|
1
|
Margueron R and Reinberg D: Chromatin
structure and the inheritance of epigenetic information. Nat Rev
Genet. 11:285–296. 2010.PubMed/NCBI View
Article : Google Scholar
|
|
2
|
Ryšlavá H, Doubnerová V, Kavan D and Vaněk
O: Effect of posttranslational modifications on enzyme function and
assembly. J Proteomics. 92:80–109. 2013.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Berger SL, Kouzarides T, Shiekhattar R and
Shilatifard A: An operational definition of epigenetics. Genes Dev.
23:781–783. 2009.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Waddington CH: Towards a Theoretical
Biology. Edinburgh University Press, Edinburgh, 1968.
|
|
5
|
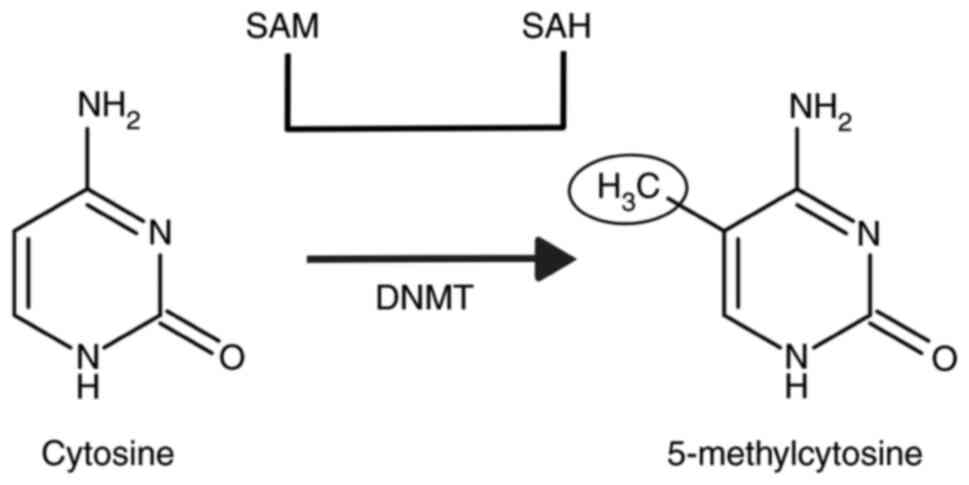
Jeltsch A: On the enzymatic properties of
Dnmt1: Specificity, processivity, mechanism of linear diffusion and
allosteric regulation of the enzyme. Epigenetics. 1:63–66.
2006.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Fischer A, Sananbenesi F, Mungenast A and
Tsai LH: Targeting the correct HDAC(s) to treat cognitive
disorders. Trends Pharmacol Sci. 31:605–617. 2010.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Zhu J, He F, Hu S and Yu J: On the nature
of human housekeeping genes. Trends Genet. 24:481–484.
2008.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Bibikova M: Chapter 2. DNA Methylation.
Microarrays. In: Epigenomics in Health and Disease. Fraga MF and
Fernández AF (eds). Academic Press, Boston, MA, pp19-46, 2016.
|
|
9
|
Goll MG and Bestor TH: Eukaryotic cytosine
methyltransferases. Annu Rev Biochem. 74:481–514. 2005.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Keil KP and Lein PJ: DNA methylation: A
mechanism linking environmental chemical exposures to risk of
autism spectrum disorders? Environ Epigenet.
2(dvv012)2016.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Mielcarek M, Zielonka D, Carnemolla A,
Marcinkowski JT and Guidez F: HDAC4 as a potential therapeutic
target in neurodegenerative diseases: A summary of recent
achievements. Front Cell Neurosci. 9(42)2015.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Saha RN and Pahan K: HATs and HDACs in
neurodegeneration: A tale of disconcerted acetylation homeostasis.
Cell Death Differ. 13:539–550. 2006.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Peserico A and Simone C: Physical and
functional HAT/HDAC interplay regulates protein acetylation
balance. J Biomed Biotechnol. 2011(371832)2011.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Varela MA, Roberts TC and Wood MJ:
Epigenetics and ncRNAs in brain function and disease: Mechanisms
and prospects for therapy. Neurotherapeutics. 10:621–631.
