Introduction
Lung cancer is one of the malignant tumors with the
highest incidence and mortality rate worldwide (1). Globally, the number of deaths caused
by lung cancer accounts for ~30% of all cancer-associated deaths.
In China, lung cancer has the highest incidence and mortality rate
among all tumors. The primary treatment for lung cancer is surgical
intervention, but with the advancement of medical technology,
various treatment methods are emerging. These methods range from
conventional chemotherapy to molecular targeted therapies and
immunotherapy, such as monoclonal antibodies against EGFR,
anaplastic lymphoma kinase (ALK), VEGFR and programmed cell death
protein 1/programmed cell death ligand 1 (2,3).
Consequently, the treatment of lung cancer is gradually
transitioning into the era of precision medicine.
The gene mutation rate in cancerous lung tissues is
higher than that of adjacent tissues, indicating that gene
mutations could lead to the occurrence of lung cancer and increase
its incidence (4,5). Early detection of high calmodulin 1
expression and EGFR mutations can be used to differentiate the
diagnosis of patients with colorectal cancer, which is conducive to
further prolonging the survival period and improving the quality of
life of patients (6). Identifying
the p53 Arg72Pro mutation in patients with primary hepatocellular
carcinoma is important for the early diagnosis of cancer in
patients with chronic hepatitis B-associated cirrhosis (7). Thus, identifying new gene mutation
sites and searching for novel targeted therapeutic targets are
necessary in the era of precision medicine. The discovery and
determination of the role of specific genes in the development of
lung cancer is critical in obtaining ideal, effective and precise
therapeutic targets (8).
Therefore, at present, gene research on lung adenocarcinoma
represents a significant focus. EGFR, KRAS, ALK, c-MET, and other
genes have been implicated in the progression of lung
adenocarcinoma (9), guiding its
clinical applications. However, several patients develop drug
resistance and gene mutations following targeted therapy for lung
adenocarcinoma (10).
23-Hydroxybetulinic acid (23-HBA) has been shown to promote the
apoptosis of tumor cells by inhibiting the proliferation of
vascular endothelial cells (11).
Hence, genetic changes related to lung adenocarcinoma and the
inhibitory effects of novel compounds must be further explored
(12).
23-HBA is a triterpenoid compound isolated from
Pulanemone (13), a traditional
Chinese medicine used for heat clearance and detoxification
(14). 23-HBA has been
demonstrated to reduce the mitochondrial membrane potential and
induce apoptosis in melanoma cells (B16), and activate the p38 and
JNK MAPK pathway-associated kinases in colon cancer cells (LoVo)
(15). The loss of mitochondrial
membrane potential in LoVo cells can trigger apoptosis through the
mitochondrial pathway (16).
Moreover, angiogenesis can be hindered by blocking human
microvascular endothelial cells (HMECs) in the S phase (17), restraining HMEC growth and inducing
apoptosis. 23-HBA exhibits a synergistic effect with antitumor
drugs such as 5-fluorouracil (18). It can also inhibit the activity of
the ATP binding cassette subfamily B member 1 ATPase (19), leading to the accumulation of drugs
(20) within cells and effectively
reversing P-glycoprotein-mediated multidrug resistance. In
addition, 23-HBA can induce beclin-1-dependent autophagy and
apoptosis (21) in HL-60 cells.
23-HBA has been structurally modified to enhance its antitumor
efficacy and bioavailability, resulting in >300 derivatives.
Some of these derivatives have shown improved antitumor activity
(22). 23-HBA can inhibit the
activation of NF-κB p65 and the MAPK pathway, and reduce the
secretion of inflammatory factors in mice with DSS-induced acute
ulcerative colitis (23).
Furthermore, 23-HBA has been shown to reduce tumorigenesis,
metastasis and immunosuppression in mouse models of hepatocellular
carcinoma by disrupting MAPK signaling pathways (20).
In the present study, the aim of this study is to
investigate the underlying mechanisms of action of Pulsatilla
compounds against lung adenocarcinoma. The Pulsatilla compounds
were combined with lung adenocarcinoma, and databases, such as
SwissTargetPrediction, were utilized to demonstrate a robust
association between the target genes of 23-HBA and the pathogenesis
of lung adenocarcinoma (24). MTT
and flow cytometry assays were conducted to validate the effect of
23-HBA on lung adenocarcinoma cells (20). Target genes of 23-HBA in lung
adenocarcinoma were identified by bioinformatics analysis and
characterized to test their functions. The current study introduces
novel perspectives for the treatment of patients with lung
adenocarcinoma (25).
Materials and methods
Cell line culture
The human lung adenocarcinoma cell lines A549, H1975
and H1299, as well as the human bronchial epithelial cell line
BEAS-2B, were used in the present study. These cell lines were
provided by The Cell of Bank of Type Culture Collection of the
Chinese Academy of Sciences. The cell lines used in the presented
study were tested for mycoplasma contamination, ensuring that the
experiments were conducted with uncontaminated cell cultures. Cells
were cultured in DMEM High Glucose (Hyclone; Cytiva) with 10% FBS
(Gibco; Thermo Fisher Scientific, Inc.). The cells were cultivated
in a cell incubator at 37˚C in a humidified atmosphere with 5%
CO2.
Antibodies and drugs
Rabbit anti-human GAPDH antibody (1:6,000 dilution;
cat. no. AP0063) was purchased from Bioworld Technology, Inc.
Peroxisome proliferator-activated receptor (PPAR)-γ rabbit
polyclonal antibody (1:1,000 dilution; cat. no. 16643-1-AP) was
purchased from Proteintech Group, Inc. HRP-conjugated goat
anti-rabbit IgG (1:6,000 dilution; cat. no. BS13278) was purchased
from Bioworld Technology, Inc. and was used as a secondary antibody
(26).
23-HBA (Shanghai Aladdin Biochemical Technology Co.,
Ltd.) was used in the present study. It is known by the chemical
names is ‘(3β)-3,23-Dihydroxylup-20(29)-en-28-oic acid’ or
‘(1R,3aS,5aR,5bR,7aR,9S,11aR,11bR,13aS,13bR)-9-hydroxy-8-(hydroxymethyl)-5a,5b,8,11a-tetramethyl-1-prop-1-en-2-yl-1,2,3,4,5,6,7,7a,9,10,11,11b,12,13,13a,13b-hexadecahydrocyclopenta[a]chrysene-3a-carboxylic
acid’. The molecular weight of 23-HBA is 472.7 g/mol, while it has
a melting point of 305˚C. 23-HBA is soluble in dimethyl sulfoxide
(DMSO), a commonly used solvent in the laboratory, at a
concentration of 2 mg/ml. 23-HBA is sensitive to light, humidity
and heat, and its sensitivity is an important point to consider
during its handling and storage (27).
Data sources and processing
A total of >8,000 lung adenocarcinoma-related
genes were screened through the DisGeNET (www.disgenet.org) and GeneCards (www.genecards.org) databases (28). Subsequently, the 23-HBA drug
targets were identified through the Traditional Chinese Medicine
Systems Pharmacology (TCMSP; old.tcmsp-e.com/tcmsp.p) and SwissTargetPrediction
(www.swisstargetprediction.ch)
databases. The 23-HBA potential targets and lung
adenocarcinoma-related genes were mapped using Venny tool version
2.1.0 (bioinfogp.cnb.csic.es/tools/venny/). The total number
of common targets was analyzed.
MTT assay for verification of the
effect of 23-HBA on lung adenocarcinoma cells (29)
H1299 cells in the logarithmic growth phase were
inoculated into 96-well plates, with 2,000 cells placed in each
well. After 16-18 h, the cells completely attached to the wells,
and were divided into two groups: Experimental group (drug group)
and control group (treated with DMSO). Several drug doses (0, 40,
60, 80 and 100 µM) were examined to determine the optimal
concentration for subsequent experiments. The cells were cultured
with 23-HBA at 37˚C for 48 h. Subsequently, the supernatant was
discarded, and 90 µl RPMI-1640 complete medium (Gibco; Thermo
Fisher Scientific, Inc.) with 10 µl MTT solution were added to each
well. The plates were incubated at 37˚C in the dark for 4 h, before
the supernatant was discarded. Subsequently, 150 µl DMSO (Sigma
Corporation) was added and gently mixed to dissolve the formazan
crystals. An ELISA microplate detector (Biosharp Life Sciences) was
used to measure the optical density values at a wavelength of 491
nm.
Cell cycle analysis by flow
cytometry
H1299 cells in the logarithmic growth phase were
inoculated at a concentration of 1x105 cells/well in a
6-well plate. After 16-18 h of culture, the cells were exposed to
varying concentrations (0, 20, 40 and 60 µM) of 23-HBA. Following
48 h of treatment (37˚C), the supernatant was collected in a 2.0 ml
Eppendorf tube. For cell collection, 300 µl trypsin (Sigma-Aldrich;
Merck KGaA) was added to the cells and incubated at 37˚C for 3 min.
When the cells became spherical in shape, an equivalent amount of
RPMI-1640 complete was added to terminate digestion. The suspension
was centrifuged (37˚C) at 300 x g, and the pellet was collected
after 5 min. The single-cell suspension was washed with PBS
(OriGene Technologies, Inc.) and then subjected to centrifugation
under the same conditions. Subsequently, the suspension was fixed
(4˚C) in 75% pre-cooled ethanol (500 µl) for a duration of 2 h to
overnight (12 h), and was stored at 4˚C. Prior to staining, the
fixed cells were washed with PBS. RNase A solution (100 µl) was
added to the cell pellet, and the cells were incubated in a water
bath at 37˚C for 30 min. Subsequently, 400 µl propidium iodide
(Sigma-Aldrich; Merck KGaA) staining solution was added and
thoroughly mixed, and the samples were incubated in the dark at 4˚C
for 30 min. The cell cycle was determined using a flow cytometer
(BD Biosciences) (30) and
analyzed using FlowJo version 10.6.2 (Tree Star).
