Introduction
Ovarian cancer (OC) ranks among the most aggressive
gynecological malignancies and remains a leading contributor to
cancer-related mortality in women worldwide (1). Despite continuous refinements in
surgical techniques and the widespread application of
platinum-based chemotherapy, clinical outcomes for patients with
advanced-stage OC remain suboptimal (2). The elevated mortality rate is largely
attributable to the absence of specific early clinical
manifestations and effective screening strategies, leading to
delayed diagnosis, with many patients presenting with extensive
intraperitoneal dissemination and distant metastasis (3). Prognosis declines substantially as
the disease progresses, and tumor relapse together with the
emergence of chemoresistance further complicates therapeutic
management (4). Collectively,
these challenges underscore the urgent need for a deeper
understanding of the molecular mechanisms driving OC progression
and for the identification of reliable biomarkers and potential
therapeutic targets.
Long non-coding RNAs (lncRNAs) have been
increasingly recognized as key regulators of cellular processes,
such as proliferation, apoptosis and epithelial-mesenchymal
transition (5,6). lncRNA dysregulation has been
implicated in cancer development, including ovarian cancer (OC),
whereby a number of lncRNAs act as oncogenic drivers (7,8).
Potassium voltage-gated channel subfamily Q member 1 opposite
strand/antisense transcript 1 (KCNQ1OT1), located on chromosome
11p15.5, exerts oncogenic effects in a number of malignancies
through pathways such as PI3K/AKT, Notch signaling and glycolysis
(9,10). KCNQ1OT1 is also upregulated in OC,
enhancing proliferation and migration, yet its precise mechanism of
action remains unclear (11,12).
MicroRNAs (miRNAs) represent another critical layer
of post-transcriptional regulation. miR-140-5p has been widely
characterized as a tumor-suppressive miRNA. In OC, miR-140-5p
inhibits cell proliferation by targeting platelet-derived growth
factor receptor α, while studies in other malignancies indicate
that it suppresses invasion and angiogenesis by modulating
VEGFA-mediated signaling and Wnt/β-catenin pathways (13-15).
Downregulation of miR-140-5p has been observed in tumor tissues and
cancer cell lines, and restoration of its expression attenuates
malignant phenotypes (16).
Kallikrein-related peptidase 10 (KLK10), a secreted
serine protease of the kallikrein family, is aberrantly expressed
in several epithelial malignancies, including ovarian and
colorectal cancer (17,18). Elevated KLK10 levels have been
detected in patient tumor specimens, and dysregulated expression
has been associated with tumor progression and clinical outcomes
(17). Functional studies further
suggest that KLK10 influences tumor cell proliferation and
migration; however, its upstream regulatory mechanisms in OC remain
unclear (19).
Given the emerging lncRNA-miRNA-mRNA regulatory
paradigm and the oncogenic role of KCNQ1OT1, it remains to be
determined whether KCNQ1OT1 promotes OC progression through
modulation of miR-140-5p and downstream targets such as KLK10.
Therefore, the present study aimed to investigate the regulatory
interactions among KCNQ1OT1, miR-140-5p and KLK10 and to elucidate
their functional relevance in OC.
Materials and methods
Bioinformatics analysis
Gene expression profiles from Gene Expression
Omnibus (GEO; https://www.ncbi.nlm.nih.gov/geo/) datasets GSE66957
(57 OC vs. 12 normal samples) and GSE47841(20) (21 OC vs. 9 normal samples) were
analyzed using RStudio (version 4.4.1; Posit Software, PBC), and
differentially expressed genes were identified with cut-off levels
of log2 (fold change) ≥1 and adjusted P<0.05.
TargetScan v8.0 (https://www.targetscan.org/vert_80/) was used to
predict miR-140-5p binding to KLK10, while starBase v3.0
(https://rnasysu.com/encori/) was used to
assess the interaction of miR-140-5p with KCNQ1OT1.
Cell culture
IOSE80 (normal ovarian epithelial) and SKOV3 (OC)
cell lines were sourced from Procell Life Science & Technology
Co., Ltd. IOSE80 cells were maintained in DMEM (Gibco; Thermo
Fisher Scientific, Inc.) and SKOV3 cells in McCoy's 5A medium
(Gibco; Thermo Fisher Scientific, Inc.), both supplemented with 10%
FBS (HyClone; Cytiva) and 1% penicillin-streptomycin, at 37˚C in a
5% CO2 humidified incubator. All cell lines were
authenticated by short tandem repeat profiling and were routinely
tested for Mycoplasma contamination using a PCR-based
detection kit (cat. no. C0301S; Beyotime Biotechnology). Only
Mycoplasma-free cultures were used for subsequent
experimentation.
Cell transfection
Logarithmic-phase SKOV3 cells were seeded into
6-well plates at a density of 2x105 cells per well and
transfected (30-50% confluence) using the riboFET CP Transfection
Kit (Guangzhou RiboBio Co., Ltd.) as per the manufacturer's
instructions. Small interfering (si)-KCNQ1OT1, si-KLK10, miR-140-5p
mimic/inhibitor and negative controls (si-NC, miR-NC and
inhibitor-NC, respectively) were provided by Guangzhou Ribo Co.,
Ltd. Cells were incubated with transfection complexes at 37˚C in a
humidified atmosphere containing 5% CO2 for 24-72 h
depending on the subsequent experimental design. Transfection
efficiency was verified using reverse transcription-quantitative
PCR (RT-qPCR) 36 h after transfection. The final transfected
oligonucleotide concentration was 50 nM, with sequences as follows:
si-KCNQ1OT1 sense, 5'-GGUAGAAUAGUUCUGUCUU-3' and antisense,
5'-AAGACAGAACUAUUCUACC-3'; si-KLK10 sense,
5'-CUGGAUCAAUAAAGUCAUA-3' and antisense, 5'-UAUGACUUUAUUGAUCCAG-3';
miR-140-5p mimic sense, 5'-CAGUGGUUUUACCCUAUGGUAG-3' and antisense,
5'-CUACCAUAGGGUAAAACCACUG-3'; and miR-140-5p inhibitor,
5'-CUACCAUAGGGUAAAACCACUG-3'. According to the manufacturer's
description, miR-140-5p inhibitor is a chemically modified
antisense oligonucleotide that primarily mediates functional
inhibition by binding to mature miR-140-5p and blocking its
interaction with target mRNA. Detailed information regarding
miR-140-5p inhibitor is available on the manufacturer's website
(https://www.ribobio.com/product-and-service/mirna-function-reagent/mirna-inhibitor/).
