Introduction
Myocardial ischemia/reperfusion (I/R) injury can
occur after various types of cardiac surgery, including open heart
surgery, coronary artery bypass grafting, coronary angioplasty and
embolic surgery, and can result in serious clinical manifestations
in numerous parts of the body, thus limiting the effectiveness of
reperfusion treatment (1).
Copine 3 (CPNE3) has recently been identified as a
risk factor for patients with acute myocardial I/R, as those with
low expression levels of CPNE3 tend to be subject to acute
myocardial infarction (2). However,
at present, there is a lack of research into the mechanism of
action of CPNE3 in myocardial I/R injury.
Receptor for activated C kinase 1 (RACK1) is a type
of multifaceted scaffold protein that mediates activated protein
kinase C translocation through the cytomembrane, and has been shown
to be a participant in the regulation of several cardiovascular
diseases, such as cardiac failure and myocardial infarction
(3). A previous study reported that
CPNE3 can interact with RACK1 in non-small cell lung cancer
(NSCLC), serving an oncogenic role in metastasis (4). Currently, the therapeutic strategies
available for patients with myocardial I/R injury are limited to
the alleviation of its clinical symptoms, such as medical
intervention with mannitol to relieve the limb swelling caused by
cellular edema (5). Therefore,
identifying an effective strategy for the prevention of myocardial
I/R injury is important. The present study aimed to evaluate the
potential protective mechanism underlying the interaction between
CPNE3 and RACK1 in myocardial I/R injury. The results of the
present study may facilitate the development of innovative and
effective strategies for the clinical treatment of myocardial I/R
injury.
Materials and methods
Animals and treatments
Sprague-Dawley rats (n=7 per group; male; age, 2-3
months, weight, 180-220 g) were purchased from the Experimental
Animal Center of Cangzhou Central Hospital. Rats were maintained in
a controlled environment at 25±3˚C with ~30% relative humidity,
12-h light/dark cycles and ad libitum access to food and
water. All animal procedures and experimental methods were approved
by the Committee on the Ethics of Animal Experiments of Cangzhou
Central Hospital. The animal maintenance and experiments complied
with the guidelines drafted by the Animal Ethics Committee of
Cangzhou Central Hospital.
Myocardial I/R was simulated in rats by performing
coronary artery ligation, with myocardial ischemia for 1 h followed
by reperfusion for 2, 4 and 12 h (6). Rats in the sham group were threaded
without ligation. The control group did not receive ligation. After
model establishment, the rats displayed lethargy and hypothermia
without anesthesia. Subsequently, the rats were anesthetized by the
intraperitoneal injection of 0.8% pentobarbital sodium (40 mg/kg)
and then sacrificed by cervical dislocation. Death was verified by
cessation of the heartbeat. Myocardial tissues of the rats were
collected for subsequent experiments.
H&E staining
Myocardial tissue was fixed with 10% formalin for 24
h at room temperature, embedded in paraffin and then and were cut
into 10-µm thick sections, embedded in paraffin and frozen at -80˚C
for storage. Subsequently, the tissue sections were collected onto
microscope slides at room temperature. The slides were incubated
with hematoxylin solution (Shanghai Aladdin Biochemical Technology
Co., Ltd.) for 10 min at 37˚C, washed with running water and then
incubated with eosin (Beyotime Institute of Biotechnology) for 3
min at 37˚C. The myocardial slices were treated with ethanol to
achieve transparency at 37˚C. Images were captured using a light
microscope camera (Keyence Corporation; magnification, x400).
ImageJ software (version 146; National Institutes of Health) was
used for quantitative analysis.
Tetrazolium chloride (TTC)
staining
Cardiac tissue from rats was isolated and rapidly
frozen at -20˚C for ~20 min. After sectioning (thickness, 1 mm),
the tissue slices were incubated in 2% TTC at 37˚C in the dark for
20 min. Subsequently, the tissue sections were fixed in 4%
paraformaldehyde for 24 h. Images were captured using a light
microscope camera (Keyence Corporation, magnification, x400).
Images of the stained tissue sections were obtained and Image-Pro
Plus software (version 6.0; Media Cybernetics, Inc.) was used for
image analysis. Healthy brain tissue was stained red and infarcted
areas were stained white.
Cell culture and treatments
H9c2 cardiomyocytes (EK-Bioscience) were obtained
for the establishment of a cellular model of myocardial I/R injury.
H9c2 cells were cultured in DMEM (Gibco; Thermo Fisher Scientific,
Inc.) supplemented with 10% FBS (Gibco; Thermo Fisher Scientific,
Inc.), 100 U/ml penicillin and 100 mg/ml streptomycin in a
humidified incubator containing 95% air and 5% CO2 at
37˚C. At 80% confluence, 0.05% trypsin was used for cell digestion.
Subsequently, hypoxia was induced in H9c2 cells for 6 h, followed
by reoxygenation for 2, 4 or 12 h. Briefly, after cellular
incubation in glucose-free DMEM at 37˚C for 4 h, H9c2 cells were
placed in a hypoxic incubator containing 1% O2, 94%
N2 and 5% CO2 at 37˚C for 6 h. Subsequently,
glucose was added to the culture media until it normalized, which
occurred after 6 h of hypoxia, but before reoxygenation. Then, the
cells were incubated in a normal environment with 95% air and 5%
CO2 at 37˚C for 2, 4 or 12 h as previously described
(6). H9c2 cells in the control
group were cultured in DMEM supplemented with 10% FBS, 100 U/ml
penicillin and 100 mg/ml streptomycin in a humidified incubator
containing 95% air and 5% CO2 at 37˚C until the end of
the experiment.
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA was extracted from the myocardial tissues
of rats and H9c2 cells using TRIzol® reagent
(Invitrogen; Thermo Fisher Scientific, Inc.), according to the
manufacturer's protocol. Briefly, the tissues and cells were lysed
using 1 ml TRIzol reagent at room temperature for 10 min, followed
by centrifugation at 300 x g at 4˚C for 20 min to obtain the
supernatant. After precipitation using isopropanol, the supernatant
was washed with 75% ethanol (1 ml) at 4˚C for 5 min and the
precipitates were dried. The quality of RNA extraction was
determined using a spectrophotometer. Total RNA was reverse
transcribed into cDNA using a PrimeScript RT reagent kit (Takara
Bio, Inc.) according to the manufacturer's instructions. The
expression levels of CPNE3, RACK1 and inflammatory cytokines were
detected via qPCR using SYBR Premix EX Taq (Takara Bio, Inc.). The
following thermocycling conditions were used for qPCR: 95˚C for 10
min; followed by 40 cycles of 95˚C for 10 sec and 60˚C for 60 sec.
mRNA expression levels were quantified using the 2-ΔΔCq
method and normalized to the internal reference gene GAPDH
(7). The sequences of the forward
and reverse primers used for qPCR: TNF-α forward,
5'-GAAACACACGAGACGCTGAA-3' and reverse, 5'-GAAAGCCCATTGGAATCCTT-3';
IL-6 forward, 5'-TGATGGATGCTTCCAAACTG-3' and reverse,
5'-GAGCATTGGAAGTTGGGGTA-3'; IL-1β forward,
5'-AGCTTCAGGAAGGCAGTGTC-3' and reverse, 5'-TCAGACAGCACGAGGCATTT-3';
CPNE3 forward, 5'-GATGGCGTGATCACAGACCTT-3' and reverse,
5'-GGCTTCCATTGTCACCGTCTA-3'; RACK1 forward,
5'-GCCACCCCAGTGTACCTCTTTG-3' and reverse,
5'-TCACCTGCCATACACGCACCAA-3'; GAPDH forward,
5'-TGGCCTTCCGTGTTCCTACC-3' and reverse,
5'-CGCCTGCTTCACCACCTTCT-3'.
