
buffer, 125 µmol/l dNTP (4X), 0.5 µmol/l of each primer, 1.0 units
of Taq DNA polymerase, 1.5 mmol/l MgCl2 and 100 ng
template DNA. PCR was performed under the following conditions:
Denaturation at 94°C for 5 min; 35 cycles of denaturation at 94°C
for 30 sec, annealing at 62°C for 30 sec and extension at 72°C for
30 sec. This was followed by an extension step at 72°C for 7 min.
The PCR products were visualized on 2% agarose gels stained with
ethidium bromide and assessed with an ultraviolet transilluminator
(Gel Doc™ XR+; Bio-Rad Laboratories Inc.). Products amplified
successfully and correctly by PCR were stored at 4°C for subsequent
STR scanning. Fluorescently tagged PCR products were examined with
an ABI3730XL DNA Analyzer system (Perkin Elmer Biosystems).
GeneMarker version 1.75 (Tianyi Juiyuan Company) was used to
quantify each fluorescent PCR product. In order to confirm whether
the new alleles exist, PCR products were purified using an ABI
BigDye Terminator v3.1 Cycle Sequencing kit (Applied Biosystems;
Thermo Fisher Scientific, Inc.), cloned into the PMD18-T vector
(Takara Bio Inc.) and then sequenced using an ABI 3730XL DNA
sequencer (Applied Biosystems; Thermo Fisher Scientific, Inc.).
Mutation analysis of exons of
TP53
According to the MSI and LOH status of TP53 intron
1, 31 CRC and their paired normal tissues were selected for further
analysis (Table I). A total of 11
primers were designed to screen for nucleotide alterations in exon
1–11 of the TP53 gene (Table SI).
