Introduction
The incidence of diabetes is increasing on a yearly
basis worldwide and is frequently attributed to changes in
lifestyle (1). Almost 50% of
patients with diabetes develop peripheral neuropathy (2). Diabetic neuropathic pain (DNP) is one
of the most serious complications of diabetic peripheral neuropathy
(3). The majority of patients with
DNP experience moderate to severe levels of pain (4). There are various dysregulated
mechanisms that may contribute to the pathogenesis of DNP,
including hyperglycemia, advanced glycation end-products, oxidative
stress, neuroinflammation and endoneural hypoxia (5-7).
However, drugs acting on these pathways have limited therapeutic
effects in clinical practice. Therefore, identifying alternative
mechanisms by which DNP manifests should be explored further.
Non-coding RNAs (ncRNAs) are RNA molecules that do
not encode proteins but functionally regulate protein expression
(8). An increasing number of
studies have indicated that ncRNAs are involved in gene
transcription and translation under physiological and pathological
conditions (9,10). ncRNAs may be divided into three
categories according to their size: Small ncRNAs, <200
nucleotides (nt); long ncRNAs (lncRNAs), >200 nt; and circular
RNAs (circRNA) consisting of a closed continuous loop (11). Several studies have indicated that
ncRNAs have a crucial role in several types of pain, including
neuropathic pain (12,13). Studies have also provided evidence
that ncRNAs regulate the occurrence of DNP. For instance, microRNA
(miRNA/miR)-190a (14) and miR-193a
(15) in the dorsal root ganglion
have been indicated to be associated with the induction of DNP.
Furthermore, lncRNA NONRATT021972 and lncRNA BC168687 have been
indicated to regulate DNP (16).
circRNAs are another type of ncRNA that interact with miRNAs
(17,18), which regulate gene expression via a
circRNA/miRNA/mRNA network (19). A
recent study also suggested that altered expression levels of
circRNAs accompany the development of neuropathic pain (20). However, to date, the regulatory
functions and underlying mechanisms of ncRNAs in DNP have not been
systematically reported, to the best of our knowledge. Thus,
comprehensive analysis of the ncRNA expression profiles and their
association with the pathogenesis of DNP may facilitate the
development of effective methods to treat this disease.
In the present study, the expression profiles of
ncRNAs in the spinal cord of mice with streptozotocin (STZ)-induced
DNP were examined and analyzed using RNA sequencing techniques. The
microarray results were subjected to bioinformatics predictions,
including Gene Ontology (GO) and Kyoto Encyclopedia of Genes and
Genomes (KEGG) pathway analyses.
Materials and methods
Animals
All experiments were approved by the Animal Use and
Care Committee for Research and Education of The First People's
Hospital of Foshan (Foshan, China) and were in accordance with the
guidelines described in the International Association for the Study
of Pain (21). The animal
experiments performed in the present study were performed in
compliance with the original ARRIVE guidelines.
Periodic changes in gonadal hormone levels may
affect pain (22). Therefore, 12
male animals were used in the present study. Experiments were
performed on adult male C57BL/6 mice (age, 8 weeks; weight, 25-30
g; obtained from the Center of Laboratory Animal Science of
Guangdong), and were housed at a constant ambient temperature of
21±2˚C and relative humidity of 55±5% with a 12-h light/dark cycle
and ad libitum access to food and water. The study lasted
for 6 weeks. During the entirety of the experimental procedure,
staff evaluated the health of the animals every day. Every effort
was made to ensure the welfare of the animals, including ensuring
sufficient water and food, clean living conditions, suitable
temperature and light conditions, and death after anesthesia. When
the animals became infected or were unable to eat, the experiment
was terminated and the animal was euthanized by administering an
overdose of pentobarbital sodium (100 mg/kg, i.p.) followed by
cervical dislocation.
STZ-induced DNP model
A total of 12 adult male mice were randomly divided
into two groups: N, control mice; and D, DNP mice (n=6 per group),
and all mice were fasted for >12 h prior to injection of
STZ/citrate buffer. Diabetes mellitus was induced by a single i.p.
injection of STZ (150 mg/kg; Sigma-Aldrich; Merck KGaA) freshly
dissolved in citrate buffer (pH=4.5). Mice in the control group
were injected with an equivalent volume of the vehicle. Diabetes
mellitus was defined as hyperglycemia with a plasma glucose
concentration of >300 mg/dl (16.7 mmol/l) 3 days after STZ
injection. STZ-injected animals were removed from the study if they
did not exhibit hyperglycemia at 3 days after injection. Blood
glucose levels were measured on a weekly basis to confirm continued
hyperglycemia. Blood samples were obtained from the caudal vein and
body weight was monitored weekly throughout the experiment.
According to a previous study (14), mice should present with mechanical
allodynia 6 weeks after STZ injection. In the present study, the
mice were left for 6 weeks to allow for the development of
neuropathic pain following the STZ injection.
Mechanical sensitivity test
An investigator blinded to the treatments of the
mice performed the behavioral tests in a dedicated quiet room under
constant conditions. To quantify the mechanical sensitivity of the
hind paws, the paw withdrawal threshold (PWT) in response to
mechanical stimuli was measured in mice. The behavioral tests were
performed 1 day prior to STZ or vehicle injection (baseline) and
then on a weekly basis for 6 weeks following STZ injection. The
method of assessing mechanical allodynia was performed as described
previously (23). Animals were
placed in a plexiglas chamber with a 4x3 mm wire mesh grid floor
and allowed to acclimatize for 30 min. Calibrated von Frey
filaments of different scales (g) were applied perpendicularly to
the plantar surface of the right hind paw with sufficient force to
bend the filament for 6 sec or until the paw was withdrawn. Rapid
withdrawal or paw flinching was interpreted as a positive response.
If there was no response, the next higher force filament was
applied. Following a response, the next lower force filament was
applied.
Tissue collection and RNA
isolation
There were 12 mice in both the STZ and vehicle
injection groups (6 per group). All animals survived during the
study. In order to ensure the stability of the experimental model,
preliminary experiments were performed prior to the formal
experiments. It has been reported that mice may die after
establishing a diabetes model (24). Therefore, the final experiments
consisted of 6 mice in each group, and the mice were numbered for
further random selection. Finally, 3 mice were randomly selected
from each group for statistical analysis. A total of 42 days after
STZ/vehicle injection, 3 mice in each group were euthanized using
pentobarbital sodium (100 mg/kg, i.p.) followed by cervical
dislocation after the final behavioral test. The L4-5 spinal cord
tissues were rapidly removed and stored at -80˚C until
required.
RNA isolation and RNA
quantification
RNA degradation and contamination was monitored on
1% agarose gels. According to the manufacturer's protocol, each
tissue sample was washed three times using cold PBS and 1 ml
TRIzol® reagent was added (Thermo Fisher Scientific,
Inc.) to extract the RNA. The RNA concentration was measured using
the Qubit® RNA assay kit in a Qubit® 2.0
Fluorometer (Thermo Fisher Scientific, Inc.). RNA integrity was
verified using an RNA Nano 1000 assay kit for the Bioanalyzer 2100
system (Agilent Technologies, Inc.). The method for determining the
levels of lncRNAs and miRNAs was the same as that used for mRNAs.
For the quantification of circRNAs, exonuclease was used to exclude
non-circRNAs. The RNA was divided into two copies. Linear RNA was
digested with RNase R (cat. no. RNR07250; Epicentre; Illumina,
Inc.) to leave only the circRNAs. The other half of the sample from
the same RNA extraction was not treated with RNase R. The two
samples of RNA were reverse transcribed according to a previous
study (25). The sample treated
with RNase R was used to examine the expression of circRNAs and the
other sample that was not treated with RNase R was used to measure
the expression of β-actin.
Library construction and RNA
sequencing
RNA sequencing was performed by Aksomics Inc. A
total of 2 µg total RNA from each sample was used for the
construction of the sequencing library. According to the
manufacturer's protocol, sequencing libraries were built using
ribosomal (r)RNA-depleted RNA with an NEB Next® Ultra™
Directional RNA Library Prep Kit for Illumina® (New
England BioLabs, Inc.). First, the NEB 3' SR Adaptor was directly
ligated to the 3' end of the miRNAs. Subsequently, the SR RT Primer
was used to hybridize the excess of 3' SR Adaptor and the
single-stranded DNA adaptor was transformed into a double-stranded
(ds)DNA molecule. dsDNA cannot ligate to the 5' SR Adaptor in the
next ligation step. The 5' end adapter was then ligated to the 5'
ends of the miRNAs. Moloney murine leukemiavirus reverse
transcriptase was used to synthesize first-strand complementary
DNA. PCR amplification was performed using LongAmp Taq 2X Master
Mix, SR Primer for Illumina and index (X) primer. PCR products were
purified by 8% SDS-PAGE (100 V, 80 min). DNA fragments corresponding
to 140-160 bp (the length of small noncoding RNA plus the 3' and 5'
adaptors) were recovered and dissolved in 8 µl elution buffer.
Finally, library quality was assessed on the Agilent Bioanalyzer
2100 system using DNA High Sensitivity Chips. The method for
identifying circRNA in each sample was conducted according to a
previous study (26).
