Introduction
Autophagy is a vital metabolic process in eukaryotic
cells, which plays a significant role in regulation of cell
survival and death (1). Excessive
autophagy disrupts cellular functions, which directly results in
autophagic programmed cell death or apoptotic cell death. In
certain cases, autophagy counteracts apoptotic cell death via a
cell survival pathway (2). Cervical
cancer is the second most prevalent type of malignancy in females
(3). The primary and most common
anticancer therapy for cervical cancer is chemotherapy. However,
tumour cells develop intrinsic resistance against chemotherapy.
Plant-derived compounds characterised by low toxicity and a wide
range of anticancer activities present as promising novel
anticancer agents (4).
Panax ginseng C.A
Meyer is a type of traditional medicinal plant that
has been widely used in Asian regions for thousands of years
(5). Ginsenosides, some of the most
important active components in Panax ginseng, exert numerous
pharmacological actions, including marked suppression of the
proliferation and migration of tumour cells (5). Different ginsenoside monomers have
different effects on autophagy in cancer cells and act on different
pathways (6,7). A recent study suggested that total
ginsenoside (TGN) extract induces autophagic cell death in
non-small cell lung cancer cells (8). However, the TGN concentrations used in
the study were considered to be high (0.125-1 mg/ml) and the
changes in p62 expression were not discussed.
Bone marrow stromal antigen 2 (BST-2), also referred
to as CD317, tetherin and HM1.24 is a multifunctional protein. It
is an interferon-inducible type II transmembrane glycoprotein that
functions as an NF-κB activator (9), host restriction factor that tethers
virions on the cell membrane (10)
and survival protein that increases cancer cell adhesion and
resistance to apoptosis (11).
Under various disease conditions, particularly malignancies, BST-2
has been reported to be upregulated (12). The increased expression of BST-2
mediates tumour growth, invasion and metastasis (12,13).
Therefore, BST-2 is upregulated in autophagy knockdown cells and
associated with CD63, which may inhibit hepatitis C virus assembly
or release (14). Another study
indicated that a non-canonical autophagy pathway reminiscent of
microtubule-associated protein light chain 3 (LC3)-associated
phagocytosis contributes to viral protein U counteraction of BST-2
restriction (15). These data
indicate that BST-2 is an autophagy-associated factor.
The effect of TGN on autophagy of cervical cancer
cells remains unclear. In the present study, the effect of TGN on
autophagy in HeLa cells was investigated and TGN treatment was
found to induce irreversible autophagy in a concentration- and
time-dependent manner. TGN increased the expression of p62 at
transcriptional and protein expression levels. Further experiments
revealed that TGN promoted autophagic cell death under
serum-deprived conditions accompanied by downregulation of BST-2,
which is important for the survival of cancer cells.
Materials and methods
Antibodies and reagents
The following antibodies were used in the present
study: Mouse anti-tubulin monoclonal antibody (mAb; cat. no.
627901; BioLegend, Inc.), rabbit anti-LC3 mAb, rabbit anti-beclin 1
mAb (cat. nos. 12741 and 3498, respectively; Cell Signaling
Technology, Inc.) and rabbit anti-BST-2 pAb (cat. no. BS5634;
Bioworld Technology, Inc.). The secondary antibodies were
HRP-conjugated goat anti-mouse and anti-rabbit immunoglobulin G
(cat. nos. 115-035-003 and 111-585-003, respectively; Jackson
ImmunoResearch Laboratories, Inc.). Ginsenoside Rb1,
Rb2, Rc, Rd, Rg1, Rg2 and Rf
(purity, >98%) were purchased from Chengdu Must Bio-Technology
Co., Ltd. Acridine orange, lysis buffer and a Braford assay kit
were purchased from Beyotime Institute of Biotechnology. Cell
counting kit-8 (CCK-8) was purchased from Boster Biological
Technology. Rapamycin, bafilomycin A1 and 3-methyladenine (3-MA)
were purchased from InvivoGen. Actinomycin D was purchased from
MedChemEpxress. Other ginsenosides were purchased from Chengdu Must
Bio-Technology Co., Ltd. TRIzol® was purchased from
Thermo Fisher Scientific, Inc. SYBR Premix Ex Taq™ and a
reverse transcription kit were purchased from Takara Biotechnology
Co., Ltd. The protein extraction buffer was purchased from Beyotime
Institute of Biotechnology. Earle's Balanced Salt Solution (EBSS)
was purchased from Beijing Solarbio Science & Technology Co.,
Ltd.
Preparation of TGN
Ginseng crude powder (1 kg) was soaked in water
overnight and extracted four times in boiling water for 3 h each
time. The water-soluble substances were collected, applied to a
D101 macroporous resin column and eluted with EtOH:H2O
(0:100, 75:25; v/v). The eluent was condensed and evaporated to
obtain TGN.
Analysis of TGN by high-performance
liquid chromatography (HPLC)
To analyse ginsenoside monomers, an Agilent 1260
series high performance liquid chromatograph, Agilent system
chemistry workstation and Agilent 1260 UV-visible wavelength
detector were used. A Sepax Bio-C18 HPLC column (5 µm; 4.6x250 mm)
was used for ginsenoside separation. The temperature of the column
was maintained at 40˚C. The mobile phase consisted of solvent A
(acetonitrile) and solvent B (water). The gradient elution program
was as follows: 0-45 min, A 19%; 45-50 min, A 19-27%; 50-60 min, A
27-31%; 60-70 min, A 31-28%; 70-85 min, A 28-35%; 85-100 min, A
35%. The flow rate was set at 1 ml/min. The quantitative method
used was an external standard method (16).
Cell culture and transfection
The following cell lines were obtained from the
American Type Culture Collection: HeLa (cat. no. CCL-2), MS751
(cat. no. HTB-34) and C-33A (cat. no. HTB-31) and were cultured in
DMEM (Invitrogen; Thermo Fisher Scientific, Inc.) supplemented with
10% fetal bovine serum at 37˚C with 5% CO2. Culture
under serum deprivation condition was culture in DMEM (Invitrogen;
Thermo Fisher Scientific, Inc.) at 37˚C with 5% CO2.
Enhanced green fluorescent protein (EGFP)-LC3, BST-2 IHA and VR1012
have been described previously (17,18).
