Introduction
Osteosarcoma (OS) is one of the most common primary
malignancies and it mainly arises in the metaphyses of long bones
in adolescents and young adults (1). OS is associated with abnormal cell
differentiation caused by genetic and epigenetic alterations
(2). Although treatment methods,
including chemo- and radiotherapy and surgery, have been developed
and are in use, the prognosis of patients with recurrent and
metastatic OS remains poor with a five-year survival rate of 50-60%
(3). It has been indicated that
multiple oncogenes and tumor suppressors are involved in the
occurrence and development of OS (4). However, the detailed molecular
mechanisms of OS and processes associated with its progression,
including migration and invasion, have remained to be fully
elucidated.
Circular RNAs (circRNAs) are a group of non-coding
RNAs, which are widely expressed in numerous species. They are
characterized as having covalent and closed-loop structures
(5). CircRNAs are considered key
epigenetic regulators in the pathogenesis of various diseases and
they may have essential roles in tumor genesis and development
(6). For instance, circ-0000658 was
downregulated in OS tissues and cells, which inhibited the cell
cycle, proliferation, invasion and migration of cells through the
microRNA (miRNA/miR)-1227/IRF2 axis (7). Silencing of circ_0032462 inhibited OS
cell proliferation, migration and invasion by upregulating kinesin
family member 3B (8). Circ_0008932
was significantly upregulated in esophageal squamous cell carcinoma
(ESCC) and knockdown of circ_0008932 reduced the viability, colony
formation ability and invasion of ESCC cells through targeting the
miR-182/Myc axis (9). Circ_0008932
suppressed vascular smooth muscle cell (VSMC) migration,
proliferation and differentiation by binding to miR-145 (10,11).
However, the roles of circ_0008932 in OS have not been previously
studied.
In the present study, it was revealed that
circ_0008932 was upregulated in osteosarcoma tissues compared to
corresponding noncancerous samples and circ_0008932 upregulation
was also associated with metastasis in patients with OS. The
survival analysis demonstrated that high circ_0008932 expression in
OS tissues was associated with an unfavorable overall survival
prognosis. Suppression of circ_0008932 significantly inhibited the
proliferation, invasion and migration of OS cells, while cell
apoptosis was promoted. Furthermore, the present data indicated
that circ_0008932 directly downregulated miR-145-5p expression,
which was negatively correlated with circ_0008932 expression in OS
tissues.
Materials and methods
OS tissues and cell lines
A total of 50 OS tissues and matched non-tumor
tissues were collected from all patients (27 males and 23 females;
mean age, 6.36±2.18 years) during surgery at the Department of
Radiation Oncology, Chongqing University Cancer Hospital (Chonqing,
China), where metastasis was detected in 29 cases. Metastatic
tissues were not analyzed in this study. Written informed consent
was provided by all patients. The protocol was approved by the
Ethics Committee of Chongqing University Cancer Hospital
(Chongqing, China). Upon collection, tissues were immediately
frozen using liquid nitrogen and stored at -80˚C prior to RNA
isolation. Cell lines, including NHOst, HOS, SOSP-9607, SaOS-2 and
U2OS, were purchased from the Cell Bank of the Chinese Academy of
Sciences and cultured using Dulbecco's modified Eagle's medium
(DMEM; Invitrogen; Thermo Fisher Scientific, Inc.) supplemented
with 10% fetal bovine serum (FBS; HyClone; Cytiva).
Cell transfection
Small interfering (si)RNA against circ_0008932
(si-circ_0008932; 5'-AACCACUGCCUGCCUCAAUCU-3') and a scrambled
control siRNA used as a negative control (si-NC;
5'-UUACGUUACAAAUCCGAUGUA-3') were purchased from Shanghai
GenePharma Co., Ltd. miR-145-5p mimics
(5'-GUCCAGUUUUCCCAGGAAUCCCU-3') and the respective control (miR-NC;
5'-AAUUGAAGUUCCCAGGAAUCCCU-3'), as well as miR-145-5p inhibitor
(5'-AGGGAUUCCUGGGAAAACUGGAC-3') and its control (in-NC;
5'-CAGUACUUUUGUGUAGUACAAA-3') were synthesized by Shanghai
GenePharma Co., Ltd. siRNAs, miR mimics, miR inhibitors and their
NC oligonucleotides (50 nM) were transfected using
Lipofectamine® 2000 reagent (Invitrogen; Thermo Fisher
Scientific, Inc.) according to the manufacturer's protocol.
RNA extraction and reverse
transcription-quantitative (RT-q)PCR
Total RNA was extracted from cells and tissues using
the TRIzol® reagent (Invitrogen; Thermo Fisher
Scientific, Inc.) according to the manufacturer's protocol.
PrimeScript™ RT kit (cat. no. RR036B; Takara Biotechnology Co.,
Ltd.) was used for reverse transcription, the sample was incubated
at room temperature for 30 min. After that, 1 cycle of PCR was
performed at 42˚C for 45 min, 99˚C for 5 min and 5˚C for 5 min in a
PCR cycler. qPCR analyses of circ_0008932 and miR-145-5p were
performed using TB Green® Premix Ex Taq™ (cat. no.
RR420A; Takara Bio, Inc.) in an ABI Prism 7700 system (Applied
Biosystems; Thermo Fisher Scientific, Inc.). The thermocycling
conditions contained initial denaturation at 95˚C for 10 min,
followed by 40 cycles of 95˚C for 10 sec, 60˚C for 30 sec and 72˚C
for 10 sec and final extension at 72˚C for another 5 min. Primers
for U6 and miR-145-5p were purchased from GeneCopoeia, Inc. The
primer sequences were as follows: circ_0008932 forward,
5'-TTTGGTCCCCTTCAACCAGCTG-3' and reverse,
5'-TAAACCAAGGTAAAATGGTCGA-3'; miR-145-5p forward,
5'-GTCCAGTTTTCCCAGGAATCC-3' and reverse,
5'-TCGGCAGGGTCCAGTTTTCCCA-3'; GAPDH forward,
5'-CATGGCCTTCCGTGTTCCTA-3' and reverse, 5'-TACTTGGCAGGTTTCTCCAGG-3'
and U6 forward, 5'-GCTTCGGCAGCACATATACTAAAAT-3' and reverse,
5'-CGCTTCACGAATTTGCGTGTCAT-3'. The expression of circ_0008932 was
normalized to that of GAPDH and the levels of miR-145-5p were
normalized to those of U6. The expression was calculated using the
2-ΔΔCq method (12).