GO annotations and KEGG pathway
analysis
Fold change (FC) and false discovery rate (FDR) were
used to filter DE genes under the following criteria: i) FC >1.5
or <0.5; and ii) FDR<0.05. GO annotations and KEGG pathway
analysis were performed to predict the roles of the DE mRNAs and
miRNAs. In brief, GO analysis was used to establish genetic
regulatory networks of interest of the differentially expressed
genes in the GO categories molecular function, cellular component
and biological process (geneontology.org). Pathway analysis was performed to
select the significant pathways of the differentially expressed
genes, according to the KEGG database (genome.jp/kegg/).
Statistical analysis
Values are expressed as the mean ± standard error of
the mean. The results of the paw withdrawal thresholds were
statistically analyzed using repeated-measures ANOVA in SPSS
version 16.0 (SPSS, Inc.). Bonferroni corrections were used for
further comparison following ANOVA. A Kolmogorov-Smirnov test and
P-P graph were used to test the sample data for normality of
distribution and datasets with P>0.05 were considered to be
normally distributed. In addition, Levene's test was used to
analyze the homogeneity of variance of the data. All measurement
data were normally distributed. The blood glucose levels were
analyzed using mixed two-way ANOVA with Bonferroni's test.
P<0.05 was considered to indicate a statistically significant
difference.
Results
Changes in blood glucose levels and
PWT of mice with DNP
Blood glucose levels were assessed weekly throughout
the study. A total of 3 days after STZ injection, diabetic mice
presented with significantly increased blood glucose levels
(5.0±1.4 mmol/l at baseline vs. 23.5±2.7 mmol/l 3 days after STZ
administration) and this was maintained throughout the experiment
