1. Introduction
The Fc fragment of IgE receptor 1G (FCER1G)
gene, mapping to chromosome 1q23.3, encodes the immunoglobulin Fc
receptor common γ chain (FcRγ), an integral signaling adaptor in
innate and adaptive immunity (1).
Initially identified as the third subunit of the high-affinity IgE
receptor complex (also known as FcεRIγ), FcRγ is currently
recognized as a universal submit for multiple immunoreceptors. It
severs as an essential signaling partner for Fc fragment of IgG
receptor (FcγRI) (CD64), FcγRIII (CD16) and FcαRI (CD89), as well
as for partner recognition receptors, including Dectin-1, Dectin-2
and the natural killer (NK) cell receptor NKp46 (2,3). Via
its highly conserved intracellular immunoreceptor tyrosine-based
activation motif (ITAM), FcRγ orchestrates canonical immune
processes, including antibody-dependent cellular cytotoxicity
(ADCC), antibody-dependent phagocytosis (ADCP), antifungal defense
and inflammatory modulation (4,5).
Accumulating evidence has established FCER1G
as a central regulator linking immune signaling to disease
pathogenesis. In inflammatory diseases, its expression is tightly
controlled by epigenetic mechanisms. Functionally, the protein
displays distinct context-dependent effects: It can exacerbate
vascular inflammation by polarizing immune cells toward a pro
inflammatory phenotype, while it also regulates inflammatory
intensity and promotes pathogen clearance during infection
(6). This bidirectional regulation
parallels mechanisms underlying tumor immune evasion, offering
clues to its role in the tumor microenvironment (TME).
In oncology, FCER1G exhibits remarkable
tissue-specific heterogeneity. Pan-cancer analyses show that its
expression is elevated across numerous malignancies, primarily
within monocyte and macrophage populations in the TME, and
correlates strongly with immune checkpoint genes (1). In solid tumors such as renal cell
carcinoma and gastric cancer, high FCER1G expression
predicts poor prognosis by fostering an immunosuppressive
microenvironment (7,8). Conversely, in hematologic
malignancies such as multiple myeloma (MM) and in endometrial
carcinoma, it may play a protective role (9,10).
Mechanistically, FCER1G drives tumor progression by
modulating immune suppression, promoting angiogenesis and
regulating metastasis-related factors. It also influences the
efficacy of antibody-based therapies through its role in
antibody-dependent ADCC and ADCP.
Although no current drugs directly target
FCER1G, several approved agents, such as kinase inhibitors,
can indirectly modulate its activity by influencing upstream or
downstream signaling nodes or altered ligand-receptor interactions,
providing potential therapeutic opportunities (11). In summary, elucidating the
mechanisms by which FCER1G governs inflammatory and
oncogenic processes will deepen the current understanding of immune
adaptor signaling in disease, and support the development of
FCER1G-based biomarkers and targeted immunotherapies.
2. Molecular structure of FCER1G
The FcRγ protein encoded by FCER1G is a core
component of the FcεRI complex (Fig.
1A). FcεRI exists as either a tetramer (αβγ2) or a
trimer (αγ2). The FcεRIα chain contains two
extracellular Ig-like domains that bind individual IgE molecules,
thereby mediating IgE recognition (2). The FcεRIβ chain facilitates the
maturation and trafficking of FcεRIα, and stabilizes the FcεRI
complex on the cell surface (12).
FcRγ is a small protein with a molecular weight of 15-20 kDa. Its
extracellular domain is remarkably short, contains a conserved
cysteine residue (Cys25) and forms homodimers via disulfide bonds,
thereby constituting the essential signaling subunit of FcεRI
(Fig. 1B) (2,13).
In addition to FcεRI, FcRγ also serves as an
indispensable signaling adaptor for various Ig Fc receptors. These
include the high-affinity IgG receptor FcγRI, involved in ADCP and
ADCC, the low-affinity IgG receptor FcγRIII, which mediates ADCC in
NK cells, and the IgA receptor FcαRI, a key regulator of neutrophil
inflammatory responses. FcRγ further associates with pattern
recognition receptors (PRRs) such as Dectin-1 and Dectin-2, as well
as with the NK cell-activating receptor NKp46, playing a vital role
in innate immune recognition.
3. Immunological function and signaling
mechanisms of FCER1G
FcRγ is a crucial signaling adaptor protein within
the immune system. It lacks intrinsic ligand-binding capacity, with
its function entirely dependent upon a highly conserved ITAM within
its intracellular domain. As a universal signaling subunit for
multiple Ig Fc receptors (such as FcεRI and FcγR) and PRRs [such as
C-type lectin receptors (CLRs)], FcRγ initiates downstream
signaling cascades via its ITAM, thereby participating in
regulating core immune processes, including allergic reactions,
anti-infective immunity, and ADCC. Crucially, the biological
effects of FcRγ are not uniform but exhibit distinct cell-type
specificity. This specificity arises from its association with
different receptors and the inherent differences in cellular
signaling networks, enabling the same FcRγ-ITAM module to be
programmed in diverse immune cells to execute differentiated
functional programs.
Signal transduction and functional
programming of FcRγ in different immune cells. Mast cells and
basophils: FcεRI-mediated immediate hypersensitivity reactions
On the surfaces of mast cells and basophils, FcRγ
constitutes the core component of the high-affinity IgE receptor
FcεRI, forming a tetrameric (αβγ2) complex (14). Upon cross-linking of the receptor
by an allergen-IgE complex, the ITAM of FcRγ rapidly recruits
spleen tyrosine kinase (SYK), subsequently phosphorylating it.
SYK-activated phospholipase Cγ (PLCγ) hydrolyses
phosphatidylinositol bisphosphate to generate inositol
trisphosphate (IP3) and diacylglycerol (14,15).
IP3 subsequently induces intracellular calcium
mobilization and extracellular calcium influx, triggering the rapid
degranulation of stored mediators such as histamine and tryptamine,
which are hallmarks of immediate-type allergic reactions (Fig. 2) (14,15).
Knockout of the FCER1G gene prevents IgE-induced mast cell
activation and allergic responses in mice (16). This research indicated that FcRγ
protein stability is critical for its signaling function (16). For instance, the deubiquitinating
enzyme USP5 stabilizes FcRγ by specifically removing K48-linked
ubiquitin chains, thereby significantly enhancing IgE-induced mast
cell activation and allergic inflammation, while inhibiting USP5
attenuates this process (17).
This reveals that targeting FcRγ protein stability represents a
novel pathway for regulating allergic responses.
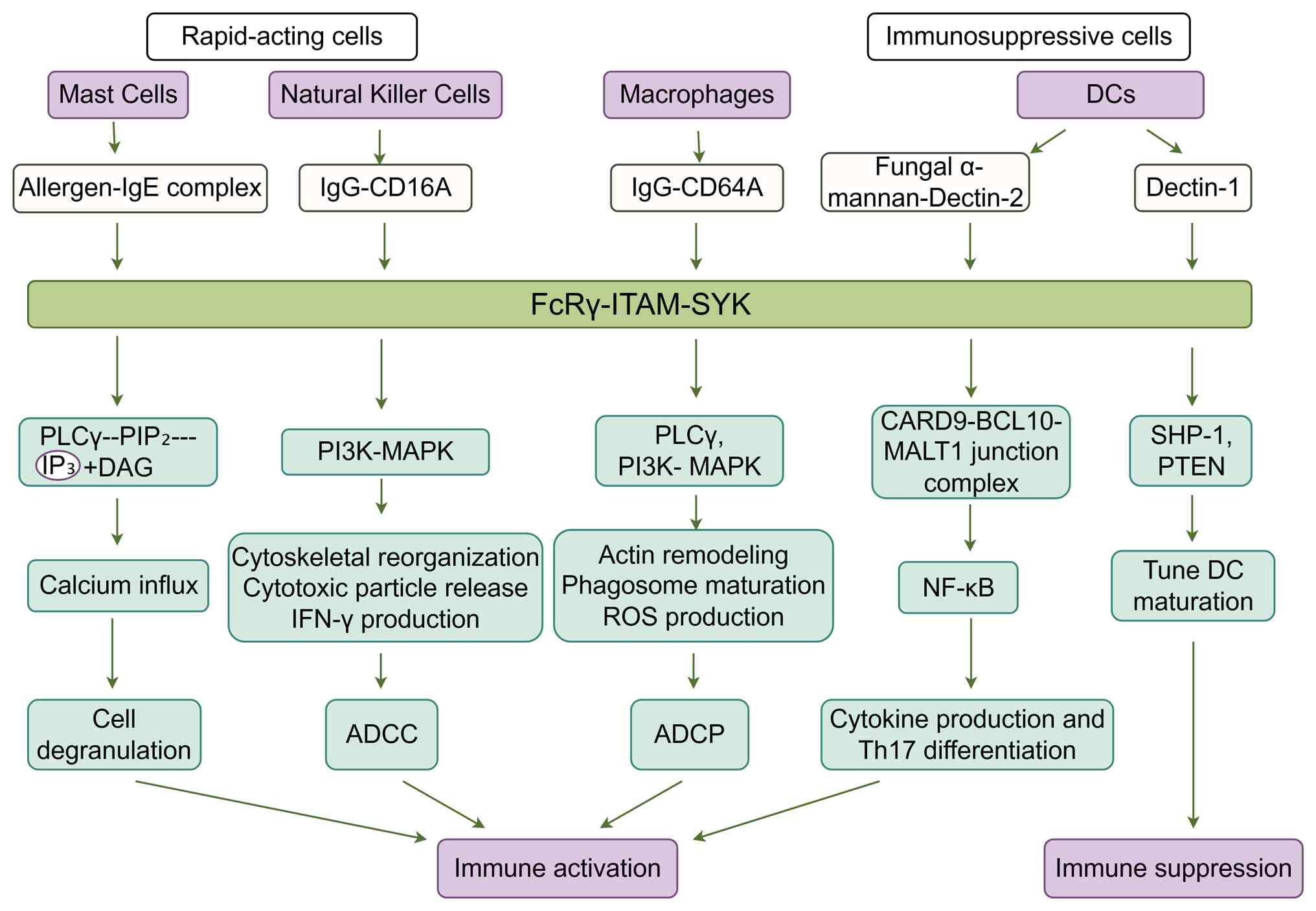 | Figure 2Divergent intracellular signaling and
functional mechanism mediated by FcRγ in different innate immune
cells. Upon engagement of distinct ligand-receptor complexes
(allergen-IgE complex, fungal α-mannan-Dectin-2, IgG-CD16A,
IgG-CD64A), signaling is initiated via the associated FcRγ subunit,
and phosphorylation of its ITAM recruits and activates SYK. This
common initiating module then diverges into cell-type-specific
downstream pathways, leading to distinct functional outputs in mast
cells, natural killer cells, DCs and macrophages. The figure was
drawn with FigDraw2.0 (supplied by Hangzhou Sifei Technology Co.,
Ltd.). FcR, Fc receptor; ITAM, immunoreceptor tyrosine-based
activation motif; SYK, spleen tyrosine kinase; PLCγ, phospholipase
Cγ; PIP2, phosphatidylinositol bisphosphate;
IP3, inositol trisphosphate; DAG, diacylglycerol; DC,
dendritic cell; ADCC, antibody-dependent cellular cytotoxicity;
ADCP, antibody-dependent phagocytosis; Th, T helper; ROS, reactive
oxygen species. |
Dendritic cells (DCs): Multireceptor-mediated
antigen uptake and immune regulation. The function of FcRγ in
DCs exhibits greater diversity and regulatory capacity than in mast
cells and basophils, contingent upon the reprogramming of signals
from the bound receptor complex FcεRI. The FcεRI expressed by DCs
typically adopts a trimeric structure lacking the β chain
(αγ2) (14). This
structural difference leads to functional reprogramming, shifting
from rapid effector responses towards antigen capture and
presentation. In allergic diseases such as atopic dermatitis (AD),
upregulation of FcεRI on cutaneous DCs enables efficient
internalization of allergens upon IgE binding. These DCs then
migrate to lymph nodes, primarily initiating T helper (Th)2-type
immune responses, which is an important mechanism in conditions
such as AD (14).
