Introduction
Retinoblastoma (Rb) is a common malignant tumour
reported mainly in children and affecting ~8,000 infants worldwide
annually (1). Rb morbidity rate in
developed countries is lower than that observed in underdeveloped
countries, suggesting that Rb morbidity is closely associated with
effective diagnosis and treatment (2,3). At
present, the main treatments of Rb include enucleation, laser
photocoagulation, chemotherapy and focal therapy (4). However, the therapeutic effects remain
limited due to accelerated metastasis formation in Rb (5). It is therefore crucial to determine
effective therapeutic targets for Rb to diminish the formation of
metastasis.
Long non-coding RNAs (lncRNAs) are defined as
transcripts of >200 nucleotides in length (6). Certain lncRNAs have been reported to
contribute considerably to the pathogenesis of Rb, including
LINC00152(7), small nucleolar RNA
host gene 16 (SNHG16) (8), Pvt1
oncogene (PVT1) (9) and TP73
antisense RNA (TP73-AS1) (10).
Yang et al (8) demonstrated
that SNHG16 downregulation can restrain Rb cell migratory and
invasive abilities. Wu et al (9) reported that PVT1 silencing not only
suppresses the proliferation and migratory and invasive abilities
of Rb cells in vitro, but also inhibits the growth of tumour
xenografts in vivo. Wang et al (10) reported increased expression of
TP73-AS1 in Rb tissues and cell lines (Y79, HXO-RB44, WERI-Rb-1 and
SO-RB50), whereas TP37-AS1 overexpression could further aggravate
the malignant phenotype of Rb in vitro (10). All the aforementioned lncRNAs serve
as oncogenes in Rb. In addition, the lncRNA myocardial
infarction-associated transcript (MIAT) was demonstrated to
facilitate the progression of numerous types of human cancer,
including cholangiocarcinoma (CCA) (11), non-small cell lung cancer (NSCLC)
(12,13), cervical cancer (CC) (14), ovarian cancer (OC) (15), colorectal cancer (CRC) (16) and gastric cancer (GC) (17). However, the potential role and
underlying mechanism of MIAT in Rb remain unclear.
MicroRNAs (miRNAs/miRs) are a class of small
endogenous RNA that can target the 3'-untranslated region to
regulate gene expression (18). The
anti-tumour roles of certain miRNAs in human cancer, especially in
Rb, including miR-22-3p (19),
miR-124(20), miR-182-5p (8), miR-128-3p (8), miR-488-3p (9), miR-506-3p (21) and miR-936(22), have received increased attention.
Furthermore, miR-665 has been reported to serve a crucial role in
inhibiting Rb function (23,24). A
recent study demonstrated that miR-665 exerts its inhibitory effect
on Rb by inactivating the Wnt/β-catenin pathway (23). In addition, miR-665 regulation by
LINC00205 has been demonstrated to restrain Rb tumorigenesis
(24). The modulation of miR-665
activity by MIAT during Rb progression requires therefore further
investigation.
LIM and SH3 protein 1 (LASP1) is an oncogene in the
pathogenesis of several human cancers, such as hepatocellular
carcinoma (HCC) (25), CRC
(26), prostate cancer (27) and breast cancer (28). In these cancers, LASP1 has been
indicated to be regulated by miRNAs involved in cancer progression
(25,29). Hu et al (25) revealed that miR-326 repressed the
cell proliferation and invasion of HCC via directly targeting
LASP1. Song et al (29)
demonstrated that LASP1 is a downstream target of miR-342-3p
affecting the progression of oral squamous cell carcinoma. In
addition, Yang et al (8)
demonstrated that lncRNA SNHG16 promoted the migration and invasion
of Rb cells by regulating LASP1. Furthermore, interestingly, Liu
et al (30) indicated that
MIAT interacted with LASP1 in the development of papillary thyroid
cancer. Nevertheless, the association between miR-665 and LASP1, as
well as the role of the MIAT/miR-665/LASP1 axis in the pathogenesis
of Rb, are largely unknown.
In the present study, the expression levels of MIAT,
miR-665 and LASP1 in Rb tissues and cells were analysed. The
detailed regulatory mechanisms of the MIAT/miR-665/LASP1 axis in
the pathogenesis of Rb were also explored. The findings from this
study may provide a possible therapeutic target for Rb.
Materials and methods
Patients with Rb
A total of 47 patients with Rb (age range, 4 months
to 14 years; mean age, 5.54±3.09 years) were enrolled in the No.
215 Hospital of Shaanxi Nuclear Industry (Xianyang, China) between
March 2017 and July 2019. The Rb tissues and adjacent normal
tissues (1 cm from the edge of the tumour tissues) were collected
following an enucleation procedure. Tumour tissues were
histologically confirmed and the adjacent tissues without
histopathological changes were considered as the normal control
group. None of the patients received any preoperative radiotherapy
and/or chemotherapy. Each patient or their parents provided written
informed consent. The protocol was approved by the Ethics Committee
of the No. 215 Hospital of Shaanxi Nuclear Industry (approval no.
EC-20200924-1017).
Cell culture and transfection
The human retinal epithelial ARPE-19 cell line and
the human Rb Y79, HXO-RB44, WERI-Rb-1 and SO-RB50 cell lines were
obtained from the American Type Culture Collection. These cell
lines were selected according to previous studies (9,31-34).
All cells were cultured in Dulbecco's modified Eagle's medium
(DMEM; Invitrogen; Thermo Fisher Scientific, Inc.) containing 10%
FBS (Gibco; Thermo Fisher Scientific, Inc.) and placed at 37˚C in a
humidified incubator containing 5% CO2.
The plasmids used in the present study were
synthesised by Hanbio Biotechnology Co., Ltd. and included the
short hairpin (sh)RNA-MIAT (sh-MIAT; 5'-UCCUCCGAACCUGGCA CGU-3'),
shRNA-negative control (sh-NC; 5'-UUCUCCGAAC GUGUCACGU-3'), miR-665
mimics (5'-ACCAGGAGGCU GAGGCCCCU-3'), mimics-NC (miR-NC;
5'-UUCUCCGAA CGUGUCACGUTT-3'), miR-665 inhibitor (5'-AGGGGCCU
CAGCCUCCUGGU-3'), inhibitor NC (5'-CAGUACUUUUGU GUAGUACAA-3'),
LASP1 overexpression vector (pcDNA-LASP1) and pcDNA-negative
control (pcDNA-NC). Transfections (all the above molecules at 20
nM) were performed using Lipofectamine® 3000 (Thermo
Fisher Scientific, Inc.) according to the manufacturers'
instructions. After 48 h, cells were collected for subsequent
experiments.
Reverse transcription-quantitative
(RT-q)PCR
Total RNA was extracted from Rb tissues and the Y79,
HXO-RB44, WERI-Rb-1 and SO-RB50 cell lines using TRIzol®
(Invitrogen; Thermo Fisher Scientific, Inc.). According to the
manufacturer's instructions, RNA was reversed transcribed into cDNA
using the First-Strand cDNA Synthesis kit (APeXBIO Technology LLC).