According to the manufacturer's policy, the exact sequences of
si-NC (cat. no. siN0000001-1-5), miR-NC (cat. no. miR1N0000001-1-5)
and inhibitor-NC (cat. no. miR2N0000001-1-5) are proprietary and
therefore not disclosed. These NCs were designed by Guangzhou
RiboBio Co., Ltd. and have no marked homology to any known human
transcripts.
Western blot analysis
Proteins from SKOV3 cells were extracted using RIPA
buffer (Beyotime Biotechnology) and quantified using a BCA assay.
Equal protein amounts (30 µg/lane) were subjected to 10% SDS-PAGE
and transferred to PVDF membranes. After blocking with 5% non-fat
milk at room temperature for 3 h, membranes were incubated
overnight at 4˚C with anti-KLK10 (cat. no. Ab229968; 1:1,000;
Abcam) or anti-β-actin (cat. no. AF5003; 1:1,000; Beyotime
Biotechnology). An HRP-conjugated goat anti-rabbit IgG secondary
antibody (cat. no. A0208; 1:3,000; Beyotime Biotechnology) was then
applied for 1 h at room temperature. Protein bands were detected
using an ECL kit [Biomiky; Maike Biotechnology (Shanghai) Co.,
Ltd.] and quantified with ImageJ software (version 1.50; National
Institutes of Health).
RNA extraction and RT-qPCR
RNA from the aforementioned cell lines was extracted
with TRIzol® (Invitrogen; Thermo Fisher Scientific,
Inc.), and 2 µg RNA were reverse-transcribed using M-MLV Reverse
Transcriptase (Promega Corporation) according to the manufacturer's
instructions. The reaction mixture contained dNTPs (Thermo Fisher
Scientific, Inc.) and oligo(dT) primers for mRNA detection. For
miRNA analysis, specific stem-loop reverse transcription primers
were used. Reverse transcription was performed at 42˚C for 60 min,
followed by enzyme inactivation at 70˚C for 15 min. qPCR was
performed with the SYBR Green Master Mix (Beijing Labgic Technology
Co., Ltd.) on a CFX Connect Real-Time PCR System (Bio-Rad
Laboratories, Inc.). Cycling conditions were as follows: 95˚C for 1
min, then 40 cycles of 95˚C for 5 sec and 60˚C for 15 sec. The
expression level of miR-140-5p was normalized to U6 and that of
KCNQ1OT1 and KLK10 to GAPDH, using the 2-ΔΔCq method
(21). The following primers were
employed: KCNQ1OT1 forward, 5'-TGCAGAAGACAGGACACTGG-3' and reverse,
5'-CTTTGGTGGGAAAGGACAGA-3'; KLK10 forward,
5'-GAGTGTGAGGTCTTCTACCCTG-3' and reverse,
5'-ATGCCTTGGAGGGTCTCGTCAC-3'; GAPDH forward,
5'-TGCACCACCAACTGCTTAGC-3' and reverse,
5'-GGCATGGACTGTGGTCATGAG-3'; miR-140-5p forward,
5'-AGGCGCCAGTGGTTTTACC-3' and reverse (universal),
5'-CAGTGCAGGGTCCGAGGT-3'; and U6 forward, 5'-CTCGCTTCGGCAGCACA-3'
and reverse, 5'-AACGCTTCACGAATTTGCGT-3'.
RNA pull-down assay
The biotin-labeled miR-140-5p (Bio-miR-140-5p) and
Bio-NC probes were provided by Guangzhou RiboBio Co., Ltd. The
Bio-miR-140-5p probe corresponded to the mature miR-140-5p sequence
(sense, 5'-CAGUGGUUUUACCCUAUGGUAG-3' and antisense,
5'-CUACCAUAGGGUAAAACCACUG-3') with a 3'-biotin modification. The
Bio-NC probe was designed by the manufacturer and had no marked
homology to human transcripts. According to the manufacturer's
policy, the specific sequence of Bio-NC is proprietary and
therefore not disclosed. SKOV3 cells were transfected with 100 nM
Bio-miR-140-5p or Bio-NC using the riboFET CP Transfection Kit
(Guangzhou RiboBio Co., Ltd.) at 37˚C in a humidified incubator
containing 5% CO2. Cells were incubated with the
transfection complexes for 48 h prior to subsequent pull-down
analysis. Pull-down assays were performed using a miRNA pull-down
kit (cat. no. Bes5108; Guangzhou Bersinbio Co., Ltd.) per the
manufacturer's protocol. At 48 h post-transfection,
~1x107 cells per reaction were harvested and lysed using
the lysis buffer provided in the kit, supplemented with protease
inhibitor, RNase inhibitor and DTT. After centrifugation, 100 µl
lysate was reserved as the input control, and the remaining ~1 ml
lysate was incubated with pre-blocked streptavidin-coated magnetic
beads at 4˚C for 4 h with gentle rotation.
After washing, the beads were resuspended in 100 µl
RNA elution buffer provided in the kit and heated at 95˚C for 2
min, and then the supernatant was collected on a magnetic stand.
The elution step was repeated once (total, 200 µl). RNA was then
purified by phenol:chloroform:isoamyl alcohol extraction (25:24:1)
followed by centrifugation at 13,000 x g for 10 min at 4˚C, and the
aqueous phase was transferred to a new RNase-free tube. RNA was
precipitated with 3 M sodium acetate, glycogen and absolute ethanol
at -80˚C for 5-16 h, pelleted by centrifugation at 16,000 x g for
30 min at 4˚C, and washed with 75% ethanol (16,000 x g for 10 min
at 4˚C). The RNA pellet was air-dried for 10 min at room
temperature and dissolved in RNase-free water for subsequent
RT-qPCR analysis of KCNQ1OT1 and KLK10 enrichment.
Proliferation assay
A Cell Counting Kit-8 (CCK-8) assay (Beyotime
Biotechnology) was used to assess SKOV3 cell proliferation. At 24 h
post-transfection, cells were trypsinized and re-seeded into
96-well plates at a density of 2,000 cells/well, and incubated for
24-72 h before CCK-8 analysis. CCK-8 solution (10 µl) was added at
24, 48 and 72 h, followed by a 2 h incubation at 37˚C. Optical
density at 450 nm was then detected with the BioTek Synergy H1
microplate reader (Agilent Technologies, Inc.).
