Introduction
Rheumatoid arthritis (RA) is a chronic autoimmune
inflammatory disorder with an incidence rate of up to 1% globally
among population, which results in irreversible cartilage and bone
damage if left untreated (1). The
efficacy of conventional anti-RA medications (such as non-steroidal
anti-inflammatory drugs) is restricted and some specific drugs
(such as methotrexate and sulfasalazine) have been associated with
adverse effects, such as nausea, alopecia, stomatitis and
hepatotoxicity (2). Although
numerous studies have explored the pathogenesis of RA, research
focusing on the specific role of macrophages in this process
remains limited. Notably, macrophages have been implicated in the
autoimmune inflammatory response of RA (3-5).
Furthermore, an important pathological characteristic of RA is the
accumulation and activation of monocytes/macrophages within the
synovial tissue (6,7).
Activated macrophages are classified into two types,
namely classically activated M1 macrophages and alternatively
activated M2 macrophages (8,9). M1
macrophages are capable of directly phagocytosing cancer cells and
are induced by TNF and IFN-γ (10). By contrast, M2 macrophages are
induced by steroids, IL-4 and IL-10 released by type 2 helper T
cells (11), and secrete
anti-inflammatory factors, diminish pro-inflammatory factors and
augment scavenger receptor levels (12).
Spermidine (SPD) is a polyamine synthesized from
putrescine, which is indispensable for DNA synthesis and cell
proliferation (13,14). In addition to its anti-inflammatory
and antioxidant capabilities SPD has a key role in cell autophagy
and transcription (15-17),
as well as in suppressing the secretion of pro-inflammatory
cytokines and related chemokines (such as CCL3 and CXCL8) (18,19).
SPD has also been shown to impede the NF-κB pathway in macrophages
within an autoimmune encephalomyelitis mouse model (19).
Paoletti et al (6) indicated that the number of M1
macrophages is elevated in the synovium of patients with RA. In
addition, macrophages tend to polarize towards the M1 phenotype,
augment the release of pro-inflammatory factors and promote
synovitis in mice with collagen-induced arthritis (20). These findings suggest that M1
macrophages are predominant in the synovium of patients with RA.
Therefore, based on the documented anti-inflammatory and
antioxidant properties of SPD, and its reported role in suppressing
macrophage pro-inflammatory cytokine secretion and regulating the
NF-κB pathway in an autoimmune encephalomyelitis mouse model, SPD
may alleviate RA symptoms by modulating macrophage polarization and
cytokine release; however, the underlying mechanisms of this remain
to be elucidated. Accordingly, the present study hypothesized that
SPD may function by regulating activation of the NF-κB pathway.
Suitably, RAW 264.7 cells were stimulated with lipopolysaccharide
(LPS) to establish a relevant inflammatory cell model, and were
subsequently treated with varying SPD concentrations. The cytokine
levels (TNF-α, IL-1β and IL-6), mRNA expression levels (iNOS
for M1 phenotype and Arg-1 for M2 phenotype) and
phosphorylation levels of NF-κB-associated proteins in RAW 264.7
macrophages from different groups were then determined to evaluate
the impact of SPD on LPS-induced inflammation and the associated
underlying mechanisms. The findings of the present study may offer
an experimental basis for exploring SPD as a potential natural
anti-inflammatory agent against RA; however, further in vivo
and clinical studies are still required.
Materials and methods
Cell culture
RAW 264.7 cells were obtained from the Institute for
Advanced Study of Central South University (Changsha, China). DMEM
(Shanghai BasalMedia Technologies Co., Ltd.) supplemented with 10%
FBS (Shanghai BasalMedia Technologies Co., Ltd.), 100 U/ml
penicillin and 25 µg/ml amphotericin B (Hyclone™;
Cytiva) was used as the cell culture medium, with incubation
conditions set at 37˚C and 5% CO2 (21).
Cell cytotoxicity assay
Cytotoxicity assays were performed using the Cell
Counting Kit-8 (CCK-8) according to the manufacturer's instructions
(NCM Biotech Co., Ltd.). Briefly, RAW 264.7 cells were seeded into
96-well plates at a density of 1.0x104 cells/well for 24
h, followed by treatment with varying concentrations (0, 100, 200,
400, 800 or 1,600 µM) of SPD (≥99% gas chromatography grade;
MilliporeSigma) for additional durations of 6, 12, 24 and 48 h at
37˚C in a 5% CO2 incubator. Subsequently, for the
cytotoxicity assay, 10 µl CCK-8 solution was added to each well and
incubated for 1 h. Finally, the absorbance was measured at a
wavelength of 450 nm and cell viability was calculated based on the
measured optical density values.
Flow cytometry
RAW 264.7 cells were inoculated into a 6-well plate
at a density of 4x105 cells/well and cultured for 24 h
until they adhered to the well walls. Pretreatment with varying
concentrations (100, 200, 400 or 800) of SPD or vehicle control
(complete DMEM) for 1 h was followed by stimulation with 1 µg/ml
LPS 24 h at 37˚C in a 5% CO2 incubator, and this cell
treatment protocol was applied uniformly prior to all subsequent
experiments in this study. Prior to antibody staining, cells were
stained with Zombie NIR™ Fixable Viability Kit (1:1,000;
cat. no. 423105; BioLegend) for 15 min at room temperature in the
dark to distinguish live/dead cells. Subsequently, the cells were
gently harvested, washed twice with PBS and stained with
CD86-phycoerythrin (1:100; clone GL-1; cat. no. 553692; BD
Biosciences) and CD206-BUV395 (1:100; clone Y17-505; cat. no.
568817; BD Biosciences) for 30 min at 4˚C in the dark. Fluorescence
intensity was assessed using the BD FACSCanto™ II flow
cytometer (BD Biosciences), and the collected data were analyzed
using BD FACSDiva™ software (Version 8.0; BD
Biosciences).
Gating hierarchy was as follows: i) Forward scatter
(FSC)/side scatter gating to exclude debris; ii) singlet gating
(FSC-area vs. FSC-height); iii) live cell gating (Zombie
NIR™-negative); and iv) CD86/CD206 double staining
gating. Compensation was performed using single-stained controls
and fluorescence minus one control and isotype-matched antibodies
(PE Rat IgG2a; 1:100; cat. no. 553930; and BUV395 Rat IgG2a; 1:100;
cat. no. 563556; both BD Biosciences) were used to set the gating
thresholds.
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA was isolated from the cells using the Cell
Total RNA Extraction Kit (cat. no. G3694; Wuhan Servicebio
Technology Co., Ltd.) according to the manufacturer's instructions.