RT-qPCR was performed using the SYBR-Green FAST Mastermix (Qiagen
GmbH). The following thermocycling conditions were used for the
qPCR: Initial denaturation at 95˚C for 3 min, followed by 40 cycles
at 95˚C for 15 sec (denaturation), 60˚C for 30 sec (annealing),
72˚C for 1 min (elongation) and a final extension at 72˚C for 5
min. GADPH and U6 were used as internal references. The relative
expression levels were normalized to endogenous control and were
expressed as 2-ΔΔCq (35). The respective sequences of primers
were as follows: MIAT forward, 5'-TCTTCATGTCAGAACACGCTTTA-3' and
reverse, 5'-AAGGTCACCCGAGGTCCAA-3'; miR-665 forward,
5'-GCCGAGACCAGGAGGCTGAG-3' and reverse, 5'-CTCAACTGGTGTCGTGGA-3';
LASP1 forward, 5'-GGT GCGGCAAGATCGTGTA-3' and reverse, 5'-TGCAGGTCT
CGCAATGGAA-3'; GAPDH forward, 5'-CCAGGTGGTCTC CTCTGA-3' and
reverse, 5'-GCTGTAGCCAAATCGTTGT-3'; and U6 forward,
5'-CTCGCTTCGGCAGCACA-3' and reverse,
5'-AACGCTTCACGAATTTGCGT-3'.
MTT assay
Cells were cultured in 96-well plates at a density
of 5x104 cells/ml for 96 h, followed by addition of 15
µl MTT (Procell Life Science & Technology, Co., Ltd.) and
incubation for 2 h at 37˚C. Subsequently, 100 µl DMSO was added to
dissolve the formazan. The optical density was measured at 570 nm
using a microplate reader (Thermo Fisher Scientific, Inc.).
Migration and invasion assays
For the migration assay, transfected Y79 and
HXO-RB44 cells (5x104 cells/ml) were re-suspended in
serum-free DMEM and seeded in the upper chamber of a Transwell
insert (8 µm pore size; BD Biosciences). Simultaneously, DMEM
containing 10% FBS was added in the lower chamber. For the invasion
assay, Matrigel matrix (Becton-Dickinson and Company) was used to
coat the membranes before cell seeding. Following overnight
incubation at 37˚C, cells in the lower chamber were fixed with 4%
paraformaldehyde at 37˚C for 1 h and stained with 0.1% crystal
violet for 15 min at 37˚C. The stained cells were imaged using an
inverted light microscope (Olympus Corporation) and analysed with
ImageJ software (version 1.46; National Institutes of Health).
Dual luciferase reporter (DLR)
assay
The targeting association between MIAT and miR-665
was analyzed using the StarBase software (version 2.0; http://starbase.sysu.edu.cn). Additionally, the
targeting interaction between miR-665 and LASP1 was predicted using
TargetScan software (v7.2; http://www.targetscan.org/vert_72/). The predicted
binding region sequences (MIAT, 5'-GGGUUCCAGGCUCCUGG-3'; LASP1,
5'-CUCCUGGU-3') were inserted into pGL3 vector (Promega
Corporation) to construct the wild-type (wt) phenotype. For
construction of the mutant (mut) phenotype, the mutation sequences
(MIAT mut, 5'-CCCUUCGUCGGAGGACC-3'; LASP1 mut, 5'-GAGGACCA-3') were
inserted into pGL3. Y79 and HXO-RB44 cells were then co-transfected
with MIAT/LASP1-wt or MIAT/LASP1-mut (80 ng) and miR-665
mimics/miR-NC (50 nM) using Lipofectamine 3000 (Invitrogen; Thermo
Fisher Scientific, Inc.) at 37˚C for 48 h, followed by detection of
the luciferase activity using a dual-luciferase reporter assay
system (Promega Corporation). The activity of firefly luciferase
was normalized to that of Renilla luciferase.
Western blotting
Transfected cells were lysed on ice with RIPA
(Beyotime Institute of Biotechnology) containing 10 mmol/l PMSF
(Beyotime Institute of Biotechnology). The protein concentration
was detected using a BCA Protein Assay Kit (Abcam). A total of 50
µg of protein/lane was separated via 10% SDS-PAGE and transferred
onto PVDF membranes. Following blocking with 5% skimmed milk for 2
h at 25˚C, membranes were incubated with the primary antibodies
against LASP1 (1:1,000; cat. no. SAB1402251; Sigma-Aldrich; Merck
KGaA) and α-tubulin (1:1,000; cat. no. T6199; Sigma-Aldrich; Merck
KGaA) at 4˚C overnight. Membranes were then incubated with the
HRP-conjugated anti-mouse IgG secondary antibody (1:5,000; cat. no.
sc-2005; Santa Cruz Biotechnology, Inc.) for 1 h at 37˚C. Bands
were detected using enhanced chemiluminescence substrate (Amersham;
Cytiva). Relative expression level of LASP1 was normalized to
endogenous control α-tubulin. The immunoblots were visualized using
an ECL detection kit (Thermo Fisher Scientific, Inc.) using Gel-Pro
analyzer (version 4.0; Media Cybernetics, Inc.).
Statistical analysis
The SPSS 20.0 software (IBM Corp.) was used to
perform statistical analysis. All cell experiments were performed
three times. Data are presented as the mean ± standard deviation.
Student's t-test was used to assess the differences between two
groups. One-way ANOVA followed by Tukey's post hoc test was used to
evaluate the differences among multiple groups. χ2 test
was used to analyse data from Table
I. The linear correlation was assessed by Pearson's correlation
analysis. P<0.05 was considered to indicate a statistically
significant difference.
 | Table IAssociation between MIAT expression
and the clinicopathological characteristics of patients with
retinoblastoma. |
Table I
Association between MIAT expression
and the clinicopathological characteristics of patients with
retinoblastoma.
