Introduction
Cystathionine γ-lyase (CSE) is one of the three
enzymes in the transsulfuration pathway that is responsible for
producing endogenous hydrogen sulfide (H2S) (1). NaHS (an H2S donor)
activates p38 and Akt, increasing the expression of angiogenic
factors and the proliferation of HUVECs (2). H2S is a gaseous endogenous
mediator that serves a potential role in modulating gastric
inflammatory responses (3). In
particular, S-propargyl-cysteine is a H2S donor that can
enhance human umbilical vein endothelial cell (HUVEC) cell
proliferation, adhesion, migration and tube formation (4). By contrast, exogenous thiosulfates,
which act in a slow manner to modulate sulfide metabolites, have
been documented to inhibit vascular endothelial growth factor
(VEGF)-dependent endothelial cell proliferation in a manner that is
associated with reductions in CSE protein levels (5). H2S has been shown to
inhibit the activation of NF-κB and the production of tumor
necrosis factor-α (TNFα) in cultured uterine smooth muscle cells
(6). Following CSE knockdown,
treatment with L-aspartate-β-hydroxamate, an aspartate
aminotransferase (AAT) inhibitor that blocks the generation of
endogenous sulfur dioxide (SO2)/AAT induced, was found
to aggravate the activation of the NF-κB pathway in endothelial
cells and its downstream inflammatory factors, including TNFα and
interleukin (IL)-6(7).
H2S exerts both pro-and antinociceptive effects through
inflammation (8). In addition, a
previous study observed an increase in inflammatory cytokine (IL-6
and TNFα) expression in the heart, and acute kidney injury (AKI)
can downregulate CSE-mediated H2S production, reduce
glutathione levels and increase oxidative stress in the heart
(9).
NF-κB is a heterodimer that is involved in a variety
of signaling pathways (10). As
such, NF-κB can regulate inflammatory responses by inducing the
expression of a number of genes, such as IL-6 and intercellular
adhesion molecule-1(11), whilst
NaHS suppresses intracellular adhesion molecule-1 expression in
TNFα-treated HUVECs (12). TNFα is
the multifunctional cytokine that is secreted primarily by
macrophages, natural killer (NK) cells and lymphocytes (13). H2S can prevent an
endothelial monolayer activation triggered by TNFα. The mechanism
of this protective effect is mainly mediated by downmodulation of
ADAM17-dependent TNF-converting enzyme activity with consequent
inhibition of soluble TNF-a shedding and its relevant MCP-1 release
in the medium (14). CSE knockdown
has been shown to protect primary hepatocytes from
D-galactosamine/TNFα-induced cell death without affecting
LPS-induced TNFα production by primary peritoneal macrophages
(15). LPS induces significant
increases in the plasma levels of multiple cytokines (TNFα, IL-1β,
IL-6, IL-10, IL-12 and interferon γ), but TNFα, IL-10 and IL-12
levels tended to be lower in male WT mice (C57/BL6) compared with
heterozygous cystathionine β-synthase (CBS) mice and CSE-knockout
mice, which express lower H2S-producing enzymes CBS, CSE
and 3-mercaptopyruvate sulfurtransferase (16). The findings indicate that CSE
ameliorates the outcome of LPS-induced endotoxemia in vivo
(16). By contrast, exogenous
H2S can attenuate angiotensin II-induced inflammation
and cytotoxicity by inhibiting the endothelin 1/NF-κB signaling
pathway in HUVECs (17).
CSE-generated H2S has also been shown to promote
prostate cancer progression and metastasis through the IL-1β/NF-κB
signaling pathway (18). In
addition, H2S can regulate lipopolysaccharide
(LPS)-induced inflammation and apoptosis of the mammary alveolar
epithelial cell lines, MAC-T by activating the PI3K/Akt/NF-κB
signaling pathway (19).
The inflammatory cytokine TNFα serves a pivotal role
in the disruption of macrovascular and microvascular circulation
both in vivo and in vitro (20). Although the H2S/CSE
signaling pathway is involved in inflammation (19,21-23),
mechanism underlying regulation of the CSE gene in HUVECs following
treatment with TNFα remains poorly understood. Therefore, the aim
of the present study was to investigate the role of the NF-κB
transcription factor binding site on the transcription of the CSE
gene in HUVECs that were treated with TNFα.
Materials and methods
Construction of the luciferase
reporter under the control of human CSE promoter
HUVECs were obtained from the School of Pharmacy of
Fudan University. The cultured cells were maintained in DMEM
(Thermo Fisher Scientific, Inc.) supplemented with 10% FBS (Thermo
Fisher Scientific, Inc.), 100 U/ml penicillin, and 0.1 mg/ml
streptomycin in a humidified atmosphere composed of 95% air and 5%
CO2 at 37˚C. HUVECs were cultured to a confluence of
80-90% and digested with trypsin at 5,000 g/5 min with RT. The
cells were collected into a 1.5 ml centrifuge tube. Subsequently,
genomic DNA was extracted from the HUVECs using a blood genomic DNA
extraction kit (Beijing Transgen Biotech Co., Ltd.). For
identification, 1% agarose gel electrophoresis was used. The
sequence of the human CSE gene promoter was searched on the GenBank
database, based on which the upstream and downstream primers were
designed. The target fragment DNA length was 710 bp (-696±16 nt).
According to the GenBank database of the human CSE (accession no.
