Introduction
Oral squamous cell carcinoma (OSCC) develops from
the epithelium lining of the oral cavity and accounts for >90%
of all cases of oral cancer worldwide (1). OSCC is the sixth most common solid
tumor malignancy worldwide and is the most common malignant
epithelial neoplasm occurring in the head and neck region in terms
of incidence and mortality (2,3). OSCC
has a tendency to cause regional lymph node metastasis and
recurrence (4). Despite advances in
diagnostic and therapeutic strategies in the past few decades, the
5-year survival rate for OSCC remains relatively low at ~50%, which
poses a great challenge for the prognosis of OSCC (5,6).
Therefore, identifying the complicated molecular signatures of OSCC
may aid in the diagnosis and treatment of the disease.
As a member of the tribbles-related family, Tribbles
pseudokinase 3 (TRIB3) contains substrate-binding domains but lacks
the conserved catalytic amino acid motifs essential for kinase
activity (7,8). A number of studies have demonstrated
that TRIB3 is involved in diverse cellular processes, including
cell proliferation and differentiation, the cellular stress
response, epithelial-to-mesenchymal transition and glucose and
lipid metabolism (9-11).
Emerging evidence suggests that TRIB3 is a crucial oncoprotein,
with the function of TRIB3 being associated with multiple different
types of cancer, including breast, colorectal and lung cancer, as
well as renal cell carcinoma (RCC) (12-15);
however, the role of TRIB3 in OSCC is not completely understood.
Additionally, the protein kinase B (AKT)/mammalian target of
rapamycin (mTOR) signaling pathway has been determined to serve an
important role in multiple cancers, including ovarian cancer,
hepatocelluar carcinoma, lung cancer and OSCC (16).
The present study focused on the importance of TRIB3
during the progression of OSCC. TRIB3 was highly expressed in human
OSCC tissues at both the mRNA and protein levels. Furthermore, the
results demonstrated the significance of TRIB3 in the proliferation
of OSCC cell lines in vitro and in vivo.
Additionally, the molecular mechanism underlying TRIB3-mediated
OSCC cell proliferation was investigated.
Materials and methods
OSCC tissue samples
After obtaining written informed consent from
patients, a total of 35 primary OSCC tissues and paired adjacent
normal tissues were obtained. All paired normal tissues were
adjacent to tumor tissues at a distance <0.5 cm. All OSCC
tissues were confirmed by two independent pathologists. RNA was
extracted from 30 paired samples for analysis via reverse
transcription-quantitative PCR (RT-qPCR) and five paired samples
were assessed via western blotting. All patients (age, 45-62 years;
mean age, 51 years; 21 male; 14 female) underwent surgery without
chemotherapy or pre-operative radiation at the Chinese People's
Liberation Army General Hospital (Beijing) from January 2017 to
January 2019. The specimens were stored at -80˚C until further
analysis. The present study was approved by the Institutional
Review Board of the Chinese People's Liberation Army General
Hospital.
Online microarray data
The relative transcript expression levels of TRIB3
and survival analyses were obtained from the Gene Expression
Profiling Interactive Analysis (GEPIA; gepia.cancer-pku.cn), an online visual tool based on
The Cancer Genome Atlas (TCGA) database (17). The online analysis of TRIB3 in GEPIA
was based on the gene expression RNAseq dataset of head and neck
squamous cell carcinoma (HNSC) in TCGA (tcga.xenahubs.net/download/TCGA.HNSC.sampleMap/HiSeqV2.gz),
which included 44 normal tissues and 520 tumor tissues. The median
value (8.1) of TRIB3 expression in tumour tissues was set as the
cut-off value to divide the 520 patients with HNSC into two groups:
TRIB3-low/medium group (n=260) and TRIB3-high group (n=260).
RNA isolation and RT-qPCR
Total RNA was extracted from the tissues of 30
patients using TRIzol® reagent (Invitrogen; Thermo
Fisher Scientific, Inc.) according to the manufacturer's
instructions. High-quality RNA (2 µg) was reverse transcribed into
cDNA using PrimeScript™ RT Reagent kit (Promega Corporation),
according to the manufacturer's protocol. Subsequently, mRNA levels
were quantified via qPCR using the PIKOREAL96 detection system
(Thermo Fisher Scientific, Inc.) and the 2X SYBR Green kit (Roche
Applied Science), according to the manufacturer's protocol. The
following thermocycling conditions were used for qPCR: Initial
denaturation for at 95˚C for 1 min, 40 cycles of denaturation for
at 95˚C for 20 sec, annealing at 60˚C for 30 sec and extension at
72˚C for 20 sec, followed by final extension at 72˚C for 10 min.
The following primers were used for qPCR: TRIB3 forward,
5'-AAGCGGTTGGAGTTGGATGAC-3' and reverse,
5'-CACGATCTGGAGCAGTAGGTG-3'; and GAPDH forward,
5'-ATGACCCCTTCATTGACCTCA-3' and reverse,
5'-GAGATGATCACCCTTTTGGCT-3'. TRIB3 mRNA expression levels were
quantified using the 2-ΔΔCq method (18) and normalized to the internal
reference gene GAPDH.
Cell culture
OSCC (SCC9, HSC3, Cal27 and SCC15), normal oral
(HOK) and 293T cell lines were purchased from The Cell Bank of Type
Culture Collection of Chinese Academy of Sciences. All cell lines
were maintained in DMEM medium (Invitrogen; Thermo Fisher
Scientific, Inc.) supplemented with 10% FBS (Invitrogen; Thermo
Fisher Scientific, Inc.) and 1% penicillin/streptomycin (Sangon
Biotech Co., Ltd.) at 37˚C in humidified atmosphere containing 5%
CO2.
Cell transfection
A TRIB3 coding sequence was constructed and inserted
into the p23-3xflag-GFP vector (Invitrogen; Thermo Fisher
Scientific, Inc.) to generate the TRIB3 overexpression vector.
Lentiviral short hairpin (sh)RNAs targeting TRIB3 were designed and
constructed into the pLKO.1-TRC vector (Sangon Biotech Co., Ltd.) A
non-targeted scrambled (SCR) oligonucleotide (Sangon Biotech Co.,
Ltd.) was used as the negative control.
To produce lentiviral particles for TRIB3
overexpression and knockdown, the core plasmid (1.0 µg) was
co-transfected with the packaging plasmids pMD2.G and psPAX2 (2.5
µg, Sangon Biotech Co., Ltd) into 293 T cells (5x105
units/well) using a calcium phosphate co-precipitation method
(19). At 12 h post-transfection,
the medium was replaced with DMEM. At 24 h post-transfection, the
supernatants containing the virus were collected and filtered
through a 0.45 µm membrane. Subsequently, the virus was
concentrated by centrifugation at 50,000 x g for 140 min at 4˚C.
