Introduction
Myocardial fibrosis (MF) represents a key
pathological change associated with the remodeling of myocardial
tissue, characterized by an increase in the interstitial space
within the heart muscle, primarily as a result of heightened
deposition of extracellular matrix (ECM) components (1). This accumulation is largely driven by
the activity of activated myofibroblasts, which serve a key role in
the fibrotic process (2). As a
consequence of MF, patients often experience a range of adverse
cardiovascular effects, including an increase in myocardial
stiffness, which can hinder the ability of the heart to relax
properly during diastole, leading to ventricular diastolic
dysfunction (3). Furthermore, MF
can contribute to a decreased coronary flow reserve, impacting the
capacity of the heart to receive an adequate blood supply during
heightened activity (4). Globally,
the annual incidence rate of MF is ~1.7% (5). Notably, MF affects >60% of
patients with chronic heart failure and is a key predictor of
adverse outcomes, including arrhythmias and sudden cardiac
mortality, with 5-year mortality rates ranging from 40 to 60%
(5,6). Current MF management involves
renin-angiotensin-aldosterone system inhibitors, β-blockers and
statins. While they may alleviate the symptoms of patients, these
agents have demonstrated a limited efficacy in halting or reversing
MF (7). Mechanistically, the
complexity of the fibroblast response to injury and the
pathophysiologic heterogeneity of MF pose challenges to the
development of antifibrotic therapeutic strategies (8). To the best of our knowledge,
currently, there is no ideal treatment for MF, thus novel
therapeutic strategies need to be developed. In previous years,
with the development of high-throughput sequencing technology,
molecular targeted therapies have become a hotspot for the
treatment of a number of diseases (9,10).
The exact etiology of MF remains unclear, highlighting the
importance of in-depth studies focusing on the molecular mechanisms
of MF to identify novel therapeutic targets.
RNA-binding proteins (RBPs) are proteins that bind
to RNA to regulate RNA metabolism (11). RBPs interact directly with single-
or double-stranded RNA and are involved in post-transcriptional
processes such as alternative splicing, RNA export, mRNA
translation, RNA degradation and RNA stabilization to regulate gene
expression (12-14).
Dysfunctions in RBPs underscore a number of diseases. A previous
study demonstrated that reducing the expression of CUGBP Elav-like
family member 1 could notably ameliorate cardiac fibrosis (15). In addition, another study
demonstrated that inhibition of cytoskeleton associated protein 4
expression led to an increase in the expression of TGF-β-stimulated
fibroblast activation-related genes, which subsequently serves a
role in the regulation of MF (16). Furthermore, it has been reported
that polypyrimidine tract binding protein 1 promotes cardiac
fibrosis by facilitating collagen deposition through the
degradation of nuclear receptor 4A1 mRNA, followed by
transcriptional repression of fatty acid binding protein 5(17).
Pumilio RNA-binding family member 2 (PUM2), a member
of the pumilio family, belongs to a group of RBPs that recognize
sequences in RNA. PUM2 functions as a translational regulator
during embryonic development and cell differentiation (18). PUM2 serves as a translational
repressor that becomes increasingly active during the process of
cellular senescence. Importantly, it functions as a negative
regulator of both lifespan and the maintenance of mitochondrial
homeostasis, meaning that the presence and activity of PUM2 can
inhibit cellular processes that are key for longevity and the
proper functioning of mitochondria, overall contributing to the
aging process at a cellular level. Understanding the mechanisms by
which PUM2 operates could provide insights into the complexities of
lifespan regulation and the preservation of mitochondrial integrity
(19). Aberrant expression of PUM2
is implicated in the progression of a range of diseases. Ding et
al (20) demonstrated that, in
cervical cancer, PUM2 promoted tumor progression by destabilizing
tumor-suppressor mRNAs. A recent study found that PUM2 also
exacerbated neuroinflammation and brain damage caused by
ischemia-reperfusion (21). These
findings demonstrate the capacity of PUM2 to drive pathological
processes across tissues, suggesting it as a high-priority target
for MF investigation. In MF-related studies, PUM2 serves as a
regulatory hub for gene translation in fibrotic hearts of patients
with dilated cardiomyopathy (22,23).
In vitro experiments have demonstrated that knockdown of
PUM2 inhibits TGF-β1-induced fibroblast activation (24). However, whether PUM2 contributes to
cardiac fibrosis by regulating the expression or alternative
splicing of downstream target genes remains unknown and warrants
investigation.
Given the established role of PUM2 in fibrotic
hearts of patients with dilated cardiomyopathy, whereby it acts as
a translational hub for profibrotic genes, we hypothesized that it
orchestrates MF through dual regulation of gene expression and
alternative splicing in cardiomyocytes. In the present study,
transcriptome data following knockdown of PUM2 in H9C2 rat
cardiomyocytes were obtained via high-throughput sequencing.
Potential transcriptional targets of PUM2 and alternative splicing
in H9C2 cells were analyzed to investigate the molecular functions
and mechanisms of PUM2 in the development of MF.
Materials and methods
Small interfering RNA (siRNA/si)
information
For the present study, siRNA duplexes were sourced
from Shanghai GenePharma Co., Ltd. A non-targeting control siRNA,
labeled siNegative, was employed, with the following sequences:
Sense, 5'-UUCUCCGAACGUGUCACGUTT-3' and antisense,
5'-ACGUGACACGUUCGGAGAATT-3'. Additionally, an siRNA designed
specifically to target the PUM2 gene was employed, labeled siPUM2,
with the following sequences: Sense,
5'-AGCAUUAGAAUCUAUUUCUUCUGAT-3' and antisense,
5-AUCAGAAGAAAUAGAUUCUAAUGCUUU-3.
Cell culture and transfection
H9C2 rat cardiomyocyte cells (cat. no. CL-0089;
Procell Life Science & Technology Co., Ltd.) were cultured in a
controlled environment at 37˚C with a humidified atmosphere
containing 5% CO2. The culture medium utilized was DMEM
(cat. no. PM150210; Gibco; Thermo Fisher Scientific, Inc.),
supplemented with 10% FBS (cat. no. 10091148; Gibco; Thermo Fisher
Scientific, Inc.) and antibiotics including 100 µg/ml streptomycin
and 100 U/ml penicillin (cat. no. SV30010; HyClone™;
Cytiva). Cells in 12-well plates were transfected with 160 pmol
siRNA using Lipofectamine™ RNAiMAX (cat. no. 13778150;
Invitrogen; Thermo Fisher Scientific, Inc.) according to the
manufacturer's protocol. Following a 48-h incubation at room
temperature, the cells were harvested and analyzed by reverse
transcription-quantitative PCR (RT-qPCR) and western blotting.
Assessment of gene expression
Total RNA was extracted from H9C2 cells by the
TRIzol™ (Thermo Fisher Scientific, Inc.). The RNA was
further purified with two phenol-chloroform treatments and then
treated with RQ1 DNase (Promega Corporation) to remove DNA. The
quality and quantity of the purified RNA were redetermined by
measuring the absorbance at 260/280 nm (A260/A280) using
NanoPhotometer N50 (Implen NanoPhotometers). The integrity of RNA
was further verified by 1.0% agarose gel electrophoresis. cDNA was
synthesized utilizing a reverse transcription kit (cat. no.
R323-01; Vazyme Biotech Co., Ltd.). The procedure entailed an
initial incubation at 42˚C for 5 min, followed by 15 min at 37˚C
and a final step at 85˚C for 5 sec, all executed using the T100
thermocycler (Bio-Rad Laboratories, Inc.). Subsequent to cDNA
synthesis, qPCR was conducted using the ABI QuantStudio™
5 (conventional) system (Applied Biosystems; Thermo Fisher
Scientific, Inc.). The amplification protocol included a
denaturation step at 95˚C for 10 min, and 40 cycles comprising
denaturation at 95˚C for 15 sec and a combined annealing and
extension step at 60˚C for 1 min. Each sample was analyzed in
triplicate to ensure data reliability. To quantify the expression
levels of each transcript, normalization was performed using GAPDH,
and the 2-ΔΔCq method was used for data analysis
(25). Detailed sequences of
primers used for qPCR analysis are listed in Table SI.
Western blotting
H9C2 cells were lysed using ice-cold RIPA lysis
buffer (cat. no. PR20001; Proteintech Group, Inc.) supplemented
with protease inhibitors (cat. no. 4693116001; Sigma-Aldrich; Merck
KGaA). The lysis process was carried out on ice for 30 min to
disrupt the cells while simultaneously inhibiting any protease
activities that might lead to protein degradation. Following lysis,
cell samples underwent a heating step, during which they were
boiled for 10 min in conjunction with a protein loading buffer
(cat. no. P1040; Beijing Solarbio Science & Technology Co.,
Ltd.). This heating step was key for denaturing the proteins,
thereby facilitating their separation during subsequent analysis.