PCR-sequencing was performed using an ABI 3730XL DNA Sequencer
(Applied Biosystems; Thermo Fisher Scientific, Inc.).
 | Table I.Clinical features of patients with
colorectal cancer (n=31). |
Table I.
Clinical features of patients with
colorectal cancer (n=31).
| Clinical feature | Value |
|---|
| Mean age (years) | 68.26±9.2 |
| Sex |
|
|
GO:Male | 20 (64.52) |
|
GO:Female | 11 (35.48) |
| Drinking |
|
|
GO:Yes | 11 (35.48) |
|
GO:No | 20 (64.52) |
| Smoking |
|
|
GO:Yes | 14 (45.16) |
|
GO:No | 17 (54.84) |
| TNM stage |
|
|
GO:II | 12 (42.86) |
|
GO:III | 16 (57.14) |
| Histologic grade |
|
|
GO:Well/moderate | 12 (41.38) |
|
GO:Poor | 17 (58.62) |
| Adjuvant therapy |
|
|
GO:Yes | 13 (41.94) |
|
GO:No | 18 (58.06) |
| Survival time
(months) |
|
|
GO:<12 | 3 (15.79) |
|
GO:12–36 | 8 (42.11) |
|
GO:>36 | 8 (42.11) |
Statistical analysis
Pearson's Chi-squared and Fisher's exact test were
performed to explore the associations of MSI, LOH and MSS within
the intron and nucleotide alterations of TP53 exons with
clinicopathological characteristics. Survival analysis was
performed for overall survival (OS) and disease-free survival (DFS)
using the Kaplan-Meier method (log-rank test). Statistical analysis
was performed using SPSS 16.0 (SPSS, Inc.). P<0.05 was
considered to indicate a statistically significant difference.
Results
MS analysis of TP53 in CRCs
MSI and LOH of TP53 intron 1 were analyzed in tumors
and their matched normal tissue samples of 256 patients. Of these,