The pellet was resuspended in 1.5 ml EP tube (Sangon Biotech Co.,
Ltd.), aliquoted and stored at -80˚C until further use. For the
transduction process, HSC3 and Cal27 cells were grown to 60%
confluence in 6-well plates and transfected with 30 µl virus
supernatant (~5x106 units per well) with 5 µg/ml
polybrene (Hanheng Biological Technology Co., Ltd.). After 24 h at
37˚C, the medium was replaced and the cells were transferred to
10-cm dishes. After 2 days at 37˚C, TRIB3-overexpression HSC3 and
Cal27 cells were screened by green fluorescence via flow cytometry
using an IX-71 flow cytometer (Olympus Corporation).
TRIB3-knockdown SCC9 and SCC15 cells were cultured and screened in
medium containing 4 µg/ml puromycin (Hanheng Biological Technology
Co., Ltd.) for 4 days. Subsequently, individual puromycin-resistant
colonies were isolated. Transfection efficiency was verified by
western blotting. Subsequent experiments were performed 24 h after
transfection. The sequences of the SCR were: Forward,
5'-TTCTCCGAACGTGTCACGT-3' and reverse, 5'-ACGTGACACGTTCGGAGAA-3'.
The sequences of the shTRIB3 were: Forward,
5'-CCGGGATCTCAAGCTGTGTCGCTTTCTCGAGAAAGCGACACAGCTTGAGATCTTTTTG-3'
and reverse,
5'-AATTCAAAAAGATCTCAAGCTGTGTCGCTTTCTCGAGAAAGCGACACAGCTTGAGATC-3'.
Western blotting
Total protein was extracted from paired tumor
tissues and cell lines by lysing cells in RIPA buffer (Thermo
Fisher Scientific, Inc.) for ≥30 min on ice. Cell lysates were
centrifuged at 10,000 x g for 15 min at 4˚C. Total protein was
quantified using Bradford reagent (Sigma-Aldrich; Merck KGaA)
according to the manufacturer's protocol. Proteins (10 µg) were
separated via 10% SDS-PAGE and transferred onto PVDF membranes. The
membranes were blocked with 5% fat-free milk for 1 h at room
temperature. Subsequently, the membranes were incubated at 4˚C
overnight with primary antibodies targeted against: TRIB3 (cat. no.
ab137526; 1:1,000; Abcam), AKT (cat. no. 4691; 1:1,000; Cell
Signaling Technology, Inc.), phosphorylated (p)-AKT (cat. no. 4060;
1:1,000; Cell Signaling Technology, Inc.), mTOR (cat. no. 2983;
1:1,000; Cell Signaling Technology, Inc.), p-mTOR (cat. no. 5536;
1:1,000; Cell Signaling Technology, Inc.), Flag (cat. no. 8164;
1:1,000; Cell Signaling Technology, Inc.) and GAPDH (cat. no. 5174;
1:1,000; Cell Signaling Technology, Inc.). Following primary
incubation, the membranes were incubated with an anti-rabbit
horseradish peroxidase-conjugated secondary antibody (cat. no.
D110103; 1:5,000; Sangon Biotech Co., Ltd.) for 2 h at room
temperature. Immunoreactive protein bands were visualized using an
enhanced chemiluminescence kit (Pierce; Thermo Fisher Scientific,
Inc.) and a Gel Dox XR system (Bio-Rad Laboratories, Inc.). GAPDH
was used as the loading control. ImageJ software (version 1.8.0,
National Institutes of Health) was used for quantification.
Immunohistochemistry
Tumor xenografts were fixed in 4% formalin (Sangon
Biotech Co., Ltd.) at room temperature for 12 h, embedded in
paraffin and cut into 5-µm-thick consecutive sections. After
deparaffinization and antigen recovery in a sodium citrate solution
(pH 6.0) for 20 min at 98˚C, the sections were washed three times
with 0.01 mol/l PBS for 5 min each time and blocked for 1 h in 0.01
mol/l PBS containing 0.3% Triton X-100 and 5% BSA (Gibco; Thermo
Fisher Scientific, Inc.). Subsequently, the sections were incubated
with anti-TRIB3 (cat. no. ab137526; 1:200; Abcam) and anti-p-AKT
(cat. no. 4060; 1:200; Cell Signaling Technology, Inc.) primary
antibodies at 4˚C overnight. After washing with 0.01 mol/l PBS, the
sections were incubated with 0.01 mol/l PBS containing a
horseradish peroxidase-conjugated anti-rabbit IgG secondary
antibody (cat. no. 7074; 1:5,000; Cell Signaling Technology, Inc.)
at room temperature for 2 h. Subsequently, the sections were
developed using 0.003% H2O2 and 0.03%
3,3'-diaminobenzidine in 0.05 mol/l Tris-HCl (pH 7.6).
Immunohistochemistry was performed in triplicate. Stained sections
were observed in five randomly selected and independent high-power
microscopic fields of view using an inverted light microscope
(magnification, x400).
Crystal violet assay
Cells were seeded (1x103 cells/well) into
6-well plates and cultured in DMEM medium supplemented with 10%
FBS. The medium was replaced every three days. After 2 weeks, the
medium was removed and the cells were fixed with 20% methanol at
room temperature for 10 min. Subsequently, the cells were stained
with 0.5% crystal violet (Sigma-Aldrich; Merck KGaA) at room
temperature or 10 min. Cells were washed with PBS and photographed
using a light microscope (magnification, x4). The absorbance values
were measured at a wavelength of 600 nm using a microplate
reader.
MTT assay
Cells were seeded (1x103 cells/well) into
96-well plates. Subsequently, 20 µl MTT solution (5 mg/ml)
was added to each well and incubated at 37˚C for 4 h. The medium
was then removed and 200 µl DMSO was added to dissolve the formazan
crystals. The microtiter plate was placed on a shaker to dissolve
the crystals. The absorbance of each well was measured at a
wavelength of 490 nm using an automated microplate reader. The MTT
assay was performed in triplicate.