Western blotting
Total protein was extracted from myocardial tissues
and H9c2 cells using RIPA reagent (Protech Technology Enterprise
Co., Ltd.). Protein concentrations were determined using a BCA kit
(Abcam). Subsequently, 20 µg proteins were separated via 10%
SDS-PAGE and transferred to PVDF membranes. Following blocking with
5% non-fat milk for 2 h at room temperature, the membranes were
incubated overnight at 4˚C with primary antibodies targeted
against: CPNE3 (1:1,000; cat. no. ab236606; Abcam), RACK1 (1:1,000;
cat. no. ab129084; Abcam), phosphorylated (p)-NF-κB P65 (1:1,000;
cat. no. ab183559; Abcam), cyclooxygenase 2 (Cox2; 1:1,000; cat.
no. ab179800; Abcam), P65 (1:1,000; cat. no. ab32536; Abcam), Bax
(1:1,000; cat. no. ab32503; Abcam), caspase-3 (1:1,000; cat. no.
ab32351; Abcam), cleaved caspase-3 (1:1,000; cat. no. ab32042;
Abcam), cleaved poly(ADP-ribose) polymerase (PARP; 1:1,000; cat.
no. ab32064; Abcam), PARP (1:1,000; cat. no. ab191217; Abcam),
Bcl-2 (1:1,000; cat. no. ab32124; Abcam) and GAPDH (1:1,000; cat.
no. ab8245; Abcam). After washing with 0.05% TBS-Tween 20 (Shanghai
Aladdin Biochemical Technology Co., Ltd.), the membranes were
incubated with HRP-conjugated anti-mouse IgG (1:5,000; cat. no.
7076S; Cell Signaling Technology, Inc.) secondary antibodies for 2
h at room temperature. Subsequently, the membranes were placed in
the dark for 1 h after the addition of color developing solution.
The signals were detected using an enhanced chemiluminescence
reagent (Cytiva). Protein expression was semi-quantified using
Image Lab software (version 3.0; Bio-Rad Laboratories, Inc.) with
GAPDH as the loading control.
Cell Counting Kit (CCK)-8
H9c2 cells were inoculated (2x103
cells/well) into a 96-well plate. After corresponding treatments in
the different groups of cells, 10 µl sterile CCK-8 solution
(Dojindo Molecular Technologies, Inc.) was added to each well for 2
h. Cell viability was detected at a wavelength of 450 nm using a
microplate reader (Bio-Rad Laboratories, Inc.) according to the
manufacturer's instructions.
Cell transfection
Before H9c2 cells were subjected to H/R treatment,
cells were inoculated (1x106 cells/well) into a six-well
plate and cultured for 12 h at 37˚C. At ~80% confluence, cells were
transfected with 20 nM overexpression (Ov)-CPNE3 plasmid, 20 nM
Ov-negative control (NC, an empty vector) plasmid, 20 nM
RACK1-small interfering (si)RNA or 20 nM non-targeting siRNA-NC
(all purchased from Shanghai GenePharma Co., Ltd.) using
Lipofectamine® 2000 (Invitrogen; Thermo Fisher
Scientific, Inc.) at 37˚C for 48 h according to the manufacturer's
protocol. At 48 h post-transfection, transfection efficiencies were
assessed via RT-qPCR and western blotting. The transfected
sequences were as follows: si-RACK1, 5'-GACATCATCATGTGGAAGC-3'; and
si-NC, 5'-GACCATCATCATGTGGAAGC-3'.
Immunoprecipitation (IP) assay
An immunoprecipitation kit (cat. no. ab206996;
Abcam) was used to detect the interaction between CPNE3 and RACK1.
Cells were harvested using IP lysis buffer and centrifuged at
>13,000 x g for 30 min at 4˚C. The resulting supernatants were
then collected for use. Magnetic beads were dissolved in IP buffer
(Abcam), after which antibodies including CPNE3 (1:1,000; cat. no.
ab236606; Abcam), RACK1 (1:1,000; cat. no. ab129084; Abcam) and
normal rabbit immunoglobulin G (negative control; cat. no.
ab172730; 1:1,000; Abcam) were added to the magnetic beads for
ligation (5 g for each reaction) overnight at 4˚C. The cells were
then mixed with the antigenic antibody complex and incubated with
Protein A/G Sepharose® for 2 h. The antigen-antibody
complex attached to Protein A/G Sepharose was eluted, incubated
with 50 µl elution solution for 5 min and then centrifuged at 1,000
x g for 5 min at 4˚C. The supernatants were collected and mixed.
The pH of the protein samples was immediately adjusted to the
physiological value. Lastly, the eluted samples were desalinized,
which was followed by protein precipitation examination via western
blotting according to the manufacturer's protocol.
Lactate dehydrogenase (LDH) activity
assay
An LDH Cytotoxicity assay kit (cat. no. C0016;
Beyotime Institute of Biotechnology) was used to determine the LDH
level, which is an indicator for cytotoxic release, in H9c2 cells.
Cells were inoculated into a 96-well plate to 80-90% confluence.
After the different treatments in the respective groups, the plate
was centrifuged at 400 x g for 5 min 4˚C. Subsequently, 150 µl
PBS-diluted LDH release reagent was added per well and the plate
was shaken for thorough mixing. Following incubation for 1 h, the
culture plate was centrifuged at 400 x g for 5 min 4˚C. Then, 120
µl supernatant from each well was added to a new 96-well plate,
followed by detection with LDH working solution. The absorbance was
measured at a wavelength of 490 nm using a microplate reader.
TUNEL staining
A colorimetric TUNEL Apoptosis Assay kit (Beyotime
Institute of Biotechnology) was used to observe H9c2 cell
apoptosis. Cells were fixed with Immunol Staining Fix Solution
(Beyotime Institute of Biotechnology) for 30 min at 37˚C, followed
by washing with PBS. Cells were then incubated with PBS containing
0.3% Triton X-100 (Sigma-Aldrich; Merck KGaA) at room temperature
for 5 min. After washing with PBS, TUNEL solution was added for 1 h
at 37˚C. DAPI was then used to stain cells for 10 min at room
temperature in the dark. Finally, five random fields of views were
selected for analysis, in which H9c2 cell apoptosis was observed
using glass coverslips with PBS as mounting medium. Images were
captured using a regular optical microscope (magnification, x200).
The number of apoptotic cells was calculated as follows: Mean
proportion of positive cells/total number of cells in five fields
of view per slide.
Data analysis
Statistical analyses were performed using GraphPad
Prism software (version 8.0.1; GraphPad Software, Inc.). Data are
presented as the mean ± SD. One-way ANOVA followed by Tukey's post
hoc test was used to analyze comparisons among multiple groups.