The concentration of total RNA was quantified using a NanoDrop
spectrophotometer (NanoDrop; Thermo Fisher Scientific, Inc.),
whereas cDNA was synthesized using the PrimeScript™ RT
kit (Takara Bio, Inc.). Gene-specific primers were synthesized by
Sangon Biotech (Shanghai) Co., Ltd. and qPCR was carried out using
the TB Green® Premix Ex Taq™ II kit as per
the manufacturer's instructions (Wuhan Servicebio Technology Co.,
Ltd.). The primer sequences are listed in Table I. RNA amplification reactions were
performed using the StepOnePlus Real-Time PCR System (Thermo Fisher
Scientific, Inc.) with an initial denaturation step at 95˚C for 30
sec, followed by 40 cycles of amplification (denaturation at 95˚C
for 5 sec and annealing at 60˚C for 34 sec), and finally, a
dissociation curve was generated. After amplification, the data
were analyzed using the 2-ΔΔCq method (22) with the mouse Gapdh gene as
the reference (23).
 | Table IPrimer sequences. |
Table I
Primer sequences.
| Primer name | Sequence
(5'-3') |
|---|
| Gapdh | Forward:
TCAACGGCACAGTCAAGG |
| | Reverse:
ACTCCACGACATACTCAGC |
| Tnf | Forward:
TACTGAACTTCGGGGTGATCGGTC |
| | Reverse:
CAGCCTTGTCCCTTGAAGAGAACC |
| Il6 | Forward:
CTCCCAACAGACCTGTCTATAC |
| | Reverse:
CCATTGCACAACTCTTTTCTCA |
| Il1b | Forward:
CACTACAGGCTCCGAGATGAAC AAC |
| | Reverse:
TGTCGTTGCTTGGTTCTCCTTGTAC |
| Nos2 | Forward:
CACCTTGGAGTTCACCCAGT |
| | Reverse:
ACCACTCGTACTTGGGATGC |
| Arg1 | Forward:
CATATCTGCCAAAGACATCGTG |
| | Reverse:
GACATCAAAGCTCAGGTGAATC |
| Il10 | Forward:
CACAAAGCAGCCTTGCAGAA |
| | Reverse:
AGAGCAGGCAGCATAGCAGTG |
ELISA
The cells were resuspended in complete DMEM, seeded
into 6-well plates and incubated for 24 h. After a 1 h pretreatment
with SPD, LPS was added to the co-culture system. Cytokine levels
in the cell supernatants were quantified using a ELISA kits (Mouse
IL-6 ELISA kit, cat. no. EMC004.96; Mouse IL-10 ELISA kit, cat. no.
EMC005.96; both Neobioscience Technology Co., Ltd.) according to
the manufacturer's instructions.
Western blotting
The cells were seeded into 6-well plates and
incubated for 24 h. Subsequently, the cells were pretreated with
SPD for 1 h and then stimulated with LPS for another 24 h.
Immediately after stimulation, total proteins were extracted on ice
using RIPA Lysis Buffer (Wuhan Servicebio Technology Co., Ltd.).
The protein concentration was determined using a BCA assay (BCA
Protein Assay Kit; cat. no. P0012; Beyotime Biotechnology). Equal
amounts of protein (20 µg per lane) were separated by 12% SDS-PAGE
(cat. no. BL522A; Labgic Technology Co., Ltd.) and transferred onto
PVDF membranes (cat. no. IPVH10100, Merckmillipore Co., Ltd.). The
membranes were blocked with 5% skimmed milk in Tris-Buffered Saline
with Tween-20 (TBST; cat. no. G0004-1L; Wuhan Servicebio Technology
Co., Ltd.) for 1 h at room temperature. The membranes were then
incubated with the following primary antibodies overnight at 4˚C:
p65 (1:1,000; cat. no. 8242; Cell Signaling Technology, Inc.),
phosphorylated (p)-p65 (1:1,000; cat. no. 3033; Cell Signaling
Technology, Inc.), IκBα (1:1,000; cat. no. 4814; Cell Signaling
Technology, Inc.), p-IκBα (1:1,000; cat. no. 2859; Cell Signaling
Technology, Inc.) and β-tubulin (1:1,000; cat. no. 10094-1-AP;
Proteintech Group, Inc.). After washing with TBST, the membranes
were incubated with the corresponding HRP-linked secondary
antibodies for 1 h at room temperature, namely anti-rabbit IgG
(1:5,000; cat. no. 7074; Cell Signaling Technology, Inc.) and
anti-mouse IgG (1:5,000; cat. no. 7076; Cell Signaling Technology,
Inc.). The protein bands were visualized using a super-sensitive
ECL chemiluminescent substrate (cat. no. BL5965A; Labgic Technology
Co., Ltd.) and detected with a chemiluminescence imaging system
(ChemiDoc Imaging System, Bio-Rad Laboratories, Inc.). The gray
values of the target bands were semi-quantified using ImageJ
software version 1.53 (National Institutes of Health) to determine
the relative protein expression levels and β-tubulin was used as an
internal control.
Statistical analysis
Statistical analysis and data presentation were
performed using SPSS 26.0 (IBM Corp.) and GraphPad Prism 8.0
(Dotmatics) software. Differences among multiple groups were
evaluated using one-way ANOVA, followed by Tukey's multiple
comparison test for pairwise comparisons. P<0.05 was considered
to indicate a statistically significant difference.