Migration assay
A wound-healing assay was performed to evaluate
migration. SKOV3 cells (2x105/well) were seeded into
6-well plates until ~80% confluence was reached, after which a
straight scratch was made with a 200-µl pipette tip. Detached cells
were removed by PBS washing and cultures were maintained in
serum-free McCoy's 5A medium for 24 h at 37˚C. Images were captured
at 0 and 24 h (magnification, x100) with an IXplore™
inverted microscope system (EVIDENT). Wound areas were quantified
using ImageJ software (version 1.50; National Institutes of
Health). Wound closure (%) was calculated using the following
formula: Wound closure (%)=[(wound area at 0 h-wound area at 24
h)/wound area at 0 h] x100.
Statistical analysis
Data are presented as the mean ± SD from at least
three independent experiments. Statistical analyses were performed
using SPSS software (version 18.0; SPSS, Inc.). Data normality was
assessed using the Shapiro-Wilk test prior to statistical analysis.
Comparisons between two groups were assessed using unpaired
Student's t-test, while differences among multiple groups were
evaluated using one-way ANOVA followed by Tukey's post hoc test.
P<0.05 was considered to indicate a statistically significant
difference.
Results
Upregulation of KCNQ1OT1 and KLK10
with concomitant downregulation of miR-140-5p is observed in
OC
To explore the transcriptional alterations of
KCNQ1OT1, miR-140-5p and KLK10 in OC and to preliminarily verify
their relevance in patient samples, two GEO datasets, GSE66957 and
GSE47841, were first analyzed. These datasets were selected based
on their complementary profiling characteristics: GSE66957 provided
mRNA and lncRNA expression data that include both KCNQ1OT1 and
KLK10, enabling evaluation of their transcript levels in OC
tissues, whereas GSE47841 contained non-coding RNA profiling data,
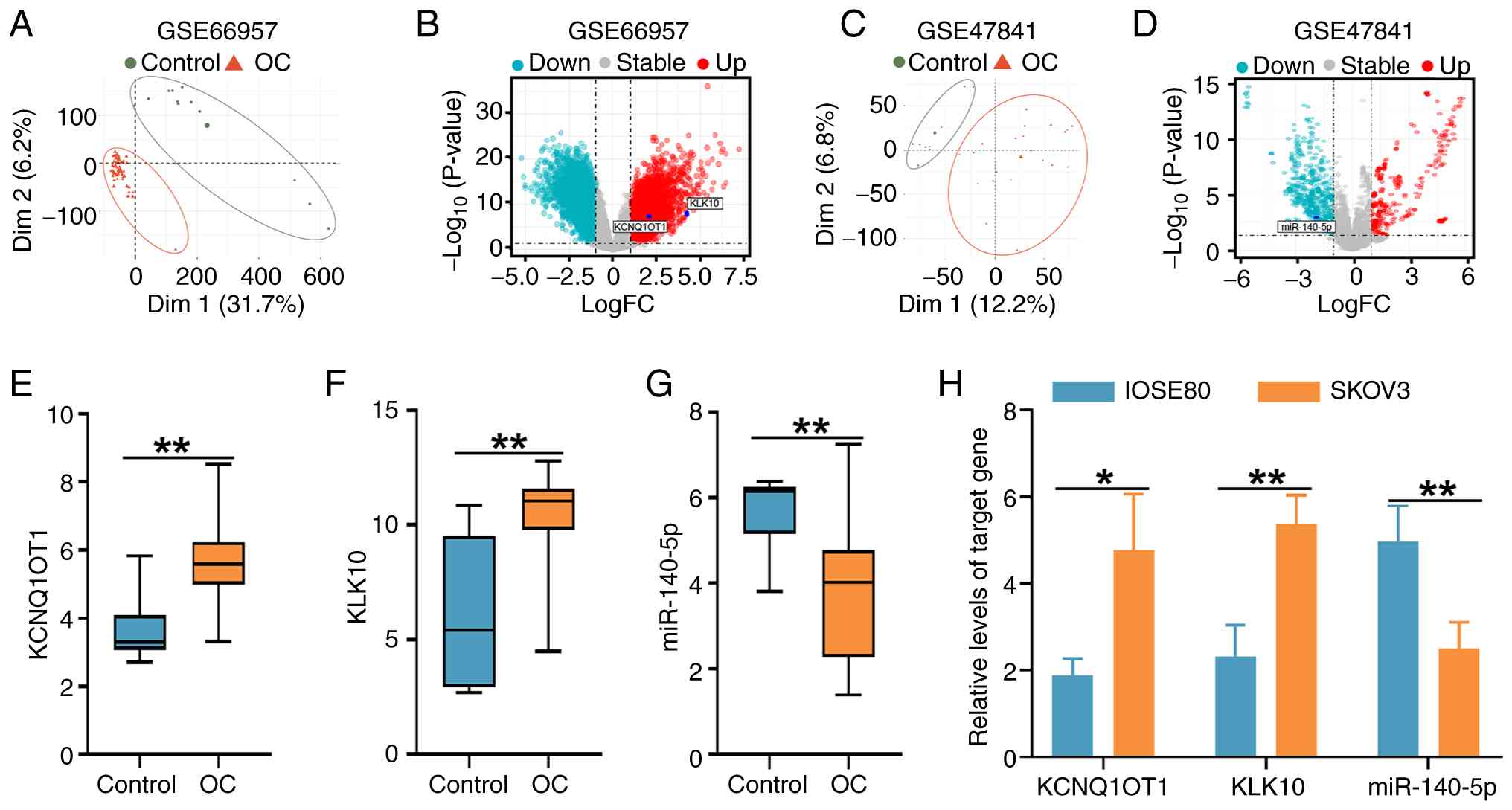
making it suitable for assessing miR-140-5p expression. Principal
component analysis of the GSE66957 and GSE47841 datasets
demonstrated clear separation between OC and control samples
(Fig. 1A and C). Differential expression analysis
further identified KCNQ1OT1, KLK10 and miR-140-5p as significantly
dysregulated genes, as shown in the corresponding volcano plots
(Fig. 1B and D). Consistently, expression profiling
revealed marked upregulation of KCNQ1OT1 and KLK10 in OC samples
from the GSE66957 dataset, whereas miR-140-5p was significantly
downregulated in the GSE47841 dataset compared with the control
group (Fig. 1E-G).
The expression levels of KCNQ1OT1, KLK10 and
miR-140-5p were examined using RT-qPCR in IOSE80 and SKOV3 cells.