Lipofectamine® 2000 (Invitrogen; Thermo Fisher
Scientific, Inc.) was used for transient plasmid transfections.
EGFP-LC3-II plasmid (500 ng) was transfected into HeLa, MS751 and
C-33A cells. After 37˚C for 24 h, the cells were treated with
dimethylsulfoxide (DMSO) or TGN for an additional 7 h and then
analysed for fluorescence. ImageJ Software (v1.8.0; National
Institutes of Health) was used for densitometric analysis. VR1012
or BST-2 IHA plasmid (50 ng) was transfected into HeLa, MS751 and
C-33A cells. After incubation at 37˚C for 24 h, cells were treated
with DMSO or TGN for an additional 24 h and then analysed via CCK-8
assay. Following overnight culture in 6-well plates, HeLa cells
were treated with Actinomycin D (80 mM) at 37˚C with 5%
CO2 for 24 h. Following overnight culture in 6-well
plates, cells were treated with 1 ml EBSS at 37˚C for 3 h.
Western blotting
After overnight culture in 6-well plates, HeLa cells
were treated with DMSO or TGN (40/60/80 µg/ml) for 24 h. Cells were
harvested by centrifugation (800 x g; 25˚C; 5 min), resuspended in
RIPA total protein extraction lysis buffer (cat. no. BD0031;
Bioworld Technology, Inc.) and BCA was used to detect the protein
content. The loading buffer was added and boiled for 15 min. A
total of 8 µg protein/lane was separated by SDS-PAGE on a 12% gel
and the separated proteins were transferred to nitrocellulose
membranes (Whatman plc; Cytiva). The membranes were blocked with 5%
dry non-fat milk (BD Biosciences) for 30 min at room temperature.
After washing in PBST 3 times, the membranes were incubated with
the primary antibodies detailed in Antibodies and reagents
(dilution, 1:1,000) overnight at 4˚C, washed with in PBST three
times and then incubated with a secondary antibody as detailed in
Antibodies and reagents (dilution, 1:1,000) for 1 h at room
temperature. Protein bands were visualised using the Ultra High
Sensitivity ECL Substrate kit (Beyotime Institute of
Biotechnology). Immunoreactivity was visualised by
chemiluminescence and densitometric analysis was performed with
ImageJ Software (v1.8.0). The western blots in Fig. 3 are from cells following treatment
with 80 µg/ml TGN for 16 h or EBSS at 37˚C for 3 h. The compounds
were washed out and proteins were extracted at the indicated time
points.
Reverse transcription-quantitative PCR
(RT-qPCR)
Following overnight culture in 6-well plates, HeLa
cells were treated with DMSO or TGN for 24 h. Following
centrifugation (800 x g; 25˚C; 5 min), the cells were collected and
washed in PBS three times. Total RNA was extracted with TRIzol and
reverse transcribed into cDNA using the reverse transcription kit.
The PCR primers used were as follows: p62 forward,
5'-GCCAGAGGAACAGATGGAGT-3' and reverse, 5'-TCCGATTCTGGCATCTGTAG-3';
BST-2 forward, 5'-CTGCAACCACACTGTGATG-3' and reverse,
5'-ACGCGTCCTGAAGCTTATG-3'; GAPDH forward,
5'-GGTGAAGGTCGGAGTCAACGGA-3' and reverse,
5'-GAGGGATCTCGCTCCTGGAAGA-3'. RT-qPCR was performed using a SYBR
Premix Ex Taq kit and the CFX Connect™ Real-Time system
(Bio-Rad Laboratories, Inc.). The thermal cycling conditions were
as follows: 95˚C for 3 min; 40 cycles of 95˚C for 10 sec and 55˚C
for 30 sec; 95˚C for 10 sec; 65˚C for 5 sec and 95˚C for 5 sec.
Data were calculated relative to a calibrator according to the
2-∆∆Cq method (19).
Acidic vesicular organelle (AVO)
staining assay
After rinsing in PBS and fixing with 4%
paraformaldehyde 25˚C for 10 min, the cells were stained for AVOs
in the dark 37˚C for 30 min. Stained cells were observed and imaged
by fluorescence microscopy (excitation, 488 nm).
Cell Counting Kit (CCK)-8 assay
Cells were cultured overnight in 96-well plates. The
next day, the culture supernatant was replaced with medium
containing DMSO or TGN and the cells were incubated 37˚C for 40
min. For analysis, CCK-8 substrate was added to the 96-well plates,
followed by incubation at 37˚C for 1 h. Absorbance was measured at
450 nm using an Infinite 200 PRO microplate reader (Tecan Group,
Ltd.). The cells in Fig. 4C were
treated with 80 µg/ml TGN and 3-MA for 24 h and cell viability
detected via Cell Counting Kit-8 assays.
Statistical analysis
All data represent at least three independent
experiments, which were evaluated statistically by one-way ANOVA
and Dunnett's post hoc test. P<0.05 was considered to indicate a
statistically significant difference. The statistical analysis
software was GraphPad Prism 7.0 (GraphPad Software Inc.).
Results
TGN induces cervical cancer cell
autophagy in a time- and concentration-dependent manner
A TGN extract of ginseng root was analysed by HPLC.
The chromatograms, including standard and sample chromatograms are
presented in Fig. S1. The main
components and contents of TGN are presented in Table I. The retention times specified are
presented in Table II. To
determine the effect of TGN on autophagy, the expression levels of
autophagy factors in TGN-treated HeLa, MS751 and C-33A cells were
investigated. TGN was observed to increase the processing of LC3-I
to LC3-II in a time- and dose-dependent manner, as well as increase
the expression of autophagy-related factor Beclin-1 in the three
types of cervical cancer cells (Fig.
1A and B).