Results
Cytotoxic effects of SPD on RAW 264.7
cells
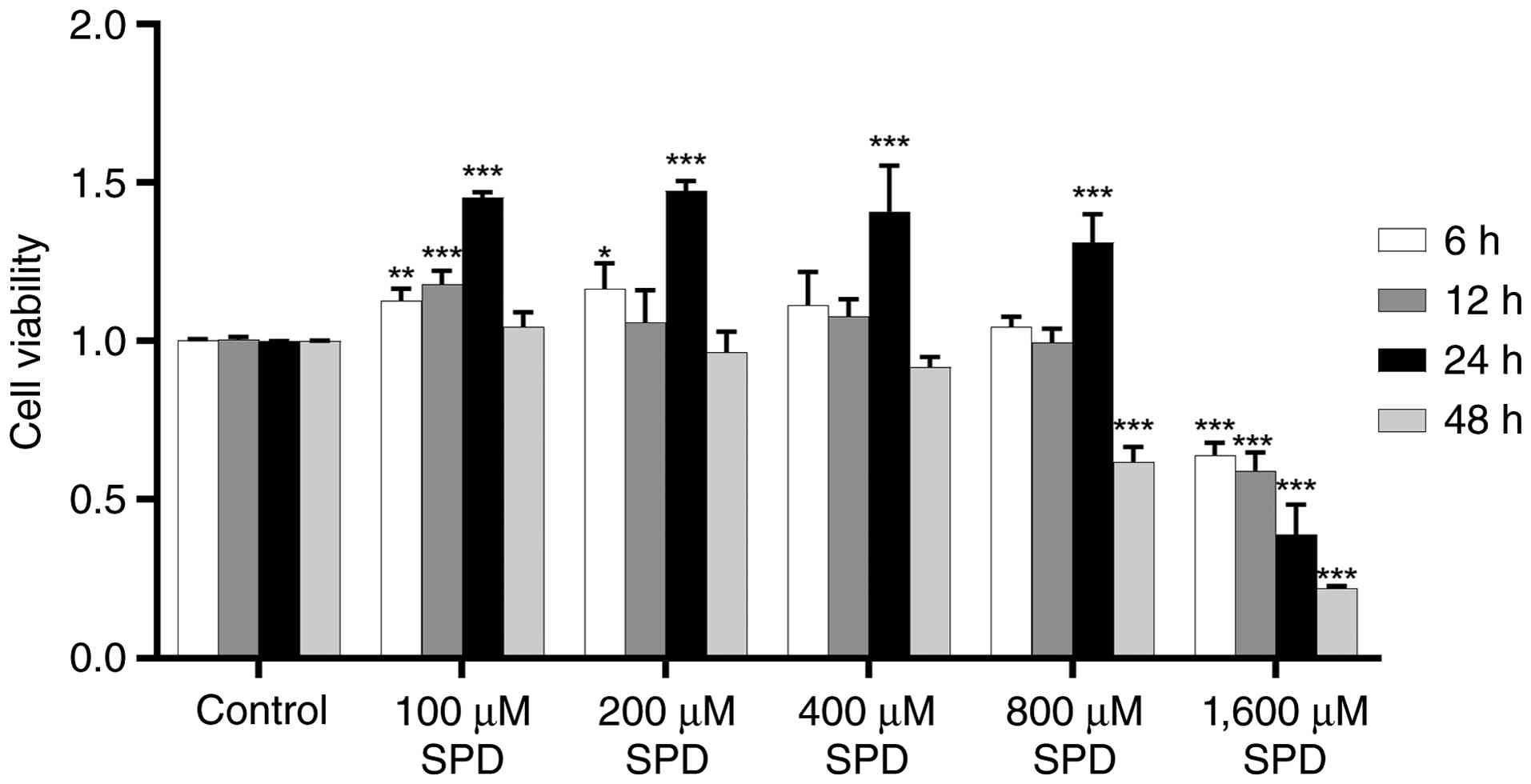
To evaluate whether SPD treatment induced
cytotoxicity, cells were incubated with different concentrations of
SPD for 6, 12, 24 and 48 h. As shown in Fig. 1, treatment with 100 and 200 µM SPD
for 6 h significantly enhanced RAW 264.7 cell viability
(P<0.05). After 12 h of treatment, only 100 µM SPD significantly
increased cell viability (P<0.001). After 24 h of 0-800 µM SPD
treatment, RAW 264.7 macrophage viability increased significantly
(P<0.001), however the difference in cell viability between
different concentration subgroups (200, 400, 600 and 800 µM) was
not significant (P>0.05). Treatment with 1,600 µM SPD induced
cytotoxicity compared with that of the control group (DMEM)
(P<0.001). After 48 h of treatment, none of the SPD
concentrations improved cell viability. Therefore, SPD
concentrations of 0-800 µM and a 24 h treatment duration were
chosen for subsequent experimentation.
Effect of SPD on LPS-induced
polarization of RAW 264.7 cells
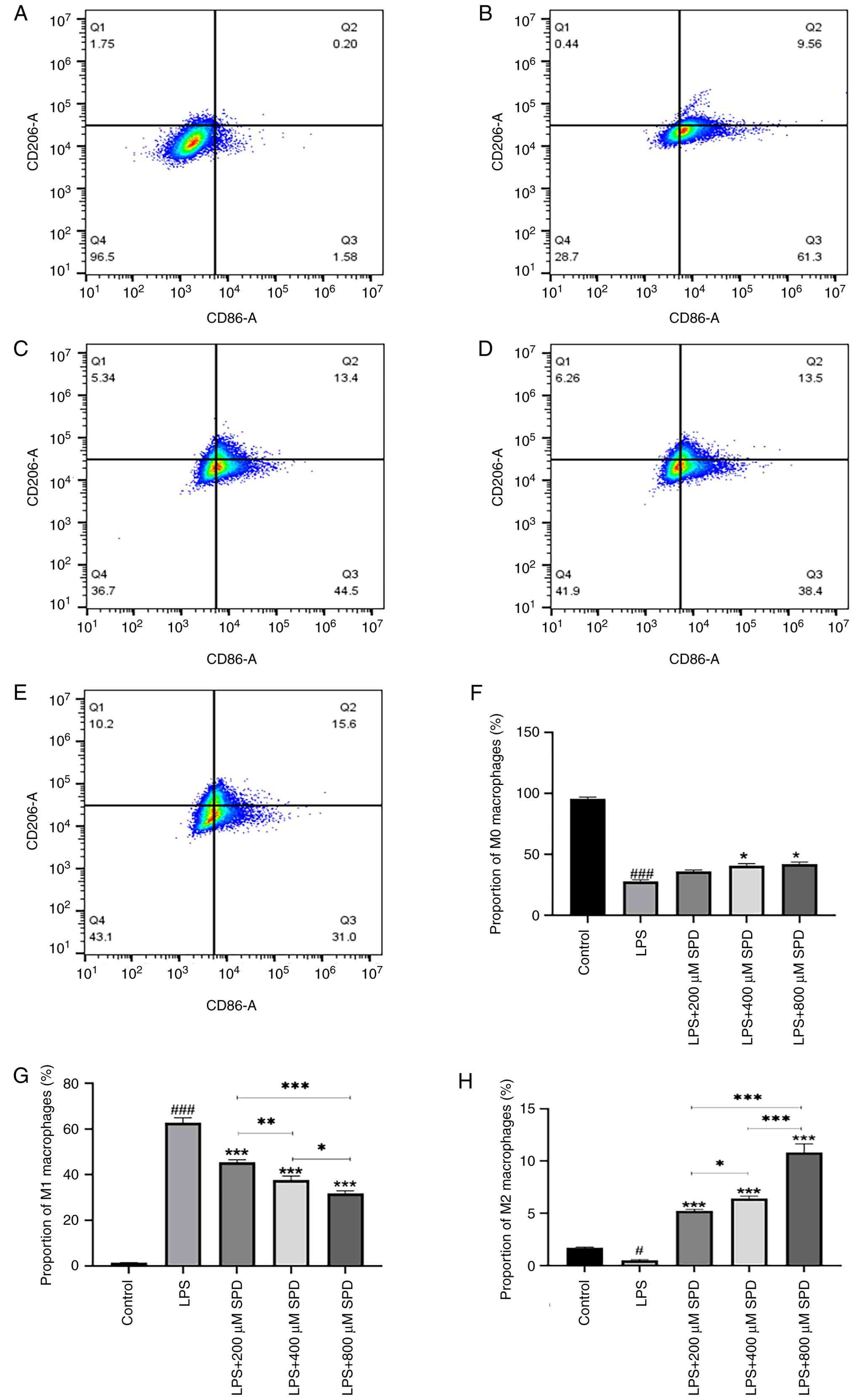
M1-type macrophages express high levels of CD86,
whereas M2-type macrophages express CD206(24). The proportions of M1 and M2 cells
after LPS-induced polarization of RAW 264.7 cells were ascertained
using flow cytometry. CD86-CD206- represents
M0 macrophages, CD86+CD206- represents M1
macrophages and CD86-CD206+ represents M2
macrophages. The findings revealed that SPD treatment diminished
macrophage CD86 molecule levels and shifted macrophage polarization
from a pro-inflammatory M1 phenotype towards an anti-inflammatory
M2 phenotype after 24 h; this effect was most notable in response
to 800 µM SPD (Fig. 2A-E). The
results revealed that LPS stimulation significantly increased the
percentage of M1 macrophages and decreased the percentage of M0 or
M2 macrophages compared with the control. Co-treatment with SPD
dose-dependently reversed this effect. Specifically, increasing SPD
concentrations (200, 400 and 800 µM) gradually decreased the M1
percentage while concurrently increasing the M2 percentage. The
percentage of M0 macrophages remained largely unchanged across SPD
groups. These data indicate that SPD can redirect LPS-induced M1
polarization toward an M2 phenotype in a dose-dependent manner
(P<0.05; Fig. 2F-H).