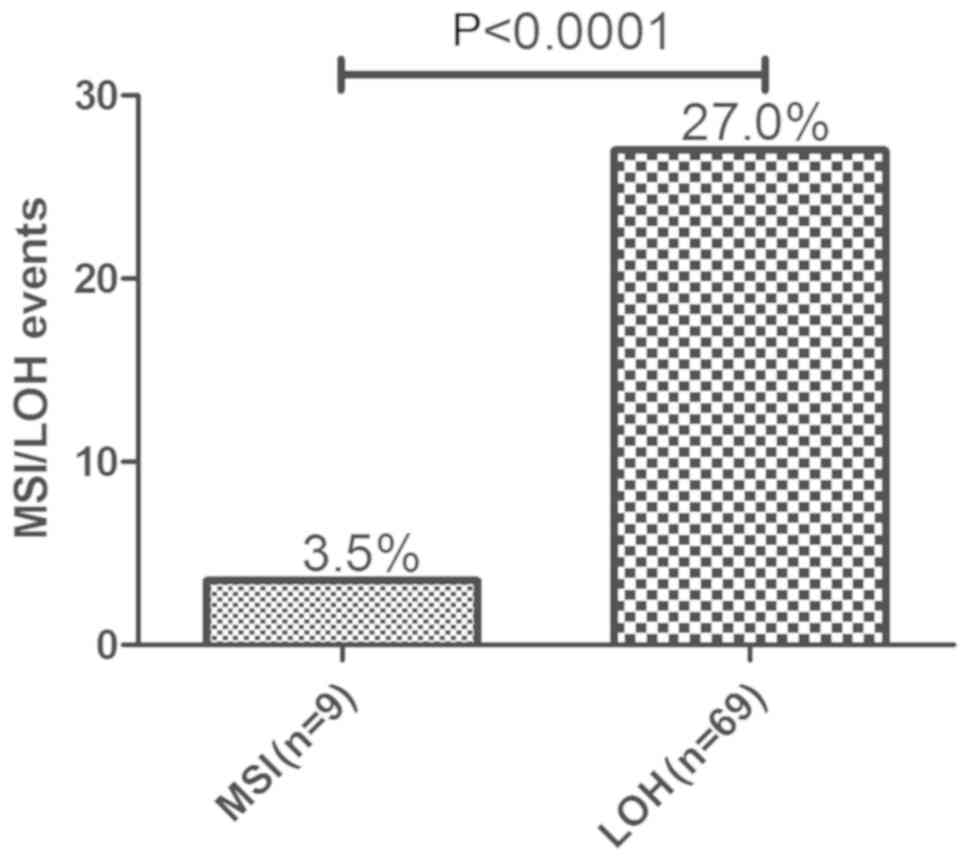
9 tumors (3.5%) were identified as having MSI and 69 (27.0%) as
having LOH in TP53 intron 1 (TP53ALU) by STR scanning. The
frequency of LOH was significantly higher than that of MSI
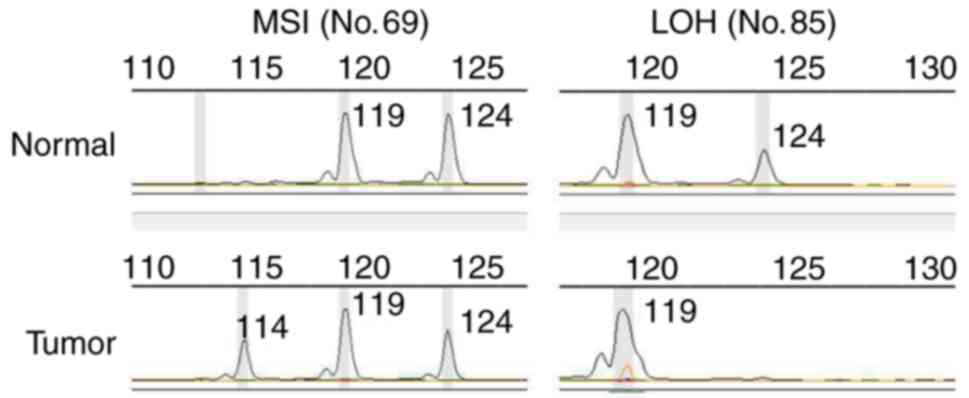
(P<0.0001; Fig. 1). MSI refers to
the presence of new fragments, whereas LOH refers to a complete or
partial loss of one of the two alleles. The two alteration patterns
of TP53ALU are provided in Fig. 2.
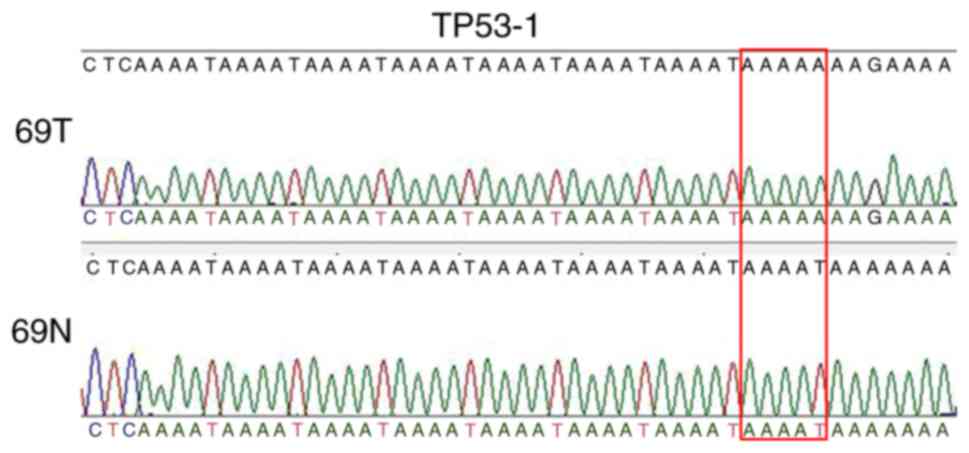
Next, clone sequencing was performed to detect the PCR product of
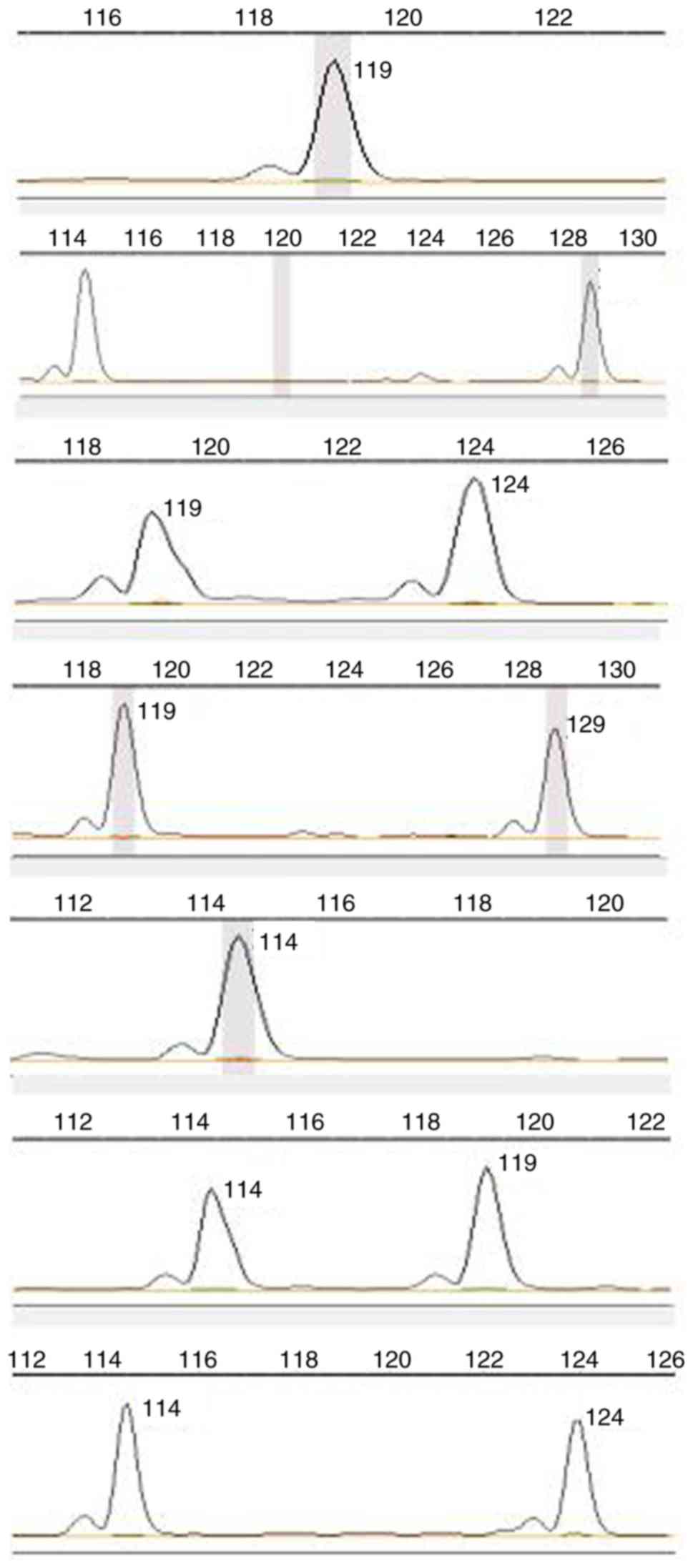
sample no. 69 and verify a new allele of 114 bp (Fig. 3). In addition, the results suggested
that MS TP53ALU, which has seven genotypes, is a polymorphic locus
in normal human tissues (Fig.
4).
The 256 patients with CRC were further stratified
based on their TP53ALU status, including MSI, LOH and MSS, and its
association with clinicopathological characteristics was determined
(Table II). No significant
differences in age, sex, smoking, drinking, depth of tumor
invasion, lymph node involvement, metastasis, pathological type,
histologic grade and survival time were observed. However, compared
to the LOH (P=0.027) and MSS (P=0.048) groups, MSI tumors exhibited
a greater association with TNM stage II.
 | Table II.Association of TP53ALU alterations
with clinical characteristics of 256 cases. |
Table II.
Association of TP53ALU alterations
with clinical characteristics of 256 cases.