Protein-protein interaction (PPI)
network
The targets associated with the pathogenesis of lung
adenocarcinoma and the 23-HBA drug targets were imported into the
STRING database (https://string-db.org/) to generate the PPI network of
overlapping targets. Subsequently, the overlapping targets were
imported into Cytoscape software version is 3.3 (https://cytoscape.org/) for visual analysis. Five core
targets with a high degree of association between 23-HBA and lung
adenocarcinoma were identified using the betweenness centrality
(BC; the degree to which nodes are independent of each other) size
sequencing (31), and the
characteristics were analyzed. The characteristic is mainly
manifested in: ACE mainly exists in vascular endothelial cells,
while tumor cells have no blood vessels. Research on PTSG2 has been
relatively clear and specific. MDM2 is highly expressed in tumors
and has been proven to bind to p53. H1299 cells have a homozygous
partial deletion of TP53 and therefore cannot express p53, and
PPAR- and PPAR-γ belong to the same category and need not be
studied. The median degree of freedom of the targets was 7.055, and
the maximum degree of freedom was 18. The inclusion criteria for
screening core targets were set at 15 and 18, and the best five
core targets were selected based on the relevant parameters of each
target (32).
Gene Expression Profiling Interactive
Analysis (GEPIA) database
The mRNA expression profile of PPAR-γ in various
tumor types, including lung adenocarcinoma, was assessed using the
GEPIA database (http://gepia.cancer-pku.cn).
Western blot analysis
Western blot analysis was performed to compare the
expression of PPAR-γ between normal and lung cancer cells and
explore the characteristics of this protein (33). BEAS-2B, A549, H1975 and H1299 cells
with robust growth profiles were collected. These cells were
digested with trypsin, and the cell concentration was adjusted to
1x106 cells/ml. Subsequently, the cells were inoculated
into a 6-well plate (1x106 cells per well) and cultured
overnight. After a further 48-h incubation, the cells were digested
with trypsin, and the cell precipitate was collected through
centrifugation (200 x g, 37˚C, 3 min). RIPA buffer (Beyotime
Institute of Biotechnology) and PMSF (Beijing Solarbio Science
& Technology Co., Ltd.) were used to lyse cells (30 min with a
10 min mixing interval), followed by centrifugation (300 x g, 37˚C,
10 min). The supernatant was collected, and 5X SDS (Xilong
Scientific Co., Ltd.) solution was added to the supernatant at a
1:4 ratio. The mixture was boiled at 95˚C for 10 min and stored at
-20˚C. The protein concentration was determined using the
bicinchoninic acid method. A total of 40 µg protein/lane were
separated using 5% SDS-PAGE transferred to PVDF. The membranes were
incubated with primary anti-GAPDH (rabbit anti-human GAPDH; cat.
no. GTX100118; 1:1,000-5,000 dilution; Bioworld Technology) and
PPAR-γ antibodies (PPAR-γ rabbit polyAb; cat. no. HY-P80872; 5
µg/ml; Protein Technology) at 4˚C for 6 h, followed by secondary
anti-rabbit antibodies (goat anti-rabbit IgG-HRP; cat. no. L27A9; 2
µg/ml; Bioworld Technology) at 25˚C for 30 min. Following washing
with 10% SDS, the membranes were incubated with ECL reagent
(Biosharp Life Sciences) for signal development. The absorbance
values of each band were calculated using ImageJ software (version
64; National Institutes of Health). GAPDH served as the internal
reference. The entire experiment was repeated three times.
Subsequently, H1299 cells were selected for subsequent experiments,
and 23-HBA with various concentrations (0, 20, 40 and 60 µM) was
applied to the cells at 37˚C for 2 h to analyze the expression of
PPAR-γ.
Kyoto Encyclopedia of Genes and
Genomes (KEGG) and Gene Ontology (GO) enrichment analysis
KEGG analysis was performed using the R (version
4.1.2) package ‘Clusterprofiler’ (https://bioconductor.org), and pathways with P<0.05
and an association with cancer were screened (34). This process resulted in the
identification of 12 pathways, which were then visualized and
further screened for pathways exhibiting significant enrichment. It
was visualized by using KEGG analysis; a graph is automatically
generated when a gene is imported into the website for
visualization purposes. GO enrichment analysis was conducted using
the R package ‘Clusterprofiler’, and P<0.05 was used as the
level of statistical significance. The relevant biological
processes (BPs), cellular components (CCs) and molecular functions
(MFs) were analyzed based on their P-value. Processes displaying
high enrichment were selected for subsequent analyses.
Further exploration of the PPAR
pathway-associated mechanism (35)
The target genes enriched in this pathway were
identified through databases, such as KEGG (https://www.kegg.jp), to explore the mechanism of the
PPAR signaling pathway. The mechanism of the PPAR pathway and its
association with reactive oxygen species (ROS) were investigated by
searching sources, such as PubMed (https://www.pubmed.ncbi.nlm.nih.gov) and Web of
Science (https://clarivate.com.cn/solutions/web-of-science/),
to predict the mechanism associated with the PPAR signaling
pathway.
Mitochondrial ROS detection
H1299 cells stably cultured for several generations
were used and diluted to 8x104 cells/ml after digestion.
Of this cell suspension, 1 ml was added to a 6-well plate in
addition to 1 ml of complete culture medium. After observing that
the cells were almost completely attached to the wall, the old
culture medium and was removed and 2 ml of 0, 20, 40 or 60 µmol/l
23-HBA culture medium was added to each well and incubated for 48
h. A ROS detection reagent (Beyotime Institute of Biotechnology)
was employed for mitochondrial ROS detection.
Dichlorodihydrofluorescein diacetate (DCFH-DA) was diluted with
serum-free medium at a ratio of 1:1,000 to achieve a final
concentration of 10 µmol/l. After the cell culture medium was
discarded, an appropriate volume of diluted DCFH-DA was added to
adequately cover the cells. The cells were then incubated in a 37˚C
incubator for 20 min, followed by washing three times with
serum-free medium to fully eliminate the DCFH-DA that did not enter
the cells. Subsequently, the cells were observed directly using a
laser confocal microscope (excitation wavelength, 488 nm; emission
wavelength, 525 nm; Shanghai Tianneng Life Science Co., Ltd.).
Clinical prognosis
The target genes enriched in the PPAR signaling
pathway were obtained through the GEPIA database (36). Kaplan-Meier curve analysis was then
used to examine the effect of these target genes on the survival
rate of patients with lung adenocarcinoma, with an aim to predict
the clinical prognosis of these patients in association with the
expression levels of the PPAR pathway target genes.
Statistical analysis
Data are presented as the mean ± SD of three
experimental repeats. R (version 4.1.2) were utilized for the
statistical analysis and required in the present study. The primary
prognostic value was assessed using Kaplan-Meier curve analysis and
the log-rank test, as well as Cox regression analysis. Other
statistical analyses were performed in GraphPad Prism 6 software
(Dotmatics). Comparisons among multiple groups were performed using
one-way ANOVA followed by Tukey's post-hoc test, while Student's
t-test comparing the experimental group with the control group one
by one was used for comparisons between two groups. P<0.05 was
considered do indicate a statistically significant difference.