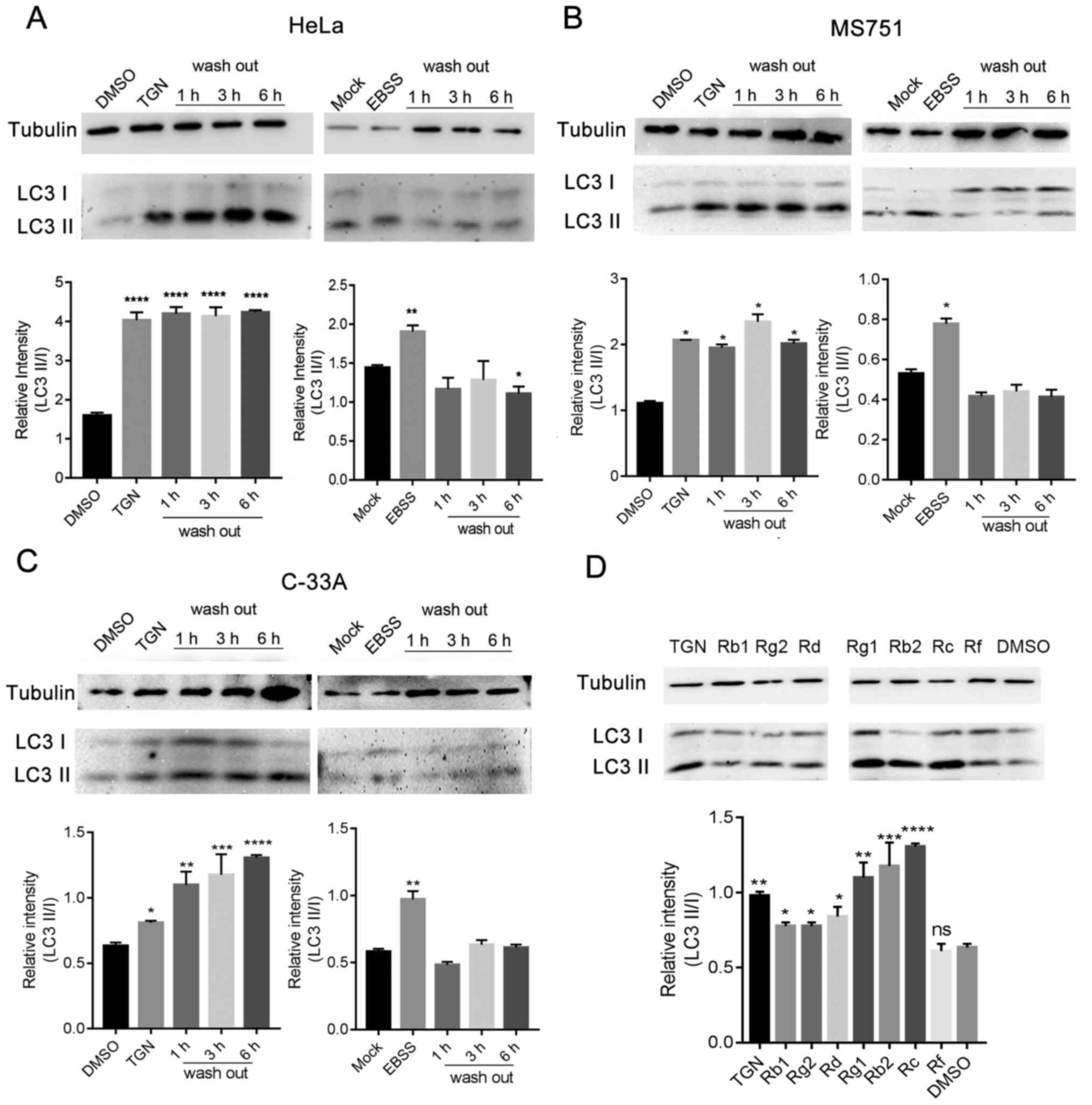
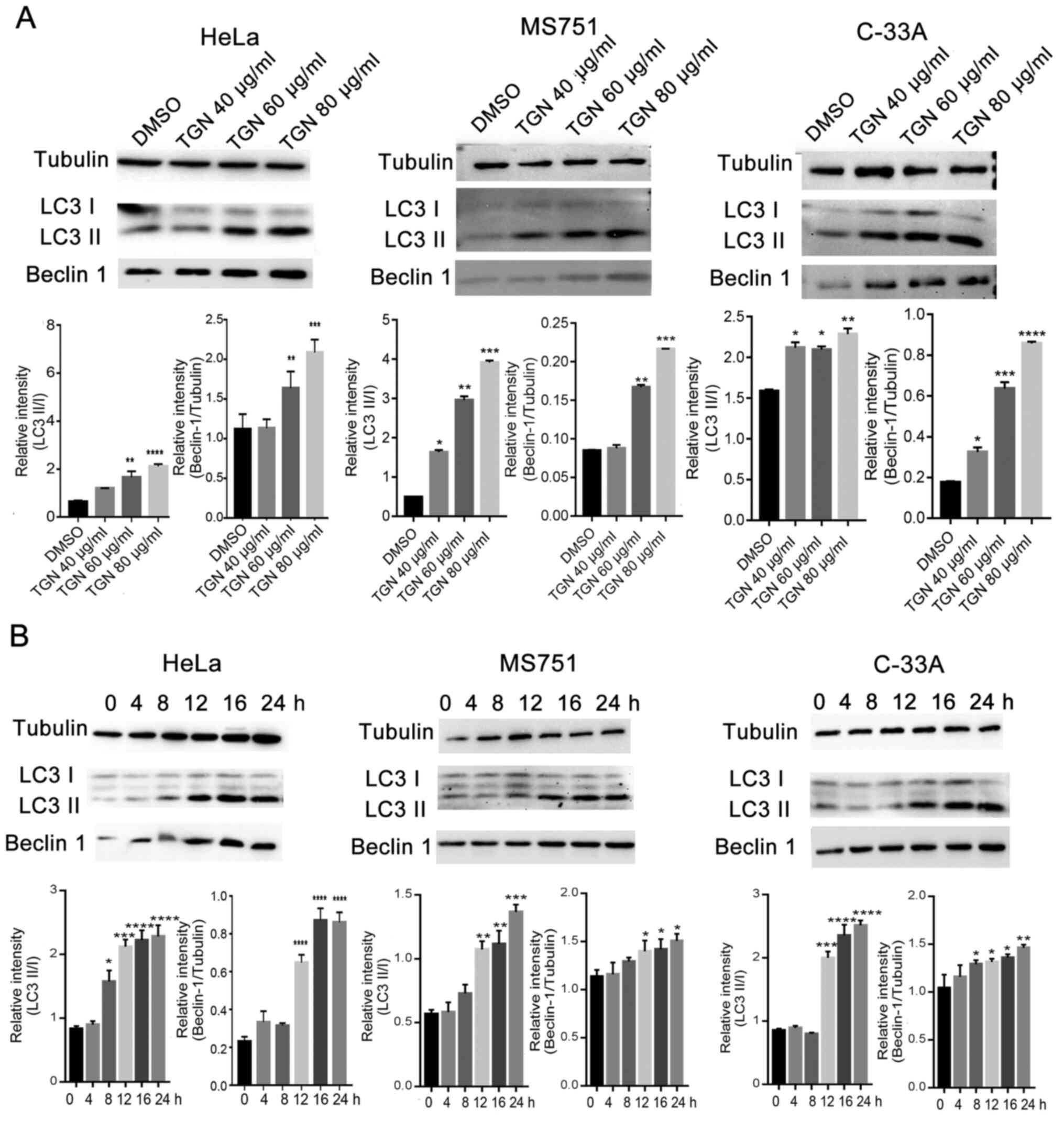 | Figure 1TGN induces cell autophagy in
cervical cancer cell lines, HeLa, MS751 and C-33A in a (A) dose-
and (B) time-dependent manner. Western blot analysis was performed
with antibodies specific for Beclin-1, LC3 I/II and control protein,
tubulin, followed by statistical analysis of the results; the
values are presented as means ± SD. *P<0.05,
**P<0.01, ***P<0.001 and
****P<0.0001 vs. DMSO. DMSO, dimethylsulfoxide; TGN,
total ginsenoside; LC3, microtubule-associated protein light chain
3. |
 | Table IMain components and contents of total
ginsenosides. |
Table I
Main components and contents of total
ginsenosides.
