Introduction
Abdominal aortic aneurysm (AAA), defined as the
regional enlargement of the abdominal aorta to a diameter of
>50% greater compared with its adjacent healthy size or an aorta
that measures >30 mm in diameter (1,2), is
a key cause of mortality among individuals aged >65 years, due
to rupture of the dilation aorta tissue. In addition, the mortality
rate of patients with AAA is >80% following rupture, despite
surgical advancements (3).
Effective pharmacotherapies for halting the growth and rupture of
AAA or delaying the requirement for surgical repair are lacking at
present.
The pathogenesis of vascular injury in aortic
aneurysms involves the apoptosis of vascular smooth muscle cells
(VSMCs) and the enhancement of extracellular matrix
metalloproteinase (MMP) activity, oxidative stress aggravated by
endothelial cell dysfunction and the increase of inflammatory cell
infiltration in the vascular wall. At present, mitochondrial
dysfunction is considered to carry out a vital role in the
development of AAA; results of a previous study using single-cell
RNA sequencing demonstrated that excessive mitochondrial
dysfunction occurs in different aortic cell types and is a key
character of aortic aneurysms (4).
The regulation of mitochondrial metabolism in VSMCs is involved in
the pathogenesis of AAA (5,6);
VSMCs derived from patients with AAA exhibited increased
mitochondrial fragmentation, elevated reactive oxygen species (ROS)
production and DNA damage, highlighting the role of mitochondrial
health in AAA progression (7). In
addition, results of a previous study demonstrated that angiotensin
II (Ang II) induced mitochondrial dysfunction in VSMCs,
contributing to AAA development; consequently, restoring
mitochondrial function in VSMCs attenuated AAA formation in mice
models infused with Ang II, emphasizing the therapeutic potential
of targeting mitochondrial pathways (8).
Wingless/β catenin (Wnt/β-catenin) signaling is
often involved in the pathological progression of vascular diseases
associated with endothelial dysfunction, proliferation (9), macrophage activation (10) and VSMC migration (11). Moreover, Wnt/β-catenin signaling
was involved not only in chronic kidney disease-associated vascular
calcification (12), but also in
the osteogenic conversion of VSMCs (13). Through forming a complex with
receptors, such as LRP5/6 and Kremen, in the cell membrane,
Dickkopf-1 (DKK1) inhibited Wnt/β-catenin signaling (14). Results of a prospective,
population-based clinical cohort study revealed that elevated
baseline levels of DKK1 were independently associated with the
occurrence of cardiovascular events, providing clinical
implications for investigating its role in the pathogenesis of
cardiovascular disease (15).
Notably, numerous investigations have highlighted the association
between elevated plasma levels of DKK1 and the presence of
cardiovascular diseases (16,17).
DKK1 was involved in vascular calcification, endothelial cell
interstitial transformation and mitochondrial disorder in vascular
endothelial cells (18). However,
it is still unknown whether DKK1 is involved in the regulation of
AAA.
While DKK1 is primarily recognized for its role in
Wnt signaling, emerging evidence suggests involvement in
mitochondrial regulation (18).
However, the specific interaction between DKK1 and single-pass
membrane protein with aspartate rich tail 1 (SMDT1), a component of
the mitochondrial calcium uniporter (MCU) complex, in the context
of Ang II-induced mitochondrial injury in VSMCs, is novel and not
extensively explored in current literature. The present study
explored whether DKK1/SMDT1 mediated Ang II-induced mitochondrial
injury in primary cultured HSMCs, thereby clarifying DKK1 whether
could serve as a therapeutic target for the prevention of AAA.
Materials and methods
Cell culture and treatment
Passage 2 human aortic smooth muscle cells (HSMCs;
cat. no. CP-H081; https://www.procell.com.cn/p/human-aortic-smooth-muscle-cells-cp-h081-69769;
Procell Life Science & Technology Co., Ltd.) which carried the
SV40T gene were used in the present study. Cells from passages 4 to
10 were cultured in smooth muscle cell medium (cat. no. 1101;
ScienCell Research Laboratories, Inc.) in a 5% CO2
incubator at 37˚C. HSMCs were stimulated by human Ang II (1 µM;
cat. no. HY-13948; MedChemExpress) for 24 h; the concentration of
Ang II was chosen based on a previous study (19). The HSMCs isolated by Procell
laboratory were identified by immunofluorescence for α-SMA, with a
purity of >90%, and free of HIV-1, HBV, HCV, mycoplasma,
bacteria, yeast and fungi (data not shown).
Western blot assay
Cell homogenates were lysed in an ice-cold
radioimmunoprecipitation lysis buffer (cat. no. AR0102; Wuhan
Boster Biological Technology Co., Ltd.) with 1 mM
phenylmethanesulphonyl fluoride (liquid, 100x, cat. no. AR1192;
Wuhan Boster Biological Technology Co., Ltd.) and then the total
protein concentration was measured by a BCA protein concentration
kit (cat. no. P0010; ProteinTech Group, Inc.). Protein extracts (30
µg per lane) were loaded and separated by 8-12% SDS-PAGE and
transferred to a PVDF membrane (cat. no. IPVH00010;
MilliporeSigma). Next the membranes were blocked in 5% skimmed milk
powder for 15 min at room temperature (RT), and then were incubated
with primary antibodies at 4˚C overnight; the primary antibodies
included anti-DKK1 (1:1,000; cat. no. ab109416; Abcam) and GAPDH
(1:5,000; cat. no. 60004-1-Ig; ProteinTech Group, Inc.). Membranes
were then incubated with secondary antibodies at RT for 2 h the
next day and included HRP-conjugated Affinipure Goat Anti-Mouse IgG
(H+L), (1:5,000; cat. no. SA00001-1; ProteinTech Group, Inc.) and
HRP-conjugated Affinipure Goat Anti-Rabbit IgG (H+L) (1:10,000;
cat. no. ZB-2301; Beijing Zhongshan Jinqiao Biotechnology Co.,
Ltd.). The bands were exposed using Tanon5200 (Tanon Science and
Technology Co., Ltd.). Densitometry analysis was performed for
semi-quantification using ImageJ 1.53t software (National
Institutes of Health), using GAPDH as the internal control. The
experiments were performed in triplicate.
RNA extraction and reverse
transcription-quantitative PCR (RT-qPCR)
Total RNA was isolated from HSMCs using an Ultrapure
RNA Kit (cat. no. CW0581S; Jiangsu CoWin Biotech Co., Ltd.)
according to the manufacturer's instructions and then converted to
cDNA using an Evo M-MLV RT Master Mix (cat. no. AG11706; Accurate
Biology Co., Ltd.) at 37˚C for 15 min and 85˚C for 5 sec. After RT,
qPCR was performed with a CFX connect Real-Time PCR System (Bio-Rad
Laboratories, Inc.) using SYBR Green Pro Taq HS Premix (cat. no.
AG11701; Accurate Biology Co., Ltd.). The thermocycling conditions
were 95˚C for 5 sec and then 60˚C for 30 sec for 40 cycles,
following a step at 95˚C for 30 sec. GAPDH was the internal
control, and the primer sequences are provided in Table I. The relative fold change was
quantified using the 2-∆∆Cq method (20); the experiments were performed in
triplicate.
 | Table IPCR primer sequences. |
Table I
PCR primer sequences.
| Gene | Accession no. | Forward primer (5'
to 3') | Reverse primer (5'
to 3') |
|---|
| Human DKK1 | NM_012242.4 |
AGGCACGCTATGTGCTG |
CAGTGTGGTTCTTCTGGGA |
| Human IL-1β | NM_000576.3 |
AGCACCTTCTTTCCCTTCATCTT |
CACCACTTGTTGCTCCATATCCT |
| Human GRP-78 | NM_005347.5 |
TATTGGAGGTGGGCAAACAAAGA |
CAGCAATAGTTCCAGCGTCTTTG |
| Human GRP-94 | NM_003299.3 |
TGACAGAATCTCCGTGTGCTTTG |
CAGCGGGTGTCTGGGATTAATTT |
| Human SMDT1 | NM_033318.5 |
TCTCCGTGTGTTCTCCATTGT |
TAGTCATCATCATCATCATCCTCTG |
| Human GAPDH | NM_002046.7 |
GGTGAAGGTCGGTGTGAACG |
CTCGCTCCTGGAAGATGGTG |
Transfection of HSMCs with small
interfering RNA (siRNA) and vector
siRNAs targeting human DKK1
(5'-GCUUCACACUUGUCAGAGATT-3') with a high silencing efficiency and
scrambled siRNAs as negative control (NC) were purchased from
Changzhou Ruibo Bio-Technology Co., Ltd. When HSMCs reached 70-80%
confluence, they were transfected with these siRNAs (50 nM) using
Lipofectamine RNAi MAX (Invitrogen; Thermo Fisher Scientific, Inc.)
according to the manufacturer's instructions. After the 6 h of
transfection at 37˚C, the medium was replaced with smooth muscle
cell medium. Sufficiently downregulated expression caused by these
siRNAs was verified by RT-qPCR 48 h after transfection. The
scrambled siRNAs were used as controls across all steps.