|
|
| P-value | Value |
|---|
|
|
|
|
|
|---|
| Clinical feature | Patients (n) | MSI | LOH | MSS | MSI vs. MSS | LOH vs. MSS | MSI+LOH vs. MSS | MSI vs. LOH |
|---|
| Mean age (years) | 256 | 72±8.9 | 67.3±8.8 | 67.4±8.2 | 0.791a | 0.322a | 0.316a | 0.879a |
| Sex |
|
|
|
| 0.736 | 0.305 | 0.252 | 1 |
|
GO:Male | 146 | 6 (4.1%) | 43 (29.5%) | 97 (66.4%) |
|
|
|
|
|
GO:Female | 108 | 3 (2.8%) | 26 (24.1%) | 79 (73.1%) |
|
|
|
|
| Smoking |
|
|
|
| 0.742 | 0.392 | 0.552 | 0.535 |
|
GO:Yes | 88 | 2 (2.3%) | 27 (30.7%) | 59 (67.0%) |
|
|
|
|
|
GO:No | 167 | 7 (4.2%) | 42 (25.1%) | 118 (70.7%) |
|
|
|
|
| Drinking |
|
|
|
| 1.000 | 0.592 | 0.610 | 1.000 |
|
GO:Yes | 57 | 2 (3.5%) | 17 (29.8%) | 38 (66.7%) |
|
|
|
|
|
GO:No | 198 | 7 (3.5%) | 52 (26.3%) | 139 (70.2%) |
|
|
|
|
| TNM stage |
|
|
|
| 0.048 | 0.421 | 0.965 | 0.027 |
|
GO:II | 127 | 7 (5.5%) | 36 (28.3%) | 84 (66.1%) |
|
|
|
|
|
GO:III | 93 | 1 (1.07%) | 23 (24.7%) | 69 (74.1%) |
|
|
|
|
| Depth of tumor
invasion |
|
|
|
| 0.443 | 0.868 | 0.719 | 0.667 |
|
GO:pT2 | 20 | 1 (5%) | 6 (30%) | 13 (65%) |
|
|
|
|
|
GO:pT3 | 190 | 8 (4.2%) | 51 (26.8%) | 131 (68.9%) |
|
|
|
|
|
GO:pT4 | 32 | 0 (0.0%) | 8 (25.0%) | 24 (75.0%) |
|
|
|
|
| Lymph node
involvement |
|
|
|
| 0.122 | 0.288 | 0.105 | 0.281 |
|
GO:pN0 | 144 | 8 (5.6%) | 42 (29.2%) | 94 (65.3%) |
|
|
|
|
|
GO:pN1 | 73 | 1 (1.4%) | 19 (26.0%) | 53 (72.6%) |
|
|
|
|
|
GO:pN2 | 22 | 0 (0.0%) | 4 (18.2%) | 18 (81.8%) |
|
|
|
|
| Metastasis |
|
|
|
| 1.000 | 1.000 | 1.000 | 1.000 |
|
GO:M0 | 252 | 9 (3.6%) | 68 (26.9%) | 175 (69.4%) |
|
|
|
|
|
GO:M1 | 3 | 0 (0.0%) | 1 (33.3%) | 2 (66.7%) |
|
|
|
|
| Pathological
type |
|
|
|
| 1.000 | 0.514 | 0.449 | 0.412 |
|
GO:Adenocarcinoma | 216 | 8 (3.7%) | 62 (28.7%) | 146 (67.6%) |
|
|
|
|
|
GO:Mucinous carcinoma | 17 | 1 (5.9%) | 3 (17.6%) | 13 (76.5%) |
|
|
|
|
| Histologic
grade |
|
|
|
| 0.682 | 0.245 | 0.185 | 1.000 |
|
GO:Well | 50 | 1 (2.0%) | 11 (22.0%) | 38 (76.0) |
|
|
|
|
|
GO:Moderate/poor | 168 | 7 (4.2%) | 50 (29.8%) | 111 (66.1%) |
|
|
|
|
| Survival time
(months) |
|
|
|
|
|
|
|
|
|
GO:OS | 210 | 37.09 | 27.9 | 30 | 0.307b | 0.760 b | 0.560 b | 0.326 b |
|
GO:PFS | 218 | 36.23 | 25.68 | 28.22 | 0.315b | 0.415b | 0.290b | 0.310b |
Mutation analysis of TP53 exons in the
CRCs
DNA sequencing of the exons of the TP53 gene in 31
CRC samples (MSI, n=4; LOH, n=7; MSS, n=20) revealed a total of 6
mutations, which were mainly distributed in 4 exons, in 4 samples
(4/31, 12.9%). Of all of the mutations, 66 (C>T) were in exon 2
(codon 13), 12 (G>A) in exon 3 (codon 36), 108 (G>A) in exon
3 (codon 68), 25 (T>C) in exon 5 (codon 195), 71 (G>G+A) in
exon 7 (codon 273) and 36 (G>G+A) in exon 7 (codon 285). All of
these mutation sites have been defined as single nucleotide
polymorphism (SNP) loci and previously registered in the PubMed
database. The tumor samples T7 and T11 were regarded as the most
unstable cases, owning to two mutations. Exon 3 and exon 7 were
prone to mutations, as they were harboring 2 mutation loci
(Table III).
 | Table III.Mutations of tumor protein 53 exons
detected in colorectal cancer samples (n=31). |
Table III.
Mutations of tumor protein 53 exons
detected in colorectal cancer samples (n=31).