2013.PubMed/NCBI View Article : Google Scholar
|
|
15
|
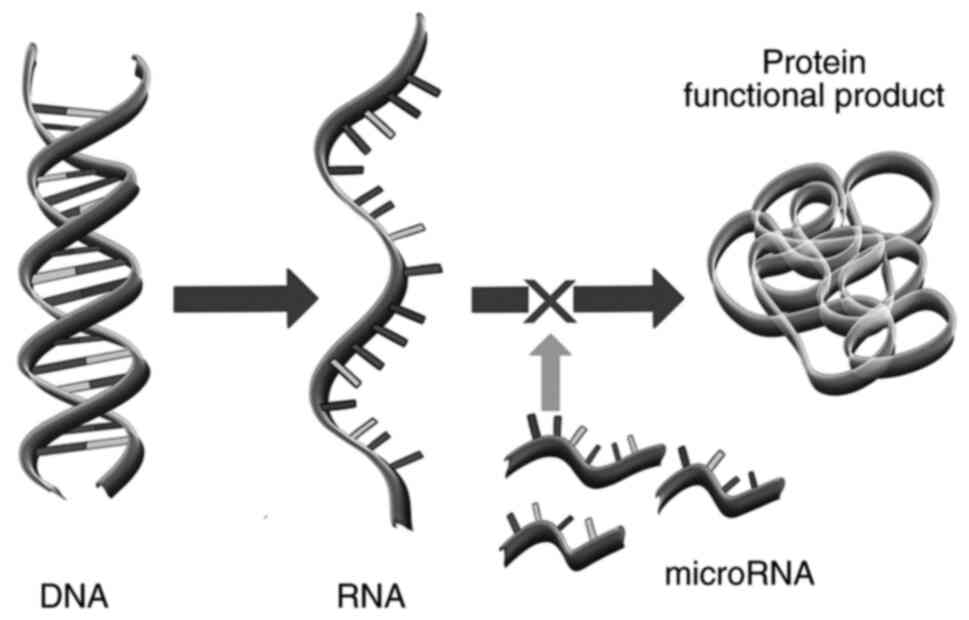
Kim VN: MicroRNA biogenesis: Coordinated
cropping and dicing. Nat Rev Mol Cell Biol. 6:376–385.
2005.PubMed/NCBI View
Article : Google Scholar
|
|
16
|
Pasquinelli AE: MicroRNAs and their
targets: Recognition, regulation and an emerging reciprocal
relationship. Nat Rev Genet. 13:271–282. 2012.PubMed/NCBI View
Article : Google Scholar
|
|
17
|
Isac S, Panaitescu AM, Iesanu MI, Zeca V,
Cucu N, Zagrean L, Peltecu G and Zagrean AM: Maternal
citicoline-supplemented diet improves the response of the immature
hippocampus to perinatal asphyxia in rats. Neonatology.
117:729–735. 2020.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Isac T, Isac S, Ioanitescu S, Mihaly E,
Tanasescu MD, Balan DG, Tulin A and Iliescu L: Dynamics of serum
α-fetoprotein in viral hepatitis C without hepatocellular
carcinoma. Exp Ther Med. 22(749)2021.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Wu K, Ye C, Lin L, Chu Y, Ji M, Dai W,
Zeng X and Lin Y: Inhibiting miR-21 attenuates experimental hepatic
fibrosis by suppressing both the ERK1 pathway in HSC and hepatocyte
EMT. Clin Sci (Lond). 130:1469–1480. 2016.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Popescu I, Ionescu M, Braşoveanu V,
Hrehoreţ D, Copca N, Lupaşcu C, Botea F, Dorobanţu B, Alexandrescu
S, Grigorie M, et al: The Romanian National program for liver
transplantation -852 procedures in 815 patients over 17 years
(2000-2017): A continuous evolution to success. Chirurgia (Bucur).
112:229–243. 2017.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Ahrens M, Ammerpohl O, von Schönfels W,
Kolarova J, Bens S, Itzel T, Teufel A, Herrmann A, Brosch M,
Hinrichsen H, et al: DNA methylation analysis in nonalcoholic fatty
liver disease suggests distinct disease-specific and remodeling
signatures after bariatric surgery. Cell Metab. 18:296–302.
2013.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Cao Y, Xue Y, Xue L, Jiang X, Wang X,
Zhang Z, Yang J, Lu J, Zhang C, Wang W and Ning G: Hepatic menin
recruits SIRT1 to control liver steatosis through histone
deacetylation. J Hepatol. 59:1299–1306. 2013.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Nakahata Y, Kaluzova M, Grimaldi B, Sahar
S, Hirayama J, Chen D, Guarente LP and Sassone-Corsi P: The
NAD+-dependent deacetylase SIRT1 modulates CLOCK-mediated chromatin
remodeling and circadian control. Cell. 134:329–340.