For CLRs lacking intracellular signaling domains,
such as Dectin-2, FcRγ serves as an indispensable signaling
partner. Upon recognizing fungal α-mannan, Dectin-2 activates SYK
via the ITAM of FcRγ (18,19). This subsequently activates the
NF-κB pathway through the CARD9-BCL10-MALT1 junction complex,
driving cytokine production (including IL-6 and IL-23) and
promoting Th17 differentiation (Fig.
2) (18,19). This is a critical component of
antifungal immunity.
Previous research indicates that FcRγ also exerts
negative regulatory effects on DCs. For instance, upon binding to
Dectin-1 bearing a semi-ITAM, it may attenuate signal output by
recruiting phosphatases such as SHP-1 and PTEN (16). This finely tunes DC maturation and
cytokine production, preventing excessive inflammation (Fig. 2). This demonstrates that FcRγ in
DCs functions not only as a transmitter of activation signals but
also as a precise regulator of immune responses.
NK cells: CD16A (FcγRIIIA) signaling and
antibody-dependent cytotoxicity. Within NK cells, FcRγ
primarily functions as the signal-adaptor subunit for the
low-affinity IgG receptor CD16A (FcγRIIIa). Their binding is
essential for the stable expression and functional activity of this
receptor on the cell surface. Upon NK cell recognition of tumor
cells coated with antibodies (namely therapeutic monoclonal
antibodies) via CD16A, phosphorylation of the ITAM recruits and
activates SYK. Signaling in NK cells predominantly activates the
SYK/PI3K-δ/MAPK axis (20). This
pathway drives cytoskeletal reorganization as well as polarization
and release of cytotoxic granules-a process involving the directed
trafficking of perforin- and granzyme-containing vesicles toward
the immune synapse - and promotes IFN-γ production, thereby
efficiently executing ADCC (Fig.
2) (20). This constitutes a
key mechanism underpinning the efficacy of numerous antibody
therapeutics. This study indicates that de-fucosylated antibodies,
by enhancing CD16A binding, significantly amplify downstream
signaling through SYK/PI3K/MAPK and other pathways, thereby
enhancing ADCC efficiency (4).
Consequently, the functional state of FCER1G directly
influences the efficacy of monoclonal antibody-based tumor
immunotherapies.
Macrophages: Integrated signaling regulates
phagocytosis, polarization and inflammatory balance. The
function of FcRγ in macrophages integrates effector and regulatory
roles, reflecting their immunological multifunctionality. As a
signal-transducing subunit of receptors such as FcγRI (CD64), FcγRγ
plays a pivotal role in mediating ADCP. Following immune complex
cross-linking of FcγR, SYK activated by FcRγ-ITAM synergistically
initiates multiple downstream pathways, including PLCγ, PI3K and
MAPK (5). These collectively
regulate actin remodeling, phagosome maturation and reactive oxygen
species production, which is essential for phagocytosis, thereby
efficiently eliminating antibody-coated targets (Fig. 2) (5).
Regarding bidirectional regulation of inflammatory
phenotype and function, FcRγ signaling serves as a pivotal node in
modulating macrophage polarization and inflammatory homeostasis.
For instance, in atherosclerosis models, immune complexes activate
FcγR to drive macrophage polarization towards a pro-inflammatory
M1-like phenotype (21). This
activates the NF-κB pathway, and leads to substantial release of
inflammatory mediators such as TNF-α and IL-6, exacerbating plaque
instability and vascular inflammation (21). Conversely, FCER1G deficiency
predisposes macrophages towards an anti-inflammatory M2-like
phenotype, mitigating disease progression (21). This demonstrates the pivotal role
of FcRγ in determining macrophage functional output.
Other cell types and functions. FcRγ also
participates in other immune processes, such as acting as a
co-signaling chain for the collagen receptor GPVI in platelets to
mediate thrombosis, and stabilizing cell surface receptors in type
3 innate lymphoid cells (ILC3s) to promote IL-22 production to
combat infections (22,23). Its extensive involvement further
underscores its importance as a universal ITAM-binding adaptor
protein. FcRγ also functions as a signaling partner for myeloid
receptors such as OSCAR and TREM-1, playing roles in several
processes such as osteoclast differentiation and inflammatory
amplification (24,25).
Common mechanisms and differentiation
basis of FcRγ signaling
Despite varying functions across different cell
types, the molecular mechanisms initiating FcRγ signaling remain
highly conserved. Ligand-induced receptor clustering leads to
mutual phosphorylation and activation of adjacent Src family
kinases, which subsequently phosphorylate two tyrosine residues on
the FcRγ-ITAM. The doubly phosphorylated ITAM recruits and
activates SYK with high affinity, establishing SYK as the central
hub for downstream signal differentiation (13,26).
From SYK, signals diverge into at least three
primary pathways, with cells selectively amplifying specific
pathways according to their functional predisposition: i) The
PLCγ/Ca²+ pathway, which drives rapid degranulation
responses in effector cells such as mast cells; ii) the PI3K/MAPK
pathway, which predominantly governs ADCC/ADCP, cell migration,
cell proliferation, and partial cytokine synthesis in NK cells and
macrophages; and iii) the CARD9/NF-κB pathway, which predominantly
governs transcriptional activation of pro-inflammatory cytokines
and chemokines in DCs, macrophages and other cells following
pathogen recognition via CLRs.
Therefore, FcRγ functions as a universal signaling
molecule, receiving initial signals from upstream receptor clusters
via its ITAM module and subsequently diverting these signals
through SYK. Its ultimate functional output is co-programmed by the
specific receptor it binds to and the downstream signaling network
preferences dictated by the host cell type. This characteristic of
‘common mechanism, differential programming’ provides the molecular
basis for understanding how FcRγ serves distinct roles across
diverse pathological contexts, including allergy, infection,
autoimmunity and even tumorigenesis, while also offering a logical
explanation for its paradoxical dual function within the TME.
4. Immunoregulatory role of FCER1G in
inflammatory diseases
Previous studies have revealed the multifaceted
pathological roles of the innate immunity-related FCER1G
gene. In non-neoplastic diseases, this gene contributes to the
pathogenesis of inflammatory disorders such as eczema and AD
(27-31).
Multilayered epigenetic regulation of FCER1G
expression. The expression of FCER1G undergoes intricate
and multilevel epigenetic regulation, forming a crucial molecular
basis for its abnormal expression and functional plasticity in
disease. In inflammatory conditions, DNA methylation emerges as a
central regulatory mechanism controlling its expression, offering a
relatively clear framework for understanding its functional
regulation (Fig. 3).
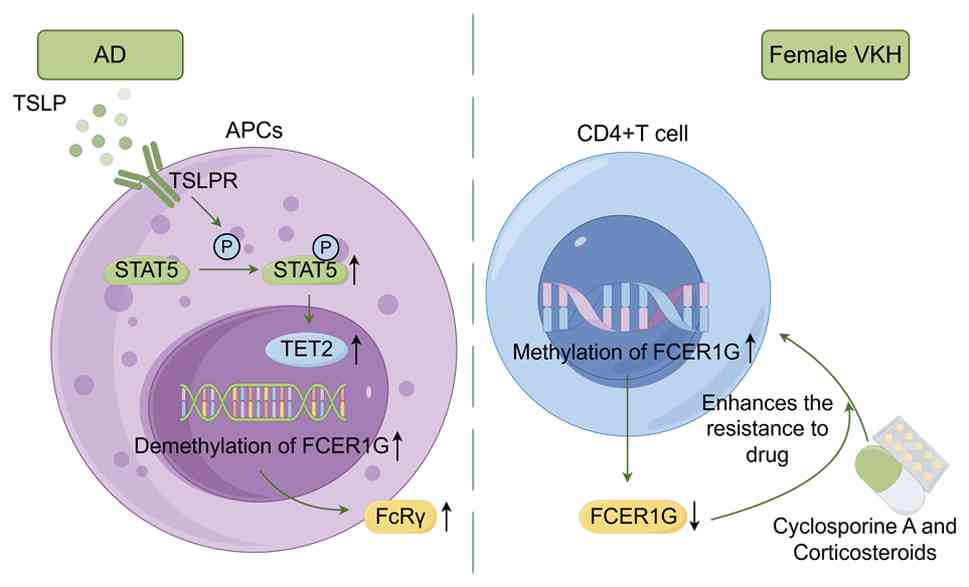 | Figure 3Examples of epigenetic regulation of
FCER1G in inflammatory diseases. In AD, the cytokine TSLP induces
active demethylation of FCER1G by activating STAT5 and recruiting
the demethylase TET2 to the FCER1G promoter region, thereby
upregulating FCER1G expression and exacerbating allergic responses.