| Case | MS status | Exon2 | Exon3 | Exon5 | Exon7 |
|---|
| T7 | LOH | – | 12:G>A | – | 71:G>G+A |
|
|
| – | rs1800370 | – | rs28934576 |
| T8 | LOH | – | – | 25:T>C | – |
|
|
| – | – | rs760043106 | – |
| T11 | LOH | 66:C>T | 108:G>A | – | – |
|
|
| rs878854070 | rs746814615 | – | – |
| T24 | MSS | – | – | – | 36:G>G+A |
|
|
| – | – | – | rs112431538 |
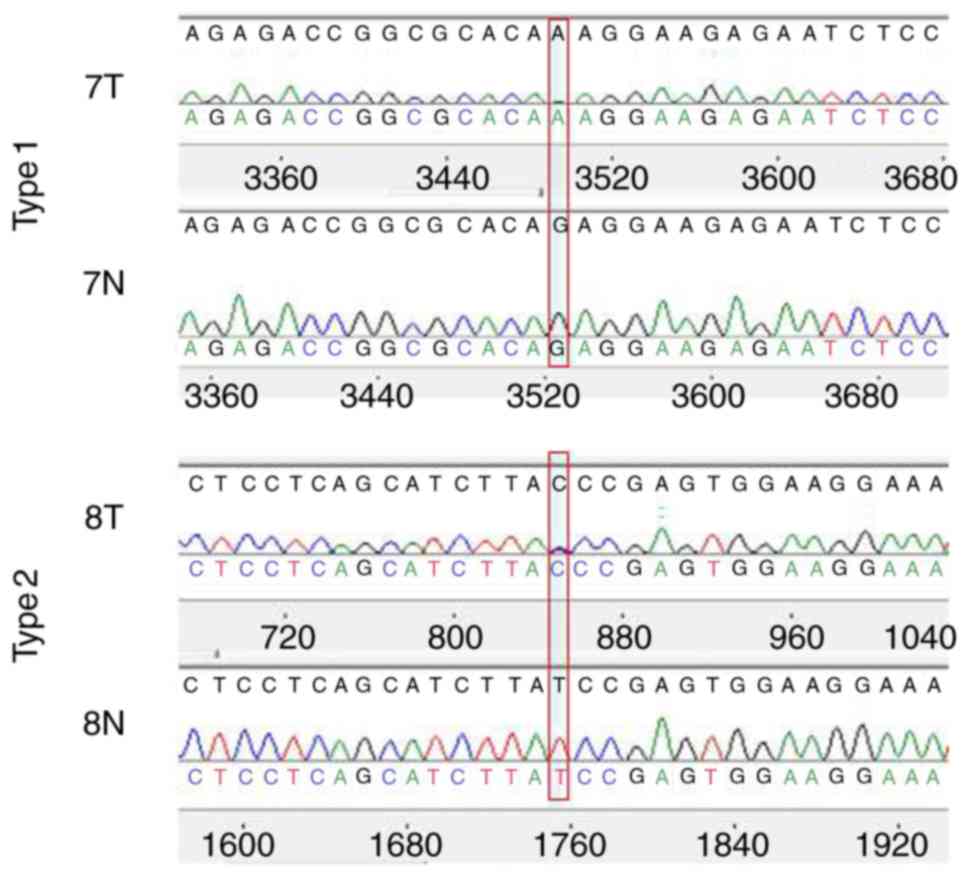
Furthermore, the mutation patterns of TP53 exons
were divided into 2 types. Among the six mutations detected, the
genotype of three mutations changed from homozygous to homozygous
and the remaining mutations ranged from homozygous to heterozygous.
For instance, compared to matched normal tissues, the genotype of
tumor sample 7 changed from G/G to A/A. These genotypes ranging
from homozygous to homozygous were classified as Type 1 mutations,
whereas the genotype of tumor sample T8 changed from T/T to T/C,
which ranged from homozygous to heterozygous, and was classified as
a Type 2 mutation (Fig. 5).
Unexpectedly, no correlation was identified between
the TP53 exon mutation and any of the clinicopathological
characteristics (Table IV). While
there was a slight trend towards a worse prognosis, no significant
association between TP53 exon mutation and survival (OS and DFS)
was detected (POS=0.093, PDFS=0.095).
 | Table IV.Association of tumor protein 53-exon
mutations with clinical characteristics of the patients (n=31). |
Table IV.
Association of tumor protein 53-exon
mutations with clinical characteristics of the patients (n=31).
| Item | Patients (n) | Mut | Non-Mut | P-value |
|---|
| Mean age
(years) | 31 | 67 | 68.32 | 0.815 |
| Sex |
|
|
| 0.115 |
|
GO:Male | 20 | 1 | 19 |
|
|
GO:Female | 11 | 3 | 8 |
|
| Drinking |
|
|
| 1 |
|
GO:Yes | 11 | 1 | 10 |
|
|
GO:No | 20 | 3 | 17 |
|
| Smoking |
|
|
| 0.304 |
|
GO:Yes | 14 | 3 | 11 |
|
|
GO:No | 17 | 1 | 16 |
|
| Histologic
grade |
|
|
| 1 |
|
GO:Well | 12 | 2 | 10 |
|
|
GO:Moderate/poor | 17 | 2 | 15 |
|
| TNM stage |
|
|
| 1 |
|
GO:II | 12 | 2 | 10 |
|
|
GO:III | 16 | 2 | 14 |
|
| Survival time
(months) |