2008.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Xu F and Guo W: The progress of
epigenetics in the development and progression of non-alcoholic
fatty liver disease. Liver Res. 4:118–123. 2020.
|
|
25
|
Lane EA, Choi DW, Garcia-Haro L, Levine
ZG, Tedoldi M, Walker S and Danial NN: HCF-1 regulates de novo
lipogenesis through a nutrient-sensitive complex with ChREBP. Mol
Cell 25:. 75:357–371.e7. 2019.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Hsu SH, Wang B, Kota J, Yu J, Costinean S,
Kutay H, Yu L, Bai S, La Perle K, Chivukula RR, et al: Essential
metabolic, anti-inflammatory, and anti-tumorigenic functions of
miR-122 in liver. J Clin Invest. 122:2871–2883. 2012.PubMed/NCBI View
Article : Google Scholar
|
|
27
|
Calo N, Ramadori P, Sobolewski C, Romero
Y, Maeder C, Fournier M, Rantakari P, Zhang FP, Poutanen M, Dufour
JF, et al: Stress-activated miR-21/miR-21* in hepatocytes promotes
lipid and glucose metabolic disorders associated with high-fat diet
consumption. Gut. 65:1871–1881. 2016.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Liu X, Zhao J, Liu Q, Xiong X, Zhang Z,
Jiao Y, Li X, Liu B, Li Y and Lu Y: MicroRNA-124 promotes hepatic
triglyceride accumulation through targeting tribbles homolog 3. Sci
Rep. 6(37170)2016.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Ying W, Riopel M, Bandyopadhyay G, Dong Y,
Birmingham A, Seo JB, Ofrecio JM, Wollam J, Hernandez-Carretero A,
Fu W, et al: Adipose tissue macrophage-derived exosomal miRNAs can
modulate in vivo and in vitro insulin sensitivity. Cell.
171:372–384.e12. 2017.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Kerkar N and Vergani D: De novo autoimmune
hepatitis -is this different in adults compared to children? J
Autoimmun. 95:26–33. 2018.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Behairy BE, El-Araby HA, Abd El Kader HH,
Ehsan NA, Salem ME, Zakaria HM and Khedr MA: Assessment of
intrahepatic regulatory T cells in children with autoimmune
hepatitis. Ann Hepatol. 15:682–690. 2016.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Bandiera S, Pfeffer S, Baumert TF and
Zeisel MB: MiR-122-a key factor and therapeutic target in liver
disease. J Hepatol. 62:448–457. 2015.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Migita K, Komori A, Kozuru H, Jiuchi Y,
Nakamura M, Yasunami M, Furukawa H, Abiru S, Yamasaki K, Nagaoka S,
et al: Circulating microRNA profiles in patients with type-1
autoimmune hepatitis. PLoS One. 10(e0136908)2015.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Chen L, Lu FB, Chen DZ, Wu JL, Hu ED, Xu
LM, Zheng MH, Li H, Huang Y, Jin XY, et al: BMSCs-derived
miR-223-containing exosomes contribute to liver protection in
experimental autoimmune hepatitis. Mol Immunol. 93:38–46.
2018.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Xia G, Wu S, Wang X and Fu M: Inhibition
of microRNA-155 attenuates concanavalin-A-induced autoimmune
hepatitis by regulating Treg/Th17 cell differentiation. Can J
Physiol Pharmacol. 96:1293–1300. 2018.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Liu Q, Li Y, Xiong M and Tang R:
Epigenetics of autoimmune liver diseases: Current progress and
future directions. J Bio-X Res. 2:46–55. 2019.
|
|
37
|
Chen M, Shirai M, Czaja AJ, Kurokohchi K,
Arichi T, Arima K, Kodama T and Nishioka M: Characterization of
anti-histone antibodies in patients with type 1 autoimmune
hepatitis. J Gastroenterol Hepatol. 13:483–489. 1998.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Zachou K, Arvaniti P, Lyberopoulou A,
Sevdali E, Speletas M, Ioannou M, Koukoulis GK, Renaudineau Y and
Dalekos GN: Altered DNA methylation pattern characterizes the
peripheral immune cells of patients with autoimmune hepatitis.