RNA sequencing (RNA-seq)
Total RNA was extracted from each group of HSMCs
transfected with siRNA using the VAMNE Magnetic Universal Total RNA
Kit (cat. no. ROA3301; Vazyme Biotech Co., Ltd.). The purity of RNA
samples was detected by NanoPhotometer® NP80 ultramicro
spectrophotometer (Implen GmbH). The concentration of RNA samples
was detected by Qubit®3.0 Flurometer (Life Technologies;
Thermo Fischer Scientific, Inc.). The integrity of RNA samples was
determined by the Agilent 2100 RNA Nano 6000 Assay Kit (Agilent
Technologies, Inc.). After the total RNA samples were qualified,
the mRNAs were purified by poly-A selection and rRNA depletion and
the library was constructed. The loading concentration of the final
library was 20 pM, as measured by iQ™ SYBR®
Green Supermix (cat. no. 1708882; Bio-Rad Laboratories, Inc.) on a
CFX 96 Real-Time PCR System (Bio-Rad Laboratories, Inc.) The
sequencing was performed using a NovaSeq 6000 S4 Reagent Kit V1.5
(cat. no. 20028312; Illumina, Inc.) using Next Generation
Sequencing on an Illumina NovaSeq (Illumina, Inc.) sequencing
platform. The nucleotide length was 150 bp and the direction of
sequencing was paired end. Raw sequencing data were transformed to
FastQ format, and the FastQ files were checked for quality
assessment by FastQC (v0.11.7; http://www.bioinformatics.babraham.ac.uk/projects/fastqc).
The Phred Quality Score (Q) of 30 indicates that the base
sequencing error rate is 0.1%. Q30 percentage >85% indicates
that the sequencing quality is up to standard. The data were
filtered by removal of adaptor sequences and low-quality reads in
order to get the clean data; the adaptor sequences (the minimum
required adapter overlap/stringency: 5bp) and low quality reads
(the cutoff of Q value=19) were removed by Trim Galore software (v
0.6.1; https://www.bioinformatics.babraham.ac.uk/projects/trim_galore/)
and Cutadapt (21). The duplicate
sequences were not removed, because removing duplicates might lead
to the loss of important biological information, especially for
highly expressed genes (22).
Before analysis, the sequence was aligned, and the abundance of
transcripts was quantified; the clean data was aligned to the
GRCh38 human reference genome in Ensemble (http://www.ensembl.org/index.html) using HISAT2
(v2.1.0.; https://github.com/DaehwanKimLab/hisat2). Reads count
for each gene in each sample was counted by HTSeq (v2.0.0,
https://htseq.readthedocs.io/en/latest/) and FPKM
(fragments per kilobase million mapped reads) was then calculated
to estimate the expression level of genes in each sample. There
were four groups from different experimental conditions for HMSCs
with two biological replicates in each group: Negative control
(NC), siDKK1, NC-Ang II and siDKK1-Ang II. The experimental groups
were based on previously published research (18).
Differentially expressed gene (DEG)
analysis
DESeq2 (v1.20.0; https://www.bioconductor.org/packages/devel/bioc/html/DESeq2.html)
was used to analyze the expression of DEGs, and the fold change
value and adjusted P-value were used as the main indicators.
|log2 fold change|≥1 and adjusted P-value <0.05 were
selected as the thresholds. The results of DEG analysis were
visualized using the ‘ggplot2’ R package (v4.2.1; https://cran.r-project.org/package=ggplot2) (23).
Venn diagram analysis
Overlapping genes between each group of data were
visualized using the ‘ggplot2’ R package and the ‘VennDiagram’ R