Tumorigenesis in vivo
A total of 8 male BALB/c nude mice (age, 5 weeks;
median weight, 19 g; weight range, 18-21 g) were obtained from
Beijing Huafukang Bioscience Co., Ltd. Mice were housed with free
access to regular chow diet under specific pathogen-free conditions
at 23±3˚C with 35±5% humidity and 12-h light/dark cycles. raised
under standard conditions. Mice were subcutaneously injected with
100 µl cell suspension with PBS as the vehicle (1x106
cells/ml) of SCR negative control or shRNA-TRIB3 SCC9 cells into
the right flank. Mice in the SCC9 SCR group (n=4) received an
injection of SCR SCC9 cells and mice in the SCC9 shRNA-TRIB3 group
(n=4) received an injection of shRNA-TRIB3 SCC9 cells. Tumor volume
was measured every 7 days according to the following equation:
Volume=0.5 x length x width2. At the end of the
experiment (5 weeks), the mice were sacrificed by cervical
dislocation and tumors were excised. The tumors were photographed
and weighed. The maximum tumor volume observed was 698
mm3. All animal experiments were approved by the
Institutional Animal Care and Use Committee of the Chinese People's
Liberation Army General Hospital.
Statistical analysis
Statistical analyses were performed using GraphPad
Prism (version 5; GraphPad Software, Inc.). Data are presented as
the mean ± standard deviation. Comparisons between two groups were
analyzed using the unpaired Student's t-test. All experiments were
performed in triplicate. P<0.05 was considered to indicate a
statistically significant difference.
Results
TRIB3 is upregulated in OSCC
tissues
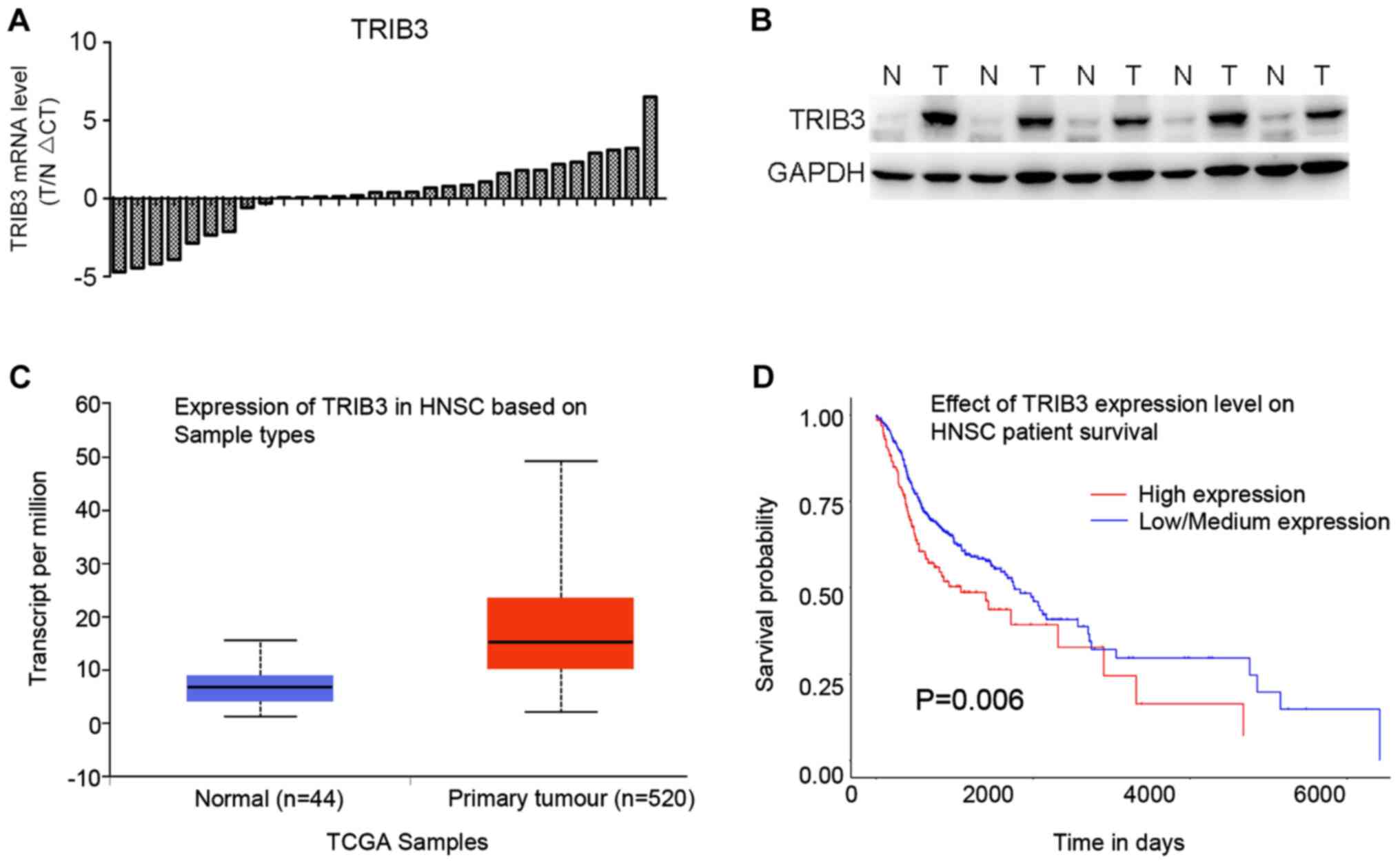
Firstly, TRIB3 mRNA expression levels in 30 paired
OSCC tissues and corresponding normal tissues were measured. TRIB3
mRNA expression was higher in 21/30 OSCC tissues compared with the
corresponding normal tissues (Fig.
1A). In addition, TRIB3 protein expression levels were also
higher in OSCC tissues compared with the corresponding normal
tissues (Fig. 1B), which was
consistent with the RT-qPCR results. As indicated by data obtained
from the GEPIA database, TRIB3 expression was higher in the 520
tumor tissues compared with the 44 normal tissues (Fig. 1C). Furthermore, the overall survival
rate of the TRIB3-low/medium group was significantly improved
compared with the TRIB3-high group (P=0.006; Fig. 1D). Based on the expression analyses
and the data obtained from the GEPIA database, the results
indicated that TRIB3 was upregulated in OSCC tissues compared with
normal tissues.
