Introduction
The novel microRNA miR-1246, first identified in
2008(1), has been investigated as
a biomarker for pancreatic adenocarcinoma, hepatocellular
carcinoma, and esophageal, breast, and lung cancers (2-6).
A fragment of U2 small nuclear RNA (U2 snRNA) that carries an
identical sequence to miR-1246, referred to as RNU2-1f (7) or miR-U2-1(8), has also been evaluated as a potential
biomarker for pancreatic and colorectal adenocarcinoma, melanoma,
central nervous system lymphoma, and ovarian and lung cancers
(7-11).
The area under the receiver-operator characteristics curve (AUC) of
miR-1246 was 0.878 (CI: 0.818-0.925) for discriminating healthy
controls (HC) (n=96) from patients with all stages of non-small
cell lung cancer (NSCLC) (n=62) (8). The AUC of miR-1246 was 0.891 (CI:
0.819-0.962) for distinguishing lung cancer patients of all
histological types (n=211) from HC (n=58) and 0.873 (CI:
0.761-0.985) for distinguishing stage 0-II patients (n=54) from
controls (12).
Various isoforms of miR-1246 have been detected in
sera and tumor samples from patients with cancer, some of which
were longer than its archetypical sequence as described in the
micro-RNA database, miRBase (7-8,13).
Sequences at both ends of these isoforms correspond to the U2 snRNA
sequence rather than the predicted pre-miR-1246 sequence (7-8,13).
These results revealed that miR-1246 is not derived from
pre-miR-1246, but from U2 snRNA. Genomic deletion of the predicted
region of the miR-1246 gene (chromosome 2: 176,600,980-176,601,052)
was shown to reduce neither miR-1246 expression nor exosomal
miR-1246 levels in Panc-1 cells (14). These findings strongly suggest that
miR-1246 is not encoded by the miR-1246 gene, but rather by the U2
snRNA gene RNU2-1 (14).
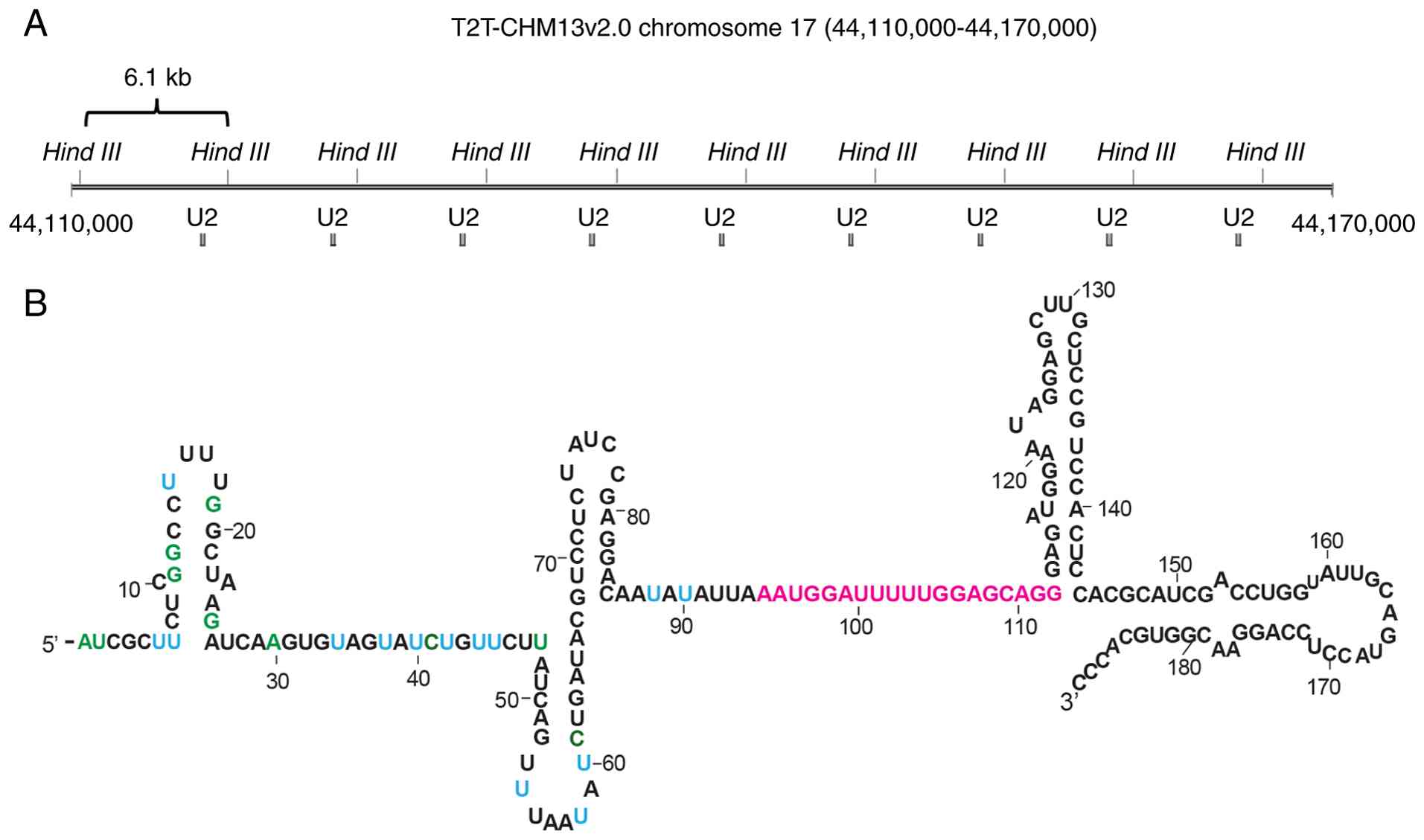
RNU2-1 is located on human chromosome
17q21.31, and is organized as a tandem array (15,16).