package (v1.7.3; https://CRAN.R-project.org/package=VennDiagram)
(24).
Robust rank aggregation (RRA)
analysis
RRA algorithm was used to sort and merge multiple
data sets, to screen out the key genes from the DEGs and visualize
them in the form of heat maps using the ‘RobustRankAggreg’ R
package (v1.2.1; https://CRAN.R-project.org/package=RobustRankAggreg)
(25) and the ‘ggplot2’ R
package.
Functional enrichment analysis
Gene Ontology (GO; https://geneontology.org) and Kyoto Encyclopedia of
Genes and Genomes (KEGG; https://www.kegg.jp) pathway enrichment analysis of
the screened genes were performed using the ‘clusterProfiler’ R
package (v4.2.0; https://www.bioconductor.org/packages/devel/bioc/html/clusterProfiler.html)
(26). The adjusted P<0.05 was
set as the cut-off criterion.
Protein-protein interaction (PPI)
network analysis
STRING (https://string-db.org/) database was utilized to
analyze the PPI data. Firstly, the ‘multiple proteins’ option was
selected and the genes in the top 60 RRA scores were then input
into the STRING database (27),
and ‘Homo sapiens’ was selected in the species option. The obtained
PPI data were illustrated into a high-level network diagram using
the ‘igraph’ R package (v1.4.0; https://cran.r-project.org/web/packages/igraph/index.html)
and the ‘ggraph’ R package (v2.2.0; https://cran.r-project.org/web/packages/ggraph/index.html).
Statistical analysis
The data in the in vitro experiments are
presented as the mean ± SE from three independent experiments. The
statistical analysis was performed by unpaired student's t-tests or
two-way ANOVA followed by post hoc Tukey's multiple comparisons
test in Figs. 2B and 5F using GraphPad Prism 9.4 software
(Dotmatics). P<0.05 was considered to indicate a statistically
significant difference. Benjamini-Hochberg multiple corrections
were applied in DEG analysis.
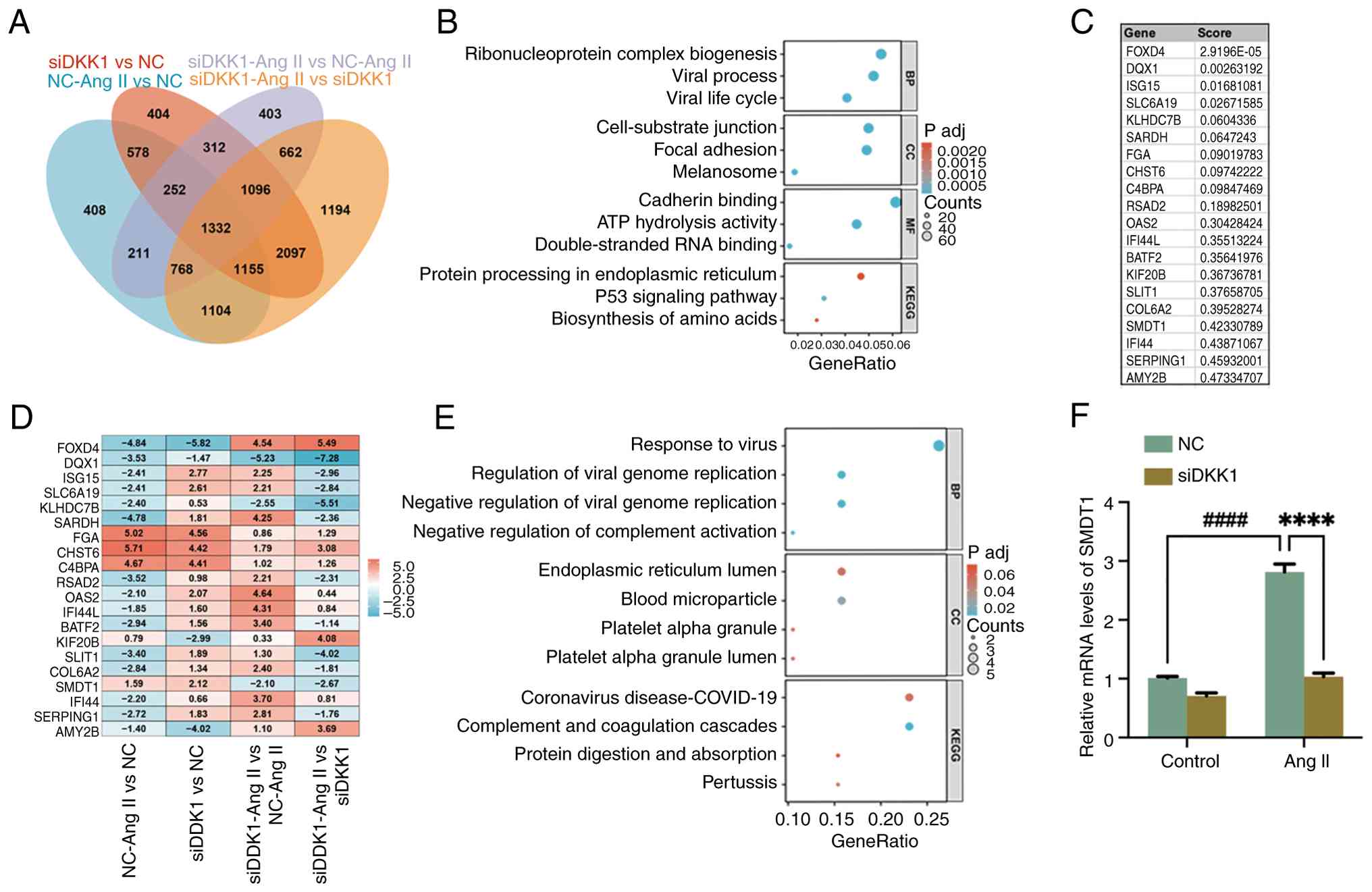 | Figure 5Identification of downstream key
genes after DKK1 silencing. (A) Venn diagram showed co-expressed
DEGs. (B) Functional enrichment analysis of GO and KEGG of
co-expressing DEGs. (C) The top 20 genes in RRA analysis. (D) Heat
map of RRA analysis. (E) Functional enrichment analysis of GO and
KEGG of the top 20 genes in RRA analysis. (F) Reverse
transcription-quantitative PCR results showed the mRNA expression
level of SMDT1. ****P<0.0001;
####P<0.0001. NC, treated with scrambled siRNAs as
negative control; siDKK1, treated with siRNAs targeting human DKK1;
Ang II, treated with Ang II (1 µM). GO, gene ontology; KEGG, Kyoto
Encyclopedia of Genes and Genomes; NC, negative control; siDKK1,
small interfering RNA for DKK1; DKK1, Dickkopf-1; Ang II,
angiotensin II; BP, biological processes; CC, cellular components;
MF, molecular functions; DEGS, differentially expressed genes; RRA,
robust rank aggregation; SMDT1, single-pass membrane protein with
aspartate rich tail 1. |
Results
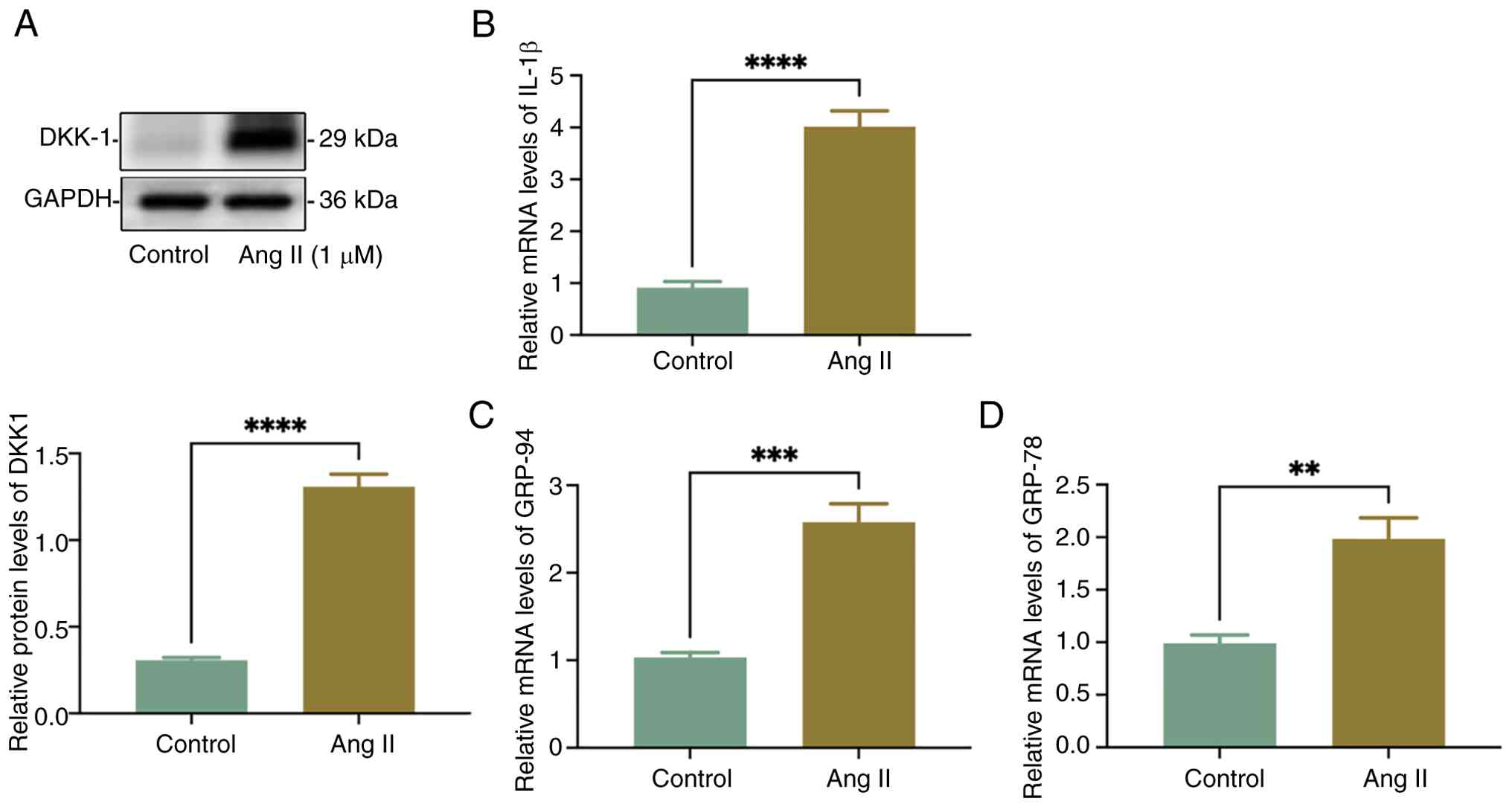
Ang II promotes DKK1 expression and
mitochondrial damage-associated mRNAs in HSMCs
HSMCs were stimulated with 1 µm/l Ang II for 24 h.
Results of the western blot analysis revealed that DKK1 protein
expression was significantly increased by 4.26-fold in HSMCs
following Ang II stimulation (Fig.
1A). Furthermore, mRNA expression levels of IL-1β and
mitochondrial damage-associated genes, GRP-94 and GRP-78, were
significantly increased in HSMCs following Ang II stimulation
(Fig. 1B-D). While, the actual
mitochondrial damage was not shown.