Prepared samples (25 µg protein/lane) were then carefully loaded
onto a 10% SDS-PAGE gel.Electrophoresis was carried out to separate
the proteins based on their molecular weights, allowing for
effective analysis of the protein profiles. Once electrophoresis
was completed, the separated proteins were transferred onto 0.45-µm
PVDF membranes (cat. no. ISEQ00010; MilliporeSigma) to enable
further examination. Following successful protein transfer, the
PVDF membrane was blocked with 5% skimmed milk to reduce
non-specific binding. This involved incubating the membranes at
room temperature for 1 h, which helped to minimize background noise
during the detection phase. Subsequently, the membranes were
incubated overnight at 4˚C with a specific primary GAPDH antibody
(1:1,000; cat. no. A19056; ABclonal Biotech Co., Ltd.) that
targeted PUM2 (1:1,000; cat. no. Ab92390; Abcam) and an actin
antibody (1:1,000; cat. no. 20536-1-AP; Proteintech Group, Inc.).
This overnight incubation allowed the primary antibodies time to
bind to their respective protein targets. Following this incubation
period, membranes were treated with an HRP-conjugated secondary
antibody, either anti-rabbit (1:10,000; cat. no. SA00001-2;
Proteintech Group, Inc.) or anti-mouse (1:10,000; cat. no. AS003;
ABclonal Biotech Co., Ltd.), for 45 min at room temperature. This
step was essential for enhancing the detection of the bound primary
antibodies. Finally, visualization of the proteins on the membranes
was achieved using an enhanced ECL reagent (BeyoECL Moon; cat. no.
P0018FM; Beyotime Biotechnology), which permitted chemiluminescent
detection of the proteins of interest.
Cell viability assay
In the present study, a cell viability assay was
conducted utilizing a Cell Counting Kit-8 (CCK-8; Shanghai Yeasen
Biotechnology Co., Ltd.). The procedure involved plating H9C2 cells
at a density of 10,000 cells/well in 96-well culture plates in a
controlled environment (37˚C and 5% CO2) for cell
viability. Cells were transfected with either a non-targeting
control siRNA [negative control (NC) group] or dicer-substrate
small interfering RNA (DSI; experimental group), while vials
lacking cells served as blank controls to ensure measurement
accuracy. Cells were incubated at 37˚C in a 5% CO2
atmosphere for designated time periods: 48 and 72 h. Following
incubation, 10 µl CCK-8 solution was added to each well for further
incubation for 3 h at 37˚C. This step allowed for the assessment of
cell viability. After incubation, the optical density (OD) was
measured using the ELx800 microplate reader (BioTek; Agilent
Technologies, Inc.) at a wavelength of 450 nm. To quantify the
viability rate of the cells, the following formaul was applied:
viability rate=(experimental OD value-blank OD value)/(control OD
value-blank OD value) x100. This formula enabled a clear assessment
of cell viability in response to varying treatments.
Flow cytometry analysis of
apoptosis
For the assessment of apoptosis, an annexin
V-allophycocyanin (APC)/7-aminoactinomycin D apoptosis detection
kit (cat. no. 40304ES60; Shanghai Yeasen Biotechnology Co., Ltd.)
was employed according to the manufacturer's instructions. H9C2
cells were cultivated in 6-well plates for 24 h at 37˚C and
subsequently transfected with siRNAs for 48 h as aforementioned.
The transfected and control siRNA cell populations were combined
with 5 µl annexin V-APC and incubated in the dark at room
temperature for 5 min, followed by the addition of 5 µl
7-amino-actinomycin for incubation for 5 min at room temperature.
Samples were then analyzed using the BD FACSCanto™
Clinical Flow Cytometry system (BD Biosciences) to evaluate the
levels of apoptosis, and the data were analyzed with
FlowJo™ software (BD Biosciences).
RNA extraction and sequencing
Total RNA was isolated from H9C2 cells using
TRIzol® reagent (cat. no. 15596026; Invitrogen; Thermo
Fisher Scientific, Inc.), following the protocol established by
Chomczynski and Sacchi (26).
Following RNA extraction, DNA digestion (37˚C for 10 min) was
carried out using RQ1 RNase-Free DNase (cat. no. M6101; Promega
Corporation). RNA quality was assessed by measuring the A260/A280
ratio with a Nanodrop™ OneC spectrophotometer
(Thermo Fisher Scientific, Inc.). Integrity was verified through
1.5% agarose gel electrophoresis. The concentration of selected RNA
samples was subsequently quantified using the Qubit™ 3.0
fluorometer (Thermo Fisher Scientific, Inc.) in combination with
the Qubit™ RNA broad range assay kit (cat. no. Q10210;
Thermo Fisher Scientific, Inc.). A total of 2 µg RNA was then used
for the preparation of stranded RNA-sequencing (RNA-seq) libraries,
utilizing the KC™ Stranded mRNA Library Prep Kit for
Illumina (cat. no. DR08402; Wuhan Kangce Technology) in accordance
with the manufacturer's protocol. PCR products with lengths between
200 and 500 base pairs were enriched, quantified and ultimately
sequenced by Novogene (Tianjin Sequencing Center and CAP Certified
Clinical Lab, Tianjin, China) using the NovaSeq 6000 sequencer
(Illumina, Inc.) and 150-bp paired-end sequencing was
performed.
RNA-seq data and differentially
expressed gene (DEG) analysis
Initially, raw sequencing reads that contained >2
ambiguous bases, denoted as ‘N’, were eliminated from the dataset
to ensure data quality. Following this step, adaptors and
low-quality base calls were systematically trimmed from the
remaining raw sequencing reads. This processing was carried out
using the FASTX-Toolkit (version 0.0.13; https://github.com/agordon/fastx_toolkit), which is
specifically designed for efficient read preprocessing.
Furthermore, any sequencing reads that were <16 nucleotides in
length were discarded to maintain the integrity of the dataset. The
clean reads that passed these quality control measures were then
aligned to the mRatBN7.2 genome using the HISAT2 tool (v2.2.1;
http://daehwankimlab.github.io/hisat2/), allowing for
the accommodation of ≥4 mismatches during the alignment process.
This alignment procedure ensured that the uniquely mapped reads
could be accurately utilized for counting gene read numbers and for
calculating the fragments per kilobase of transcript per million
fragments mapped (FPKM). During analysis of gene expression, the
‘DESeq2’ package (v 1.38.3; http://www.bioconductor.org/packages/release/bioc/html/DESeq2.html)
from R Bioconductor (v 3.22; https://www.bioconductor.org/) was employed to
identify DEGs using the thresholds P<0.01 and fold change
>1.5 or <0.67. These parameters were essential for
pinpointing variations in gene expression that could have
biological relevance.
Alternative splicing analysis
For the analysis of alternative splicing events
(ASEs) and specifically regulated ASEs (RASEs) among the samples,
the ABLas pipeline (https://github.com/ablifedev/ablas) was employed.
ABLas identifies 10 distinct forms of alternative splicing based on
the analysis of splice junction reads. These forms include exon
skipping (ES), intron retention (IntronR), alternative 5' splice
sites (A5SS), alternative 3' splice sites (A3SS) and mutually
exclusive exons (MXE), as well as variations in the 5' and 3'
untranslated regions (UTRs), specifically mutually exclusive 5'
UTRs and mutually exclusive 3' UTRs, in addition to cassette exons,
and combinations of A3SS and ES, and A5SS and ES.
To evaluate the influence of PUM2 on the ASEs,
statistical analysis was carried out using unpaired Student's
t-test to determine the significance of changes in the ratio of
ASEs across the samples. Events that demonstrated significant
alterations, with a P-value corresponding to a threshold that
aligned with a false discovery rate cut-off of 5%, were classified
as PUM2-regulated ASEs. This approach ensured that only those
events with statistical significance were recognized to be
potentially influenced by PUM2.
RT-qPCR verification
For RT-qPCR verification with GAPDH as the control
gene to measure relative target gene expression levels, total RNA
was isolated from H9C2 cells using TRIzol reagent (cat. no.
15596026; Invitrogen; Thermo Fisher Scientific, Inc.). cDNA was
synthesized with a reverse transcription kit (cat. no. R323-01;
Vazyme Biotech Co., Ltd.) on a mycycler (cat. no. T100; Bio-Rad
Laboratories, Inc.) at 42˚C for 5 min, 37˚C for 15 min and 85˚C for
5 sec. qPCR was performed on the ABI QuantStudio 5 with
Hieff® qPCR SYBR Green Master Mix, starting with
denaturing at 95˚C for 10 min, followed by 40 cycles of 95˚C for 15
sec and 60˚C for 1 min, with three technical replicates per sample.
Transcript levels were normalized to GAPDH using the
2-ΔΔCq method (25).
The primers for pre-mRNA splicing detection, detailed in Table SI, were designed to target splice
junctions between constitutive and alternative exons for precise
isoform amplification.
Functional enrichment analysis
To organize the functional classifications of DEGs,
the KOBAS 2.0 server (https://github.com/xmao/kobas) (19) was utilized to identify Gene
Ontology (GO) terms as well as Kyoto Encyclopedia of Genes and
Genomes (KEGG) pathways. The enrichment of each term was determined
using the hyper-geometric test along with the Benjamini-Hochberg
false discovery rate-controlling method.