In female patients with Vogt-Koyanagi-Harada disease,
hypermethylation of the FCER1G promoter region in CD4+ T
cells leads to gene silencing and is associated with glucocorticoid
resistance. The figure was drawn with FigDraw2.0 (supplied by
Hangzhou Sifei Technology Co., Ltd.). FCER1G, Fc fragment of IgE
receptor 1G; TSLP, thymic stromal lymphopoietin; TSLPR, thymic
stromal lymphopoietin receptor; AD, atopic dermatitis; TET2,
ten-eleven translocation 2; STAT5, signal transducer and activator
of transcription 5; APCs, antigen-presenting cells; FcRγ, Fc
receptor common γ chain; p, phosphorylation. |
Dynamic regulation of DNA
methylation/demethylation. DNA methylation constitutes a core
epigenetic mechanism in the regulation of FCER1G expression.
Previous studies have shown a significant inverse correlation
between the methylation status of CpG islands in the FCER1G
promoter region and its expression levels (28,32-35).
In patients with AD, monocytes exhibit specific hypomethylation of
the FCER1G promoter, which directly upregulates its mRNA and
protein levels. This consequently causes overexpression of
receptors such as FcεRI on antigen-presenting cell (APC) surfaces,
exacerbating allergic reactions (28). This causal association has been
directly validated by patch methylation combined with luciferase
reporter assays (28,36).
In rheumatoid arthritis, FCER1G also displays
a pattern of high expression associated with hypomethylation
(33,34). Similarly, in female patients with
Vogt-Koyanagi-Harada disease, hypermethylation of the FCER1G
promoter in CD4+ T cells silences its expression,
enhancing patient resistance to cyclosporine A and corticosteroids
(32). This suggests that
intervention targeting the methylation status of the FCER1G
promoter may represent a potential therapeutic sensitization
strategy.
Synergistic interaction between key transcription
factors and demethylases: The demethylation of FCER1G is
driven by specific cytokine signals through an active mechanism
involving transcription factor-mediated recruitment of DNA
demethylases. Within the pathological environment of AD, thymic
stromal lymphopoietin (TSLP) produced by epithelial cells serves as
the key driver. TSLP activates its receptor, leading to signal
transduction and the phosphorylation of signal transducer and
activator of transcription 5 (STAT5) (29,37).
Activated phosphorylated (p)-STAT5, acting as a transcription
factor, is recruited to the FCER1G promoter region, simultaneously
recruiting the DNA demethylase ten-eleven-translocation 2 (TET2),
which catalyzes the conversion of 5-methylcytosine to
5-hydroxymethylcytosine, thereby initiating an active demethylation
program that relieves the transcriptional repression of
FCER1G (29,37). This TSLP/p-STAT5/TET2 axis forms a
coherent epigenetic reprogramming pathway, explaining the
epigenetic basis for the sustained high expression of FCER1G
in AD.
Regulation through chromatin plasticity by
transcription factors. In addition to the aforementioned core
mechanisms, the regulation of chromatin plasticity by specific
transcription factors also contributes to the control of
FCER1G expression. For instance, in hematopoietic stem cells
(HSCs), the transcription factor Bcl11a directly represses
FCER1G transcription by suppressing chromatin accessibility
at its promoter region, a process that is critical for maintaining
HSC quiescence (38). This
mechanism illustrates how transcription factors can precisely
control FCER1G expression at the epigenetic level by
modifying chromatin architecture.
The aforementioned complex epigenetic regulatory
mechanisms determine that FCER1G expression exhibits a high
degree of microenvironmental dependency and dynamic plasticity.
This characteristic enables it to perform differentiated roles
across distinct pathological contexts, where both its expression
levels and functional outputs (pro-inflammatory or
anti-inflammatory) must be interpreted within specific diseases
(39-41).
Context-dependent regulation of FcRγ
in anti-infection immunity
In anti-infection immunity, the FcRγ exhibits a dual
function, with its presence or absence influencing pathogen
clearance and host prognosis differently across various infection
models, thus profoundly demonstrating its context-dependent nature.
In a chronic lymphocytic choroid plexus meningitis virus infection
model, FcRγ expressed in NK cells delays pathogen clearance by
suppressing virus-specific CD8+ T-cell responses
(6). In addition, deletion of FcRγ
significantly reduces mortality in mouse models of sepsis induced
by lipopolysaccharide or Escherichia coli, which is
characterized by lower serum levels of TNF-α, IL-6 and
IL-10(39). These models suggest
that, under certain circumstances, FcRγ-mediated signaling may
prove detrimental to infection control or host survival by
promoting excessive inflammation or suppressing adaptive immunity.
However, in other infection models, FcRγ plays an indispensable
role in defensive mechanisms. Conversely, the absence of FcRγ in
ILC3s impairs JAK-STAT pathway activation and reduces IL-22 and
IL-17A secretion, thereby increasing mortality during fungal
infection (22). Similarly, during
Pneumocystis pneumonia, FcRγ deficiency decreases the
production of inflammatory cytokines (TNF-α, IL-6 and IL-1β) but
compromises pathogen clearance efficiency (40). This indicates that FcRγ-mediated
immune responses are essential for the effective clearance of the
aforementioned pathogens.
The dual outcomes of FcRγ during infection
(beneficial or detrimental) are likely determined by two major
factors. On the one hand, the type of pathogen and the nature of
the infection are critical. For intracellular chronic viruses (such
as lymphocytic choriomeningitis virus) or systemic bacterial toxins
(sepsis), excessive inflammation or inappropriate immune regulation
may lead to immunopathology or tissue damage, where FcRγ activity
often contributes to detrimental outcomes. Conversely, combating
certain fungi and opportunistic pathogens (such as
Pneumocystis) necessitates FcRγ-dependent rapid initiation
of innate immune effector programs in specific cells (namely
ILC3s), where its function is beneficial. On the other hand, maybe
the dominant immune cells and effector mechanisms govern the
response. FcRγ's function is entirely dependent on its cellular
environment. In NK cells, it may modulate immune crosstalk,
occasionally suppressing T cell function, whereas, in ILC3s, it
directly stimulates antimicrobial cytokine production. Thus, the
cell type dictates whether FcRγ-mediated signaling leads to
inhibitory regulation or effector activation.
In summary, the presence of FcRγ is not inherently
positive or negative in infectious immunity. Instead, its value
depends on the immune equilibrium set by the specific infection
microenvironment. FcRγ dynamically modulates the intensity and
quality of immune responses by influencing two critical balances:
i) Pro-inflammation vs. anti-inflammation and ii) immune activation
vs. suppression. This capacity for flexible role-switching in
response to microenvironmental signals and precise regulation of
immune balance bears striking similarity to the functional
plasticity of immune checkpoint molecules within the TME, and
provides a core logical framework for understanding the complex
role of FCER1G in tumor immunity.
5. Expression of FCER1G in
tumors
Pan-cancer expression profile and
characteristics associated with the immune microenvironment
Pan-cancer analysis based on the TIMER 2.0 database
(https://compbio.cn/timer2/) shows that
FCER1G is significantly upregulated in 9 types of malignant tumor
compared with normal tissues (Fig.
4). These include esophageal carcinoma, glioblastoma multiforme
(GBM), head and neck squamous cell carcinoma, kidney clear cell
carcinoma, kidney papillary cell carcinoma, stomach adenocarcinoma
(STAD) and thyroid carcinoma. Gene enrichment analyses further
reveal that FCER1G is predominantly involved in cell
proliferation-related pathways across multiple tumor types,
particularly STAD, TGCT, acute myeloid leukemia and GBM (P<0.05,
false discovery rate <0.25) (41). Furthermore, FCER1G expression
positively correlates with >50% of immune checkpoint genes,
suggesting that it functions as a hub regulator within the tumor
immunoregulatory network (41).
Single-cell transcriptomic data from the TISCH database indicate
that FCER1G is preferentially expressed in monocyte/macrophage
populations within the TME across ~85% of the tumors analyzed, thus
providing cellular-level evidence for its involvement in TME
remodeling.