Consistent with the aforementioned in silico predictions,
the expression levels of KCNQ1OT1 and KLK10 were significantly
elevated, while those of miR-140-5p were significantly reduced in
SKOV3 cells compared with those in IOSE80 cells (Fig. 1H), suggesting the involvement of
these genes in OC progression.
miR-140-5p directly binds to KCNQ1OT1
lncRNA and KLK10 mRNA
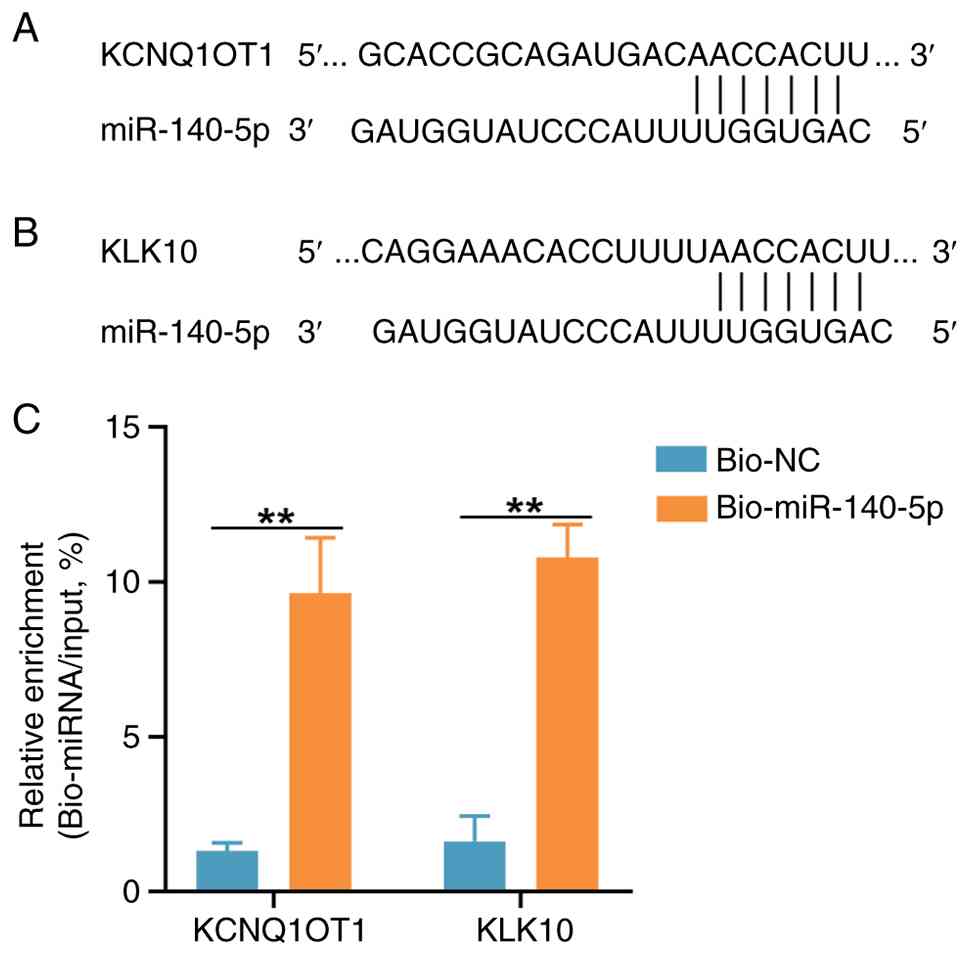
The starBase and TargetScan databases were used to
predict the potential binding of miR-140-5p to KCNQ1OT1 and KLK10,
respectively. Complementary sequences were found between miR-140-5p
and KCNQ1OT1 lncRNA and between miR-140-5p and the 3'-untranslated
region of KLK10 mRNA (Fig. 2A and
B). A miRNA pull-down assay
further determined that both KCNQ1OT1 lncRNA and KLK10 mRNA were
enriched after pull-down with the miR-140-5p probe in SKOV3 cells.
RT-qPCR was performed to quantify the enriched RNA levels and
subsequent calculation of pull-down/input ratios suggested a direct
binding of miR-140-5p to KCNQ1OT1 and KLK10 (Fig. 2C). These results suggest that
miR-140-5p, KCNQ1OT1 and KLK10 may functionally interact to promote
OC progression.
miR-140-5p suppresses OC cell
proliferation and migration by regulating KLK10
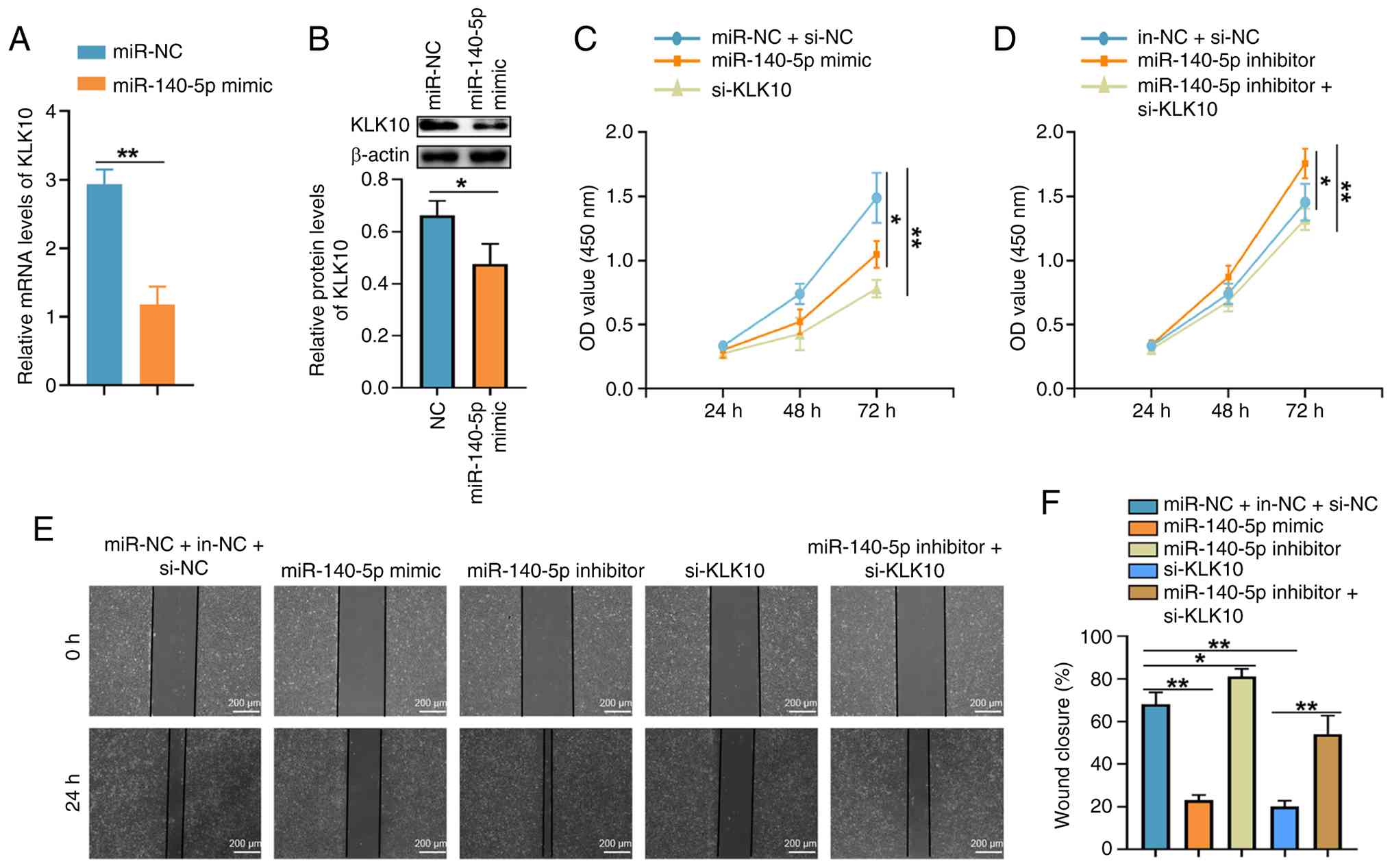
The association between miR-140-5p and KLK10 was
examined by transfecting SKOV3 cells with miR-140-5p mimic,
miR-140-5p inhibitor and/or si-KLK10, and the transfection
efficiency of each molecule was verified via RT-qPCR compared with
the corresponding NC (Fig.