Statistical analysis
All plots were conducted in R (version 4.2.3;
https://cran.r-project.org/bin/windows/base/old/4.2.3/)
as implemented in RStudio, in addition to pattern diagrams and
stacked bar charts. Pattern diagrams and stacked bar charts were
generated using GraphPad Prism (version 8.0; Dotmatics). Data are
presented as the mean ± standard error of the mean (SEM), with
three independent biological replicates. The statistical difference
between two groups for the RT-qPCR, cell viability and apoptosis
assays was calculated using an unpaired Student's t-test, where
P<0.05 was considered to indicate a statistically significant
difference. Other statistical methods used for the bioinformatics
analyses have been described in the corresponding sections; for
example, DEGs were calculated using DESeq2, and ASEs were analyzed
using unpaired t-tests.
Results
Inhibition of viability in H9C2 cells
by PUM2 knockdown
To explore the role of PUM2 in MF, H9C2 cell lines
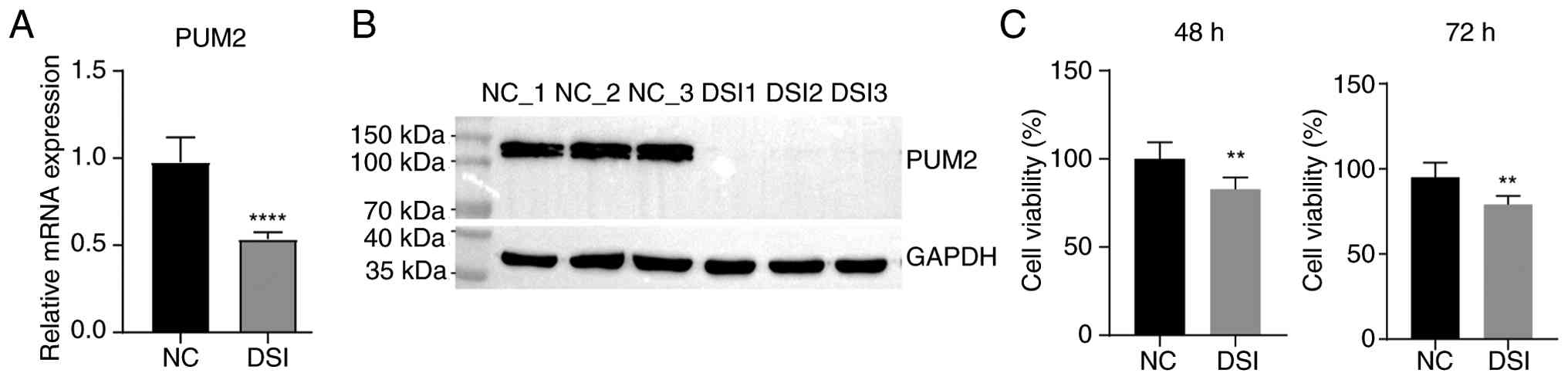
with PUM2 knockdown were constructed using siRNA transfection. Both
RT-qPCR and western blotting confirmed successful knockdown of PUM2
in H9C2 cells (Fig. 1A and
B). These efficiency validation
experiments demonstrated that mRNA expression was significantly
reduced (by ~50%) but not eliminated, whereas protein expression
was knocked down; however, this was not statistically assessed. The
rat PUM2 gene encodes five protein-coding transcripts, and the DSI
sequence used may specifically target only a subset of these
transcripts (27), while the
RT-qPCR assay measured total PUM2 expression at the gene level.
This selective targeting may explain why a larger reduction was
observed at the protein level compared with the mRNA level.
Consequently, expression from non-targeted transcripts masks the
extent of knockdown in the PCR results, leading to the observed
discrepancy between the RT-qPCR and western blot analyses. The
effect of PUM2 knockdown on viability and apoptosis of H9C2 cells
was detected using a CCK-8 assay and flow cytometry. The CCK-8
assay demonstrated that the viability of H9C2 cells was
significantly inhibited at both 48 and 72 h after transfection
following PUM2 knockdown (Fig.
1C). Flow cytometry revealed that PUM2 knockdown did not
significantly affect apoptosis at 48 h post-transfection (Fig. S1). The results indicated that PUM2
is key for the regulation of viability in H9C2 cells.
PUM2 knockdown regulates gene
expression in H9C2 cells
To explore the regulatory role of PUM2 in gene
expression, RNA-seq was conducted to assess and compare the overall
gene expression profiles between samples subjected to siPUM2
transfection and those in NC group. Utilizing the principal
component method, the FPKM values of all DEGs were analyzed, with
the first principal component (dimension 1) successfully
distinguishing the DSI group from the control group (Fig. 2A). The volcano plot illustrates the
DEGs identified between DSI and NC samples (Fig. 2B). A total of 147 DEGs influenced
by PUM2 were identified, encompassing 95 upregulated and 52
downregulated genes. Additionally, analysis of the heat map
depicting DEG expression patterns in the RNA-seq samples (Fig. 2C) revealed a high level of
consistency among the three biological replicates regarding
siPUM2-mediated transcription. Notably, the number of upregulated
genes markedly exceeded that of downregulated genes, implying that
PUM2 may exert a repressive effect on gene expression.
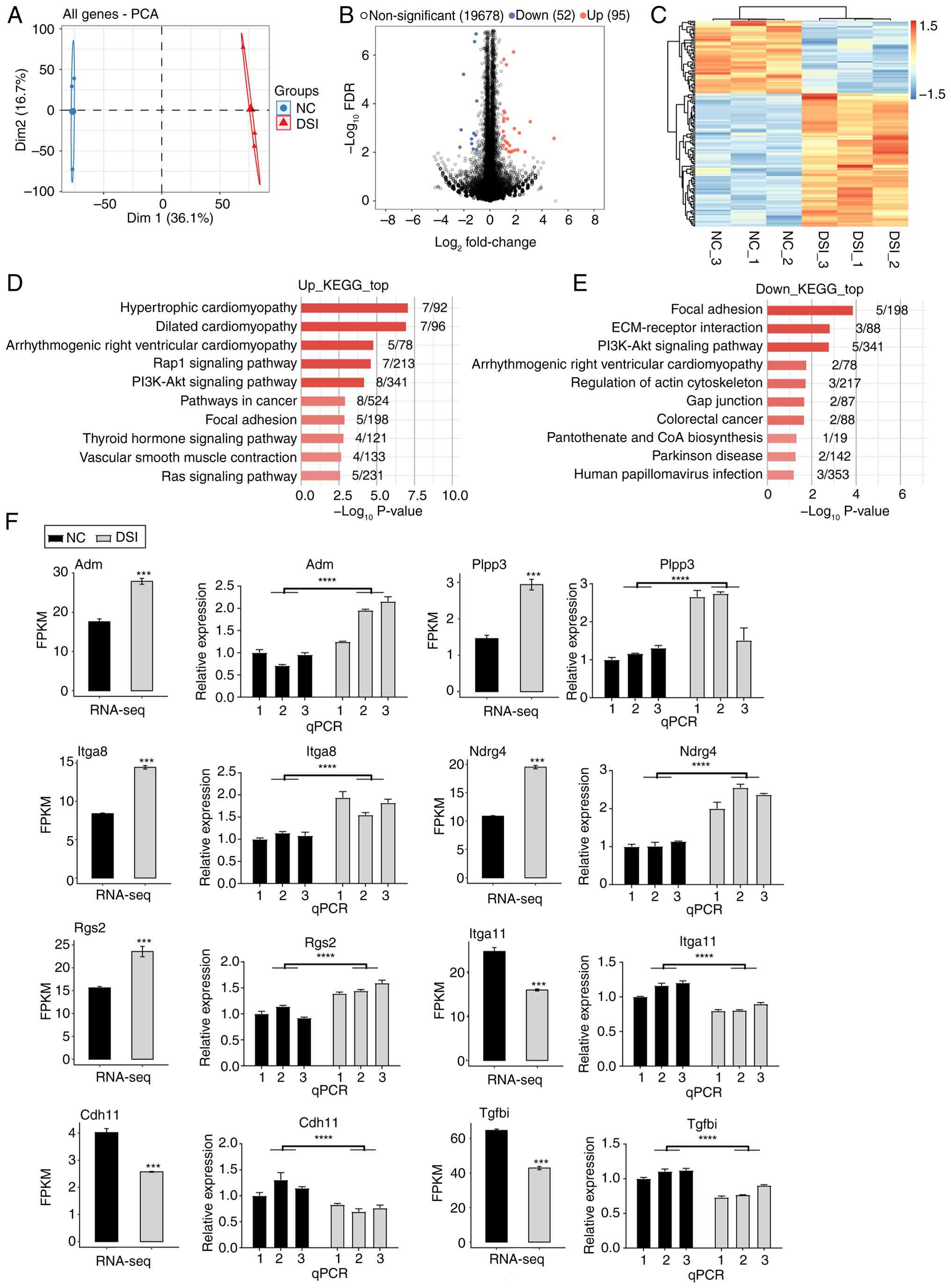 | Figure 2PUM2 knockdown regulates gene
expression in H9C2 cells. (A) PCA based on the FPKM value of all
genes. The ellipse for each group is the confidence ellipse. (B)
Volcano plot showing all DEGs between DSI and NC samples. (C)
Hierarchical clustering heat map showing expression levels of all
DEGs. (D) Bar plot showing the enriched KEGG pathways of
upregulated DEGs. (E) Bar plot showing the enriched KEGG pathways
of downregulated DEGs. (F) Bar plots showing the expression pattern
and statistical difference of DEGs with qPCR validation (numbers
represent three biological replicate samples). Data are presented
as the mean ± standard error of the mean. ***P<0.001
and ****P<0.0001. DSI, dicer-substrate small
interfering RNA; PUM2, pumilio RNA-binding family member 2; PCA,
principal component analysis; FPKM, fragments per kilobase of
transcript per million fragments mapped; DEGs, differentially
expressed genes; NC, negative control; KEGG, Kyoto Encyclopedia of
Genes and Genomes; Adm, adrenomedullin; Itga8, integrin subunit α
8; RNA-seq, RNA-sequencing; Dim1, dimension 1; Dim2, dimension 2;
FDR, false discovery rate; ECM, extracellular matrix; Rgs2,
regulator of G protein signaling 2; Cdh11, cadherin 11; Plpp3,
phospholipid phosphatase 3; Ndrg4, NDRG family member 4; Itga11,
integrin subunit α 11; Tgfbi, transforming growth factor β-induced;
qPCR, quantitative PCR. |
To evaluate the possible biological functions of
these DEGs, GO and KEGG enrichment analyses were conducted. GO
enrichment analysis indicated that the genes exhibiting increased
expression were predominantly associated with two pathways:
‘Positive regulation of GTPase activity’ and ‘positive regulation
of transcription from RNA polymerase II promoter’ (Fig. S2). KEGG analysis provided
comprehensive insights into the pathways associated with the
upregulated genes, indicating their notable involvement in a
multitude of key biological processes. The genes were heavily
represented in pathways related to various forms of cardiomyopathy,
such as ‘hypertrophic cardiomyopathy’, ‘dilated cardiomyopathy’ and
‘arrhythmogenic right ventricular cardiomyopathy’. Additionally,
analysis highlighted the inclusion of key signaling pathways,
including the ‘Rap1 signaling pathway’ and the ‘PI3K-Akt signaling
pathway’, both of which serve vital roles in cellular responses.