| | MIAT expression,
n | |
|---|
|
Characteristics | Total, n | Low (n=23) | High (n=24) | P-value |
|---|
| Age, years | | | | 0.192 |
|
<5 | 20 | 12 | 8 | |
|
≥5 | 27 | 11 | 16 | |
| Sex | | | | 0.891 |
|
Male | 22 | 11 | 11 | |
|
Female | 25 | 12 | 13 | |
| Tumor size, mm | | | | 0.440 |
|
<10 | 28 | 15 | 13 | |
|
≥10 | 19 | 8 | 11 | |
| Optic nerve
invasion | | | | 0.044 |
|
No | 30 | 18 | 12 | |
|
Yes | 17 | 5 | 12 | |
| TNM stage | | | | 0.012 |
|
I/II | 18 | 13 | 5 | |
|
III/IV | 29 | 10 | 19 | |
| IIRC stage | | | | 0.0035 |
|
Early stages
(A-C) | 15 | 12 | 3 | |
|
Advanced
stages (D and E) | 32 | 11 | 21 | |
Results
High MIAT expression is observed in Rb
tissues and cells
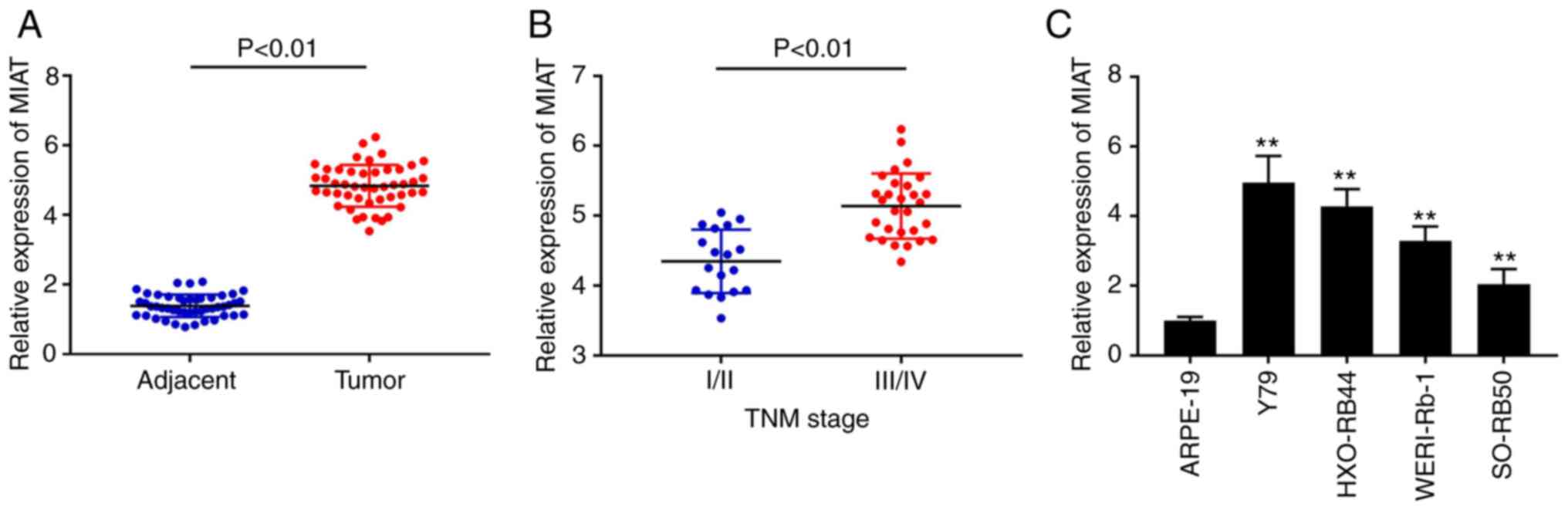
The expression of MIAT was initially detected in Rb
tissues using RT-qPCR. The results demonstrated that MIAT
expression was upregulated in Rb tissues compared with that in
adjacent tissues (Fig. 1A;
P<0.01). Furthermore, a significantly increased expression level
of MIAT was observed in the tissues from patients with
Tumour-Node-Metastasis (TNM) stage III/IV compared with that in the
tissues from patients with TNM stage I/II (Fig. 1B; P<0.01). As shown in Table I, MIAT expression was significantly
associated with intraocular international retinoblastoma
classification (IIRC) stage (P=0.0035), TNM stage (P=0.012) and
optic nerve invasion (P=0.044). In addition, MIAT mRNA expression
was detected in the Rb WERI-Rb-1, SO-RB50, HXO-RB44 and Y79 cell
lines, and the normal ARPE-19 cell line. The results from RT-qPCR
demonstrated that MIAT expression level was significantly increased
in all Rb cell lines compared with that in the ARPE-19 cell line
(Fig. 1C; P<0.01). Since the Y79
and HXO-RB44 cell lines exhibited the highest MIAT expression
levels amongst all Rb cell lines, they were selected for subsequent
experiments.
MIAT knockdown inhibits the
proliferation and migratory and invasive abilities of Rb cells in
vitro
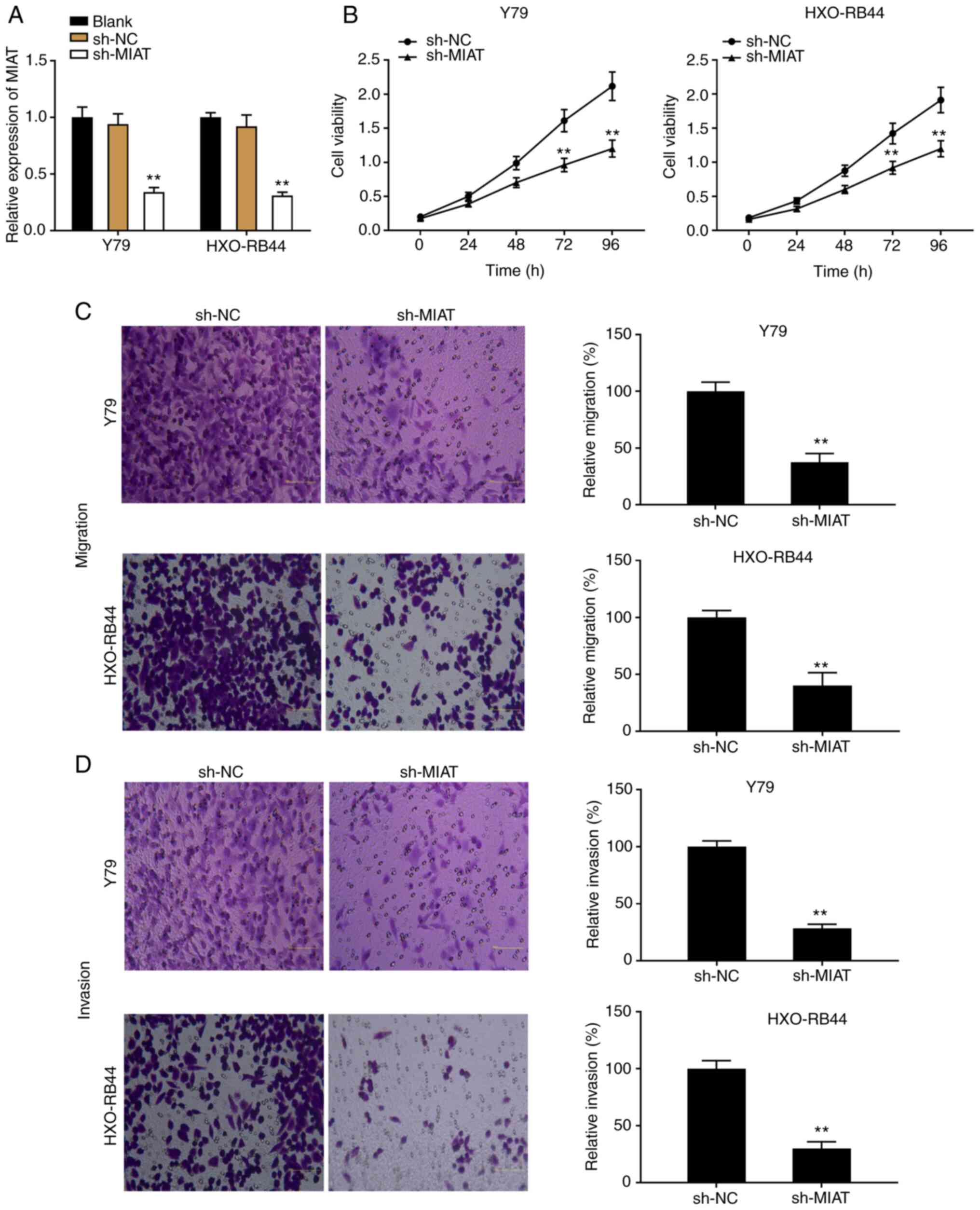
To explore the effects of MIAT on Rb progression
in vitro, Y79 and HXO-RB44 cells were transfected with
sh-MIAT or sh-NC and the transfection efficiency was detected by
RT-qPCR. The results demonstrated that the expression level of MIAT
was significantly decreased following transfection with sh-MIAT
compared with sh-NC, which confirmed the successful transfection
into Y79 and HXO-RB44 cells (Fig.