S1A-C). As miR-140-5p inhibitor primarily blocks miR-140-5p
activity rather than altering its expression level, its inhibitory
efficiency was demonstrated by RT-qPCR analysis of the downstream
target gene KLK10, which exhibited a significant increase upon
miR-140-5p inhibition (Fig. S1B).
Transfection with miR-140-5p mimic resulted in a significant
decrease in KLK10 expression at both the mRNA and protein levels
compared with that in the control group (Fig. 3A and B). Furthermore, transfection with either
miR-140-5p mimic or si-KLK10 significantly reduced SKOV3 cell
proliferation and migration (Fig.
3C, E and F). Conversely, miR-140-5p inhibition
promoted proliferation and migration of SKOV3 cells, whereas
co-transfection with si-KLK10 reversed these effects (Fig. 3D-F). Overall, these data suggest
that miR-140-5p inhibits SKOV3 OC cell proliferation and migration
by targeting KLK10.
KCNQ1OT1 silencing inhibits
proliferation and migration of OC cells through miR-140-5p
regulation
To assess the involvement of KCNQ1OT1 in OC,
siRNA-mediated silencing was performed in SKOV3 cells. Knockdown of
KCNQ1OT1 led to a significant increase in miR-140-5p expression
levels but a significant decrease in KLK10 mRNA levels (Fig. 4A). Similarly, KLK10 protein levels
were reduced following KCNQ1OT1 depletion, yet this effect was
reversed by co-transfection with miR-140-5p inhibitor (Fig. 4B). Functionally, KCNQ1OT1 silencing
suppressed proliferation and migration of SKOV3 cells, whereas
inhibition of miR-140-5p counteracted these effects (Fig. 4C-E). Together, these observations
underscore the role of KCNQ1OT1 in promoting OC progression by
suppressing miR-140-5p.
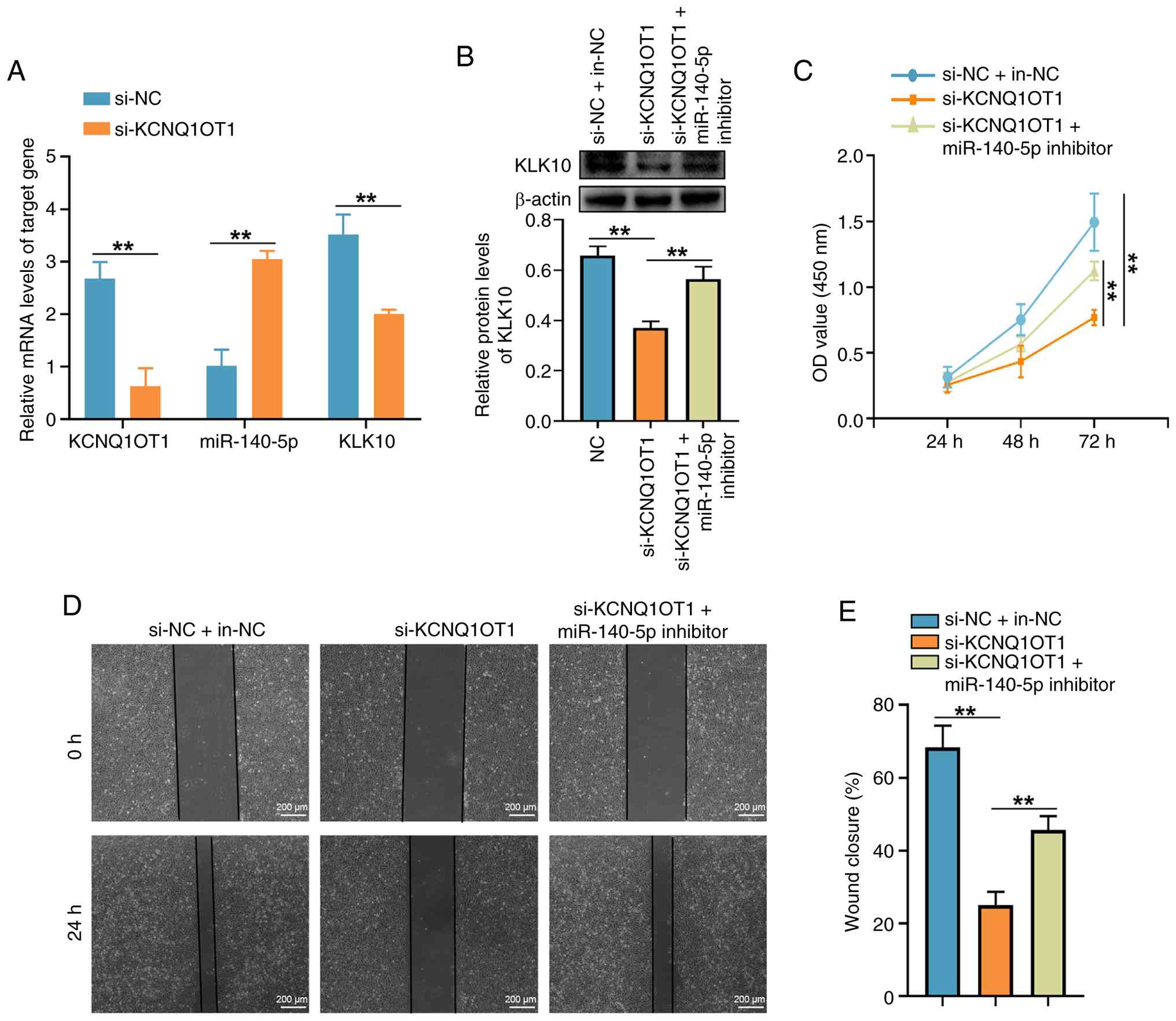 | Figure 4KCNQ1OT1 knockdown inhibits OC cell
proliferation and migration by upregulating miR-140-5p. (A)
Transfection with si-KCNQ1OT1 elevated miR-140-5p expression levels
while reducing KLK10 mRNA levels in OC cells. (B) si-KCNQ1OT1
decreased KLK10 protein expression and this effect was
significantly reversed by co-transfection with miR-140-5p
inhibitor. (C) si-KCNQ1OT1 suppressed cell proliferation, whereas
co-transfection with miR-140-5p inhibitor ameliorated this effect.