NG_008041.1) gene promoter sequences and the Primer-BLAST online
application (https://www.ncbi.nlm.nih.gov/tools/primer-blast/),
a 710-bp DNA transcription start site of human CSE gene was
amplified by PCR using the HUVEC genomic DNA as the template
(forward, 5'-CGGGGTACCCATTAGGGGGAGTTTCTCTCTGT-3' and reverse,
5'-CCGCTCGAGCTGCAGTCTCACGATCACAGT-3'; Promega Corporation). The
PrimeSTAR HS DNA Polymerase PCR kit (cat. no. R010A) was purchased
from Takara Bio, Inc. The thermocycling conditions were as follows:
Initial denaturation at 94˚C for 3 min; followed by 30 cycles at
95˚C for 30 sec, 60˚C for 45 sec and 72˚C for 90 sec; and a final
step at 72˚C for 10 min. The PCR product was digested with the
restriction enzymes KpnI and XhoI (Takara
Biotechnology Co., Ltd.) and cloned into the pGL4.12 vector
(Promega Corporation) that containing a firefly luciferase gene
driven by the inserted promoter (pGL4.12). The resultant construct
was designated as pGL4.12-HuCSE710. The inserted DNA fragment was
confirmed by Sanger sequencing by Sangon Biotech Co., Ltd. The
reporter with the mutant CSE promoter was the same as that
aforementioned, except that an alternative forward primer
(5'-CGGGGTACCCATTAGGATCTGTTTCTCTCTGT-3') was used during PCR
amplification. Both the Wild-type promoter and the Mutant promoter
were transformed into Trans5α chemically competent cells (A strain
of Escherichia coli (E. coli); TransGen Biotech Co.,
Ltd.), cultured overnight at 37˚C before a single bacterial colony
was selected, followed by culture overnight in a shaking bed at
37˚C for amplification. Plasmids were extracted using the EasyPure
Plasmid MiniPrep kit (cat. no. EM101-01; TransGen Biotech Co.,
Ltd.), which was analyzed by restriction enzyme digestion. The
fragment size was identified and sequenced by Sanger sequencing by
Sangon Biotech Co., Ltd. The sequencing results were compared with
the published sequences on the GenBank database for analysis.
Cell culture and treatments
293T cell lines (cat. no. GNHu17) were purchased
from The Cell Bank of Type Culture Collection of the Chinese
Academy of Sciences. Cell culture reagents were purchased from
Thermo Fisher Scientific, Inc. The 293T cells and HUVECs were
cultured in a 5% CO2/balance air incubator at 37˚C in
DMEM supplemented with 10% heat-inactivated FBS and with the
addition of 100 U/ml; penicillin G, 100 µg/ml streptomycin and 6.5
mM L-glutamine. According to the different cell lines, the
transfected 293T cells and HUVECs were cultured at a density of
0.5-1x106 cells in 35-mm dishes (Corning, Inc.) prior to
treatment. For treatment with TNFα (Recombinant Human 4-1BB
Receptor, 5 µg; Beyotime Institute of Biotechnology), all tested
cells were incubated with TNFα (10, 30 and 50 ng/ml) for 1, 3 and 6
h at 37˚C. The controls received the same volume of saline as the
TNFα-treated cells. Following incubation and the removal of the
cell medium, luciferase assay, reverse transcription-quantitative
(RT-qPCR) and western blot analysis were performed.
Luciferase assay
For transfection, the 293T cells were grown to
70-80% confluency. Subsequently, 5 µg pGL4.12-HuCSE710 or
pGL4.12-HuCSE710 m (the mutant promoter) together with 0.028 µg the
pRL-CMV control vector were transfected into the cells per 35-mm
dishes using Xfect™ transfection reagent (Takara Bio, Inc.)
according to the manufacturer's protocols. After 12 h, the
transfected cells were sub-cultured in several 35-mm dishes at a
proportion of 1:3 for 24 h. Following treatment with TNFα for 1, 3,
and 6 h after 36 h of transfection, both Firefly luciferase and
Renilla luciferase activities were assayed using the
TransDetect Double-Luciferase Reporter Assay kit (Beijing Transgen
Biotech Co., Ltd.) according to the manufacturer's protocol using
the Multimode Microplate Reader (Berthold Technologies GmbH &
Co., K.G.). Firefly luciferase activity was normalized against
Renilla luciferase activity.
Isolation of RNA and RT-qPCR
According to the manufacturer's protocol, total RNA
of HUVECs was isolated using the TransZol Up reagent (Beijing
Transgen Biotech Co., Ltd.). Reaction Mix and
TransScript® RT/RI Enzyme mix (Transgen Biotech Co.,
Ltd.) was used for reverse transcription. The temperature protocol
for reverse transcription was as follows: Incubation at 42˚C for 30
min and inactivation at 85˚C for 5 sec. TransStart®
Green qPCR SuperMix Real-Time PCR kit (Beijing Transgen Biotech
Co., Ltd.) was used for qPCR. The following thermocycling
conditions were used: Preincubation at 94˚C for 30 sec, followed by
a 2-step amplification of 94˚C for 5 sec and 60˚C for 30 sec. An
oligo-dT primer (Beijing Transgen Biotech Co., Ltd.) was used for
mRNA reverse transcription. The sequences of the primers used for
qPCR are summarized in Table I. To
design the primer pairs, CSE Forward Primer/CSE Reversed Primer
(Table I) was used to determine the
relative expression of CSE. By verifying that both ACTB
(β-actin gene) and CSE mRNA primers had similar amplifying
efficiencies, the comparative 2-ΔΔCq method was used for
performing relative quantification analysis of the mRNA levels
(24). Roche
LightCycler® 96 System (Roche Molecular Diagnostics) was
used for cDNA amplification and detection.