Patients were divided into circ_0008932 high- or low-expression
groups according to the mean circ_0008932 expression value. All
experiments were performed in triplicate.
Cell proliferation assay
SaOS-2 and U2OS cells were seeded in 96-well plates
at a density of 1x103 cells per well. The cells were
transfected with si-NC or si-circ_0008932 or co-transfected with
miR-145-5p mimics/inhibitors and si-circ_0008932. Cell viability
was evaluated for 4 consecutive days by using a Cell Counting Kit
(CCK)-8 assay (Dojindo Molecular Technologies, Inc.). A total of 10
µl CCK-8 reagent was added to each well and the cells were
incubated at 37˚C for another 2 h. The absorbance at 450 nm was
measured using a microplate reader (Bio-Rad Laboratories). All
experiments were independent and performed in triplicate.
Cell migration and invasion
assays
For the cell invasion and migration assays,
1.0x105 SaOS-2 or U2OS cells were collected after
transfection and used for each experiment. Cell migration and
invasion were determined using Transwell assays. In brief, the
suspension of cells containing no FBS was seeded into the upper
chamber of the Transwell insert with a pore size of 8 µm (BD
Biosciences). In addition, for the cell invasion assay, cells were
seeded onto the upper chambers pre-coated with Matrigel®
(Sigma-Aldrich; Merck KGaA). DMEM with 10% FBS was added into lower
chamber. Cells were incubated at 37˚C for 12 h and those that
migrated or invaded through the membrane were fixed with 4%
polyoxymethylene at room temperature for 10 min, stained with 0.2%
crystal violet at room temperature for 10 min and then counted
under an IX83 inverted fluorescent microscope (magnification, x200;
Olympus Corporation).
Luciferase assay
Bioinformatics analysis was performed using
Targetscan (release 5.2; www.targetscan.org) and miRNA.org (version
10.0; www.microrna.org/microrna). To investigate the
interaction between circ_0008932 and miR-145-5p, wild-type fragment
of the 3'-untranslated region (UTR) on circ_0008932 with
complementary binding sites of miR-145-5p were obtained from
Shanghai GenePharma Co., Ltd. A circ_0008932 3'UTR-mutant (MUT)
vector containing the mutant binding site of miR-145-5p was
produced using a QuikChange Multi Site-Directed Mutagenesis kit
(Stratagene; Agilent Technologies, Inc.) using the restriction
digestion sites of SpeI and HindIII (Promega
Corporation). The pmirGLO Dual-Luciferase plasmids (Promega
Corporation) were used to construct the luciferase reporter
vectors. Mutant circ_0008932 was used as a control. For
circ_0008932 (https://circinteractome.nia.nih.gov/api/v2/circsearch?circular_rna_query=hsa_circ_0008932&gene_symbol_query=&submit=circRNA+Search),
Homo sapiens full open reading frame cDNA of circ_0008932
was selected. The products were amplified using primers with
flanking SpeI and HindIII restriction digestion sites
and the amplified DNA was then inserted into the pmirGLO
Dual-Luciferase reporter vector (Promega Corporation). The 293T
cells (The Cell Bank of Type Culture Collection of the Chinese
Academy of Sciences) were cultured with serum-free DMEM at 37˚C
with 5% of CO2. Cells were transfected with 25 nM
wild-type or mutant pMIR-circ_0008932, as well as Renilla
luciferase control vector (pRL-TK; Promega Corporation) using
Lipofectamine® 2000. Cells were co-transfected with 25
nM miR-NC or miR-145-5p mimics/inhibitors, respectively. Luciferase
activity was examined 48 h after transfection using the
Dual-Luciferase reporter assay system (Promega Corporation) using
an LB 960 Centro XS3 luminometer (Molecular Devices, LLC).
Flow cytometry
Flow cytometry was used to determine cell apoptosis.
Transfected cells (1x105 per well) were assayed using
the Annexin V-FITC/propidium iodide (PI) apoptosis detection kit
(Beijing Solarbio Science & Technology Co., Ltd.) according to
the manufacturer's protocol. Cells were stained with annexin V-FITC
and propidium iodide (PI). After a 10 min incubation at room
temperature, the cells were analyzed using a BD FACSAria III
cytometer (BD Biosciences) and interpreted by ModFit software
(version 2.0; BD Biosciences).
Statistical analysis
Values are expressed as the mean ± standard
deviation and analyzed using SPSS 19.0 (IBM Corporation).
Statistical significance was examined using Student's t-test or
one-way ANOVA. Tukey's test was conducted as a post-hoc test
following ANOVA. Patient survival was assessed using the
Kaplan-Meier method and the log-rank test was applied to determine
significant differences between groups. Pearson's correlation
analysis was used to evaluate the relationship between the
expression levels of circ_0008932 and miR-145-5p. P<0.05 was
considered to indicate statistical significance.