(D) Wound-healing assay and (E) quantification showing that
si-KCNQ1OT1 inhibited cell migration with co-transfection with
miR-140-5p inhibitor counteracting this effect. Scale bar, 200 µm.
**P<0.01. miR, microRNA; OC, ovarian cancer; si,
small interfering; KCNQ1OT1, potassium voltage-gated channel
subfamily Q member 1 opposite strand/antisense transcript 1; KLK10,
kallikrein-related peptidase 10; NC, negative control; in,
inhibitor; OD, optical density. |
Discussion
OC is one of the most harmful gynecological
malignancies, characterized by complex etiology and poor prognosis,
with contributions from inherited susceptibility (e.g., BRCA1/2
alterations), marked histological and molecular heterogeneity, and
a tumor microenvironment that facilitates early intraperitoneal
dissemination and treatment resistance (22-25).
Owing to non-specific clinical manifestations, such as abdominal
bloating, pelvic or abdominal pain, early satiety and urinary
urgency or frequency, together with the lack of reliable early
diagnostic biomarkers, OC is frequently diagnosed at advanced
stages, where the 5-year survival rate remains as low as ~30%
(26-28).
Standard treatments typically involve cytoreductive surgery
combined with chemotherapy, yet therapeutic efficacy is often
hindered by chemoresistance, immune evasion, metastasis and
recurrence (29,30). Thus, unraveling the molecular
events that govern OC progression is important in improving early
diagnosis and therapeutic strategies.
lncRNAs have emerged as important regulators of
tumor biology, influencing proliferation, migration, invasion and
apoptosis through diverse mechanisms, including acting as competing
endogenous RNAs to sponge microRNAs, modulating transcriptional
activity, interacting with epigenetic modifiers, and regulating key
oncogenic signaling pathways such as PI3K/AKT, Wnt/β-catenin and
Notch (31-34).
Among them, KCNQ1OT1 has been reported to be upregulated in
hepatocellular, colorectal, gastric and breast cancer, as well as
non-small cell lung cancer and OC (11,34).
Consistent with prior evidence, the present study revealed KCNQ1OT1
to be upregulated in OC cells, while silencing KCNQ1OT1
significantly reduced cell proliferation and migration, indicative
of its oncogenic function.
A number of studies have previously investigated the
role of KCNQ1OT1 in OC and reported its oncogenic functions through
distinct molecular mechanisms. For example, He et al
(10) demonstrated that KCNQ1OT1
promotes OC metastasis by increasing the methylation level of the
EIF2B5 promoter, thereby suppressing EIF2B5 transcription and
reducing its expression, highlighting an epigenetic regulatory
mechanism. In another study, Chen et al (11) reported that KCNQ1OT1 accelerates OC
progression through the miR-125b-5p/CD147 axis, supporting a
competing endogenous RNA-mediated mode of action. Although these
studies consistently indicated that KCNQ1OT1 enhances malignant
phenotypes in OC such as proliferation and migration, the
downstream targets and regulatory pathways proposed differ among
studies, underscoring the complex and multifaceted role of KCNQ1OT1
in OC progression.
In comparison with these previously reported
mechanisms, the present study identified a distinct
KCNQ1OT1/miR-140-5p/KLK10 regulatory axis. While the
pro-tumorigenic effects of KCNQ1OT1 observed in the present study
are consistent with previous reports, the involvement of miR-140-5p
and KLK10 expands the mechanistic landscape of KCNQ1OT1 in OC and
suggests that this lncRNA may exert its oncogenic functions in a
context-dependent manner, influenced by factors such as
tissue-specific gene expression patterns, availability of
interacting microRNAs, epigenetic status and the molecular subtype
of the tumor.
As key regulators of gene expression, miRNAs
function either as oncogenes or tumor suppressors (35,36).
miR-140-5p has been reported as a tumor suppressor in OC and
non-small cell lung cancer, where it modulates cell proliferation,
invasion and migration (13,37).
The present findings demonstrated that KCNQ1OT1 directly interacted
with miR-140-5p and suppressed its expression. In addition,
inhibition of miR-140-5p negated the inhibitory effects of KCNQ1OT1
knockdown on OC cell proliferation and migration, establishing
miR-140-5p as a pivotal downstream target of KCNQ1OT1. Although
silencing KCNQ1OT1 significantly elevated miR-140-5p levels, the
precise regulatory mechanism remains unclear. Previous evidence has
indicated that a number of lncRNAs can induce miRNA decay through
target-directed miRNA degradation (TDMD), driven by highly
complementary lncRNA-miRNA pairing (38,39).
Due to the complementarity between KCNQ1OT1 and miR-140-5p,
KCNQ1OT1 may potentially modulate miR-140-5p through a TDMD-like
mechanism. Further experiments assessing miRNA stability or
argonaute RNA-induced silencing complex catalytic component
2-dependent decay are needed to further investigate this
possibility.