 | Table IPrimers used for reverse
transcription-quantitative PCR. |
Table I
Primers used for reverse
transcription-quantitative PCR.
| Gene | GenBank Accession
number | Primer
sequence | Exon | Amplicon size |
|---|
| CSE | NM_001902.5 |
F:5'-GGCTCTACCTGCGTGCTTTA-3' | 1 | 118 bp |
| | |
R:5'-CGCGAAAGAAGAAGAGAGGA-3' | 1 | |
| ACTB | NM_001101.3 |
F:5'-CTCTTCCAGCCTTCCTTCCT-3' | 2 | 109 bp |
| | |
R:5'-TGTTGGCGTACAGGTCTTTG-3' | 2 | |
Western blot analysis
Western blot analysis was performed according to a
previously described method (25).
For total protein extraction, 0.5x106 HUVECs were
incubated in 120 µl mild RIPA lysis buffer (Beijing Transgen
Biotech Co., Ltd.) supplemented with 1 mM PMSF proteinase, 0.25
U/µl Benzonase and inhibitor cocktail (Takara Bio, Inc.). The level
of protein was determined using the bicinchoninic acid assay
method. For 10% SDS-PAGE, the protein samples (30 µg) were mixed
with loading buffer and boiled at 98˚C for 10 min. The separated
proteins were transferred onto a 0.45 µm PVDF membrane (EMD
Millipore). The membrane was incubated at 4˚C with anti-CSE (cat.
no. D199513, Sangon Biotech Co., Ltd.) mouse monoclonal antibodies
(1:1,000 dilution) or anti-ACTB (cat. no. D191047, Sangon Biotech
Co., Ltd.) mouse monoclonal antibodies (1:2,000 dilution) for 12 h.
After another wash, the membrane was incubated with HRP-conjugated
goat anti-mouse antibodies (1:5,000 dilution; cat. no. D110087;
Sangon Biotech Co., Ltd.) for 2 h, The results were scanned for the
detection of CSE and ACTB with BeyoECL Plus (cat. no. P0018S;
Beyotime Institute of Biotechnology) and quantified using the
AlphaView System (ProteinSimple) sensitive chemiluminescent imaging
system with a separate instrument software (FluorChem HD2; v3.4.0;
ProteinSimple).
Statistical analysis
All data are expressed as the means ± SEM from ≥
four experiments. Shapiro-Wilk test is used to confirm the
normality of the data distribution. Multiple group comparisons were
performed using one-way ANOVA followed by Tukey's post hoc test.
P<0.05 was considered to indicate a statistically significant
difference.
Results
Effects of TNFα treatment on the CSE
mRNA level
To analyze the effects of TNFα on the transcription
of the CSE gene, the expression level of CSE mRNA in HUVECs was
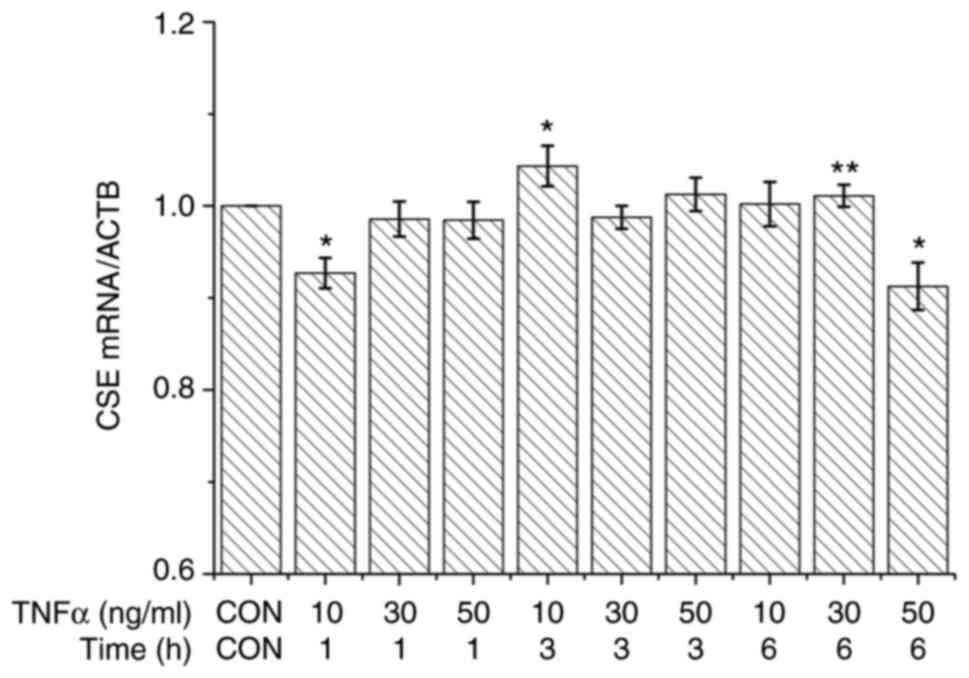
examined. As shown in Fig. 1,
following treatment with TNFα (10, 30 and 50 ng/ml) for 1, 3 and 6
h, the CSE mRNA levels in HUVECs decreased with 10 ng/ml TNFα for 1
h compared to all other groups except 50 ng/ml for 6 h. In
addition, CSE mRNA levels in HUVECs increased following treatment
with TNFα at 10 ng/ml for 3 h compared to all other groups.