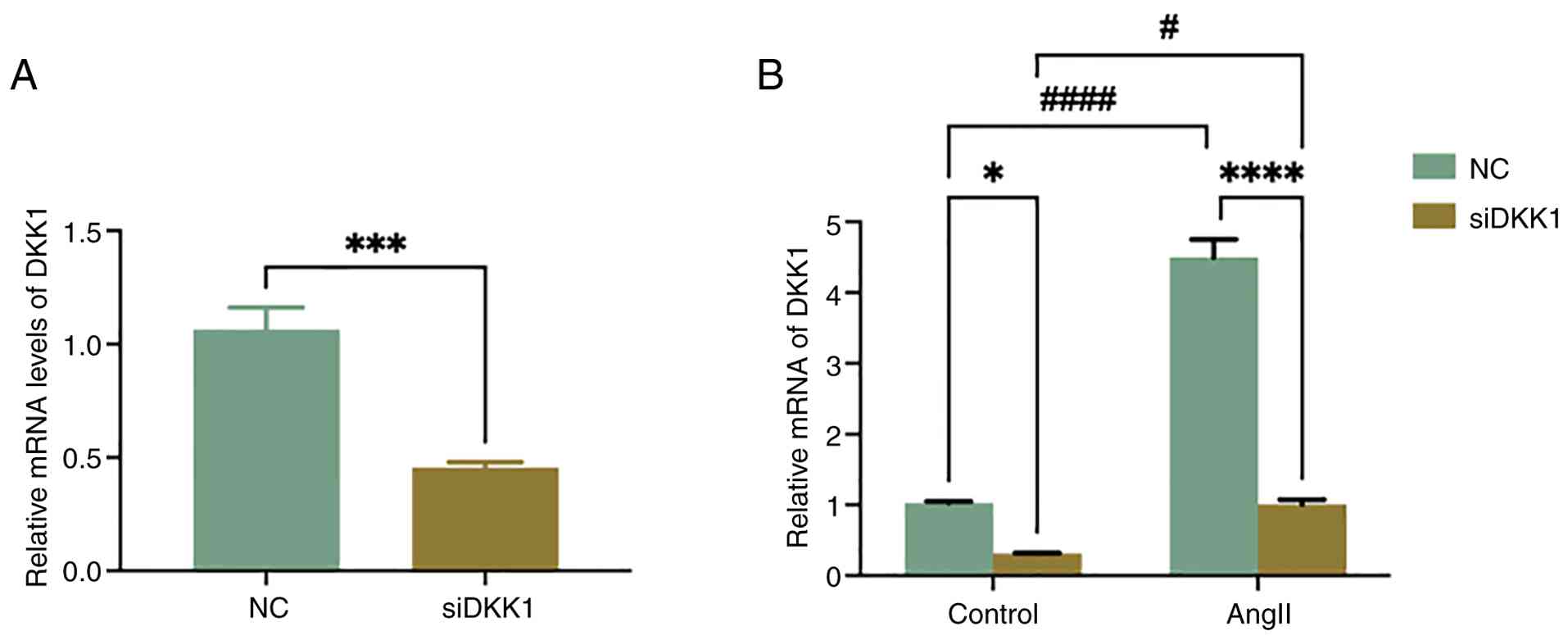
Transfection efficiency of siDKK1
prior to RNA-seq
Following transfection with siDKK1, expression
levels of DKK1 in HSMCs were significantly decreased (Fig. 2A), indicating that siDKK1
effectively reduced the expression of DKK1. Subsequently, cells
were divided into four groups for RNA sequencing: NC, siDKK1,
NC-Ang II and siDKK1-Ang II. Prior to sequencing, the expression
levels of DKK1 were detected in samples. Results of the RT-qPCR
revealed that the levels of DKK1 expression was significantly
decreased in the Ang II stimulated group following transfection
with siDKK1 (Fig. 2B).
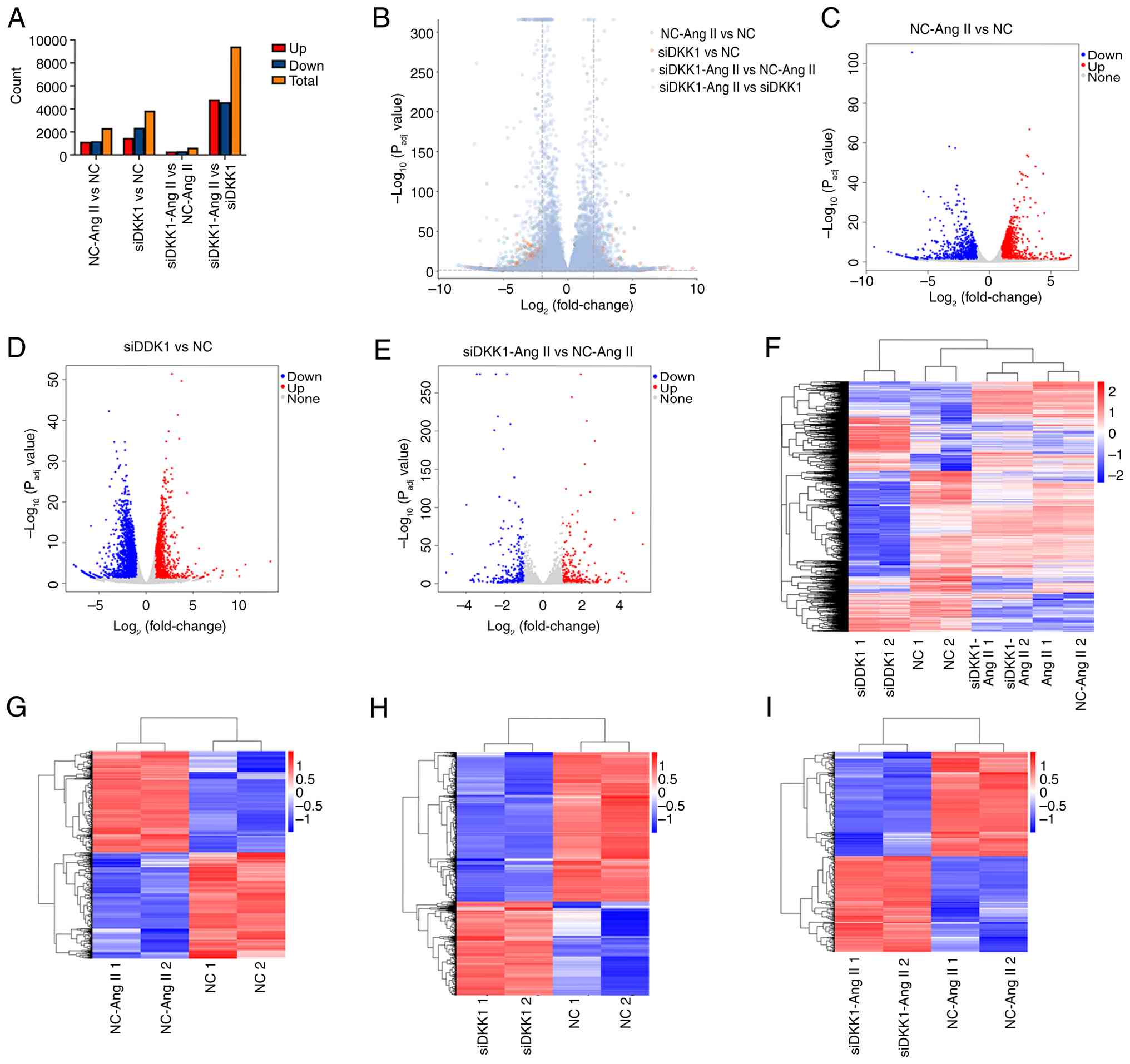
DEGs analysis following DKK1
silencing
DEGs were analyzed using RNA-seq, and statistical
analysis of DEGs are displayed in Fig.
3A; notably, ‘up’ is used to indicates genes that are
upregulated in expression in the treatment group, compared with the
control group, ‘down’ is used to indicate genes that are
downregulated in expression in the treatment group, compared with
the control group and total is used to indicate the total number of
genes with significant differences between the two groups.
According to the upregulated genes of each comparison group, a
volcano map of DEGs was produced (Fig.
3B). The volcano map of DEGs in NC-Ang II and NC groups is
displayed in Fig. 3C. There were
1,143 genes upregulated and 1,198 genes downregulated. The volcano
map of DEGs in siDKK1 and NC groups is displayed in Fig. 3D. There were 1,478 genes
upregulated and 2,366 genes downregulated. The volcano map of DEGs
in siDKK1-Ang II and NC-Ang II groups is displayed in Fig. 3E. There were 304 genes upregulated
and 332 genes downregulated. Genes that were significantly
upregulated, significantly downregulated or exhibited no
significant differential expression were marked with different
colors; blue indicated downregulation, whereas red indicated
upregulation. Cluster heat maps of all DEGs were also established,
and these are displayed in Fig.
3F. DEG clustering heat maps of NC-Ang II vs. NC, siDKK1 vs. NC
and siDKK1-Ang II vs. NC-Ang II are displayed in Fig. 3G and I, respectively. Heat maps of all
differentially expressed genes were drawn, including each
biological repetition. Rows represent genes, columns represent
groups for comparative analysis. Genes that were significantly
upregulated or significantly downregulated were marked with
different colors; blue indicated downregulation and red indicated
upregulation. The clustering tree diagrams on the left and top show
the similarities between genes or samples. The closer the branches,
the closer the expression patterns.
GO and KEGG functional enrichment
analysis
Results of the GO and KEGG functional enrichment
analyses revealed that the top 600 DEGs of NC-Ang II vs. NC were
associated with ‘Negative regulation of the immune system process’,
‘Collagen-containing extracellular matrix’, ‘Extracellular matrix
structural constituent’ and ‘Circadian entrainment’ (Fig. 4A). Moreover, the top 600 DEGs of
siDKK1 vs. NC were associated with ‘Neutrophil extracellular trap
formation’, ‘Negative regulation of viral genome replication’,
‘Protein-DNA complex’ and ‘Oxygen carrier activity’ (Fig. 4B). Results of the present study
also demonstrated that the top 600 DEGs of siDKK1-Ang II vs. NC-Ang
II were associated with the ‘Cytokine-cytokine receptor
interaction’, ‘Defense response to virus’, ‘Collagen-containing
extracellular matrix’ and ‘Unfolded protein binding’ (Fig. 4C).