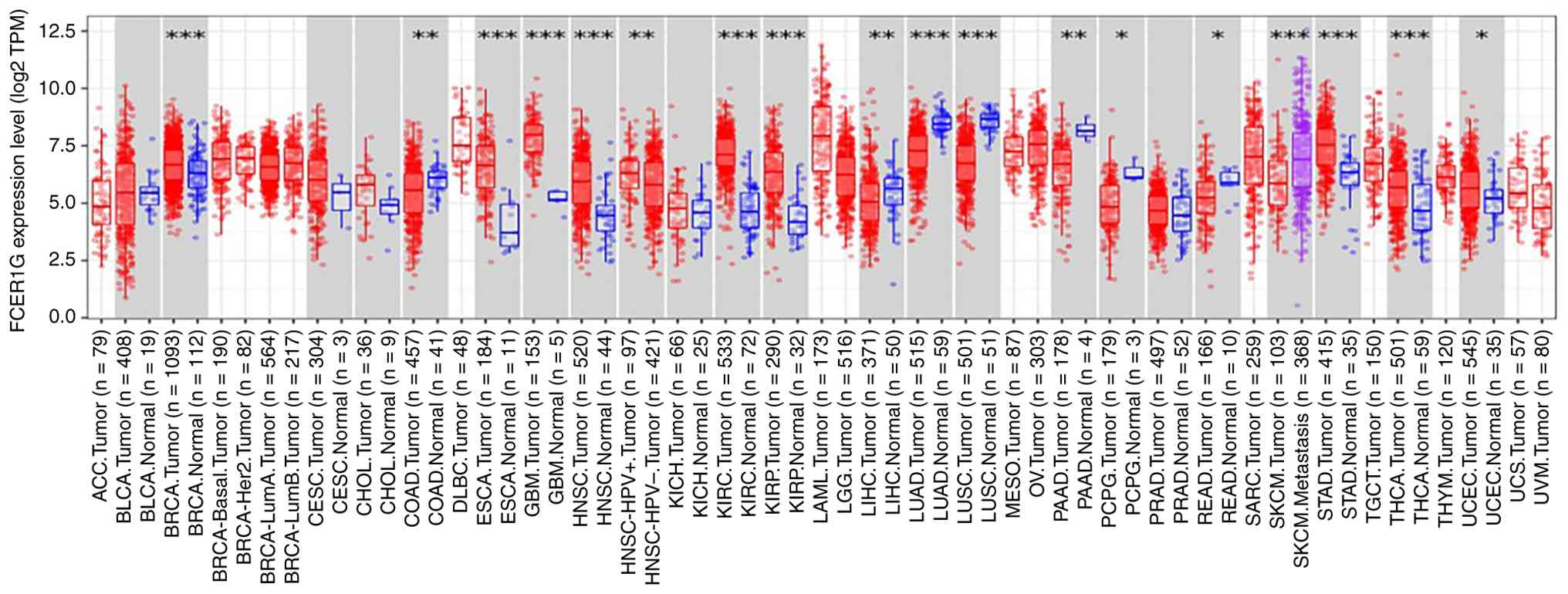 | Figure 4Expression of FCER1G across 33
cancer tissue types and 21 paired normal tissues based on the
TIMER2.0 database (*P<0.05, **P<0.01
and ***P<0.001). The figure was from the TIMER2.0
database (https://compbio.cn/timer2/).
FCER1G, Fc fragment of IgE receptor 1G; TPM, transcripts per
million. ACC, adrenocortical carcinoma; BLCA, bladder urothelial
carcinoma; BRCA, breast invasive carcinoma; CESC, cervical squamous
cell carcinoma and endocervical adenocarcinoma; CHOL,
cholangiocarcinoma; COAD, colon adenocarcinoma; DLBC, diffuse large
B-cell lymphoma; ESCA, esophageal carcinoma; GBM, glioblastoma
multiforme; HNSC, head and neck squamous cell carcinoma; KICH,
kidney chromophobe; KIRC, kidney renal clear cell carcinoma; KIRP,
Kidney Renal Papillary Cell Carcinoma; LAML, acute myeloid
leukemia; LGG, brain lower grade glioma; LIHC, liver hepatocellular
carcinoma; LUAD, lung adenocarcinoma; LUSC, lung squamous cell
carcinoma; MESO, mesothelioma; OV, ovarian serous
cystadenocarcinoma; PAAD, pancreatic adenocarcinoma; PCPG,
pheochromocytoma and paraganglioma; PRAP, prostate adenocarcinoma;
READ, rectum adenocarcinoma; SARC, sarcoma; SKCM, skin cutaneous
melanoma; STAD, stomach adenocarcinoma; TGCT, testicular germ cell
tumors; THCA, thyroid carcinoma; THYM, thymoma; UCEC, uterine
corpus endometrial carcinoma; UCS, uterine carcinosarcoma; UVM,
uveal melanoma. |
Dual nature of tissue-specific
expression patterns and prognostic value. FCER1G expression
correlates with poor prognosis in tumors
Comprehensive analysis indicates that FCER1G
is highly expressed in multiple solid tumors, including clear cell
renal cell carcinoma (ccRCC) (presented as KIRC in Fig. 4), esophageal squamous cell
carcinoma (ESCC) (included within the esophageal cancer group when
presented in Fig. 4) and STAD
(7,8,42)
(Fig. 4). Its expression is
predominantly enriched in myeloid immune cells, where it promotes
the formation of an immunosuppressive microenvironment, thereby
contributing to poor prognosis.
Data from the Human Protein Atlas (HPA) show
markedly stronger immunohistochemical staining of FCER1G in
renal carcinoma tissues than in adjacent normal tissues (Fig. 5). According to Wang et al
(7), analysis of the ONCOMINE
database and three validation datasets identified significantly
elevated FCER1G expression in ccRCC, where high expression
correlated with reduced overall survival (OS). Similarly, Dong
et al (43) observed that,
in ccRCC, FCER1G expression exhibited a strong correlation
with CD68 and co-localization with macrophages. Their
concurrent overexpression portends an unfavorable prognosis. Joint
assessment of FCER1G and CD68 expression levels can
optimize prognostic stratification models for patients.
Furthermore, gene set enrichment analysis revealed that high
FCER1G expression is associated with suppression of T-cell
activation and proliferation (43). FCER1G shows moderate
diagnostic accuracy (area under the curve, 0.74) in distinguishing
localized (stage I/II) from advanced (stage III/IV) ccRCC, and
functions as an independent prognostic biomarker (44).
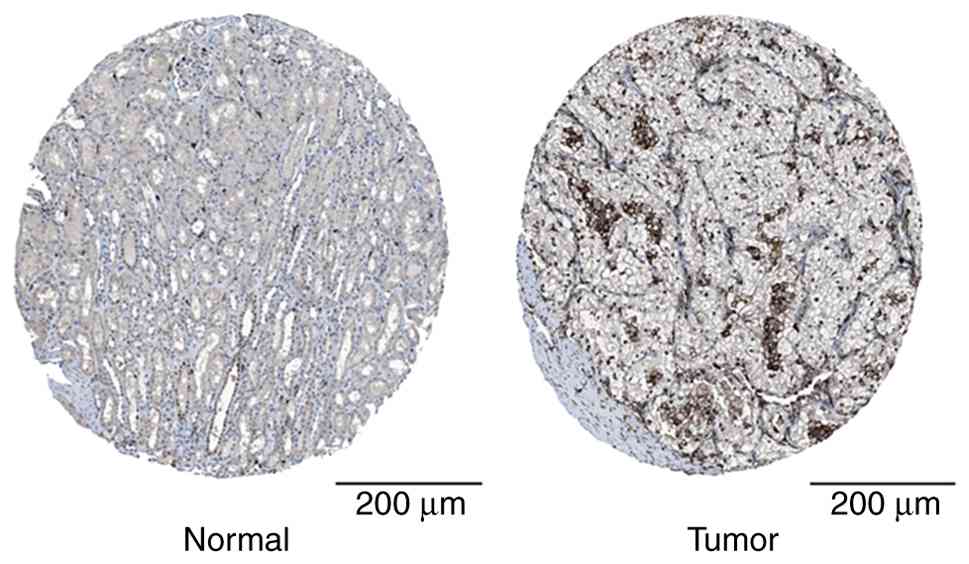 | Figure 5FCER1G protein is highly
expressed in ccRCC tissues. Immunohistochemical staining from the
Human Protein Atlas database revealed significantly elevated FCER1G
expression in ccRCC tissue compared with that in normal renal
tissue (increased staining intensity and proportion of positive
cells). The normal kidney sample corresponds to a 16-year-old male
donor (sample ID, 1,767; staining, medium; intensity, low; quantity
of FCER1G-positive cells <25%; location of FCER1G protein,
nuclear), while the renal cancer sample corresponds to a
70-year-old female patient with ccRCC (sample ID, 1,498; staining,
medium; intensity, medium; quantity of FCER1G-positive cells
>75%; location of FCER1G protein, nuclear). This figure is from
the public database HPA (Human Protein Atlas), for which patient
consent for publication is not required in accordance with the
database's usage guidelines (https://www.proteinatlas.org/). Scale bar, 200 µm.
FCER1G, Fc fragment of IgE receptor 1G; ccRCC, clear cell renal
cell carcinoma. |
In ESCC, FCER1G is predominantly enriched
within the tumor stroma. Double immunofluorescence staining clearly
demonstrates that FCER1G-positive cells highly co-express
the M2 macrophage marker CD163, meaning that the majority of
infiltrating M2 macrophages simultaneously express FCER1G
(42). The infiltration density of
these FCER1G+ M2 macrophages correlates directly
with worse prognosis, and FCER1G itself constitutes an
independent risk factor affecting patients' OS (42). In STAD, analyses from multiple
transcriptomic cohorts (The Cancer Genome Atlas, GSE13195 and
GSE15459) consistently showed higher FCER1G expression in
tumor tissues than in adjacent normal tissues (8,41,45).
Its overexpression is linked to poorer OS and increased
infiltration of M2 macrophages (8). Furthermore, FCER1G expression
progressively increases with advancing tumor stage, suggesting a
dynamic upregulation trend during tumor progression. This finding,
similar to that observed for ESCC, suggests that FCER1G may
be primarily expressed in M2-type tumor-associated macrophages
(TAMs) within STAD and participate in shaping the immunosuppressive
microenvironment.
Additionally, in gliomas, diffuse large B-cell
lymphoma, bladder cancer and papillary thyroid carcinoma,
FCER1G acts as a progression-associated gene, where high
expression of FCER1G correlates with aggressive tumor
behavior, poor prognosis and distinct immune infiltration patterns
(11,46-51).
In osteosarcoma, a dual-gene risk model incorporating FCER1G
and SPI1, developed based on Cox regression analysis, holds
independent prognostic significance. Patients in the low-expression
group for this signature exhibit an immunosuppressive
microenvironment, worse clinical outcomes and a higher risk of
metastasis, highlighting the potential of FCER1G as a
prognostic biomarker and immunotherapeutic target (52). It is noteworthy that bioinformatics
analysis suggests that FCER1G may be a hypermethylated gene
in osteosarcoma, offering a potential epigenetic hypothesis to
explain its low expression in this tumor type. However, the precise
regulatory mechanisms require experimental validation (35).