2A; P<0.01). Subsequently, transfected Y79 and HXO-RB44
cells were used to determine the proliferation and migratory and
invasive abilities of Rb cells using MTT assay and Transwell assay.
As presented in Fig. 2B, Y79 and
HXO-RB44 cell proliferation was significantly decreased following
transfection with sh-MIAT compared with that in the sh-NC group
(P<0.01). Furthermore, both migratory and invasive abilities of
Rb cell lines were decreased in the sh-MIAT group compared with
those in the sh-NC group (Fig. 2C
and D; P<0.01).
MIAT acts as an endogenous sponge of
miR-665
lncRNAs can act as competing endogenous RNAs or
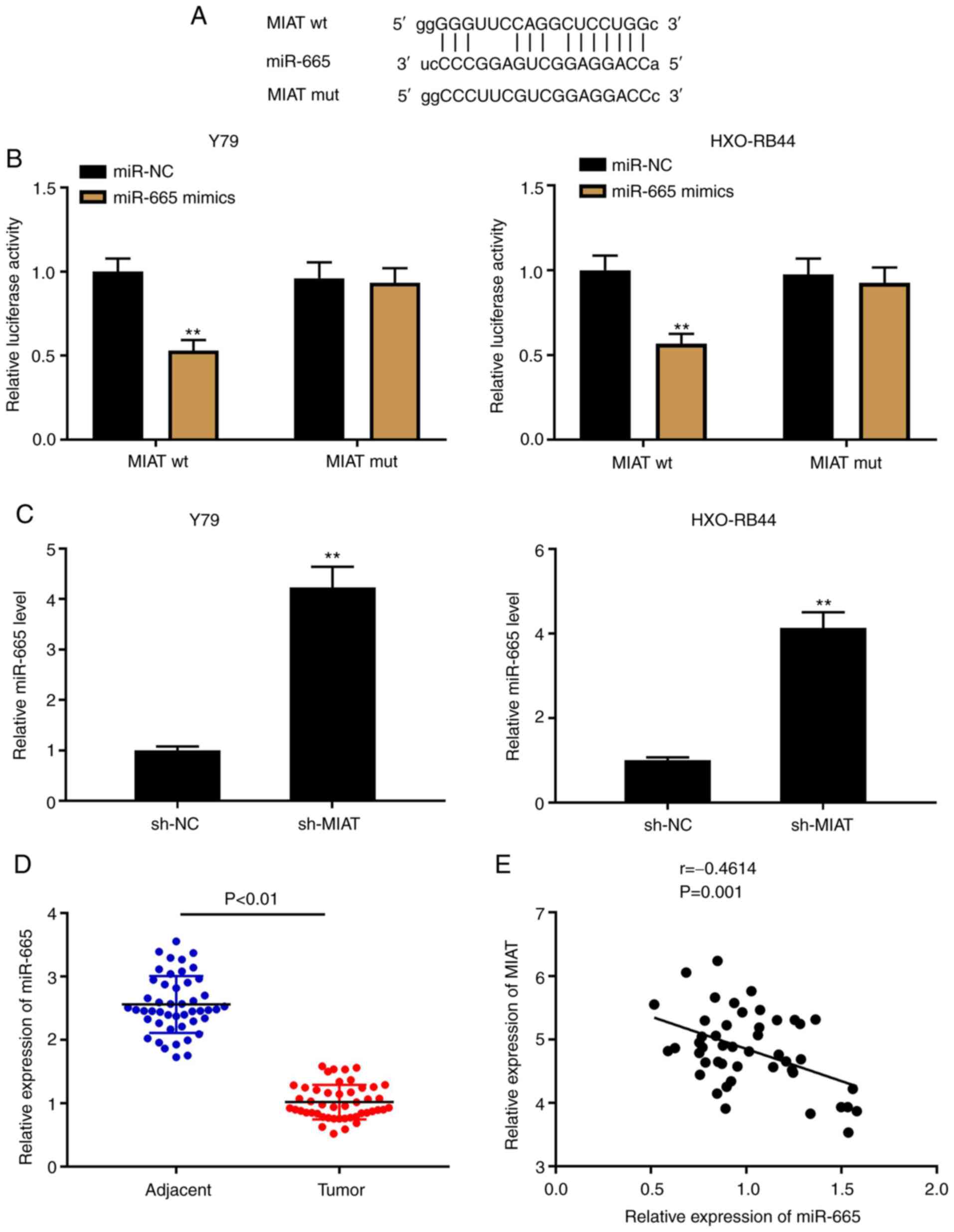
sponges of miRNAs to modulate cancer progression (36). Through the StarBase software, 58
target miRNAs of MIAT were predicted (data not shown). In the
present study, miR-665 was selected due to its important role in Rb
(23,24) (Fig.
3A). A DLR assay was used to confirm the target relationship
between MIAT and miR-665 in Y79 and HXO-RB44 cells. The results
demonstrated that the luciferase activity in MIAT wt/miR-665 mimics
was decreased, which was not the case in the MIAT mut (Fig. 3B; P<0.01). The results from
RT-qPCR indicated that the expression levels of MIAT and miR-665 in
Rb cell lines (Fig. 3C; P<0.01)
and Rb tissues (Fig. 3E; P=0.001,
r=-0.4614) were negatively correlated. In addition, a decreased
expression level of miR-665 was detected in Rb tissues compared
with that in the adjacent tissues (Fig.
3D; P<0.01).