Furthermore, the involvement of these genes in ‘pathways in
cancer’, ‘focal adhesion’, the ‘thyroid hormone signaling pathway’,
‘vascular smooth muscle contraction’ and the ‘Ras signaling
pathway’ underscores their relevance in the complex interplay of
genetic and biochemical signals (Fig.
2D). Conversely, the examination of downregulated genes using
GO enrichment analysis revealed a primary connection to ‘positive
regulation of cell proliferation’ and ‘positive regulation of
transcription from RNA polymerase II promoter’ (Fig. S2). KEGG analysis identified
notable associations between the downregulated genes and a diverse
array of biological pathways, including ‘focal adhesion’ and
‘ECM-receptor interaction’, both of which are key for cellular
communication and structural integrity (28,29).
Additionally, the ‘PI3K-Akt signaling pathway’ was revealed to
serve a role in the regulation of these genes, along with pathways
associated with ‘arrhythmogenic right ventricular cardiomyopathy’
and the ‘regulation of the actin cytoskeleton’. Other notable
associations included pathways involved in ‘colorectal cancer’,
‘pantothenate and CoA biosynthesis’, ‘Parkinson disease’ and ‘human
papillomavirus infection’ (Fig.
2E). Specific pathways such as ‘hypertrophic cardiomyopathy’,
‘dilated cardiomyopathy’, ‘arrhythmogenic right ventricular
cardiomyopathy’, ‘vascular smooth muscle contraction’, ‘focal
adhesion’ (Fig. 2D), ‘ECM-receptor
interaction’ and ‘regulation of actin cytoskeleton’ (Fig. 2E) are particularly noteworthy as
they relate to molecular function. RNA-seq data for a
representative set of five upregulated genes, including
adrenomedullin (Adm), NDRG family member 4 (Ndrg4), phospholipid
phosphatase 3 (Plpp3), integrin subunit α 8 (Itga8) and regulator
of G protein signaling 2 (Rgs2), and three downregulated genes,
including cadherin 11 (Cdh11), integrin subunit α 11 (Itga11) and
transforming growth factor β-induced (Tgfbi), can be found in
Fig. 2F.
Pathway analysis revealed that several significantly
dysregulated genes are involved in key molecular functions.
Notably, the elevated genes Itga8 and Adm, along with the
downregulated gene Itga11 (identified in Tables I and II), were central components of the
enriched pathways related to cell adhesion and cardiovascular
regulation. To confirm the findings of RNA-seq, RT-qPCR was
conducted to assess the relative expression patterns of the genes
Adm, Itga8, Rgs2, Plpp3, Ndrg4, Itga11, Tgfbi and Cdh11 in both the
control and experimental groups. Results from the RT-qPCR
validation aligned with those obtained from the RNA-seq analysis,
affirming the reliability of the sequencing data (Fig. 2F). RT-qPCR amplification and
melting curves for all targets confirmed assay specificity and
reproducibility, validating the RNA-seq findings (data not
shown).
 | Table ISpecific gene information for
myocardial fibrosis-related pathways, as identified through Kyoto
Encyclopedia of Genes and Genomes analysis of upregulated
differentially expressed genes. |
Table I
Specific gene information for
myocardial fibrosis-related pathways, as identified through Kyoto
Encyclopedia of Genes and Genomes analysis of upregulated
differentially expressed genes.
| Term | Corrected
P-value | Input |
|---|
| Hypertrophic
cardiomyopathy | 0.00000884 | Itga8, Tnnt2,
Atp2a1, Itga1, Sgcd, Prkaa2, Itga2b |
| Dilated
cardiomyopathy | 0.00000884 | Itga8, Tnnt2,
Atp2a1, Itga1, Sgcd, Itga2b, Pln |
| Arrhythmogenic
right ventricular cardiomyopathy | 0.00085372 | Itga8, Itga1, Sgcd,
Atp2a1, Itga2b |
| Vascular smooth
muscle contraction | 0.03787322 | Myh1, Adm, Plcb4,
Prkcg |
 | Table IISpecific gene information linked to
myocardial fibrosis-related pathways, as identified through Kyoto
Encyclopedia of Genes and Genomes analysis of downregulated
differentially expressed genes. |
Table II
Specific gene information linked to
myocardial fibrosis-related pathways, as identified through Kyoto
Encyclopedia of Genes and Genomes analysis of downregulated
differentially expressed genes.
| Term | Corrected
P-value | Input |
|---|
| Focal adhesion | 0.00968071 | Col6a3, Pdgfb,
Itga11, Mylpf, Tnc |
| ECM-receptor
interaction | 0.03893884 | Col6a3, Itga11,
Tnc |
| Arrhythmogenic
right ventricular cardiomyopathy | 0.21945932 | Gja1, Itga11 |
| Regulation of actin
cytoskeleton | 0.21945932 | Pdgfb, Itga11,
Mylpf |
PUM2 regulates gene alternative
splicing in H9C2 cells
To explore how PUM2 knockdown influences the
regulation of alternative splicing, RNA-seq data were assessed to
identify PUM2-regulated RASEs in the H9C2 cell line. The
aforementioned ABLas pipeline was employed to characterize and
quantify ASEs and RASEs across the different samples. The types and
quantities of all notable RASEs between the NC and DSI samples are
illustrated in Fig. 3A. The
primary RASEs detected included IntronR (183 events), A3SS (129
events) and A5SS (100 events; Table
SII). These findings indicate that PUM2 regulates ASEs globally
in the H9C2 cell line. The heat map of splicing ratios for RASEs
demonstrated high reproducibility across biological triplicates in
siPUM2-treated cells (Fig. 3B). GO
analysis of PUM2-regulated alternative splicing genes (PUM2-RAS)
was carried out to explore the potential biological functions of
PUM2-RAS. Analysis revealed that the enriched pathways included
‘brain development’, ‘peptidyl-tyrosine dephosphorylation’,
‘axonogenesis’, ‘neural tube closure’, ‘mRNA processing’, ‘protein
stabilization’, ‘positive regulation of GTPase activity’, ‘negative
regulation of cell proliferation’, ‘protein phosphorylation’ and
‘cell proliferation’ (Fig. 3C).
Splicing ratios for several ASEs were significantly altered upon
PUM2 knockdown. Representative examples are presented in Figs. 3D and S3, including tropomyosin 1 (Tpm1; MXE),
actinin a 1 (Actn1; ES) and PPFIA binding protein 1 (Ppfibp1; ES).
The reads distribution diagram showed the distribution of splicing
junctions for Tpm1 and Actn1, with significant changes in the
splicing ratio (Fig. 3D). In
addition, the distribution of Ppfibp1 is also represented as a
reads distribution diagram, with the splicing ratio of Ppfibp1
presented as a bar plot (Fig.
S3). RT-qPCR validation of ASEs in the Ppfibp1, Actn1 and Tpm1
genes aligned with the RNA-seq results (Figs. 3D and S3). RT-qPCR amplification and melting
curves for all targets confirmed assay specificity and
reproducibility, therefore validating the RNA-seq findings (data
not shown).