Results
Association of 23-HBA with lung
adenocarcinoma
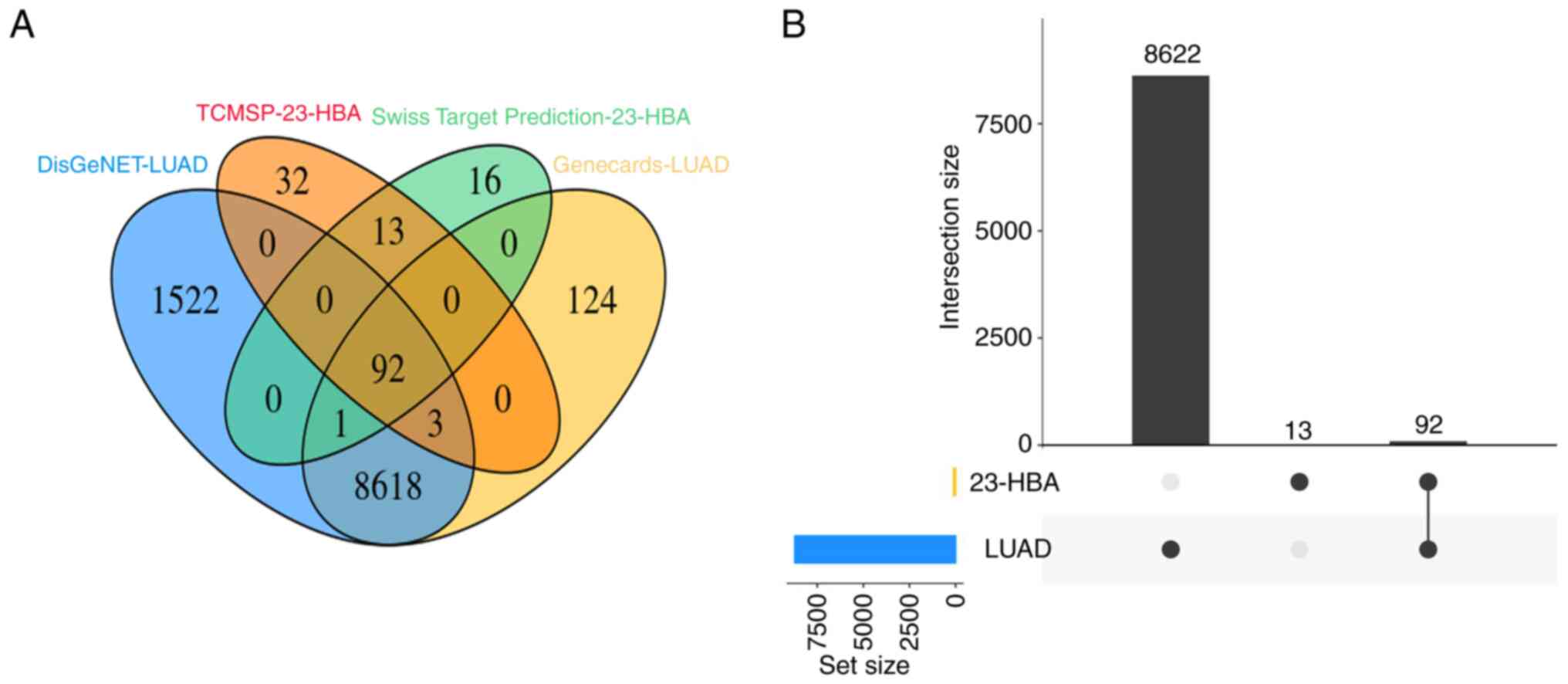
Initially, genes associated with the pathogenesis of
lung adenocarcinoma in the DisGeNET and Genecards databases and
23-HBA target genes in the SwissTargetPrediction and TCMSP
databases were examined. Subsequently, a Venn diagram was
constructed, revealing 92 overlapping genes (Fig. 1A). Further analysis demonstrated
that these overlapping genes constituted 87.6% of the genes
associated with 23-HBA (Fig. 1B).
This finding provides evidence that the target genes of 23-HBA are
closely linked to the pathogenesis of lung adenocarcinoma.
Inhibitory effect of 23-HBA on H1299
lung adenocarcinoma cells
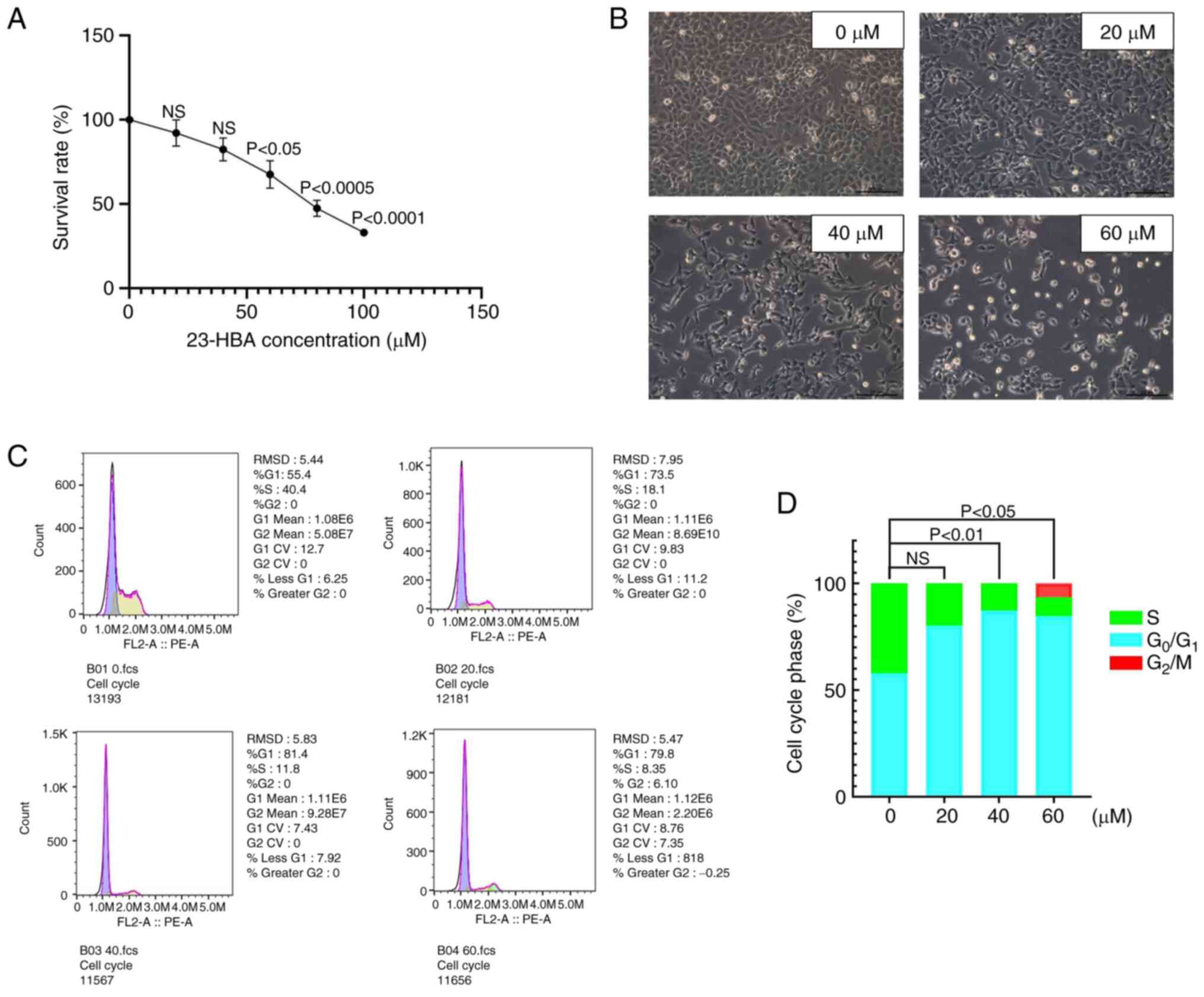
First, the H1299 cell line, which was selected due
to being cost-effective and producing reliable results, was
cultured for MTT assays. The cells were divided into two groups,
the control group and the experimental group, and various drug
concentrations (0, 20, 40, 60, 80 and 100 µM) were used to explore
the IC50 value of 23-HBA. The cell survival rate
exhibited a steady decline with increasing concentrations of 23-HBA
(Fig. 2A). At a drug concentration
of 60 µM, which is close to the IC50 value, the cell
survival rate was 54.4%. This finding provided evidence that 23-HBA
had an inhibitory effect on the viability of H1299 cells.
Therefore, in the follow-up experiments, drug doses between 0-60 µM
were considered to be representative of the effect of 23-HBA.
Microscopic analysis (fluorescence microscope) after treatment of
the cells with 0-60 µM 23-HBA revealed a significant decrease in
the number of adherent cells with an increase in drug
concentration, accompanied by an increase in the number of floating
dying cells (Fig. 2B). Finally,
changes in the cell cycle of H1299 cells treated with various
concentrations of 23-HBA were analyzed using flow cytometry
(Fig. 2C), and the distribution of
cells in each phase of the cell cycle was quantified (Fig. 2D). The proportion of H1299 cells in
the G1 phase notably increased with increasing
concentrations of 23-HBA, and, apart from the 20 µm group, this
difference was statistically significant compared with the control
group. However, fewer cells were in G2 and S phases.
This finding indicated that 23-HBA affects H1299 cells by arresting
the cell cycle in the G1 phase. All these results
collectively demonstrated that 23-HBA inhibited the proliferation
of lung adenocarcinoma H1299 cells.
PPAR-γ is the optimal target for the
action of 23-HBA on lung adenocarcinoma cells
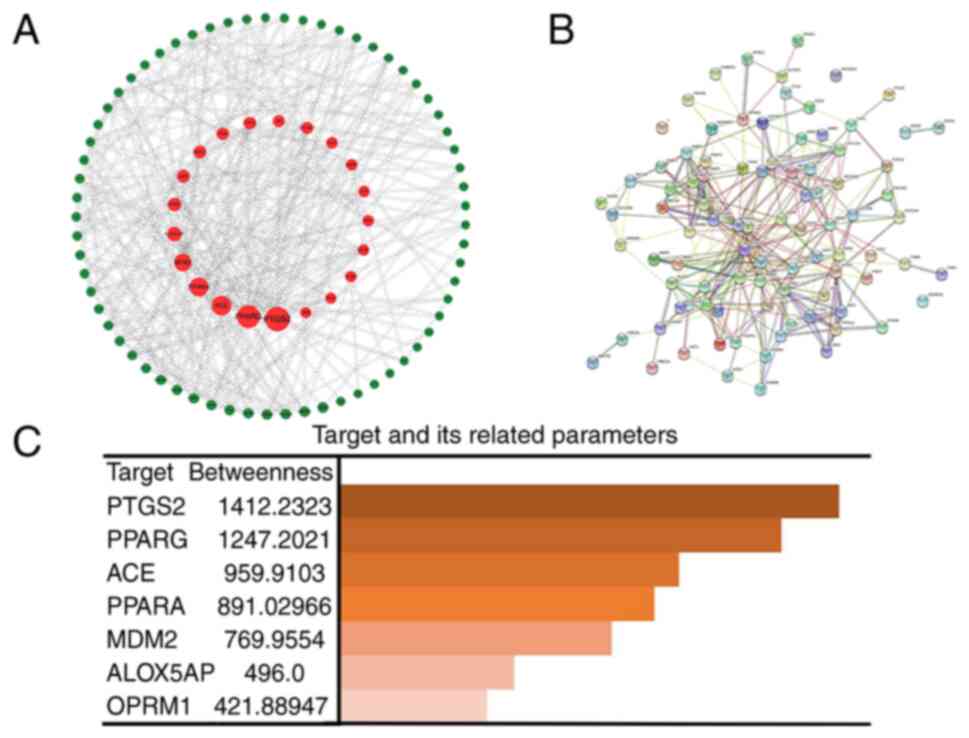
Firstly, the genes from the Venn diagram were
imported into Cytoscape software, and topological parameter
diagrams were created on the basis of the BC association of each
gene for preliminary gene selection (Fig. 3A). Second, the related genes with
lung adenocarcinoma pathogenesis genes and targets of the
23-HBA-associated domain were imported into STRING software to
generate a PPI network diagram (Fig.