High expression of miR-665 serves as a
suppressor of the proliferation and migratory and invasive
abilities of Rb cells in vitro
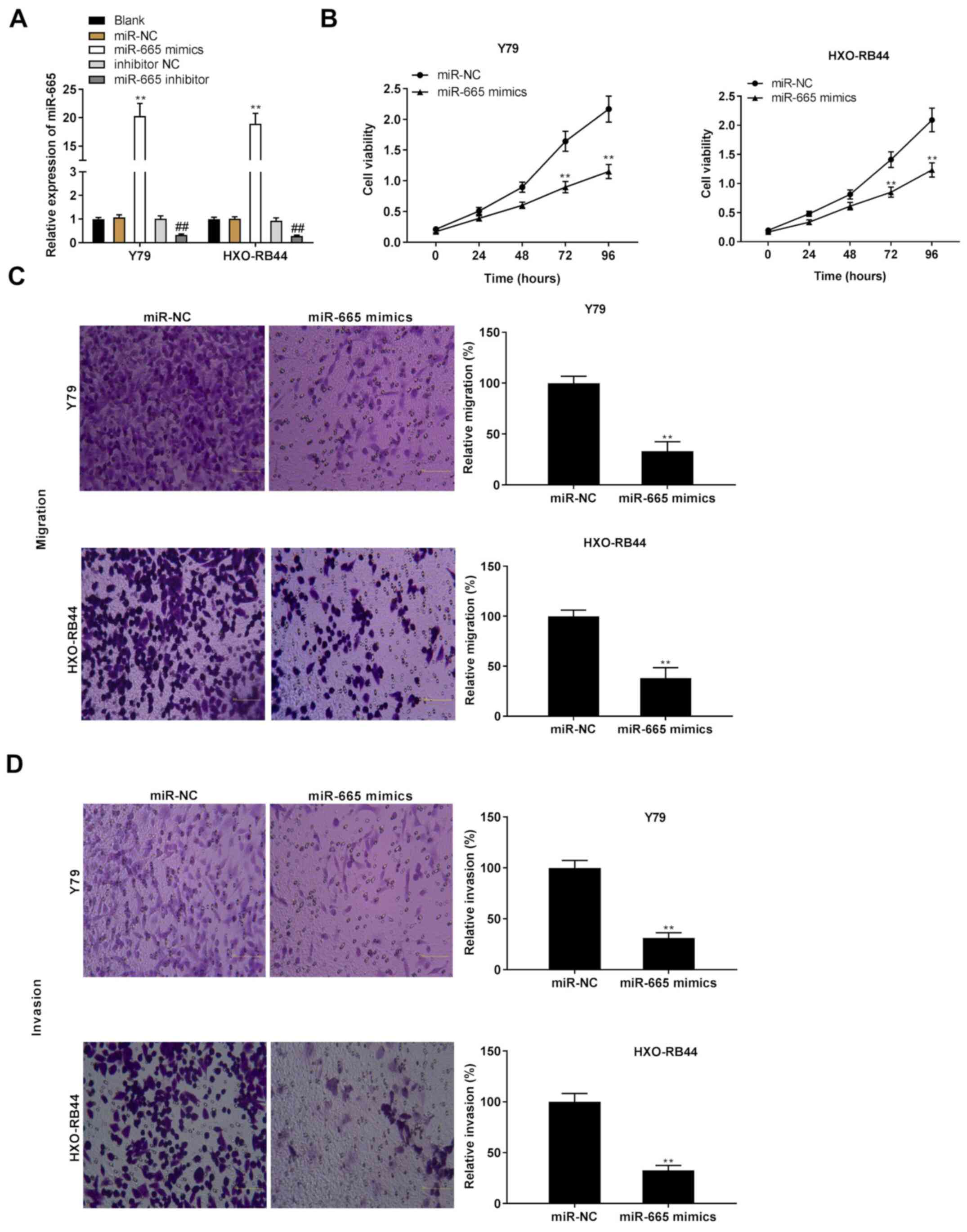
Both miR-665 mimics and miR-665 inhibitor were
transfected into Y79 and HXO-RB44 cells, and the effects of miR-665
overexpression on Rb progression in vitro were subsequently
evaluated. The results demonstrated that miR-665 was significantly
upregulated following transfection with miR-665 mimics, whereas it
was significantly downregulated following transfection with miR-665
inhibitor (Fig. 4A; P<0.01).
Similar to the effects of MIAT knockdown on the progression of Rb
in vitro, the results from the MTT assay and Transwell assay
revealed that the proliferation and migratory and invasive
abilities of Rb cells were inhibited following miR-665
overexpression (Fig. 4B-D;
P<0.01).
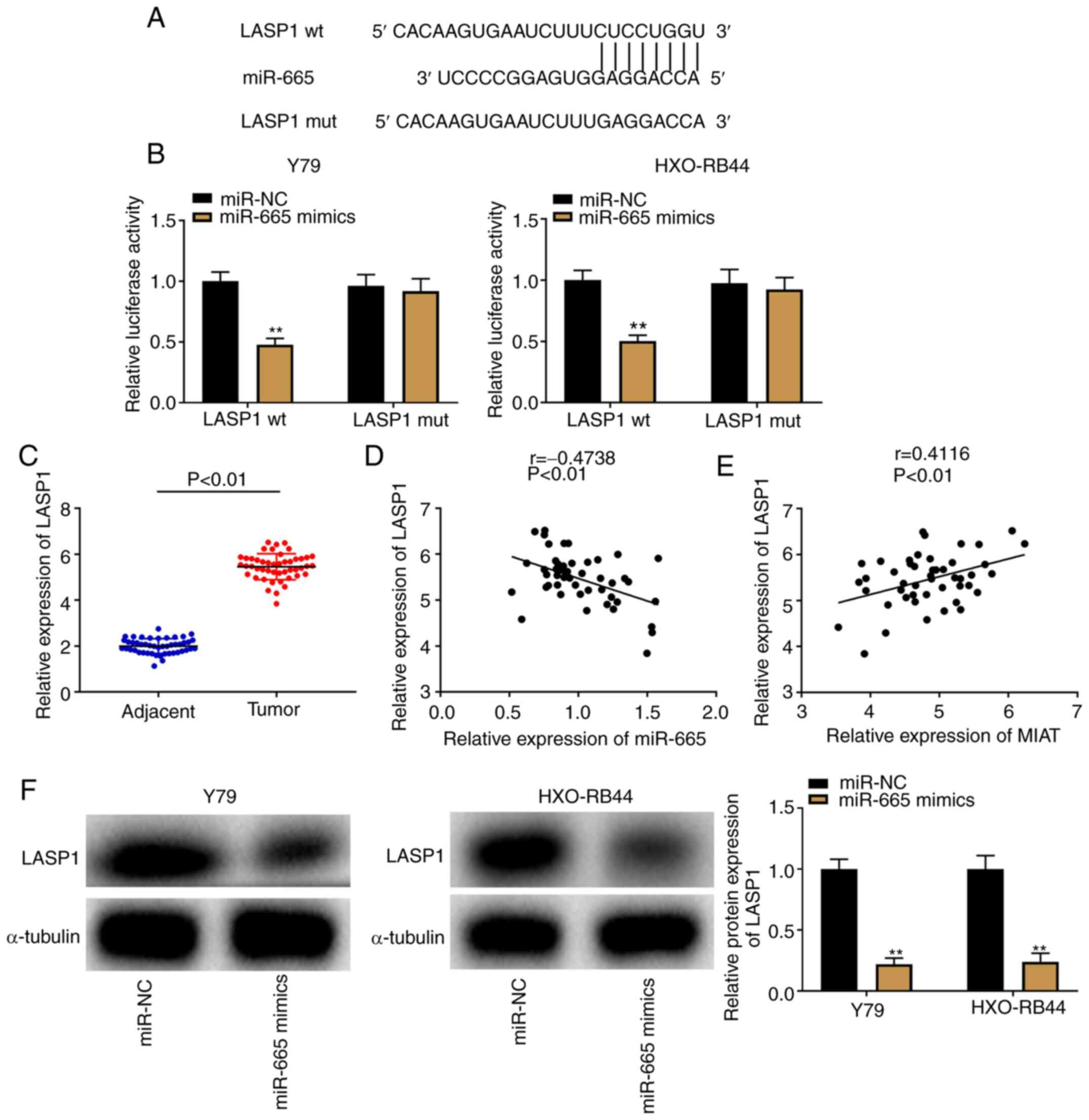
LASP1 is a downstream target gene of
miR-665
As illustrated in Fig.
5A, the TargetScan software predicted a binding site between
miR-665 and LASP1. A DLR assay was subsequently used to confirm the
binding relationship, and a decrease in luciferase activity in the
LASP1 wt/miR-665 mimics groups was observed in both Y79 and
HXO-RB44 cells (Fig. 5B;
P<0.01). Furthermore, LASP1 was overexpressed in Rb tissues
compared with that in adjacent tissues (Fig. 5C; P<0.01). In addition, the
results from Pearson's correlation analysis demonstrated a negative
correlation between LASP1 and miR-665 expression (Fig. 5D; P<0.01, r=-0.4738); however, a
positive correlation was observed between LASP1 and MIAT expression
(Fig. 5E; P<0.01, r=0.4116) in
Rb tissues. To further verify the interaction between the
expression of miR-665 and LASP1, western blotting was performed to
determine the protein expression of LASP1 following transfection of
Y79 and HXO-RB44 cells with miR-665 mimics. The results revealed
that LASP1 protein expression was decreased following miR-665
upregulation (Fig. 5F;
P<0.01).