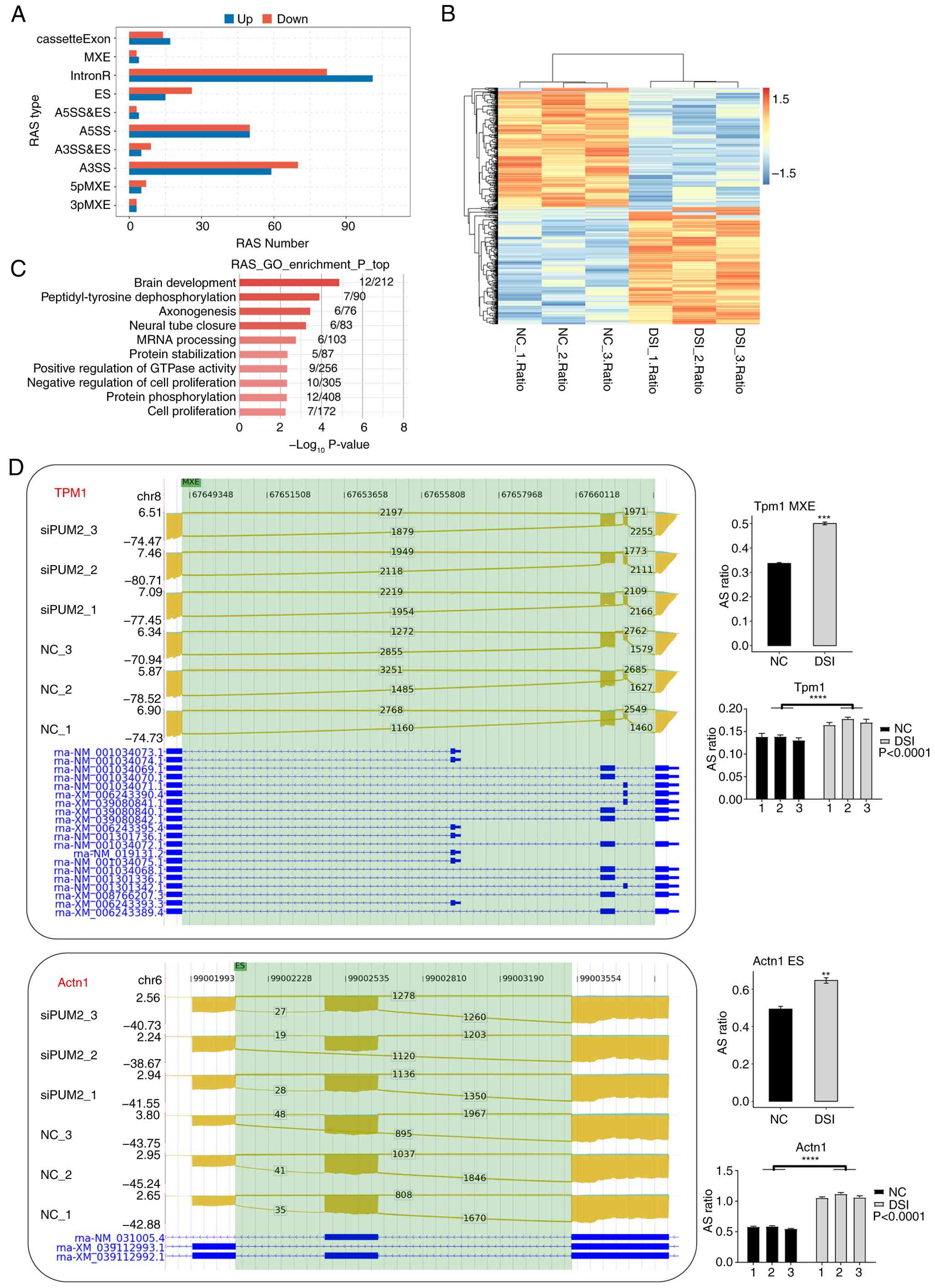 | Figure 3PUM2 regulates gene alternative
splicing in H9C2 cells. (A) Bar plot showing the number of all
notable regulated ASEs between DSI and NC samples. (B) Heat map of
splicing ratios for ASEs with hierarchical clustering. (C) Bar plot
showing the enriched GO biological process terms of RASG. (D) Reads
distribution diagram showing Actn1 (ES) and Tpm1 (MXE). Green
blocks indicate the positions where AS occurs (the connection lines
above are junction reads and splicing sites). The top shows the
absolute position of the chromosome base; the left side represents
the samples, and the vertical coordinate is the normalized reads;
the bottom provides transcript information. Bar plots showing
splicing ratio of Actn1 (ES) and Tpm1 (MXE) and quantitative PCR
validation. **P≤0.01, ***P≤0.001 and
****P<0.0001. DSI, dicer-substrate small interfering
RNA; PUM2, pumilio RNA-binding family member 2; NC, negative
control; ASE, alternative splicing event; GO, Gene Ontology, Actn1,
actinin α1; ES, exon skipping; Tpm1, tropomyosin 1; MXE, mutually
exclusive exon; intronR, intron retention; A5SS, alternative 5'
splice sites; A3SS, alternative 3' splice sites; 3pMXE, mutually
exclusive 3'UTRs; 5pMXE, mutually exclusive 5'UTRs; AS, alternative
splicing; RASG, regulated alternative splicing gene. |
Discussion
PUM2 is an abnormally expressed RBP in various
diseases (30). However, its
function and the corresponding mechanism of MF have not been fully
explored. To the best of our knowledge, the present study was the
first to investigate how PUM2 globally regulates the expression and
alternative splicing of genes associated with MF in H9C2 cells. The
present study examined the effects of PUM2 knockdown on gene
expression levels and alternative splicing patterns in rat H9C2
cells using high-throughput transcriptome sequencing. Cytological
experiment results indicated that PUM2 knockdown significantly
inhibited the viability of H9C2 cells. RNA-seq results demonstrated
that PUM2 knockdown led to 147 DEGs. Upregulation of the expression
levels of Adm, Rgs2, Itga8, Plpp3 and Ndrg4 genes, as well as
downregulation of the expression levels of Tgfbi, Itga11 and Cdh11
genes, was associated with the development of MF. Additionally,
numerous ASEs were found in genes associated with fibrosis, such as
Tpm1 (MXE) and Actn1 (ES). These results suggest that PUM2 can
influence the development of MF by regulating the expression of
associated genes and alternative splicing. PUM2 is a
post-transcriptional regulator that controls numerous genes
involved in key cellular pathways, including cell proliferation and
differentiation (31). PUM2 has
been shown to effectively inhibit translation and promote
G1 to S transition, resulting in a notable increase in
cell proliferation (32). In
addition, PUM2 has been identified as key in facilitating
cardiomyocyte apoptosis induced by hypoxia or reoxygenation
(33). By knocking down PUM2 in
rat H9C2 cells and carrying out cytological analyses, the present
study demonstrated that PUM2 is vital for the regulation of cell
viability. Results indicated that silencing PUM2 markedly decreased
cell viability without causing a notable change in apoptosis rates.
These observations suggest that PUM2 is fundamental in modulating
cell viability, primarily serving to enhance it, aligning with a
previous study (32).
We hypothesized that PUM2 may contribute to the
progression of MF by regulating the expression of MF-related genes.
Therefore, the aim was to identify key genes involved in MF
progression. RNA-seq analysis was conducted to identify DEGs
between the DSI and control groups. The results demonstrated that
PUM2 knockdown upregulated the expression levels of Adm, Itga8,
Rgs2 and Plpp3 genes, which have been reported to be involved in
the development of MF.
The present study revealed notable upregulation of
Adm in the ‘vascular smooth muscle contraction pathway’. Adm, a
multifunctional vasodilator peptide, reduces vascular tone and
participates in hormone secretion and inflammatory responses
(34). Adm may be a regulator of
fibrosis development via multiple pathways (35-37).
Furthermore, Adm expression has a protective effect against MF. Adm
has been reported to inhibit fibroblast proliferation and
ameliorate pressure overload-induced cardiac hypertrophy and MF
(38). Igta8 is a cell surface
protein belonging to the α-integrin family of transmembrane cell
surface receptors (39). Igta8 was
significantly enriched in the pathways of ‘hypertrophic
cardiomyopathy’, ‘dilated cardiomyopathy’ and ‘arrhythmogenic right
ventricular cardiomyopathy’. Previous studies have found that
aberrant Igta8 expression was associated with fibrosis (39,40).
In animals with angiotensin II (Ang II)-induced MF,
the expression of integrin α8β1 has been shown to be increased in
the left ventricle and arteries, suggesting that integrin α8β1 may
contribute to the fibrotic process in the heart (41). In addition, Rgs2 is a
multifunctional RGS protein involved in several G-protein signaling
pathways (42). It regulates
multiple cellular functions and behaviors in a range of
cardiovascular cells (43). It has
been shown that the expression of the Rgs2 gene is markedly
associated with the pro-fibrotic effect induced by TGF-β1 in
fibroblasts (44). In addition,
Rgs2 serves a key role as a negative regulator of Ang II-induced
cardiac fibroblast responses, thereby contributing to the
development of Ang II-induced fibrosis (45). In addition, the Plpp3 gene is
responsible for encoding lipid phosphate phosphatase 3 (LPP3), an
enzyme that is integral to the membrane and functions to
dephosphorylate glycerophospholipid and sphingolipid phosphate
esters (46). LPP3 activity is key
in vascular and cardiac development (47), and its expression regulates the
differentiation state of vascular smooth muscle cells and
influences fibroblast-like phenotypic transformation (48). Experimental evidence has
demonstrated that, in a myeloid-specific Plpp3-deficient mouse
model, the scar size and fibrosis area were markedly greater
compared with those in controls 30 days after myocardial infarction
(49). These findings indicate
that PLPP3, responsible for encoding LPP3, may have a protective
effect against MF.