| Type | Content (%) |
|---|
| Rb1 | 16.60 |
| Rb2 | 11.53 |
| Rc | 9.90 |
| Rd | 6.80 |
| Rg1 | 3.30 |
| Rg2 | 0.98 |
| Rf | 1.40 |
| Total | 50.51 |
 | Table IIRetention time of total
ginsenosides. |
Table II
Retention time of total
ginsenosides.
| Type | Retention time
(min) |
|---|
| Rg1 | 31.863 |
| Re | 33.853 |
| Rf | 60.595 |
| Rg2 | 65.816 |
| Rb1 | 70.277 |
| Rb2 | 82.749 |
| Rb3 | 84.051 |
| Rd | 88.941 |
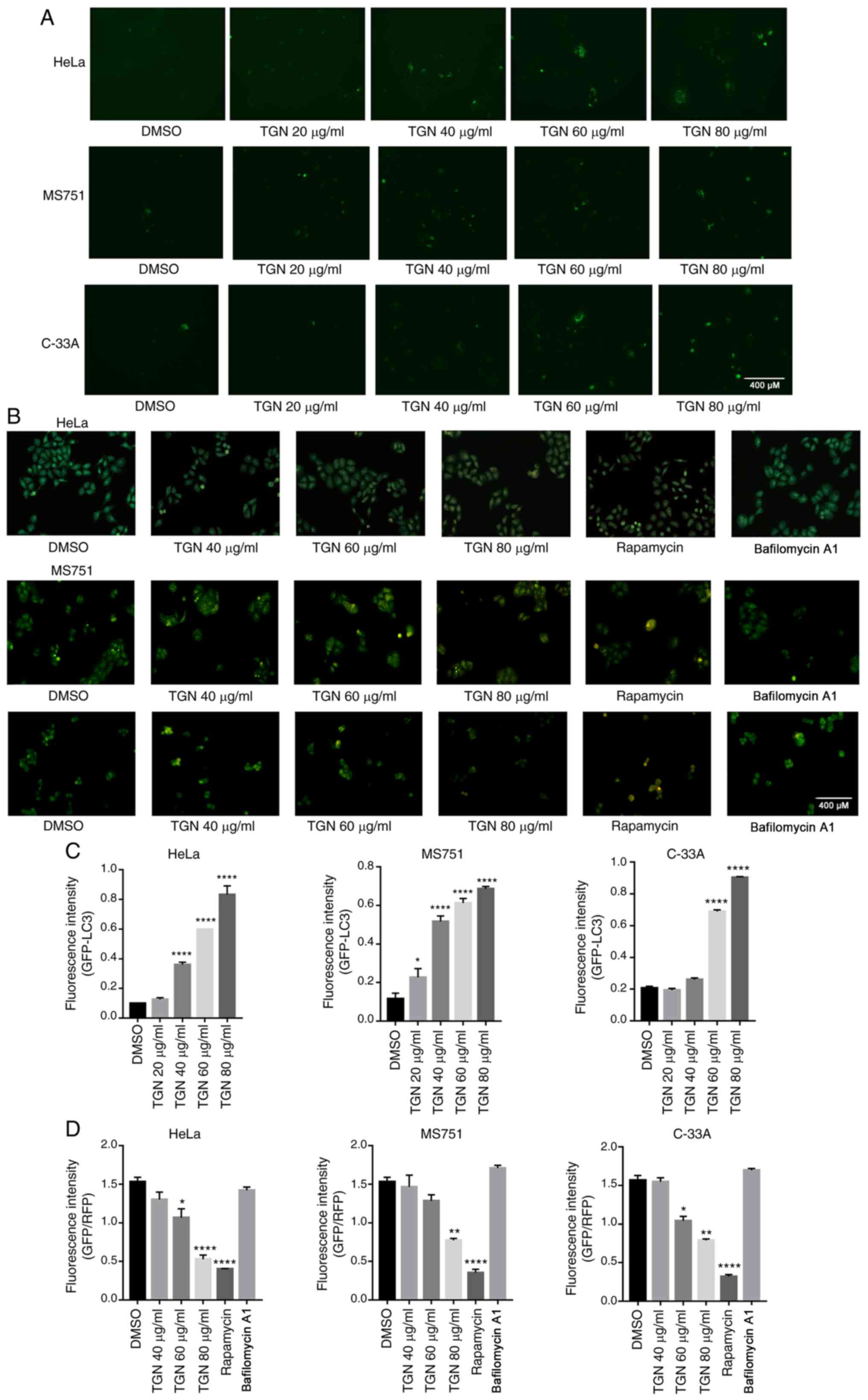
Next, an EGFP-LC3-II plasmid (500 ng) was
transfected into HeLa, MS751 and C-33A cells. After 24 h, the cells
were treated with DMSO or TGN for an additional 7 h and then
analysed for fluorescence. As shown in Fig. 2A, TGN promoted an increase in
EGFP-LC3-II puncta. To further detect autophagy activation, the
formation of AVOs in the three types of TGN-treated cervical cancer
cells was analysed by AVO staining. Fig. 2B shows that a large number of AVOs
appeared following treatment in a dose-dependent manner. These
results suggested that TGN induced cervical cancer cell autophagy
in a time- and concentration-dependent manner.