Results
Circ_0008932 is upregulated in OS
tissues and cells and high levels are associated with metastasis
and prognosis
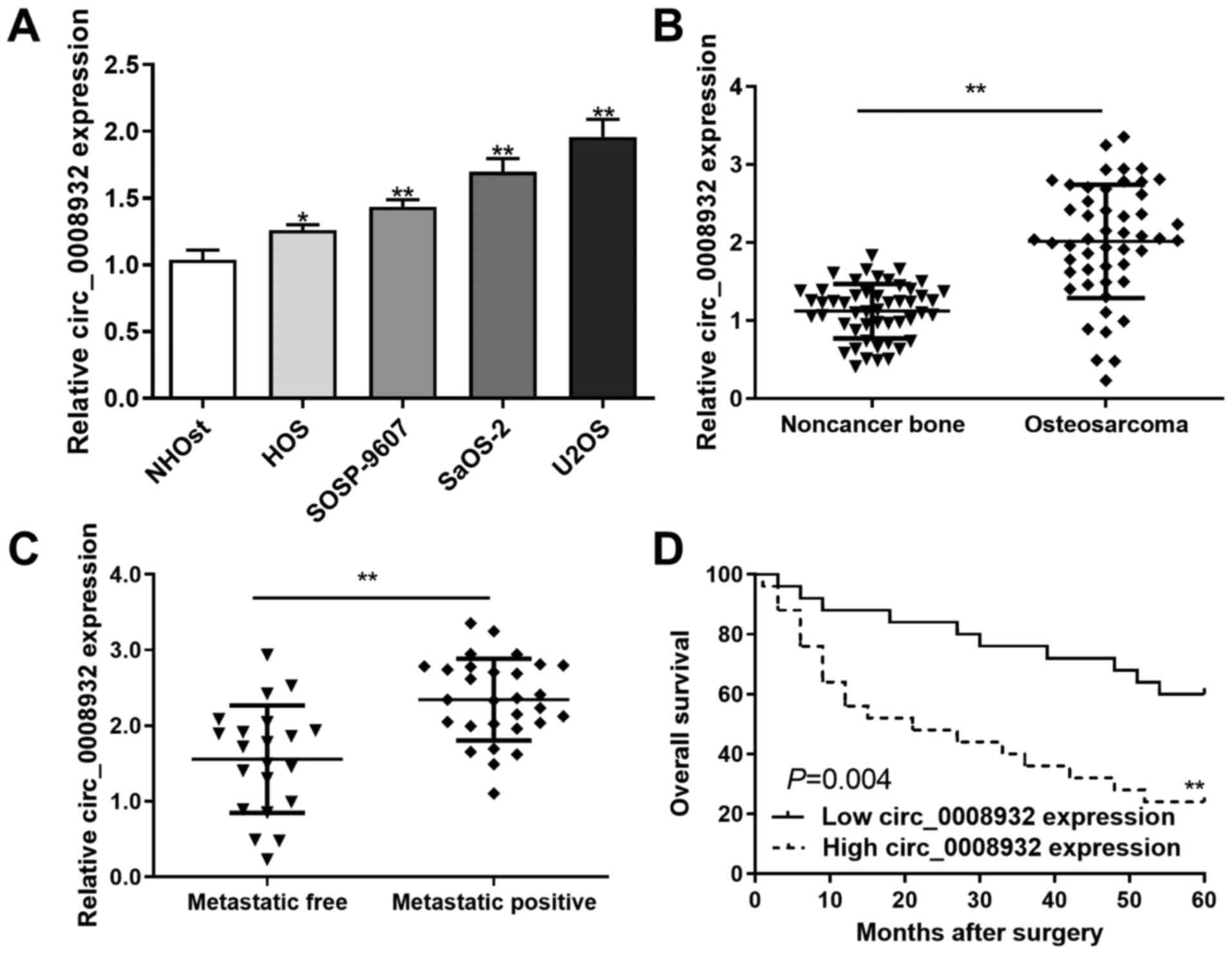
RT-qPCR was performed to detect the levels of
circ_0008932 in OS tissues and cells. The expression of
circ_0008932 was assessed in five cell lines. The results indicated
that the expression of circ_0008932 was increased in the four OS
cell lines (HOS, SOSP-9607, SaOS-2 and U2OS) compared with that in
NHOst normal human osteoblast cells (Fig. 1A). Furthermore, the expression of
circ_0008932 in OS and paired non-tumor tissues was also evaluated.
The results indicated that circ_0008932 expression was
significantly increased in OS samples (Fig. 1B). In addition, circ_0008932
expression was increased in primary tumors of patients with
metastasis compared with those without metastasis (Fig. 1C). In addition, the overall survival
of patients with OS with high circ_0008932 expression was poor
compared with that of patients with low circ_0008932 expression
(P=0.004; Fig. 1D).