PUM2 knockdown also upregulated the expression of
the Ndrg4 gene. Ndrg4 is a member of the NDRG gene family, which is
involved in cell proliferation, differentiation, development and
the stress response (50).
Research has revealed upregulation of Ndrg4 expression in liver
fibrosis (51); however, there are
limited studies on Ndrg4 in MF. DEGs that were downregulated, such
as Itga11, Tgfbi and Cdh11, require further research regarding
their involvement in the development of MF. The present study found
that the downregulated gene Itga11 was a component of several
fibrosis-related pathways identified by KEGG enrichment analysis,
including ‘focal adhesion’, ‘ECM-receptor interaction’,
‘arrhythmogenic right ventricular cardiomyopathy’ and ‘regulation
of actin cytoskeleton. Additionally, research shows that Itgα11 is
the primary collagen receptor on fibroblasts (52). Furthermore, the interaction between
Itgα11 and TGF-β2 signaling has been found to promote myofibroblast
differentiation in cardiac fibroblasts (53).
Tgfbi is an ECM protein that is upregulated by
TGF-β1(54) and is associated with
the TGF-β1 pathway and cardiac fibrosis (55). In addition, Cdh11 is a type II
cadherin that mediates calcium-dependent cell-cell adhesion
(56). Cdh11 deficiency suppresses
the activation and pro-fibrotic function of atrial fibroblasts,
resulting in a reduction of atrial fibrosis (57). A previous study has demonstrated
that cells expressing Cdh11 contribute to inflammation-driven
fibrotic remodeling following myocardial infarction (58).
In the present study, after PUM2 knockdown, the
expression levels of the Adm, Plpp3, Rgs2 and Itga8 genes were
upregulated. The expression of Adm, Plpp3 and Rgs2 genes tends to
be protective against MF, whereas Igta8 may exacerbate MF. Based on
the findings of the present study, it is proposed that the effects
of PUM2 on MF are the result of a combination of global gene
regulatory networks of PUM2. Future research endeavors should focus
on a comprehensive analysis of the molecular functions associated
with each specific target gene. Additionally, it is essential to
examine the context-dependent interactions that arise from
PUM2-mediated global modifications within a signaling pathway. Such
an approach will contribute to a deeper understanding of the
mechanisms through which PUM2 influences the development of MF. By
integrating these two perspectives, the intricate relationships and
dynamics at play can be explored, ultimately leading to a more
complete evaluation of the role of PUM2 in MF progression.
In the regulation of RBPs at the
post-transcriptional level, alternative splicing is essential. This
entails rearranging the exons of pre-mRNA in various configurations
to generate mRNA and protein variants that differ structurally and
functionally (59). This mechanism
is closely monitored, as non-coding regions of pre-mRNA are
eliminated while protein-coding parts are combined (60). Consequently, this approach leads to
the formation of proteins that exhibit distinct or even opposing
functionalities. The present study proposes that PUM2 could have a
notable influence on post-transcriptional regulation, with a
specific focus on alternative splicing. Analysis revealed that PUM2
affected a large number of ASEs in the H9C2 cell line.
Specifically, ≥530 transcripts were found to be regulated by PUM2.
The majority of the RASEs identified in the present study were
classified as IntronR (183 events), A3SS (129 events) and A5SS (100
events). Importantly, a number of ASEs were found in genes
associated with fibrosis, including Tpm1 and Actn1. Tpm1 is
involved in the development of various cardiac diseases, such as
hypertrophic and dilated cardiomyopathies (61,62),
and it has been suggested that cardiac fibrosis is notably reduced
after specific mutations in the lysine crotonylation locus of
Tpm1(63). In addition, Actn1 is a
cytoskeletal protein that serves an important role in mediating
sarcomere function (64). Actn1
could be an additional potential marker for myofibroblasts
(65); however, its role in MF
remains to be investigated. Ppfibp11, also known as liprin-β1, is a
ubiquitously expressed liprin (66) that may be associated with an
inflammatory myofibroblastic tumor (67). To the best of our knowledge,
limited reports exist regarding the involvement of Ppfibp1 in
fibrosis.
The present study, based on the alternative splicing
data, suggested that PUM2 may affect the progression of MF by
regulating the alternative splicing of genes such as Tpm1 and
Actn1. The aim of the present study was to investigate
PUM2-mediated ASEs, which are important post-transcriptional
regulatory mechanisms involved in the development of MF. These ASEs
may serve as important biomarkers and potential therapeutic targets
for MF. Although the global catalog of PUM2-RASEs was identified,
further studies are necessary to investigate the molecular
consequences of the affected transcripts. This will aid in
understanding the molecular functions of the affected genes and the
PUM2-mediated context-dependent regulatory network of ASEs.
PUM2 knockdown orchestrates multi-pathway
antifibrotic effects through coordinated transcriptional and
post-transcriptional regulation. Specifically, suppression of the
TGF-β pathway occurs through regulator of G-protein signaling 2
upregulation, which inhibits G q protein α subunit-mediated TGF-β
receptor signaling (45), combined
with Tgfbi downregulation that reduces ECM-bound TGF-β1
bioavailability (55),
collectively attenuating SMAD2/3 phosphorylation and
fibroblast-to-myofibroblast transition. Concurrently,
integrin-cytoskeleton integrity is compromised through Itga11
downregulation, impairing αvβ6 integrin-dependent latent TGF-β
activation, collagen I binding (68) and aberrant Actn1 splicing that
generates isoforms deficient in Z-disc binding domains, ultimately
disrupting mechanical force transmission and focal adhesion complex
assembly (64). Furthermore,
coordinated reduction of Cdh11 (diminishing cadherin-mediated
fibroblast aggregation) and Tgfbi (suppressing matricellular
signaling) synergistically limits fibrotic niche formation,
indicating the regulation of ECM remodeling by PUM2(56).
A key limitation of the present study is that while
H9C2 cells provide a tractable model for initial mechanistic
insights, they lack the multicellular interactions (such as
cardiomyocyte-fibroblast crosstalk) and hemodynamic stressors
present in vivo. Future studies should aim to validate the
role of PUM2 in pressure-overload murine models (such as transverse
aortic constriction) and human cardiac fibroblasts. Additionally,
the functional impact of PUM2-regulated splicing events (such as
Tpm1 MXE) on protein isoform activity warrants investigation
through isoform-specific knockdown. Despite these limitations, the
identification of PUM2 as a global regulator of MF-associated
transcripts and splicing events nominates it as a potential
therapeutic node. Pharmacological inhibition of PUM2 (for example,
using antisense oligonucleotides) could simultaneously target
multiple fibrotic pathways, overcoming the redundancy that limits
single-target approaches.
In conclusion, the properties of PUM2 as an RBP
suggest that it may have a multifunctional role in
post-transcriptional RNA regulation. In the present study, RNA-seq
was used to investigate the role of PUM2 in the H9C2 cell line and
it was concluded that it may contribute to the development of MF by
regulating the expression and alternative splicing of related
genes. The present study indicated that PUM2 was associated with
the development of MF and partially explained its molecular
mechanisms. Further molecular investigations of the target genes of
PUM2, as identified by the present RNA-seq experiment, could aid in
the development of PUM2 as a therapeutic target for MF.
Supplementary Material
Flow cytometry (A) plots and (B)
analysis demonstrating that PUM2 knockdown did not significantly
affect apoptosis at 48 h post-transfection. NC, negative control;
DSI, dicer-substrate small interfering RNA; ns, not significant;
PE-H, phycoerythrin fluorescence height; APC-H, allophycocyanin
height.
Bar plot showing the enriched GO terms
of upregulated and downregulated differentially expressed genes.
GO, Gene Ontology.
Reads distribution diagram showing
Ppfibp1 (ES) and bar plots showing splicing ratio of Ppfibp1 (ES)
and quantitative PCR validation. *P≤0.05 and
****P<0.0001. Ppfibp1, PPFIA binding protein 1; AS,
alternative splicing; ES, exon skipping; chr4, chromosome 4; NC,
negative control; DSI, dicer-substrate small interfering RNA; PUM2,
pumilio RNA-binding family member 2.
Primer sets related to the
experimental procedures: Reverse transcription-quantitative PCR
primers for gene expression quantification.
Classification of all regulated
alternative splicing events between siPUM2 and NC samples.
Acknowledgements
Not applicable.
Funding
Funding: The People's Hospital of Xinjiang Uygur Autonomous
Region (grant no. 20210208) funded the present study.
Availability of data and materials
The data generated in the present study may be found
in the (Gene Expression Omnibus database) under accession number
(GSE302720) or at the following URL: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE302720.
The remaining data generated in the present study may be requested
from the corresponding author.
Authors' contributions
The present study was conceived and designed by both
JL and ZW. JL and ZW confirm the authenticity of all the raw data.