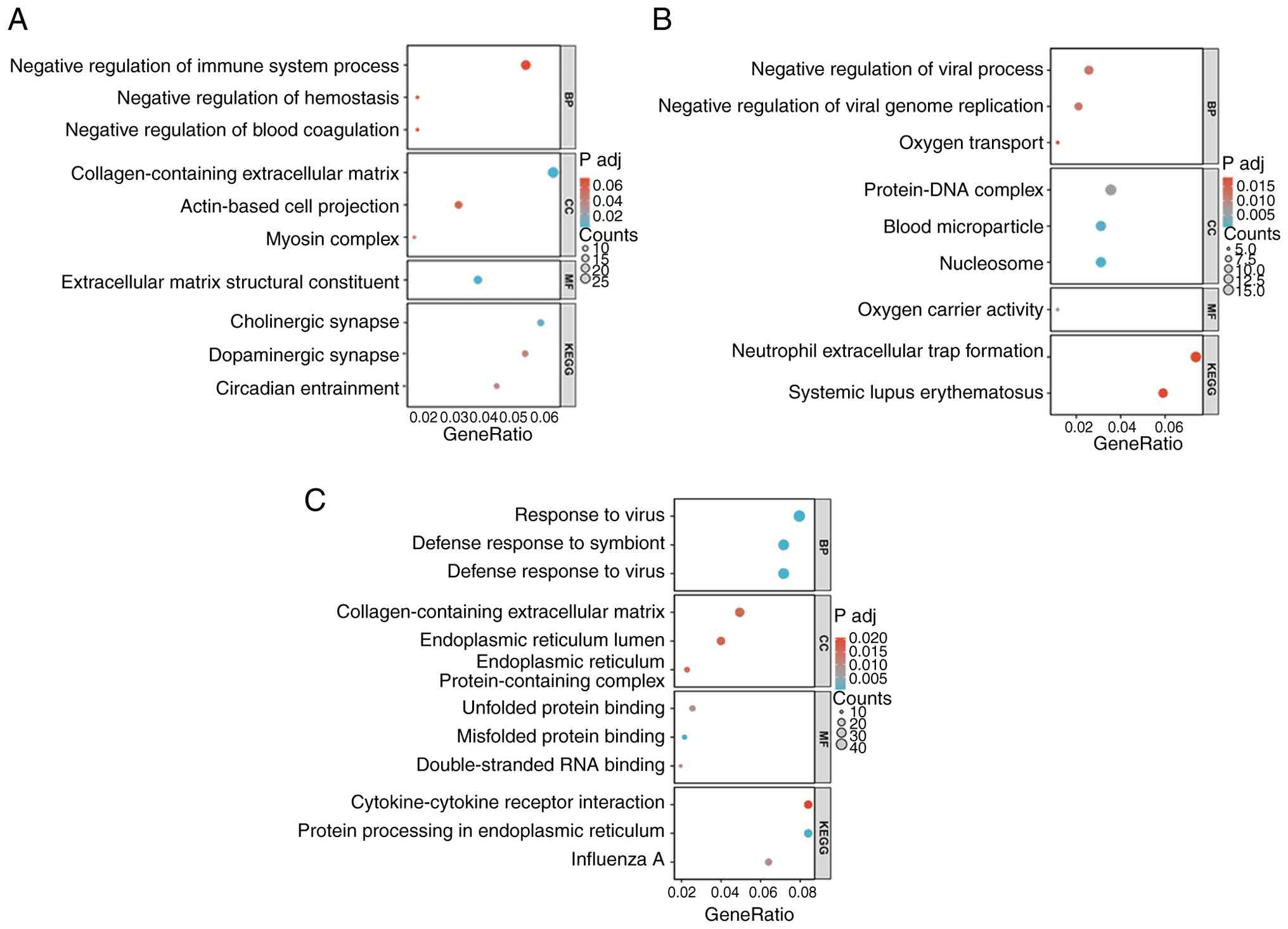 | Figure 4GO and KEGG functional enrichment
analysis. (A) Functional enrichment analysis of GO and KEGG of DEGs
in NC-Ang II vs. NC. (B) Functional enrichment analysis of GO and
KEGG of DEGs in siDKK1 vs. NC. (C) Functional enrichment analysis
of GO and KEGG of DEGs in siDKK1-Ang II vs. NC-Ang II. NC, treated
with scrambled siRNAs as negative control; siDKK1, treated with
siRNAs targeting human DKK1; Ang II, treated with Ang II (1 µM).
GO, gene ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes;
NC, negative control; siDKK1, small interfering RNA for DKK1; DKK1,
Dickkopf-1; Ang II, angiotensin II; BP, biological processes; CC,
cellular components; MF, molecular functions; P adj, adjusted
P. |
Key downstream genes following DKK1
silencing
A Venn diagram was used to identify 1,332 DEGs that
were co-expressed across the different groups. The Venn diagram
shows the intersection relationship of four groups of DEGs (NC-Ang
II vs. NC, siDKK1 vs. NC, siDKK1-Ang II vs. NC-Ang II and
siDKK1-Ang II vs. siDKK1). Each color represents one group. The
number of DEGs co-expressed in all four groups was 1332 (Fig. 5A). Functional enrichment analyses
using GO and KEGG revealed that the co-expressed DEGs were
associated with ‘Ribonucleoprotein complex biogenesis’,
‘Cell-substrate junction’, ‘Cadherin binding’ and ‘Protein
processing in the endoplasmic reticulum’ pathway (Fig. 5B). RRA analysis was performed using
1,332 co-expressed DEGs in all four groups in the aforementioned
Venn diagram, and the top 20 genes obtained are displayed in
Fig. 5C. The heat map analyzed
using RRA revealed the Log2 fold change value (Fig. 5D). Rows represent genes, columns
represent groups for comparative analysis. Genes that were
significantly upregulated or significantly downregulated were
marked with different colors; blue indicated downregulation and red
indicated upregulation. Functional enrichment analysis using GO and
KEGG demonstrated the potential association with ‘Negative
regulation of viral genome replication’, ‘Endoplasmic reticulum
lumen’ and ‘Coronavirus disease-COVID-19’ (Fig. 5E). Notably, SMDT1 was identified in
the RRA analysis, this is a mitochondrial calcium ion regulatory
protein and was therefore investigated further. Results of the
RT-qPCR assay revealed that the mRNA expression levels of SMDT1
were increased 2.78-fold following Ang II stimulation. In addition,
following DKK1 silencing, mRNA expression levels of SMDT1 were
downregulated by 0.37-fold (Fig.
5F).
Discussion
The present study aimed to elucidate the potential
association between DKK1 and AAA. Following stimulation of Ang II
in primary HSMCs, the results of the present study revealed that
the expression of DKK1 in HSMCs was elevated. Following the
successful knockdown of DKK1 using targeted siRNA, RNA-seq was used
to further investigate the downstream DEGs. Results of RNA-seq and
RT-qPCR analyses demonstrated that SMDT1 was significantly
downregulated following DKK1 silencing, indicating that DKK1 may
participate in the Ang II-induced mitochondrial injury of
HSMCs.
AAA is a complex pathological disease associated
with the artery walls, and is a key cause of mortality in the
elderly population, due to the rupture of dilated aorta tissue
(3). Effective pharmacotherapies
for halting the growth and rupture of AAA or delaying the
requirement for surgical repair are lacking at present. Risk
factors for this condition include smoking, obesity, aging, VSMC
phenotype transitions, metabolic disorders, nuclear receptors,
apoptosis and inflammation (1,8).
Notably, progressive inflammation is the focus of numerous studies
and is a well-established hallmark of AAA progression (28,29).
Yuan et al (30) revealed
that various inflammatory cell-associated mechanisms participate in
AAA development, such as T cells, macrophages, dendritic cells,
neutrophils, B cells and mast cells. Notably, AAA may begin with an
immune response to various antigens within the aortic wall, which
leads to the chemotaxis and infiltration of immune cells and,
ultimately, the release of different cytokines. This may cause
damage to the vascular endothelial cells and aggregation of
platelets, leading to aortic dilation; notably, this process is
comparable to the initiation of atherosclerosis (30). In addition, thrombosis may
participate in the pathogenesis of AAA; results of a previous study
revealed that a greater thrombus burden was associated with a
higher rupture risk even at lower AAA diameters, which was strongly
interconnected with inflammation and hemodynamics (31,32).
The study showed that antithrombotic therapies could prevent and
stabilize AAA; thus, the inhibition of platelets and coagulation
factors may exhibit potential in reducing the risk of AAA
development (31). However,
further investigations are required to determine key biomarkers
that may influence AAA pathogenesis, and to develop novel
pharmacological strategies for the clinical management of AAA.