Silencing of circ_0008932 inhibits the
proliferation, invasion and migration of OS cells, whereas cell
apoptosis is enhanced
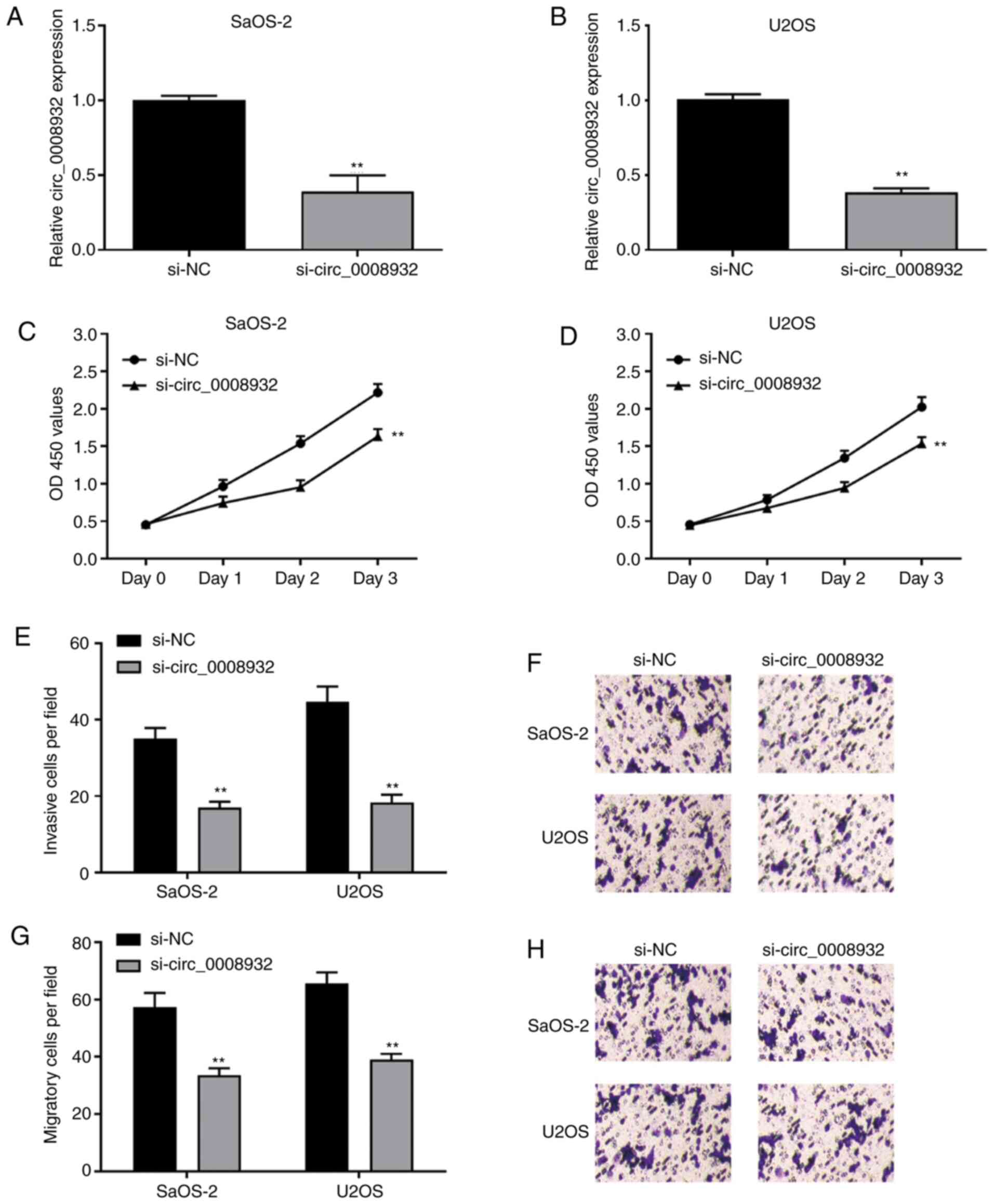
To determine whether the inhibition of circ_0008932
is able to suppress the proliferation, migration and invasion of OS
cells, SaOS-2 and U2OS cells were transfected with si-NC or
si-circ_0008932 for 24 h and subsequently, cell proliferation,
invasion and migration were examined. It was confirmed that the
expression of circ_0008932 was significantly decreased in SaOS-2
and U2OS cells transfected with si-circ_0008932 (Fig. 2A and B). The results revealed that knockdown of
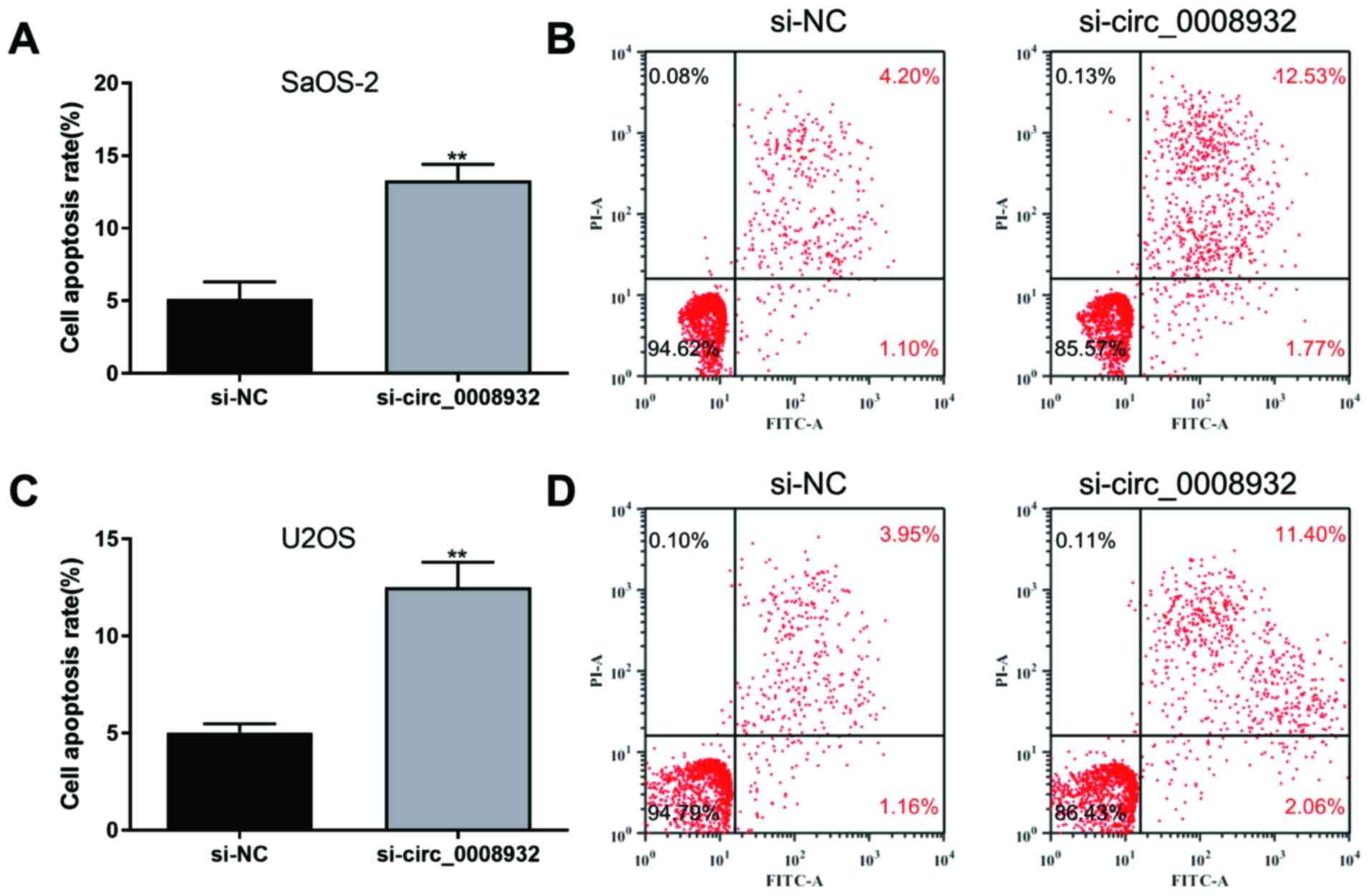
circ_0008932 markedly suppressed cell proliferation (Fig. 2C and D), invasion (Fig. 2E and F) and migration (Fig. 2G and H), while cell apoptosis was promoted
(Fig. 3). These data suggested that
knockdown of circ_0008932 suppressed the proliferation/motility and
promoted apoptosis of OS cells.