3B), which allowed for the analysis of gene associations to
identify closely acting genes and exclude marginal genes of the
network. Finally, the five most relevant genes associated with lung
adenocarcinoma pathogenesis genes and targets of the
23-HBA-associated domain, prostaglandin-endoperoxide synthase 2,
PPAR-γ, angiotensin I-converting enzyme (ACE), PPAR-α and mouse
double minute 2 (MDM2), were selected and sorted in accordance with
their association (Fig. 3C). ACE
is a zinc metallopeptidase that is widely distributed in
endothelial and epithelial cells. It plays a crucial role in the
renin-angiotensin-aldosterone system. Consequently, ACE is mainly
found in vascular endothelial cells (37). For PTSG2, the related studies have
been relatively clear and specific, so we excluded it from further
analysis. MDM2, as a key negative regulator of p53, exhibits high
expression in tumors and serves an important role in tumor
initiation and development. MDM2 has been demonstrated to bind to
p53, thereby blocking its tumor-suppressor transactivation domain
(38). Furthermore, MDM2 functions
as an E3 ligase, marking p53 for degradation by the proteasome.
H1299 cells possess a homozygous partial deletion of TP53(39), rendering them unable to express the
tumor-suppressor protein p53, which contributes to their
proliferative phenotype. Therefore, MDM2 was excluded from further
analysis. PPAR-α and PPAR-γ belong to the same category. PPAR-γ has
been rarely studied in the literature and it represents a novel
finding. Therefore, it was selected for exploration in the present
study.
PPAR-γ functions as a tumor suppressor
in lung adenocarcinoma
The GEPIA database was utilized to analyze the
expression of the target gene PPAR-γ in various tumor types
(40), including lung
adenocarcinoma (Fig. 4A). The
results demonstrated that PPAR-γ was significantly downregulated in
lung adenocarcinoma compared with normal tissues (Fig. 4B), while being highly expressed in
most tumors, thereby indicating that PPAR-γ may function as a
tumor-suppressor gene. Western blot assay was performed to measure
PPAR-γ expression in normal cells (BEAS-2B) and multiple lung
adenocarcinoma cell lines (A549, H1975 and H1299), to further
confirm its role as a tumor suppressor. The results showed that
PPAR-γ was highly expressed in normal cells, whereas its expression
was notably lower in the lung adenocarcinoma cell lines (Fig. 4C). Subsequently, western blotting
was conducted to examine the protein expression of PPAR-γ after
treatment of the cells with different concentrations of 23-HBA. The
results indicated that the expression of PPAR-γ in the 23-HBA
treatment groups was higher than that in the control group
(41), with statistical
significance observed at a drug concentration of 60 µM (Fig. 4D). These results confirmed that
23-HBA treatment increases the expression of PPAR-γ. Given the
earlier conclusion that 23-HBA can inhibit the survival of lung
adenocarcinoma cells, PPAR-γ may act as a tumor suppressor, which
is consistent with the results from the GEPIA database.
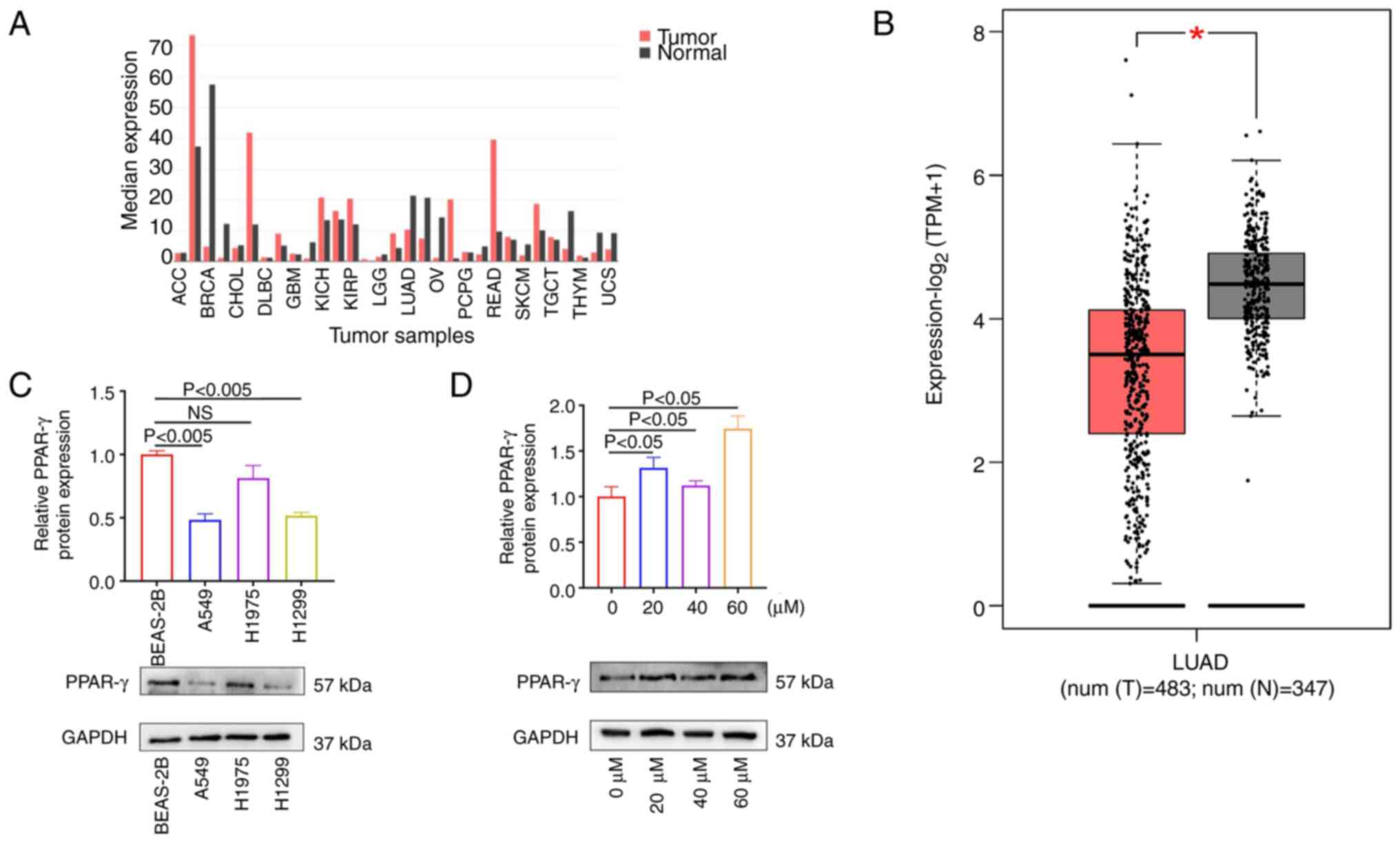 | Figure 4PPAR-γ is tumor-suppressor gene in
lung adenocarcinoma. (A) Analysis of the GEPIA database revealed
the expression levels of the target gene PPAR-γ in various tumor
types. (B) The GEPIA database analysis results indicated that
PPAR-γ expression levels are significantly decreased in lung
adenocarcinoma compared with normal tissue. *P<0.05.