However, CSE mRNA levels of cells treated with higher
concentrations of TNFα (50 ng/ml) were reduced at 3 h compared with
that at 6 h, where it probably exerted toxic effects on the cells.
At 3 h, a ‘U’ curve in CSE mRNA expression was observed in the
cells treated with ascending concentrations of TNFα. At 6 h, an
inverted ‘U’ curve was observed in the expression of CSE mRNA in
the cells treated with ascending concentrations of TNFα. This
result indicated that HUVECs treated with various concentrations of
TNFα differed significantly among the different treatment
times.
Effects of TNFα on the CSE protein
level
The effect of TNFα on CSE protein expression in
HUVECs was next investigated. As shown in Fig. 2, following treatment with TNFα (10
ng/ml) for 1 h, CSE protein expression decreased significantly
compared with that in the control group. Following treatment with
TNFα (10, 30 and 50 ng/ml) for 1, 3 and 6 h, the CSE protein level
in the HUVECs increased, particularly at the concentration of 10
ng/ml for 3 h of treatment compared with that for 1 h. Compared
with that in the control, CSE protein expression in the HUVECs
increased with TNFα at 30 ng/ml for 1 h and 50 ng/ml for 3 h but
declined slightly in other conditions compared to all other groups
except 10 ng/ml for 3 h. These results suggested that the same
concentration of TNFα exerted differential effects on the CSE
protein level in a manner that was dependent on the treatment
duration. Although TNFα-induced upregulation of CSE expression
within 3 h at 50 ng/ml TNFα was evident compared to all other
groups, TNFα can affect CSE expression in a concentration and
duration-dependent manner.
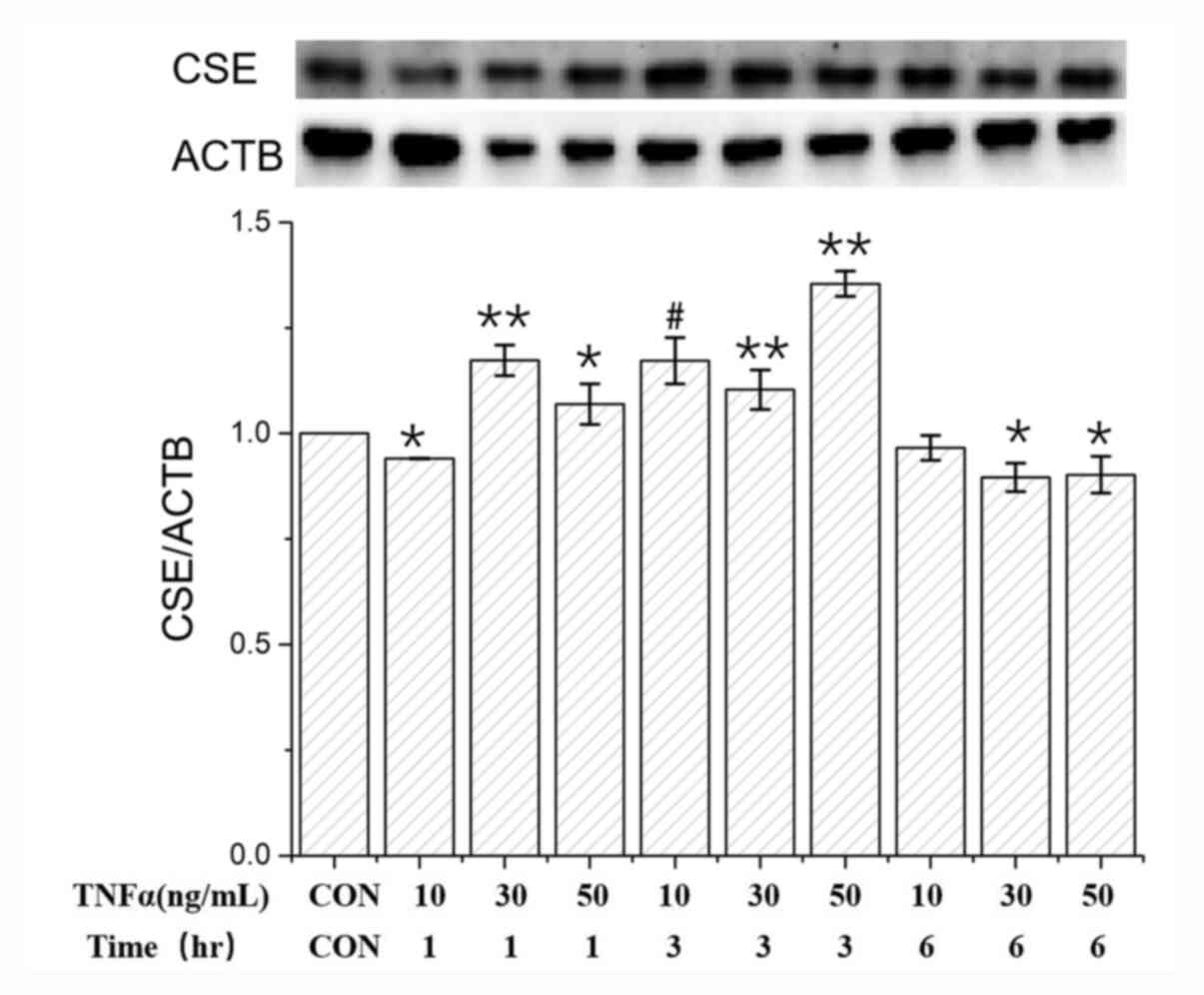 | Figure 2Effects of TNFα on the CSE expression