In summary, FCER1G is highly expressed in
multiple solid tumors, including ccRCC, ESCC, STAD and glioma. By
promoting M2 macrophage infiltration and shaping an
immunosuppressive TME, it contributes to tumor progression and
serves as a consistent indicator of poor prognosis across various
cancer types (Table I).
 | Table IExpression patterns, cellular types,
prognostic correlations and immune microenvironment characteristics
of FCER1G in different tumors. |
Table I
Expression patterns, cellular types,
prognostic correlations and immune microenvironment characteristics
of FCER1G in different tumors.
| Tumor | FCER1G
expression | Prognosis | Primary cell types
expressing FCER1G | Key associated
immune cells/ pathways |
|---|
| ccRCC | Upregulated | Poor | Tumor-associated
macrophages | Co-expressed with
CD68, associated with suppression of T-cell activation |
| ESCC | Upregulated | Poor | M2 macrophages | Positively
correlated with CD163+ M2 macrophage infiltration |
| STAD | Upregulated | Poor | Hematopoietic
immune cells (such as macrophages) | Associated with
enhanced M2 macrophage infiltration |
| MM | Downregulated | Favorable | NK and other
effector immune cells | Associated with NK
cell-mediated cytotoxicity pathway |
| UCEC | Downregulated | Favorable | Multiple immune
cells (including B, CD8+ T and dendritic cells) | Associated with
increased infiltration of B, CD8+ T and other cells |
FCER1G correlates with favorable patient
prognosis. In contrast to its generally pro-tumor pattern,
FCER1G displays a protective role in certain malignancies,
where high expression is associated with prolonged survival and
enhanced immune activation. In MM, analyses of several datasets
(including GSE39754, GSE5900 and GSE2113) reveal a progressive
decline in FCER1G expression with disease advancement
(9,53). Multivariate Cox regression analysis
identifies high FCER1G expression as an independent
predictor of both event-free survival and OS, establishing it as a
favorable prognostic biomarker in MM (9). Furthermore, FCER1G
overexpression correlates with NK cell-mediated cytotoxic pathways
(9). Notably, previous research
has identified a functionally distinct FcεRIγ- NK cell
subset (termed g-NK cells) (54).
Compared to conventional NK cells, these cells exhibit enhanced
ADCC following CD16 cross-linking and significantly amplify the
efficacy of monoclonal antibodies such as daratumumab in
preclinical models (54). This
offers a novel perspective on the association between FCER1G
expression in MM, NK cell function and favorable prognosis.
Similarly, in uterine corpus endometrial carcinoma (UCEC), high
FCER1G expression predicts improved prognosis, and
correlates with increased infiltration of immune cells, including B
cells, CD8+ T cells and DCs (Table I) (10).
In lung adenocarcinoma (LUAD), FCER1G
expression exhibits stage-specific dynamics, being downregulated in
early stages and restored at advanced stages (55). Network analyses further indicate
that FCER1G consistently functions as an immune regulatory
hub throughout LUAD progression, and that its functional loss
impairs antitumor immune activity (55).
Overall, the prognostic significance of
FCER1G shows clear tumor-type specificity.
In malignancies such as MM and UCEC, as well as in
specific stages of LUAD, elevated FCER1G expression is not
associated with tumor promotion. Instead, it may exert a protective
effect by sustaining or activating antitumor immune responses,
ultimately predicting more favorable clinical outcomes (Table I).
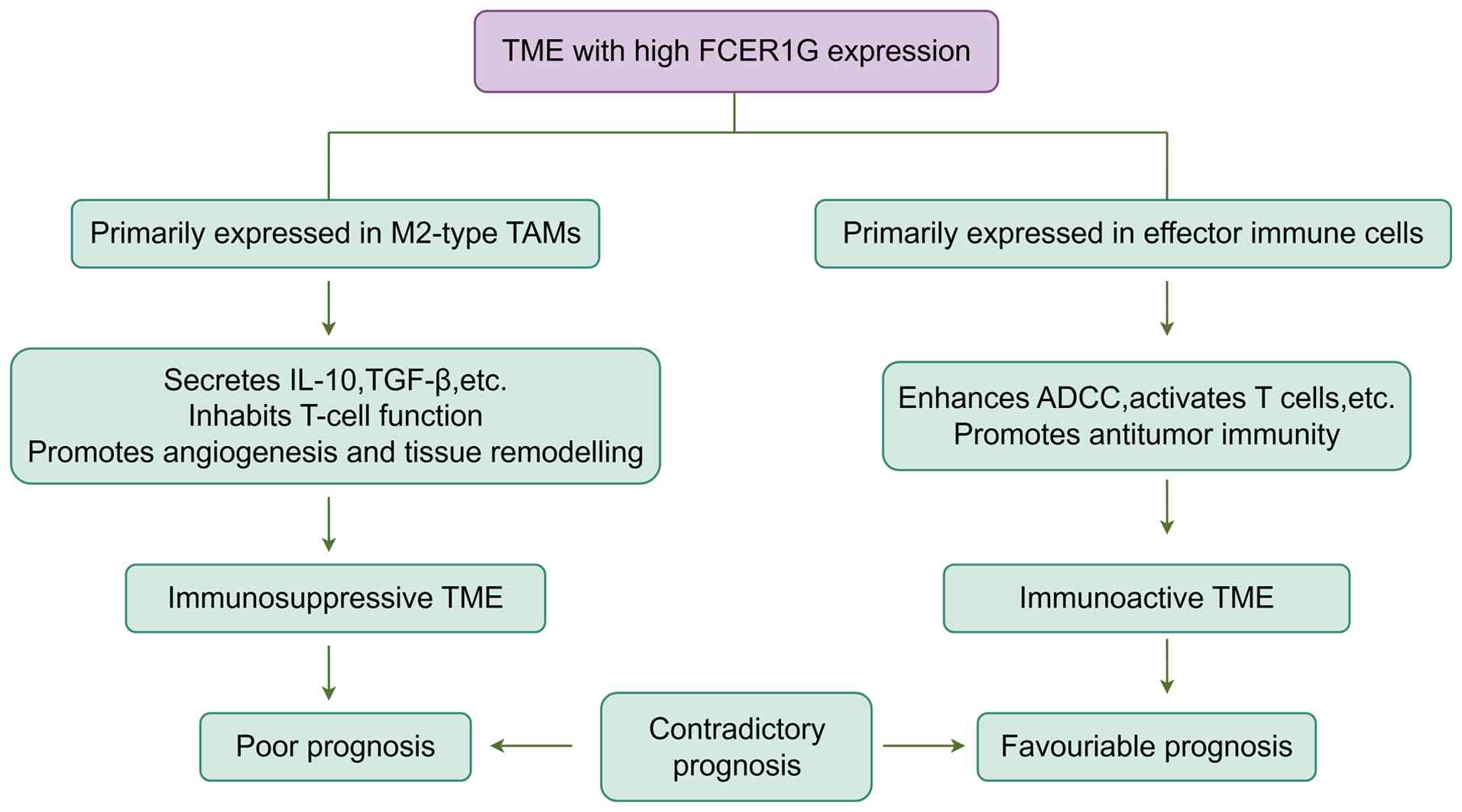
Contradictory prognostic value of FCER1G arises
from heterogeneity in the TME. In summary, the prognostic value
of FCER1G exhibits marked inconsistencies, fundamentally
stemming from the cellular types expressing FCER1G, which
determine the nature of the specific, coordinated immune processes
in which it participates (Fig.
6).
Regarding its expression in immunosuppressive
myeloid cells (pro-tumor function), in solid tumors such as ccRCC
and ESCC, FCER1G is primarily expressed in M2-type TAMs.
These cells shape an immunosuppressive TME by secreting
immunosuppressive factors (such as IL-10 and TGF-β), depleting T
cells and promoting angiogenesis. This drives tumor progression and
immune evasion, leading to poor prognosis (42,43).
Regarding its expression in cytotoxic cells or APCs (antitumor
function), in tumors such as MM and UCEC, high expression of
FCER1G correlates with the active state of effector cells or
APCs, including DC, and NK, cytotoxic T and B cells. In this
context, FCER1G functions as a signaling adaptor for
activation receptors (CD16A) on these cells, participating in the
initiation of antitumor immune responses such as ADCC and T-cell
activation, thereby correlating with favorable prognosis (9,10,41).
Therefore, assessing the clinical relevance of
FCER1G necessitates moving beyond mere expression levels to
deeply analyze its cellular localization and the associated
functional state of the overall immune microenvironment. This
cellular context determinism lies at the core of understanding its
complexity as a biomarker.
Core mechanisms driving tumor
progression and immune remodeling. Regulation of immune
microenvironment structure and function
FCER1G exhibits distinct immunoregulatory
patterns across different tumor types. In STAD, its expression
correlates positively with the infiltration of M1/M2 macrophages,
quiescent mast cells and DCs, but negatively with plasma and
CD8+ T cells (45).
This opposing infiltration pattern indicates that FCER1G may
facilitate tumor progression by reshaping an immunosuppressive
immune microenvironment. In glioma, FCER1G expression is
strongly associated with the infiltration of T cells, macrophages
and B cells, further implicating it in disease progression and
immune landscape remodeling (47).
Notably, previous single-cell RNA sequencing studies have
identified a tumor-specific, innate-like cytotoxic T-cell subset
characterized by FCER1G+ αβTCR+ cells
(56,57). FCER1G serves as a definitive
lineage marker for this population. Unlike conventional
tumor-reactive T cells, these cells recognize unmutated
self-antigens presented by MHC-I molecules rather than
tumor-derived neoantigens. Their intratumoral activation and
effector functions depend strictly on the IL-15 signaling axis,
underscoring a unique FCER1G-mediated mechanism of cytotoxic
immune regulation within the TME (56,57).
This identification implies that FCER1G is
not merely a participant in immunosuppression but also acts as a
phenotypic identity tag for specific antitumor immune cell subsets,
highlighting its multifaceted roles within the TME. Collectively,
the evidence suggests that FCER1G possesses dual regulatory
properties. It can promote immunosuppressive phenotypes in certain
contexts while also functioning as a key regulatory node for
antitumor responses in distinct immune populations. Dysregulation
of its expression or signaling may therefore disrupt immune
homeostasis and drive the pathological remodeling of the TME.