P<0.05 was considered to indicate a statistically significant
difference. Each experiment was repeated three times.
Results
Decreased expression levels of CPNE3
and RACK1 in I/R-induced rat myocardial tissues
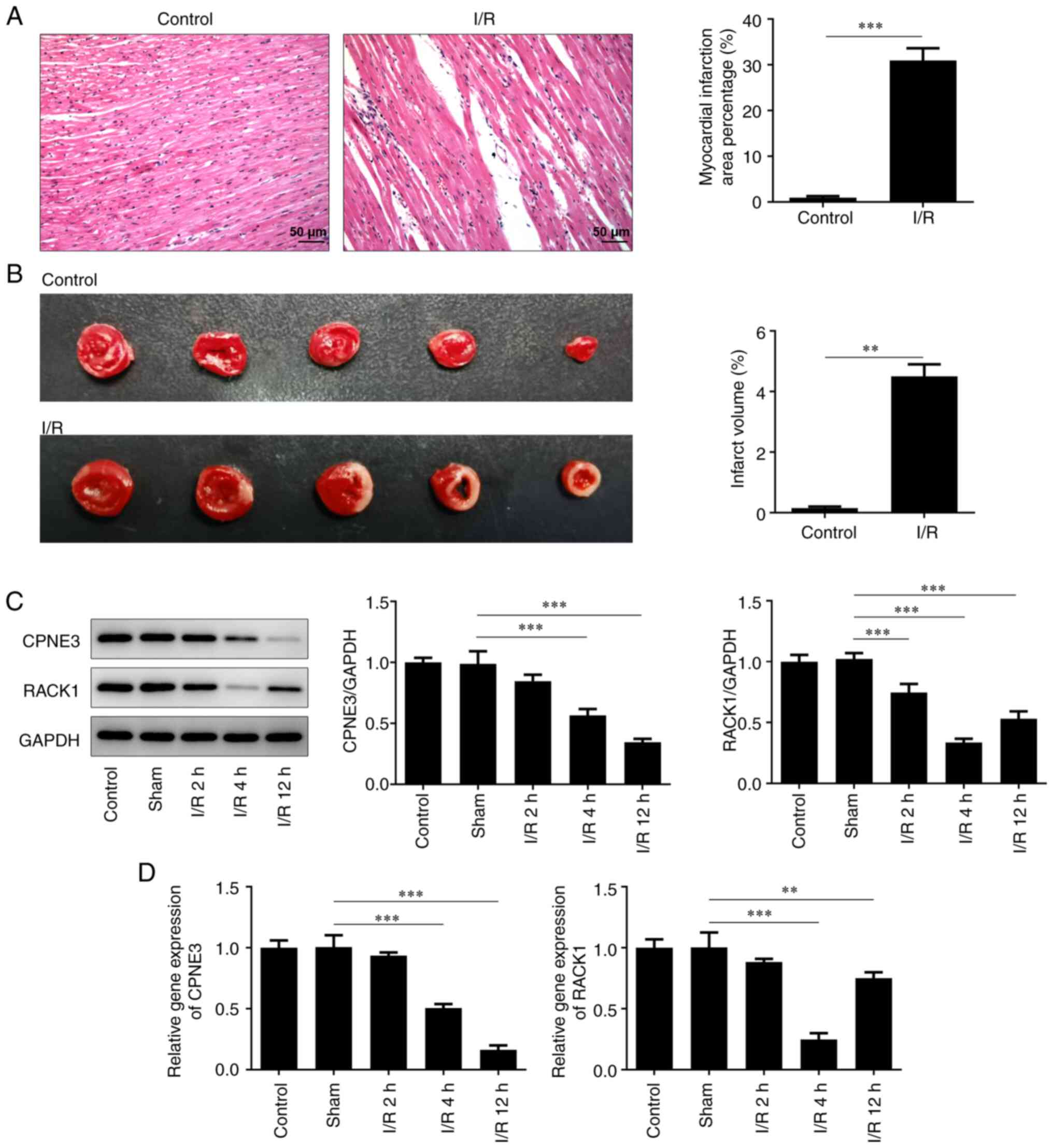
Pathological infiltration and the ischemic area of
the myocardial tissues of I/R-induced rats were observed via
H&E staining. Compared with the control group, the I/R-induced
myocardial tissues displayed severe pathological infiltration and a
significantly larger ischemic area (Fig. 1A and B). Additionally, the relative gene and
protein expression levels of CPNE3 and RACK1 were detected via
RT-qPCR and western blotting, respectively. CPNE3 and RACK1
expression levels were significantly lower compared with those in
the control and sham groups exposed to I/R for 2, 4 and 12 h
(Fig. 1C and D). The results demonstrated that the
expression level of CPNE3 decreased with increasing reperfusion
time. Moreover, although the expression level of RACK1 started to
significantly rise from 4 to 12 h of I/R, it remained lower
compared with that in the control group.
Decreased expression levels of CPNE3
and RACK1 in hypoxia/reoxygenation (H/R)-induced
cardiomyocytes
Cell viability was detected after H/R for 2, 4 or 12
h. The results demonstrated that the viability of H9c2 cells was
gradually decreased with increasing reoxygenation time (Fig. 2A). The western blotting (Fig. 2B and C) and RT-qPCR (Fig. 2D) results indicated decreased
expression levels of CPNE3 and RACK1 in H/R-induced H9c2 cells with
increasing durations of H/R; however, RACK1 only increased between
4 and 12 h of H/R. Therefore, H/R for 4 h was selected for
subsequent experiments.