The majority of the experiments and data analysis were carried out
by ZZ and HZ, who also drafted the initial manuscript. Data
collection was carried out by WZhu, DA, YL and WZho, who also
contributed to the drafting of the manuscript. All authors read and
approved the final version of the manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Tian G and Ren T: Mechanical stress
regulates the mechanotransduction and metabolism of cardiac
fibroblasts in fibrotic cardiac diseases. Eur J Cell Biol.
102(151288)2023.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Frangogiannis NG: Cardiac fibrosis.
Cardiovasc Res. 117:1450–1488. 2021.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Zou Y, Shi H, Li Y, Li T, Liu N and Liu B:
Heat shock protein 27 downregulation attenuates
isoprenaline-induced myocardial fibrosis and diastolic dysfunction
by modulating the endothelial-mesenchymal transition. Biochem
Pharmacol. 230(116612)2024.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Li C, Wang N, Rao P, Wang L, Lu D and Sun
L: Role of the microRNA-29 family in myocardial fibrosis. J Physiol
Biochem. 77:365–376. 2021.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Disertori M, Masè M and Ravelli F:
Myocardial fibrosis predicts ventricular tachyarrhythmias. Trends
Cardiovas Med. 27:363–372. 2017.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Zhao X, Kwan JYY, Yip K, Liu PP and Liu
FF: Targeting metabolic dysregulation for fibrosis therapy. Nat Rev
Drug Discov. 19:57–75. 2020.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Fang Z, Raza U, Song J, Lu J, Yao S, Liu
X, Zhang W and Li S: Systemic aging fuels heart failure: Molecular
mechanisms and therapeutic avenues. ESC Heart Fail. 12:1059–1080.
2025.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Liu M, López de Juan Abad B and Cheng K:
Cardiac fibrosis: Myofibroblast-mediated pathological regulation
and drug delivery strategies. Adv Drug Deliver Rev. 173:504–519.
2021.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Galeș LN, Păun MA, Butnariu I, Simion L,
Manolescu LSC, Trifănescu OG and Anghel RM: Next-Generation
sequencing in oncology-a guiding compass for targeted therapy and
emerging applications. Int J Mol Sci. 26(3123)2025.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Waarts MR, Stonestrom AJ, Park YC and
Levine RL: Targeting mutations in cancer. J Clin Invest.
132(e154943)2022.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Wang S, Sun Z, Lei Z and Zhang HT:
RNA-binding proteins and cancer metastasis. Semin Cancer Biol.
86:748–768. 2022.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Gebauer F, Schwarzl T, Valcárcel J and
Hentze MW: RNA-binding proteins in human genetic disease. Nat Rev
Genet. 22:185–198. 2021.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Chen X, Wu J, Li Z, Han J, Xia P, Shen Y,
Ma J, Liu X, Zhang J and Yu P: Advances in the study of RNA-binding
proteins in diabetic complications. Mol Metab.
62(101515)2022.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Smith JM, Sandow JJ and Webb AI: The
search for RNA-binding proteins: A technical and interdisciplinary
challenge. Biochem Soc Trans. 49:393–403. 2021.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Hu X, Wu P, Liu B, Lang Y and Li T:
RNA-binding protein CELF1 promotes cardiac hypertrophy via
interaction with PEBP1 in cardiomyocytes. Cell Tissue Res.
387:111–121. 2022.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Zhu H, Zhang Y, Zhang C and Xie Z:
RNA-binding profiles of CKAP4 as an RNA-binding protein in
myocardial tissues. Front Cardiovasc Med. 8(773573)2021.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Chen Z, He C, Gao Z, Li Y, He Q, Wang Y
and Cai C: Polypyrimidine tract binding protein 1 exacerbates
cardiac fibrosis by regulating fatty acid-binding protein 5. ESC
Heart Fail. 10:1677–1688. 2023.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Wang D, Ruan X, Liu X, Xue Y, Shao L, Yang
C, Zhu L, Yang Y, Li Z, Yu B, et al: SUMOylation of PUM2 promotes
the vasculogenic mimicry of glioma cells via regulating CEBPD. Clin
Transl Med. 10(e168)2020.PubMed/NCBI View Article : Google Scholar
|
|
19
|
D'Amico D, Mottis A, Potenza F, Sorrentino
V, Li H, Romani M, Lemos V, Schoonjans K, Zamboni N, Knott G, et
al: The RNA-binding protein PUM2 impairs mitochondrial dynamics and
mitophagy during aging. Mol Cell. 73:775–787.e10. 2019.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Ding Y, Yuan X and Gu W: Circular RNA
RBM33 contributes to cervical cancer progression via modulation of
the miR-758-3p/PUM2 axis. J Mol Histol. 52:173–185. 2021.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Liu Q, Liu Y, Li Y, Hong Z, Li S and Liu
C: PUM2 aggravates the neuroinflammation and brain damage induced
by ischemia-reperfusion through the SLC7A11-dependent inhibition of
ferroptosis via suppressing the SIRT1. Mol Cell Biochem.
478:609–620. 2023.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Gibb AA, Lazaropoulos MP and Elrod JW:
Myofibroblasts and fibrosis: Mitochondrial and metabolic control of
cellular differentiation. Circ Res. 127:427–447. 2020.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Tsoy S and Liu J: Regulation of protein
synthesis at the translational level: Novel findings in
cardiovascular biology. Biomolecules. 15(692)2025.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Chothani S, Schäfer S, Adami E,
Viswanathan S, Widjaja AA, Langley SR, Tan J, Wang M, Quaife NM,
Jian Pua C, et al: Widespread translational control of fibrosis in
the human heart by RNA-binding proteins. Circulation. 140:937–951.
2019.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Chomczynski P and Sacchi N: Single-step
method of RNA isolation by acid guanidinium
thiocyanate-phenol-chloroform extraction. Anal Biochem.
162:156–159. 1987.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Zhou Y, Ng DY, Richards AM and Wang P:
Loss of full-length pumilio 1 abrogates miRNA-221-induced gene p27
silencing-mediated cell proliferation in the heart. Mol Ther
Nucleic Acids. 27:456–470. 2021.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Tapial Martinez P, López Navajas P and
Lietha D: FAK structure and regulation by membrane interactions and
force in focal adhesions. Biomolecules. 10(179)2020.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Martisova A, Sommerova L, Krejci A,
Selingerova I, Kolarova T, Zavadil Kokas F, Holanek M, Podhorec J,
Kazda T and Hrstka R: Identification of AGR2 gene-specific
expression patterns associated with epithelial-mesenchymal
transition. Int J Mol Sci. 23(10845)2022.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Smialek MJ, Ilaslan E, Sajek MP and
Jaruzelska J: Role of PUM RNA-binding proteins in cancer. Cancers
(Basel). 13(129)2021.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Galgano A, Forrer M, Jaskiewicz L, Kanitz
A, Zavolan M and Gerber AP: Comparative analysis of mRNA targets
for human PUF-family proteins suggests extensive interaction with
the miRNA regulatory system. PLoS One. 3(e3164)2008.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Lin K, Qiang W, Zhu M, Ding Y, Shi Q, Chen
X, Zsiros E, Wang K, Yang X, Kurita T and Xu EY: Mammalian Pum1 and
Pum2 control body size via translational regulation of the cell
cycle inhibitor Cdkn1b. Cell Rep. 26:2434–2450.e6. 2019.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Cao Y, Liu C, Wang Q, Wang W, Tao E and
Wan L: Pum2 mediates Sirt1 mRNA decay and exacerbates
hypoxia/reoxygenation-induced cardiomyocyte apoptosis. Exp Cell
Res. 393(112058)2020.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Wong HK, Tang F, Cheung TT and Cheung BM:
Adrenomedullin and diabetes. World J Diabetes. 5:364–371.
2014.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Wei Y, Tanaka M, Sakurai T, Kamiyoshi A,
Ichikawa-Shindo Y, Kawate H, Cui N, Kakihara S, Zhao Y, Aruga K, et
al: Adrenomedullin ameliorates pulmonary fibrosis by regulating
TGF-ß-smads signaling and myofibroblast differentiation.
Endocrinology. 162(bqab090)2021.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Kach J, Sandbo N, Sethakorn N, Williams J,
Reed EB, La J, Tian X, Brain SD, Rajendran K, Krishnan R, et al:
Regulation of myofibroblast differentiation and bleomycin-induced
pulmonary fibrosis by adrenomedullin. Am J Physiol Lung Cell Mol
Physiol. 304:L757–L764. 2013.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Nagae T, Mori K, Mukoyama M, Kasahara M,
Yokoi H, Suganami T, Sawai K, Yoshioka T, Koshikawa M, Saito Y, et
al: Adrenomedullin inhibits connective tissue growth factor
expression, extracellular signal-regulated kinase activation and
renal fibrosis. Kidney Int. 74:70–80. 2008.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Niu P, Shindo T, Iwata H, Iimuro S, Takeda
N, Zhang Y, Ebihara A, Suematsu Y, Kangawa K, Hirata Y and Nagai R:
Protective effects of endogenous adrenomedullin on cardiac
hypertrophy, fibrosis, and renal damage. Circulation.
109:1789–1794. 2004.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Hung CF, Wilson CL, Chow YH and Schnapp
LM: Role of integrin alpha8 in murine model of lung fibrosis. PLoS
One. 13(e0197937)2018.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Levine D, Rockey DC, Milner TA, Breuss JM,
Fallon JT and Schnapp LM: Expression of the integrin alpha8beta1
during pulmonary and hepatic fibrosis. Am J Pathol. 156:1927–1935.