Effect of SPD on LPS-activated mRNA
expression of inflammatory factors in RAW 264.7 cells
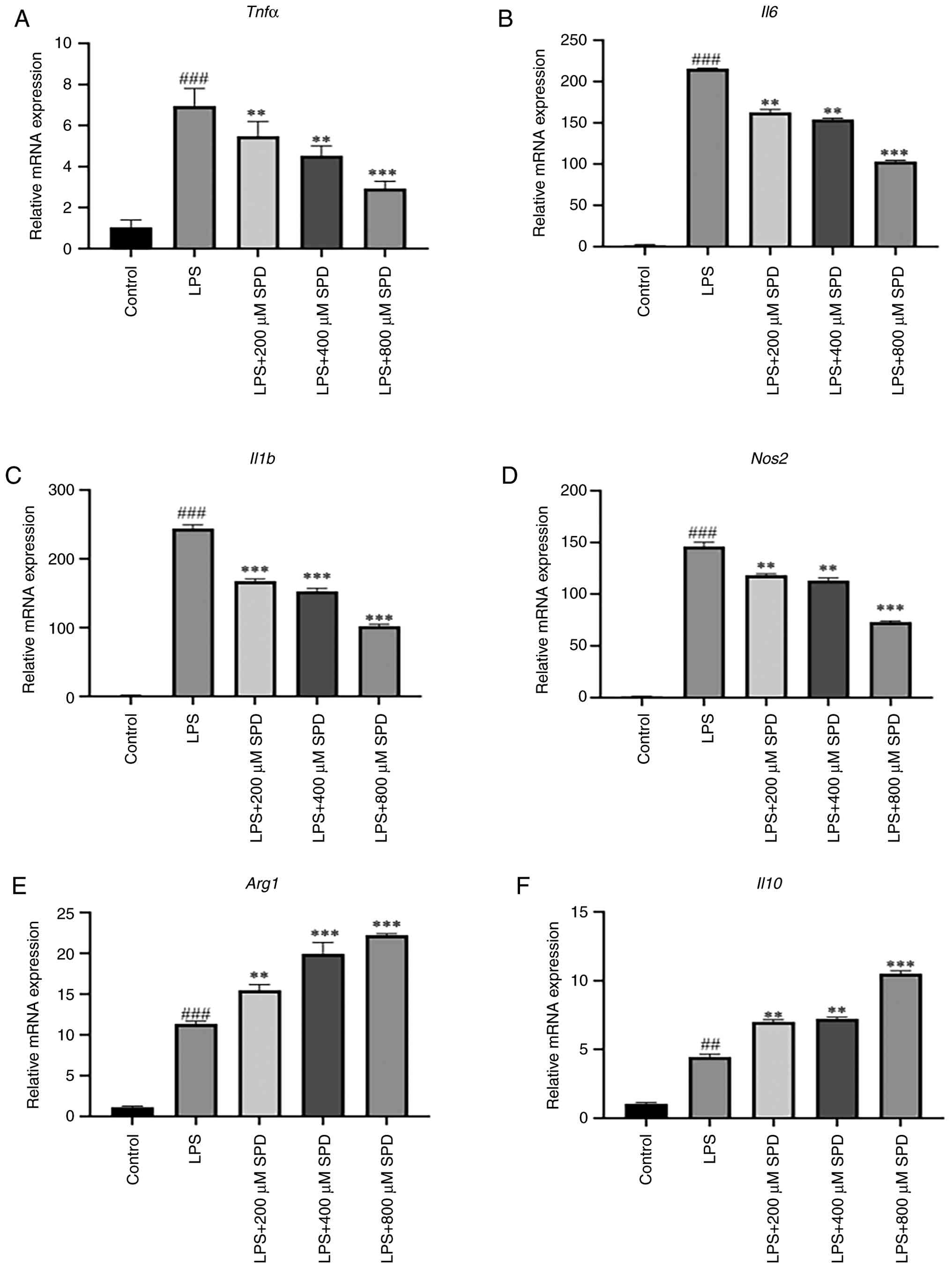
RT-qPCR was performed to assess the ability of LPS
to suppress the expression of inflammatory cytokines. LPS
significantly elevated the mRNA expression levels of Tnfα,
Il6, Il1b and nitric oxide synthase (Nos2)
compared with those in the control group (Fig. 3A-D). The mRNA levels of
Tnfα, Il6, Il1b and Nos2 then gradually
diminished with increasing SPD concentrations, thereby suggesting a
significant dose-dependent effect (P<0.01). Relative to those in
the control group, LPS significantly elevated the mRNA expression
levels of arginase 1 (Arg1) and Il10. Compared with
those in the LPS group, the mRNA expression levels of arginase 1
(Arg1) and Il10 were significantly increased in the
200 and 400 µM SPD groups (P<0.01), and peaked after treatment
with 800 µM SPD (Fig. 3E and
F). Furthermore, SPD treatment
reduced the mRNA expression levels of Tnfα, Il6,
Il1b and Nos2 at all tested SPD concentrations. These
results implied that SPD inhibited autoinflammation.