TRIB3 overexpression and knockdown in
OSCC cell lines
Based on the results obtained from the clinical
data, it was hypothesized that TRIB3 might influence OSCC cell
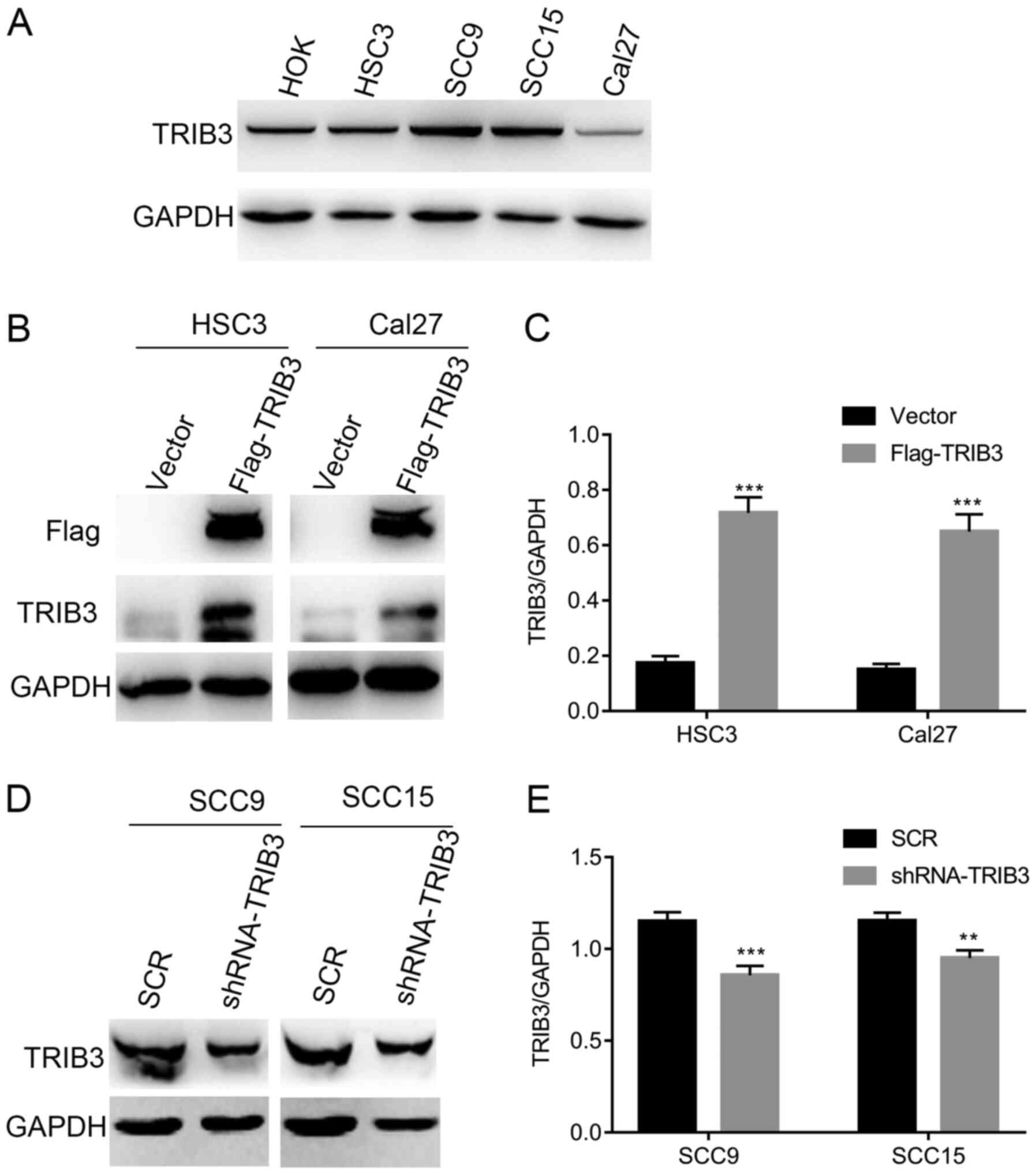
proliferation. Firstly, endogenous TRIB3 expression levels in
several human OSCC cell lines (SCC9, HSC3, Cal27 and SCC15) and a
normal oral cell line (HOK) were investigated. TRIB3 expression was
higher in SCC9 and SCC15 cells compared with HSC3, Cal27 and HOK
cells (Fig. 2A). To identify the
function of TRIB3 in OSCC cells, HSC3 and Cal27 cells were
transduced with empty p23 vector or TRIB3 overexpression vector
(Flag-TRIB3) plasmids. In addition, shRNA-TRIB3 was used to
knockdown TRIB3 expression in SCC9 and SCC15 cell lines.
Transfection efficiency was verified via western blotting (Fig. 2B-E).
TRIB3 overexpression promotes the
proliferation of OSCC cell lines
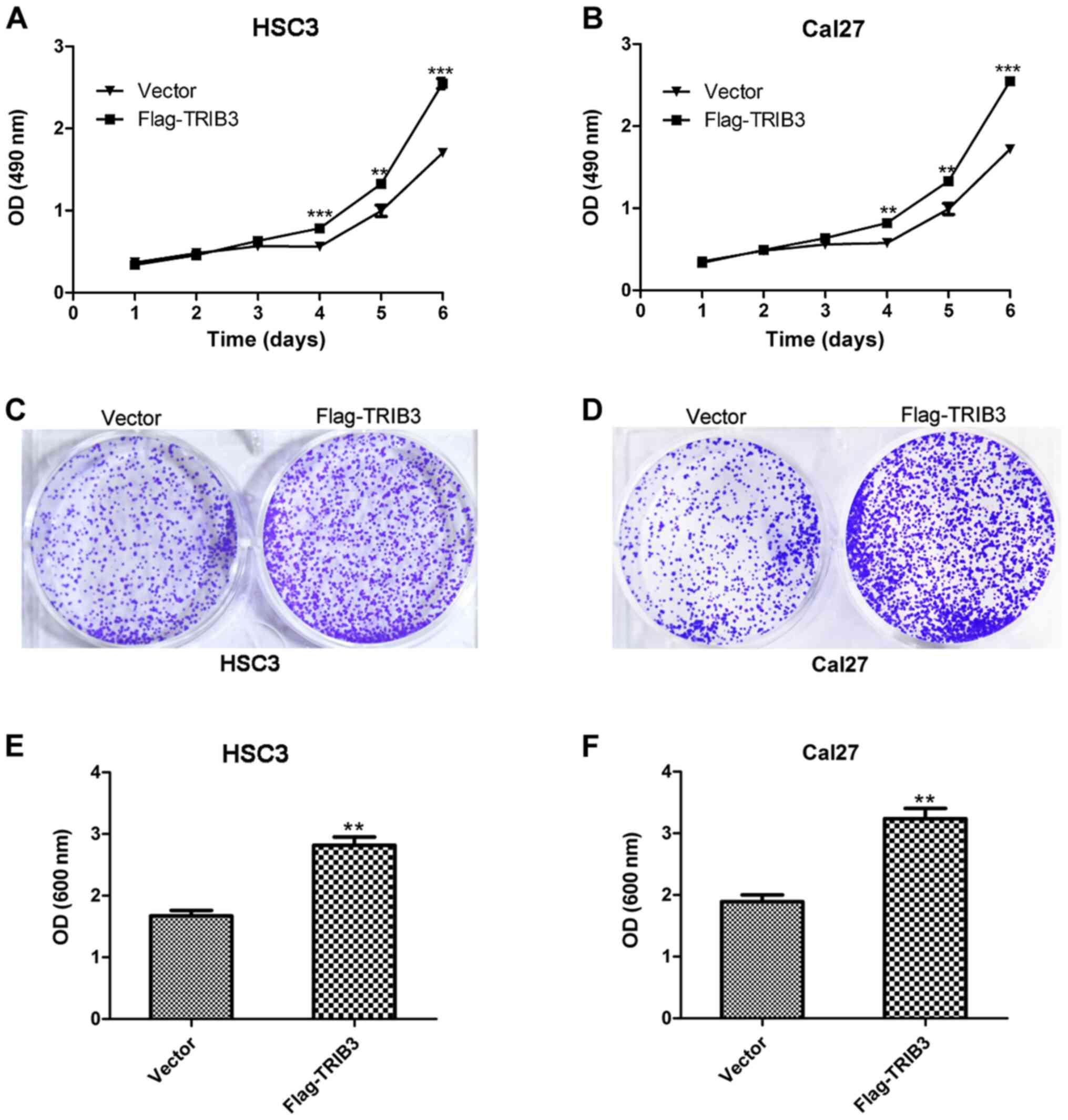
The MTT assay indicated that HSC3 and Cal27 cell
proliferation was significantly increased at 4, 5 and 6 days
post-transfection in the TRIB3 overexpression group compared with
the untreated group (P<0.01; Fig.