(C) Western blotting results demonstrated high expression of PPAR-γ
in normal bronchial epithelial cells and low expression in lung
adenocarcinoma cell lines. (D) Western blotting results revealed a
notable increase in PPAR-γ expression levels in lung adenocarcinoma
cells treated with 23-hydroxybetulinic acid compared with the
control group. PPAR-γ, peroxisome proliferator-activated
receptor-γ; LUAD, lung adenocarcinoma; ACC, adrenocortical cancer;
BRCA, breast invasive carcinoma; CHOL, cholangiocarcinoma; DLBC,
diffuse large B-cell lymphoma; GBM, glioblastoma multiforme; KICH,
kidney chromophobe; KIRP, kidney papillary cell carcinoma; LGG,
brain lower grade glioma; OV, ovarian serous cystadenocarcinoma;
PCPG, pheochromocytoma; READ, rectum adenocarcinoma; SKCM, skin
cutaneous melanoma; TGCT, testicular germ cell tumor; THYM,
thymoma; UCS, uterine carcinosarcoma; T, tumor; N, normal tissue;
GEPIA, Gene Expression Profiling Interactive Analysis; TPM,
transcripts per million; ns, not significant. |
Mitochondrial ROS is associated with
the mechanism of inhibition of lung adenocarcinoma cell
proliferation by 23-HBA
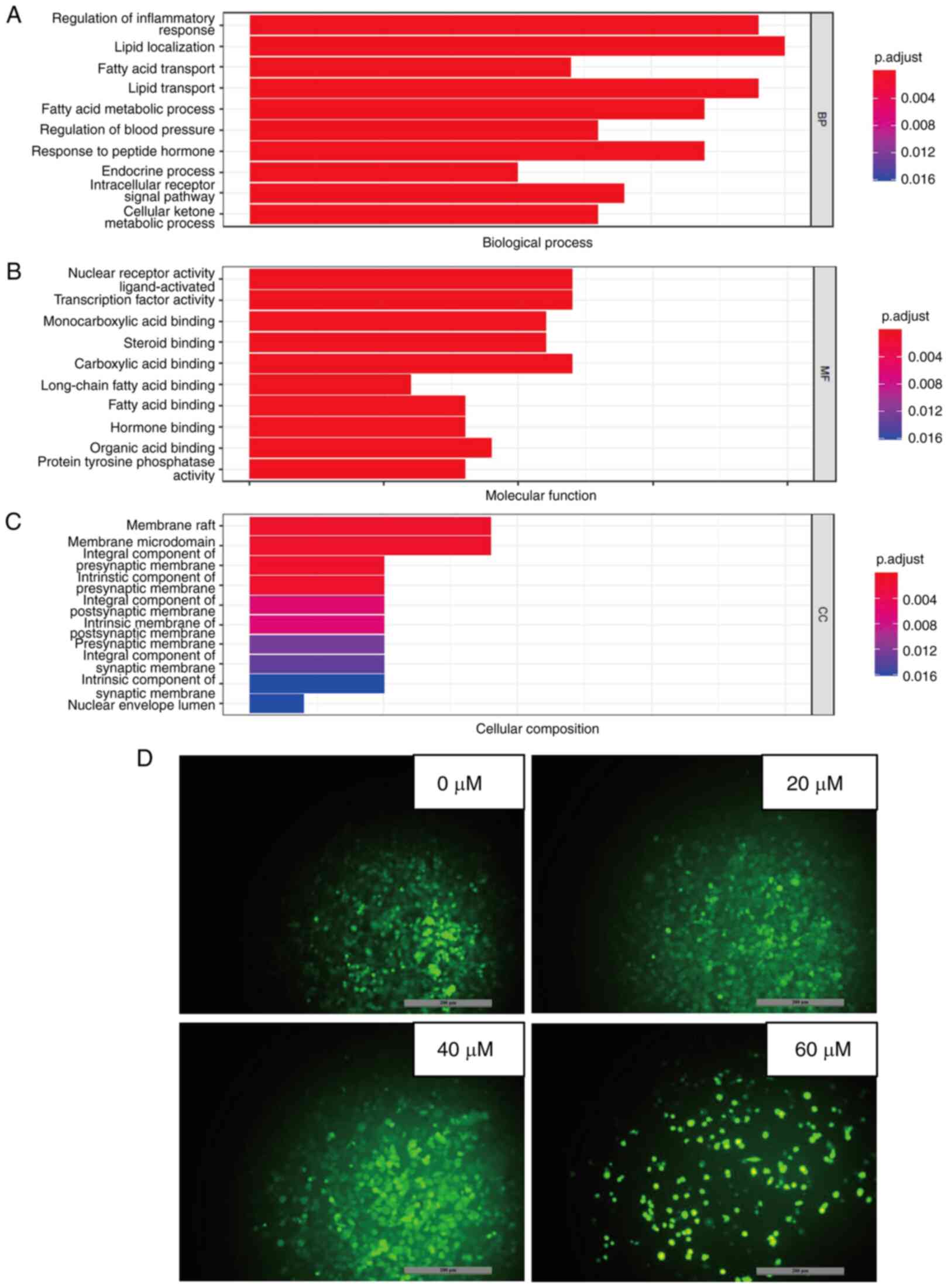
GO analysis was conducted to investigate the mode of
action of 23-HBA on lung adenocarcinoma (42). P<0.05 was used as the screening
criterion for selected functional terms in the categories BP, MF
and CC (Fig. 5A-C). The results
showed that most of the enriched BP terms require the consumption
of ATP, which is primarily produced by mitochondria. From Fig. 5A, it can be concluded that most
enriched BP terms are associated with regulation of inflammatory
response, which require the consumption of ATP, such as lipid
transporting and fatty acid metabolic processing. The most
important mechanisms of action in the category MF are associated
with nuclear receptor activity, which also contains
ligand-activated transcription factor activity and carboxylic acid
binding. The most dominant mechanism of action in the category CC
is membrane raft, which also contains membrane microdomain and
integral component of presynaptic membrane. Therefore, the
mechanism of action of the PPAR pathway is related to mitochondria
due to increasing the content of mitochondrial ROS, thereby
inhibiting cell growth. Further investigation through PubMed into
the PPAR signaling pathway revealed that it induces physiological
changes, including inflammation and alterations in the
mitochondrial membrane potential (43). Therefore, the mechanism by which
23-HBA inhibits the viability and/or proliferation of lung
adenocarcinoma cells may be related to mitochondrial ROS.
Subsequently, the mitochondrial ROS levels were detected, and the
results showed that the production of intracellular mitochondrial
ROS notably increased with increasing concentrations of 23-HBA
(Fig. 5D), which is consistent
with the aforementioned mechanism.
Clinical prognosis of patients with
lung adenocarcinoma is associated with the levels of PPAR signaling
pathway-related genes
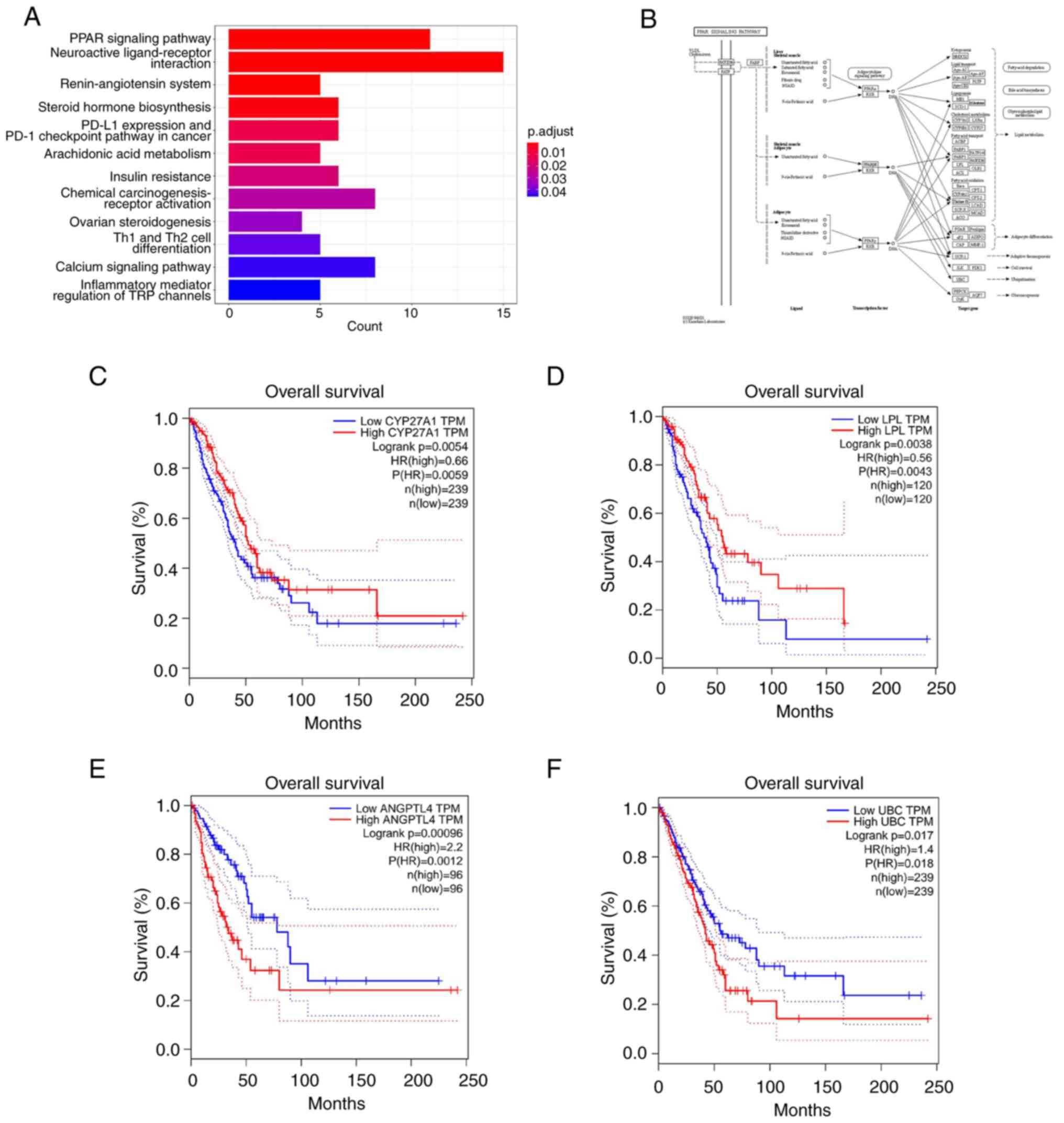
The PPAR signaling pathway is considered to be an
important signal transduction pathway in several conditions, such
as inflammation and cancer (44).
Previous studies have demonstrated the inhibitory effects of 23-HBA
and its derivatives on inflammation, as well as liver, lung and
breast cancer (45,46). The KEGG analysis revealed that the
pathway exhibiting the highest degree of enrichment was for the
‘PPAR signaling pathway’ (Fig.