A 6.1 kb repeat unit includes a188 bp U2 snRNA coding region, and
the repeat number of the unit exhibits certain polymorphisms [i.e.,
copy number variations (CNVs)] (17) (Fig.
1A). The identified copy number range was 6-82 per haploid
genome, as determined through direct visualization of the
polymorphic RNU2-1 gene copy number using fluorescence in
situ hybridization (FISH) on 46 unrelated chromosomes (18). In addition, 53 alleles were
identified via Southern blot analysis (19).
The present study aimed to clarify whether the
polymorphic copy number of RNU2-1 affects serum miR-1246
levels in both HC and patients with lung adenocarcinoma.
RNU2-1 copy numbers have traditionally been measured using
Southern blot analysis (19) or
FISH (18). Recently, the depth of
coverage values of sequences from over 1,000 individuals from
various genome projects were used to determine the RNU2-1
copy number per diploid genome (20). In this study, we used digital PCR
(dPCR) as a relatively simple method to determine the extremely
wide-ranging RNU2-1 copy number per diploid genome, and
analyzed the correlation between RNU2-1 copy number and
serum miR-1246 levels.
Materials and methods
Patients and clinical specimens
This study was approved by the Ethics Committee of
the Faculty of Health Sciences, Kyorin University for the HC
(approval number: 2022-30) and the patients with stage IV lung
adenocarcinoma whose samples were obtained from Kanagawa Cancer
Center (approval number: 2023-22). The study protocol for the
patients was also approved by the Ethics Committee of Kanagawa
Cancer Center (approval number: 2023 epidemiology-84). Samples from
the HC were collected at Kyorin University in October 2014 and
October 2020. The HC consented to the use of their samples in the
present study, and they provided written informed consent. Samples
from patients were collected from the biobank of the Kanagawa
Cancer Center between April 2021 and March 2023 and provided to us
in November 2023. The patients consented to the use of their
samples in comprehensive cancer research, including genetic
analysis. The exclusion criteria for patients with lung
adenocarcinoma were as follows: participants who were pregnant, had
a history of other malignancies, or had received prior
treatment.
Extraction of serum RNAs
All serum samples were centrifuged at 20,000 x g for
10 min at 4˚C to remove cell debris, divided into 200 µl aliquots,
and stored at -80˚C until further use. An miRNeasy Serum/Plasma kit
(Qiagen GmbH, cat. no. 217184) was used for small RNA extraction.
Briefly, 3.5 µl of 0.16 fmol/µl 5'-phosphorylated cel-miR-39-3p was
added to 200 µl of serum sample as a spike-in control for RT-qPCR.
RNA was extracted according to the manufacturer's instructions,
with the only minor modification being that the volume of
RNase-free H2O used to elute the RNA was changed to 28
µl (21).
RT-qPCR
The experimental protocol was essentially identical
to the one described in our previous study (21). MiR-X miRNA First-Strand Synthesis
and TB Green qRT-PCR systems (Takara Bio Inc., cat. no. 638313)
were used to quantify miR-1246. The cDNA was synthesized according
to the manufacturer's instructions. Briefly, 5 µl of 2x mRQ buffer,
3.75 µl of RNA sample, and 1.25 µl of mRQ Enzyme Mix were added to
0.2 ml tubes, incubated at 37˚C for 1 h, then inactivated at 85˚C
for 5 min. Thereafter, 90 µl of DNase-RNase-free H2O was
added to the solution. The qPCR reaction mixture consisted of 7.8
µl of DNase/RNase-free H2O, 10 Μl of 2x TB Green
Advantage Premix (Takara Bio Inc., cat. no. 638314), 0.4 Μl of 50x
ROX Reference Dye LMP, 0.4 µl of the primers (10 µM), and 1 µl of
cDNA. A two-step qPCR was performed in duplicate on a 7500 Fast
Real-Time PCR System (Thermo Fisher Scientific Inc.), using the
cycling protocol: 95˚C for 10 sec, 40 cycles of 95˚C for 4 sec, and
60˚C for 32 sec. The forward primer sequences are presented in
Table I. A reverse transcription
primer and a reverse primer were provided as part of the MiR-X
miRNA First-Strand Synthesis and TB Green qRT-PCR systems. The
2-ΔCq method was used to perform relative quantification
of the miRNAs as follows: ΔCq=(Cq of miR-1246)-(Cq of spike-in
control cel-miR-39-3p).
 | Table IOligonucleotides used in the present
study. |
Table I
Oligonucleotides used in the present
study.