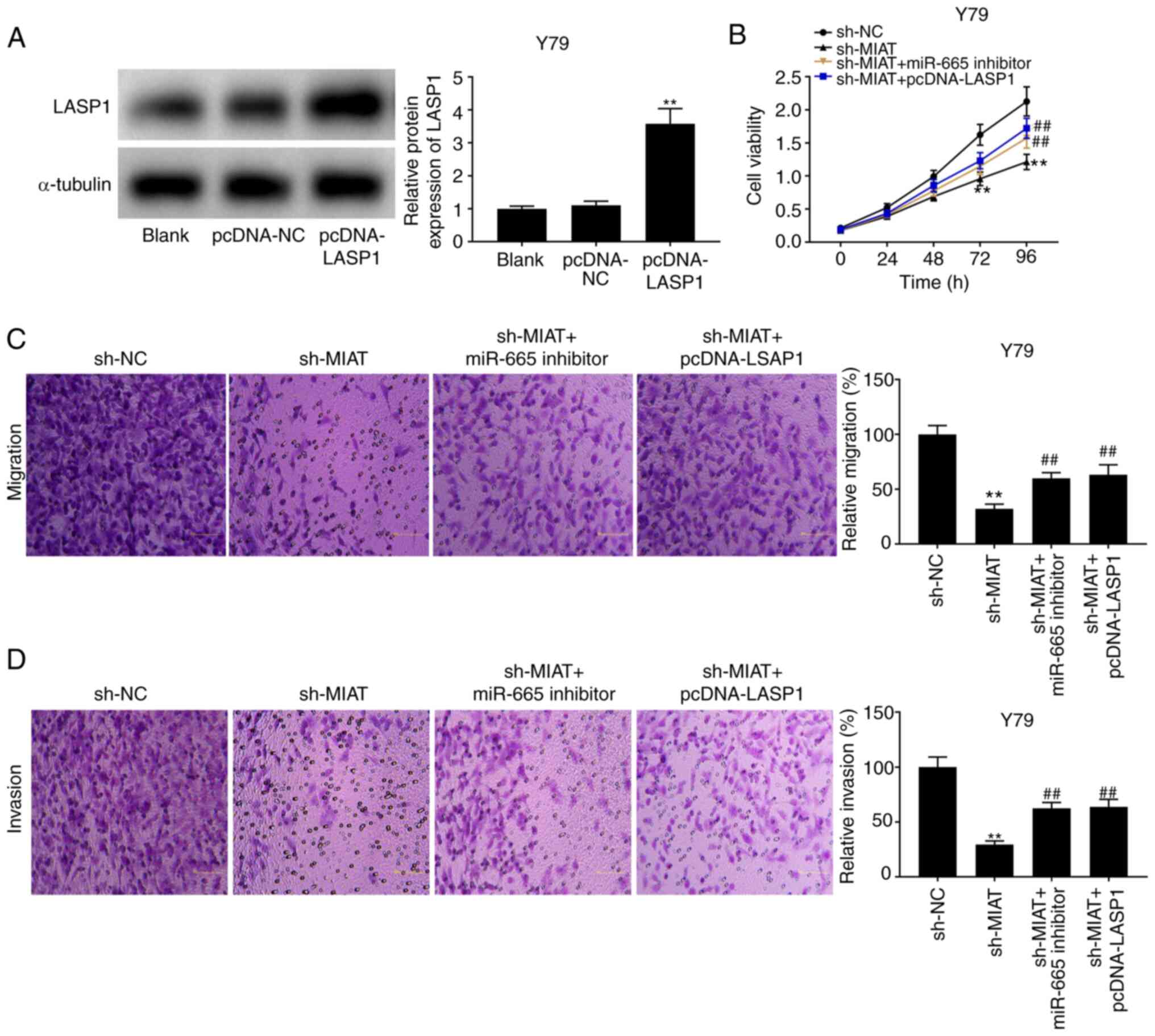
MIAT knockdown decreases the
development of Rb by sponging miR-665 and regulating LASP1
The results from western blotting revealed that
LASP1 expression was increased following transfection with
pcDNA-LASP1 (Fig. 6A; P<0.01).
The results from MTT assay and Transwell assay demonstrated that
the suppressive effects of sh-MIAT on the proliferation and
migratory and invasive abilities of Y79 and HXO-RB44 cells
(Fig. 2) were reversed following
transfection with miR-665 inhibitor or pcDNA-LASP1 (Fig. 6B-D; P<0.01).
Discussion
Chemotherapy and radiation are not the optimal
therapy strategies for Rb as most of the patients with Rb are
children or infants and these therapies can cause serious physical
injury (8). Determining a novel
therapeutic target for Rb is therefore essential. Growing evidence
has demonstrated that certain lncRNAs are upregulated in Rb and are
associated with some pathological features of Rb, such as the
association between XIST and TNM stage (34), the association of AFAP1-AS1 with
choroidal invasion and optic nerve invasion (37), the association of SNHG16 with
choroidal invasion, optic nerve invasion and TNM stage (8), and the association of PVT1 with optic
nerve invasion and IIRC stage (9).
The present study demonstrated that MIAT was overexpressed in Rb
tissues and cell lines. Furthermore, MIAT expression was
significantly associated with IIRC stage, TNM stage and optic nerve
invasion. These findings suggested that MIAT may be considered as a
risk factor in Rb and therefore a potential target for treating
Rb.
To further explore the possible effect of MIAT on
the pathogenesis of Rb, sh-MIAT was transfected into Y79 and
HXO-RB44 cells. The results demonstrated that the proliferation and
migratory and invasive abilities of Rb cells were suppressed
following MIAT knockdown. Similarly, MIAT has been reported to slow
tumorigenesis in several types of human cancer, such as CCA, NSCLC
and CC (11-14).
Chang et al (11)
demonstrated that MIAT silencing can inhibit the proliferation and
accelerate the apoptosis of CCA cells. Similar to the findings
reported by Zhou et al (12), Li et al (13) reported that transfection with
sh-MIAT inhibits the proliferation, migratory and invasive
abilities of NSCLC cells. Similar results were reported in a study
by Zhang et al (14), which
revealed that MIAT downregulation has an inhibitory effect on the
proliferation and migration of CC cells in vitro (14). The present study therefore
hypothesized that MIAT silencing may inhibit the progression of
Rb.
Increasing research efforts have been engaged to
elucidate the anti-tumour roles of miR-665 in human cancer
(38-40).
For example, miR-665 downregulation was reported to reverse the
suppressive effects of the lncRNA RHPN1-AS1 on the proliferation
and migratory and invasive abilities of OC cells (38). Furthermore, transfection with
miR-665 inhibitor partly aggravates the tumorigenicity of CRC in
vitro (39). A recent study by
Wu et al (40) revealed that
miR-665 expression is downregulated in GC tissues, and that miR-665
overexpression exhibits a visible effect on tumour inhibition. In
the present study, a decreased expression of miR-665 was found in
Rb tissues, and miR-665 overexpression significantly reduced the
proliferation and migratory and invasive abilities of Rb cells.