(23.9±3.1 mmol/l on day 42) (Table
I). The mice in the control group did not exhibit
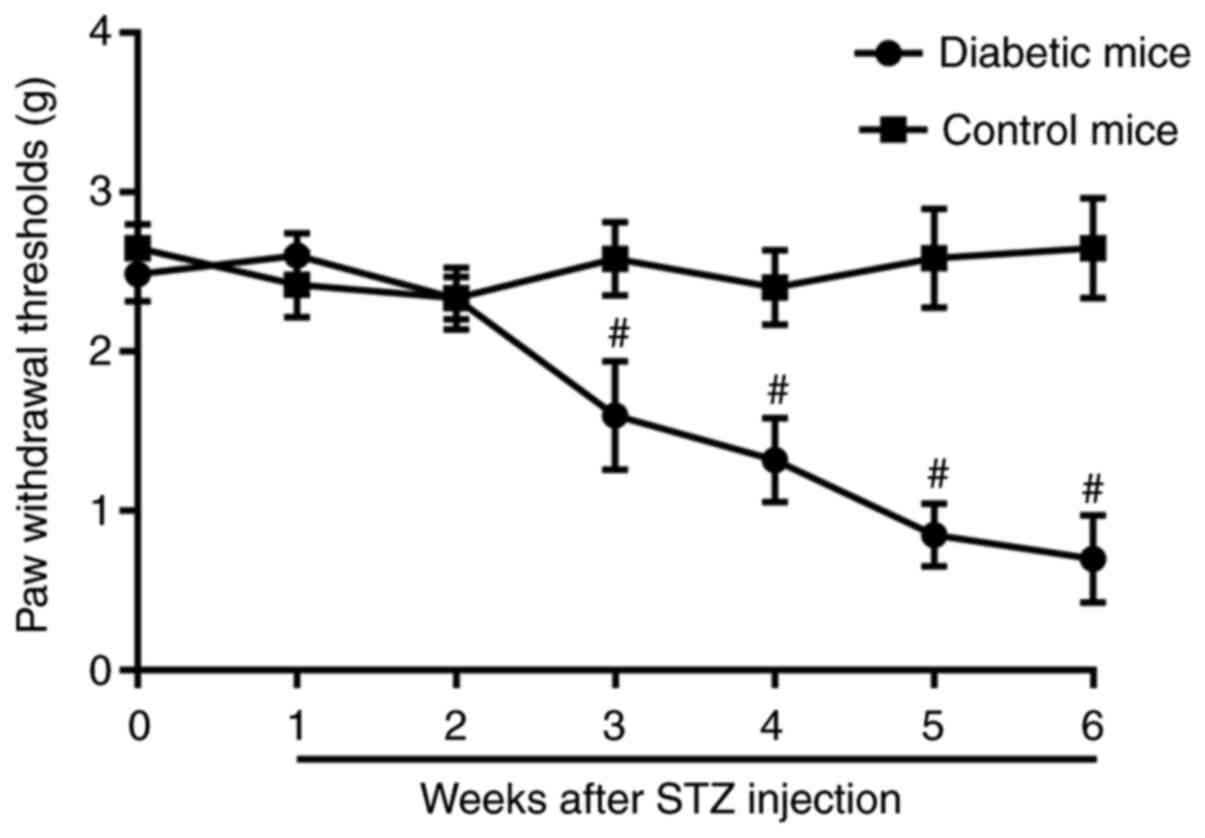
hyperglycemia. Mice with STZ-induced diabetes exhibited gradually
decreasing PWT values over the 6-week period compared with the
baseline (Fig. 1). In the
non-diabetic mice, the PWT did not vary during the 6-week period
(Fig. 1).
 | Table IBlood glucose levels of the mice
during the experiment (mmol/l). |
Table I
Blood glucose levels of the mice
during the experiment (mmol/l).
| | Days after
streptozotocin injection |
|---|
| Group | Baseline | 3 | 7 | 14 | 21 | 28 | 35 | 42 |
|---|
| C | 5.6±0.1.1 | 5.00±1.4 | 6.6±1.3 | 6.5±1.5 | 6.9±1.2 | 6.4±1.5 | 5.5±1.6 | 5.9±1.3 |
| D | 5.8±1.4 |
23.5±2.7a |
23.0±2.5a |
21.4±2.6a |
26.4±2.8a |
27.5±2.8a |
27.4±2.9a |
23.9±3.1a |
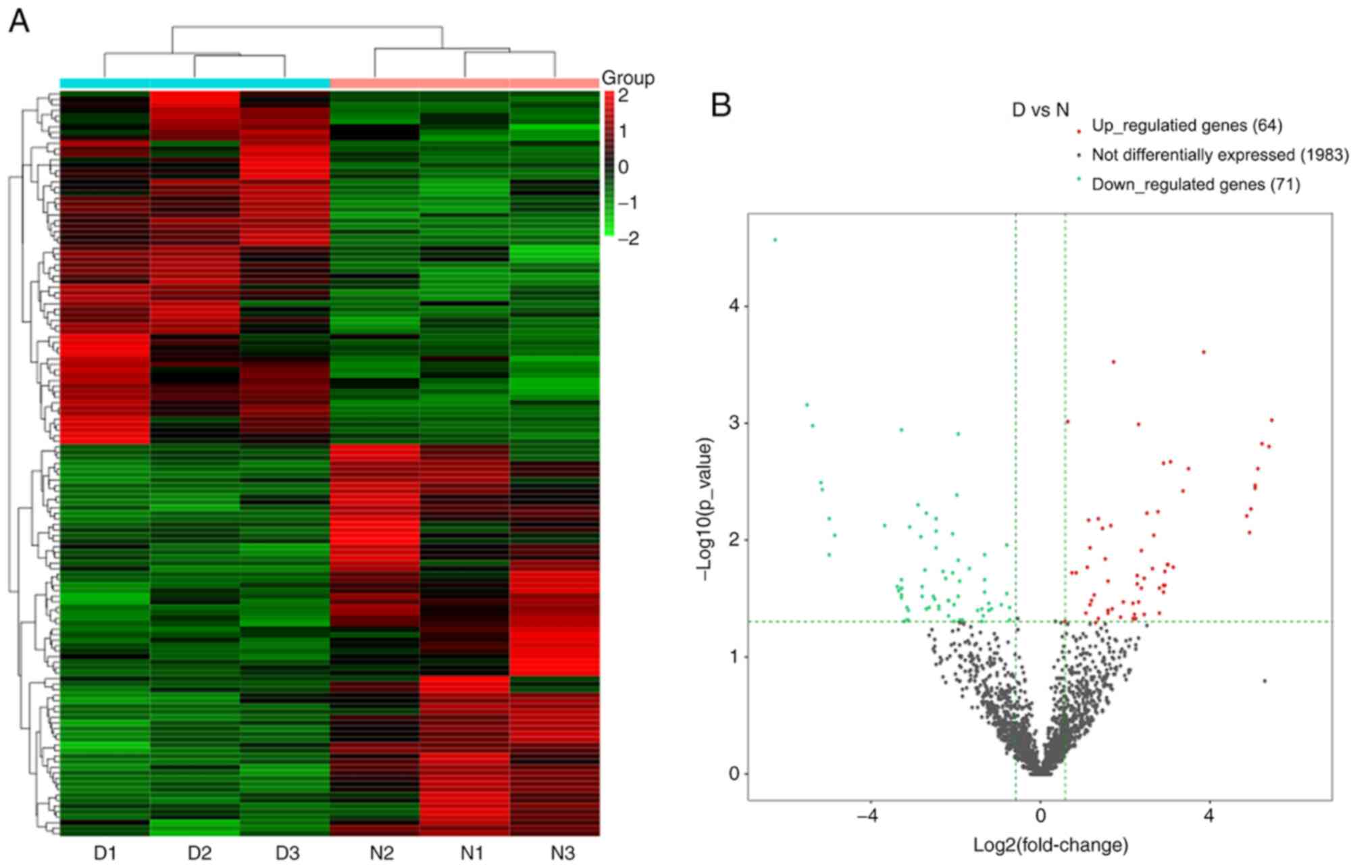
Expression profile of the coding
genes
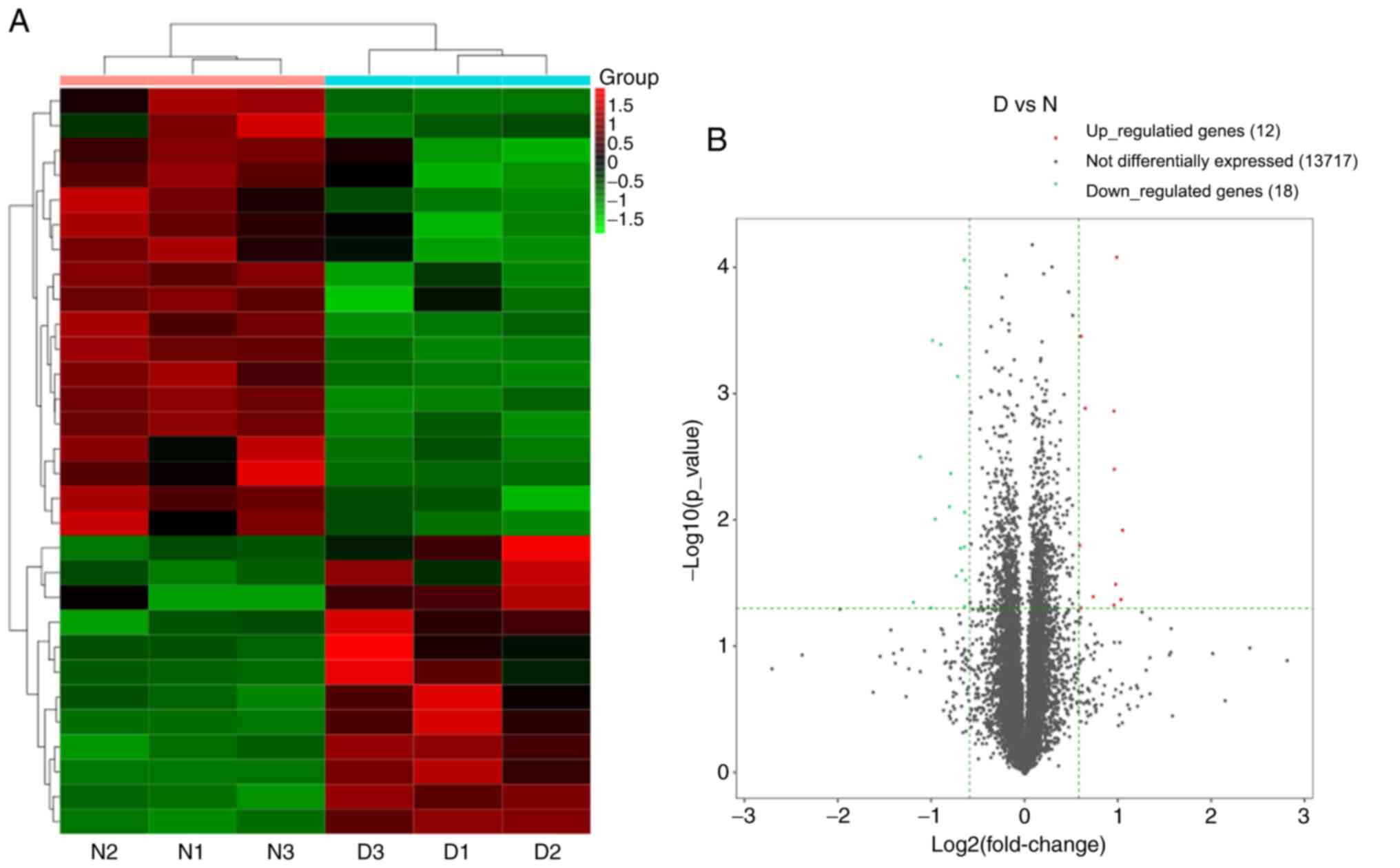
A total of 13,747 mRNAs were detected and 30 were
differentially expressed in the spinal cord tissues between the DNP
and control groups. At 42 days after STZ injection, there were 12
upregulated mRNAs and 18 downregulated mRNAs in the DNP group
compared with those in the control group. The top 10 upregulated
and downregulated mRNAs in the DNP group compared with the control
group 42 days after STZ injection are listed in Table II. Fig.
2A and B presents the heat map
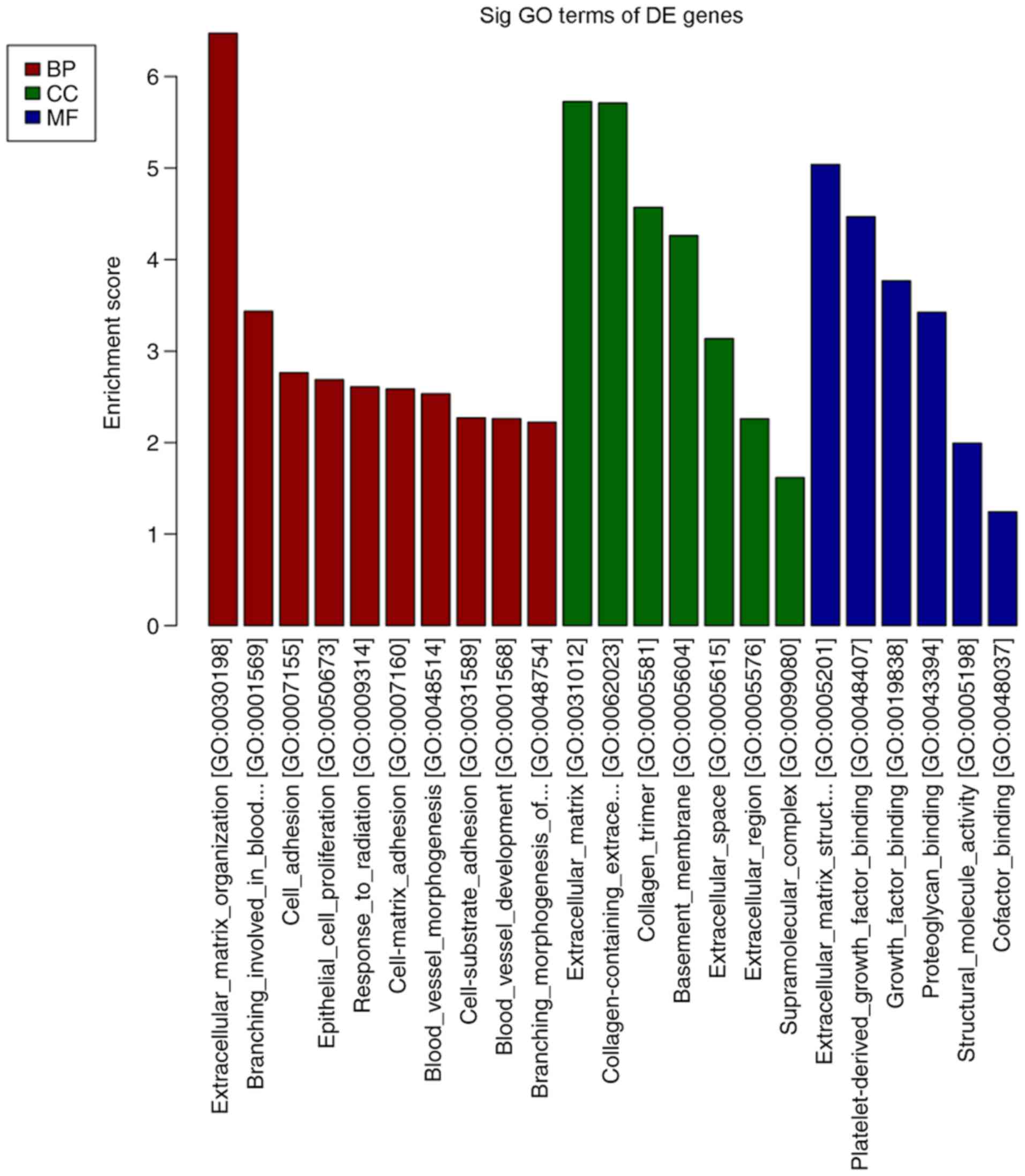
and volcano plot of the DE mRNAs, respectively. Differentially
expressed genes were primarily involved in GO molecular function
terms of ‘receptor ligand activity’, ‘growth factor binding’ and
‘extracellular matrix structural constituent’ based on the GO
analyses (Fig. 3). The products of
DE genes were primarily located in the extracellular matrix based
on GO cellular component analyses (Fig.