Liver Int: Feb 2, 2022 (Epub ahead of print).
|
|
39
|
Nehme Z, Pasquereau S and Herbein G:
Control of viral infections by epigenetic-targeted therapy. Clin
Epigenetics. 11(55)2019.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Jain S, Chang TT, Chen S, Boldbaatar B,
Clemens A, Lin SY, Yan R, Hu CT, Guo H, Block TM, et al:
Comprehensive DNA methylation analysis of hepatitis B virus genome
in infected liver tissues. Sci Rep. 5(10478)2015.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Zhang Y, Li C, Zhang Y, Zhu H, Kang Y, Liu
H, Wang J, Qin Y, Mao R, Xie Y, et al: Comparative analysis of CpG
islands among HBV genotypes. PLoS One. 8(e56711)2013.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Hong X, Kim ES and Guo H: Epigenetic
regulation of hepatitis B virus covalently closed circular DNA:
Implications for epigenetic therapy against chronic hepatitis B.
Hepatology. 66:2066–2077. 2017.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Pollicino T, Belloni L, Raffa G, Pediconi
N, Squadrito G, Raimondo G and Levrero M: Hepatitis B virus
replication is regulated by the acetylation status of hepatitis B
virus cccDNA-bound H3 and H4 histones. Gastroenterology.
130:823–837. 2006.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Wang TZ, Lin DD, Jin BX, Sun XY and Li N:
Plasma microRNA: A novel non-invasive biomarker for HBV-associated
liver fibrosis staging. Exp Ther Med. 17:1919–1929. 2019.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Perez S, Kaspi A, Domovitz T, Davidovich
A, Lavi-Itzkovitz A, Meirson T, Alison Holmes J, Dai CY, Huang CF,
Chung RT, et al: Hepatitis C virus leaves an epigenetic signature
post cure of infection by direct-acting antivirals. PLoS Genet.
15(e1008181)2019.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Zheng Y, Hlady RA, Joyce BT, Robertson KD,
He C, Nannini DR, Kibbe WA, Achenbach CJ, Murphy RL, Roberts LR and
Hou L: DNA methylation of individual repetitive elements in
hepatitis C virus infection-induced hepatocellular carcinoma. Clin
Epigenetics. 11(145)2019.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Weis A, Marquart L, Calvopina DA, Genz B,
Ramm GA and Skoien R: Serum MicroRNAs as biomarkers in hepatitis C:
Preliminary evidence of a MicroRNA panel for the diagnosis of
hepatocellular carcinoma. Int J Mol Sci. 20(864)2019.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Shrivastava S, Petrone J, Steele R, Lauer
GM, Di Bisceglie AM and Ray RB: Up-regulation of circulating
miR-20a is correlated with hepatitis C virus-mediated liver disease
progression. Hepatology. 58:863–871. 2013.PubMed/NCBI View Article : Google Scholar
|
|
49
|
Matsuura K, De Giorgi V, Schechterly C,
Wang RY, Farci P, Tanaka Y and Alter HJ: Circulating let-7 levels
in plasma and extracellular vesicles correlate with hepatic
fibrosis progression in chronic hepatitis C. Hepatology.
64:732–745. 2016.PubMed/NCBI View Article : Google Scholar
|
|
50
|
El-Diwany R, Wasilewski LN, Witwer KW,
Bailey JR, Page K, Ray SC, Cox AL, Thomas DL and Balagopal A: Acute
hepatitis C virus infection induces consistent changes in
circulating MicroRNAs that are associated with nonlytic hepatocyte
release. J Virol. 89:9454–9464. 2015.PubMed/NCBI View Article : Google Scholar
|
|
51
|
Liu XL, Pan Q, Zhang RN, Shen F, Yan SY,
Sun C, Xu ZJ, Chen YW and Fan JG: Disease-specific miR-34a as
diagnostic marker of non-alcoholic steatohepatitis in a Chinese
population. World J Gastroenterol. 22:9844–9852. 2016.PubMed/NCBI View Article : Google Scholar
|
|
52
|
Isac S, Panaitescu AM, Iesanu M, Grigoras
IF, Totan A, Udriste A, Cucu N, Peltecu G, Zagrean L and Zagrean
AM: Maternal high-fat diet modifies the immature hippocampus
vulnerability to perinatal asphyxia in rats. Neonatology.