Supporting its functional relevance, analyses of
the Gene Expression Profiling Interactive Analysis and HPA
databases showed that FCER1G expression is significantly
higher in PAAD tissues than in normal tissues at both the mRNA and
protein levels. In animal models, FcRγ-deficient
[FCER1G-/-, FcRγ knockout (KO)] mice
bearing pancreatic ductal adenocarcinoma exhibit reduced tumor
growth and attenuated desmoplasia, accompanied by a complete loss
of FcγRI/III expression within tumors (58). These findings indicate that
FCER1G likely modulates the TME and drives malignant
progression through the regulation of Fc receptor-mediated
signaling pathways.
Promotion of angiogenesis. Andreu et
al (59), using an
HPV16+/FcRγ-/- mouse model,
revealed a key role for FcRγ in tumorigenesis. Compared with
wild-type (WT) mice, FcRγ KO mice showed a lower incidence of SCC.
This effect was associated with inhibited angiogenesis and
decreased expression of major oncogenic factors, including vascular
endothelial growth factor (VEGF) and matrix metalloproteinase-9
(MMP-9) (59). Further experiments
demonstrated that mast cells and macrophages promote endothelial
cell migration via FcRγ-dependent pathways, thereby enhancing the
in vivo tumorigenicity of the PDSC5 SCC cell line (59). Collectively, these findings
indicate that FcRγ promotes tumor angiogenesis by mediating
interactions between immune and stromal cells within the TME.
Promotion of tumor metastasis. The
expression status of FCER1G is frequently implicated in
tumor metastasis, although its effects vary among cancer types. In
hepatocellular carcinoma (HCC), FCER1G shows a downregulated
trend, and its low expression is associated with the activation of
apoptotic and ferroptotic pathways (60). Previous in vitro experiments
have shown that silencing FCER1G enhances the proliferation and
migration of HCC cells by upregulating Snail1, TWIST1 and
N-cadherin while suppressing E-cadherin expression (60). These results suggest that
FCER1G may restrain HCC invasiveness by modulating
epithelial-mesenchymal transition (EMT) processes.
In malignant melanoma (MEL), the role of FcRγ
exhibits a more intricate profile, characterized by
context-dependent effects. On the one hand, lung metastasis can be
suppressed by treatment with the monoclonal antibody TA99 or
intravenous Ig (IVIG) (61,62).
This antimetastatic effect depends entirely on FcRγ, as it is
abolished in FcRγ KO mice, indicating that ADCC serves as a key
mechanism. On the other hand, FcRγ deficiency itself exerts dual
and opposing effects on metastasis (Fig. 7). Specifically, the absence of FcRγ
enhances platelet adhesion to circulating tumor cells, and
stimulates platelets to release the chemokines C-X-C motif
chemokine ligand (CXCL)5 and CXCL7. This cascade promotes
neutrophil recruitment to lung tissue and creates a pro-transfer
microenvironment that accelerates lung metastasis without affecting
primary tumor growth (3).
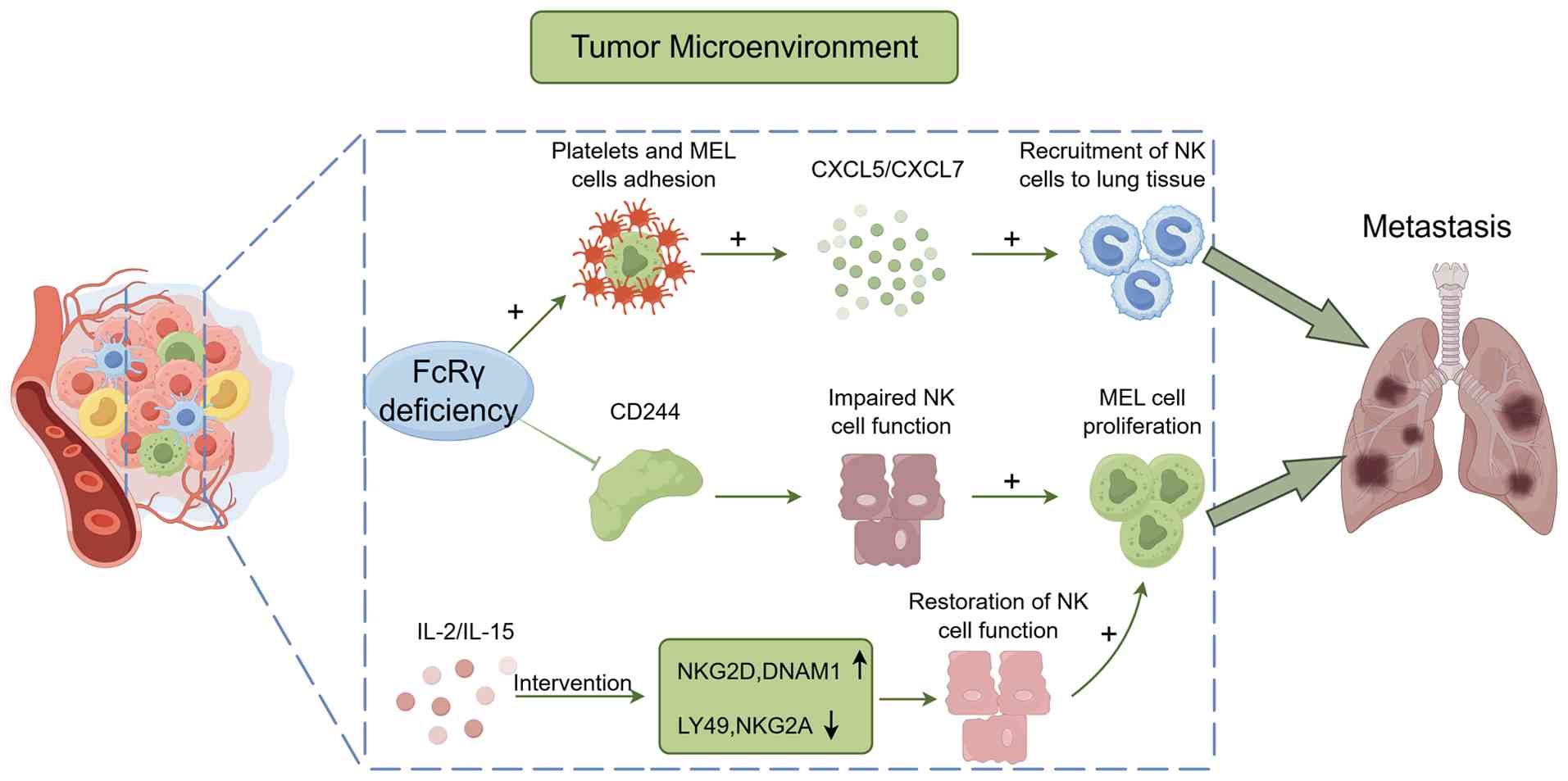 | Figure 7Two mechanisms by which FcRγ
deficiency promotes tumor lung metastasis in MEL: i) Enhances
adhesion between platelets and circulating tumor cells, inducing
platelets to release chemokines CXCL5/7 to recruit neutrophils and
establish a pre-metastatic microenvironment; and ii) reduces the
expression of CD244, a key maturation molecule for NK cells,
thereby impairing NK cell function. The figure was drawn with
FigDraw2.0 (supplied by Hangzhou Sifei Technology Co., Ltd.). MEL,
malignant melanoma; NK, natural killer; FcRγ, Fc receptor common γ
chain; CXCL, C-X-C motif chemokine ligand; NKG2, natural killer
cell group 2; LY49, Lymphocyte Antigen 49 Complex; DNAM-1, DNAX
Accessory Molecule-1. |
Furthermore, FcRγ KO impairs CD244-mediated
inhibitory signaling during NK cell development, leading to
diminished NK cell function (63).
This defect can be reversed by treatment with IL-2 or IL-15, which
upregulates activation receptors [natural killer cell group 2
(NKG2)D and DNAX accessory molecule-1] and downregulates inhibitory
receptors (LY49 and NKG2A), thereby restoring NK cell cytotoxicity
and suppressing metastasis (63).
Therefore, therapeutic strategies aimed at modulating FcRγ
signaling must carefully balance the benefits of enhancing
antibody-dependent tumor clearance against the potential risk of
facilitating metastatic dissemination.
FcRγ-dependent therapeutic responses
and resistance mechanisms. Essential role of the FcγRI/III
signaling pathway
FcRγ serves as a critical molecular determinant for
the efficacy of multiple antibody-based therapies. In colorectal
cancer (CRC), FCER1G expression is paradoxically higher in
adjacent non-tumorous tissues than in tumor tissues (64). Within this setting, IVIG treatment
fails to produce antitumor effects in FcRγ KO mice.
Mechanistically, the therapeutic activity of IVIG depends on
FcγRI/III signaling, which drives the reprogramming of TAMs from an
M2-like, pro-tumor state toward an M1-like, antitumor phenotype
(64).
In adult T-cell leukemia/lymphoma (ATL) models,
anti-CD2 monoclonal antibody MEDI-507 and anti-CD25 monoclonal
antibodies significantly suppress tumor progression and prolong
survival, with their efficacy strictly dependent on FcRγ (65,66).
Specifically, polymorphonuclear leukocytes and monocytes eliminate
CD2+/CD25+ tumor cells through
FcγRIII-mediated ADCC (65,66).
The absence of FcRγ disrupts this pathway, resulting in therapeutic
resistance despite normal antibody pharmacokinetics.
It is noteworthy that not all antibody therapies
rely on FcγR signaling. In anaplastic large-cell lymphoma (ALCL),
the anti-CD30 monoclonal antibody HeFi-1 suppresses tumor growth
primarily by inducing G1-phase cell-cycle arrest,
thereby exerting its antitumor effect independently of FcγRIII
expression (67).
In general, these findings underscore a crucial
principle: For a broad range of antibody therapies targeting tumors
such as CRC and ATL, the FcγRI/III signaling pathway represents an
indispensable axis for antitumor efficacy. This pathway mediates
the key effector functions of myeloid cells, including macrophages
and neutrophils, in achieving therapeutic success.