3). GO analysis in the category biological process indicated
that ‘hormone metabolic process’, ‘regulation of hormone levels’,
‘regulation of signaling receptor activity’, ‘cell adhesion’,
‘extracellular matrix organization’ and ‘branching involved in
blood vessel morphogenesis’ were the most enriched processes
amongst the DE mRNAs (Fig. 3). The
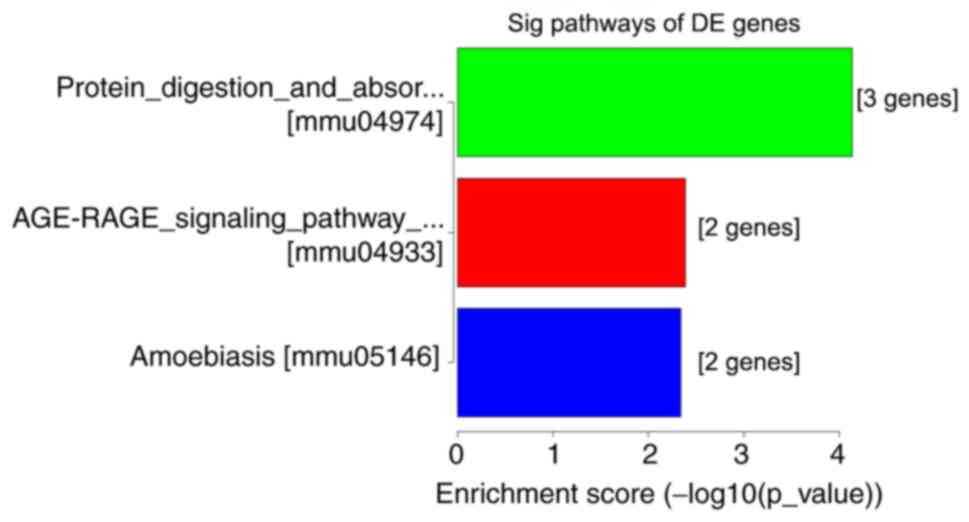
above-mentioned functional terms were all closely associated with
neuropathic pain. KEGG pathway analyses suggested that ‘protein
digestion and absorption pathway’, ‘amoebiasis’ and the ‘AGE-RAGE
signaling pathway in diabetic complications’ were most enriched
amongst the DE genes (Fig. 4).
 | Table IIDetailed information of the top 10
upregulated and 10 downregulated mRNAs. |
Table II
Detailed information of the top 10
upregulated and 10 downregulated mRNAs.
| mRNA | Fold change | P-value | Direction of
regulation |
|---|
| Mup3 | 2.072998732 | 0.011983219 | Up |
| Ttr | 2.041955888 | 0.042207033 | Up |
| BC030500 | 1.972604837 | 8.2096E-05 | Up |
| Gm21320 | 1.961106079 | 0.032707565 | Up |
| Gm28036 | 1.944253915 | 0.003957426 | Up |
| Ppp1cb | 1.942959398 | 0.001371697 | Up |
| Srd5a2 | 1.94003704 | 0.046905962 | Up |
| Ccl21b | 1.657051771 | 0.040197116 | Up |
| Gm45713 | 1.565600467 | 0.001308144 | Up |
| Tomm40l | 1.523968534 | 0.000350659 | Up |
| Col3a1 | 0.438021721 | 0.045039029 | Down |
| Sema5a | 0.461710683 | 0.003190532 | Down |
| Pcdhga2 | 0.497370882 | 0.049717412 | Down |
| Mfap4 | 0.50342587 | 0.000375246 | Down |
| Gm27029 | 0.513535783 | 0.009984364 | Down |
| Vkorc1l1 | 0.533680568 | 0.000410166 | Down |
| Slc38a5 | 0.573498342 | 0.007919411 | Down |
| Igfbp4 | 0.577354961 | 0.004239532 | Down |
| Col4a1 | 0.60148317 | 0.027653137 | Down |
| Btbd2 | 0.607906603 | 0.000738708 | Down |
miRNA expression in DNP
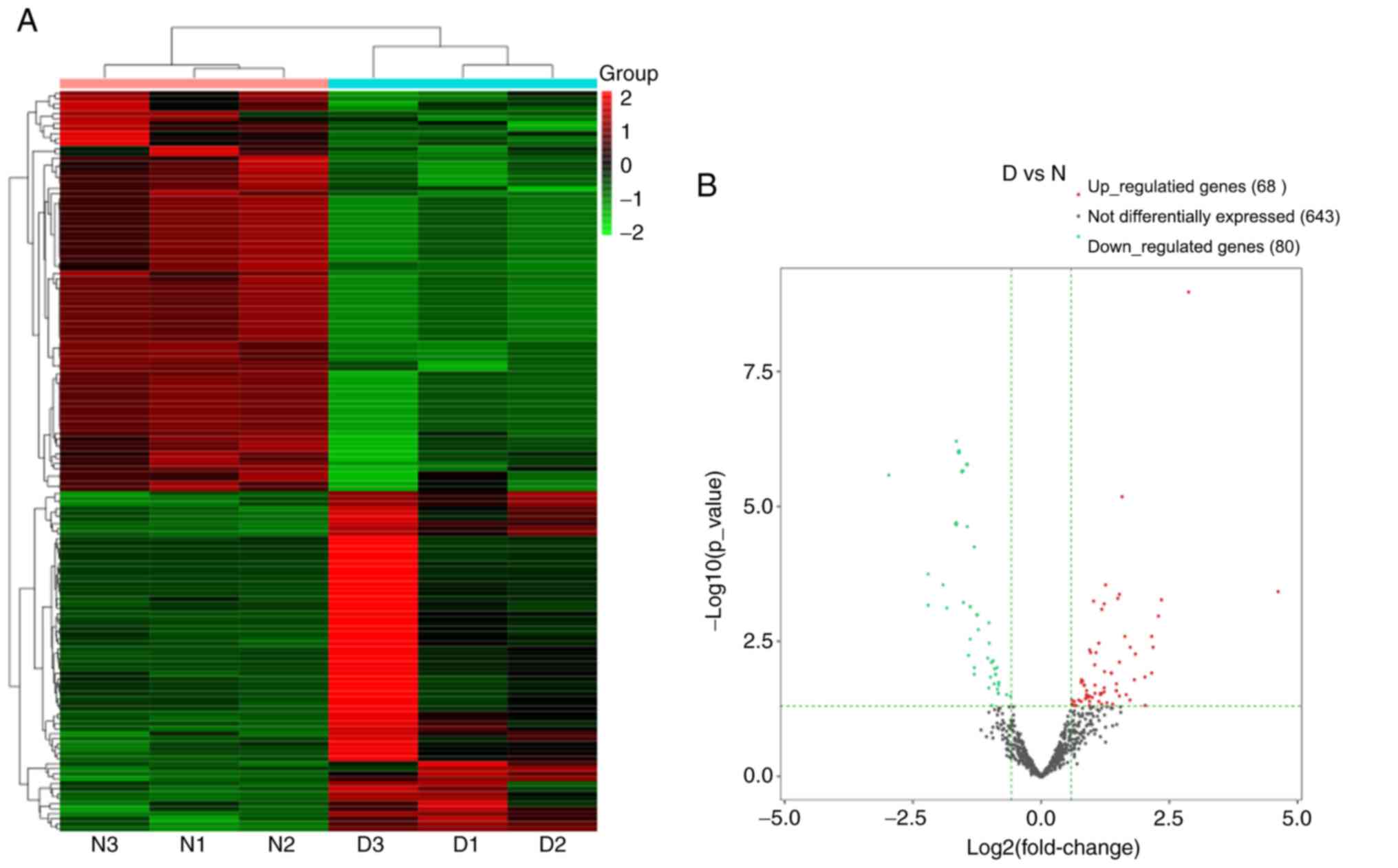
A total of 791 miRNAs detected and 148 miRNAs were
differentially expressed in the DNP group compared with those in
the control group. At 42 days after STZ injection, 68 upregulated
and 80 downregulated miRNAs were detected. Fig. 5A and B present the heat map and volcano plot of
the DE miRNAs, respectively. The top 10 upregulated and
downregulated miRNAs in the DNP group compared with the control
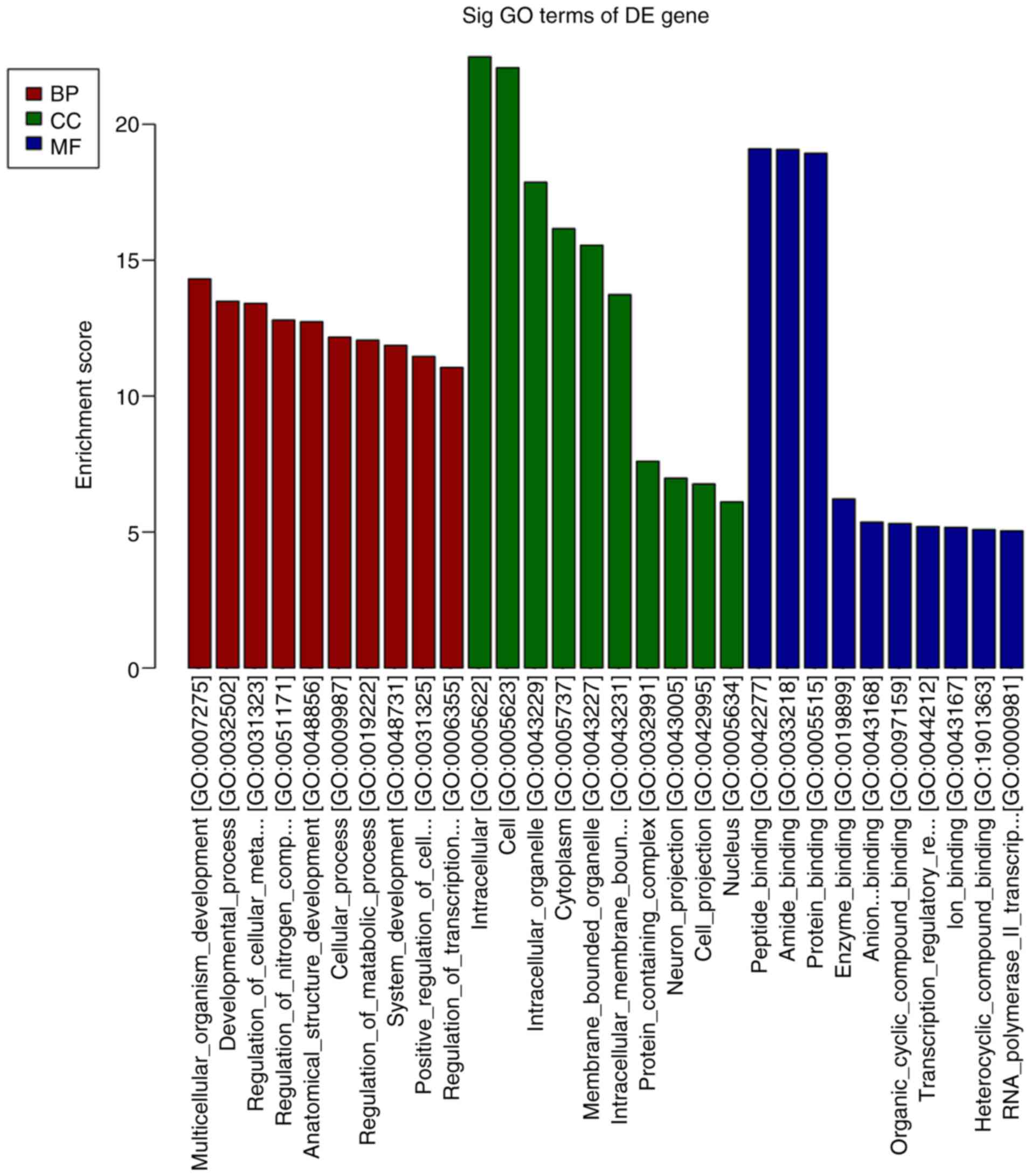