| A, Primers for
digital PCR |
|---|
| Name | Sequence
(5'-3') | (Refs.) |
|---|
| RNU2-1F97 |
GGATTTTTGGAGCAGGGAGAT | - |
| RNU2-1R172 |
GAGGTACTGCAATACCAGGTCGAT | - |
| CEP17-F |
GCTGATGATCATAAAGCCACAGGTA | (22) |
| CEP17-R19 |
AGCTGGTGCTCAGGCAGTG | (22) |
| B, Primers for
real-time PCR |
| Name | Sequence
(5'-3') | (Refs.) |
| cel-39 miR-XF |
CACCGGGTGTAAATCAGCTTG | (21) |
| U2 miR-XF |
CCAATGGATTTTTGGAGCAGG | (21) |
| C, TaqMan probes
for digital PCR |
| Name | Sequence
(5'-3') | (Refs.) |
| RNU2-1TM132 |
FAM-CTCCGTCCACTCCAC | - |
| CEP17-p
FAM |
FAM-TGCTGCAATAGGCGG | (22) |
Quantification of gene copy number via
dPCR
Genomic DNA was extracted from peripheral venous
blood using a QIAamp DNA Mini kit (Qiagen GmbH, cat. no. 51304),
according to the manufacturer's instructions. The restriction
enzyme HindIII was used to divide RNU2-1 tandem repeats into
repeat units (17) (Fig. 1A). A total of 35-75 ng of DNA was
digested using 10.5 U of HindIII (Takara Bio Inc., cat. no. 1060A)
in a 20 µl reaction mixture at 37˚C for 60 min, after which HindIII
was inactivated at 65˚C for 20 min. The HindIII-digested DNA was
then centrifuged at 20,630 x g for 3 min before being added to the
dPCR reaction mixture. A 10 µl aliquot of reaction mixture,
containing 1 µl of HindIII-digested DNA, 0.8 µl of 2.5 µM TaqMan
probe, 0.45 µl each of 10 µM primers, 0.5 µl of TaqMan™ Copy Number
Reference Assay RNase P (Thermo Fisher Scientific Inc., cat. no.
4403328), 2 µl of 5x Absolute Q DNA Digital PCR Master Mix (Thermo
Fisher Scientific Inc., cat. no. A52490), and 4.8 µl of ultra-pure
H2O was prepared. After centrifuging the reaction
mixture at 20,630 x g for 1 min, 9 µl of the mixture and 15 µl of
Absolute Q Isolation Buffer were applied to each well of the
Absolute Q MAP16 Plate (Thermo Fisher Scientific Inc., cat. No.
A52865), according to the manufacturer's instructions. The plate
was centrifuged at 160 x g for 1 min, after which PCR was performed
in duplicate according to the cycling protocol: 96˚C for 10 min, 40
cycles of 96˚C for 5 sec, and 61˚C for 30 sec, on a QuantStudio
Absolute Q Digital PCR system (Thermo Fisher Scientific Inc.). The
sequences of all TaqMan probes and primers are listed in Table I. The primers and TaqMan probe for
RNU2-1 were designed using Primer Express software 3.0.1
(Thermo Fisher Scientific Inc.) based on the human RNU2-1
sequence (NCBI Gene ID: 6066). Similarly, the copy number of
CEP-17, a marker for the centromere of chromosome 17, was
quantified using the RPPH1 gene as a reference to evaluate
any chromosome 17 polysomy. The CEP17-R19 primer was modified from
CEP17-R (22) by adding three
nucleotides (AGC) at the 5' end.
Statistical analysis
All statistical analyses were conducted using JMP
13.2.1 software (SAS Institute Inc.). The ages of the study
participants were expressed as means ± standard deviations (SDs).
The Shapiro-Wilk test indicated that each experimental dataset was
non-normally distributed (P<0.005). Therefore, the Mann-Whitney
U test was used to compare miR-1246 serum levels or RNU2-1
copy numbers between the patient and control groups. Fisher's exact
test was used to assess the significance of any differences in age,
and the chi-squared test was used to assess differences in sex.
Spearman's rank correlation coefficient test was used to perform a
correlation analysis. Stepwise linear regression analysis was used
to identify factors influencing serum miR-1246 levels. Differences