TGN increases transcription of p62
independent of autophagy
SQSTM1/p62 is a substrate for autophagy, which
should be degraded following autophagy activation. However, p62
changes can be specific to the cell type and context. Occasionally,
the expression level of p62 changes independent of autophagy
(20-22).
Furthermore, p62 may be transcriptionally upregulated under certain
conditions (23). In the present
study, western blotting demonstrated increases in p62 protein
levels following TGN treatment (Fig.
S2A). RT-qPCR analysis was performed to evaluate the
transcriptional level of p62 after TGN treatment, which confirmed
that p62 accumulation was due to transcriptional activation
(Fig. S2B). To assess the change
in p62 protein level, transcription inhibitor actinomycin D was
used to eliminate the interference caused by increased
transcription. TGN markedly decreased p62 protein levels in the
presence of actinomycin D (Fig.
S2C and D). These results
further confirmed that TGN induced autophagy and increased
transcription of p62 independent of autophagy.
The effect of TGN on autophagy
promotion is irreversible
Subsequently, whether autophagy activation was
reversible in the three cervical cancer cell lines was determined.
EBSS is used for the short-term maintenance of cells in a
CO2 environment and induces reversible autophagy
(6). As shown in Fig. 3A-C, the activation of autophagy was
not relieved even at 6 h after removal of TGN. Furthermore, the
ratio of LC3-II/I continued to increase. However, EBSS-induced
autophagy was rapidly relieved after replacing the medium. These
data indicated that the effect of TGN on autophagy was irreversible
over a short time period. The effects of ginsenoside monomers on
autophagy showed that major components of TGN promoted autophagy,
except ginsenoside Rf (Fig. 3D).
These results differed from previous studies (8), which may be due to the use of
different cell lines.
Reduction of BST-2 enhances cervical
cancer cell death
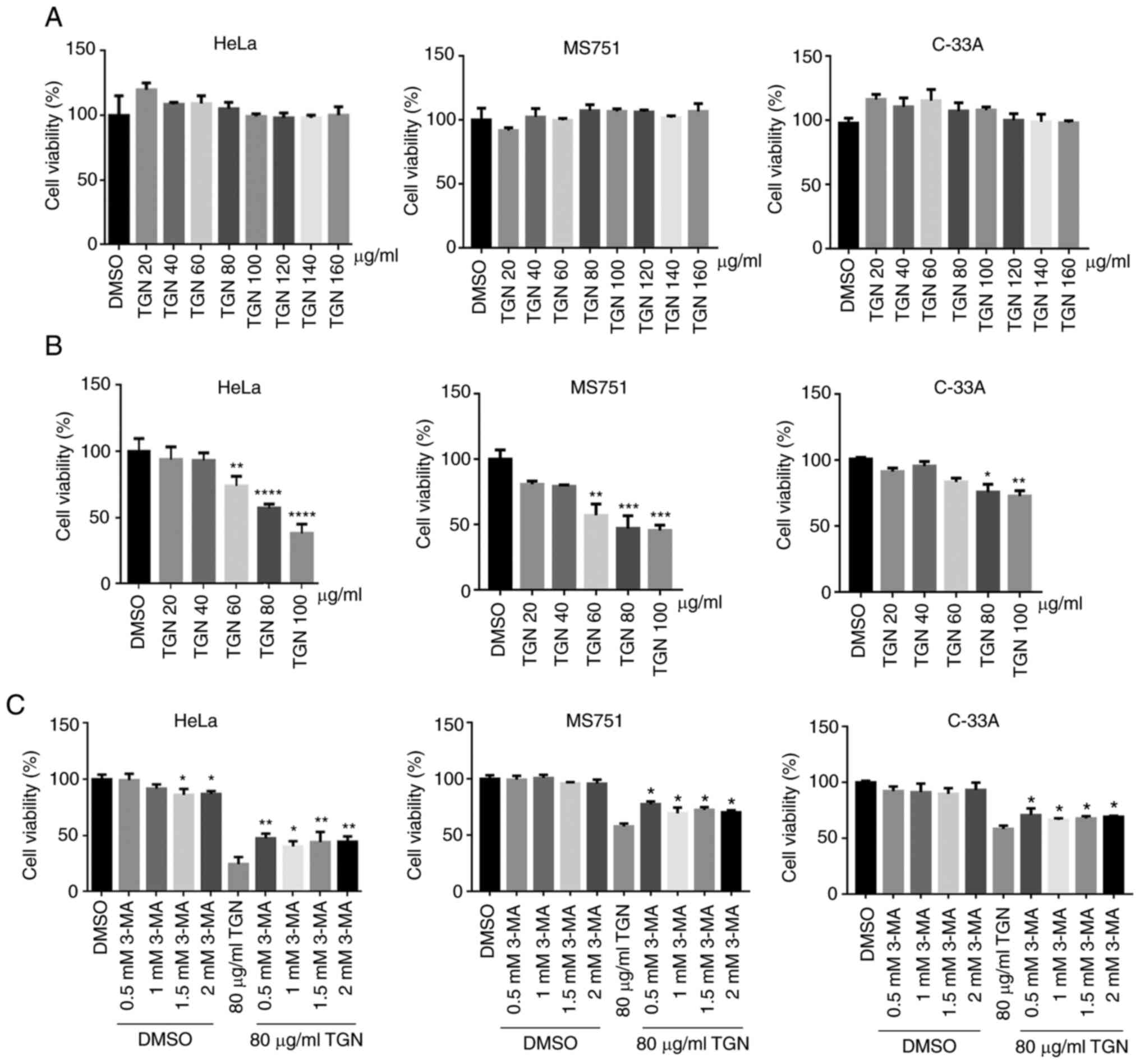
Next, the cytotoxicity of TGN in the three types of
cervical cancer cells was assessed via CCK-8 assay. TGN (~120
µg/ml) had no effect on the proliferation of HeLa, MS751 or C-33A
cells (Fig. 4A). Serum deprivation
is often used to emulate the tumour microenvironment. Therefore,
the cells were exposed to TGN under serum deprivation and then cell
viability was assessed. As a result, TGN notably suppressed cell
growth in a dose- and time-dependent manner (Fig. 4B). To determine whether cell death
was caused by the combination of nutrient deficiency and TGN,
autophagy inhibitor 3-MA was introduced into the experiment.