miR-145-5p is a direct target of
circ_0008932
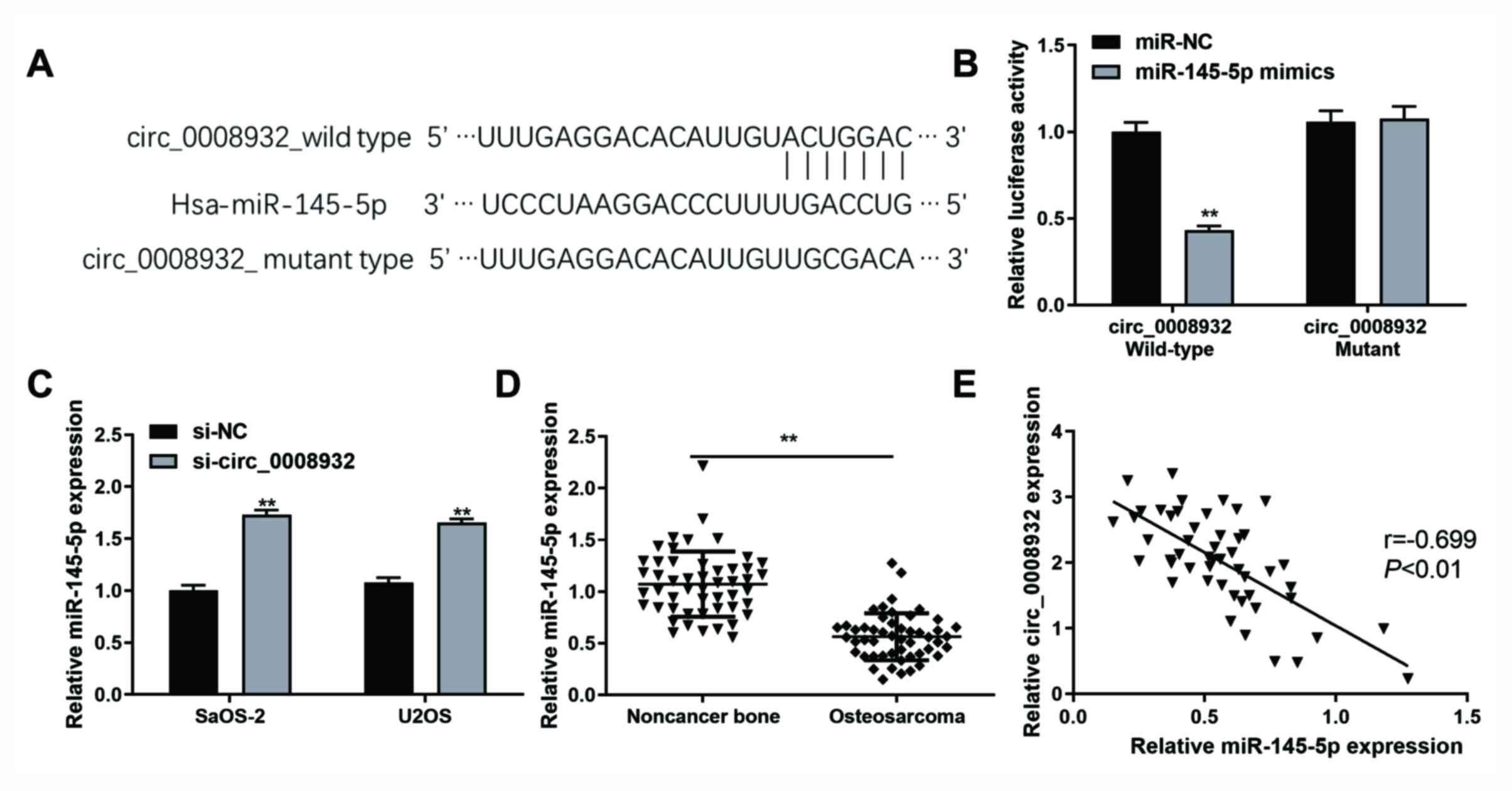
Bioinformatics analysis using Targetscan and
miRNA.org indicated that circ_0008932 contains
potential binding sites for miR-145-5p. To confirm whether
miR-145-5p is a direct target of circ_0008932, luciferase vectors
containing the wild-type or mutant circ_0008932 sequence were
constructed (Fig. 4A). For the
luciferase assays, wild-type or mutant plasmids were co-transfected
into 293T cells with miR-145-5p mimics or inhibitors. As presented
in Fig. 4B, miR-145-5p mimics
significantly reduced the luciferase activity of wild-type
circ_0008932, while that of the mutant circ_0008932 was not
affected. The results also suggested that knockdown of circ_0008932
significantly increased the expression of miR-145-5p (Fig. 4C). To evaluate the interaction
between circ_0008932 and miR-145-5p, their expression levels were
determined in OS and paired adjacent normal tissues. The results
indicated that the expression of miR-145-5p was significantly
decreased in OS tissues (Fig. 4D).