KLK10 is an additional cancer-associated molecule
with reported involvement in tumor growth, invasion and apoptosis
in ovarian and colorectal cancer (17,18).
In the present study, miR-140-5p was demonstrated to directly bind
to KLK10 and downregulate its expression. Thus, KLK10 may serve as
a functional downstream target of the KCNQ1OT1/miR-140-5p pathway,
whose dysregulation may promote OC development.
In conclusion, the present findings showed that
KCNQ1OT1 drives OC cell proliferation and migration through the
miR-140-5p/KLK10 axis. This regulatory pathway may serve as a
promising target for future diagnostic and therapeutic
interventions. However, the present study was limited by the
absence of patient tissue samples and the lack of in vivo
experiments. Future investigations using clinical specimens and
animal models are therefore required to further validate these
findings and to clarify the clinical relevance of the
KCNQ1OT1/miR-140-5p/KLK10 axis in OC.
Supplementary Material
Validation of oligonucleotide
transfection efficiency using reverse transcription-quantitative
PCR. (A) miR-140-5p expression levels in OC cells transfected with
miR-140-5p mimic. (B) miR-140-5p and KLK10 expression levels in OC
cells transfected with miR-140-5p inhibitor. (C) KLK10 expression
levels in OC cells transfected with si-KLK10.
**P<0.01. ns, not significant; miR, microRNA; OC,
ovarian cancer; si, small interfering; KLK10, kallikrein-related
peptidase 10; NC, negative control; in, inhibitor.
Acknowledgements
Not applicable.
Funding
Funding: The present study was supported by The Key Scientific
Research Foundation of Wannan Medical College (grant nos.
WK2024ZZD03 and WK2024ZZD09), The Student Research Funding Project
of Wannan Medical College (grant no. WK2024XS07) and The National
College Student Innovation Training Program (grant no.
202210368022).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
ShaT, ZT and RD conceived and designed the present
study. RD, ZT, ShuT, AY, XY, CL, XZ, KZ and ZL performed the
experiments and collected the data. ShaT wrote the manuscript. RD
and ZT performed the statistical analysis. ShaT and RD confirm the
authenticity of all the raw data. All authors read and approved the
final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Hu Z, Cai M, Zhang Y, Tao L and Guo R:
miR-29c-3p inhibits autophagy and cisplatin resistance in ovarian
cancer by regulating FOXP1/ATG14 pathway. Cell Cycle. 19:193–206.
2020.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Xiong T, Wang Y, Zhang Y, Yua J, Zhu C and
Jiang W: lncRNA AC005224.4/miR-140-3p/SNAI2 regulating axis
facilitates the invasion and metastasis of ovarian cancer through
epithelial-mesenchymal transition. Chin Med J (Engl).
136:1098–1110. 2023.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Li F, Zhao C, Diao Y, Wang Z, Peng J, Yang
N, Qiu C, Kong B and Li Y: MEX3A promotes the malignant progression
of ovarian cancer by regulating intron retention in TIMELESS. Cell
Death Dis. 13(553)2022.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Gaba F, Blyuss O, Chandrasekaran D,
Bizzarri N, Refky B, Barton D, Ind T, Nobbenhuis M, Butler J, Heath
O, et al: Prognosis following surgery for recurrent ovarian cancer
and diagnostic criteria predictive of cytoreduction success: A
Systematic review and meta-analysis. Diagnostics (Basel).
13(3484)2023.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Cheng JT, Wang L, Wang H, Tang FR, Cai WQ,
Sethi G, Xin HW and Ma Z: Insights into biological role of LncRNAs
in epithelial-mesenchymal transition. Cells. 8(1178)2019.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Kuang D, Zhang X, Hua S, Dong W and Li W:
Long non-coding RNA TUG1 regulates ovarian cancer proliferation and
metastasis via affecting epithelial-mesenchymal transition. Exp Mol
Pathol. 101:267–273. 2016.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Salamini-Montemurri M, Lamas-Maceiras M,
Lorenzo-Catoira L, Vizoso-Vázquez Á, Barreiro-Alonso A,
Rodríguez-Belmonte E, Quindós-Varela M and Cerdán ME:
Identification of lncRNAs deregulated in epithelial ovarian cancer
based on a gene expression profiling meta-analysis. Int J Mol Sci.
24(10798)2023.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Zhu Y, Zhao Y, Dong S, Liu L, Tai L and Xu
Y: Systematic identification of dysregulated lncRNAs associated
with platinum-based chemotherapy response across 11 cancer types.
Genomics. 112:1214–1222. 2020.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Zheng ZH, You HY, Feng YJ and Zhang ZT:
LncRNA KCNQ1OT1 is a key factor in the reversal effect of curcumin
on cisplatin resistance in the colorectal cancer cells. Mol Cell
Biochem. 476:2575–2585. 2021.PubMed/NCBI View Article : Google Scholar
|
|
10
|
He SL, Chen YL, Chen QH, Tian Q and Yi SJ:
LncRNA KCNQ1OT1 promotes the metastasis of ovarian cancer by
increasing the methylation of EIF2B5 promoter. Mol Med.
28(112)2022.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Chen P, Sun LS, Shen HM and QU B: LncRNA
KCNQ1OT1 accelerates ovarian cancer progression via
miR-125b-5p/CD147 axis. Pathol Res Pract.
239(154135)2022.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Liu Y, Wang Y, Yao D and Cui D: LncSOX4
serves an oncogenic role in the tumorigenesis of epithelial ovarian
cancer by promoting cell proliferation and inhibiting apoptosis.
Mol Med Rep. 17:8282–8288. 2018.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Lan H, Chen W, He G and Yang S: miR-140-5p
inhibits ovarian cancer growth partially by repression of PDGFRA.
Biomed Pharmacother. 75:117–122. 2015.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Lu Y, Qin T, Li J, Wang L, Zhang Q, Jiang
Z and Mao J: MicroRNA-140-5p inhibits invasion and angiogenesis
through targeting VEGF-A in breast cancer. Cancer Gene Ther.
24:386–392. 2017.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Cha Y, He Y, Ouyang K, Xiong H, Li J and
Yuan X: MicroRNA-140-5p suppresses cell proliferation and invasion
in gastric cancer by targeting WNT1 in the WNT/β-catenin signaling
pathway. Oncol Lett. 16:6369–6376. 2018.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Wu Y, Li J, Chen S and Yu Z: The effects
of miR-140-5p on the biological characteristics of ovarian cancer
cells through the Wnt signaling pathway. Adv Clin Exp Med.