6A). Investigation of the research mechanism within this
pathway resulted in the identification of target genes associated
with it, including CYP27, LPL, PGAR/ANGPTL4 and UBC (Fig. 6B). Subsequently, the GEPIA database
was used to analyze the association of the expression levels of
these target genes with the survival of patients with lung
adenocarcinoma. The results indicated that high (ANGPTL4 and UBC)
or low (CYP27 and LPL) expression levels of these genes were
significantly associated with a lower patient survival (Fig. 6C-F), thereby demonstrating the
clinical prognostic significance of the PPAR pathway.
Effect of 23-HBA on lung
adenocarcinoma and its mechanism of action
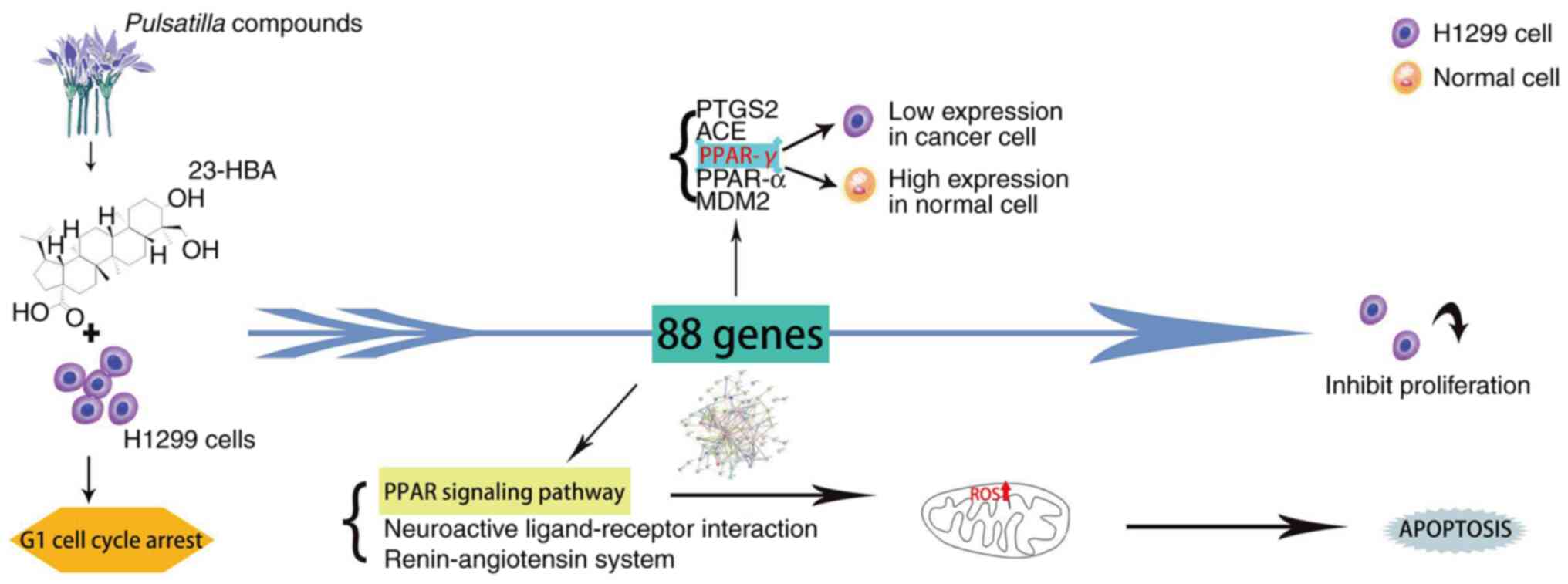
Collectively, the results of the present study
demonstrated that 23-HBA inhibits cell proliferation by blocking
lung adenocarcinoma cells in the G1 phase. The results
indicated that PPAR-γ may function as a tumor-suppressor gene in
lung adenocarcinoma. Further analysis unveiled the relationship
among this finding, the PPAR signaling pathway and mitochondrial
ROS. The relationship is that the survival rate of tumor cells
decreases with the increase of drug concentration, and at the same
time, the expression of ROS increases with the increase of drug
concentration. Finally, a clinical prognosis association was
demonstrated between the expression levels of PPAR pathway-related
genes and the survival of patients with lung adenocarcinoma
(Fig. 7).
Discussion
PPAR-γ is a nuclear receptor involved in various
cellular processes, including regulation of gene expression that
serves an important regulatory function in the process of lipid
metabolism (47). Studies have
indicated that PPAR-γ plays a pivotal role in antitumor mechanisms
by governing cell cycle arrest, apoptosis, proliferation, invasion
and migration, among other functions (48,49).
Western blot analysis in the present study indicated that PPAR-γ
functions as a tumor-suppressor gene by inhibiting the growth of
lung adenocarcinoma cells. In a previous study, PPAR-γ gene
silencing and overexpression vectors were constructed for the study
of human squamous cell carcinoma. These vectors were transfected
into human squamous cell carcinoma Colo16 cells to assess the
effects of PPAR-γ on cell proliferation and apoptosis. The results
indicated that PPAR-γ silencing promoted Colo16 cell proliferation
and reduced apoptosis, whereas PPAR-γ overexpression exhibited the
opposite effects. Therefore, PPAR-γ serves a role in inhibiting
tumor cell proliferation and promoting apoptosis in human squamous
cell carcinoma Colo16 cells (50).
The conclusions of the present study align with these findings.
However, no specific mechanism concerning the involvement of
PPAR-γ-associated genes in lung adenocarcinoma has been reported.
In the present study, PPAR-γ was shown, to be downregulated in lung
adenocarcinoma cells and function as a tumor-suppressor gene
(51,52).
KEGG analysis was conducted on the selected
intersecting genes to delve further into the mechanism of 23-HBA in
lung adenocarcinoma (20). The
pathway exhibiting the highest degree of enrichment was the ‘PPAR
signaling pathway’, which plays a pivotal regulatory role in
immunity (53), inflammation,
vascular function (54), cell
proliferation, differentiation, development, apoptosis and cancer.
This pathway is not only implicated in immune function,
carbohydrate and lipid metabolism, but is also closely associated
with the proliferation, death, invasion and metastasis of tumor
cells during the course of tumor development. Activation of the
PPAR pathway affects angiogenesis and tumor suppression, and
regulates the proliferation and differentiation of cancer cells
(55).
Previous studies reported that Pulsatilla
compounds play a crucial role in inhibiting the growth of tumor
cells, particularly in liver, lung and breast cancer (56,57).
B4G6, a derivative of 23-HBA, has been demonstrated to induce
mitochondrial apoptosis by triggering ROS accumulation and the
opening of calcium-dependent permeability transition pores, thereby
inhibiting the proliferation of HepG2, Hep3B, Bel-7402 and
HepG2/ADM cell lines (58). B4G2
exhibits a potent inhibitory effect on HepG2 cell proliferation and
induces mitochondrial autophagy. Furthermore, the use of autophagy
inhibitors could enhance the apoptosis rate of HepG2 cells
(59). Experiments involving
mitochondrial ROS were conducted to further explore the mechanism
of 23-HBA in lung adenocarcinoma. The results showed that the
accumulation of intracellular mitochondrial ROS significantly
increased with an increase in drug concentration.
Furthermore, MTT and flow cytometry analyses were
employed in the present study to assess the effect of 23-HBA on
lung adenocarcinoma cells (60).
The MTT results indicated that 23-HBA had an inhibitory effect on
lung adenocarcinoma cell survival. Moreover, 23-HBA caused the
arrest of H1299 cells in the G1 cell cycle phase, as
determined by flow cytometry. In summary, 23-HBA was shown to
effectively inhibit the survival and proliferation of lung
adenocarcinoma cells. This finding establishes a theoretical
foundation for the development and clinical application of new
antitumor drugs based on Pulsatilla compounds (61). Developing high-efficiency and
low-toxicity drugs with Pulsatilla compounds as the primary
constituents may aid in inhibiting tumor growth in the future
(62). In this context, 23-HBA may
represent a promising therapeutic strategy for lung adenocarcinoma
by interacting with the PPAR-γ and mitochondrial ROS pathways
(63).
In summary 23-HBA was shown to inhibit cell
proliferation by arresting lung adenocarcinoma cells in the
G1 phase, and PPAR-γ was indicated to be a
tumor-suppressor gene in lung adenocarcinoma. In addition, the PPAR
signaling pathway had the highest degree of enrichment among genes
associated with 23-HBA and lung adenocarcinoma; however, the
mechanism involved was not clarified, and further animal
experiments are required to elucidate the inhibitory effects of
23-HBA on lung adenocarcinoma in vivo. 23-HBA is the active
ingredient extracted from the Radix Pulsatilla, a plant used
in traditional Chinese medicine. However, the use of traditional
Chinese medicine in cancer treatment is uncommon. Therefore, the
present study could set the foundation for the potential
application of traditional Chinese medicine in lung adenocarcinoma
treatment, thereby expanding new avenues for the development of
traditional Chinese and Western medicines.
Acknowledgements
Not applicable.