Furthermore, the expression of miR-145-5p was inversely correlated
with the levels of circ_0008932 (Fig.
4E). Taken together, circ_0008932 was indicated to downregulate
miR-145-5p through direct binding, consequently affecting tumor
progression in OS.
Si-circ_0008932 inhibits OS cell
growth and motility by targeting miR-145-5p
To investigate whether circ_0008932 regulates the
proliferation, invasion and migration of OS cells via targeting
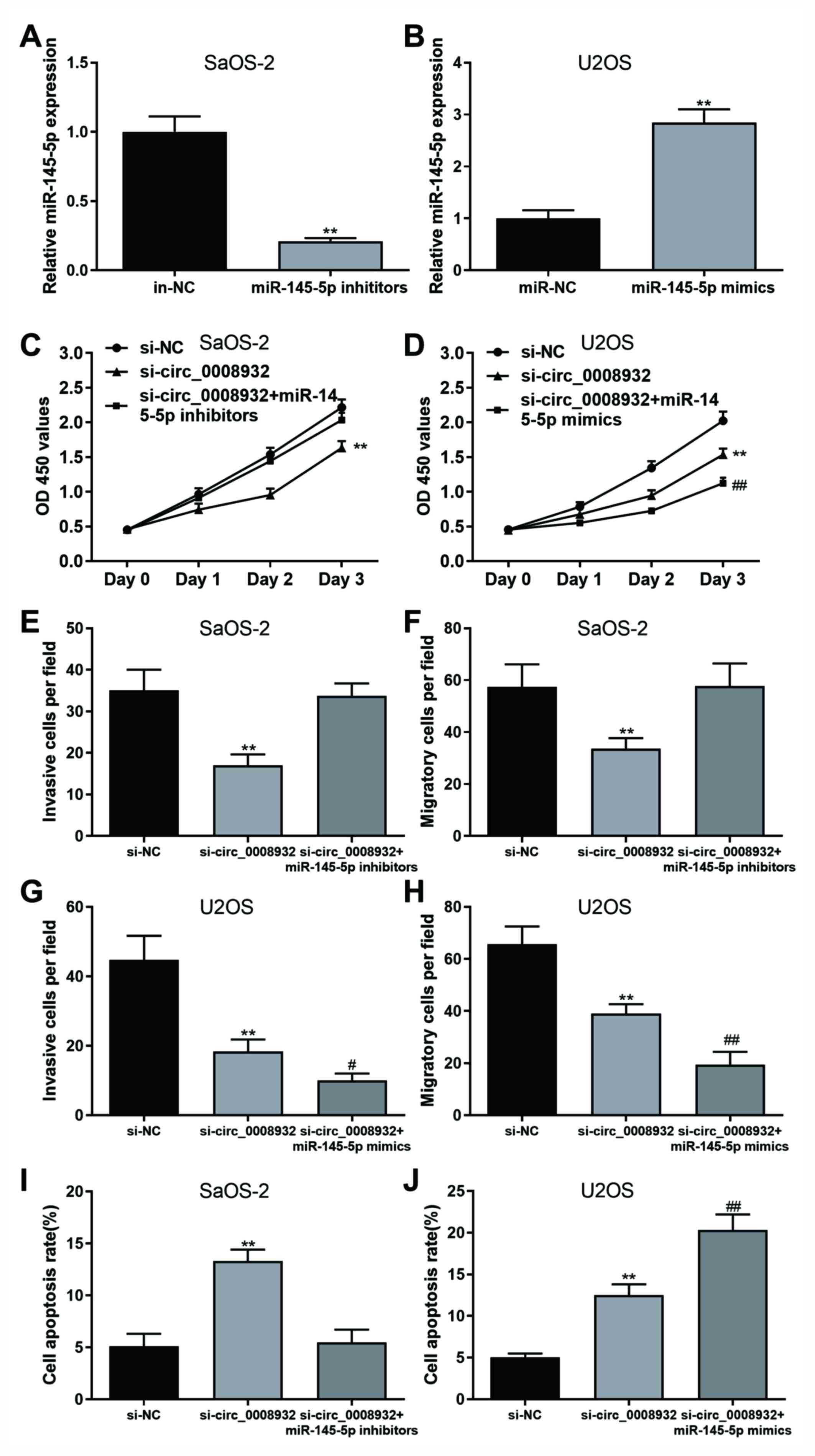
miR-145-5p, further experiments were performed (Fig. 5). OS cells were transfected with
si-NC and si-circ_0008932, or co-transfected with si-circ_0008932,
miR-145-5p mimics or inhibitors. The transfection efficiencies of
miR-145-5p inhibitors and mimics were confirmed compared with their
respective negative controls in both of the OS cell lines (Fig. 5A and B). The results revealed that the
proliferation, invasion and migration of OS cells were
significantly inhibited, while cell apoptosis was promoted after
transfection with si-circ_0008932, and these effects were abolished
by miR-145-5p inhibitors (Figs. 5C,
E, F, I,
S1A and S2A). Furthermore, the changes in the
biological behaviour of OS cells caused by si-circ_0008932 were
enhanced by co-transfection with miR-145-5p mimics (Figs. 5D, G, H,
J, S1B and S2B). In summary, silencing of
circ_0008932 suppressed the proliferation/motility and promoted
apoptosis of OS cells by targeting miR-145-5p.