TGN-induced cell death was partly weakened by 3-MA, indicating that
TGN induced autophagic cell death (Fig.
4C).
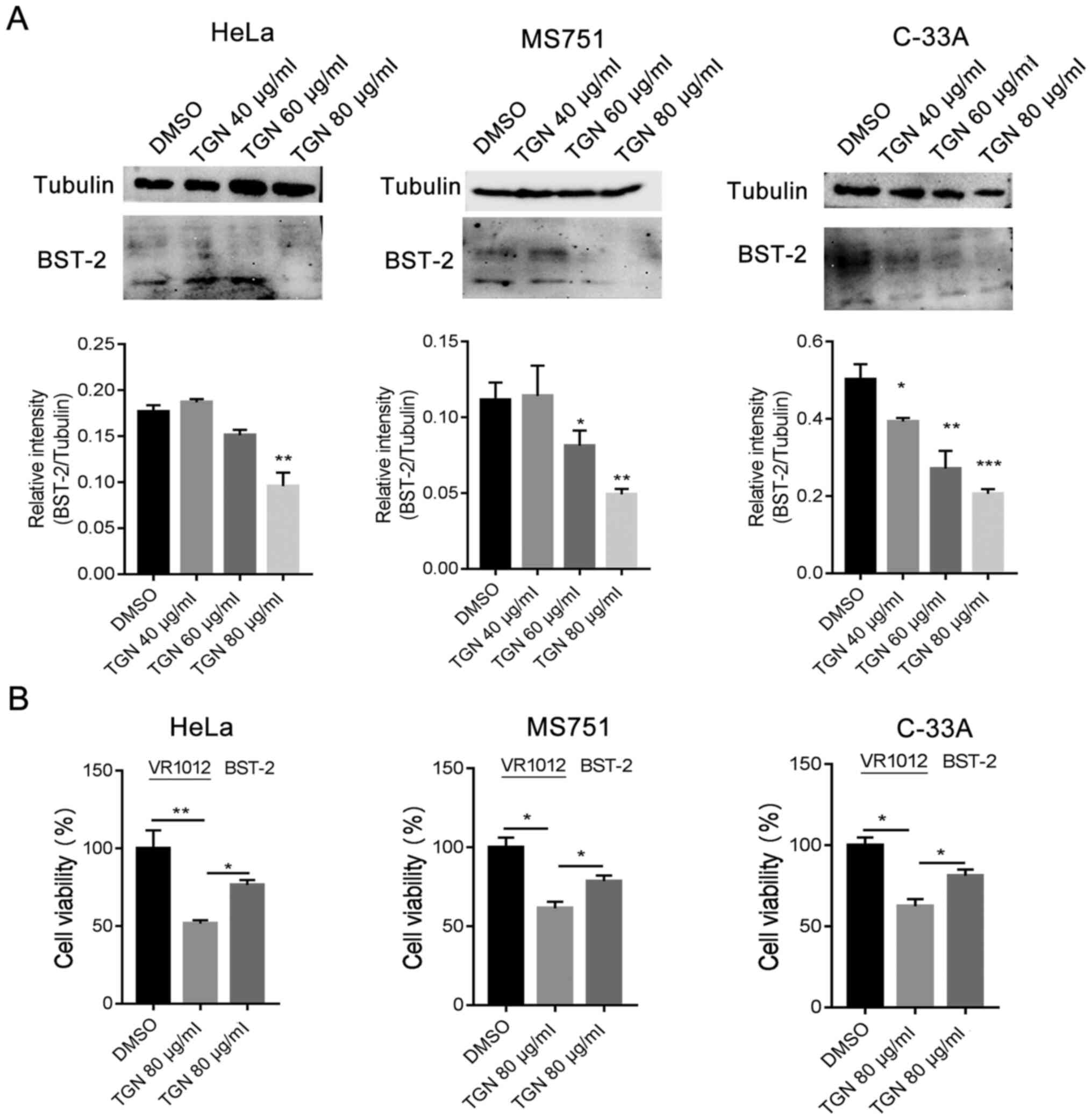
The expression of BST-2 is associated with various
types of cancer. Therefore, the protein level of BST-2 was analysed
in TGN-treated cells. The transcription and protein expression
levels of BST-2 were observed to be downregulated in TGN-treated
cervical cancer cells in normal culture (Fig. 5A). To further confirm the
association of BST-2 downregulation with cell death, a BST-2
expression plasmid was introduced into the experiment and a CCK-8
assay was used to evaluate cell viability. The result demonstrated
that upregulation of BST-2 reduced cell death in a serum-deprived
culture (Fig. 5B). These results
indicated that TGN induced downregulation of BST-2 and promoted
cervical cancer cell death in serum-deprived cultures.
Discussion
In recent years, there has been a rise in the
morbidity and mortality of cancer, presenting a global health issue
(24). Botanical medicines have
been used to treat various types of disease in Asia for thousands
of years, and ginseng is one of the most well-known and widely used
oriental medicinal plants (24).
Dried, steamed or heated ginseng is distributed in 35 countries in
various forms (24). Previous
studies have reported the beneficial effects of ginseng on
diseases, such as cancer, immune disorders, diabetes, as well as on
liver, nervous system, cardiovascular and infectious diseases
(25-31).
BST-2 is an innate immune gene that is upregulated
in various types of cancer, such as breast cancer, mammary tumours
and bladder cancer (32-34).
Dimers of BST-2 promote cell-cell and cell-matrix adhesions, cell
motility, survival and growth (35). It also protects cervical cancer
cells from serum deprivation-induced death (35). A BST-2-based peptide, known as B49
and its analogue, B49Mod1 inhibit adhesion and growth of breast
cancer cells (35,36). Therefore, targeting BST-2 presents a
potential therapeutic strategy against cancer (37-39).