were considered statistically significant when their P-values were
<0.05.
Results
RNU2-1 copy numbers of healthy
individuals and patients with lung adenocarcinoma
We first quantified serum miR-1246 levels in 51
healthy individuals and 45 patients with stage IV lung
adenocarcinoma using RT-qPCR. The characteristics of the
participants are described in Table
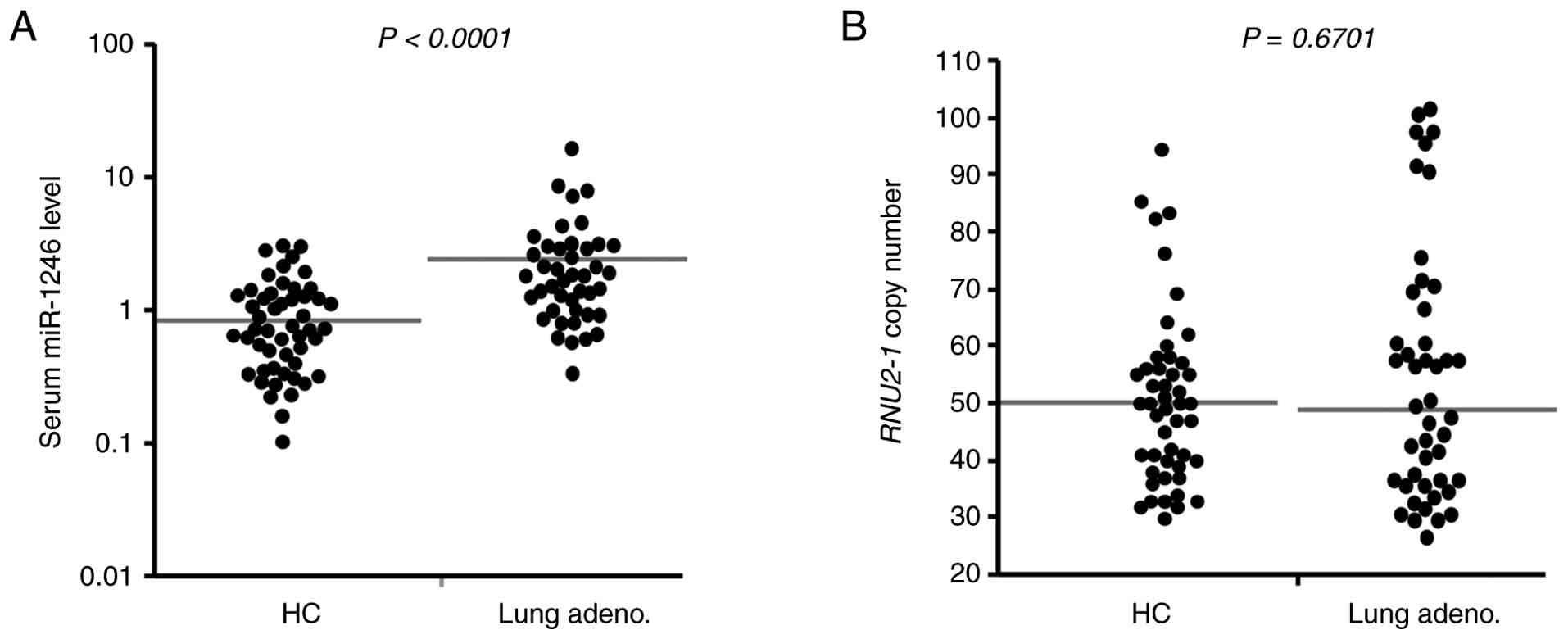
II. Serum miR-1246 levels were significantly higher in patients
with stage IV lung adenocarcinoma than in the HC (P<0.0001;
Fig. 2A). These results obtained
from patients with stage IV adenocarcinoma were consistent with the
results in our previous report on patients with stage III or IV
adenocarcinoma (21). There was a
significant difference in the age distribution between the patients
and HC (P<0.0001; Table II).
The age (mean ± standard deviation) was 68.56±7.31 years in the
patient group and 36.19±14.74 years in the HC group. Therefore, we
examined whether age affected serum miR-1246 levels. We first
performed a stepwise multiple regression analysis, with age as a
covariate. Age was the only retained variable (P<0.0001) in the
current cohort, because the ages of the HC were significantly lower
than those of patients with lung adenocarcinoma. To investigate the
potential confounding effect of age, we performed the same analysis
using data from our previous study (21). Results showed that disease status
was the strongest factor associated with serum miR-1246 levels
(P<0.05). In the current cohort, a correlation between age and
serum miR-1246 level was not observed in patients with lung
adenocarcinoma (Fig. S1A).
Although a moderate age-related correlation was observed (R=0.569)
in the HCs (Fig. S2B), this
correlation was largely driven by a subset of HC. This subset was a
young healthy cohort (20-22 years old;) whose samples were
collected on the same day. The correlation was no longer
significant after excluding this young healthy subset (Fig. S1C).
 | Table IICharacteristics of patients with lung
adenocarcinoma and control subjects. |
Table II
Characteristics of patients with lung
adenocarcinoma and control subjects.
|
Characteristics | No. of patients
with lung adenocarcinoma | No. of control
subjects | P-value |
|---|
| Total | 45 | 51 | |
| Age | | | <0.0001 |
|
≥50 | 45 | 17 | |
|
<50 | 0 | 34 | |
| Sex | | | <0.001 |
|
Male | 27 | 12 | |
|
Female | 18 | 39 | |
| Stage | | | |
|
IV | 45 | | |
Next, the RNU2-1 copy numbers per diploid
were measured via dPCR using genomic DNA derived from peripheral
blood samples (Fig. 2B). Each
RNU2-1 gene in the tandem array was digested using HindIII,
as described in the Materials and Methods section (Fig. 1A). The range of the RNU2-1
copy numbers was 30-94 (median: 50) in the HC group, and 26-101
(median: 49) in the stage IV lung adenocarcinoma group, with no
significant differences observed between the two groups (P=0.4349;
Fig. 2B). We also investigated the
possibility of chromosome 17 polysomy in seven patients with stage
IV lung adenocarcinoma who had particularly high RNU2-1 copy
numbers (Table III), using a
TaqMan probe for CEP17 (Table I).
The CEP17 copy numbers per diploid genome ranged between 1.97-2.37,
revealing that there was no chromosome 17 polysomy (Table III).
 | Table IIINumber of CEP17 signals in
seven patients with lung adenocarcinoma found to have particularly
high RNU2-1 copy numbers. |
Table III
Number of CEP17 signals in
seven patients with lung adenocarcinoma found to have particularly
high RNU2-1 copy numbers.