Effect of SPD on inflammatory factor
levels in RAW 264.7 cells
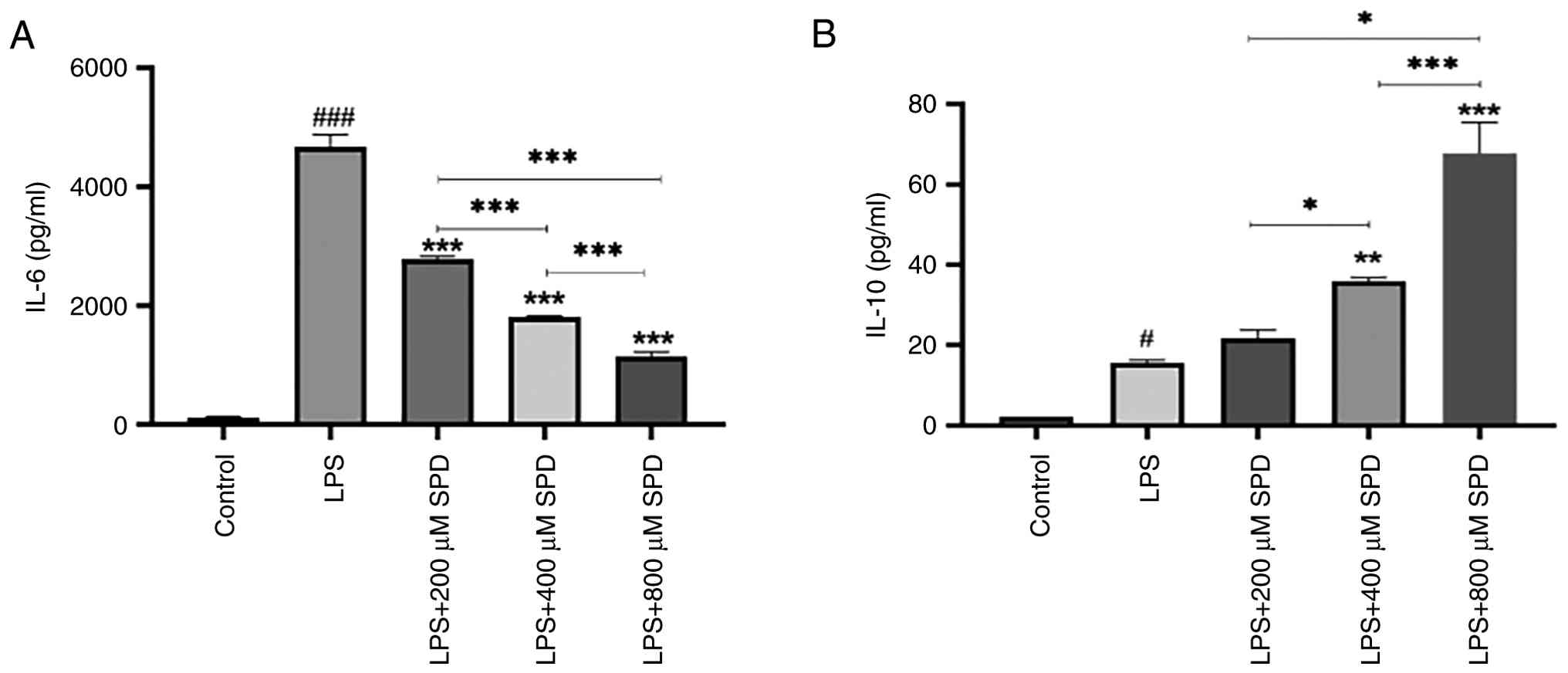
A major function of M1-polarized macrophages is the
production of pro-inflammatory cytokines. Initially, IL-6 secreted
by RAW 264.7 cells into the conditioned medium was examined using
ELISA. LPS notably stimulated IL-6 secretion by macrophages
compared with that in the control group (Fig. 4A). This suggested that RAW 264.7
cells were activated by LPS stimulation, thereby inducing an
inflammatory response. In addition, SPD markedly reduced IL-6
production in a dose-dependent manner compared with that in the LPS
group. LPS stimulation led to an elevation in IL-10 levels compared
with those in the control group. IL-10 was not significantly
induced at the lower SPD concentration (200 µM) but was
significantly enhanced compared with that in the LPS group at
higher concentrations (400 and 800 µM) (Fig. 4B). These results indicated that SPD
may promote anti-inflammatory effects by both inhibiting IL-6 and
inducing IL-10 secretion.
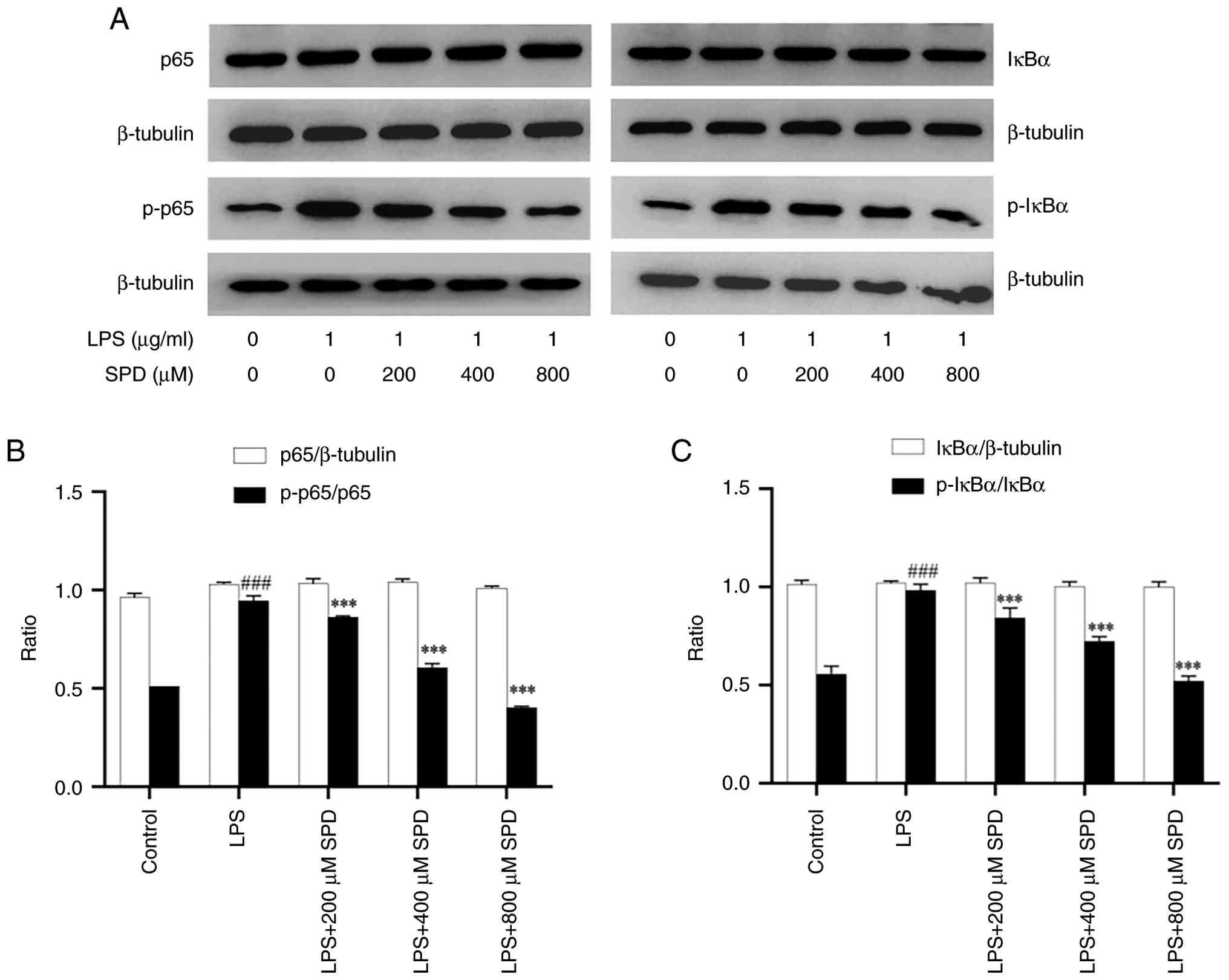
Influence of SPD on LPS-induced
phosphorylation of p65 and IκBα in macrophages
NF-κB is a key target pathway activated by
inflammation (25). Therefore,
NF-κB activation was investigated through western blotting to
determine whether SPD inhibited its activation (Fig. 5A). The levels of the NF-κB
pathway-associated proteins p65, p-p65, IκBα and p-IκBα were
analyzed after 24 h of treatment with LPS and SPD. The expression
levels of p-p65 and p-IκBα were significantly increased in
macrophages after 24 h of LPS stimulation compared with that in the
control group (Fig. 5B and
C). Notably, in cultured RAW 264.7
cells, three SPD concentrations (200, 400 and 800 µM) inhibited the
LPS-induced increase in p-p65 and p-IκBα protein expression after
24 h of incubation in a dose-dependent manner. Treatment with all
tested SPD concentrations for 24 h markedly inhibited the
activation of both p-p65 and p-IκBα proteins. These results
suggested that the mechanism underlying the anti-inflammatory
effect of SPD may involve inhibition of NF-κB activation, which in
turn inhibits the expression of inflammatory genes.
Discussion
Within the present study, SPD treatment was
demonstrated to impede the shift of RAW 264.7 cells towards the M1
phenotype upon LPS stimulation. This was evidenced by the
downregulation of the M1 phenotype marker CD86 and upregulation of
the M2 phenotype marker CD206, along with the modulation of
relevant cytokine production. Compared with previous studies on the
anti-inflammatory effects of SPD (18-20),
the present study systematically provided dose-response data
regarding the effects of SPD (0-800 µM) on macrophage polarization,
combining transcriptional (RT-qPCR) and secretory (ELISA) analyses
to demonstrate the regulatory effect of SPD on pro-inflammatory and
anti-inflammatory factors. The findings of the present study
further provide a mechanistic understanding of the
anti-inflammatory properties of SPD. Alongside the reduction in the
M1 phenotype-specific surface marker, SPD reduced M1 phenotype
cytokine secretion, as determined by measuring the downregulation