2000.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Bouzeghrane F, Mercure C, Reudelhuber TL
and Thibault G: Alpha8beta1 integrin is upregulated in
myofibroblasts of fibrotic and scarring myocardium. J Mol Cell
Cardiol. 36:343–353. 2004.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Kehrl JH and Sinnarajah S: RGS2: A
multifunctional regulator of G-protein signaling. Int J Biochem
Cell Biol. 34:432–438. 2002.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Zhang P and Mende U: Functional role,
mechanisms of regulation, and therapeutic potential of regulator of
G protein signaling 2 in the heart. Trends Cardiovasc Med.
24:85–93. 2014.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Wu X, Gou H, Zhou O, Qiu H, Liu H, Fu Z
and Chen L: Human umbilical cord mesenchymal stem cells combined
with pirfenidone upregulates the expression of RGS2 in the
pulmonary fibrosis in mice. Respir Res. 23(270)2022.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Zhang P, Su J, King ME, Maldonado AE, Park
C and Mende U: Regulator of G protein signaling 2 is a functionally
important negative regulator of angiotensin II-induced cardiac
fibroblast responses. Am J Physiol Heart Circ Physiol.
301:H147–H156. 2011.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Mao G, Smyth SS and Morris AJ: Regulation
of PLPP3 gene expression by NF-κB family transcription factors. J
Biol Chem. 294:14009–14019. 2019.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Busnelli M, Manzini S, Parolini C,
Escalante-Alcalde D and Chiesa G: Lipid phosphate phosphatase 3 in
vascular pathophysiology. Atherosclerosis. 271:156–165.
2018.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Van Hoose PM, Yang L, Kraemer M, Ubele M,
Morris AJ and Smyth SS: Lipid phosphate phosphatase 3 in smooth
muscle cells regulates angiotensin II-induced abdominal aortic
aneurysm formation. Sci Rep. 12(5664)2022.PubMed/NCBI View Article : Google Scholar
|
|
49
|
Tripathi H, Shindo K, Donahue RR, Gao E,
Kuppa A, ElKammar M, Morris AJ, Smyth SS and Abdel-Latif A:
Myeloid-specific deletion of lipid Plpp3 (phosphate phosphatase 3)
increases cardiac inflammation after myocardial infarction.
Arterioscler Thromb Vasc Biol. 43:379–381. 2023.PubMed/NCBI View Article : Google Scholar
|
|
50
|
Melotte V, Qu X, Ongenaert M, van
Criekinge W, de Bruïne AP, Baldwin HS and van Engeland M: The N-myc
downstream regulated gene (NDRG) family: Diverse functions,
multiple applications. FASEB J. 24:4153–4166. 2010.PubMed/NCBI View Article : Google Scholar
|
|
51
|
Gibriel AA, Ismail MF, Sleem H, Zayed N,
Yosry A, El-Nahaas SM and Shehata NI: Diagnosis and staging of HCV
associated fibrosis, cirrhosis and hepatocellular carcinoma with
target identification for miR-650, 552-3p, 676-3p, 512-5p and 147b.
Cancer Biomark. 34:413–430. 2022.PubMed/NCBI View Article : Google Scholar
|
|
52
|
Popova SN, Barczyk M, Tiger CF, Beertsen
W, Zigrino P, Aszodi A, Miosge N, Forsberg E and Gullberg D:
Alpha11 beta1 integrin-dependent regulation of periodontal ligament
function in the erupting mouse incisor. Mol Cell Biol.
27:4306–4316. 2007.PubMed/NCBI View Article : Google Scholar
|
|
53
|
Talior-Volodarsky I, Connelly KA, Arora
PD, Gullberg D and McCulloch CA: α11 integrin stimulates
myofibroblast differentiation in diabetic cardiomyopathy.
Cardiovasc Res. 96:265–275. 2012.PubMed/NCBI View Article : Google Scholar
|
|
54
|
Lee SG, Chae J, Woo SM, Seo SU, Kim HJ,
Kim SY, Schlaepfer DD, Kim IS, Park HS, Kwon TK and Nam JO: TGFBI
remodels adipose metabolism by regulating the Notch-1 signaling
pathway. Exp Mol Med. 55:520–531. 2023.PubMed/NCBI View Article : Google Scholar
|
|
55
|
Gao QY, Zhang HF, Chen ZT, Li YW, Wang SH,
Wen ZZ, Xie Y, Mai JT, Wang JF and Chen YX: Construction and
analysis of a ceRNA network in cardiac fibroblast during fibrosis
based on in vivo and in vitro data. Front Genet.
11(503256)2021.PubMed/NCBI View Article : Google Scholar
|
|
56
|
Ruan W, Pan R, Shen X, Nie Y and Wu Y:
CDH11 promotes liver fibrosis via activation of hepatic stellate
cells. Biochem Biophys Res Commun. 508:543–549. 2019.PubMed/NCBI View Article : Google Scholar
|
|
57
|
Cao W, Song S, Fang G, Li Y, Wang Y and
Wang QS: Cadherin-11 deficiency attenuates Ang-II-induced atrial
fibrosis and susceptibility to atrial fibrillation. J Inflamm Res.
14:2897–2911. 2021.PubMed/NCBI View Article : Google Scholar
|
|
58
|
Schroer AK, Bersi MR, Clark CR, Zhang Q,
Sanders LH, Hatzopoulos AK, Force TL, Majka SM, Lal H and Merryman
WD: Cadherin-11 blockade reduces inflammation-driven fibrotic
remodeling and improves outcomes after myocardial infarction. JCI
Insight. 4(e131545)2019.PubMed/NCBI View Article : Google Scholar
|
|
59
|
Fu XD and Ares M Jr: Context-dependent
control of alternative splicing by RNA-binding proteins. Nat Rev
Genet. 15:689–701. 2014.PubMed/NCBI View Article : Google Scholar
|
|
60
|
Li WJ, Huang Y, Lin YA, Zhang BD, Li MY,
Zou YQ, Hu GS, He YH, Yang JJ, Xie BL, et al: Targeting
PRMT1-mediated SRSF1 methylation to suppress oncogenic exon
inclusion events and breast tumorigenesis. Cell Rep.
42(113385)2023.PubMed/NCBI View Article : Google Scholar
|
|
61
|
Lorenzini M, Norrish G, Field E, Ochoa JP,
Cicerchia M, Akhtar MM, Syrris P, Lopes LR, Kaski JP and Elliott
PM: Penetrance of hypertrophic cardiomyopathy in sarcomere protein
mutation carriers. J Am Coll Cardiol. 76:550–559. 2020.PubMed/NCBI View Article : Google Scholar
|
|
62
|
Rajan S, Jagatheesan G, Petrashevskaya N,
Biesiadecki BJ, Warren CM, Riddle T, Liggett S, Wolska BM, Solaro
RJ and Wieczorek DF: Tropomyosin pseudo-phosphorylation results in
dilated cardiomyopathy. J Biol Chem. 294:2913–2923. 2019.PubMed/NCBI View Article : Google Scholar
|
|
63
|
Cai W, Xu D, Zeng C, Liao F, Li R, Lin Y,
Zhao Y, Dong W, Wang Q, Yang H, et al: Modulating lysine
crotonylation in cardiomyocytes improves myocardial outcomes. Circ
Res. 131:456–472. 2022.PubMed/NCBI View Article : Google Scholar
|
|
64
|
Xie GF, Zhao LD, Chen Q, Tang DX, Chen QY,
Lu HF, Cai JR and Chen Z: High ACTN1 is associated with poor
prognosis, and ACTN1 silencing suppresses cell proliferation and
metastasis in oral squamous cell carcinoma. Drug Des Devel Ther.
14:1717–1727. 2020.PubMed/NCBI View Article : Google Scholar
|
|
65
|
Wettlaufer SH, Scott JP, McEachin RC,
Peters-Golden M and Huang SK: Reversal of the transcriptome by
prostaglandin E2 during myofibroblast dedifferentiation. Am J
Respir Cell Mol Biol. 54:114–127. 2016.PubMed/NCBI View Article : Google Scholar
|
|
66
|
Dong C, Li X, Yang J, Yuan D, Zhou Y,
Zhang Y, Shi G, Zhang R, Liu J, Fu P and Sun M: PPFIBP1 induces
glioma cell migration and invasion through FAK/Src/JNK signaling
pathway. Cell Death Dis. 12(827)2021.PubMed/NCBI View Article : Google Scholar
|
|
67
|
Yoshida A, Shibata T, Tsuta K, Watanabe SI
and Tsuda H: Inflammatory myofibroblastic tumour of the lung with a
novel PPFIBP1-ALK fusion variant. Histopathology. 63:881–883.
2013.PubMed/NCBI View Article : Google Scholar
|
|
68
|
Diehl P, Fricke A, Sander L, Stamm J,
Bassler N, Htun N, Ziemann M, Helbing T, El-Osta A, Jowett JB and
Peter K: Microparticles: Major transport vehicles for distinct
microRNAs in circulation. Cardiovasc Res. 93:633–644.
2012.PubMed/NCBI View Article : Google Scholar
|