| Case ID | RNU2-1 copy
number | CEP17 copy
number |
|---|
| LC no.1 | 91.91±2.08 | 2.27±0.05 |
| LC no.13 | 98.68±0.91 | 2.27±0.03 |
| LC no.27 | 110.00±7.19 | 1.99±0.04 |
| LC no.32 | 84.26±2.51 | 2.26±0.05 |
| LC no.38 | 85.25±2.40 | 2.28±0.07 |
| LC no.39 | 87.56±1.65 | 1.97±0.04 |
| LC no.43 | 99.66±1.11 | 2.37±0.04 |
Correlation analysis between serum
miR-1246 levels and RNU2-1 copy number
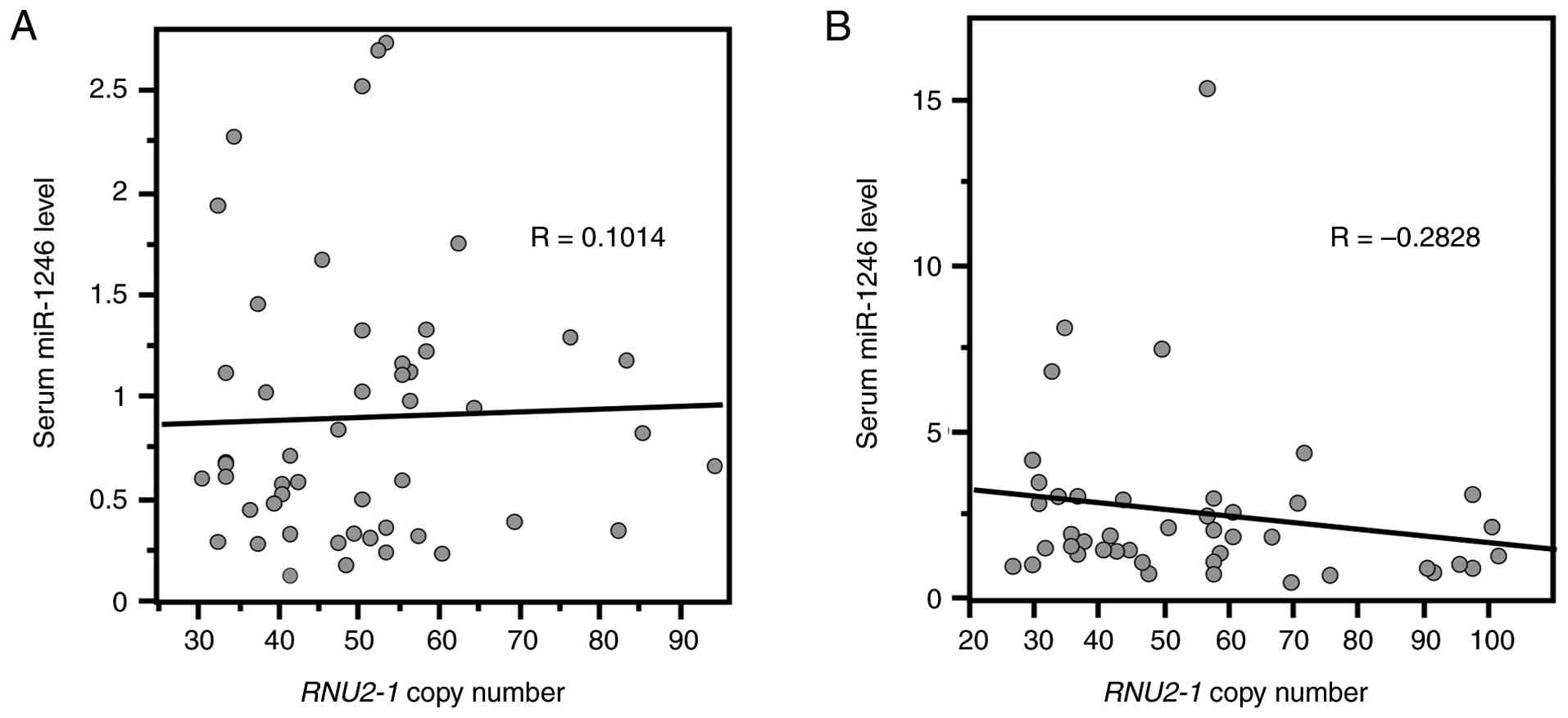
Finally, we analyzed the correlation between serum
miR-1246 levels and RNU2-1 copy numbers in the control and
stage IV lung adenocarcinoma groups. Spearman's rank correlation
coefficients between serum miR-1246 levels and RNU2-1 copy
numbers were 0.0692 in the control group (P=0.6594) and -0.2828 in
the stage IV lung adenocarcinoma one (P=0.0598), indicating no
significant correlation (Fig.
3).
Discussion
In this study, we investigated whether polymorphisms
in RNU2-1 copy number affect serum miR-1246 levels in
patients with stage IV lung adenocarcinoma and healthy individuals.
First, we confirmed that serum miR-1246 levels were significantly
higher in patients with stage IV lung adenocarcinoma than in
healthy individuals, as has been reported previously (8,12,21).
We then analyzed the RNU2-1 copy number per diploid genome
using dPCR with DNA derived from peripheral blood samples. The
median RNU2-1 copy numbers in HC and patients with stage IV
lung adenocarcinoma were 50 and 49, respectively, with no
significant difference being observed between them. We also did not
identify any correlation between RNU2-1 copy number and
serum miR-1246 level in either group. These results suggest that
RNU2-1 CNVs have no effect on serum levels of the lung
adenocarcinoma biomarker miR-1246. Other studies have previously
investigated correlations between gene copy numbers and serum
levels of the corresponding gene products, for some other genes.
Serum amylase levels, for example, were not found to correlate with
AMY2 or AMY2B copy numbers (range: 1-4), but did
correlate with that of AMY1A (range: 2-27) (23). A correlation was also demonstrated
between the copy numbers of the FCGR3B gene (range: 0-3) and
serum levels of Fc gamma receptor III-B (FCGR3B) (24). The RNU2-1 copy numbers,
which range from 26 to 101, had no effect on serum miR-1246 levels
in both healthy individuals and patients with lung
adenocarcinoma.