Similarly, Wang et al (23)
reported that miR-665 is minimally expressed in Rb tissues and cell
lines, and that the proliferation and migratory and invasive
abilities of Rb cells are inhibited following transfection with
miR-665 mimics. In the present study, MIAT was shown to target and
negatively modulate miR-665 expression, suggesting that miR-665 may
be involved in Rb tumorigenesis via MIAT regulation. The results of
the present study further demonstrated that the decreased
expression of miR-665 reversed the inhibitory effects of MIAT
silencing on the proliferation and migratory and invasive abilities
of Y79 cells. These findings indicated that MIAT knockdown may
attenuate Rb malignancy by modulating miR-665.
LASP1, an oncogene involved in cancer aggressiveness
(41), was found to be upregulated
in numerous types of cancer, including gallbladder cancer (42), NSCLC (43), CRC (44) and OC (45). In the present study, LASP1 was
demonstrated to be highly expressed in Rb tissues compared with
that in adjacent tissues. Consistent with these findings, Yang
et al (8) reported a high
expression level of LASP1 in Rb tissues. These results suggested
that LASP1 may act as an oncogene in Rb pathogenesis. Furthermore,
a positive correlation between MIAT and LASP1 expression was
observed in the present study. Similarly, Liu et al
(46) confirmed that the expression
of LASP1 is positively correlated with MIAT expression in papillary
thyroid cancer tissues. Furthermore, the present study demonstrated
that LASP1 may be a target gene of miR-665 and was negatively
regulated by miR-665. These findings suggested that miR-665 may be
involved in the tumorigenesis of Rb via MIAT regulation. We
hypothesized that LASP1 may be modulated by MIAT and be involved in
Rb progression. The feedback verification experiments demonstrated
that the enhancement of proliferation and migratory and invasive
abilities of Y79 cells caused by MIAT silencing were restrained
following transfection with pcDNA-LASP1, which confirmed this
hypothesis. These findings indicated that MIAT downregulation may
reduce the progression of Rb by sponging miR-665 and regulating
LASP1.
In summary, the present study reported an elevated
expression level of MIAT in Rb tissues and demonstrated that MIAT
silencing significantly reduced the tumorigenicity of Rb by
modulating the miR-665/LASP1 axis in vitro. The results from
this study provide a novel target for treating Rb and may be
applied in the clinical setting. One limitation of this study is
the absence of in vivo experiments to confirm the
interactions among MIAT, miR-665 and LASP1. Further investigation
will therefore be performed in the future.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
XX and BD made significant contributions to the
overall structure design of the study, data analysis, methodology,
project management, funding and draft writing. YZ and GD are mainly
responsible for resource collection and integration, experiments,
experimental data analysis, software processing and paper
modification and editing. All authors confirmed the authenticity of
all the raw data and have read and approved the final
manuscript.
Ethics approval and consent to
participate
This study was performed in line with the principles
of the Declaration of Helsinki. Approval was granted by the Ethics
Committee of No. 215 Hospital of Shaanxi Nuclear Industry (approval
no. EC-20200924-1017). Each patient or their parents provided
written informed consent.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Pascual-Pasto G, Bazan-Peregrino M,
Olaciregui NG, Restrepo-Perdomo CA, Mato-Berciano A, Ottaviani D,
Weber K, Correa G, Paco S, Vila-Ubach M, et al: Therapeutic
targeting of the RB1 pathway in retinoblastoma with the oncolytic
adenovirus VCN-01. Sci Transl Med. 11(11)2019.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Chantada GL, Qaddoumi I, Canturk S, Khetan
V, Ma Z, Kimani K, Yeniad B, Sultan I, Sitorus RS, Tacyildiz N, et
al: Strategies to manage retinoblastoma in developing countries.
Pediatr Blood Cancer. 56:341–348. 2011.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Canturk S, Qaddoumi I, Khetan V, Ma Z,
Furmanchuk A, Antoneli CB, Sultan I, Kebudi R, Sharma T,
Rodriguez-Galindo C, et al: Survival of retinoblastoma in
less-developed countries impact of socioeconomic and health-related
indicators. Br J Ophthalmol. 94:1432–1436. 2010.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Errico A: Cancer therapy: Retinoblastoma -
chemotherapy increases the risk of secondary cancer. Nat Rev Clin
Oncol. 11(623)2014.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Shields CL, Say EA, Pointdujour-Lim R, Cao
C, Jabbour PM and Shields JA: Rescue intra-arterial chemotherapy
following retinoblastoma recurrence after initial intra-arterial
chemotherapy. J Fr Ophtalmol. 38:542–549. 2015.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Rao AKDM, Rajkumar T and Mani S:
Perspectives of long non-coding RNAs in cancer. Mol Biol Rep.
44:203–218. 2017.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Li S, Wen D, Che S, Cui Z, Sun Y, Ren H
and Hao J: Knockdown of long noncoding RNA 00152 (LINC00152)
inhibits human retinoblastoma progression. OncoTargets Ther.
11:3215–3223. 2018.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Yang L, Zhang L, Lu L and Wang Y: Long
noncoding RNA SNHG16 sponges miR-182-5p and miR-128-3p to promote
retinoblastoma cell migration and invasion by targeting LASP1.
OncoTargets Ther. 12:8653–8662. 2019.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Wu XZ, Cui HP, Lv HJ and Feng L: Knockdown
of lncRNA PVT1 inhibits retinoblastoma progression by sponging
miR-488-3p. Biomed Pharmacother. 112(108627)2019.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Wang L, Wang C, Wu T and Sun F: Long
non-coding RNA TP73-AS1 promotes TFAP2B-mediated proliferation,
metastasis and invasion in retinoblastoma via decoying of
miRNA-874-3p. J Cell Commun Signal. 14:193–205. 2020.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Chang W, Wang Y, Li W and Geng Z: Long
non-coding RNA myocardial infarction associated transcript promotes
the proliferation of cholangiocarcinoma cells by targeting
miR-551b-3p/CCND1 axis. Clin Exp Pharmacol Physiol. 47:1067–1075.
2020.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Zhou Z, Zhang S and Xiong Y: Long
noncoding RNA MIAT promotes non-small cell lung cancer progression
by sponging miR-149-5p and regulating FOXM1 expression. Cancer Cell
Int. 20(348)2020.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Li F, Li H, Li S, Lv B, Shi J, Yan H,
Zhang H and He Y: Long Non-coding RNA MIAT mediates non-small cell
lung cancer development through regulating the miR-128-3p/PELI3
Axis. Biochem Genet. 58:867–882. 2020.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Zhang L, Ge S and Cao B: Long non-coding
RNA MIAT promotes cervical cancer proliferation and migration. J
Biochem. 168:183–190. 2020.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Zhou S, Xu A, Song T, Gao F, Sun H and
Kong X: lncRNA MIAT regulates cell growth, migration, and invasion
through sponging miR-150-5p in ovarian cancer. Cancer Biother
Radiopharm. 35:650–660. 2020.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Liu Y, Peng H, Shen Y, Da R, Tian A and
Guo X: Downregulation of long noncoding RNA myocardial infarction
associated transcript suppresses cell proliferation, migration,
invasion, and glycolysis by regulation of miR-488-3p/IGF1R pathway
in colorectal cancer. Cancer Biother Radiopharm: Oct 21, 2020 (Epub
ahead of print). doi: 10.1089/cbr.2020.3671.
|
|
17
|
Sha M, Lin M, Wang J, Ye J, Xu J, Xu N and
Huang J: Long non-coding RNA MIAT promotes gastric cancer growth
and metastasis through regulation of miR-141/DDX5 pathway. J Exp
Clin Cancer Res. 37(58)2018.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Lu TX and Rothenberg ME: MicroRNA. J
Allergy Clin Immunol. 141:1202–1207. 2018.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Liu Y, Li H, Liu Y and Zhu Z: MiR-22-3p
targeting alpha-enolase 1 regulates the proliferation of
retinoblastoma cells. Biomed Pharmacother. 105:805–812.