3A and B). Furthermore, the
crystal violet assay suggested that TRIB3 overexpression markedly
promoted the colony formation ability of HSC3 and Cal27 cells
compared with the untreated group (Fig.
3C and D), which reflected the
enhanced proliferative ability of the TRIB3-overexpression HSC3 and
Cal27 cells. Furthermore, the absorbance values of
TRIB3-overexpression HSC3 and Cal27 cells were significantly higher
compared with untreated cells (P<0.01; Fig. 3E and F).
TRIB3 knockdown suppresses the
proliferation of OSCC cell lines
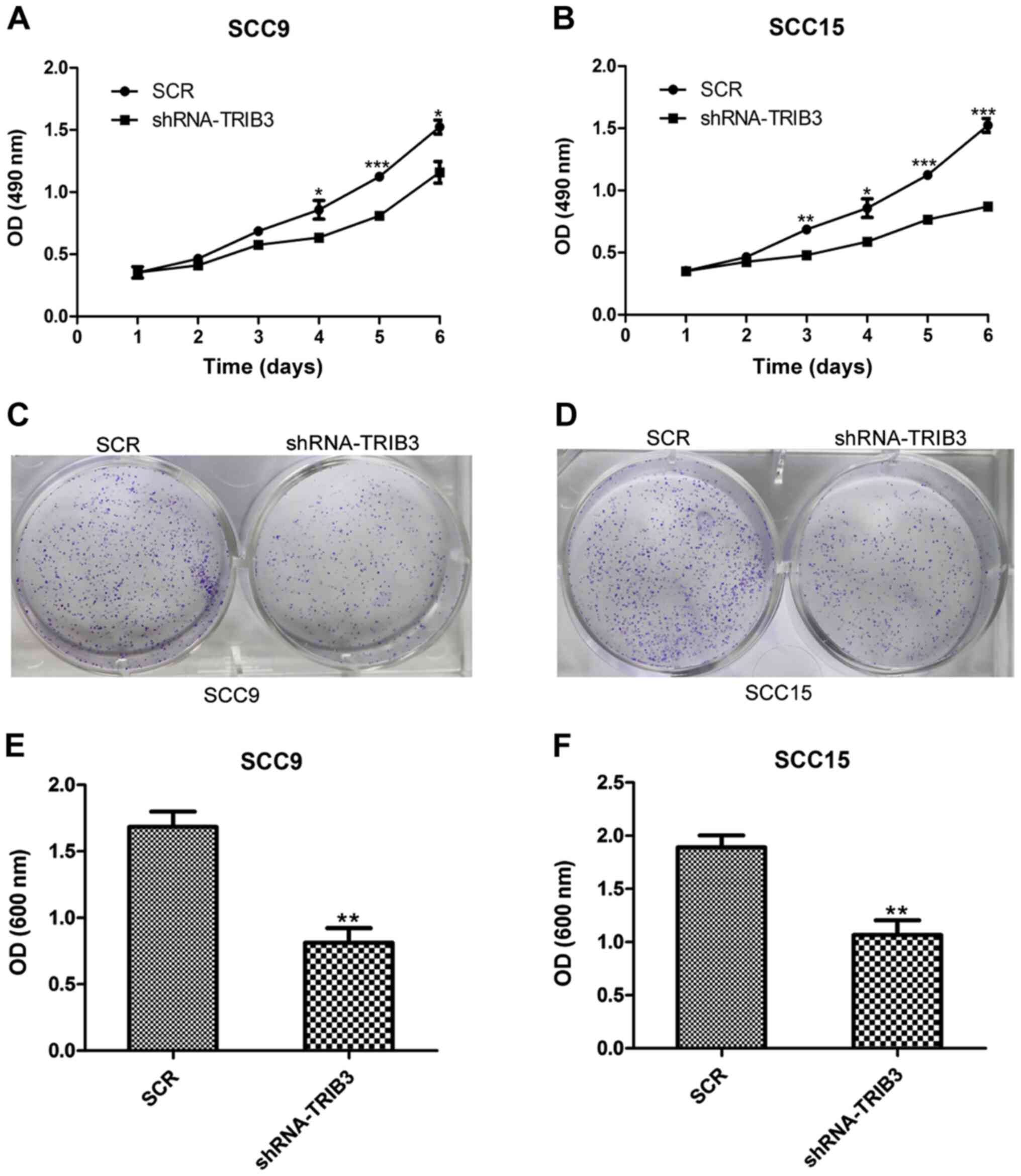
TRIB3-knockdown SCC9 and SCC15 cell lines were
established by transfection with shRNA-TRIB3 and a non-targeting
control lentivirus (SCR). Similar to the overexpression
experiments, the proliferation of the SCR- and
shRNA-TRIB3-transfected cell lines was assessed. The MTT assay
results indicated that SCC9 cell proliferation was significantly
reduced in the TRIB3 knockdown group at 4, 5 and 6 days
post-transfection compared with the untreated group (P<0.05;
Fig. 4A). Similarly, SCC15 cell
proliferation at 3, 4, 5 and 6 days post-transfection was
significantly reduced in the TRIB3 knockdown group compared with
the untreated group (P<0.05; Fig.