in HUVECs at the protein level. Following treatment with TNFα (10
ng/ml) for 1 h, CSE protein expression significantly decreased
compared with that in the control. Following treatment with TNFα
(10, 30 and 50 ng/ml) for 1, 3 and 6 h, CSE protein expression in
HUVECs increased at the concentration of 10 ng/ml with 3 h of
treatment compared with 10 ng/ml after 1 h, whilst the CSE protein
level in the HUVECs were increased at the concentration of 50 ng/ml
for 3 h treatment compared with 50 ng/ml after 6 h. Compared to the
control, CSE protein expression in the HUVECs markedly increased
with TNFα at 30 ng/ml for 1 h and 50 ng/ml for 3 h but declined
slightly in other cases (*P<0.05 and
**P<0.01 vs. CON; #P<0.05, 10 ng/ml for
3 h vs. 10 ng/ml for 1 h). HUVECs, human umbilical vein endothelial
cells; TNFα, tumor necrosis factor α; CSE, cystathionine
γ-lyase. |
Effect of TNFα on the wild-type or
mutant promoter activity of the CSE gene
The effects of TNFα on the transactivation
activities of the promoter of the CSE gene were subsequently
analyzed by transient transfection experiments. By bioinformatics
analysis, a potential NF-κB binding site was identified on the
human CSE gene promoter with the DNA sequence of 5'-GGGACATTCC-3'.
The reporter luciferase expression vector was constructed, which
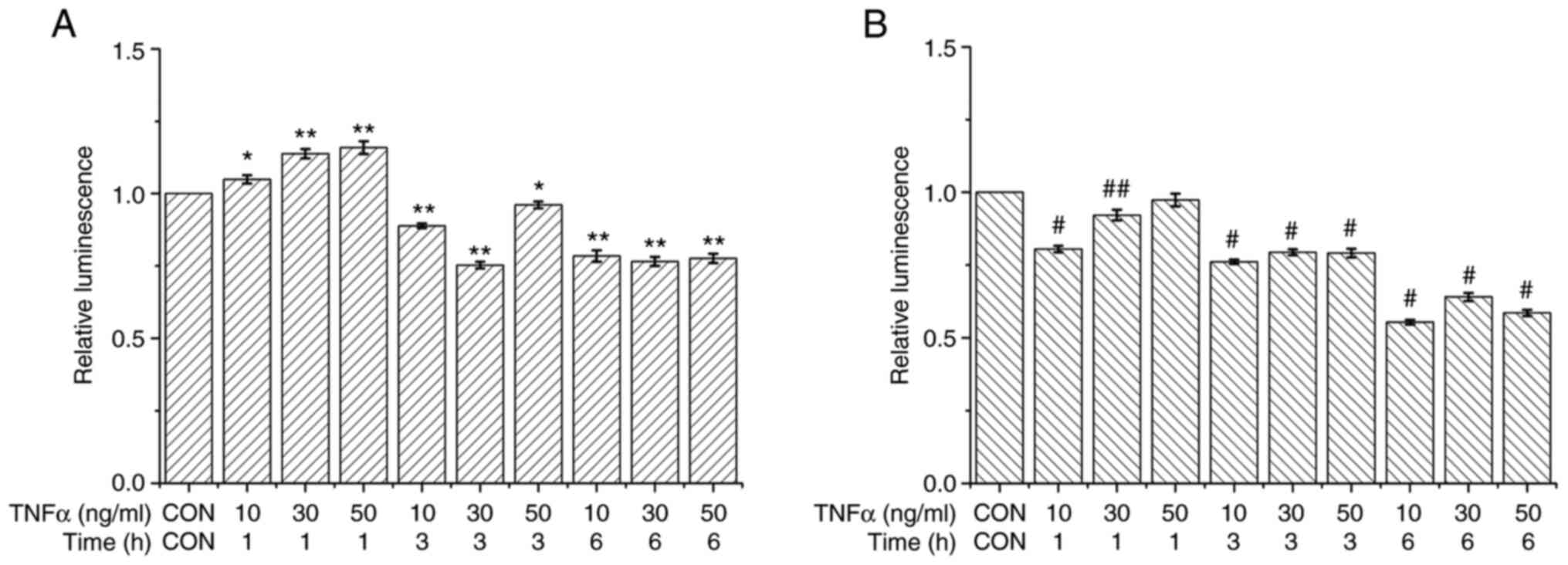
was either controlled either by the wild-type (Fig. 3A) or mutant (Fig. 3B) promoter region of the CSE gene,
as shown. As shown in Fig. 3A,
following treatment with TNFα (10, 30 and 50 ng/ml) for 1, 3 and 6
h, the wild-type promoter activity of the CSE gene in transfected
293T cells increased with the increment in the concentration of
TNFα at 1 h compared to all other group.