Funding
Funding: The present study was supported by the Shandong Science
and Technology Committee (grant no. ZR2023MH223), the National
Natural Science Foundation of China (grant no. 81800169) and
College Students' Innovation and Entrepreneurship Training Program
(grant nos. 202310440061, 202310440029 and S202310440012).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
BT designed and performed the cell experiments. XL
screened the final drug through cell experiments. YZ performed the
database search and data processing, and assisted in the
experimental design. PL and QJ analyzed the data. ZW, ZL, WS and YX
assisted in the cell experiments. YL and YS designed the
experiments and revised the manuscript. All authors read and
approved the final manuscript. All authors confirm the authenticity
of all the raw data.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Sung H, Ferlay J, Siegel RL, Laversanne M,
Soerjomataram I, Jemal A and Bray F: Gloal cancer statistics 2020:
GLOBOCAN estimates of incidence and mortality worldwide for 36
cancers in 185 countries. CA Cancer J Clin. 71:209–249.
2021.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Shigeta K, Datta M, Hato T, Kitahara S,
Chen IX, Matsui A, Kikuchi H, Mamessier E, Aoki S, Ramjiawan RR, et
al: Dual programmed death receptor-1 and vascular endothelial
growth factor receptor-2 blockade promotes vascular normalization
and enhances antitumor immune responses in hepatocellular
carcinoma. Hepatology. 71:1247–1261. 2020.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Aoki T, Kudo M, Ueshima K, Morita M,
Chishina H, Takita M, Hagiwara S, Ida H, Minami Y, Tsurusaki M and
Nishida N: Incidence of hyper progressive disease in combination
immunotherapy and anti-programmed cell death protein 1/programmed
death-ligand 1 monotherapy for unresectable hepatocellular
carcinoma. Liver Cancer. 13:56–69. 2023.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Cheng X: The effect of CDKN2A gene
mutation on lung cancer and its relationship with prognosis. J Appl
Cancer. 875-878(883)2021.
|
|
5
|
Oudkerk M, Liu S, Heuvelmans MA, Walter JE
and Field JK: Lung cancer LDCT screening and mortality
reduction-evidence, pitfalls and future perspectives. Nat Rev Clin
Oncol. 18:135–151. 2021.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Zhang L, Shao F, Li L, et al: CALM1 gene
expression and EGFR mutation in colorectal cancer and its clinical
significance. Gen Med Edu. 21:18–21. 2023.DOI: 10.13558/j. carol
carroll nki issn1672-3686.2023.001.005.
|
|
7
|
Brown A and Goodman Z: Hepatitis
B-associated fibrosis and fibrosis/cirrhosis regression with
nucleoside and nucleotide analogs. Expert Rev Gastroenterol
Hepatol. 6:187–198. 2012.PubMed/NCBI View
Article : Google Scholar
|
|
8
|
Clough E and Barrett T: The gene
expression omnibus database. Methods Mol Biol. 1418:93–110.
2016.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Huang J, Zhang J, Zhang F, Lu S, Guo S,
Shi R, Zhai Y, Gao Y, Tao X, Jin Z, et al: Identification of a
disulfidptosis-related genes signature for prognostic implication
in lung adenocarcinoma. Comput Biol Med. 165(107402)2023.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Liang Q, Gong M, Zou JH, Luo MY, Jiang LL,
Wang C, Shen NX, Zhang MC, Xu L, Lei HM, et al: A phosphoglycerate
mutase 1 allosteric inhibitor overcomes drug resistance to
EGFR-targeted therapy via disrupting IL-6/JAK2/STAT3 signaling
pathway in lung adenocarcinoma. Drug Resist Updat.
68(100957)2023.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Jiang M, Yang M, Zhou YY, et al: Effects
of 23-hydroxybetulinic acid on proliferation and apoptosis of
vascular endothelial cells. Cancer Prev Res. 911–913+985. 2007.
|
|
12
|
Molina JR, Yang P, Cassivi SD, Schild SE
and Adjei AA: Non-small cell lung cancer: Epidemiology, risk
factors, treatment, and survivorship. Mayo Clin Proc. 83:584–594.
2008.PubMed/NCBI View
Article : Google Scholar
|
|
13
|
Tang Z, Kang B, Li C, Chen T and Zhang Z:
GEPIA2: An enhanced web server for large-scale expression profiling
and interactive analysis. Nucleic Acids Res. 47 (W1):W556–W560.
2019.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Caligiuri G: CD31 as a therapeutic target
in atherosclerosis. Circ Res. 126:1178–1189. 2020.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Liao D, Sundlov J, Zhu J, Mei H, Hu Y,
Newman DK and Newman PJ: Atomic level dissection of the platelet
endothelial cell adhesion molecule 1 (PECAM-1) homophilic binding
interface: Implications for endothelial cell barrier function.
Arterioscler Thromb Vasc Biol. 42:193–204. 2022.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Hu M, Zhang H, Liu Q and Hao Q: Structural
basis for human PECAM-1-mediated trans-homophilic cell adhesion.
Sci Rep. 6(38655)2016.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Winneberger J, Schöls S, Lessmann K,
Rández-Garbayo J, Bauer AT, Mohamud Yusuf A, Hermann DM, Gunzer M,
Schneider SW, Fiehler J, et al: Platelet endothelial cell adhesion
molecule-1 is a gatekeeper of neutrophil transendothelial migration
in ischemic stroke. Brain Behav Immun. 93:277–287. 2021.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Zhu H, Lu L, Zhu W, Tan Y, Duan Y, Liu J,
Ye W, Zhu Z, Xu J and Xu S: Design and synthesis of NAD(P)H:
Quinone oxidoreductase (NQO1)-activated prodrugs of
23-hydroxybetulinic acid with enhanced antitumor properties. Eur J
Med Chem. 240(114575)2022.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Chen SY, Tsuneyama K, Yen MH, Lee JT, Chen
JL and Huang SM: Hyperbaric oxygen suppressed tumor progression
through the improvement of tumor hypoxia and induction of tumor
apoptosis in A549-cell-transferred lung cancer. Sci Rep.
11(12033)2021.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Tian D, Yu Y, Zhang L, Sun J and Jiang W:
23-Hydroxybetulinic acid reduces tumorigenesis, metastasis and
immunosuppression in a mouse model of hepatocellular carcinoma via
disruption of the MAPK signaling pathway. Anticancer Drugs.
33:815–825. 2022.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Ye B and Ji ZN: 23-Hydroxybetulinic
acid-induced HL-60 cell autophagic apoptosis and its molecular
mechanism. Nat Prod Res. 26:1063–1068. 2012.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Doi H, Kida T, Nishino K, Nakatsuji M,
Sakamoto S, Shimizu S, Teraoka Y, Tamura Y, Kataoka Y and Inui T:
Solubility-improved 10-O-substituted SN-38 derivatives with
antitumor activity. ChemMedChem. 12:1715–1722. 2017.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Xiang S: Effect and mechanism of
23-hydroxybetulcholic acid on ulcerative colitis induced by DSS.
Zunyi Medical Coll, 2022. DOI: 10.27680 /, dc nki. Gzyyc.
2022.000496.
|
|
24
|
Wang C, Yu Q, Song T, Wang Z, Song L, Yang
Y, Shao J, Li J, Ni Y, Chao N, et al: The heterogeneous immune
landscape between lung adenocarcinoma and squamous carcinoma
revealed by single-cell RNA sequencing. Signal Transduct Target
Ther. 7(289)2022.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Kobayashi Y and Mitsudomi T: Not all
epidermal growth factor receptor mutations in lung cancer are
created equal: Perspectives for individualized treatment strategy.
Cancer Sci. 107:1179–1186. 2016.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Li Q, Zhang H, Yan X, Zhao Z, Qiu J, Hu L,
Jiang S, Kong Q, Sun J and Li L: Up-regulation of PPAR-γ involved
in the therapeutic effect of icariin on cigarette smoke-induced
inflammation. Pulm Pharmacol Ther. 79(102197)2023.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Chen JJ, Patel A, Sodani K, Xiao ZJ,
Tiwari AK, Zhang DM, Li YJ, Yang DH, Ye WC, Chen SD and Chen ZS:
BBA, a synthetic derivative of 23-hydroxybutulinic acid, reverses
multidrug resistance by inhibiting the efflux activity of MRP7
(ABCC10). PLoS One. 8(e74573)2013.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Chen H, Bai L, Shi Y, Zhang X, Wang X,
Wang Y, Hu J and Zhou P: Investigation of the molecular mechanism
underlying the therapeutic effect of perilla frutescens L.
Essential oil on acute lung injury using gas chromatography mass
spectrometry and network pharmacology. Comb Chem High Throughput
Screen: Oct 10, 2023 (Epub ahead of print).
|
|
29
|
Huang Z, Gao Y and Hou D: Interleukin-22
enhances chemoresistance of lung adenocarcinoma cells to
paclitaxel. Hum Cell. 33:850–858. 2020.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Feng Z, Chen Y, Cai C, Tan J, Liu P, Chen
Y, Shen H, Zeng S and Han Y: Pan-cancer and single-cell analysis
reveals CENPL as a cancer prognosis and immune infiltration-related
biomarker. Front Immunol. 13(916594)2022.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Cao J, Sun S, Min R, Li R, Fan X, Han Y,
Feng Z and Li N: Prognostic significance of CCNB2 expression in
triple-negative breast cancer. Cancer Manag Res. 13:9477–9487.
2021.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Zhang W, Tian W, Wang Y, Jin X, Guo H,
Wang Y, Tang Y and Yao X: Explore the mechanism and substance basis
of Mahuang FuziXixin Decoction for the treatment of lung cancer
based on network pharmacology and molecular docking. Comput Biol
Med. 151(106293)2022.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Du Y, Lv Z, Sun D, Li Y, Sun L and Zhou J:
Physcion 8-O-β-glucopyranoside exerts anti-tumor activity against
non-small cell lung cancer by targeting PPARγ. Anat Rec (Hoboken).