114:355–361. 2018.PubMed/NCBI View Article : Google Scholar
|
|
53
|
Lieber CS, Leo MA, Wang X and Decarli LM:
Effect of chronic alcohol consumption on Hepatic SIRT1 and
PGC-1alpha in rats. Biochem Biophys Res Commun. 370:44–48.
2008.PubMed/NCBI View Article : Google Scholar
|
|
54
|
Lee YJ and Shukla SD: Histone H3
phosphorylation at serine 10 and serine 28 is mediated by p38 MAPK
in rat hepatocytes exposed to ethanol and acetaldehyde. Eur J
Pharmacol. 573:29–38. 2007.PubMed/NCBI View Article : Google Scholar
|
|
55
|
Pal-Bhadra M, Bhadra U, Jackson DE,
Mamatha L, Park PH and Shukla SD: Distinct methylation patterns in
histone H3 at Lys-4 and Lys-9 correlate with up- &
down-regulation of genes by ethanol in hepatocytes. Life Sci.
81:979–987. 2007.PubMed/NCBI View Article : Google Scholar
|
|
56
|
Lu SC, Huang ZZ, Yang H, Mato JM, Avila MA
and Tsukamoto H: Changes in methionine adenosyltransferase and
S-adenosylmethionine homeostasis in alcoholic rat liver. Am J
Physiol Gastrointest Liver Physiol. 279:G178–G185. 2000.PubMed/NCBI View Article : Google Scholar
|
|
57
|
Dannenberg LO, Chen HJ, Tian H and
Edenberg HJ: Differential regulation of the alcohol dehydrogenase
1B (ADH1B) and ADH1C genes by DNA methylation and histone
deacetylation. Alcohol Clin Exp Res. 30:928–937. 2006.PubMed/NCBI View Article : Google Scholar
|
|
58
|
Mandrekar P: Epigenetic regulation in
alcoholic liver disease. World J Gastroenterol. 17:2456–2464.
2011.PubMed/NCBI View Article : Google Scholar
|
|
59
|
Bala S, Marcos M, Kodys K, Csak T,
Catalano D, Mandrekar P and Szabo G: Up-regulation of microRNA-155
in macrophages contributes to increased tumor necrosis factor
{alpha} (TNF{alpha}) production via increased mRNA half-life in
alcoholic liver disease. J Biol Chem. 286:1436–1444.
2011.PubMed/NCBI View Article : Google Scholar
|
|
60
|
Tang Y, Banan A, Forsyth CB, Fields JZ,
Lau CK, Zhang LJ and Keshavarzian A: Effect of alcohol on miR-212
expression in intestinal epithelial cells and its potential role in
alcoholic liver disease. Alcohol Clin Exp Res. 32:355–364.
2008.PubMed/NCBI View Article : Google Scholar
|
|
61
|
Medici V and LaSalle JM: Genetics and
epigenetic factors of Wilson disease. Ann Transl Med. 7 (Suppl
2)(S58)2019.PubMed/NCBI View Article : Google Scholar
|
|
62
|
Li M, Li Y, Chen J, Wei W, Pan X, Liu J,
Liu Q, Leu W, Zhang L, Yang X, et al: Copper ions inhibit
S-adenosylhomocysteine hydrolase by causing dissociation of NAD+
cofactor. Biochemistry. 46:11451–11458. 2007.PubMed/NCBI View Article : Google Scholar
|
|
63
|
Yin R, Mo J, Dai J and Wang H: Nickel(II)
inhibits tet-mediated 5-methylcytosine oxidation by high affinity
displacement of the cofactor iron(II). ACS Chem Biol. 12:1494–1498.
2017.PubMed/NCBI View Article : Google Scholar
|
|
64
|
Le A, Shibata NM, French SW, Kim K,
Kharbanda KK, Islam MS, LaSalle JM, Halsted CH, Keen CL and Medici
V: Characterization of timed changes in hepatic copper
concentrations, methionine metabolism, gene expression, and global
DNA methylation in the Jackson toxic milk mouse model of Wilson
disease. Int J Mol Sci. 15:8004–8023. 2014.PubMed/NCBI View Article : Google Scholar
|
|
65
|
Medici V, Kieffer DA, Shibata NM, Chima H,
Kim K, Canovas A, Medrano JF, Islas-Trejo AD, Kharbanda KK, Olson
K, et al: Wilson disease: Epigenetic effects of choline
supplementation on phenotype and clinical course in a mouse model.
Epigenetics. 11:804–818. 2016.PubMed/NCBI View Article : Google Scholar
|