As shown in Fig. 3B,
the CSE gene mutant promoter activity decreased with the
concentrations of TNFα at 1 h compared to TNFα (30 and 50 ng/ml)
for 1, 3 and 6 h. These results revealed that TNFα regulated CSE
gene transcription via the NF-κB binding site on the CSE gene
promoter in HUVECs. Just as CSE mRNA expression was different
following treatment with different concentrations of TNFα, the
activity of CSE promoter was also similar at varying concentrations
of TNFα. These results suggest that the level of TNFα at different
treatment times affected the expression of CSE.
Discussion
The H2S/CSE signaling pathway is involved
in various inflammatory conditions, whereas TNFα is one of the
inflammatory cytokines that is activated during sepsis (26-29).
The concentration of H2S is enhanced when abdominal
sepsis or endotoxemia occurs, and administration of H2S
leads to exacerbation of these conditions, mainly because of its
pro-inflammatory effect (30,31).
It has also been reported that the concentration of H2S
rose in response to the presence of pancreatitis, which is ascribed
to its pro-inflammatory effect (32) NaHS treatment also exerts
anti-inflammatory effects through the inhibition of nitric oxide
and TNFα production in MC3T3-E1 osteoblastic cells, suggesting an
anti-inflammatory effect of H2S (33). The anti-inflammatory effect of
H2S has been previously reported, particularly on TNF
production, in addition to the molecular mechanisms involving NF-κB
inhibition and the reduced expression of IL-6, TNF and IL-1β by
H2S (34). For example,
H2S production facilitates the pathogenesis of severe
acute pancreatitis (35).
Additionally, NaHS treatment has been shown to improve wound
healing in ob/ob mice (hyperphagic, obese, hyperinsulinemic and
hyperglycemic mice), which is associated with the decreased
production of TNFα and IL-6(36).
The inflammatory role of H2S remains controversial due
to findings that it exerts both proinflammatory and
anti-inflammatory effects (37).
TNF-α induces the gene expression of various
inflammatory cytokines and chemokines, such as IL-6, IL-8 and
MCP-1, either dependently or independently to activate
transcriptional factors, such as NF-κB and activator protein
1(20). Results from the present
study demonstrated that transcriptional regulation of the CSE gene
was possibly mediated on the NF-κB binding site on the promoter in
HUVECs after treatment with TNFα. After the transfected 293T cells
were incubated with TNFα (10 ng/ml) for 1, 3 and 6 h, the wild-type
promoter activity of the CSE gene significantly increased at 1 h.
However, the mutant-type promoter activity of the CSE gene
considerably decreased at 1 h. When comparing the action of the
wild-type promoter and the mutant-type, it was found that the CSE
gene promoter activity significantly decreased with the mutation at
10, 30 and 50 ng/ml for 1 h. CSE expression was upregulated at a
low concentration of TNFα over a short time period, whereas it was
suppressed with the high concentration of TNFα and a longer
treatment time. This suggest that TNFα exerted a concentration- and
time-dependent effect on CSE expression. In HUVECs, the high
concentration (50 ng/ml) of TNFα and a longer treatment duration (6
h) downregulated CSE expression, which may reduce endogenous
H2S production and induce the inhibition of vascular
smooth muscle relaxation (38)
resulting in reduced local blood flow (39). This may lead to the chronic
insufficiency of blood supply to local tumor tissue, resulting in
the induction of apoptosis (19)
and in the prevention of tumor recurrence, this suggests a
potential therapeutic target for tumor (40). These results demonstrated that the
DNA sequence, GGGACATTCC, on the CSE gene promoter was directly
associated with the transcriptional regulation of the CSE gene in
mammalian cells treated with TNFα. In the present study, the effect
of TNFα on the transcription regulation and expression of the CSE
gene in HUVECs and 293T cells. However, further studies in animal
models are required in future experiments.
In conclusion, results from the present study
suggest that the NF-κB binding site on the CSE promoter is
potentially important for TNFα-induced CSE expression. TNFα can
potentially affect CSE gene expression, where vascular endothelial
cells can respond to TNFα in the blood by regulating the CSE gene
expression. Consequently, the regulatory mechanisms associated with
the effects of TNFα on the transcriptional regulation of the CSE
gene in HUVECs via the NF-κB pathway warrant further
investigation.
Acknowledgements
The author thanks Dr Fei Zhou of Hanshan Normal
University (Guangdong, China) and Madame Rong-Lan Shi of Hanshan
Normal University (Guangdong, China) for their technical
assistance.
Funding
Funding: Natural Science Foundation of Guangdong Province (grant
no. 2016A030307039) supported this project.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
MW contributed to this study independently; read and
approved the final version of the manuscript; and confirmed the
authenticity of all the raw data.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Wang R: Hydrogen sulfide: The third
gasotransmitter in biology and medicine. Antioxid Redox Signal.