U2 snRNA is transcribed by RNA polymerase II. Each
repeat unit in the RNU2-1 tandem repeat contains a proximal
sequence element (PSE) and an enhancer-like distal sequence element
(DSE), located ~55 and ~220 bp, respectively, upstream of the U2
snRNA coding region. RNU2-1 transcription is regulated by
Oct-1 and snRNA-activating protein complex, which bind to DSE and
PSE, respectively (25,26).
Intracellular U2 snRNA levels were not changed by
either the overexpression or knockdown of RNU2-1 in Panc-1
cells, in a previous study. In both cases, exosomal miR-1246 levels
(i.e., RNU2-1f), an intermediate degradation product of U2 snRNA,
were found to be elevated (14).
These results suggest that the transcription and degradation of U2
snRNA are tightly regulated under such circumstances. The miR-1246
sequence within the U2 snRNA sequence contains a binding sequence
for the RNA-binding protein, SmB/B'. Exosomal miR-1246 levels were
found to be reduced in SmB/B' knockdown cells, suggesting that
SmB/B' binds to miR-1246 to protect it from degradation (14,27).
Exosomal miR-1246 was also found to be reduced by knockdown of the
RNA-binding protein, SRSF1(28);
therefore, SRSF1 may be involved in the sorting of miR-1246 into
exosomes. In the present study, no correlation was observed between
RNU2-1 copy number and serum miR-1246 levels in our HC group
(Fig. 3A). These results imply
that a significant difference in RNU2-1 copy number does not
affect circulating miR-1246 level, which may instead be regulated
by other mechanisms such as transcription of RNU2-1, or the
stability or releasing efficiency of miR-1246.
MiR-1246 expression was elevated in lung cancer
(6) and colorectal cancer
(13) tissues, and its serum
levels decreased after surgery (29). U2 snRNAs are upregulated in certain
subtypes of breast cancer, suggesting that U2 snRNA expression may
vary under specific cancer types or physiological conditions
(30). Cancer tissues may
represent one of the major sources of circulating miR-1246.
Although this study focused on germline RNU2-1 CNV,
circulating miR-1246 levels may be influenced by other
tumor-derived regulatory factors (e.g., RNU2-1
transcriptional activity and regulation of miR-1246 release via
exosomal packaging), and these mechanisms should be investigated in
future studies.
The estimated total length of the RNU2-1
tandem repeat ranges 30-492 kb when the copy number of
RNU2-1 ranges 5-82. It is impossible to amplify the entire
region of the tandem repeat even when using enzymes for long PCR.
In this study, we used dPCR to analyze RNU2-1 copy numbers.
This approach is a relatively simple method that facilitates the
quantification of gene copy number per diploid genome. The
RNU2-1 copy number per diploid genome ranged from 30 to 94
(median, 50) in the 51 healthy individuals we analyzed. An
estimated RNU2-1 copy number per diploid, based on depth of
coverage value from the 1000 Genomes Project, ranged from 2.5 to
160 (mean, 40.6) (20). The copy
number of each allele, as revealed through Southern blot analysis
and a fiber FISH approach, ranged from 5 to 63(19) and 6 to 82(18), respectively.
The major limitation of this study is relatively
small sample size. At least 53 alleles on RNU2-1 copy
numbers have been identified thus far (19); therefore, our sample size was too
small to analyze all possible copy numbers. Tessereau et al
estimated RNU2-1 copy numbers based on data from public
genomics databases (20). Schaap
et al analyzed RNU2-1 copy numbers using DNA from 270
individuals by pulsed-field gel electrophoresis and Southern blot
(19). We drew histograms of
RNU2-1 copy numbers using their data supplied as the
additional file 3 in reference (19) or our dataset and then reconfirmed
whether the sample size impacted on distribution range of copy
number. The distribution range of copy number in this study was
consistent with that in these two reports, although our sample size
was quite smaller than that in these two reports.
The second major limitation of this study is that
age was significantly different between the patient and HC group.
We performed stepwise regression analysis and correlation analysis
to examine whether age affected serum miR-1246 levels. No
significant correlation was observed between age and serum miR-1246
levels in both HC and patients with lung adenocarcinoma; thus, we
concluded that the significant difference in age distribution
between the patients and HC did not affect the results of this
study. However, several age-dependent serum microRNAs were reported
in large cohort studies (31), and
further studies with increased sample size will be needed.
In this study, we analyzed serum levels of miR-1246
and copy number of RNU2-1 and found that copy number
polymorphism of RNU2-1 has no effect on serum levels of
miR-1246 as a biomarker for lung adenocarcinoma.
Supplementary Material
(A) Correlation between serum miR-1246
levels and age in patients with lung adenocarcinoma. (B)
Correlation between serum miR-1246 levels and age in all healthy
controls of the present study. Young participants (20-22 years
old), whose samples were collected on the same day, are shown as
black dots. (C) Correlation between serum miR-1246 levels and age
in healthy controls, excluding young participants (shown as black
dots in B). (D) Correlation between serum miR-1246 levels and age
in healthy controls (64-91 years old) from our previous study (21).
Lung adeno., lung adenocarcinoma; HC, healthy controls; miR,
microRNA.
(A) Histogram of RNU2-1 copy number
per diploid in the current healthy controls determined by digital
PCR. (B) Histogram of RNU2-1 copy number per diploid in the current
patients with lung adenocarcinoma determined by digital PCR. (C)
Histogram of RNU2-1 copy numbers per diploid was generated using
the data supplied as the Additional File 3 in Schaap et al
(19), which included 270 individuals and was analyzed by
pulsed-field gel electrophoresis and Southern blot analysis. Lung
adeno., lung adenocarcinoma; HC, healthy controls; miR, microRNA;
PCR, polymerase chain reaction.