Biomarkers associated with cardiovascular disease
may also exhibit potential as predictors for aortic disease, as
atherosclerosis plays a key role in the progression of AAA
(29). Notably, atherosclerosis
contributes to the progression of AAA in various ways, including
the release of pro-inflammatory cytokines, shear stress and
arterial remodeling. Thus, the regulation of atherosclerotic risk
factors is of importance in patients with AAA (31), and aneurysms of the abdominal aorta
represent a notable cardiovascular risk. A previous study revealed
that the renin angiotensin system plays a key role in atherogenesis
(33); notably, Ang II is an
important active peptide of the renin angiotensin system, and is
widely involved in the development of cardiovascular diseases, such
as atherosclerosis and hypertension (34). Ang II may indirectly influence the
atherogenic process via hemodynamic effects, as a result of
increased arterial blood pressure. In addition, Ang II has also
been shown to participate in the stimulation of monocyte
recruitment, recruitment and activation of macrophages and enhanced
oxidative stress, which are directly associated with the
progression of atherogenesis; these factors are consistent with the
chemoattractant property of Ang II (34).
Results of previous studies revealed the association
between Ang II and the development of AAA. Blocking Ang II using
either angiotensin-converting enzyme inhibitors or angiotensin
receptor blockers was associated with a reduction of mortality in
patients with AAA (34,35). Furthermore, within days of Ang II
infusion into mice, medial accumulation of macrophages and elastin
degradation in infrarenal aortic segments occur in the lesion of
AAA (36). The characteristics of
Ang II-induced AAA were comparable with those of atherogenesis,
such as activation of an inflammatory response, stimulation of
metabolic disorders, nuclear receptors and apoptosis (34). Ang II infusion of Apo
E-/- mice is a widely established
modeling method to induce AAA formation; in this model, an osmotic
pump is inserted into hyperlipidemic mice to continually inject Ang
II, this induces aneurysm phenotypes without surgical intervention
(34,37). A key principle of the Ang II
infusion model is systemic atherosclerosis, mobilization of
systemic inflammation through changing the hemodynamics of mice and
aggravation of artery wall damage to induce aneurysm formation. In
the present study, to the best of our knowledge, for the first time
HSMCs were stimulated with Ang II to mimic the pathological
environment of AAA.
VSMCs are the main component of the blood vessel
wall and exhibit numerous physiological functions, and the membrane
of the blood vessel wall is mainly composed of VSMCs and
extracellular matrix synthesized by VSMCs. In the pathological
course of AAA, the release of MMPs produce high levels of ROS to
damage VSMCs. VSMCs with oxidative damage exhibit abnormal
proliferation and apoptosis, promote the decline of elastin and
collagen synthesis and lead to the degeneration or even rupture of
the artery wall (38).
DKK1 is a conserved secreted glycoprotein that
mediates endocytosis and regulates the activity of the Wnt
signaling pathway through binding to LRP5/6, Kremen and other
receptors on the cell membrane (14). Results of a previous study revealed
that serum DKK1 plays an important role in the development of
vascular diseases, such as coronary atherosclerosis and acute
ischemic stroke (39). In 2009,
Ueland et al (40)
initially reported that patients with coronary artery disease or
advanced carotid plaques exhibited increased levels of DKK1, both
in serum and within the atherosclerotic lesion. DKK1 is involved in
platelet-induced endothelial cell activation, and both endothelial
and platelet-derived DKK1 could increase the inflammatory response
between them; these results indicate that DKK1 may increase the
inflammatory response in atherosclerotic plaques thus, playing a
role in promoting atherosclerosis (40). Di et al (41) revealed that DKK1 overexpression led
to the formation of enlarged and destabilized atherosclerotic
lesions, high levels of apoptosis in the aorta and the development
of carotid plaque. Moreover, DKK1-mediated apoptosis in human
umbilical vein endothelial cells was promoted through activation of
endoplasmic reticulum stress via the JNK pathway and canonical Wnt
signaling (41). Thus, DKK1 plays
a key role as a biomarker during the pathological development of
atherosclerosis, and is associated with inflammation facilitation,
platelet activation and endothelial dysfunction. When exposed to
atherogenic factors, such as oxidized low-density lipoprotein
receptor or disturbed flow, platelets or endothelial cells release
DKK1, resulting in vascular inflammation, endothelial-mesenchymal
transition and apoptosis (17). In
addition, DKK1 directly facilitates the expression of biomarkers
that participate in the pathogenesis of atherosclerosis, such as
plasminogen activator inhibitor type 1, clusterin/apolipoprotein J
and pentraxin 3(42). Results of
previous studies demonstrated the association between DKK1 and
mitochondrial disorders; DKK1 knockdown in human pulmonary artery
endothelial cells cultured under hypoxic conditions increased the
levels of ROS and the extent of mitochondrial DNA damage and
inhibited mitochondrial membrane hyperpolarization (18,43).
Moreover, DKK1 exacerbated doxorubicin-induced cardiomyocyte
apoptosis and mitochondrial dysfunction (44). Studies focused on tumors revealed
that DKK1 induced the apoptosis of JEG3 and BeWo trophoblastic
tumor cells through the mitochondrial apoptotic pathway (45), and DKK1 may inhibit
Wnt/Ca2+-CaMKII-NF-κB signaling (46). Although DKK1 exhibits potential as
a biomarker in cardiovascular disease, it is still unknown whether
DKK1 is involved in the regulation of AAA. Further investigations
into the potential role of DKK1 in AAA are required.
MCU complex consists of at least four main
components: The pore-forming subunit (MCU) capable of higher-order
oligomerization, the essential MCU regulator (EMRE) and two
membrane gate-keeping factors (MICU1 and MICU2) (47,48).
The molecular identification of MCU led to further studies focused
on mitochondrial calcium homeostasis (49,50).
Results of previous studies provided novel insights into the role
of MCU in physiological and pathological events (51), including AAA. Hypertension, induced
by chronic infusion of angiotensin II in the Micu2-depleted mice,
led to a progressive increase in the aortic diameter and eventually
to aneurysms (52). MCU
dysfunction alters the dynamics of mitochondrial Ca2+,
which in turn provokes pathological conditions affecting multiple
organs, such as the heart, skeletal muscle and brain (53). MCU also plays key roles in the
development of atherosclerosis, through modulating intracytoplasmic
Ca2+ concentration; MCU inhibition improved impaired
macrophage efferocytosis, and decreased ROS generation. Macrophage
efferocytosis eliminated apoptotic cells and prevented the release
of inflammatory factors and the formation of foam cells, thereby
alleviating atherosclerosis (54).
Moreover, Patel et al (55)
revealed that different shear stress patterns in vascular
endothelial cells modulated MCU complex subunit expression, which,
in turn, impacted the pathological process of atherosclerosis.
SMDT1, a subunit of the MCU complex, essential for
bridging the calcium-sensing role of MICU1 and MICU2, is involved
in regulating calcium influx into the mitochondria of almost all
mammalian tissues, which is of importance for regulating energy
production and cell signaling due to ATP synthesis and apoptosis
(56,57). SMDT1 variants impair EMRE-mediated
mitochondrial calcium uptake in patients with muscle involvement
(58). Results of a previous study
revealed that SMDT1 overexpression promoted mitochondrial fission
in PDAC cells. SMDT1-driven change in mitochondrial dynamics
morphology mediates apoptosis in pancreatic ductal adenocarcinoma
(59).
A recent Mendelian randomization analysis revealed a
causal association between elevated levels of SMDT1 and increased
risks of cardiovascular disease, including coronary
atherosclerosis, myocardial infarction and cardiomyopathy (60). In the present study, Ang II
stimulated HSMCs to simulate AAA pathological damage, and following
stimulation of Ang II, the concentration of DKK1 was increased and
the expression levels of SMDT1 were increased concurrently.
Notably, both DKK1 and SMDT1 levels were reduced following DKK1
silencing; thus, we hypothesize that DKK1/SMDT1 may participate in
Ang II-induced mitochondrial injury of human smooth muscle cells,
which may act as a novel pathological process in AAA.
Notably, the present study exhibits limitations. For
example, only in vitro experiments were performed in the
absence of in vivo validation, and only one cell type,
namely HSMCs, were used, in the absence of endothelial/macrophage
co-culturing. Therefore, the physiological relevance of these
findings remains to be fully elucidated. The present study focused
on understanding potential changes in the expression of downstream
mitochondrial function-associated protein, SMDT1, following DKK1
silencing under Ang II stimulation. Further functional experiments
are required.