group 42 days after STZ injection are listed in Table III. The products of the target
genes of the DE miRNAs were primarily located ‘intracellular and
cell’ in the GO cellular component analysis (Fig. 6). GO analysis in the category
biological process indicated that ‘multicellular organism
development’, ‘developmental process’ and ‘regulation of cellular
metabolic process’ were the most enriched processes amongst the DE
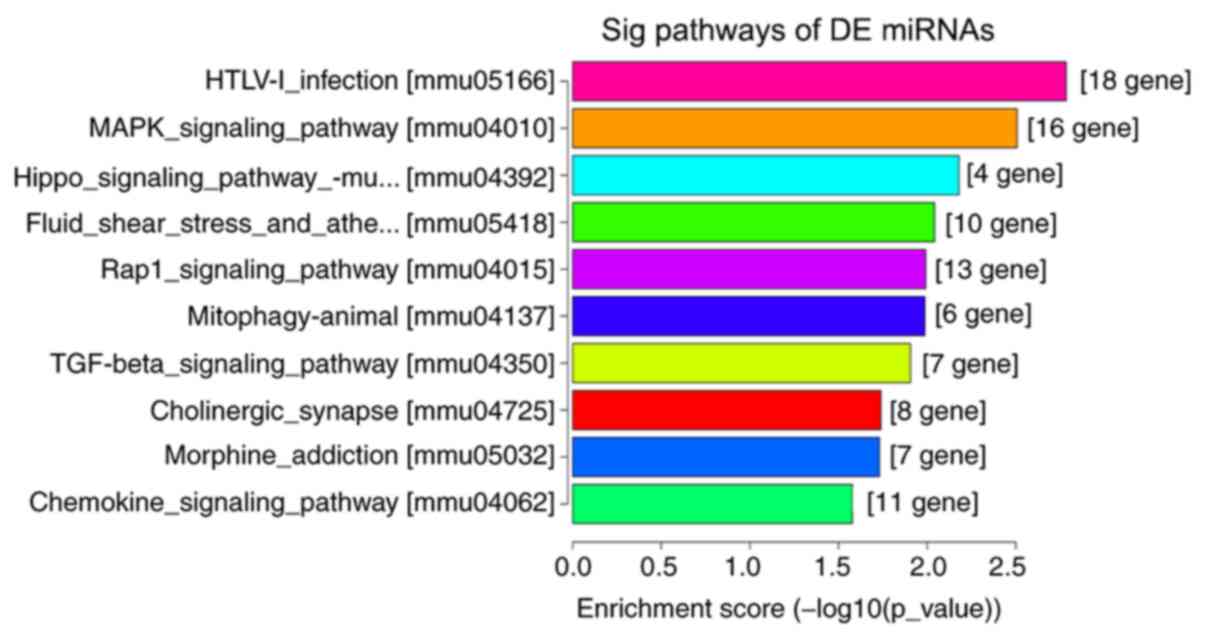
miRNA target genes (Fig. 6). KEGG
pathway analyses suggested that ‘regulation of actin cytoskeleton’,
‘cell adhesion molecules’, ‘Rap1 signaling pathway’, ‘human
T-lymphotropic virus-I infection’ and the ‘MAPK signaling pathway’
were the most enriched pathways among the DE genes (Fig. 7).
 | Table IIIDetailed information of the top 10
upregulated and 10 downregulated miRNAs. |
Table III
Detailed information of the top 10
upregulated and 10 downregulated miRNAs.
| miRNA | Fold change | P-value | Direction of
regulation |
|---|
| mmu-miR-122-5p | 24.70639724 | 0.000371545 | Up |
| mmu-miR-3474 | 7.377313224 | 0.000407508 | Up |
| mmu-miR-342-5p | 5.132108479 | 0.000523721 | Up |
|
mmu-miR-376a-3p | 4.896686217 | 0.001069652 | Up |
| mmu-miR-664-5p | 4.514300346 | 0.004094473 | Up |
| mmu-miR-29a-5p | 4.45171166 | 0.002624442 | Up |
|
mmu-miR-200a-5p | 4.440473679 | 0.011984776 | Up |
| mmu-miR-378b | 4.102131532 | 0.048569546 | Up |
| mmu-miR-491-5p | 4.064537867 | 0.01467785 | Up |
|
mmu-miR-218-1-3p | 3.584196558 | 0.005336551 | Up |
|
mmu-miR-669b-3p | 0.128138311 | 0.000262216 | Down |
|
mmu-miR-467c-3p | 0.217508283 | 0.000678429 | Down |
|
mmu-miR-467e-3p | 0.218846498 | 0.000182836 | Down |
| mmu-miR-215-5p | 0.267746723 | 0.000290494 | Down |
|
mmu-miR-3083-5p | 0.278361768 | 0.000762342 | Down |
|
mmu-miR-467d-3p | 0.31631178 | 0.000199364 | Down |
|
mmu-miR-467a-3p | 0.316889919 | 0.000205482 | Down |
|
mmu-miR-466a-3p | 0.317690341 | 0.000612554 | Down |
|
mmu-miR-466e-3p | 0.330358608 | 0.000939574 | Down |
|
mmu-miR-466b-3p | 0.330809385 | 0.000958902 | Down |
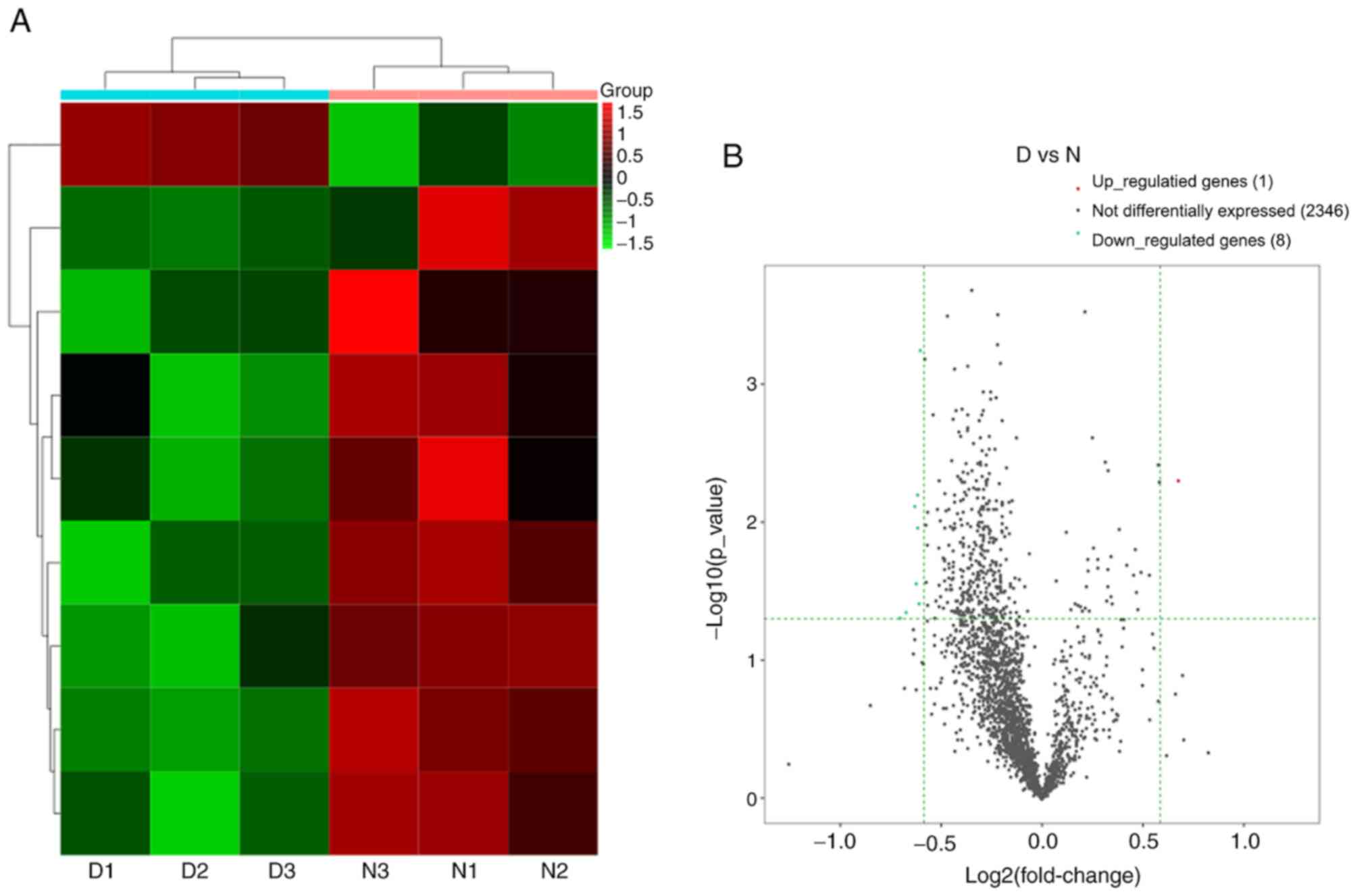
lncRNA expression in DNP
A total of 2,355 lncRNAs detected and 9 lncRNAs were
differentially expressed in the DNP group compared with the control
group. There were 1 upregulated lncRNA and 8 downregulated lncRNAs