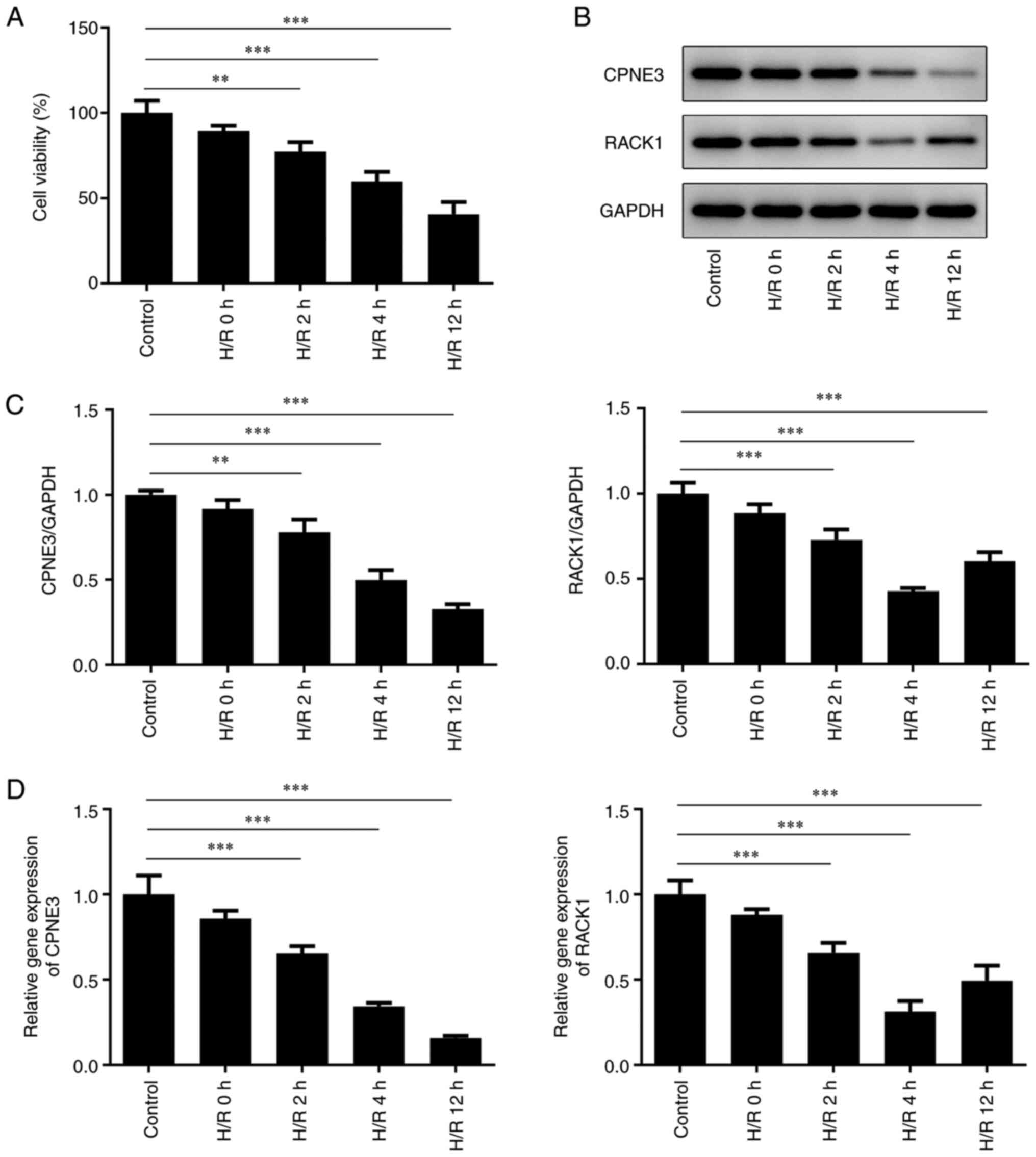 | Figure 2(A) H9c2 cell viability after H/R for
0, 2, 4 or 12 h was detected by performing Cell Counting Kit-8
assays. CPNE3 and RACK1 protein expression levels in H9c2 cells
after H/R for 0, 2, 4 or 12 h were (B) determined by western
blotting and (C) semi-quantified. (D) CPNE3 and RACK1 mRNA
expression levels in H9c2 cells after H/R for 0, 2, 4 or 12 h.
**P<0.01, ***P<0.001. H/R,
hypoxia/reoxygenation; CPNE3, copine 3; RACK1, receptor for
activated C kinase 1. |
CPNE3 overexpression activates RACK1
expression in H/R-induced cardiomyocytes
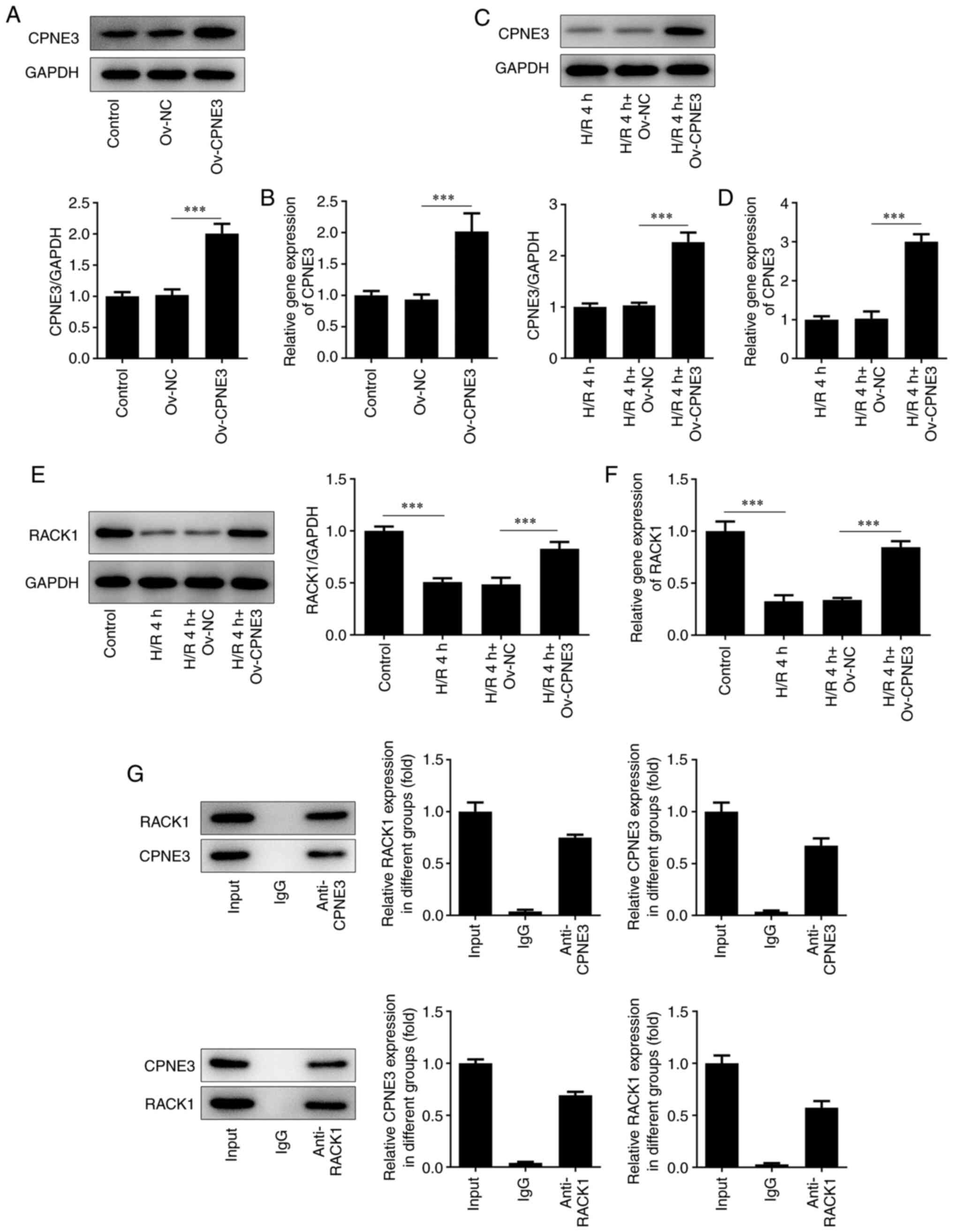
To investigate how CPNE3 affected or interacted with
RACK1 in myocardial I/R injury, untreated H9c2 cells and H9c2 cells
exposed to 4 h of H/R were transfected with Ov-CPNE3. CPNE3
expression was detected via RT-qPCR and western blotting. The
Ov-CPNE3 group displayed significantly increased CPNE3 expression
levels compared with those in the Ov-NC group (Fig. 3A and B). Similarly, CPNE3 expression levels were
significantly elevated in the H/R 4 h + Ov-CPNE3 group compared
with those in the H/R 4 h + Ov-NC group (Fig. 3C and D). Subsequently, RT-qPCR and western
blotting were performed to detect the expression levels of RACK1 in
different groups. The results demonstrated that RACK1 expression
was significantly decreased in the H/R 4 h group compared with that
in the control group. Moreover, RACK1 expression was significantly
increased in H9c2 cells subjected to H/R for 4 h that were
transfected with Ov-CPNE3 compared with those transfected with
Ov-NC (Fig. 3E and F). Furthermore, the IP assay results
revealed that the relative enrichment of CPNE3 and RACK1 was
statistically enhanced after treatment with the CPNE3 polyclonal
antibody compared with that in the IgG group, which indicated the
interaction between CPNE3 and RACK1 (Fig. 3G). Taken together, these results
suggested that CPNE3 interacted with RACK1, and CPNE3
overexpression activated RACK1 expression in H9c2 cells induced by
4 h of H/R.