Impact of drug intervention on the prognostic
value of FCER1G. The prognostic importance of FCER1G in
ccRCC appears to be treatment specific. It shows strong prognostic
value in patients treated with the anti-PD-1 antibody nivolumab,
but not in those receiving the mTOR inhibitor everolimus (43). This treatment-specific difference
may reflect the regulatory role of FCER1G in modulating
responses to immune checkpoint inhibitors such as nivolumab,
whereas its association with mTOR pathway-targeted therapies
appears to be relatively weak (41). Furthermore, pharmacological
interventions can dynamically influence the immunoregulatory
activity of FcRγ. For instance, peripheral blood NK cells from lung
transplant recipients treated with rapamycin (an mTOR inhibitor)
exhibit markedly reduced FcRγ expression. This observation suggests
that drug-induced modulation of FcRγ may indirectly affect immune
homeostasis and therapeutic outcomes (68).
6. Existing drugs indirectly regulating
FCER1G-related pathways
Despite the pivotal role of FCER1G in
multiple tumor immune microenvironments, no clinical drugs directly
targeting this molecule currently exist. Through an integrated
analysis of the DrugBank website and the DGIdb database, four
classes of drugs have been identified that indirectly influence
FCER1G-related pathways by regulating upstream and
downstream signaling molecules (Table
II). These include aspirin, benzylpenicilloyl polylysine,
omalizumab and fostamatinib. For instance, omalizumab functions as
an IgE monoclonal antibody. By blocking the binding of IgE to its
high-affinity receptor FcεRI, it is widely employed in treating
asthma and chronic spontaneous urticaria. Similarly, fostamatinib,
as a SYK inhibitor, targets SYK, the core downstream signaling
molecule of FcRγ. Although these drugs do not directly target
FCER1G, their mechanisms of action demonstrate that
intervention in pathways upstream (ligand-receptor binding) or
downstream (ITAM signaling) of FCER1G can produce distinct
biological effects.
 | Table IICandidate drugs that may indirectly
modulate the FCER1G-associated signaling pathway. |
Table II
Candidate drugs that may indirectly
modulate the FCER1G-associated signaling pathway.
| Potential
drugs | Drug category | Status | Primary mechanism
of action | Target |
|---|
| Aspirin | Nonsteroidal
anti-inflammatory drug | Approved | Non-selective COX
inhibitor | COX |
| Benzylpenicilloyl
polylysine | Diagnostic
reagent | Approved | FCER1A agonist | FCER1A |
| Omalizumab | Monoclonal
antibody | Approved | IgE inhibitor | IgE |
| Fostamatinib | Small-molecule
inhibitor | Approved | SYK inhibitor | SYK |
Although existing pharmacological agents offer
valuable insights into modulating the ITAM-SYK pathway activated by
FCRγ, directly targeting FCER1G for cancer therapy remains
considerably challenging. Therefore, a more feasible current
approach may be to leverage FCER1G not as a direct
therapeutic target, but as a key interpretative factor for
understanding tumor immune microenvironment heterogeneity and as a
biomarker for patient stratification. For instance, evaluating
FCER1G expression and its associated immune cell
infiltration patterns could help identify patients most likely to
benefit from antibody therapies or immune checkpoint inhibitors
that rely on FcγR effects. Future research should elucidate the
precise molecular switches determining whether FCER1G
function shifts towards a pro-tumor or antitumor role across
different tumor types. This is a prerequisite for its safe
translation into an effective therapeutic target.
7. Conclusion and outlook
As a core signaling adaptor, FCER1G mediates
activation signals for multiple immune receptors through its ITAM,
playing an indispensable role in both innate and adaptive immune
responses. The present review systematically summarizes the
multidimensional functions of FCER1G, spanning fundamental
immune regulation to its involvement in disease pathogenesis.
Beyond its classical roles in allergic reactions and anti-infection
immunity, FCER1G shows profound pathological importance in
inflammatory diseases and tumors.
In inflammation, FCER1G expression is finely
regulated by epigenetic mechanisms. Acting as a sensor of the
immune microenvironment, it dynamically modulates the magnitude and
outcome of inflammatory responses by influencing the polarization
of immune cells such as macrophages. Its functional plasticity
provides essential clues to its behavior within the more complex
TME.
FCER1G is highly expressed in several
malignancies, including ccRCC, STAD and GBM, and correlates
positively with immune checkpoint molecules such as PD-L1,
suggesting its role as a central immune regulatory hub. Its
tumor-promoting mechanisms are multifaceted, manifested in the
formation of an immunosuppressive milieu through M2 macrophage
recruitment and limited CD8+ T-cell infiltration (as in
STAD). FCER1G also mediates signaling crosstalk between FcγR
and EMT pathways, thus shaping tumor-immune interactions. Beyond
immunity, it participates in various non-immune regulatory
processes, including angiogenesis (via VEGF/MMP-9 in SCC),
platelet-mediated metastasis (via CXCL5/CXCL7 in MEL) and
cell-cycle progression (in ALCL).
Clinically, the impact of FCER1G is context
dependent. High expression predicts favorable prognosis in MM and
UCEC, but correlates with poor outcomes in ccRCC and STAD,
underscoring its tissue-specific prognostic importance.
Furthermore, antibody therapies that rely on FcRγ show pronounced
therapeutic efficacy in ATL and MEL models. Although several drugs
targeting FCER1G-related pathways have been identified, such
as aspirin and penicillanoyl polylysine (11), their translational value in
oncology remains to be thoroughly investigated.
Despite substantial progress has been achieved in
FCER1G research, several critical directions demand further
investigation. Future studies should first prioritize elucidating
the cross-cancer synergistic mechanisms between FCER1G and
immune checkpoints (such as PD-1/PD-L1) as well as other
immunoregulatory molecules, providing a conceptual basis for
rationally designed combination therapies. Particular attention
should be given to elucidating the molecular mechanisms underlying
FCER1G's dual roles in promoting or suppressing tumor
progression across distinct microenvironmental contexts.
Additionally, clinical translation faces notable challenges,
especially concerning the dose-dependent toxicity and limited
tissue selectivity of existing FCER1G-related drugs (such as
aspirin and penicillanoyl polylysine). Thus, well-designed,
multicenter clinical trials are urgently needed to validate
therapeutic efficacy and safety profiles of interventions targeting
FCER1G or its associated pathways (FcγR and SYK) in both
solid tumors and hematological malignancies.
Furthermore, optimizing therapeutic strategies
through tissue-targeted drug delivery represents a promising
avenue. Developing delivery systems capable of specifically
targeting FCER1G-expressing immune subsets or
tumor-associated stromal cells could improve treatment precision.
Such strategies may also help minimize off-target effects and
mitigate potential pro-metastatic risks such as unintended platelet
activation, which can arise from the broad distribution of
FcγRs.
In summary, as a multifunctional molecule with both
fundamental immunological importance and translational potential,
FCER1G represents a promising target for next-generation
cancer immunotherapy. Continuous elucidation of its complex
regulatory networks and development of innovative therapeutic
strategies are expected to further expand its clinical value and
ultimately improve patient outcomes.
Acknowledgements
Not applicable.
Funding
Funding: The present study was supported by the Science and
Technology Program of Gansu Province (grant no. 23JRRA1015).
Availability of data and materials
Not applicable.
Authors' contributions
YZ and JW wrote the original draft. JW, WH and TL
conceptualized the topic, reviewed and edited the review. All
authors read and approved the final manuscript. Data authentication
is not applicable.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Zhang X, Cai J, Song F and ang Z:
Prognostic and immunological role of FCER1G in pan-cancer. Pathol
Res Pract. 240(154174)2022.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Deng M, Du S, Hou H and Xiao J: Structural
insights into the high-affinity IgE receptor FcεRI complex. Nature.
633:952–959. 2024.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Wang Y, Tian W, Li R, Zhou D, Ding K, Feng
S, Ge Y, Luo Y, Chen Z and Hou H: Platelet FcRγ inhibits tumor
metastasis by preventing the colonization of circulating tumor
cells. Eur J Pharmacol. 990(177286)2025.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Wang W, Erbe AK, Hank JA, Morris ZS and
Sondel PM: NK cell-mediated antibody-dependent cellular
cytotoxicity in cancer immunotherapy. Front Immunol.
6(368)2015.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Vallières F and Girard D: Mechanism
involved in interleukin-21-induced phagocytosis in human monocytes
and macrophages. Clin Exp Immunol. 187:294–303. 2017.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Duhan V, Hamdan TA, Xu HC, Shinde P, Bhat
H, Li F, Al-Matary Y, Häussinger D, Bezgovsek J, Friedrich SK, et
al: NK cell-intrinsic FcεRIγ limits CD8+ T-cell expansion and
thereby turns an acute into a chronic viral infection. PLoS Pathog.
15(e1007797)2019.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Wang L, Lin Y, Yuan Y, Liu F and Sun K:
Identification of TYROBP and FCER1G as Key genes with prognostic
value in clear cell renal cell carcinoma by bioinformatics
analysis. Biochem Genet. 59:1278–1294. 2021.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Liu R, Liu J, Cao Q, Chu Y, Chi H, Zhang
J, Fu J, Zhang T, Fan L, Liang C, et al: Identification of crucial
genes through WGCNA in the progression of gastric cancer. J Cancer.
15:3284–3296. 2024.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Fu L, Cheng Z, Dong F, Quan L, Cui L, Liu
Y, Zeng T, Huang W, Chen J, Pang Y, et al: Enhanced expression of
FCER1G predicts positive prognosis in multiple myeloma. J Cancer.
11:1182–1194. 2020.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Chen Q, Wang S and Lang JH: Development
and validation of nomogram with tumor microenvironment-related
genes and clinical factors for predicting overall survival of
endometrial cancer. J Cancer. 12:3530–3538. 2021.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Jiang H, He Y, Lan X and Xie X:
Identification and validation of potential common biomarkers for
papillary thyroid carcinoma and Hashimoto's thyroiditis through
bioinformatics analysis and machine learning. Sci Rep.