Discussion
CircRNA is a type of non-coding RNA widely
distributed in eukaryotes. CircRNA is highly stable as it is
resistant to exonuclease- or ribonuclease-mediated degradation
(13). It has been reported that
circRNA is a key regulator of tumor gene expression and serves key
roles in the signaling networks in oral squamous cell carcinoma,
including the Wnt/β-catenin and PI3K/AKT pathways (14). Furthermore, upregulation of
circ_0008932 was detected in ESCC and knockdown of circ_0008932
suppressed the proliferation, migration and invasion of ESCC cells
via the miR-182/Myc axis (9). In
addition, circ_0008932 inhibited the migration, proliferation and
differentiation of VSMCs through targeting miR-145(10). However, the detailed functions of
circ_0008932 in OS have remained to be fully elucidated.
The results of the present study revealed that
circ_0008932 was increased in tumor tissues of patients with OS. In
addition, upregulation of circ_0008932 was associated with
metastasis in patients with OS. Furthermore, the survival analysis
demonstrated that patients with OS with high circ_0008932
expression exhibited poor overall survival. In addition,
transfection with si-circ_0008932 inhibited the proliferation,
invasion and migration of OS cells, while cell apoptosis was
promoted, suggesting that circ_0008932 affected the biological
behavior of OS cells. Furthermore, the expression of circ_0008932
was significantly increased in OS cell lines, including HOS, MG-63,
SaOS-2 and U2OS.
Previous studies indicated that circRNA acts as an
miRNA sponge through competitive binding, consequently leading to
loss of function of target miRNA and affect gene expression at the
post-transcriptional level (15-19).
In the present study, the results of the luciferase reporter assay
revealed the interaction between circ_0008932 and miR-145-5p.
Furthermore, a negative correlation between the expression of
circ_0008932 and miR-145-5p in OS tissues was observed, suggesting
that the circ_0008932 and miR-145-5p axis may be involved in the
pathogenesis of OS.
Recent studies have revealed the dysregulation of
miRNAs in numerous types of cancer, including OS, which is closely
associated with tumor cell proliferation, differentiation,
apoptosis, autophagy, metastasis and invasion (20,21).
Emerging evidence has revealed the involvement of miRNA in the
growth and metastasis of OS cells, such as miR-32, miR-142-3p,
miR-194, miR-202, miR-217 and let-7a. Furthermore, miR-145-5p was
downregulated in OS, which was associated with the metastasis and
prognosis of patients with OS (22,23).
The regulatory roles of miR-145-5p in the occurrence and
development of OS have also been confirmed (24). In addition, Feng et al
(25) revealed that miR-145-5p
suppressed colorectal cancer cell proliferation, invasion and
migration by targeting Fascin-1. Furthermore, Tang et al
(26) suggested that the
downregulation of miR-145-5p is associated with tumor
aggressiveness/metastasis of OS and may be an independent
prognostic marker for patients with OS.
In consistency with these results, the present data
revealed the downregulation of miR-145-5p in OS specimens, where
the expression of miR-145-5p and circ_0008932 were inversely
correlated. Furthermore, the present study also indicated that
miR-145-5p inhibitors abolished the inhibitory effects caused by
si-circ_0008932 on OS progression, whereas miR-145-5p mimics
enhanced the changes in biological behavior induced by
si-circ_0008932 in OS cells. These results suggested that
miR-145-5p is involved in circ_0008932-modulated OS cell
proliferation, invasion, migration and apoptosis. However, there
were certain limitations to this study. For instance, small RNAs in
the samples could have been enriched using a commercial kit prior
to the experiments and both SaOS-2 and U2OS cells should be
transfected with miR-145-5p mimics and miR-145 inhibitors.
In conclusion, the present results indicated that
circ_0008932 may be an oncogenic factor in OS and the inhibitory
effects of si-circ_0008932 on the growth and motility of OS cells
was through miR-145-5p. To the best of our knowledge, the present
study is the first to suggest that the circ_0008932/miR-145-5p axis
regulates the proliferation, migration, invasion and apoptosis of
OS cells. Of note, this novel signaling pathway of circ_0008932 may
be a putative target in future therapies for patients with OS.
Supplementary Material
Si-circ_0008932 inhibitsosteosarcoma
cell invasion and migration by targeting miR-145-5p. (A) A
Transwell assay was used to examine the invasion and migration of
SaOS-2 cells after co-transfection with si-circ_0008932 and
miR-145-5p inhibitors. (B) A Transwell assay was used to determine
the invasion and migration of U2OS cells co-transfected with
si-circ_0008932 and miR-145-5p mimics. Magnification, x200. miR,
microRNA; si, short interfering RNA; NC, negative control.
Si-circ_0008932 promotesosteosarcoma
cell invasion and migration by targeting miR-145-5p. (A) Apoptosis
was measured in SaOS-2 cells after co-transfection with
si-circ_0008932 and miR-145-5p inhibitors. (B) The apoptosis was
examined in U2OS cells co-transfected with si-circ_0008932 and
miR-145-5p mimics. PI, propidium iodide; A, absorbance; miR,
microRNA; si, short interfering RNA; NC, negative control.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the present
study are available from the corresponding author on reasonable
request.
Authors' contributions
XS designed the study. CC and XS performed the
experiments and analyzed the data. Both authors checked and
approved the authenticity of the data. Both authors read and
approved the final version of the manuscript.
Ethics approval and consent to
participate
The present study was approved by the Ethics
Committee of Chongqing University Cancer Hospital (Chongqing,
China). Written informed consent was obtained from all patients for
the use of their clinical tissues.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Benjamin RS: Adjuvant and neoadjuvant
chemotherapy for osteosarcoma: A historical perspective. Adv Exp
Med Biol. 1257:1–10. 2020.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Garcia MB, Ness KK and Schadler KL:
Exercise and physical activity in patients with osteosarcoma and
survivors. Adv Exp Med Biol. 1257:193–207. 2020.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Anderson PM: Radiopharmaceuticals for
treatment of osteosarcoma. Adv Exp Med Biol. 1257:45–53.