at 42 days after STZ injection. Fig.
8A and B provides the heat map
and volcano plot of the DE lncRNAs, respectively. Detailed
information on the DE lncRNAs is listed in Table IV.
 | Table IVDetailed information on the
upregulated and downregulated long non-coding RNAs. |
Table IV
Detailed information on the
upregulated and downregulated long non-coding RNAs.
| Track ID | Gene name | Locus | Log2 (fold
change) | Fold change | P-value | Direction of
regulation |
|---|
|
ENSMUSG00000099759.1 | 1700030C10Rik |
chr12:20804381-20815779 | 0.675036717 | 1.596637407 | 0.005038816 | Up |
|
ENSMUSG00000084894.1 | Gm13834 |
chr6:31087609-31087912 | -0.703372524 | 0.614134892 | 0.049325442 | Down |
|
ENSMUSG00000099521.1 | Gm28309 |
chr2:74683446-74694194 | -0.670740279 | 0.628184269 | 0.045688601 | Down |
|
ENSMUSG00000109359.1 | Gm44797 |
chr8:9595109-9596945 | -0.627678166 | 0.647217191 | 0.007719935 | Down |
|
ENSMUSG00000108123.1 | Gm43884 |
chr6:45329238-45329796 | -0.623406223 | 0.649136497 | 0.028225881 | Down |
|
ENSMUSG00000102296.1 | Gm37543 |
chr1:25284218-25285248 | -0.614160855 | 0.653309781 | 0.006376142 | Down |
|
ENSMUSG00000105791.1 | Gm43341 |
chr5:48978278-48979585 | -0.613953524 | 0.653403676 | 0.011026114 | Down |
|
ENSMUSG00000103331.1 | Gm37995 |
chr6:40026894-40028607 | -0.608815799 | 0.655734725 | 0.039158326 | Down |
|
ENSMUSG00000085638.1 | Gm15521 |
chr9:29590484-29592510 | -0.605274191 | 0.657346436 | 0.000570587 | Down |
|
ENSMUSG00000084894.1 | Gm13834 |
chr6:31087609-31087912 | -0.703372524 | 0.614134892 | 0.049325442 | Down |
|
ENSMUSG00000099521.1 | Gm28309 |
chr2:74683446-74694194 | -0.670740279 | 0.628184269 | 0.045688601 | Down |
|
ENSMUSG00000109359.1 | Gm44797 |
chr8:9595109-9596945 | -0.627678166 | 0.647217191 | 0.007719935 | Down |
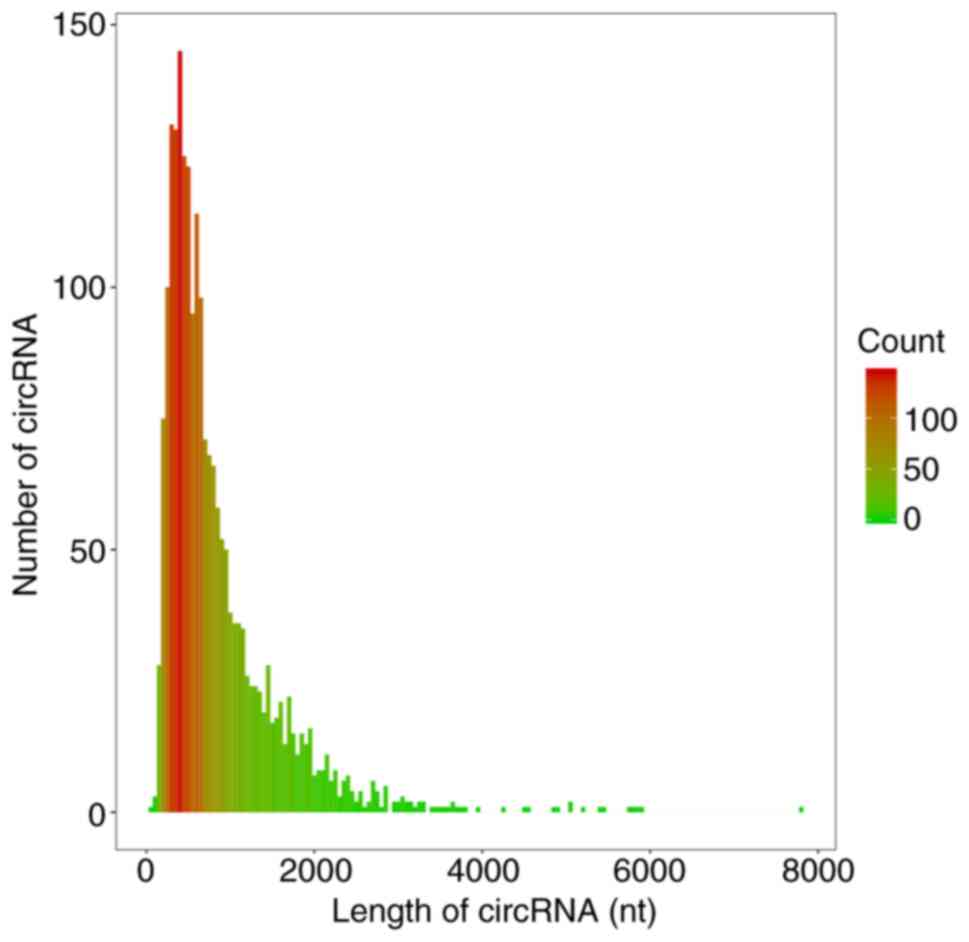
circRNA expression in DNP
The circRNA prediction algorithm identified 2,118
distinct circRNA candidates (≥2 back-spliced reads). The length of
1,552 circRNAs was <1,000 nt and the median length was 620 nt
(Fig. 9), consistent with a
previous study (27). According to
the filtration criteria (fold change ≥1.5 and P≤0.05), 135 DE
circRNAs (64 upregulated and 71 downregulated) between the DNP and
control group were identified. A heat map and volcano plot for the
DE circRNAs are presented in Fig.