12:1061–1064. 2010.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Choi KS, Song H, Kim EH, Choi JH, Hong H,
Han YM and Hahm KB: Inhibition of hydrogen sulfide-induced
angiogenesis and inflammation in vascular endothelial cells:
Potential mechanisms of gastric cancer prevention by Korean red
ginseng. J Ginseng Res. 36:135–145. 2012.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Aboubakr EM, Taye A, El-Moselhy MA and
Hassan MK: Protective effect of hydrogen sulfide against cold
restraint stress-induced gastric mucosal injury in rats. Arch Pharm
Res. 36:1507–1515. 2013.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Kan J, Guo W, Huang C, Bao G, Zhu Y and
Zhu YZ: S-propargyl-cysteine, a novel water-soluble modulator of
endogenous hydrogen sulfide, promotes angiogenesis through
activation of signal transducer and activator of transcription 3.
Antioxid Redox Signal. 20:2303–2316. 2014.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Leskova A, Pardue S, Glawe JD, Kevil CG
and Shen X: Role of thiosulfate in hydrogen sulfide-dependent redox
signaling in endothelial cells. Am J Physiol Heart Circ Physiol.
313:H256–H264. 2017.PubMed/NCBI View Article : Google Scholar
|
|
6
|
You X, Chen Z, Zhao H, Xu C, Liu W, Sun Q,
He P, Gu H and Ni X: Endogenous hydrogen sulfide contributes to
uterine quiescence during pregnancy. Reproduction. 153:535–543.
2017.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Zhang D, Wang X, Tian X, Zhang L, Yang G,
Tao Y, Liang C, Li K, Yu X, Tang X, et al: The increased endogenous
sulfur dioxide acts as a compensatory mechanism for the
downregulated endogenous hydrogen sulfide pathway in the
endothelial cell inflammation. Front Immunol. 9(882)2018.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Garattini EG, Santos BM, Ferrari DP, Capel
CP, Francescato HDC, Coimbra TM, Leite-Panissi CRA, Branco LGS and
Nascimento GC: Propargylglycine decreases neuro-immune interaction
inducing pain response in temporomandibular joint inflammation
model. Nitric Oxide. 93:90–101. 2019.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Wijerathne CUB, Madduma Hewage S, Siow YL
and O K: Kidney ischemia-reperfusion decreases hydrogen sulfide and
increases oxidative stress in the heart. Biomolecules.
10(1565)2020.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Pahl HL: Activators and target genes of
Rel/NF-kappaB transcription factors. Oncogene. 18:6853–6866.
1999.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Baeuerle PA and Baltimore D: NF-kappa B:
Ten years after. Cell. 87:13–20. 1996.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Wang Y, Zhao X, Jin H, Wei H, Li W, Bu D,
Tang X, Ren Y, Tang C and Du J: Role of hydrogen sulfide in the
development of atherosclerotic lesions in apolipoprotein E knockout
mice. Arterioscler Thromb Vasc Biol. 29:173–179. 2009.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Clyde K and Glaunsinger BA: Chapter 1 -
Getting the Message: Direct Manipulation of Host mRNA Accumulation
During Gammaherpesvirus Lytic. Infection. In: Advances in Virus
Research. Maramorosch K, Shatkin AJ and Murphy FA (eds). Vol 78.
Academic Press, Burlington, MA, pp1-42, 2010.
|
|
14
|
Perna AF, Sepe I, Lanza D, Capasso R,
Zappavigna S, Capasso G, Caraglia M and Ingrosso D: Hydrogen
sulfide reduces cell adhesion and relevant inflammatory triggering
by preventing ADAM17-dependent TNF-α activation. J Cell Biochem.
114:1536–1548. 2013.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Shirozu K, Tokuda K, Marutani E, Lefer D,
Wang R and Ichinose F: Cystathionine γ-lyase deficiency protects
mice from galactosamine/lipopolysaccharide-induced acute liver
failure. Antioxid Redox Signal. 20:204–216. 2014.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Ahmad A, Gero D, Olah G and Szabo C:
Effect of endotoxemia in mice genetically deficient in
cystathionine-γ-lyase, cystathionine-β-synthase or
3-mercaptopyruvate sulfurtransferase. Int J Mol Med. 38:1683–1692.
2016.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Hu HJ, Jiang ZS, Zhou SH and Liu QM:
Hydrogen sulfide suppresses angiotensin II-stimulated endothelin-1
generation and subsequent cytotoxicity-induced endoplasmic
reticulum stress in endothelial cells via NF-κB. Mol Med Rep.
14:4729–4740. 2016.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Wang YH, Huang JT, Chen WL, Wang RH, Kao
MC, Pan YR, Chan SH, Tsai KW, Kung HJ, Lin KT and Wang LH:
Dysregulation of cystathionine γ-lyase promotes prostate cancer
progression and metastasis. EMBO Rep. 20(e45986)2019.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Wang Y, Zhang C, Xu C, Feng L, Li A, Jin
X, Guo S, Jiao X, Liu J, Guo Y, et al: H2S mediates apoptosis in
response to inflammation through PI3K/Akt/NFkB signaling pathway.
Biotechnol Lett. 42:375–387. 2020.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Zhang H, Park Y, Wu J, Chen XP, Lee S,
Yang J, Dellsperger KC and Zhang C: Role of TNF-alpha in vascular
dysfunction. Clin Sci (Lond). 116:219–230. 2009.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Wu C, Xu Z and Huang K: Effects of dietary
selenium on inflammation and hydrogen sulfide in the
gastrointestinal tract in chickens. Biol Trace Elem Res.