|
|
|
|
|
GO:OS | 31 | 10.65 | 37.53 | 0.093 |
|
GO:PFS | 31 | 10.64 | 36.24 | 0.095 |
Association between TP53 exon mutation
and TP53ALU alterations
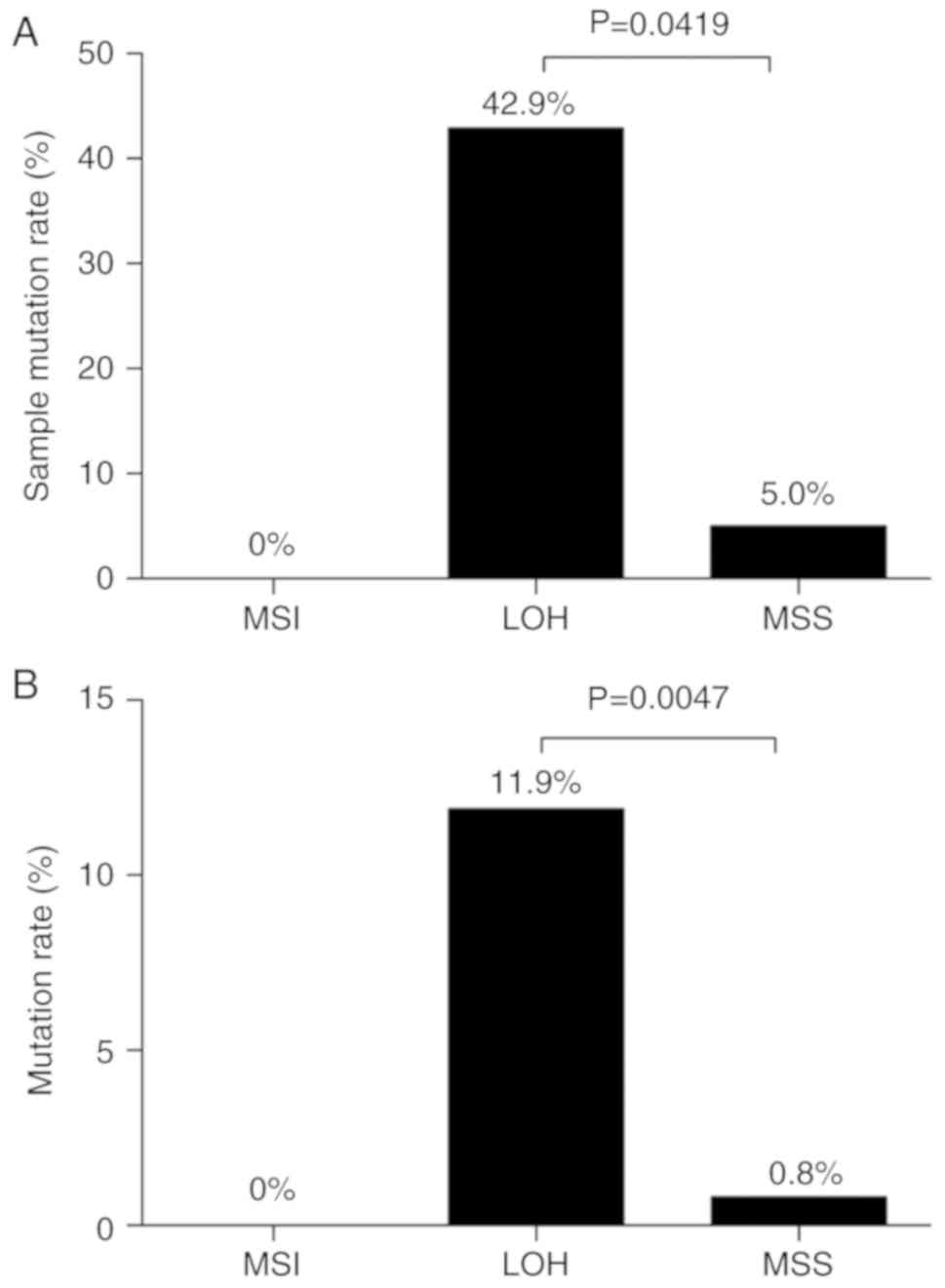
A total of 6 mutations were identified in 3 LOH
samples (3/7, 42.9%) and in 1 MSS sample (1/20, 5.0%). None of the
exon mutations in TP53 was detected in the MSI samples (0/4, 0%).
The mutation rate in the LOH group was significantly higher than
that in the MSS group (P=0.0419; Fig.
6A). In addition, regarding the 6 mutation positions in 31
tumor tissues, the mutation rate in the MSI group was 0% [0
mutations/(6 positions × 4 MSI-tumor cases)]; however, it was 11.9%
[5 mutations/(6 positions × 7 LOH-tumor cases)] in the LOH group,
which was significantly higher than that in the MSS group [1
mutation/(6 positions × 20 MSS-tumor cases), 0.8%; P=0.0047;
Fig. 6B].
Discussion
CRC is the second most common cancer type with 1.2
million novel cases per year worldwide (14). Chromosomal and MS alterations
constitute the major genetic instability events in CRC (15,16).
However, few studies have explored the correlation between the MS
status of a single gene and the exon mutations in the same gene. In
the present study, the MSI and LOH status of the MS in an intron of
the TP53 gene was detected and the association between MSI/LOH
alterations and the mutations in TP53 exons was analyzed. The
results suggested a positive association between LOH alterations in
the MS of the TP53 intron and mutations in TP53 exons in CRCs.
These results indicated that MS analysis is not sufficient to
predict the mutation status of TP53 exons, but was higher than the
probability for MSS or MSI when LOH was present.
MS are widely abundant in the genome and certain MS
are highly polymorphic in the normal population. The MS TP53ALU was
initially identified to be a polymorphic locus and has been
frequently used as a marker in previous studies (17,18). In
the present study, 7 genotypes of TP53 ALU were detected in the
matched normal tissues, and it was confirmed that it is a
polymorphic MS.