CPNE3 overexpression alleviates
H/R-induced decreases in H9c2 cell viability and inhibits LDH
cytotoxic release via RACK1 activation
To examine the potential effect of the interaction
between CPNE3 and RACK1, H9c2 cells were transfected with
RACK1-targeted siRNAs. siRNA-RACK1-1 transfection resulted in the
lowest expression levels of RACK1 compared with the siRNA-NC and
siRNA-RACK1-2 groups (Fig. 4A and
B). Thus, siRNA-RACK1-1 was
selected for subsequent experiments. The viability of H/R-induced
H9c2 cells in different groups was assessed using a CCK-8 assay.
Cell viability that was weakened by H/R for 4 h was significantly
enhanced by CPNE3 overexpression. Furthermore, siRNA-RACK1
significantly downregulated the viability of H/R-induced H9c2 cells
co-transfected with Ov-CPNE3 compared with those co-transfected
with siRNA-NC (Fig. 4C).
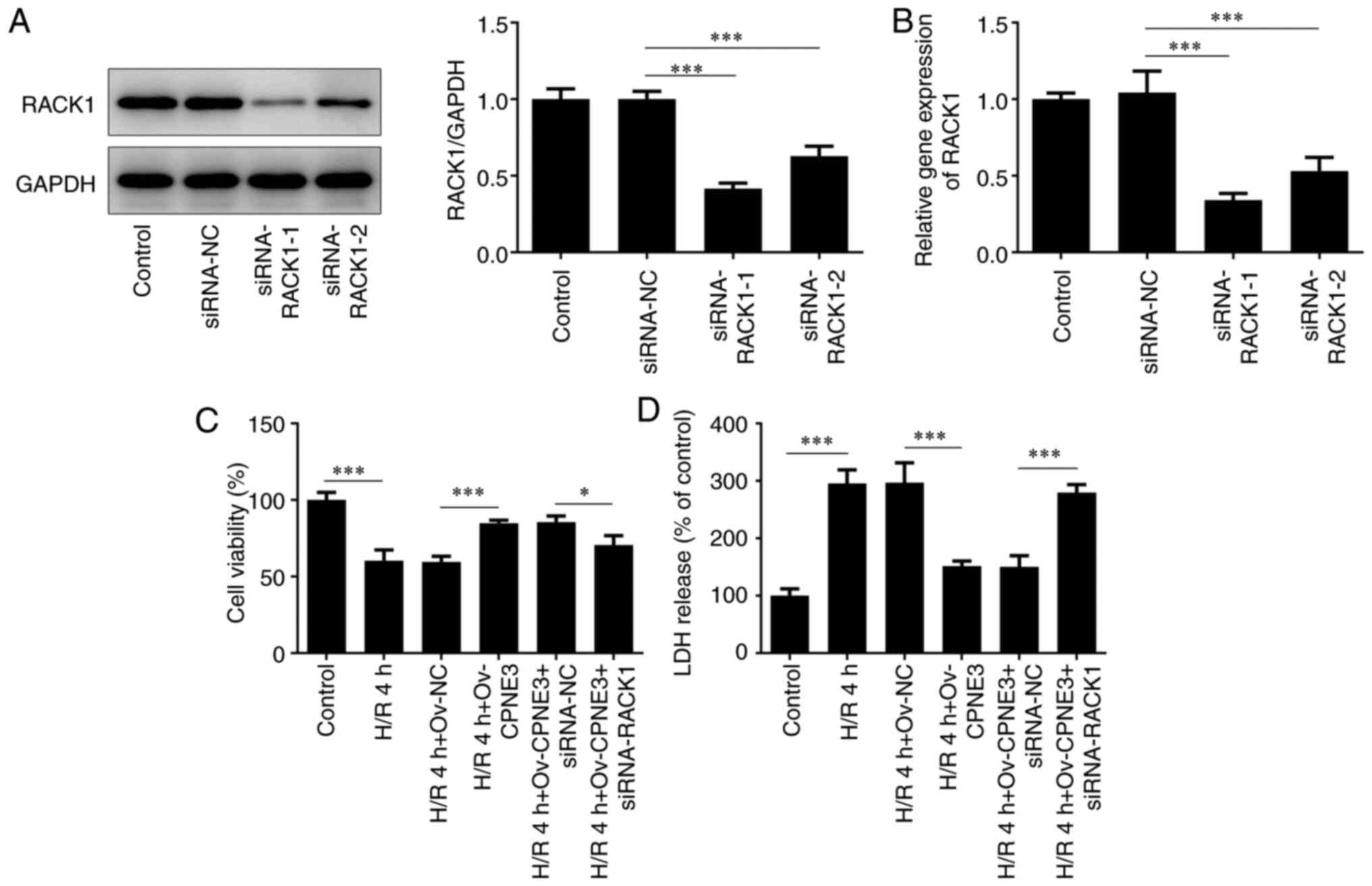 | Figure 4Transfection efficiencies of
siRNA-RACK1-1 and siRNA-RACK1-2 in H9c2 cells were detected by (A)
western blotting and (B) reverse transcription-quantitative PCR.
(C) Effect of siRNA-RACK1 on H9c2 cell viability in H/R-induced,
CPNE3-overexpressing H9c2 cells was detected by performing Cell
Counting Kit-8 assays. (D) Effect of siRNA-RACK1 on LDH release in
H/R-induced, CPNE3-overexpressing H9c2 cells was detected using an
LDH assay kit. *P<0.05, ***P<0.001.
siRNA, small interfering RNA; RACK1, receptor for activated C
kinase 1; H/R, hypoxia/reoxygenation; CPNE3, copine 3; LDH, lactate
dehydrogenase; NC, negative control; Ov, overexpression. |
The release of cytotoxic LDH in H/R-induced H9c2
cells was assessed using a LDH detecting commercial kit. It was
found that, compared with the control group, 4 h of H/R
significantly increased the release of LDH in H9c2 cells, which was
significantly decreased by CPNE3 overexpression. However, CPNE3
overexpression-induced effects on LDH release were significantly
inhibited in H/R-induced H9c2 cells co-transfected with siRNA-RACK1
compared with those co-transfected with siRNA-NC (Fig. 4D). These findings indicated that
CPNE3 overexpression alleviated H/R-induced decreases in H9c2 cell
viability and inhibited the release of cytotoxic LDH by activating
RACK1 expression.