174:428–435. 2016.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Xu C, Wan Y, Guo T and Chen X: Effect of
hydrogen sulfide on the expression of CSE, NF-kB, and IL-8 mRNA in
GES-1 cells with Helicobacter pylori infection. Zhong Nan Da Xue
Xue Bao Yi Xue Ban. 38:977–983. 2013.PubMed/NCBI View Article : Google Scholar : (In Chinese).
|
|
23
|
Xu Y, Du HP, Li J, Xu R, Wang YL, You SJ,
Liu H, Wang F, Cao YJ, Liu CF and Hu LF: Statins upregulate
cystathionine γ-lyase transcription and H2S generation via
activating Akt signaling in macrophage. Pharmacol Res. 87:18–25.
2014.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Schmittgen TD and Livak KJ: Analyzing
real-time PCR data by the comparative C(T) method. Nat Protoc.
3:1101–1108. 2008.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Wang M, Guo Z and Wang S: The binding site
for the transcription factor, NF-kB, on the cystathionine γ-lyase
promoter is critical for LPS-induced cystathionine γ-lyase
expression. Int J Mol Med. 34:639–645. 2014.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Zhang H, Zhi L, Moochhala S, Moore PK and
Bhatia M: Hydrogen sulfide acts as an inflammatory mediator in
cecal ligation and puncture-induced sepsis in mice by upregulating
the production of cytokines and chemokines via NF-kappaB. Am J
Physiol Lung Cell Mol Physiol. 292:L960–L971. 2007.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Tokuda K, Kida K, Marutani E, Crimi E,
Bougaki M, Khatri A, Kimura H and Ichinose F: Inhaled hydrogen
sulfide prevents endotoxin-induced systemic inflammation and
improves survival by altering sulfide metabolism in mice. Antioxid
Redox Signal. 17:11–21. 2012.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Marko L, Szijártó IA, Filipovic MR,
Kaßmann M, Balogh A, Park JK, Przybyl L, N'diaye G, Krämer S,
Anders J, et al: Role of cystathionine gamma-lyase in immediate
renal impairment and inflammatory response in acute ischemic kidney
injury. Sci Rep. 6(27517)2016.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Ivanciuc T, Sbrana E, Ansar M, Bazhanov N,
Szabo C, Casola A and Garofalo RP: Hydrogen sulfide is an antiviral
and antiinflammatory endogenous gasotransmitter in the airways.
Role in respiratory syncytial virus infection. Am J Respir Cell Mol
Biol. 55:684–696. 2016.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Wallace JL: Hydrogen sulfide-releasing
anti-inflammatory drugs. Trends Pharmacol Sci. 28:501–505.
2007.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Ganster F, Burban M, de la Bourdonnaye M,
Fizanne L, Douay O, Loufrani L, Mercat A, Calès P, Radermacher P,
Henrion D, et al: Effects of hydrogen sulfide on hemodynamics,
inflammatory response and oxidative stress during resuscitated
hemorrhagic shock in rats. Crit Care. 14(R165)2010.PubMed/NCBI View
Article : Google Scholar
|
|
32
|
Guo W, Cheng ZY and Zhu YZ: Hydrogen
sulfide and translational medicine. Acta Pharmacol Sin.
34:1284–1291. 2013.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Xu ZS, Wang XY, Xiao DM, Hu LF, Lu M, Wu
ZY and Bian JS: Hydrogen sulfide protects MC3T3-E1 osteoblastic
cells against H2O2-induced oxidative damage-implications for the
treatment of osteoporosis. Free Radic Biol Med. 50:1314–1323.
2011.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Huang CW, Feng W, Peh MT, Peh K, Dymock BW
and Moore PK: A novel slow-releasing hydrogen sulfide donor,
FW1256, exerts anti-inflammatory effects in mouse macrophages and
in vivo. Pharmacol Res. 113:533–546. 2016.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Liu Y, Liao R, Qiang Z and Zhang C:
Pro-inflammatory cytokine-driven PI3K/Akt/Sp1 signalling and H2S
production facilitates the pathogenesis of severe acute
pancreatitis. Biosci Rep. 37(BSR20160483)2017.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Zhao H, Lu S, Chai J, Zhang Y, Ma X, Chen
J, Guan Q, Wan M and Liu Y: Hydrogen sulfide improves diabetic
wound healing in ob/ob mice via attenuating inflammation. J
Diabetes Complications. 31:1363–1369. 2017.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Castelblanco M, Lugrin J, Ehirchiou D,
Nasi S, Ishii I, So A, Martinon F and Busso N: Hydrogen sulfide
inhibits NLRP3 inflammasome activation and reduces cytokine
production both in vitro and in a mouse model of inflammation. J
Biol Chem. 293:2546–2557. 2018.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Kimura H: Production and physiological
effects of hydrogen sulfide. Antioxid Redox Signal. 20:783–793.
2014.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Wallace JL and Wang R: Hydrogen
sulfide-based therapeutics: Exploiting a unique but ubiquitous
gasotransmitter. Nat Rev Drug Discov. 14:329–345. 2015.PubMed/NCBI View
Article : Google Scholar
|
|
40
|
Sogutdelen E, Pacoli K, Juriasingani S,
Akbari M, Gabril M and Sener A: Patterns of expression of
H2S-producing enzyme in human renal cell carcinoma specimens:
Potential avenue for future therapeutics. In Vivo. 34:2775–2781.
2020.PubMed/NCBI View Article : Google Scholar
|