10A and B, respectively. The
top 10 upregulated and downregulated circRNAs in the DNP group
compared with those in the control group at 42 days after STZ
injection are listed in Table
V.
 | Table VDetailed information on the top 10
upregulated and 10 downregulated circRNAs. |
Table V
Detailed information on the top 10
upregulated and 10 downregulated circRNAs.
| circRNA ID | Locus | Gene name | Length | Fold change | P-value | Direction of
regulalion |
|---|
| |
chr4:41226279-41229848:- | Ubap2 | 287 | 43.96713505 | 0.00092825 | Up |
|
mmu_circ_0010794 |
chr3:79195934-79215027:- | U6 | 212 | 41.81604666 | 0.001562029 | Up |
|
mmu_circ_0006623 |
chr17:26142617-26143576:+ | Axin1 | 959 | 37.6311112 | 0.001492146 | Up |
|
mmu_circ_0006175 |
chr16:29469240-29479902:- | Atp13a4 | 574 | 35.26138418 | 0.002452781 | Up |
|
mmu_circ_0007095 |
chr17:90362759-90362941:- | Nrxn1 | 182 | 33.46733596 | 0.003380647 | Up |
| |
chr9:9984062-10172122:- | Cntn5 | 1104 | 33.13425687 | 0.003546726 | Up |
|
mmu_circ_0005297 |
chr14:56748834-56764244:- | Pspc1 | 487 | 31.31885528 | 0.005412182 | Up |
|
mmu_circ_0012840 |
chr5:88934748-88954957:+ | Slc4a4 | 254 | 30.36278728 | 0.00845595 | Up |
| |
chr19:37044521-37126388:- | Cpeb3 | 755 | 29.27386362 | 0.006105253 | Up |
|
mmu_circ_0001580 |
chr7:66125241-66125737:+ | Chsy1 | 496 | 14.39983645 | 0.000247404 | Up |
| |
chr16:4655009-4655634:- | Coro7 | 170 | 0.012940375 | 0.000268838 | Down |
| |
chr13:119381675-119404754:- | Nnt | 1016 | 0.022155209 | 0.000693194 | Down |
|
mmu_circ_0016083 |
chrX:113139335-113140786:- | Chm | 198 | 0.02393434 | 0.001058838 | Down |
| |
chr18:23535161-23545766:+ | Dtna | 149 | 0.027664966 | 0.003177893 | Down |
|
mmu_circ_0006471 |
chr16:93799906-93800247:+ | Dopey2 | 257 | 0.028359819 | 0.00367879 | Down |
|
mmu_circ_0008757 |
chr1:5095614-5124469:+ | Atp6v1h | 564 | 0.031315099 | 0.006494722 | Down |
| |
chr6:115244145-115263981:+ | Syn2 | 624 | 0.031522436 | 0.013407635 | Down |
|
mmu_circ_0004843 |
chr13:8697619-8731971:+ | Adarb2 | 672 | 0.034389942 | 0.009121529 | Down |
|
mmu_circ_0013996 |
chr7:141588285-141605010:+ | Ap2a2 | 638 | 0.078133536 | 0.007429339 | Down |
| |
chr1:105640664-105649407:- | Pign | 474 | 0.095120485 | 0.024887204 | Down |
Discussion
In the present study, the DE mRNAs, miRNAs, lncRNAs
and circRNAs in the spinal cord of mice with DNP were
comprehensively analyzed using rRNA-depleted RNA sequencing. A
total of 30 mRNAs, 148 miRNAs, 9 lncRNAs and 135 circRNAs were
determined to be differentially expressed in the DNP mice compared
with the control mice. In addition, the potential functions of the
DE ncRNAs were determined using GO and KEGG pathway analysis. Based
on these results, it was hypothesized that ncRNAs serve a vital
role in the development of DNP and that they may serve as
potentially novel therapeutic targets for the management of
DNP.
The pathogenesis of DNP is complex and remains
poorly understood. The spinal cord is the relay station of
nociceptive stimuli, which serves an important role in the
development of pain (28). However,
the underlying mechanisms by which the spinal dorsal horn processes
nociceptive stimuli are complex. ncRNAs are genetic, epigenetic and
translational regulators. Previous studies have indicated that
dysregulation of ncRNAs is associated with a variety of diseases,
including neuropathic pain (12,13).
However, the role of ncRNAs in the pathogenesis of DNP has remained
largely elusive. Thus, in the present study, the DE ncRNAs in DNP
were determined and their functions and regulatory interactions
were analyzed.
The causal roles of miRNAs in chronic pain have
previously been established (29).
In the present study, numerous DE miRNAs were detected and miR-122
was the most notably upregulated miRNA. It was previously reported
that miR-122 is involved in the regulation of neuropathic pain
(30); however, whether miR-122
regulates DNP remains elusive and further studies are required to
determine this. Although DE miRNAs in the spinal cord of mice with
DNP were screened and the differential expression of certain miRNAs
was confirmed in the present study, the underlying mechanisms of
miRNAs in DNP are poorly understood. GO analysis may be used to
unify the representation of genes and gene product attributes in
all species (31). GO terms and GO
annotations are good predictors of gene functions and trends
(32). The KEGG pathway database is
the most widely used enrichment analysis platform and it stores
higher-order functional information for systematic analysis of gene
functions (33). In the present
study, the miRNA-related gene functions and the corresponding
pathways in mice with DNP were predicted using GO term and KEGG
pathway enrichment analyses. The results indicated that the most
significantly involved pathways in the pathogenesis of DNP were the
MAPK signaling pathway, Rap1 signaling pathway and TGF-β signaling
pathway. Previous studies have demonstrated that these signaling
pathways are closely associated with neuropathic pain (34-36).
The function of miRNAs in the pathogenesis of DNP should be
examined in more detail in future studies.
A growing number of studies have also indicated that
noxious stimuli may result in dysregulated expression of lncRNAs
and this may be involved in the pain hypersensitivity underlying NP
(37,38). To the best of our knowledge, there
are no comprehensive studies of the lncRNAs associated with DNP.
Thus, second-generation sequencing was used to analyze the DE
lncRNAs in the spinal cord of mice with DNP. The results indicated
that a total of 9 lncRNAs were significantly dysregulated in mice
with DNP compared to control mice.
CircRNAs are a type of highly stable, circularized
lncRNA. A previous study indicated that circRNAs are conserved
across species and are primarily enriched in the nervous system
(39). Various circRNAs have been
identified; however, the biological functions of the majority of
these circRNAs remain elusive. In the present study, 135 DE
circRNAs that may be involved in the pathogenesis of DNP were
identified, which may provide further insight into the underlying
mechanisms of circRNAs in DNP. The majority of circRNAs detected
were derived from exons, similar to the results of a previous study
where most circRNAs were derived from coding sequences (39). In the present study, it was also
indicated that multiple circRNAs were able to be generated from one
host gene. Regulating synaptic membrane exocytosis 2 is able to
generate 17 distinct circRNAs and the gene encoding protein
tyrosine kinase 2 is able to generate 47 distinct circRNAs. The
median length of circRNAs was 620 nt, similar to a previous study
in which the median length of circRNAs was around 500 nt (40,41).
The present study had certain limitations.
Sequencing analysis was used to investigate the expression patterns
of coding genes, miRNAs, lncRNAs and circRNAs in the spinal cord of
mice with STZ-induced DNP. In order to verify the effectiveness of
the preliminary screening approach, the effects of these DE ncRNAs
in DNP will be further assessed in animals and in humans to verify
the related functions and pathways of these ncRNAs. In addition,
the differential expression of mRNAs, miRNAs, lncRNAs and circRNAs
in the spinal cord of DNP mice was assessed in the present study.
However, whether these results translate to humans remains unknown.
Thus, whether these ncRNAs are of relevance to DNP in humans will
next be determined. According to the theory of biological
evolution, RNA is conserved to a certain extent. Conservation
analysis will be performed on RNAs to identify the human ncRNAs
similar to those identified in the mice for further mechanistic
research. In addition, the complications and other physiological
indicators following STZ injection were not addressed in the
present study, and thus, future studies should take this limitation
into account. According to a previous study (14), mice exhibit notable chronic DNP 6
weeks after STZ injection; however, whether different time-points
affect the results is unknown but worthy of further study.
In conclusion, the differential expression profiles
of mRNAs, miRNAs, lncRNAs and circRNAs in the spinal cord of DNP
mice were determined using rRNA-depleted RNA sequencing. A total of
30 mRNAs, 148 miRNAs, 9 lncRNAs and 135 circRNAs were
differentially expressed between the DNP and control mice. ‘Rap1
signaling pathway’ and ‘MAPK signaling pathway’ were the most
enriched pathways among the DE genes. The complete proteomic and
relevant signaling pathway of this differential expression ncRNAs
is worthy of further study, which may ultimately enable full
disclosure of the mechanisms underlying DNP.
Acknowledgements
Not applicable.
Funding
Funding: This study was funded by The National Natural Science
Funds of China (grant no. 81771357).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request. The sequencing data have been submitted to https://bigd.big.ac.cn/databases with the
accession ID CRA003943.
Authors' contributions
ZSQ, JH and HBW designed the present study. JH, JJH,
LZ and DLL performed the experiments. JH, WYH and QMX analyzed data
and wrote the manuscript draft. JH and ZSQ revised the final
manuscript. HBW and ZSQ confirm the authenticity of all the raw
data. All authors read and approved the final manuscript.