BST-2 is an autophagy-associated protein. It is hypothesized to be
a substrate of autophagy. In the present study, the expression
level of BST-2 was identified to be decreased in TGN-treated cells
and accompanied by increasing cell death. The underlying mechanism
may be that BST-2 was degraded through induced autophagy and
increased the death of serum-deprived cells.
Therefore, TGN enhanced irreversible autophagy in
cervical cancer cell lines and caused significant autophagic cell
death in serum-deprived cells. TGN markedly increased the
expression of p62 at the transcriptional level, but decreased p62
protein levels in the presence of actinomycin D. Furthermore,
protein expression levels of BST-2 were downregulated by TGN and
upregulation of BST-2 reduced the cell death that was caused by
TGN. These results provide the molecular basis to develop TGN as a
promising candidate for cancer therapy.
Supplementary Material
Analysis of total ginsenoside extract
of ginseng root. (A) Standard chromatogram. (B) Sample
chromatogram.
TGN increased transcription of p62
independent of autophagy. (A) Western blot analysis with antibodies
specific for p62 and tubulin serving as the control. (B) After HeLa
cells were treated with 80 μg/ml TGN, the expression of p62 was
determined by reverse transcription-quantitative PCR. (C) After
HeLa cells were treated with TGN and DMSO or 80 nM actinomycin D
for 16 h, the expression of p62 was determined by western blotting.
(D) Statistical analysis of western blot results.
*P<0.05, **P<0.01,
***P<0.001 and ****P<0.0001 vs. DMSO.
TGN, total ginsenoside; 3-MA, 3-methyladenine; DMSO,
dimethylsulfoxide.
Acknowledgements
Not applicable.
Funding
Funding: The present study was supported by the National Key
Research and Development Program of China (grant no.
2017YFC1702104), the Key Project at Central Government Level: The
ability establishment of sustainable use for valuable Chinese
medicine resources (grant no. 2060302) and the Cultivation Fund
Project of Changchun University of Chinese Medicine (grant no.
2018KJ03).
Availability of data and materials
The datasets used and/or analysed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
JW, ML and SB designed the experiments. SB, YZ, FL,
SL, ZH and SW performed the experiments. ML, SW, XB, DZ analysed
the experiments. JW wrote the paper. SB and JW confirm the
authenticity of all the raw data. All authors read and approved the
final version of the manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Yu L, Chen Y and Tooze SA: Autophagy
pathway: Cellular and molecular mechanisms. Autophagy. 14:207–215.
2018.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Wang K: Autophagy and apoptosis in liver
injury. Cell Cycle. 14:1631–1642. 2015.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Finocchario-Kessler S, Wexler C, Maloba M,
Mabachi N, Ndikum-Moffor F and Bukusi E: Cervical cancer prevention
and treatment research in Africa: A systematic review from a public
health perspective. BMC Womens Health. 16(29)2016.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Wong AS, Che CM and Leung KW: Recent
advances in ginseng as cancer therapeutics: A functional and
mechanistic overview. Nat Prod Rep. 32:256–272. 2015.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Ahuja A, Kim JH, Kim JH, Yi YS and Cho JY:
Functional role of ginseng-derived compounds in cancer. J Ginseng
Res. 42:248–254. 2018.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Zheng K, Li Y, Wang S, Wang X, Liao C, Hu
X, Fan L, Kang Q, Zeng Y, Wu X, et al: Inhibition of
autophagosome-lysosome fusion by ginsenoside Ro via the
ESR2-NCF1-ROS pathway sensitizes esophageal cancer cells to
5-fluorouracil-induced cell death via the CHEK1-mediated DNA damage
checkpoint. Autophagy. 12:1593–1613. 2016.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Zhuang J, Yin J, Xu C, Mu Y and Lv S:
20(S)-ginsenoside Rh2 induce the apoptosis and autophagy in U937
and K562 cells. Nutrients. 10(328)2018.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Zhao M, Chen Q, Xu W, Wang H, Che Y, Wu M,
Wang L, Lijuan C and Hao H: Total ginsenosides extract induce
autophagic cell death in NSCLC cells through activation of
endoplasmic reticulum stress. J Ethnopharmacol.
243(112093)2019.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Tokarev A, Suarez M, Kwan W, Fitzpatrick
K, Singh R and Guatelli J: Stimulation of NF-κB activity by the HIV
restriction factor BST2. J Virol. 87:2046–2057. 2013.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Neil SJ, Zang T and Bieniasz PD: Tetherin
inhibits retrovirus release and is antagonized by HIV-1 Vpu.
Nature. 451:425–430. 2008.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Mahauad-Fernandez WD, DeMali KA, Olivier
AK and Okeoma CM: Bone marrow stromal antigen 2 expressed in cancer
cells promotes mammary tumor growth and metastasis. Breast Cancer
Res. 16(493)2014.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Fang KH, Kao HK, Chi LM, Liang Y, Liu SC,
Hseuh C, Liao CT, Yen TC, Yu JS and Chang KP: Overexpression of
BST2 is associated with nodal metastasis and poorer prognosis in
oral cavity cancer. Laryngoscope. 124:E354–E360. 2014.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Mukai S, Oue N, Oshima T, Mukai R,
Tatsumoto Y, Sakamoto N, Sentani K, Tanabe K, Egi H, Hinoi T, Ohdan
H and Yasui W: Overexpression of transmembrane protein BST-2 is
associated with poor survival of patients with esophageal, gastric,
or colorectal cancer. Ann Surg Oncol. 24:594–602. 2017.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Shrivastava S, Devhare P, Sujijantarat N,
Steele R, Kwon YC, Ray R and Ray RB: Knockdown of autophagy
inhibits infectious Hepatitis C Virus release by the exosomal
pathway. J Virol. 90:1387–1396. 2015.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Madjo U, Leymarie O, Frémont S, Kuster A,
Nehlich M, Gallois-Montbrun S, Janvier K and Berlioz-Torrent C:
LC3C contributes to Vpu-mediated antagonism of BST2/Tetherin
restriction on HIV-1 release through a non-canonical autophagy
pathway. Cell Rep. 17:2221–2233. 2016.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Qi AN, Mei G, Ya-Jun S, Ying Z, Rui-Luan
W, Long G, Yu-Guang Z and Dan Z: Comparative study on changes of
ginsenosides and activities of American ginseng before and after
steaming. Zhongguo Zhong Yao Za Zhi. 45:4404–4410. 2020.PubMed/NCBI View Article : Google Scholar : (In Chinese).
|
|
17
|
Bian S, Zhao Y, Li F, Lu S, Wang S, Bai X,
Liu M, Zhao D, Wang J and Guo D: 20(S)-Ginsenoside Rg3 promotes
HeLa cell apoptosis by regulating autophagy. Molecules.