To the best of our knowledge, the present study is
the first to determine the role of DKK1/SMDT1 in AAA. In
conclusion, the present study provided novel insights into the role
of DKK1/SMDT1 in inflammatory responses in vascular diseases, which
may aid in further understanding the pathophysiological mechanisms
that are associated with AAA.
Acknowledgements
Not applicable.
Funding
Funding: The present study was funded by The Projects of
Technological Innovation Development Program in Yantai City (grant
no. 2022YD071), Traditional Chinese medicine science and technology
project of Shandong Province (grant no. M-2023017), Binzhou Medical
University research start-up fund (grant no. BY2020KJ46) and
Introduction of Talent Research Initiation Fund of Central Hospital
Affiliated to Shandong First Medical University (grant no.
YJRC2022017).
Availability of data and materials
The data generated in the present study may be found
in the National Center for Biotechnology Information under the
login number of the data set: PRJNA1234348 or at the following URL:
http://www.ncbi.nlm.nih.gov/bioproject/1234348.
Authors' contributions
XC, BY and ZL were responsible for performing the
bioinformatics analysis using software. data collection,
statistical analysis and manuscript writing. ZG, PQ and XL helped
with data collection, statistical analysis and literature search.
LW and AL supervised the project and were involved in the study
design, data analysis and writing of the manuscript. LW and AL
confirm the authenticity of all the raw data. All authors have read
and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Ullery BW, Hallett RL and Fleischmann D:
Epidemiology and contemporary management of abdominal aortic
aneurysms. Abdom Radiol (NY). 43:1032–1043. 2018.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Kent KC: Clinical practice. Abdominal
aortic aneurysms. N Engl J Med. 371:2101–2108. 2014.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Zhao G, Fu Y, Cai Z, Yu F, Gong Z, Dai R,
Hu Y, Zeng L, Xu Q and Kong W: Unspliced XBP1 confers VSMC
homeostasis and prevents aortic aneurysm formation via FoxO4
interaction. Circ Res. 121:1331–1345. 2017.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Li Y, Ren P, Dawson A, Vasquez HG, Ageedi
W, Zhang C, Luo W, Chen R, Li Y, Kim S, et al: Single-cell
transcriptome analysis reveals dynamic cell populations and
differential gene expression patterns in control and aneurysmal
human aortic tissue. Circulation. 142:1374–1388. 2020.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Wang H, Kazaleh M, Gioscia-Ryan R, Millar
J, Temprano-Sagrera G, Wood S, Van Den Bergh F, Blin MG, Wragg KM,
Luna A, et al: Deficiency of mitophagy mediator Parkin in aortic
smooth muscle cells exacerbates abdominal aortic aneurysm. bioRxiv
[Preprint]: 2024.10.30.621201, 2025.
|
|
6
|
Sun LY, Lyu YY, Zhang HY, Shen Z, Lin GQ,
Geng N, Wang YL, Huang L, Feng ZH, Guo X, et al: Nuclear receptor
NR1D1 regulates abdominal aortic aneurysm development by targeting
the mitochondrial tricarboxylic acid cycle enzyme aconitase-2.
Circulation. 146:1591–1609. 2022.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Tavris BS, Peters AS, Böckler D and
Dihlmann S: Mitochondrial dysfunction and increased DNA damage in
vascular smooth muscle cells of abdominal aortic aneurysm
(AAA-SMC). Oxid Med Cell Longev. 2023(6237960)2023.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Liu Y, Yu M, Wang H, Dorsey KH, Cheng Y,
Zhao Y, Luo Y, Zhao G, Zhao Y, Lu H, et al: restoring vascular
smooth muscle cell mitochondrial function attenuates abdominal
aortic aneurysm in mice. Arterioscler Thromb Vasc Biol. 45:523–540.
2025.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Azhdari M and Zur Hausen A: Wnt/β-catenin
and notch signaling pathways in cardiovascular disease: Mechanisms
and therapeutics approaches. Pharmacol Res.
211(107565)2025.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Methatham T, Tomida S, Kimura N, Imai Y
and Aizawa K: Inhibition of the canonical Wnt signaling pathway by
a β-catenin/CBP inhibitor prevents heart failure by ameliorating
cardiac hypertrophy and fibrosis. Sci Rep. 11(14886)2021.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Rong Z, Li F, Zhang R, Niu S, Di X, Ni L
and Liu C: Ant-neointimal formation effects of SLC6A6 in preventing
vascular smooth muscle cell proliferation and migration via
Wnt/β-catenin signaling. Int J Mol Sci. 24(2018)2023.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Schunk SJ, Floege J, Fliser D and Speer T:
WNT-β-catenin signalling-a versatile player in kidney injury and
repair. Nat Rev Nephrol. 17:172–184. 2021.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Xu L, Liu B, Ma H, Qi E, Ma J, Chang T,
Zhang J, Zhang W, Chen W, Cao X and Xiong X: O-GlcNAc transferase
promotes vascular smooth muscle calcification through modulating
Wnt/β-catenin signaling. Faseb J. 38(e70271)2024.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Boucher P, Matz RL and Terrand J:
atherosclerosis: Gone with the Wnt? Atherosclerosis. 301:15–22.
2020.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Klingenschmid G, Tschiderer L, Himmler G,
Rungger G, Brugger S, Santer P, Willeit J, Kiechl S and Willeit P:
Associations of serum dickkopf-1 and sclerostin with cardiovascular
events: results from the prospective bruneck study. J Am Heart
Assoc. 9(e014816)2020.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Xu H, Ding Z, Chen J, Zhang Y, Shan W,
Chen X, Liu X, Gao Y and Han G: Correlation between serum
Dickkopf-1 (DKK1) levels and coronary artery stenosis. Nutr Metab
Cardiovasc Dis. 33:168–176. 2023.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Baetta R and Banfi C: Dkk (dickkopf)
proteins. Arterioscler Thromb Vasc Biol. 39:1330–1342.
2019.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Wang Q, Tian J, Li X, Liu X, Zheng T, Zhao
Y, Li X, Zhong H, Liu D, Zhang W, et al: Upregulation of
endothelial DKK1 (dickkopf 1) promotes the development of pulmonary
hypertension through the sp1 (specificity protein 1)/SHMT2 (serine
hydroxymethyltransferase 2) pathway. Hypertension. 79:960–973.
2022.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Huo C, Liu Y, Li X, Xu R, Jia X, Hou L and
Wang X: LRRC8A contributes to angiotensin II-induced cardiac
hypertrophy by interacting with NADPH oxidases via the C-terminal
leucine-rich repeat domain. Free Radic Biol Med. 165:191–202.
2021.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Martin M: Cutadapt removes adapter
sequences from high-throughput sequencing reads. EMBnet J.
17(3)2011.
|
|
22
|
Stark R, Grzelak M and Hadfield J: RNA
sequencing: The teenage years. Nat Rev Genet. 20:631–656.
2019.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Wickham H: ggplot2: Elegant graphics for
data analysis. New York, NY, Springer, 2009.
|
|
24
|
Chen H and Boutros PC: VennDiagram: A
package for the generation of highly-customizable Venn and Euler
diagrams in R. BMC Bioinformatics. 12(35)2011.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Kolde R, Laur S, Adler P and Vilo J:
Robust rank aggregation for gene list integration and
meta-analysis. Bioinformatics. 28:573–580. 2012.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Yu G, Wang LG, Han Y and He QY:
clusterProfiler: An R package for comparing biological themes among
gene clusters. OMICS. 16:284–287. 2012.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Szklarczyk D, Gable AL, Nastou KC, Lyon D,
Kirsch R, Pyysalo S, Doncheva NT, Legeay M, Fang T, Bork P, et al:
The STRING database in 2021: Customizable protein-protein networks,
and functional characterization of user-uploaded gene/measurement
sets. Nucleic Acids Res. 49 (D1):D605–D612. 2021.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Trollope AF and Golledge J: Angiopoietins,
abdominal aortic aneurysm and atherosclerosis. Atherosclerosis.
214:237–243. 2011.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Khan H, Abu-Raisi M, Feasson M, Shaikh F,
Saposnik G, Mamdani M and Qadura M: Current prognostic biomarkers
for abdominal aortic aneurysm: A comprehensive scoping review of
the literature. Biomolecules. 14(661)2024.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Yuan Z, Lu Y, Wei J, Wu J, Yang J and Cai
Z: Abdominal aortic aneurysm: Roles of inflammatory cells. Front
Immunol. 11(609161)2021.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Bontekoe J, Matsumura J and Liu B:
Thrombosis in the pathogenesis of abdominal aortic aneurysm. JVS
Vasc Sci. 4(100106)2023.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Liu X, Weng Y, Lou J, Chen X, Du C and
Tang L: Combinational therapy with aspirin and ticagrelor
alleviates vascular inflammation and angiotensin II-driven
abdominal aortic aneurysm formation in mice. Austin J Cardiovasc
Dis Atherosclerosis. 9(1048)2022.
|
|
33
|
Lu H, Rateri DL, Bruemmer D, Cassis LA and
Daugherty A: Involvement of the renin-angiotensin system in
abdominal and thoracic aortic aneurysms. Clin Sci (Lond).