of Tnfα, Il6 and Il1b gene expression. All of
these genes are implicated in RA (26).
RA is an autoimmune inflammatory disease that is
primarily characterized by bilateral symmetrical joint damage and
synovial inflammation (24,27).
During chronic inflammation, numerous types of immune cells
contribute to the development and progression of synovitis, among
which macrophages are important. The enhanced pro-inflammatory
capacity of macrophages is associated with their hyperactivation
(5,28). They subsequently secrete TNF-α,
IL-6, and IL-1β, and drive inflammation by engaging other immune
cells, thereby activating fibroblasts and promoting T-cell
polarization (5). Activated
macrophages include M1 and M2 phenotypes, with M1 macrophages being
the primary type present in patients with RA (29). Previous studies have highlighted
the key pathogenic effect of M1-type macrophages in RA (6,20).
Specifically, an imbalance between M1 and M2 macrophages is one of
the principal drivers of synovial inflammation progression in RA
(29). Therefore, macrophages are
considered potentially novel therapeutic targets for RA (30). This potential has also been
demonstrated by altered macrophage numbers and inflammatory product
levels in the synovial membrane of patients with RA (20). Following macrophage activation by
LPS, a number of signaling pathways are activated, including the
NF-κB pathway (31). In addition,
macrophage activation is tightly controlled by specific signaling
pathways such as NF-κB to maintain immune homeostasis (32).
LPS-induced inflammatory RAW 264.7 cell models have
been utilized to screen anti-inflammatory drugs in vitro
(33). Notably, LPS upregulates
CD86, a specific marker of M1-type macrophages (34). Consistent with these findings, in
the present study, the increase in CD86 protein levels following
LPS activation demonstrated its ability to induce the M1 phenotype
in RAW 264.7 cells.
SPD, a trivalent cationic polyamine compound present
in eukaryotic cells, particularly sperm (18), interacts with nucleic acids,
proteins, ATP and other polyanions to regulate autophagy and
apoptosis, foster lipid metabolism, enhance anti-inflammatory and
antioxidant activities, and augment mitochondrial metabolism and
respiration (17). Furthermore,
SPD reduces inflammation-induced damage in the body by suppressing
the production of inflammatory cytokines (35).
SPD treatment reduces joint swelling and synovitis
in mice with collagen-induced arthritis (20). In the present study, flow cytometry
revealed that LPS facilitated the conversion of RAW 264.7 cells
into M1-type macrophages, whereas SPD treatment suppressed this
effect and promoted the conversion of RAW 264.7 cells into M2-type
macrophages. Thus, SPD may exert its anti-inflammatory effects by
suppressing the LPS-driven transition of RAW 264.7 cells to the M1
phenotype. However, a limitation of the present study was the lack
of SPD-only control groups, which prevented the exclusion of
potential direct effects of SPD on macrophage polarization or NF-κB
signaling in resting cells. Future studies should aim to include
these controls to further clarify the basal regulatory effects of
SPD.