Acknowledgements
The authors thank Dr. Hiroaki Ohnishi (Kyorin
University, Mitaka, Tokyo, Japan), Dr. Haruhiro Saito, Dr. Hiroyuki
Ito, Dr. Chie Morohashi (Kanagawa Cancer Center, Yokohama,
Kanagawa, Japan), and Dr. Yataro Daigo (University of Tokyo, Tokyo,
Japan) for their assistance with sample collection.
Funding
Funding: This study was supported in part by JSPS KAKENHI [grant
nos. JP20K07791 and JP22H04923 (CoBiA)].
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
MU and TA conducted the experiments and wrote the
manuscript. MU and TA designed the study and interpreted the
experimental results. SS performed the pathological diagnoses of
the patients. SS and YM prepared the specimens. MU and TA confirmed
the authenticity of the raw data. All of the authors have read and
approved the final version of the manuscript.
Ethics approval and consent to
participate
This study's protocol was approved by the Ethics
Committees of the Faculty of Health Sciences, Kyorin University
(approval nos. 2022-30 and 2023-22) and Kanagawa Cancer Center
(approval no. 2023epidemiology-84). Signed informed consent was
obtained from all of the participants.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Morin RD, O'Connor MD, Griffith M,
Kuchenbauer F, Delaney A, Prabhu AL, Zhao Y, McDonald H, Zeng T,
Hirst M, et al: Application of massively parallel sequencing to
microRNA profiling and discovery in human embryonic stem cells.
Genome Res. 18:610–621. 2008.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Takeshita N, Hoshino I, Mori M, Akutsu Y,
Hanari N, Yoneyama Y, Ikeda N, Isozaki Y, Maruyama T, Akanuma N, et
al: Serum microRNA expression profile: miR-1246 as a novel
diagnostic and prognostic biomarker for oesophageal squamous cell
carcinoma. Br J Cancer. 108:644–652. 2013.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Shimomura A, Shiino S, Kawauchi J,
Takizawa S, Sakamoto H, Matsuzaki J, Ono M, Takeshita F, Niida S,
Shimizu C, et al: Novel combination of serum microRNA for detecting
breast cancer in the early stage. Cancer Sci. 107:326–334.
2016.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Xu YF, Hannafon BN, Zhao YD, Postier RG
and Ding WQ: Plasma exosome miR-196a and miR-1246 are potential
indicators of localized pancreatic cancer. Oncotarget.
8:77028–77040. 2017.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Moshiri F, Salvi A, Gramantieri L,
Sangiovanni A, Guerriero P, De Petro G, Bassi C, Lupini L, Sattari
A, Cheung D, et al: Circulating miR-106b-3p, miR-101-3p and
miR-1246 as diagnostic biomarkers of hepatocellular carcinoma.
Oncotarget. 9:15350–15364. 2018.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Zhang WC, Chin TM, Yang H, Nga ME, Lunny
DP, Lim EKH, Sun LL, Pang YH, Leow YN, Malusay SRY, et al:
Tumour-initiating cell-specific miR-1246 and miR-1290 expression
converge to promote non-small cell lung cancer progression. Nat
Commun. 7(11702)2016.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Baraniskin A, Nöpel-Dünnebacke S, Ahrens
M, Jensen SG, Zöllner H, Maghnouj A, Wos A, Mayerle J, Munding J,
Kost D, et al: Circulating U2 small nuclear RNA fragments as a
novel diagnostic biomarker for pancreatic and colorectal
adenocarcinoma. Int J Cancer. 132:E48–E57. 2013.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Mazières J, Catherinne C, Delfour O, Gouin
S, Rouquette I, Delisle MB, Prévot G, Escamilla R, Didier A,
Persing DH, et al: Alternative processing of the U2 small nuclear
RNA produces a 19-22nt fragment with relevance for the detection of
non-small cell lung cancer in human serum. PLoS One.
8(e60134)2013.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Kuhlmann JD, Baraniskin A, Hahn SA, Mosel
F, Bredemeier M, Wimberger P, Kimmig R and Kasimir-Bauer S:
Circulating U2 small nuclear RNA fragments as a novel diagnostic
tool for patients with epithelial ovarian cancer. Clin Chem.
60:206–213. 2014.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Kuhlmann JD, Wimberger P, Wilsch K, Fluck
M, Suter L and Brunner G: Increased level of circulating U2 small
nuclear RNA fragments indicates metastasis in melanoma patients.
Clin Chem Lab Med. 53:605–611. 2015.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Baraniskin A, Zaslavska E,
Nöpel-Dünnebacke S, Ahle G, Seidel S, Schlegel U, Schmiegel W, Hahn
S and Schroers R: Circulating U2 small nuclear RNA fragments as a
novel diagnostic biomarker for primary central nervous system
lymphoma. Neuro Oncol. 18:361–367. 2016.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Köhler J, Schuler M, Gauler TC,
Nöpel-Dünnebacke S, Ahrens M, Hoffmann AC, Kasper S, Nensa F, Gomez
B, Hahnemann M, et al: Circulating U2 small nuclear RNA fragments
as a diagnostic and prognostic biomarker in lung cancer patients. J
Cancer Res Clin Oncol. 142:795–805. 2016.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Zhang Q, Jeppesen DK, Higginbotham JN,
Graves-Deal R, Trinh VQ, Ramirez MA, Sohn Y, Neininger AC, Taneja
N, McKinley ET, et al: Supermeres are functional extracellular
nanoparticles replete with disease biomarkers and therapeutic
targets. Nat Cell Biol. 23:1240–1254. 2021.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Xu YF, Hannafon BN, Khatri U, Gin A and
Ding WQ: The origin of exosomal miR-1246 in human cancer cells. RNA
Biol. 16:770–784. 2019.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Van Arsdell SW and Weiner AM: Human genes
for U2 small nuclear RNA are tandemly repeated. Mol Cell Biol.