4B). Furthermore, the crystal violet assay results indicated
that the colony formation ability of SCC9 and SCC15 cells was
significantly lower in TRIB3-knockdown cells compared with
untreated cells (P<0.01; Fig.
4C-F).
TRIB3 knockdown suppresses OSCC cell
tumorigenesis in vivo
To verify the in vitro results in
vivo, control and TRIB3-knockdown SCC9 cells were
subcutaneously injected into the right flank of nude mice and tumor
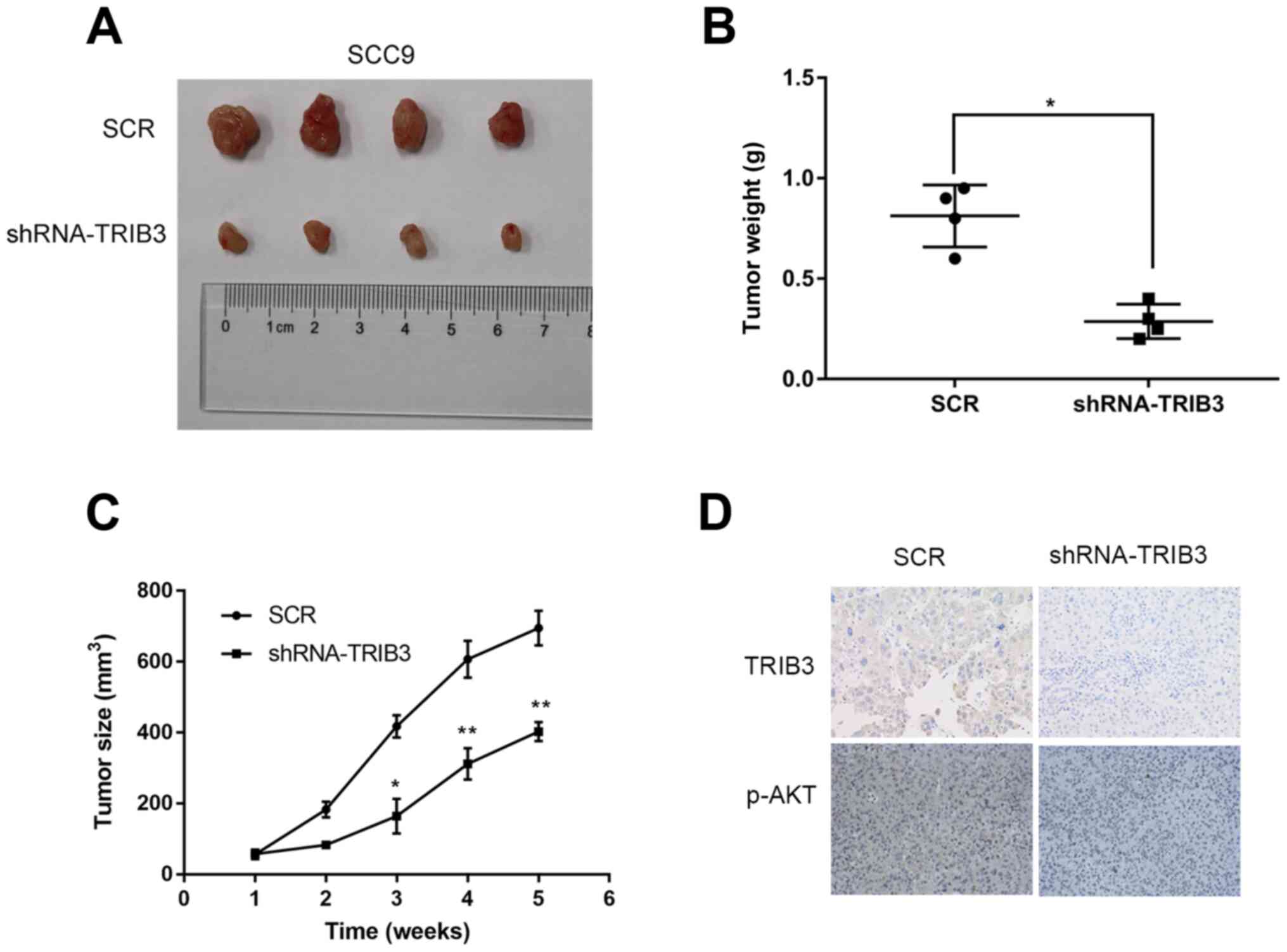
growth was monitored (Fig. 5A). The
mean tumor weight in the shRNA-TRIB3 group was significantly lower
compared with the SCR group (n=4; P=0.001; Fig. 5B). Similarly, the mean tumor volume
of the shRNA-TRIB3 group was significantly lower compared with the
SCR group at 3, 4 and 5 weeks post-injection (n=4; P<0.05;
Fig. 5C), which suggested that
TRIB3 knockdown suppressed OSCC cell-mediated tumour growth in
vivo. The results were consistent with the in vitro
results, which indicated that TRIB3 knockdown inhibited OSCC cell
proliferation. In addition, AKT phosphorylation was notably
increased in the shRNA-TRIB3 group compared with the SCR group
(Fig. 5D).