CRCs with MSI have unique clinicopathological
features. Studies have indicated that patients with MSI in their
tumors frequently exhibit localization in the right colon, female
sex, mucinous histology, larger tumor size and less advanced
disease stage (19–22). In the present study, the TP53ALU MSI
phenotype was significantly associated with the TNM stage, as
TP53ALU MSI tumors were more frequently stage II than LOH (P=0.027)
and MSS tumors (P=0.048). These results indicated that the status
of TP53ALU may be of prognostic value regarding the TNM stage of
CRC.
A total of 6 different mutations in the TP53 exons
were identified in 4 unrelated CRC patients by comparing the
sequence of the tumor's TP53 with those of the matched normal
tissues. Although all of the mutations (6/6) are known as SNP loci,
position 71 in exon 7 (codon 273) is a hotspot that has been
previously reported (23). All of
the mutations were in coding sequences and only three missense
mutations were detected: Codon 13 (p. Pro13Leu), codon 195 (p.
Ile195Thr) and codon 273 (p. Arg273His).
The role of SNPs in the regulation of different
functions of proteins is an important genetic research field aimed
at understanding the molecular basis of disease (24). Among the SNPs, non-synonymous coding
SNPs in the coding regions result in an amino acid variation in the
protein products of genes, which are thought to have an impact on
the phenotype (25). The TP53
rs28934576 variation (p.Arg273His) is a known pathological mutation
that is important in the initiation and progression of CRC, which
has been previously described in a public database. The rs1800370
(CCG to CCA, both encoding proline) is a synonymous SNP, which does
not change the amino acid sequence, but this silent mutation has
been indicated to reduce the ability of p53 to activate apoptosis
by lowering its synthesis through a reduction in the affinity of
its mRNA to MDM2 (26). The
rs112431538 and rs760043106 variations of TP53 are likely
pathogenic according to the records in the PubMed database
(https://www.ncbi.nlm.nih.gov/clinvar/variation/420133/#clinical-assertions;
http://www.ncbi.nlm.nih.gov/clinvar/?linkname=snp_clinvar&from_uid=760043106).
However, no correlation was identified between TP53
exon mutations and any of the clinicopathological characteristics;
this may be due to the limited number of cases examined. However,
patients with TP53 exon mutation exhibited a slight but
insignificant tendency to have shorter OS (P=0.093) and DFS
(P=0.095). This is in accordance with the mutations in tumor
suppressor genes that are associated with highly malignant tumors
in a previous study (27). Driver
mutations in CRC include intracellular KRAS, B-Raf proto-oncogene,
serine/threonine kinase, phosphatidylinositol-4,5-bisphosphate
3-kinase catalytic subunit alpha, TP53, F-box and WD repeat
domain-containing 7 and NRAS gene mutations, and the first four of
these mutations have been proposed as indicators of CRC prognosis.
Therefore, the correlation between TP53 gene mutation and
chemotherapy requires further exploration.
The present study had several limitations that
require further discussion. The number of samples tested was small
due to the exclusion of the effects of mutations at other sites.
The small sample size may have affected the accuracy of the
results, and therefore, the present results require to be verified
using a larger sample size. Still, the results of the present study
provided a basis for further in-depth research. Due to the limited
number of specimens, no histological control of the samples used
for DNA extraction was performed. Although exon mutations were not
detected, it cannot be ruled out that these mutations may affect
protein sequence/transcription.
In conclusion, the present study indicated that the
prevalence of TP53 exon mutations was significantly higher in CRC
tumors with LOH of TP53 than in CRC tumors with MSI and MSS of
TP53. Exonmutations of TP53 were associated with TP53ALULOH.
Furthermore, an MSI status was closely associated with stage II
CRCs.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
The present study was supported by the National
Science Foundation of China (grant no. 31772545) and the Support
Project of High-level Teachers in Beijing Municipal Universities in
the Period of the 13th five-year Plan (grant no. IDHT
20170516).
Availability of data and materials
All data generated or analyzed during the present
study are included in this published article.
Authors' contributions
ZC, XL, XH and DF contributed to the conception and
design of the present study. XL and DF performed the experiments.
XL, DF, XH and XX analyzed the data. XL, DF and XH drafted and
revised the manuscript. All authors critically revised the
manuscript and approved the final version.
Ethics approval and consent to
participate
The study was reviewed and approved by China
National center for Biotechnology development (Beijing, China).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Kuzminov A: Inhibition of DNA synthesis
facilitates expansion of low-complexity repeats: Is strand slippage
stimulated by transient local depletion of specific dNTPs?
Bioessays. 35:306–313. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Liu X and Meltzer SJ: Gastric cancer in
the era of precision medicine. Cell Mol Gastroenterol Hepatol.