29:777–784. 2020.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Batra J, Tan OL, O'Mara T, Zammit R, Nagle
CM, Clements JA, Kedda MA and Spurdle AB: Kallikrein-related
peptidase 10 (KLK10) expression and single nucleotide polymorphisms
in ovarian cancer survival. Int J Gynecol Cancer. 20:529–236.
2010.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Alexopoulou DK, Papadopoulos IN and
Scorilas A: Clinical significance of kallikrein-related peptidase
(KLK10) mRNA expression in colorectal cancer. Clin Biochem.
46:1453–1461. 2013.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Kato K, Noda T, Kobayashi S, Sasaki K,
Iwagami Y, Yamada D, Tomimaru Y, Takahashi H, Uemura M, Asaoka T,
et al: KLK10 derived from tumor endothelial cells accelerates colon
cancer cell proliferation and hematogenous liver metastasis
formation. Cancer Sci. 115:1520–1535. 2024.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Vilming Elgaaen B, Olstad OK, Haug KB,
Brusletto B, Sandvik L, Staff AC, Gautvik KM and Davidson B: Global
miRNA expression analysis of serous and clear cell ovarian
carcinomas identifies differentially expressed miRNAs including
miR-200c-3p as a prognostic marker. BMC Cancer.
14(80)2014.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Rakina M, Kazakova A, Villert A, Kolomiets
L and Larionova I: Spheroid formation and peritoneal metastasis in
ovarian cancer: The role of stromal and immune components. Int J
Mol Sci. 23(6215)2022.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Chang L, Ni J, Zhu Y, Pang B, Graham P,
Zhang H and Li Y: Liquid biopsy in ovarian cancer: Recent advances
in circulating extracellular vesicle detection for early diagnosis
and monitoring progression. Theranostics. 9:4130–4140.
2019.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Cheng Z, Chen Y and Huang H:
Identification and validation of a novel prognostic signature based
on ferroptosis-related genes in ovarian cancer. Vaccines (Basel).
11(205)2023.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Xu X, Zhuang X, Yu H, Li P, Li X, Lin H,
Teoh JP, Chen Y, Yang Y, Cheng Y, et al: FSH induces EMT in ovarian
cancer via ALKBH5-regulated Snail m6A demethylation. Theranostics.
14:2151–2166. 2024.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Ojasalu K, Lieber S, Sokol AM, Nist A,
Stiewe T, Bullwinkel I, Finkernagel F, Reinartz S,
Müller-Brüsselbach S, Grosse R, et al: The lysophosphatidic
acid-regulated signal transduction network in ovarian cancer cells
and its role in actomyosin dynamics, cell migration and entosis.
Theranostics. 13:1921–1948. 2023.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Žilovič D, Vaicekauskaitė I, Čiurlienė R,
Sabaliauskaitė R and Jarmalaitė S: Uterine cavity lavage mutation
analysis in lithuanian ovarian cancer patients. Cancers (Basel).
15(868)2023.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Fan W, Xiong R, Zhou Z, Zhang C, Han Y,
Shi T, Qiu J and Zhang R: ZFP57 promotes ovarian cancer progression
by transcriptionally regulating BRCA1 and managing G1 checkpoint. J
Cancer. 14:2039–2050. 2023.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Zhang Y and Pei L: Machine learning
constructs a T cell-related signature for predicting prognosis and
drug sensitivity in ovarian cancer. Aging (Albany NY).
16:3332–3349. 2024.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Chudecka-Głaz A, Strojna A, Michalczyk K,
Wieder-Huszla S, Safranow K, Skwirczyńska E and Jurczak A:
Evaluation of He4 Use in the diagnosis of ovarian cancer: First and
second recurrence, and an analysis of HE4 concentration during
second- and third-line chemotherapy. Diagnostics (Basel).
13(452)2023.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Luo M, Lei R, Zhao Q, Shen Y, He Z and Xu
J: LINC00662 promotes melanoma progression by competitively binding
miR-107 and activating the β-catenin signaling pathway. Int J Med
Sci. 21:265–276. 2024.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Yang YY, Deng YX, Yao XT, Luo HH, He WG,
Cao XL, Chen RC, He BC, Jiang HT and Wang J: lncRNA MEG3 promotes
PDK4/GSK-3β/β-catenin axis in MEFs by targeting miR-532-5p. Oxid
Med Cell Longev. 2023(3563663)2023.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Zeng Z, Wang J, Xu F, Hu P, Hu Y, Zhuo W,
Chen D, Han S, Wang F, Zhao Y, et al: The m6A modification-mediated
positive feedback between glycolytic lncRNA SLC2A1-DT and c-Myc
promotes tumorigenesis of hepatocellular carcinoma. Int J Biol Sci.
20:1744–1762. 2024.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Zhan K, Pan H, Zhou Z, Tang W, Ye Z, Huang
S and Luo L: Biological role of long non-coding RNA KCNQ1OT1 in
cancer progression. Biomed Pharmacother. 169(115876)2023.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Yang S, Luo J, Zhang L, Feng L, He Y, Gao
X, Xie S, Gao M, Luo D, Chang K and Chen M: A smart
nano-theranostic platform based on dual-microRNAs guided
self-feedback tetrahedral entropy-driven DNA circuit. Adv Sci
(Weinh). 10(e2301814)2023.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Ye Q, Raese R, Luo D, Cao S, Wan YW, Qian
Y and Guo NL: MicroRNA, mRNA, and proteomics biomarkers and
therapeutic targets for improving lung cancer treatment outcomes.
Cancers (Basel). 15(2294)2023.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Yang P, Xiong J, Zuo L, Liu K and Zhang H:
miR-140-5p regulates cell migration and invasion of non-small cell
lung cancer cells through targeting VEGFA. Mol Med Rep.
18:2866–2872. 2018.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Ameres SL, Horwich MD, Hung JH, Xu J,
Ghildiyal M, Weng Z and Zamore PD: Target RNA-directed trimming and
tailing of small silencing RNAs. Science. 328:1534–1539.
2010.PubMed/NCBI View Article : Google Scholar
|
|
39
|
de la Mata M, Gaidatzis D, Vitanescu M,
Stadler MB, Wentzel C, Scheiffele P, Filipowicz W and Großhans H:
Potent degradation of neuronal miRNAs induced by highly
complementary targets. EMBO Rep. 16:500–511. 2015.PubMed/NCBI View Article : Google Scholar
|