302:785–793. 2019.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Kim S and Bae S: In vitro and in vivo
human body odor analysis method using GO:PANI/ZNRs/ZIF-8 adsorbent
followed by GC/MS. Molecules. 27(4795)2022.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Lei Y: Anti-liver cancer activity
evaluation and mechanism of 23-hydroxybetulinic acid derivative
B4G2. Jinan University, 2017.
|
|
36
|
Ma J, Cai X, Kang L, Chen S and Liu H:
Identification of novel biomarkers and candidate small-molecule
drugs in cutaneous melanoma by comprehensive gene microarrays
analysis. J Cancer. 12:1307–1317. 2021.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Liu L, Zhou S, Yang Z, et al: Study on
population distribution of ACE gene polymorphism and its high
expression in essential hypertension. J Pract Lab Phys. 10:132–134.
2018.
|
|
38
|
Ding H and Hu M: Research progress of
anti-tumor small molecule inhibitors targeting p53-MDM2. Adv Clin
Med. 13:13474–13483. 2023.(In Chinese).
|
|
39
|
Xiong Y and Li B: The cause of deletion of
P53 protein in H1299 cells. Macau Med J. 4:8–9. 2004.
|
|
40
|
Dourado KMC, Baik J, Oliveira VKP,
Beltrame M, Yamamoto A, Theuer CP, Figueiredo CAV, Verneris MR and
Perlingeiro RCR: Endoglin: A novel target for therapeutic
intervention in acute leukemias revealed in xenograft mouse models.
Blood. 129:2526–2536. 2017.PubMed/NCBI View Article : Google Scholar
|
|
41
|
liu xiao mei, first hai-li wang, etc:
Research progress on the role of PPAR-γ/AP-1 signaling pathway in
related diseases. J Yichun Univ, 21,43: 12-16.
|
|
42
|
Moloney JN and Cotter TG: ROS signalling
in the biology of cancer. Semin Cell Dev Biol. 80:50–64.
2018.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Cao XJ, Zhang MJ, Zhang LL, Yu K, Xiang Y,
Ding X, Fan J, Li JC and Wang QS: TLR4 mediates high-fat diet
induced physiological changes in mice via attenuating PPARγ/ABCG1
signaling pathway. Biochem Biophys Res Commun. 503:1356–1363.
2018.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Alqahtani S and Mahmoud AM:
Gamma-Glutamylcysteine Ethyl Ester Protects against
Cyclophosphamide-Induced Liver Injury and Hematologic Alterations
via Upregulation of PPAR and Attenuation of Oxidative Stress,
Inflammation, and Apoptosis. Oxidative Medicine and Cellular
Longevity, 2016.
|
|
45
|
Laganà AS, Vitale SG, Nigro A, Sofo V,
Salmeri FM, Rossetti P, Rapisarda AM, La Vignera S, Condorelli RA,
Rizzo G and Buscema M: Pleiotropic actions of peroxisome
proliferator-activated receptors (PPARs) in dysregulated metabolic
homeostasis, inflammation and cancer: Current evidence and future
perspectives. Int J Mol Sci. 17(999)2016.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Jiang MJ, Yang M, Zhou YY, Zhang RJ, Cao
GX, Cai GM and Wang GJ: In vitro inhibitory effect of 23-HBA on
angiogenesis. Xi Bao Yu Fen Zi Mian Yi Xue Za Zhi. 22:88–91.
2006.PubMed/NCBI(In Chinese).
|
|
47
|
Zhang DM, Shu C, Chen JJ, Sodani K, Wang
J, Bhatnagar J, Lan P, Ruan ZX, Xiao ZJ, Ambudkar SV, et al: BBA, a
derivative of 23-hydroxybetulinic acid, potently reverses
ABCB1-mediated drug resistance in vitro and in vivo. Mol Pharm.
9:3147–3159. 2012.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Nahlé N: PPAR trilogy from metabolism to
cancer. Curr Opin Clin Nutr Metab Care. 7:397–402. 2004.PubMed/NCBI View Article : Google Scholar
|
|
49
|
Elix CC, Salgia MM, Otto-Duessel M,
Copeland BT, Yoo C, Lee M, Tew BY, Ann D, Pal SK and Jones JO:
Peroxisome proliferator-activated receptor gamma controls prostate
cancer cell growth through AR-dependent and independent mechanisms.
Prostate. 80:162–172. 2020.PubMed/NCBI View Article : Google Scholar
|
|
50
|
Zhang J and Bai WP: C1q/tumor necrosis
factor related protein 6 (CTRP6) regulates the phenotypes of high
glucose-induced gestational trophoblast cells via peroxisome
proliferator-activated receptor gamma (PPARγ) signaling.
Bioengineered. 13:206–216. 2022.PubMed/NCBI View Article : Google Scholar
|
|
51
|
Zhou P: Preliminary study on the function
of PPARG gene in human skin squamous cell carcinoma Colol6 cells.
Peking Union Medical College, 2019.
|
|
52
|
Zhan L, Zhang H, Zhang Q, Woods CG, Chen
Y, Xue P, Dong J, Tokar EJ, Xu Y, Hou Y, et al: Regulatory role of
KEAP1 and NRF2 in PPARγ expression and chemoresistance in human
non-small-cell lung carcinoma cells. Free Radic Biol Med.
53:758–768. 2012.PubMed/NCBI View Article : Google Scholar
|
|
53
|
Li H, Sorenson AL, Poczobutt J, Amin J,
Joyal T, Sullivan T, Crossno JT Jr, Weiser-Evans MC and Nemenoff
RA: Activation of PPARgamma in myeloid cells promotes lung cancer
progression and metastasis. PLoS One. 6(e28133)2011.PubMed/NCBI View Article : Google Scholar
|
|
54
|
Liu S, Zhang H, Li Y, Zhang Y, Bian Y,
Zeng Y, Yao X, Wan J, Chen X, Li J, et al: S100A4 enhances protumor
macrophage polarization by control of PPAR-γ-dependent induction of
fatty acid oxidation. J Immunother Cancer.
9(e002548)2021.PubMed/NCBI View Article : Google Scholar
|
|
55
|
Wagner N and Wagner KD: Pharmacological
utility of PPAR modulation for angiogenesis in cardiovascular
disease. Int J Mol Sci. 24(2345)2023.PubMed/NCBI View Article : Google Scholar
|
|
56
|
Fang C, Liu Y, Chen L, Luo Y, Cui Y, Zhang
N, Liu P, Zhou M and Xie Y: α-Hederin inhibits the growth of lung
cancer A549 cells in vitro and in vivo by decreasing SIRT6
dependent glycolysis. Pharm Biol. 59:11–20. 2021.PubMed/NCBI View Article : Google Scholar
|
|
57
|
Reka AK, Goswami MT, Krishnapuram R,
Standiford TJ and Keshamouni VG: Molecular cross-regulation between
PPAR-γ and other signaling pathways: Implications for lung cancer
therapy. Lung Cancer. 72:154–159. 2011.PubMed/NCBI View Article : Google Scholar
|
|
58
|
Łaska G, Sieniawska E, Maciejewska-Turska
M, Świątek Ł, Pasco DS and Balachandran P: Pulsatilla vulgaris
inhibits cancer proliferation in signaling pathways of 12 reporter
genes. Int J Mol Sci. 24(1139)2023.PubMed/NCBI View Article : Google Scholar
|
|
59
|
Cheung EC and Vousden KH: The role of ROS
in tumour development and progression. Nat Rev Cancer. 22:280–297.
2022.PubMed/NCBI View Article : Google Scholar
|
|
60
|
Zhang S, Wang C, Tang S, Deng S, Zhou Y,
Dai C, Yang X and Xiao X: Inhibition of autophagy promotes
caspase-mediated apoptosis by tunicamycin in HepG2 cells. Toxicol
Mech Methods. 24:654–665. 2014.PubMed/NCBI View Article : Google Scholar
|
|
61
|
Yao N, Li YJ, Lei YH, Hu N, Chen WM, Yao
Z, Yu M, Liu JS, Ye WC and Zhang DM: A piperazidine derivative of
23-hydroxy betulinic acid induces a mitochondria-derived ROS burst
to trigger apoptotic cell death in hepatocellular carcinoma cells.
J Exp Clin Cancer Res. 35(192)2016.PubMed/NCBI View Article : Google Scholar
|
|
62
|
Liu M, Zhao X, Xiao L, Liu G, Liu H, Wang
X, Feng X and Lin X: Cytotoxicity of the compounds isolated from
Pulsatilla chinensis saponins and apoptosis induced by
23-hydroxybetulinic acid. Pharm Biol. 53:1–9. 2015.PubMed/NCBI View Article : Google Scholar
|
|
63
|
Yu C, Chen F, Jiang J, Zhang H and Zhou M:
Screening key genes and signaling pathways in colorectal cancer by
integrated bioinformatics analysis. Mol Med Rep. 20:1259–1269.
2019.PubMed/NCBI View Article : Google Scholar
|
|
64
|
Zhang Z, Zhang X, Meng L, Gong M, Li J,
Shi W, Qiu J, Yang Y, Zhao J, Suo Y, et al: Pioglitazone inhibits
diabetes-induced atrial mitochondrial oxidative stress and improves
mitochondrial biogenesis, dynamics, and function through the
PPAR-γ/PGC-1α signaling pathway. Front Pharmacol.
12(658362)2021.PubMed/NCBI View Article : Google Scholar
|