Macrophages are the primary TNF-α- and
IL-6-secreting cells in the joints of patients with RA. TNF-α and
IL-6 are central components of the RA synovial cytokine network,
stimulating osteoclast synthesis, causing bone and cartilage
degradation and damage, and inducing the release of other
pro-inflammatory mediators, leading to exacerbation of the
inflammatory response (36). The
degree of release of inflammatory factors is an important indicator
of the severity of inflammation. In the present study, LPS
stimulated RAW 264.7 cells, which increased the mRNA levels of
Tnf, Il6, Il1b and Nos2, and enhanced
IL-6 production, similar to the findings of Jeong et al
(19) and Huang et al
(37). By contrast, SPD markedly
diminished IL-6 production in RAW 264.7 cells by downregulating
Il6 expression at the mRNA level without inducing
cytotoxicity. This was consistent with previous findings in mice
with collagen-induced arthritis (20). In addition, SPD increased the
levels of M2 macrophage-related factors (Arg1 and
Il10) and promoted the secretion of IL-10, with the most
notable effect at 800 µM. This suggests that SPD may attenuate the
LPS-induced inflammatory response in macrophages by inhibiting the
secretion of pro-inflammatory factors and promoting the secretion
of the anti-inflammatory cytokine. The present study only detected
Tnf and Il1b mRNA levels and their secretion levels
remain unclear. Differences between transcriptional and secretory
levels may exist and future studies should aim to supplement ELISA
to further demonstrate these results.
In unstimulated cells, the NF-κB dimer and IκB
protein associate tightly to form a complex that resides in the
cytoplasm of inactive cells. When cells are stimulated by LPS, the
IκB protein is phosphorylated and the NF-κB dimer is no longer
inhibited. NF-κB enters the nucleus under the guidance of the
nuclear localization sequence, where it interacts with specific DNA
sequences and participates in the control of gene transcription,
thereby encoding multiple functions, such as inflammation, cell
survival and cell division (38,39).
In the present study, LPS-activated phosphorylation of p65 and IκBα
was elevated in RAW 264.7 macrophages. SPD decreased LPS-activated
phosphorylation of p65 and IκBα without altering total p65 and IκBα
protein levels. Overall, SPD mediated the NF-κB pathway by
preventing the phosphorylation of IκB-α. The present results also
suggest that deactivation of the NF-κB pathway by SPD may
downregulate pro-inflammatory gene expression. NF-κB activation
occurs rapidly (within minutes to hours) and the single time-point
measurement at 24 h in the present study may neglect the early
dynamic changes of the pathway. The 24 h time-point was chosen to
align with the detection of macrophage polarization and cytokine
secretion, which are key downstream events of NF-κB signaling in
inflammatory responses. The present study did not perform
nuclear/cytosolic fractionation or immunofluorescence to directly
confirm p65 nuclear translocation, which further limits the depth
of the NF-κB mechanism analysis. Although the present data imply
that the potential anti-inflammatory effect of SPD is achieved by
blocking the NF-κB pathway, the precise mechanism by which SPD
influences the production of cell factors that activate macrophages
warrants further investigation.
In conclusion, the present study established that
SPD exerts potent anti-inflammatory effects on RAW 264.7
macrophages. Specifically, SPD decreased the mRNA expression levels
of pro-inflammatory mediators, thereby reducing the synthesis of
inflammatory factors in LPS-activated RAW 264.7 cells. These
effects were associated with the suppression of LPS-induced
activation of the NF-κB signaling pathway, as evidenced by the
inhibition of p65 and IκBα phosphorylation. Therefore, these
findings indicate the potential of SPD as a natural drug for the
treatment of RA.
Acknowledgements
Not applicable.
Funding
Funding: The present study was supported by the Natural Science
Foundation of Hunan Provincial Department of Science and Technology
(grant no. 2025JJ90184), the Key Project of Hunan Provincial Health
Commission (grant no. C202311000138), the Key Cultivation Project
of Renshu Fund of Hunan Provincial People's Hospital (grant no.
RS2022A14) and the Natural Science Foundation of Changsha Science
and Technology Bureau (grant no. kq2403140).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
HY and FP conceptualized the present study. HY, FP,
YFZ and YZZ established the methodology. FP, ZYL, SXW, YMP and YZZ
conducted the present investigation. HY verified the experimental
design and data analysis. FP, YMP and SXW performed the data
analysis, and FP, SXW and ZYL provided the reagents. ZYL and FP
prepared the figures and tables. FP and HY conducted the formal
analysis. HY, FP and YFZ conducted data curation. FP wrote the
original draft. HY and YFZ wrote, reviewed and edited the
manuscript. HY supervised the study, conducted project
administration and acquired the funding. HY and FP confirm the
authenticity of all the raw data. All authors have read and
approved the final version of the manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
McInnes IB and Schett G: Pathogenetic
insights from the treatment of rheumatoid arthritis. Lancet.
389:2328–2337. 2017.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Smolen JS, Aletaha D, Barton A, Burmester
GR, Emery P, Firestein GS, Kavanaugh A, McInnes IB, Solomon DH,
Strand V and Yamamoto K: Rheumatoid arthritis. Nat Rev Dis Primers.
4(18001)2018.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Calabresi E, Petrelli F, Bonifacio AF,
Puxeddu I and Alunno A: One year: Pathogenesis of rheumatoid
arthritis. Clin Exp Rheumatol. 36:175–184. 2018.PubMed/NCBI
|
|
4
|
Roszkowski L and Ciechomska M: Tuning
monocytes and macrophages for personalized therapy and diagnostic
challenge in rheumatoid arthritis. Cells. 10(1860)2021.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Cutolo M, Soldano S, Gotelli E, Montagna
P, Campitiello R, Paolino S, Pizzorni C, Sulli A, Smith V and
Tardito S: CTLA4-Ig treatment induces M1-M2 shift in cultured
monocyte-derived macrophages from healthy subjects and rheumatoid
arthritis patients. Arthritis Res Ther. 23(306)2021.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Paoletti A, Rohmer J, Ly B, Pascaud J,
Rivière E, Seror R, Le Goff B, Nocturne G and Mariette X:
Monocyte/macrophage abnormalities specific to rheumatoid arthritis
are linked to miR-155 and are differentially modulated by different
TNF inhibitors. J Immunol. 203:1766–1775. 2019.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Tu J, Hong W, Guo Y, Zhang P, Fang Y, Wang
X, Chen X, Lu S and Wei W: Ontogeny of synovial macrophages and the
roles of synovial macrophages from different origins in arthritis.
Front Immunol. 10(1146)2019.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Wynn TA and Vannella KM: Macrophages in
tissue repair, regeneration, and fibrosis. Immunity. 44:450–462.
2016.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Huen SC and Cantley LG: Macrophages in
renal injury and repair. Annu Rev Physiol. 79:449–469.
2017.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Mosser DM and Edwards JP: Exploring the
full spectrum of macrophage activation. Nat Rev Immunol. 8:958–969.
2008.PubMed/NCBI View
Article : Google Scholar
|
|
11
|
Meng XM, Mak TSK and Lan HY: Macrophages
in renal fibrosis. Adv Exp Med Biol. 1165:285–303. 2019.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Han C, Yang Y, Sheng Y, Wang J, Zhou X, Li
W, Guo L, Zhang C and Ye Q: Glaucocalyxin B inhibits cartilage
inflammatory injury in rheumatoid arthritis by regulating M1
polarization of synovial macrophages through NF-κB pathway. Aging
(Albany, NY). 13:22544–22555. 2021.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Fan J, Feng Z and Chen N: Spermidine as a
target for cancer therapy. Pharmacol Res.