Ethics approval and consent to
participate
All experiments were approved by the Animal Use and
Care Committee for Research and Education of The First People's
Hospital of Foshan (Foshan, China) and were performed in accordance
with the guidelines described in the International Association for
the Study of Pain, as well as in compliance with the original
ARRIVE guidelines.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Kesavadev J, Saboo B, Sadikot S, Das AK,
Joshi S, Chawla R, Thacker H, Shankar A, Ramachandran L and Kalra
S: Unproven therapies for diabetes and their implications. Adv
Ther. 34:60–77. 2017.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Sun J, Wang Y, Zhang X, Zhu S and He H:
Prevalence of peripheral neuropathy in patients with diabetes: A
systematic review and meta-analysis. Prim Care Diabetes.
14:435–444. 2020.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Shillo P, Sloan G, Greig M, Hunt L,
Selvarajah D, Elliott J, Gandhi R, Wilkinson ID and Tesfaye S:
Painful and painless diabetic neuropathies: What is the difference?
Curr Diab Rep. 19(32)2019.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Paisley P and Serpell M: Improving pain
control in diabetic neuropathy. Practitioner. 261:23–26.
2017.PubMed/NCBI
|
|
5
|
Dewanjee S, Das S, Das AK, Bhattacharjee
N, Dihingia A, Dua TK, Kalita J and Manna P: Molecular mechanism of
diabetic neuropathy and its pharmacotherapeutic targets. Eur J
Pharmacol. 833:472–523. 2018.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Sloan G, Shillo P, Selvarajah D, Wu J,
Wilkinson ID, Tracey I, Anand P and Tesfaye S: A new look at
painful diabetic neuropathy. Diabetes Res Clin Pract. 144:177–191.
2018.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Yang XD, Fang PF, Xiang DX and Yang YY:
Topical treatments for diabetic neuropathic pain. Exp Ther Med.
17:1963–1976. 2019.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Mattick JS and Makunin IV: Non-coding RNA.
Hum Mol Genet. 15:R17–R29. 2006.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Schmitt AM and Chang HY: Long noncoding
RNAs in cancer pathways. Cancer Cell. 29:452–463. 2016.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Zaratiegui M, Irvine DV and Martienssen
RA: Noncoding RNAs and gene silencing. Cell. 128:763–776.
2007.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Thum T: Noncoding RNAs and myocardial
fibrosis. Nat Rev Cardiol. 11:655–663. 2014.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Li G, Jiang H, Zheng C, Zhu G, Xu Y, Sheng
X, Wu B, Guo J, Zhu S, Zhan Y, et al: Long noncoding RNA MRAK009713
is a novel regulator of neuropathic pain in rats. Pain.
158:2042–2052. 2017.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Liu C, Li C, Deng Z, Du E and Xu C: Long
Non-coding RNA BC168687 is involved in TRPV1-mediated diabetic
neuropathic pain in rats. Neuroscience. 374:214–222.
2018.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Yang D, Yang Q, Wei X, Liu Y, Ma D, Li J,
Wan Y and Luo Y: The role of miR-190a-5p contributes to diabetic
neuropathic pain via targeting SLC17A6. J Pain Res. 10:2395–2403.
2017.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Wu B, Guo Y, Chen Q, Xiong Q and Min S:
MicroRNA-193a Downregulates HMGB1 to alleviate diabetic neuropathic
pain in a mouse model. Neuroimmunomodulat. 26:250–257.
2019.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Yu W, Zhao GQ, Cao RJ, Zhu ZH and Li K:
LncRNA NONRATT021972 was associated with neuropathic pain scoring
in patients with Type 2 diabetes. Behav Neurol.
2017(2941297)2017.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Wilusz JE and Sharp PA: Molecular biology.
A circuitous route to noncoding RNA. Science. 340:440–441.
2013.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Hansen TB, Jensen TI, Clausen BH, Bramsen
JB, Finsen B, Damgaard CK and Kjems J: Natural RNA circles function
as efficient microRNA sponges. Nature. 495:384–388. 2013.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Memczak S, Jens M, Elefsinioti A, Torti F,
Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer
M, et al: Circular RNAs are a large class of animal RNAs with
regulatory potency. Nature. 495:333–338. 2013.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Wang L, Luo T, Bao Z, Li Y and Bu W:
Intrathecal circHIPK3 shRNA alleviates neuropathic pain in diabetic
rats. Biochem Biophys Res Commun. 505:644–650. 2018.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Zimmermann M: Ethical guidelines for
investigations of experimental pain in conscious animals. Pain.
16:109–110. 1983.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Stoffel EC, Ulibarri CM, Folk JE, Rice KC
and Craft RM: Gonadal hormone modulation of mu, kappa, and delta
opioid antinociception in male and female rats. J Pain. 6:261–274.
2005.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Chaplan SR, Bach FW, Pogrel JW, Chung JM
and Yaksh TL: Quantitative assessment of tactile allodynia in the
rat paw. J Neurosci Methods. 53:55–63. 1994.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Furman BL: Streptozotocin-Induced diabetic
models in mice and rats. Curr Protoc Pharmacol. 70:5.47.1–5.47.20.
2015.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Jeck WR and Sharpless NE: Detecting and
characterizing circular RNAs. Nat Biotechnol. 32:453–461.
2014.PubMed/NCBI View
Article : Google Scholar
|
|
26
|
You X, Vlatkovic I, Babic A, Will T,
Epstein I, Tushev G, Akbalik G, Wang M, Glock C, Quedenau C, et al:
Neural circular RNAs are derived from synaptic genes and regulated
by development and plasticity. Nat Neurosci. 18:603–610.
2015.PubMed/NCBI View
Article : Google Scholar
|
|
27
|
Ding X, Zhang S, Li X, Feng C, Huang Q,
Wang S, Wang S, Xia W, Yang F, Yin R, et al: Profiling expression
of coding genes, long noncoding RNA, and circular RNA in lung
adenocarcinoma by ribosomal RNA-depleted RNA sequencing. FEBS Open
Bio. 8:544–555. 2018.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Prescott SA, Ma Q and De Koninck Y: Normal
and abnormal coding of somatosensory stimuli causing pain. Nat
Neurosci. 17:183–191. 2014.PubMed/NCBI View
Article : Google Scholar
|
|
29
|
López-González MJ, Landry M and Favereaux
A: MicroRNA and chronic pain: From mechanisms to therapeutic
potential. Pharmacol Ther. 180:1–15. 2017.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Ma F, Wang C, Yoder WE, Westlund KN,
Carlson CR, Miller CS and Danaher RJ: Efficacy of herpes simplex
virus vector encoding the human preproenkephalin gene for treatment
of facial pain in mice. J Oral Facial Pain Headache. 30:42–50.
2016.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Altshuler D, Daly MJ and Lander ES:
Genetic mapping in human disease. Science. 322:881–888.
2008.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Li L, Zhang K, Lee J, Cordes S, Davis DP
and Tang Z: Discovering cancer genes by integrating network and
functional properties. BMC Med Genomics. 2(61)2009.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Du J, Li M, Yuan Z, Guo M, Song J, Xie X
and Chen Y: A decision analysis model for KEGG pathway analysis.
BMC Bioinformatics. 17(407)2016.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Ni HD, Yao M, Huang B, Xu LS, Zheng Y, Chu
YX, Wang HQ, Liu MJ, Xu SJ and Li HB: Glial activation in the
periaqueductal gray promotes descending facilitation of neuropathic
pain through the p38 MAPK signaling pathway. J Neurosci Res.
94:50–61. 2016.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Singhmar P, Huo X, Eijkelkamp N, Berciano
SR, Baameur F, Mei FC, Zhu Y, Cheng X, Hawke D, Mayor F Jr, et al:
Critical role for Epac1 in inflammatory pain controlled by
GRK2-mediated phosphorylation of Epac1. Proc Natl Acad Sci USA.
113:3036–3041. 2016.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Chen NF, Huang SY, Chen WF, Chen CH, Lu
CH, Chen CL, Yang SN, Wang HM and Wen ZH: TGF-β1 attenuates spinal
neuroinflammation and the excitatory amino acid system in rats with
neuropathic pain. J Pain. 14:1671–1685. 2013.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Wu J, Wang C and Ding H: LncRNA MALAT1
promotes neuropathic pain progression through the miR-154-5p/AQP9
axis in CCI rat models. Mol Med Rep. 21:291–303. 2020.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Ren X, Yang R, Li L, Xu X and Liang S:
Long non coding RNAs involved in MAPK pathway mechanism mediates
diabetic neuropathic pain. Cell Biol Int. 44:2372–2379.
2020.PubMed/NCBI View Article : Google Scholar
|
|
39
|
van Rossum D, Verheijen BM and Pasterkamp
RJ: Circular RNAs: Novel regulators of neuronal development. Front
Mol Neurosci. 9(74)2016.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Zheng Q, Bao C, Guo W, Li S, Chen J, Chen
B, Luo Y, Lyu D, Li Y, Shi G, et al: Circular RNA profiling reveals
an abundant circHIPK3 that regulates cell growth by sponging
multiple miRNAs. Nat Commun. 7(11215)2016.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Guo JU, Agarwal V, Guo H and Bartel DP:
Expanded identification and characterization of mammalian circular
RNAs. Genome Biol. 15(409)2014.PubMed/NCBI View Article : Google Scholar
|