2018.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Wang L, Yang D, Tian R and Zhang H: NEAT1
promotes retinoblastoma progression via modulating miR-124. J Cell
Biochem. 120:15585–15593. 2019.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Han N, Zuo L, Chen H, Zhang C, He P and
Yan H: Long non-coding RNA homeobox A11 antisense RNA (HOXA11-AS)
promotes retinoblastoma progression via sponging miR-506-3p.
OncoTargets Ther. 12:3509–3517. 2019.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Xu L, Li W, Shi Q, Wang M, Li H, Yang X
and Zhang J: MicroRNA 936 inhibits the malignant phenotype of
retinoblastoma by directly targeting HDAC9 and deactivating the
PI3K/AKT pathway. Oncol Rep. 43:635–645. 2020.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Wang S, Du S, Lv Y, Zhang F and Wang W:
MicroRNA-665 inhibits the oncogenicity of retinoblastoma by
directly targeting high-mobility group box 1 and inactivating the
Wnt/β-catenin pathway. Cancer Manag Res. 11:3111–3123.
2019.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Zhang S, Long J and Hu Y: Long noncoding
RNA LINC00205 enhances the malignant characteristics of
retinoblastoma by acting as a molecular sponge of microRNA-665 and
consequently increasing HMGB1 expression. Biochem Biophys Res
Commun. 526:396–403. 2020.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Hu S, Ran Y, Chen W, Zhang Y and Xu Y:
MicroRNA-326 inhibits cell proliferation and invasion, activating
apoptosis in hepatocellular carcinoma by directly targeting LIM and
SH3 protein 1. Oncol Rep. 38:1569–1578. 2017.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Yan P, Liu J, Zhou R, Lin C, Wu K, Yang S,
Yang S, Zhou J, Xu L, Wang H, et al: LASP1 interacts with N-WASP to
activate the Arp2/3 complex and facilitate colorectal cancer
metastasis by increasing tumour budding and worsening the pattern
of invasion. Oncogene. 39:5743–5755. 2020.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Wang S, Qiu J, Wang L, Wu Z, Zhang X, Li Q
and Jiang F: Long non-coding RNA LINC01207 promotes prostate cancer
progression by downregulating microRNA-1972 and upregulating LIM
and SH3 protein 1. IUBMB Life. 72:1960–1975. 2020.PubMed/NCBI View
Article : Google Scholar
|
|
28
|
Sui Y, Zhang X, Yang H, Wei W and Wang M:
MicroRNA-133a acts as a tumour suppressor in breast cancer through
targeting LASP1. Oncol Rep. 39:473–482. 2018.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Song X, Jin Y, Yan M, Zhang Y and Chen B:
MicroRNA-342-3p functions as a tumor suppressor by targeting LIM
and SH3 protein 1 in oral squamous cell carcinoma. Oncol Lett.
17:688–696. 2019.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Liu W, Wang ZL, Wang C and Ai ZL: Long
non-coding RNA MIAT promotes papillary thyroid cancer progression
through upregulating LASP1. Cancer Cell Int. 19(194)2019.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Wang JX, Yang Y and Li K: Long noncoding
RNA DANCR aggravates retinoblastoma through miR-34c and miR-613 by
targeting MMP-9. J Cell Physiol. 233:6986–6995. 2018.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Shang W, Yang Y, Zhang J and Wu Q: Long
noncoding RNA BDNF-AS is a potential biomarker and regulates cancer
development in human retinoblastoma. Biochem Biophys Res Commun.
497:1142–1148. 2018.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Yan G, Su Y, Ma Z, Yu L and Chen N: Long
Noncoding RNA LINC00202 promotes tumor progression by sponging
miR-3619-5p in retinoblastoma. Cell Struct Funct. 44:51–60.
2019.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Hu C, Liu S, Han M, Wang Y and Xu C:
Knockdown of lncRNA XIST inhibits retinoblastoma progression by
modulating the miR-124/STAT3 axis. Biomed Pharmacother.
107:547–554. 2018.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) Method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Du Z, Sun T, Hacisuleyman E, Fei T, Wang
X, Brown M, Rinn JL, Lee MG, Chen Y, Kantoff PW, et al: Integrative
analyses reveal a long noncoding RNA-mediated sponge regulatory
network in prostate cancer. Nat Commun. 7(10982)2016.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Hao F, Mou Y, Zhang L, Wang S and Yang Y:
LncRNA AFAP1-AS1 is a prognostic biomarker and serves as oncogenic
role in retinoblastoma. Biosci Rep. 38(38)2018.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Zhao J, Yang T, Ji J, Zhao F, Li C and Han
X: RHPN1-AS1 promotes cell proliferation and migration via
miR-665/Akt3 in ovarian cancer. Cancer Gene Ther. 28:33–41.
2021.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Ouyang S, Zhou X, Chen Z, Wang M, Zheng X
and Xie M: LncRNA BCAR4, targeting to miR-665/STAT3 signaling,
maintains cancer stem cells stemness and promotes tumorigenicity in
colorectal cancer. Cancer Cell Int. 19(72)2019.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Wu KZ, Zhang CD, Zhang C, Pei JP and Dai
DQ: miR-665 suppresses the epithelial-mesenchymal transition and
progression of gastric cancer by targeting CRIM1. Cancer Manag Res.
12:3489–3501. 2020.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Orth MF, Cazes A, Butt E and Grunewald TG:
An update on the LIM and SH3 domain protein 1 (LASP1): A versatile
structural, signaling, and biomarker protein. Oncotarget. 6:26–42.
2015.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Li Z, Chen Y, Wang X, Zhang H, Zhang Y,
Gao Y, Weng M, Wang L, Liang H, Li M, et al: LASP-1 induces
proliferation, metastasis and cell cycle arrest at the G2/M phase
in gallbladder cancer by down-regulating S100P via the PI3K/AKT
pathway. Cancer Lett. 372:239–250. 2016.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Zheng J, Wang F, Lu S and Wang X: LASP-1,
regulated by miR-203, promotes tumor proliferation and
aggressiveness in human non-small cell lung cancer. Exp Mol Pathol.
100:116–124. 2016.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Zhao L, Wang H, Liu C, Liu Y, Wang X, Wang
S, Sun X, Li J, Deng Y, Jiang Y, et al: Promotion of colorectal
cancer growth and metastasis by the LIM and SH3 domain protein 1.
Gut. 59:1226–1235. 2010.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Grunewald TG, Kammerer U, Winkler C,
Schindler D, Sickmann A, Honig A and Butt E: Overexpression of
LASP-1 mediates migration and proliferation of human ovarian cancer
cells and influences zyxin localisation. Br J Cancer. 96:296–305.
2007.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Liu W, Wang Z, Wang C and Ai Z: Long
non-coding RNA MIAT promotes papillary thyroid cancer progression
through upregulating LASP1. Cancer Cell Int. 19(194)2019.PubMed/NCBI View Article : Google Scholar
|