4:492–499. 1984.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Lindgren V, Ares M Jr, Weiner AM and
Francke U: Human genes for U2 small nuclear RNA map to a major
adenovirus 12 modification site on chromosome 17. Nature.
314:115–116. 1985.PubMed/NCBI View
Article : Google Scholar
|
|
17
|
Pavelitz T, Rusché L, Matera AG, Scharf JM
and Weiner AM: Concerted evolution of the tandem array encoding
primate U2 snRNA occurs in situ, without changing the cytological
context of the RNU2 locus. EMBO J. 14:169–177. 1995.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Tessereau C, Buisson M, Monnet N, Imbert
M, Barjhoux L, Schluth-Bolard C, Sanlaville D, Conseiller E, Ceppi
M, Sinilnikova OM and Mazoyer S: Direct visualization of the highly
polymorphic RNU2 locus in proximity to the BRCA1 gene. PLoS One.
8(e76054)2013.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Schaap M, Lemmers RJLF, Maassen R, van der
Vliet PJ, Hoogerheide LF, van Dijk HK, Baştürk N, de Knijff P and
van der Maarel SM: Genome-wide analysis of macrosatellite repeat
copy number variation in worldwide populations: Evidence for
differences and commonalities in size distributions and size
restrictions. BMC Genomics. 14(143)2013.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Tessereau C, Lesecque Y, Monnet N, Buisson
M, Barjhoux L, Léoné M, Feng B, Goldgar DE, Sinilnikova OM, Mousset
S, et al: Estimation of the RNU2 macrosatellite mutation rate by
BRCA1 mutation tracing. Nucleic Acids Res. 42:9121–9130.
2014.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Aiso T and Ueda M: 5'-isomiR is the most
abundant sequence of miR-1246, a candidate biomarker of lung
cancer, in serum. Mol Med Rep. 27(92)2023.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Wang X, Xing D, Liu Z, Zhang Y, Cheng B,
Sun S, Wang Q and Dong L: Establishment and evaluation of digital
PCR methods for HER2 copy number variation in breast cancer. Anal
Bioanal Chem. 415:725–733. 2023.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Nayema Z, Sato T, Kannon T, Tsujiguchi H,
Hosomichi K, Nakamura H and Tajima A: Genetic factors associated
with serum amylase in a Japanese population: Combined analysis of
copy-number and single-nucleotide variants. J Hum Genet.
68:313–319. 2023.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Willcocks LC, Lyons PA, Clatworthy MR,
Robinson JI, Yang W, Newland SA, Plagnol V, McGovern NN, Condliffe
AM, Chilvers ER, et al: Copy number of FCGR3B, which is associated
with systemic lupus erythematosus, correlates with protein
expression and immune complex uptake. J Exp Med. 205:1573–1582.
2008.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Jawdekar GW and Henry RW: Transcriptional
regulation of human small nuclear RNA genes. Biochim Biophys Acta.
1779:295–305. 2008.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Su Y, Wu J, Chen W, Shan J, Chen D, Zhu G,
Ge S and Liu Y: Spliceosomal snRNAs, the essential players in
pre-mRNA processing in eukaryotic nucleus: From biogenesis to
functions and spatiotemporal characteristics. Adv Biol (Weinh).
8(e2400006)2024.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Tosar JP, Cayota A and Witwer K: Exomeres
and supermeres: Monolithic or diverse? J Extracell Biol.
1(e45)2022.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Xu YF, Xu X, Gin A, Nshimiyimana JD,
Mooers BHM, Caputi M, Hannafon BN and Ding WQ: SRSF1 regulates
exosome microRNA enrichment in human cancer cells. Cell Commun
Signal. 18(130)2020.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Ogata-Kawata H, Izumiya M, Kurioka D,
Honma Y, Yamada Y, Furuta K, Gunji T, Ohta H, Okamoto H, Sonoda H,
et al: Circulating exosomal microRNAs as biomarkers of colon
cancer. PLoS One. 9(e92921)2014.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Dvinge H, Guenthoer J, Porter PL and
Bradley RK: RNA components of the spliceosome regulate tissue- and
cancer-specific alternative splicing. Genome Res. 29:1591–1604.
2019.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Ameling S, Kacprowski T, Chilukoti RK,
Malsch C, Liebscher V, Suhre K, Pietzner M, Friedrich N, Homuth G,
Hammer E and Völker U: Associations of circulating plasma microRNAs
with age, body mass index and sex in a population-based study. BMC
Med Genomics. 8(61)2015.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Goldstraw P, Chansky K, Crowley J,
Rami-Porta R, Asamura H, Eberhardt WEE, Nicholson AG, Groome P,
Mitchell A, Bolejack V, et al: The IASLC lung cancer staging
project: Proposals for revision of the TNM stage groupings in the
forthcoming (eighth) edition of the TNM classification for lung
cancer. J Thorac Oncol. 11:39–51. 2016.PubMed/NCBI View Article : Google Scholar
|