123:531–543. 2012.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Daugherty A, Manning MW and Cassis LA:
Angiotensin II promotes atherosclerotic lesions and aneurysms in
apolipoprotein E-deficient mice. J Clin Invest. 105:1605–1612.
2000.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Kristensen KE, Torp-Pedersen C, Gislason
GH, Egfjord M, Rasmussen HB and Hansen PR: Angiotensin-converting
enzyme inhibitors and angiotensin II receptor blockers in patients
with abdominal aortic aneurysms: Nation-wide cohort study.
Arterioscler Thromb Vasc Biol. 35:733–740. 2015.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Daugherty A and Cassis LA: Mouse models of
abdominal aortic aneurysms. Arterioscler Thromb Vasc Biol.
24:429–434. 2004.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Sawada H, Lu HS, Cassis LA and Daugherty
A: Twenty years of studying AngII (angiotensin II)-induced
abdominal aortic pathologies in mice: Continuing questions and
challenges to provide insight into the human disease. Arterioscler
Thromb Vasc Biol. 42:277–288. 2022.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Quintana RA and Taylor WR: Cellular
mechanisms of aortic aneurysm formation. Circ Res. 124:607–618.
2019.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Zhu Z, Guo D, Zhong C, Wang A, Xie X, Xu
T, Chen CS, Peng Y, Peng H, Li Q, et al: Serum Dkk-1 (dickkopf-1)
Is a potential biomarker in the prediction of clinical outcomes
among patients with acute ischemic stroke. Arterioscler Thromb Vasc
Biol. 39:285–293. 2019.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Ueland T, Otterdal K, Lekva T, Halvorsen
B, Gabrielsen A, Sandberg WJ, Paulsson-Berne G, Pedersen TM,
Folkersen L, Gullestad L, et al: Dickkopf-1 enhances inflammatory
interaction between platelets and endothelial cells and shows
increased expression in atherosclerosis. Arterioscler Thromb Vasc
Biol. 29:1228–1234. 2009.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Di M, Wang L, Li M, Zhang Y, Liu X, Zeng
R, Wang H, Chen Y, Chen W, Zhang Y and Zhang M: Dickkopf1
destabilizes atherosclerotic plaques and promotes plaque formation
by inducing apoptosis of endothelial cells through activation of ER
stress. Cell Death Dis. 8(e2917)2017.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Magrini E, Mantovani A and Garlanda C: The
dual complexity of PTX3 in health and disease: A balancing Act?
Trends Mol Med. 22:497–510. 2016.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Kim IG, Kim SY, Kim HA, Kim JY, Lee JH,
Choi SI, Han JR, Kim KC and Cho EW: Disturbance of DKK1 level is
partly involved in survival of lung cancer cells via regulation of
ROMO1 and γ-radiation sensitivity. Biochem Biophys Res Commun.
443:49–55. 2014.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Liang L, Tu Y, Lu J, Wang P, Guo Z, Wang
Q, Guo K, Lan R, Li H and Liu P: Dkk1 exacerbates
doxorubicin-induced cardiotoxicity by inhibiting the Wnt/β-catenin
signaling pathway. J Cell Sci. 132(jcs228478)2019.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Cui H, Li H, Li QL, Chen J, Na Q and Liu
CX: Dickkopf-1 induces apoptosis in the JEG3 and BeWo trophoblast
tumor cell lines through the mitochondrial apoptosis pathway. Int J
Oncol. 46:2555–2561. 2015.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Zhuang X, Zhang H, Li X, Li X, Cong M,
Peng F, Yu J, Zhang X, Yang Q and Hu G: Differential effects on
lung and bone metastasis of breast cancer by Wnt signalling
inhibitor DKK1. Nat Cell Biol. 19:1274–1285. 2017.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Oxenoid K, Dong Y, Cao C, Cui T, Sancak Y,
Markhard AL, Grabarek Z, Kong L, Liu Z, Ouyang B, et al:
Architecture of the mitochondrial calcium uniporter. Nature.
533:269–273. 2016.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Wang Y, Nguyen NX, She J, Zeng W, Yang Y,
Bai XC and Jiang Y: Structural mechanism of EMRE-dependent gating
of the human mitochondrial calcium uniporter. Cell.
177:1252–1261.e13. 2019.PubMed/NCBI View Article : Google Scholar
|
|
49
|
Patron M, Granatiero V, Espino J, Rizzuto
R and De Stefani D: MICU3 is a tissue-specific enhancer of
mitochondrial calcium uptake. Cell Death Differ. 26:179–195.
2019.PubMed/NCBI View Article : Google Scholar
|
|
50
|
Shamseldin HE, Alasmari A, Salih MA,
Samman MM, Mian SA, Alshidi T, Ibrahim N, Hashem M, Faqeih E,
Al-Mohanna F and Alkuraya FS: A null mutation in MICU2 causes
abnormal mitochondrial calcium homeostasis and a severe
neurodevelopmental disorder. Brain. 140:2806–2813. 2017.PubMed/NCBI View Article : Google Scholar
|
|
51
|
Murgia M and Rizzuto R: Molecular
diversity and pleiotropic role of the mitochondrial calcium
uniporter. Cell Calcium. 58:11–17. 2015.PubMed/NCBI View Article : Google Scholar
|
|
52
|
Tarasova NV, Vishnyakova PA, Logashina YA
and Elchaninov AV: Mitochondrial calcium uniporter structure and
function in different types of muscle tissues in health and
disease. Int J Mol Sci. 20(4823)2019.PubMed/NCBI View Article : Google Scholar
|
|
53
|
Alevriadou BR, Patel A, Noble M, Ghosh S,
Gohil VM, Stathopulos PB and Madesh M: Molecular nature and
physiological role of the mitochondrial calcium uniporter channel.
Am J Physiol Cell Physiol. 320:C465–C482. 2021.PubMed/NCBI View Article : Google Scholar
|
|
54
|
Lu N, Zhu JF, Lv HF, Zhang HP, Wang PL,
Yang JJ and Wang XW: Modulation of oxidized low-density
lipoprotein-affected macrophage efferocytosis by mitochondrial
calcium uniporter in a murine model. Immunol Lett. 263:14–24.
2023.PubMed/NCBI View Article : Google Scholar
|
|
55
|
Patel A, Pietromicca JG, Venkatesan M, et
al: Modulation of the mitochondrial Ca(2+) uniporter complex
subunit expression by different shear stress patterns in vascular
endothelial cells. Physiol Rep. 11(e15588)2023.PubMed/NCBI View Article : Google Scholar
|
|
56
|
Sancak Y, Markhard AL, Kitami T,
Kovács-Bogdán E, Kamer KJ, Udeshi ND, Carr SA, Chaudhuri D, Clapham
DE, Li AA, et al: EMRE is an essential component of the
mitochondrial calcium uniporter complex. Science. 342:1379–1382.
2013.PubMed/NCBI View Article : Google Scholar
|
|
57
|
Arduino DM, Goh V, Mokranjac D and
Perocchi F: Drug discovery assay to identify modulators of the
mitochondrial Ca2+ uniporter. Methods Mol Biol.
2277:69–89. 2021.PubMed/NCBI View Article : Google Scholar
|
|
58
|
Bulthuis EP, Adjobo-Hermans MJW, de Potter
B, Hoogstraten S, Wezendonk LHT, Tutakhel OAZ, Wintjes LT, van den
Heuvel B, Willems PHGM, Kamsteeg EJ, et al: SMDT1 variants impair
EMRE-mediated mitochondrial calcium uptake in patients with muscle
involvement. Biochim Biophys Acta Mol Basis Dis.
1869(166808)2023.PubMed/NCBI View Article : Google Scholar
|
|
59
|
Xie KF, Guo DD and Luo XJ: SMDT1-driven
change in mitochondrial dynamics mediate cell apoptosis in PDAC.
Biochem Biophys Res Commun. 511:323–329. 2019.PubMed/NCBI View Article : Google Scholar
|
|
60
|
Meng H, He Y, Rui Y, Cai M, Fu D, Bi W,
Luo B and Gao Y: Genetic predisposition and mitochondrial
dysfunction in sudden cardiac death: Role of MCU complex genetic
variations. Cells. 14(728)2025.PubMed/NCBI View Article : Google Scholar
|