24(3655)2019.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Lv M, Wang J, Zhang J, Zhang B, Wang X,
Zhu Y, Zuo T, Liu D, Li X, Wu J, et al: Epitope tags beside the
N-terminal cytoplasmic tail of human BST-2 alter its intracellular
trafficking and HIV-1 restriction. PLoS One.
9(e111422)2014.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Nakasa K, Yoshimoto Y, Nakano T, Takeshima
T, Fukuhara Y, Yasui K, Araga S, Yanagawa T, Ishii T and Nakashima
K: Transcriptional activation of p62/A170/ZIP during the formation
of the aggregates: Possible mechanisms and the role in Lewy body
formation in Parkinson's disease. Brain Res. 1012:42–51.
2004.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Bardag-Gorce F, Francis T, Nan L, Li J, He
Lue Y, French BA and Franch SW: Modifications in P62 occur due to
proteasome inhibition in alcoholic liver disease. Life Sci.
77:2594–2602. 2005.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Kuusisto E, Suuronen T and Salminen A:
Ubiquitin-binding protein p62 expression is induced during
apoptosis and proteasomal inhibition in neuronal cells. Biochem
Biophys Res Commun. 280:223–228. 2001.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Klionsky DJ, Abdelmohsen K, Abe A, Abedin
MJ, Abeliovich H, Acevedo Arozena A, Adachi H, Adams CM, Adams PD,
Adeli K, et al: Guidelines for the use and interpretation of assays
for monitoring autophagy (3rd edition). Autophagy. 12:1–222.
2016.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Wang W, Nishioka Y, Ozaki S, Jalili A, Abe
S, Kakiuchi S, Kishuku M, Minakuchi K, Matsumoto T and Sone S:
HM1.24 (CD317) is a novel target against lung cancer for
immunotherapy using anti-HM1.24 antibody. Cancer Immunol
Immunother. 58:967–976. 2009.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Shigematsu Y, Oue N, Nishioka Y, Sakamoto
N, Sentani K, Sekino Y, Mukai S, Teishima J, Matsubara A and Yasui
W: Overexpression of the transmembrane protein BST-2 induces Akt
and Erk phosphorylation in bladder cancer. Oncol Lett. 14:999–1004.
2017.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Li X, Zhang G, Chen Q, Lin Y, Li J, Ruan
Q, Chen Y, Yu G and Wan X: CD317 Promotes the survival of cancer
cells through apoptosis-inducing factor. J Exp Clin Cancer Res.
35(117)2016.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Mahauad-Fernandez WD and Okeoma CM: B49, a
BST-2-based peptide, inhibits adhesion and growth of breast cancer
cells. Sci Rep. 8(4305)2018.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Lyu Y, Mahauad-Fernandez WD and Okeoma CM:
Development and characterization of the shortest anti-adhesion
peptide analogue of B49Mod1. Molecules. 25(1188)2020.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Yokoyama T, Enomoto T, Serada S, Morimoto
A, Matsuzaki S, Ueda Y, Yoshino K, Fujita M, Kyo S, Iwahori K, et
al: Plasma membrane proteomics identifies bone marrow stromal
antigen 2 as a potential therapeutic target in endometrial cancer.
Int J Cancer. 132:472–484. 2013.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Schliemann C, Roesli C, Kamada H, Borgia
B, Fugmann T, Klapper W and Neri D: In vivo biotinylation of the
vasculature in B-cell lymphoma identifies BST-2 as a target for
antibody-based therapy. Blood. 115:736–744. 2010.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Silveira NJ, Varuzza L, Machado-Lima A,
Lauretto MS, Pinheiro DG, Rodrigues RV, Severino P and Nobrega FG:
Head and Neck Genome Project GENCAPO. Silva WA Jr, et al: Searching
for molecular markers in head and neck squamous cell carcinomas
(HNSCC) by statistical and bioinformatic analysis of larynx-derived
SAGE libraries. BMC Med Genomics. 1(56)2008.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Im K, Kim J and Min H: Ginseng, the
natural effectual antiviral: Protective effects of Korean Red
Ginseng against viral infection. J Ginseng Res. 40:309–314.
2016.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Kim YR and Yang CS: Protective roles of
ginseng against bacterial infection. Microb Cell. 5:472–481.
2018.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Benzie IFF and Wachtel-Galor S: Herbal
medicine: Biomolecular and clinical aspects. Taylor & Francis
Group, Chapter. 1:1–11. 2011.PubMed/NCBI
|
|
35
|
Lee JS, Ko EJ, Hwang HS, Lee YN, Kwon YM,
Kim MC and Kang SM: Antiviral activity of ginseng extract against
respiratory syncytial virus infection. Int J Mol Med. 34:183–190.
2014.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Radad K, Gille G, Liu LL and Rausch WD:
Use of ginseng in medicine with emphasis on neurodegenerative
disorders. J Pharmacol Sci. 100:175–186. 2006.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Yoo DG, Kim MC, Park MK, Song JM, Quan FS,
Park KM, Cho YK and Kang SM: Protective effect of Korean red
ginseng extract on the infections by H1N1 and H3N2 influenza
viruses in mice. J Med Food. 15:855–862. 2012.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Kim S, Lee Y and Cho J: Korean red ginseng
extract exhibits neuroprotective effects through inhibition of
apoptotic cell death. Biol Pharm Bull. 37:938–946. 2014.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Mahauad-Fernandez WD, Naushad W, Panzner
TD, Bashir A, Lal G and Okeoma CM: BST-2 promotes survival in
circulation and pulmonary metastatic seeding of breast cancer
cells. Sci Rep. 8(17608)2018.PubMed/NCBI View Article : Google Scholar
|