CPNE3 overexpression reduces the
release of inflammatory cytokines in H/R-induced H9c2 cells via
RACK1 activation
To evaluate whether CPNE3 exerted an
anti-inflammatory effect in myocardial I/R injury by interacting
with RACK1, the expression levels of inflammatory response-related
cytokines (TNF-α, IL-1β and IL-6) in different treatment groups
were detected via RT-qPCR and the expression levels of
proinflammatory factors (p-NF-κB P65, P65 and Cox2) were detected
via western blotting. The relative gene expression levels of TNF-α,
IL-1β and IL-6 were significantly increased by H/R compared with
those in the control group. CPNE3 overexpression significantly
attenuated these increased expression levels in H/R-induced H9c2
cells, an effect that was significantly inhibited by
co-transfection with siRNA-RACK1 (Fig.
5A). The expression levels of p-NF-κB P65/P65 and Cox2
displayed similar trends; compared with the control group, the
protein phosphorylation and expression levels, respectively, of
these proinflammatory factors were significantly elevated by H/R.
CPNE3 overexpression significantly attenuated these effects in
H/R-induced H9c2 cells, whereas interference with siRNA-RACK1
significantly inhibited the effect of CPNE3 overexpression
(Fig. 5B). These findings indicated
that CPNE3 overexpression decreased the release of inflammatory
cytokines in H/R-induced H9c2 cells by upregulating RACK1
expression.
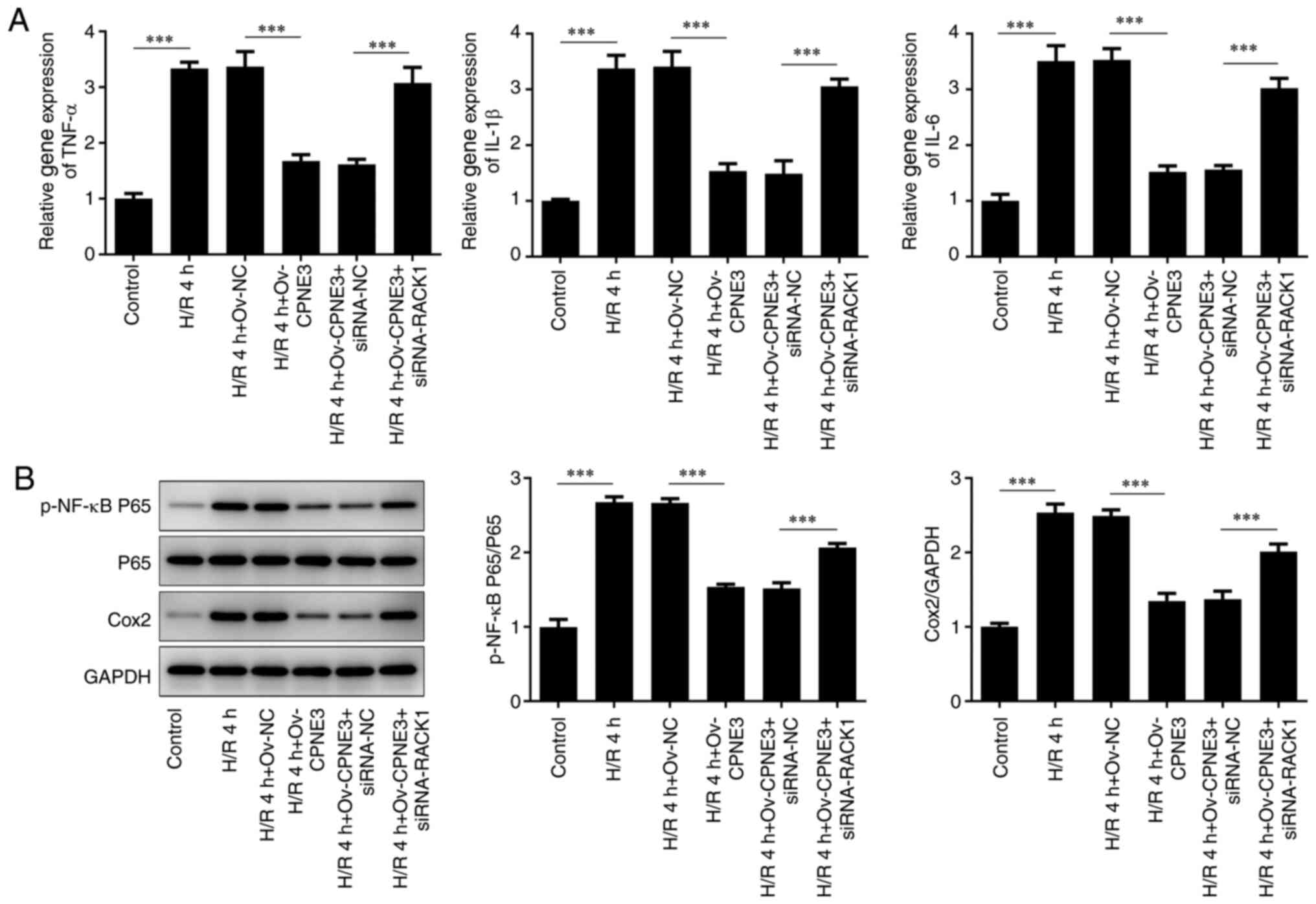 | Figure 5(A) Expression levels of TNF-α, IL-1β
and IL-6 in H/R-induced, CPNE3-overexpressing H9c2 cells
co-transfected with siRNA-RACK1 were detected by reverse
transcription-quantitative PCR. (B) Expression levels of p-NF-κB
P65, P65 and Cox2 in H/R-induced, CPNE3-overexpressing H9c2 cells
co-transfected with siRNA-RACK1 were detected by western blotting.