2020.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Kiany S, Harrison D and Gordon N: The
histone deacetylase inhibitor Entinostat/Syndax 275 in
osteosarcoma. Adv Exp Med Biol. 1257:75–83. 2020.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Li Z, Liu S, Li X, Zhao W, Li J and Xu Y:
Circular RNA in Schizophrenia and depression. Front psychiatry.
11(392)2020.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Meng L, Ding P, Liu S, Li Z, Sang M and
Shan B: The emerging prospects of circular RNA in tumor immunity.
Ann Transl Med. 8(1091)2020.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Jiang X and Chen D: Circular RNA
hsa_circ_0000658 inhibits osteosarcoma cell proliferation and
migration via the miR-1227/IRF2 axis. J Cell Mol Med. 25:510–520.
2021.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Gu R, Li X, Yan X, Feng Z and Hu A:
Circular RNA circ_0032462 enhances osteosarcoma cell progression by
promoting KIF3B expression. Technol Cancer Res Treat: Nov 6, 2020
(Epub ahead of print). doi: 10.1177/1533033820943217.
|
|
9
|
Wang J, Zhu W, Tao G and Wang W: Circular
RNA circ-LRP6 facilitates Myc-driven tumorigenesis in esophageal
squamous cell cancer. Bioengineered. 11:932–938. 2020.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Hall IF, Climent M, Quintavalle M, Farina
FM, Schorn T, Zani S, Carullo P, Kunderfranco P, Civilini E,
Condorelli G and Elia L: Circ_Lrp6, a Circular RNA enriched in
vascular smooth muscle cells, acts as a sponge regulating miRNA-145
function. Circ Res. 124:498–510. 2019.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Prats AC, David F, Diallo LH, Roussel E,
Tatin F, Garmy-Susini B and Lacazette E: Circular RNA, the key for
translation. Int J Mol Sci. 21(8591)2020.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Tsitsipatis D and Gorospe M: Practical
guide forcircular RNA analysis: Steps, tips, and resources. Wiley
Interdiscip Rev RNA. 12(e1633)2021.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Saikishore R, Velmurugan P, Ranjithkumar
D, Latha R, Sathiamoorthi T, Arun A, Ravi AV and Sivakumar S: The
circular RNA-miRNA axis: A special RNA signature regulatory
transcriptome as a potential biomarker for OSCC. Mol Ther Nucleic
Acids. 22:352–361. 2020.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Wen J, Liao J, Liang J, Chen X, Zhang B
and Chu L: Circular RNA HIPK3: A key circular RNA in a variety of
human cancers. Front Oncol. 10(773)2020.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Su Q and Lv X: Revealing new landscape of
cardiovascular disease through circular RNA-miRNA-mRNA axis.
Genomics. 112:1680–1685. 2020.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Gong C, Zhou X, Lai S, Wang L, Liu J,
Yanming X and Xu Y: Long noncoding RNA/Circular RNA-miRNA-mRNA axes
in Ischemia-Reperfusion injury. Biomed Res Int.
2020(8838524)2020.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Wong RR, Abd-Aziz N, Affendi S and Poh CL:
Role of microRNAs in antiviral responses to dengue infection. J
Biomed Sci. 27(4)2020.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Boca S, Gulei D, Zimta AA, Onaciu A, Magdo
L, Tigu AB, Ionescu C, Irimie A, Buiga R and Berindan-Neagoe I:
Nanoscale delivery systems for microRNAs in cancer therapy. Cell
Mol Life Sci. 77:1059–1086. 2020.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Ebrahimi N, Aslani S, Babaie F,
Hemmatzadeh M, Pourmoghadam Z, Azizi G, Jadidi-Niaragh F and
Mohammadi H: MicroRNAs implications in the onset, diagnosis, and
prognosis of osteosarcoma. Curr Mol Med: Dec 3, 2020 (Epub ahead of
print).
|
|
21
|
Fu Y, Wang Y, Bi K, Yang L, Sun Y, Li B,
Liu Z, Zhang F, Li Y, Feng C and Bi Z: MicroRNA-208a-3p promotes
osteosarcoma progression via targeting PTEN. Exp Ther Med.
20(255)2020.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Kushlinskii NE, Fridman MV and Braga EA:
Molecular mechanisms and microRNAs in osteosarcoma pathogenesis.
Biochemistry (Mosc). 81:315–328. 2016.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Zhou Y, Feng D, Gu X, Gao A and Liu Y: The
role and clinical significance of long noncoding RNA zinc finger
E-box-binding homeobox two antisense RNA 1 in promoting
osteosarcoma cancer cell proliferation, inhibiting apoptosis and
increasing migration by regulating miR-145. Anticancer Drugs.
32:168–177. 2021.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Li X, Liu Y, Zhang X, Shen J, Xu R, Liu Y
and Yu X: Circular RNA hsa_circ_0000073 contributes to osteosarcoma
cell proliferation, migration, invasion and methotrexate resistance
by sponging miR-145-5p and miR-151-3p and upregulating NRAS. Aging
(Albany NY). 12:14157–14173. 2020.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Feng Y, Zhu J, Ou C, Deng Z, Chen M, Huang
W and Li L: MicroRNA-145 inhibits tumour growth and metastasis in
colorectal cancer by targeting fascin-1. Brit J Cancer.
110:2300–2309. 2014.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Tang M, Lin L, Cai H, Tang J and Zhou Z:
MicroRNA-145 downregulation associates with advanced tumor
progression and poor prognosis in patients suffering osteosarcoma.
Onco Targets Ther. 6:833–838. 2013.PubMed/NCBI View Article : Google Scholar
|