159(104943)2020.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Madeo F, Hofer SJ, Pendl T, Bauer MA,
Eisenberg T, Carmona-Gutierrez D and Kroemer G: Nutritional aspects
of spermidine. Annu Rev Nutr. 40:135–159. 2020.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Hofer SJ, Liang Y, Zimmermann A, Schroeder
S, Dengjel J, Kroemer G, Eisenberg T, Sigrist SJ and Madeo F:
Spermidine-induced hypusination preserves mitochondrial and
cognitive function during aging. Autophagy. 17:2037–2039.
2021.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Ni YQ and Liu YS: New insights into the
roles and mechanisms of spermidine in aging and age-related
diseases. Aging Dis. 12:1948–1963. 2021.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Madeo F, Eisenberg T, Pietrocola F and
Kroemer G: Spermidine in health and disease. Science.
359(eaan2788)2018.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Liu R, Li X, Ma H, Yang Q, Shang Q, Song
L, Zheng Z, Zhang S, Pan Y, Huang P, et al: Spermidine endows
macrophages anti-inflammatory properties by inducing mitochondrial
superoxide-dependent AMPK activation, Hif-1α upregulation and
autophagy. Free Radic Biol Med. 161:339–350. 2020.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Jeong JW, Cha HJ, Han MH, Hwang SJ, Lee
DS, Yoo JS, Choi IW, Kim S, Kim HS, Kim GY, et al: Spermidine
protects against oxidative stress in inflammation models using
macrophages and Zebrafish. Biomol Ther (Seoul). 26:146–156.
2018.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Yuan H, Wu SX, Zhou YF and Peng F:
Spermidine inhibits joints inflammation and macrophage activation
in mice with collagen-induced arthritis. J Inflamm Res.
14:2713–2721. 2021.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Mathiesen CBK, Rudjord-Levann AM, Gad M,
Larsen J, Sellebjerg F and Pedersen AE: Cladribine inhibits
secretion of pro-inflammatory cytokines and phagocytosis in human
monocyte-derived M1 macrophages in-vitro. Int Immunopharmacol.
91(107270)2021.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Bian H, Li F, Wang W, Zhao Q, Gao S, Ma J,
Li X, Ren W, Qin C and Qi J: MAPK/p38 regulation of cytoskeleton
rearrangement accelerates induction of macrophage activation by
TLR4, but not TLR3. Int J Mol Med. 40:1495–1503. 2017.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Yang X, Chang Y and Wei W: Emerging role
of targeting macrophages in rheumatoid arthritis: Focus on
polarization, metabolism and apoptosis. Cell Prolif.
53(e12854)2020.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Castejón ML, Alarcón-de-la-Lastra C,
Rosillo MÁ, Montoya T, Fernández-Bolaños JG, González-Benjumea A
and Sánchez-Hidalgo M: A new peracetylated oleuropein derivative
ameliorates joint inflammation and destruction in a murine
collagen-induced arthritis model via activation of the Nrf-2/Ho-1
antioxidant pathway and suppression of MAPKs and NF-κB
activation/Ho-1 antioxidant pathway and suppression of MAPKs and
NF-κB activation. Nutrients. 13(311)2021.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Zhao JM, Chen X, Cheng K, Shi Q and Peng
K: Anserine and glucosamine supplementation attenuates the levels
of inflammatory markers in rats with rheumatoid arthritis. AMB
Express. 10(57)2020.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Tanaka Y: Rheumatoid arthritis. Inflamm
Regen. 40(20)2020.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Li J, Hsu HC and Mountz JD: Managing
macrophages in rheumatoid arthritis by reform or removal. Curr
Rheumatol Rep. 14:445–454. 2012.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Tsubaki T, Arita N, Kawakami T,
Shiratsuchi T, Yamamoto H, Takubo N, Yamada K, Nakata S, Yamamoto S
and Nose M: Characterization of histopathology and gene-expression
profiles of synovitis in early rheumatoid arthritis using targeted
biopsy specimens. Arthritis Res Ther. 7:R825–R836. 2005.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Tardito S, Martinelli G, Soldano S,
Paolino S, Pacini G, Patane M, Alessandri E, Smith V and Cutolo M:
Macrophage M1/M2 polarization and rheumatoid arthritis: A
systematic review. Autoimmun Rev. 18(102397)2019.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Han Y, Yuan C, Zhou X, Han Y, He Y, Ouyang
J, Zhou W, Wang Z, Wang H and Li G: Anti-inflammatory activity of
three triterpene from Hippophae rhamnoides L. in
lipopolysaccharide-stimulated RAW264.7 cells. Int J Mol Sci.
22(12009)2021.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Dorrington MG and Fraser IDC: NF-κB
signaling in macrophages: Dynamics, crosstalk, and signal
integration. Front Immunol. 10(705)2019.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Byun J, Kim SK and Ban JY:
Anti-inflammatory and anti-oxidant effects of Korean ginseng berry
extract in LPS-activated RAW264.7 macrophages. Am J Chin Med.
49:719–735. 2021.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Liu L, Guo H, Song A, Huang J, Zhang Y,
Jin S, Li S, Zhang L, Yang C and Yang P: Progranulin inhibits
LPS-induced macrophage M1 polarization via NF-кB and MAPK pathways.
BMC Immunol. 21(32)2020.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Ma L, Ni L, Yang T, Mao P, Huang X, Luo Y,
Jiang Z, Hu L, Zhao Y, Fu Z and Ni Y: Preventive and therapeutic
spermidine treatment attenuates acute colitis in mice. J Agric Food
Chem. 69:1864–1876. 2021.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Zhong Z, Sanchez-Lopez E and Karin M:
Autophagy, NLRP3 inflammasome and auto-inflammatory/immune
diseases. Clin Exp Rheumatol. 34 (4 Suppl 98):S12–S16.
2016.PubMed/NCBI
|
|
37
|
Huang P, Hong J, Mi J, Sun B, Zhang J, Li
C and Yang W: Polyphenols extracted from Enteromorpha clathrata
alleviates inflammation in lipopolysaccharide-induced RAW 264.7
cells by inhibiting the MAPKs/NF-κB signaling pathways. J
Ethnopharmacol. 286(114897)2022.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Barger SW, Moerman AM and Mao X: Molecular
mechanisms of cytokine-induced neuroprotection: NFkappaB and
neuroplasticity. Curr Pharm Des. 11:985–998. 2005.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Syama HP, Sithara T, Lekshmy Krishnan S
and Jayamurthy P: Syzygium cumini seed attenuates LPS induced
inflammatory response in murine macrophage cell line RAW264.7
through NF-κB translocation. J Funct Foods. 44:218–226. 2018.
|