TRIB3 activates the AKT signaling
pathway in OSCC cells
It has been reported that constitutive AKT
activation serves an important role during the development and
progression of OSCC (20). To
elucidate the mechanism underlying TRIB3-mediated OSCC
tumorigenesis, western blotting was performed to investigate the
protein expression levels of p-AKT and total AKT in TRIB3
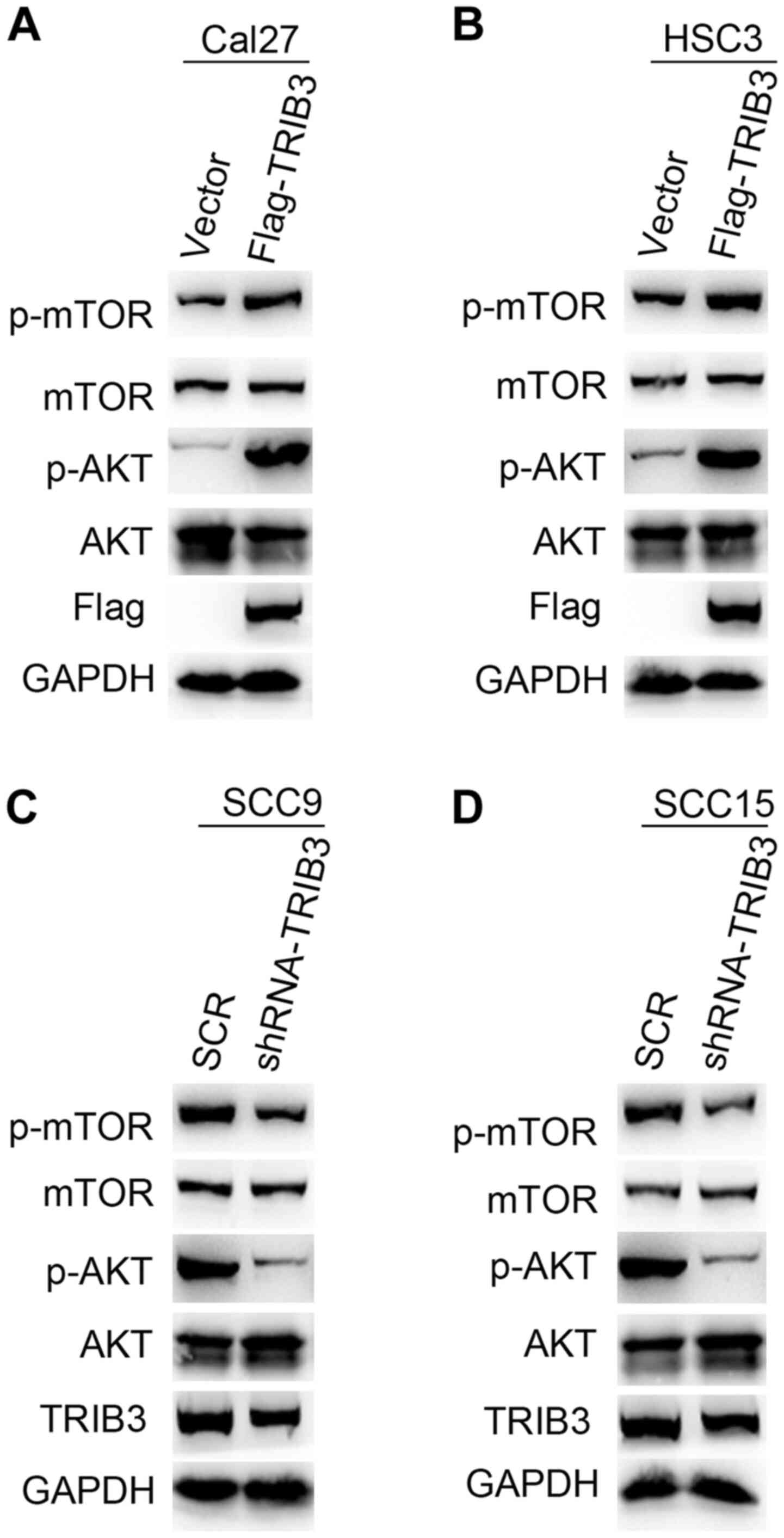
overexpression and knockdown cell lines. TRIB3 overexpression
markedly increased AKT and mTOR phosphorylation levels in HSC3 and
Cal27 cells compared with control cells, without notably altering
the total AKT and mTOR expression levels (Fig. 6A and B). By contrast, TRIB3 knockdown in SCC9
and SCC15 cells notably reduced AKT and mTOR phosphorylation
compared with SCR-transfected cells, without markedly altering the
total AKT and mTOR expression levels (Fig. 6C and D). In summary, the results indicated that
TRIB3 promoted AKT activation.
Discussion
In previous years, emerging evidence has indicated
that TRIB3 is involved in tumorigenesis and cancer progression
(12-15).
Upregulated TRIB3 expression has been associated with a suboptimal
prognosis in colon and breast cancer (12,21).
In addition, the expression level of TRIB3 was reported to display
an inverse relationship with the prognosis of breast cancer
(22). Conversely, TRIB3
upregulation was significantly correlated with tumor size and lymph
node or distal metastasis in patients with non-small cell lung
cancer (NSCLC), suggesting that TRIB3 upregulation was associated
with the poor prognosis of NSCLC (13). Furthermore, Hong et al
(15) demonstrated the potential
oncogenic role of TRIB3 in RCC, reporting that TRIB3 promoted RCC
cell proliferation, migration and invasion.
In the present study, TRIB3 mRNA and protein
expression levels were markedly higher in human OSCC tissues
compared with normal tissues. Cell proliferation was significantly
enhanced in TRIB3-overexpression cells compared with control cells.
In addition, TRIB3 knockdown inhibited OSCC cell proliferation
compared with control cells. Moreover, TRIB3 knockdown suppressed
tumor growth and decreased tumor volume in vivo compared
with control cells. The mechanism underlying TRIB3-mediated OSCC
tumorigenesis was also investigated. The results indicated that
compared with control cells, AKT and mTOR phosphorylation was
significantly increased following TRIB3 overexpression, whereas
TRIB3 knockdown decreased AKT and mTOR phosphorylation, which is
crucial for tumor progression (23). Based on the results, it was
hypothesized that TRIB3 may promote OSCC cell proliferation by
activating the AKT signaling pathway. The AKT signaling pathway
serves an important role in multiple types of cancer, including
ovarian cancer, hepatocelluar carcinoma and lung cancer (24,25).
The results of the present study were consistent with a previous
study, which reported that TRIB3 inhibited tumorigenesis by
targeting AKT (26). Restelli et
al (26) demonstrated that
TRIB3 could inhibit several cancer-related processes, such as cell
proliferation and invasion, by binding to Ser473 of the AKT protein
kinase. To the best of our knowledge, the present study was the
first study to demonstrate the role of TRIB3 in OSCC cell
proliferation. However, the present study had a number of
limitations, including the lack of analysis of mTOR expression
levels via immunohistochemistry, as well as the lack of prognostic
analysis of TRIB3 in clinical OSCC samples by performing a tissue
microarray.
In summary, the present study indicated that TRIB3
may serve a critical role in OSCC cell proliferation. The results
may improve the current understanding of the mechanisms underlying
the biological role of TRIB3 during tumor development and might
provide a potential therapeutic target for OSCC.
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
The datasets generated and/or analyzed during the
current study are available in the tcga.xenahubs.net/download/TCGA.HNSC.sampleMap/HiSeqV2.gz.
Authors' contributions
PS and TYZ conducted the experiments. PS designed
the study. SYW analyzed data. PS and SYW drafted the manuscript.
All authors read and approved the final manuscript.