***P<0.001. H/R, hypoxia/reoxygenation; CPNE3, copine
3; siRNA, small interfering RNA; RACK1, receptor for activated C
kinase 1; p, phosphorylated; NC, negative control; Ov,
overexpression. |
CPNE3 overexpression decreases the
apoptosis of H/R-induced H9c2 cells via RACK1 activation
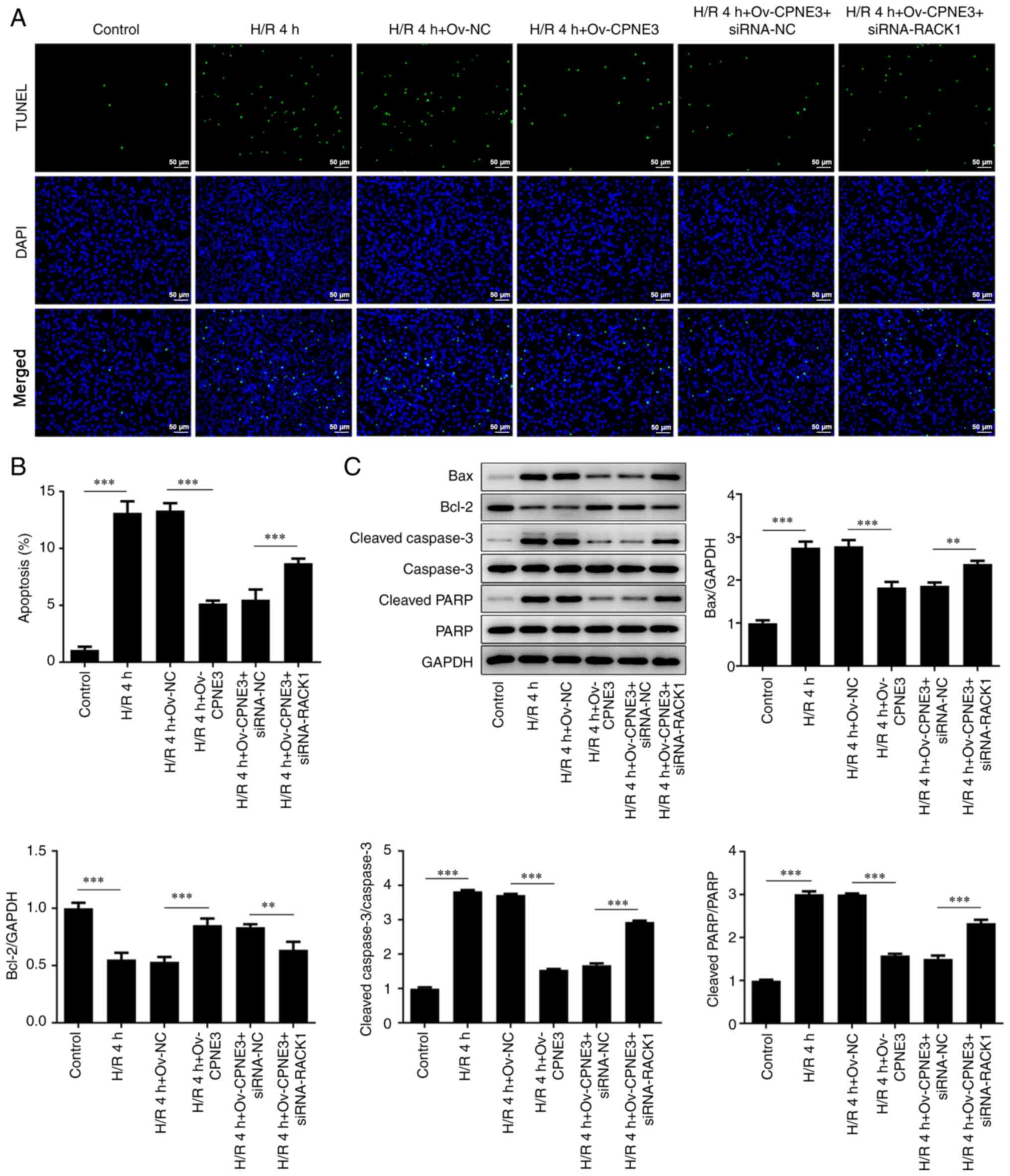
To further investigate the effect induced by the
interaction between CPNE3 and RACK1, cell apoptosis was assessed
via TUNEL staining and western blotting. A significantly increased
number of apoptotic cells was observed in the H/R group compared
with that in the control group. A significant decrease in the
number of apoptotic cells was identified after CPNE3 overexpression
in H/R-induced H9c2 cells, whereas siRNA-RACK1 co-transfection
significantly inhibited this effect (Fig. 6A and B). Moreover, the expression levels of the
apoptosis-related proteins Bax, cleaved caspase-3 and cleaved PARP
were significantly elevated in the H/R 4 h group compared with
those in the control group. However, H/R-induced effects on
apoptosis-related protein expression levels were significantly
decreased by CPNE3 overexpression. By contrast, siRNA-RACK1
significantly increased these expression levels in H/R-induced H9c2
cells co-transfected with Ov-CPNE3 (Fig. 6C). Bcl-2 expression displayed the
opposite trend. Moreover, total caspase-3 and PARP expression
levels were not markedly altered among the groups. These results
suggested that CPNE3 overexpression facilitated the decline in
H/R-induced H9c2 cell apoptosis by activating RACK1 expression.
Discussion
Reperfusion therapy is a typical approach used for
the restoration of blood and oxygen supply to ischemic tissues, and
is widely used in the treatment of acute myocardial infarction,
pulmonary embolism, deep vein thrombosis and peripheral artery
disease (8). Unfortunately, I/R has
some negative side effects, resulting in severe dysfunction of the
organism and even death (9). The
pathological mechanism underlying I/R injury is often characterized
by inflammation, abnormal microvascular function and cell death
(10). A previous review described
hyperbaric oxygen therapy as a protective strategy against I/R
injury (11). Therefore, the
present study established I/R-induced H9c2 cells to generate an
in vitro myocardial ischemia-reperfusion injury model. The
H9c2 cell line has the ability of cell division, and numerous
studies have used H/R to induce H9c2 cells to establish myocardial
ischemia injury model (12-14).
In addition, due to the weak proliferative ability of primary
cardiomyocytes (generally considered as telophase cells), the
survival rate of primary cardiomyocytes is not high and the culture
is difficult. Therefore, the present study used H9c2 cells to
conduct the experiments. Future studies should verify the results
of the present study in primary cardiomyocytes.
As previously mentioned, it has been shown that
CPNE3 expression is closely associated with the risk of
experiencing acute myocardial infarction that requires reperfusion
therapy, and there is novel evidence that supports the interaction
between CPNE3 and RACK1 in NSCLC (4). Therefore, the present study aimed to
investigate the possible protective effect of CPNE on I/R injury
via an interaction with RACK1. It has been reported that
upregulation of CPNE3 suppresses the proliferation of glioblastoma
cells via focal adhesion kinase signaling pathway inactivation
(15). These results indicated that
CPNE3 may serve an important regulatory effect in cell apoptosis.
In addition, according to previous research, the expression level
of RACK1 is downregulated after myocardial I/R and is closely
associated with the apoptosis of cardiomyocytes (5). The experimental I/R injury rat model
established in the present study showed consistent results with the
aforementioned previous research, displaying significantly
downregulated CPNE3 and RACK1 expression levels after I/R
induction. Moreover, upregulated expression levels of RACK1 were
observed in H/R-induced H9c2 cells after transfection with
Ov-CPNE3, and the IP assay results further validated the
interaction between CPNE3 and RACK1.
The pathology underlying I/R injury can be partially
represented by decreased cell viability, as a previous study has
shown that thrombolysis and recanalization therapies on patients
with acute ischemic stroke induce rapid loss of cell viability in
the tissues (16). Therefore, the
present study examined cell viability in H/R-induced H9c2 cells to
assess the effect of CPNE3 interacting with RACK1. The results of
the present study demonstrated that CPNE3 overexpression increased
cell viability via RACK1 activation. Reperfusion therapy affects
cell viability and causes cell death, largely due to being a toxic
process itself (17). Moreover, the
release of LDH is implicated in the I/R process and is reported to
be dependent on the duration of I/R (18). The present study demonstrated that
CPNE3 overexpression could effectively inhibit the release of
cytotoxic LDH by regulating RACK1 expression, which further
indicated the potential role of CPNE3 in I/R injury.