3:348–358. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Yan WY, Hu J, Xie L, Cheng L, Yang M, Li
L, Shi J, Liu BR and Qian XP: Prediction of biological behavior and
prognosis of colorectal cancer patients by tumor MSI/MMR in the
Chinese population. Onco Targets Ther. 9:7415–7424. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Lech G, Słotwiński R, Słodkowski M and
Krasnodębski IW: Colorectal cancer tumour markers and biomarkers:
Recent therapeutic advances. World J Gastroenterol. 22:1745–1755.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Worthley DL and Leggett BA: Colorectal
cancer: Molecular features and clinical opportunities. Clin Biochem
Rev. 31:31–38. 2010.PubMed/NCBI
|
|
6
|
Parikh N, Hilsenbeck S, Creighton CJ,
Dayaram T, Shuck R, Shinbrot E, Xi L, Gibbs RA, Wheeler DA and
Donehower LA: Effects of TP53 mutational status on gene expression
patterns across 10 human cancer types. J Pathol. 232:522–533. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Silwal-Pandit L, Vollan HK, Chin SF, Rueda
OM, McKinney S, Osako T, Quigley DA, Kristensen VN, Aparicio S,
Børresen-Dale AL, et al: TP53 mutation spectrum in breast cancer is
subtype specific and has distinct prognostic relevance. Clin Cancer
Res. 20:3569–3580. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Köbel M, Piskorz AM, Lee S, Lui S, LePage
C, Marass F, Rosenfeld N, Mes Masson AM and Brenton JD: Optimized
p53 immunohistochemistry is an accurate predictor of TP53 mutation
in ovarian carcinoma. J Pathol Clin Res. 2:247–258. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Saha MN, Qiu L and Chang H: Targeting p53
by small molecules in hematological malignancies. J Hematol Oncol.
6:232013. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Smith G, Carey FA, Beattie J, Wilkie MJ,
Lightfoot TJ, Coxhead J, Garner RC, Steele RJ and Wolf CR:
Mutations in APC, Kirsten-ras, and p53-alternative genetic pathways
to colorectal cancer. Proc Natl Acad Sci USA. 99:9433–9438. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Liu Y, Chen C, Xu Z, Scuoppo C, Rillahan
CD, Gao J, Spitzer B, Bosbach B, Kastenhuber ER, Baslan T, et al:
Deletions linked to TP53 loss drive cancer through p53-independent
mechanisms. Nature. 531:471–475. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Futreal PA, Barrett JC and Wiseman RW: An
Alu polymorphism intragenic to the TP53 gene. Nucleic Acids Res.
19:69771991. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Du X, Chen Z, Li W, Tan Y, Lu J, Zhu X,
Zhao T, Dong G and Zeng L: Development of novel microsatellite DNA
markers by cross-amplification and analysis of genetic variation in
gerbils. J Hered. 101:710–716. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Siegel R, Naishadham D and Jemal A: Cancer
statistics, 2012. CA Cancer J Clin. 62:10–29. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Fodde R, Smits R and Clevers H: APC,
signal transduction and genetic instability in colorectal cancer.
Nat Rev Cancer. 1:55–67. 2001. View
Article : Google Scholar : PubMed/NCBI
|
|
16
|
Müller MF, Ibrahim AE and Arends MJ:
Molecular pathological classification of colorectal cancer.
Virchows Arch. 469:125–134. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Kamat N, Khidhir MA, Jaloudi M, Hussain S,
Alashari MM, Al Qawasmeh KH and Rannug U: High incidence of
microsatellite instability and loss of heterozygosity in three loci
in breast cancer patients receiving chemotherapy: A prospective
study. BMC Cancer. 12:3732012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Hahn M, Fislage R and Pingoud A:
Polymorphism of the pentanucleotide repeat d(AAAAT) within intron 1
of the human tumor suppressor gene p53 (17p13.1). Hum Genet.
95:471–472. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Wangefjord S, Brändstedt J, Lindquist KE,
Nodin B, Jirström K and Eberhard J: Associations of beta-catenin
alterations and MSI screening status with expression of key cell
cycle regulating proteins and survival from colorectal cancer.
Diagn Pathol. 8:102013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Jass JR: Classification of colorectal
cancer based on correlation of clinical, morphological and
molecular features. Histopathology. 50:113–130. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Salahshor S, Kressner U, Fischer H,
Lindmark G, Glimelius B, Påhlman L and Lindblom A: Microsatellite
instability in sporadic colorectal cancer is not an independent
prognostic factor. Br J Cancer. 81:190–193. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Samowitz WS, Curtin K, Neuhausen S,
Schaffer D and Slattery ML: Prognostic implications of BAX and
TGFBRII mutations in colon cancers with microsatellite instability.
Genes Chromosomes Cancer. 35:368–371. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Bougeard G, Renaux-Petel M, Flaman JM,
Charbonnier C, Fermey P, Belotti M, Gauthier-Villars M,
Stoppa-Lyonnet D, Consolino E, Brugières L, et al: Revisiting
Li-Fraumeni syndrome from TP53 mutation carriers. J Clin Oncol.
33:2345–2352. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Dantzer J, Moad C, Heiland R and Mooney S:
MutDB services: Interactive structural analysis of mutation data.
Nucleic Acids Res. 33:W311–W314. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Ramensky V, Bork P and Sunyaev S: Human
non-synonymous SNPs: Server and survey. Nucleic Acids Res.
30:3894–3900. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Krammer PH: CD95's deadly mission in the
immune system. Nature. 407:789–795. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Tamura G: Genetic and epigenetic
alterations of tumor suppressor and tumor-related genes in gastric
cancer. Histol Histopathol. 17:323–329. 2002.PubMed/NCBI
|