Ethics approval and consent to
participate
The present study was approved by the Institutional
Review Board of the Chinese People's Liberation Army General
Hospital (Beijing, China). Written informed consent was obtained
from all subjects.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Rastogi B, Kumar A, Raut SK, Panda NK,
Rattan V, Joshi N and Khullar M: Downregulation of miR-377 promotes
oral squamous cell carcinoma growth and migration by targeting
HDAC9. Cancer Investigation. 35:152–162. 2017.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Warnakulasuriya S: Causes of oral
cancer-an appraisal of controversies. Br Dent J. 207:471–475.
2009.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Gharat SA, Momin M and Bhavsar C: Oral
squamous cell carcinoma: Current treatment strategies and
nanotechnology-based approaches for prevention and therapy. Crit
Rev Ther Drug Carrier Syst. 33:363–400. 2016.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Pollaers K, Hinton-Bayre A, Friedland PL
and Farah CS: AJCC 8th edition oral cavity squamous cell carcinoma
Staging-is it an improvement on the AJCC 7th edition? Oral Oncol.
82:23–28. 2018.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Lambert R, Sauvaget C, de Camargo Cancela
M and Sankaranarayanan R: Epidemiology of cancer from the oral
cavity and oropharynx. Eur J Gastroenterol Hepatol. 23:633–641.
2011.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Yokoyama T and Nakamura T: Tribbles in
disease: Signaling pathways important for cellular function and
neoplastic transformation. Cancer Sci. 102:1115–1122.
2011.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Li K, Wang F, Cao WB, Lv XX, Hua F, Cui B,
Yu JJ, Zhang XW, Shang S, Liu SS, et al: TRIB3 promotes APL
progression through stabilization of the oncoprotein PML-RARα and
inhibition of p53-mediated senescence. Cancer Cell. 31:697–710.e7.
2017.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Miyoshi N, Ishii H, Mimori K, Takatsuno Y,
Kim H, Hirose H, Sekimoto M, Doki Y and Mori M: Abnormal expression
of TRIB3 in colorectal cancer: A novel marker for prognosis. Br J
Cancer. 101:1664–1670. 2009.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Izrailit J, Jaiswal A, Zheng W, Moran MF
and Reedijk M: Cellular stress induces TRB3/USP9x-dependent Notch
activation in cancer. Oncogene. 36:1048–1057. 2017.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Hua F, Mu R, Liu J, Xue J, Wang Z, Lin H,
Yang H, Chen X and Hu Z: TRB3 interacts with SMAD3 promoting tumor
cell migration and invasion. J Cell Sci. 124:3235–3246.
2011.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Wennemers M, Bussink J, Scheijen B,
Nagtegaal ID, van Laarhoven HW, Raleigh JA, Varia MA, Heuvel JJ,
Rouschop KM, Sweep FC and Span PN: Tribbles homolog 3 denotes a
poor prognosis in breast cancer and is involved in hypoxia
response. Breast Cancer Res. 13(R82)2011.PubMed/NCBI View
Article : Google Scholar
|
|
13
|
Zhou H, Luo Y, Chen JH, Hu J, Luo YZ, Wang
W, Zeng Y and Xiao L: Knockdown of TRB3 induces apoptosis in human
lung adenocarcinoma cells through regulation of Notch 1 expression.
Mol Med Rep. 8:47–52. 2013.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Hua F, Shang S, Yang YW, Zhang HZ, Xu TL,
Yu JJ, Zhou DD, Cui B, Li K, Lv XX, et al: TRIB3 interacts With
β-catenin and TCF4 to increase stem cell features of colorectal
cancer stem cells and tumorigenesis. Gastroenterology.
156:708–721.e15. 2019.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Hong B, Zhou J, Ma K, Zhang J, Xie H,
Zhang K, Li L, Cai L, Zhang N, Zhang Z and Gong K: TRIB3 promotes
the proliferation and invasion of renal cell carcinoma cells via
activating mapk signaling pathway. Int J Biol Sci. 15:587–597.
2019.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Song M, Bode AM, Dong Z and Lee MH: AKT as
a therapeutic target for cancer. Cancer Res. 79:1019–1031.
2019.PubMed/NCBI View Article : Google Scholar
|
|
17
|
http://tcga.xenahubs.net/download/TCGA.HNSC.sampleMap/HiSeqV2.gz.
|
|
18
|
Wang Y, Liu DP, Chen PP, Koeffler HP, Tong
XJ and Xie D: Involvement of IFN regulatory factor (IRF)-1 and
IRF-2 in the formation and progression of human esophageal cancers.
Cancer Res. 67:2535–2543. 2007.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Vatandoost J and Kafi Sani K: A Study of
recombinant Factor IX in drosophila insect S2 cell lines through
transient gene expression technology. Avicenna J Med Biotechnol.
10:265–268. 2018.PubMed/NCBI
|
|
20
|
Simpson DR, Mell LK and Cohen EE:
Targeting the PI3K/AKT/mTOR pathway in squamous cell carcinoma of
the head and neck. Oral Oncol. 51:291–298. 2015.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Zhang J, Wen HJ, Guo ZM, Zeng MS, Li MZ,
Jiang YE, He XG and Sun CZ: TRB3 overexpression due to endoplasmic
reticulum stress inhibits AKT kinase activation of tongue squamous
cell carcinoma. Oral Oncol. 47:934–939. 2011.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Wennemers M, Bussink J, Grebenchtchikov N,
Sweep FC and Span PN: TRIB3 protein denotes a good prognosis in
breast cancer patients and is associated with hypoxia sensitivity.
Radiother Oncol. 101:198–202. 2011.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Revathidevi S and Munirajan AK: Akt in
cancer: Mediator and more. Semin Cancer Biol. 59:80–91.
2019.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Ikink GJ, Boer M, Bakker ER and Hilkens J:
IRS4 induces mammary tumorigenesis and confers resistance to
HER2-targeted therapy through constitutive PI3K/AKT-pathway
hyperactivation. Nat Commun. 7(13567)2016.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Ediriweera MK, Tennekoon KH and Samarakoon
SR: Role of the PI3K/AKT/mTOR signaling pathway in ovarian cancer:
Biological and Therapeutic Significance. Semin Cancer Biol.
59:147–160. 2019.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Restelli M, Magni M, Ruscica V, Pinciroli
P, De Cecco L, Buscemi G, Delia D and Zannini L: A novel crosstalk
between CCAR2 and AKT pathway in the regulation of cancer cell
proliferation. Cell Death Dis. 7(e2453)2016.PubMed/NCBI View Article : Google Scholar
|