I/R therapy can contribute to or is not sufficient
to overcome the inflammatory cascade in patients with
atherosclerotic cardiovascular disease (19). A previous study revealed that
inhibiting the inflammatory response at the stage of early
reperfusion may serve as a feasible strategy to prevent injury
(20). The present study not only
demonstrated an increased release of proinflammatory cytokines in
H/R-induced H9c2 cells, but also confirmed the anti-inflammatory
effect of CPNE3 on I/R injury by activating RACK1 expression. Signs
of cell death are observed in patients after post-ischemic
reperfusion, and such a tendency is difficult to alleviate
(21,22). I/R-induced apoptosis is considered a
trigger for ultimate cell death (23). The level of apoptosis and the
expression levels of apoptosis-related proteins were detected in
the present study. An overall decline in cell apoptosis and the
expression of apoptosis-related proteins was observed in
H/R-induced H9c2 cells following transfection with Ov-CPNE3, and
Ov-CPNE3-mediated effects were inhibited by siRNA-RACK1
interference. Thus, the results suggested that the interaction
between CPNE3 and RACK1 regulated cardiomyocyte apoptosis.
Collectively, the present study suggested that CPNE3
may serve an important role in preventing I/R injury by interacting
with RACK1. Therefore, the results of the present study may
facilitate the development of advanced preventive strategies
against myocardial I/R injury and novel clinical therapeutic
strategies for patients with I/R.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
The datasets used and/or analyzed generated during
the current study are available from the corresponding author on
reasonable request.
Authors' contributions
XZ wrote the manuscript and analyzed the data. XH
and YZ carried out the experiments, supervised the present study,
searched the literature and revised the manuscript. All authors
read and approved the final manuscript. XZ and XH confirm the
authenticity of all the raw data.
Ethics approval and consent to
participate
The present study was approved by the Committee on
the Ethics of Animal Experiments of Cangzhou Central Hospital. All
animal experiments comply with the ethical requirements of the
animal council.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Binder A, Ali A, Chawla R, Aziz HA, Abbate
A and Jovin IS: Myocardial protection from ischemia-reperfusion
injury post coronary revascularization. Expert Rev Cardiovasc Ther.
13:1045–1057. 2015.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Tan B, Liu L, Yang Y, Liu Q, Yang L and
Meng F: Low CPNE3 expression is associated with risk of acute
myocardial infarction: A feasible genetic marker of acute
myocardial infarction in patients with stable coronary artery
disease. Cardiol J. 26:186–193. 2019.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Katanasaka Y: Development of targeted
pharmacotherapy for cardiovascular disease. Yakugaku Zasshi.
137:1349–1353. 2017.PubMed/NCBI View Article : Google Scholar : (In Japanese).
|
|
4
|
Lin H, Zhang X, Liao L, Yu T, Li J, Pan H,
Liu L, Kong H, Sun L, Yan M and Yao M: CPNE3 promotes migration and
invasion in non-small cell lung cancer by interacting with RACK1
via FAK signaling activation. J Cancer. 9:4215–4222.
2018.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Willerson JT, Watson JT, Hutton I, Fixler
DE, Curry GC and Templeton GH: The influence of hypertonic mannitol
on regional myocardial blood flow during acute and chronic
myocardial ischemia in anesthetized and awake intact dogs. J Clin
Invest. 55:892–902. 1975.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Qian L, Shi J, Zhang C, Lu J, Lu X, Wu K,
Yang C, Yan D, Zhang C, You Q and Liu X: Downregulation of RACK1 is
associated with cardiomyocyte apoptosis after myocardial
ischemia/reperfusion injury in adult rats. In Vitro Cell Dev Biol
Anim. 52:305–313. 2016.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Bhaskar S, Stanwell P, Cordato D, Attia J
and Levi C: Reperfusion therapy in acute ischemic stroke: Dawn of a
new era? BMC Neurol. 18(8)2018.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Kalogeris T, Baines CP, Krenz M and
Korthuis RJ: Ischemia/Reperfusion. Compr Physiol. 7:113–170.
2016.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Granger DN and Kvietys PR: Reperfusion
therapy-What's with the obstructed, leaky and broken capillaries?
Pathophysiology. 24:213–228. 2017.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Hentia C, Rizzato A, Camporesi E, Yang Z,
Muntean DM, Săndesc D and Bosco G: An overview of protective
strategies against ischemia/reperfusion injury: The role of
hyperbaric oxygen preconditioning. Brain Behav.
8(e00959)2018.PubMed/NCBI View
Article : Google Scholar
|
|
12
|
Ma K, Qiu J, Zhou M, Yang Y and Ye X:
Cox-2 negatively affects the protective role of propofol against
hypoxia/reoxygenation induced cardiomyocytes apoptosis through
suppressing akt signaling. Biomed Res Int.
2019(7587451)2019.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Ge L, Cai Y, Ying F, Liu H, Zhang D, He Y,
Pang L, Yan D, Xu A, Ma H and Xia Z: MiR-181c-5p exacerbates
hypoxia/reoxygenation-induced cardiomyocyte apoptosis via targeting
PTPN4. Oxid Med Cell Longev. 2019(1957920)2019.PubMed/NCBI View Article : Google Scholar
|
|
14
|
He F, Wu Q, Xu B, Wang X, Wu J, Huang L
and Cheng J: Suppression of Stim1 reduced intracellular calcium
concentration and attenuated hypoxia/reoxygenation induced
apoptosis in H9C2 cells. Biosci Rep. 37(BSR20171249)2017.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Shi D, Lin B, Lai J, Li K and Feng Y:
Upregulation of CPNE3 suppresses invasion, migration and
proliferation of glioblastoma cells through FAK pathway
inactivation. J Mol Histol. 52:589–596. 2021.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Carati CJ, Rambaldo S and Gannon BJ:
Changes in macromolecular permeability of microvessels in rat small
intestine after total occlusion ischemia/reperfusion. Microcirc
Endothelium Lymphatics. 4:69–86. 1988.PubMed/NCBI
|
|
17
|
Hacker TA, Diarra G, Fahl BL, Back S,
Kaufmann E and Fahl WE: Significant reduction of
ischemia-reperfusion cell death in mouse myocardial infarcts using
the immediate-acting PrC-210 ROS-scavenger. Pharmacol Res Perspect.
7(e00500)2019.PubMed/NCBI View
Article : Google Scholar
|
|
18
|
Rossello X, Hall AR, Bell RM and Yellon
DM: Characterization of the langendorff perfused isolated mouse
heart model of global ischemia-reperfusion injury: Impact of
ischemia and reperfusion length on infarct size and LDH release. J
Cardiovasc Pharmacol Ther. 21:286–295. 2016.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Goldfine AB and Shoelson SE: Therapeutic
approaches targeting inflammation for diabetes and associated
cardiovascular risk. J Clin Invest. 127:83–93. 2017.PubMed/NCBI View
Article : Google Scholar
|
|
20
|
Bonaventura A, Montecucco F and Dallegri
F: Cellular recruitment in myocardial ischaemia/reperfusion injury.
Eur J Clin Invest. 46:590–601. 2016.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Heusch G: Critical issues for the
translation of cardioprotection. Circ Res. 120:1477–1486.
2017.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Hausenloy DJ and Yellon DM: Ischaemic
conditioning and reperfusion injury. Nat Rev Cardiol. 13:193–209.
2016.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Jiang X, Lew KS, Chen Q, Richards AM and
Wang P: Human mesenchymal stem cell-derived exosomes reduce
ischemia/reperfusion injury by the inhibitions of apoptosis and
autophagy. Curr Pharm Des. 24:5334–5341. 2018.PubMed/NCBI View Article : Google Scholar
|