14(15578)2024.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Kraft S and Kinet JP: New developments in
FcepsilonRI regulation, function and inhibition. Nat Rev Immunol.
7:365–378. 2007.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Hamerman JA, Ni M, Killebrew JR, Chu CL
and Lowell CA: The expanding roles of ITAM adapters FcRgamma and
DAP12 in myeloid cells. Immunol Rev. 232:42–58. 2009.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Novak N: New insights into the mechanism
and management of allergic diseases: atopic dermatitis. Allergy.
64:265–275. 2009.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Lieberman P and Garvey LH: Mast cells and
anaphylaxis. Curr Allergy Asthma Rep. 16(20)2016.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Pan YG, Yu YL, Lin CC, Lanier LL and Chu
CL: FcεRI γ-Chain negatively modulates dectin-1 responses in
dendritic cells. Front Immunol. 8(1424)2017.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Zhou ZW, Xu XT, Liang QN, Zhou YM, Hu WZ,
Liu S, Jiao YX, Zhang SC, Ji K and Chen JJ: USP5 deubiquitylates
and stabilizes FcεRIγ to enhance IgE-induced mast cell activation
and allergic inflammation. Sci Signal. 18(eadr3411)2025.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Saijo S, Ikeda S, Yamabe K, Kakuta S,
Ishigame H, Akitsu A, Fujikado N, Kusaka T, Kubo S, Chung SH, et
al: Dectin-2 recognition of alpha-mannans and induction of Th17
cell differentiation is essential for host defense against Candida
albicans. Immunity. 32:681–691. 2010.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Walsh NM, Wuthrich M, Wang H, Klein B and
Hull CM: Characterization of C-type lectins reveals an unexpectedly
limited interaction between Cryptococcus neoformans spores and
Dectin-1. PLoS One. 12(e0173866)2017.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Liu SD and Lowe JB: Implications of
understanding the signaling, cellular, and cytotoxic mechanisms
afforded by afucosylated antibodies. Oncoimmunology.
4(e1009288)2015.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Mallavia B, Oguiza A, Lopez-Franco O,
Recio C, Ortiz-Muñoz G, Lazaro I, Lopez-Parra V, Egido J and
Gomez-Guerrero C: Gene deficiency in activating Fcγ Receptors
influences the macrophage phenotypic balance and reduces
atherosclerosis in mice. PLoS One. 8(e66754)2013.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Huang C, Zhu W, Li Q, Lei Y, Chen X, Liu
S, Chen D, Zhong L, Gao F, Fu S, et al: Antibody Fc-receptor FcεR1γ
stabilizes cell surface receptors in group 3 innate lymphoid cells
and promotes anti-infection immunity. Nat Commun.
15(5981)2024.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Oishi S, Tsukiji N, Otake S, Oishi N,
Sasaki T, Shirai T, Yoshikawa Y, Takano K, Shinmori H, Inukai T, et
al: Heme activates platelets and exacerbates rhabdomyolysis-induced
acute kidney injury via CLEC-2 and GPVI/FcRγ. Blood Adv.
5:2017–2026. 2021.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Barrow AD, Raynal N, Andersen TL, Slatter
DA, Bihan D, Pugh N, Cella M, Kim T, Rho J, Negishi-Koga T, et al:
OSCAR is a collagen receptor that costimulates osteoclastogenesis
in DAP12-deficient humans and mice. J Clin Invest. 121:3505–3516.
2011.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Yabe R, Chung SH, Murayama MA, Kubo S,
Shimizu K, Akahori Y, Maruhashi T, Seno A, Kaifu T, Saijo S and
Iwakura Y: TARM1 contributes to development of arthritis by
activating dendritic cells through recognition of collagens. Nat
Commun. 12(94)2021.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Bashore FM, Katis VL, Du Y, Sikdar A, Wang
D, Bradshaw WJ, Rygiel KA, Leisner TM, Chalk R, Mishra S, et al:
Characterization of covalent inhibitors that disrupt the
interaction between the tandem SH2 domains of SYK and FCER1G
phospho-ITAM. PLoS One. 19(e0293548)2024.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Han S, Lan Q, Park AK, Lee KM, Park SK,
Ahn HS, Shin HY, Kang HJ, Koo HH, Seo JJ, et al: Polymorphisms in
innate immunity genes and risk of childhood leukemia. Hum Immunol.
71:727–730. 2010.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Liang Y, Wang P, Zhao M, Liang G, Yin H,
Zhang G, Wen H and Lu Q: Demethylation of the FCER1G promoter leads
to FcεRI overexpression on monocytes of patients with atopic
dermatitis. Allergy. 67:424–430. 2012.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Liang Y, Yu B, Chen J, Wu H, Xu Y, Yang B
and Lu Q: Thymic stromal lymphopoietin epigenetically upregulates
Fc receptor γ subunit-related receptors on antigen-presenting cells
and induces T(H)2/T(H)17 polarization through dectin-2. J Allergy
Clin Immunol. 144:1025–1035.e7. 2019.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Mahachie John JM, Baurecht H, Rodríguez E,
Naumann A, Wagenpfeil S, Klopp N, Mempel M, Novak N, Bieber T,
Wichmann HE, et al: Analysis of the high affinity IgE receptor
genes reveals epistatic effects of FCER1A variants on eczema risk.
Allergy. 65:875–882. 2010.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Rajaraman P, Brenner AV, Neta G, Pfeiffer
R, Wang SS, Yeager M, Thomas G, Fine HA, Linet MS, Rothman N, et
al: Risk of meningioma and common variation in genes related to
innate immunity. Cancer Epidemiol Biomarkers Prev. 19:1356–1361.
2010.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Chang R and Yang P: Molecular and
Mechanistic Study of Resistance to Glucocorticoid Combined with
Cyclosporine A Therapy in Female Patients with Vogt-Koyanagi-Harada
Syndrome. The 23rd International Ophthalmology Conference 2023.
Haikou, Hainan, China.
|
|
33
|
Podgórska D, Cieśla M and Kolarz B: FCER1G
gene hypomethylation in patients with rheumatoid arthritis. J Clin
Med. 11(4664)2022.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Yang M, Zheng H, Su Y, Xu K, Yuan Q,
Aihaiti Y, Cai Y and Xu P: Bioinformatics analysis identified the
Hub Genes, mRNA-miRNA-lncRNA axis, and signaling pathways involved
in rheumatoid arthritis pathogenesis. Int J Gen Med. 15:3879–3893.
2022.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Zhang K, Gao J and Ni Y: Screening of
candidate key genes associated with human osteosarcoma using
bioinformatics analysis. Oncol Lett. 14:2887–2893. 2017.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Liang Y, Zhao M, Liang G, Yin H and Lu Q:
Construction of special reporter to detect DNA methylation
regulatory activity in FCER1G gene promoter through
patch-methylation. Zhong Nan Da Xue Xue Bao Yi Xue Ban. 38:120–124.
2013.PubMed/NCBI View Article : Google Scholar : (In Chinese).
|
|
37
|
Liang Y: Molecular Mechanism of STAT5/TET2
Regulating DNA Methylation of FCER1G Promoter in Monocyte and its
role in the Pathogenesis of Atopic Dermatitis. Central South
University, 2013.
|
|
38
|
Wang J, Zhang L, Cui X, Xu X, Guo R, Li K,
Zhang L, Xu B, Jiang C and Yu Y: Bcl11a maintains hematopoietic
stem cell function but accelerates inflammation-driven exhaustion
during aging. Sci Immunol. 10(eadr2041)2025.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Wei ZM, Wang Z, Wan XJ, Li XJ, Li YX, Bai
Y, Yang X, Yang Y, Jiao SC and Liu ZF: FcRγ deficiency improves
survival in experimental sepsis by down-regulating TLR4 signaling
pathway. Immunol Res. 67:77–83. 2019.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Kottom TJ, Carmona EM, Schaefbauer K,
Stelzig KE, Pellegrino MR, Bindzus M and Limper AH: The importance
of Fcγ and C-type lectin receptors in host immune responses during
Pneumocystis pneumonia. Infect Immun. 93(e0027624)2025.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Yang R, Chen Z, Liang L, Ao S, Zhang J,
Chang Z, Wang Z, Zhou Y, Duan X and Deng T: Fc Fragment of IgE
Receptor Ig (FCER1G) acts as a key gene involved in cancer immune
infiltration and tumour microenvironment. Immunology. 168:302–319.
2023.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Peng W, Zhao Y, Yang N, Fang Y, Wu Y, Feng
Z, Wu Q and Wang X: Prognostic value of FCER1G expression and M2
macrophage infiltration in esophageal squamous cell carcinoma.
Discov Oncol. 16(113)2025.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Dong K, Chen W, Pan X, Wang H, Sun Y, Qian
C, Chen W, Wang C, Yang F and Cui X: FCER1G positively relates to
macrophage infiltration in clear cell renal cell carcinoma and
contributes to unfavorable prognosis by regulating tumor immunity.
BMC Cancer. 22(140)2022.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Chen L, Yuan L, Wang Y, Wang G, Zhu Y, Cao
R, Qian G, Xie C, Liu X, Xiao Y and Wang X: Co-expression network
analysis identified FCER1G in association with progression and
prognosis in human clear cell renal cell carcinoma. Int J Biol Sci.
13:1361–1372. 2017.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Ding W, Jiang H, Ye N, Zhuang L, Yuan Z,
Tan Y, Xue W and Xu X: Identification and analysis of crucial genes
in H. Pylori-Associated gastric cancer using an integrated
bioinformatics approach. J Oncol. 2023(8538240)2023.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Xu H, Zhang A and Lou M: The role of
FCER1G gene in evaluating the prognosis of glioma. Chin J Minim
Invasive Neurosurg. 26:363–366. 2021.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Xu H, Zhu Q, Tang L, Jiang J, Yuan H,
Zhang A and Lou M: Prognostic and predictive value of FCER1G in
glioma outcomes and response to immunotherapy. Cancer Cell Int.
21(103)2021.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Tian Z, Meng L, Long X, Diao T, Hu M, Wang
M, Liu M and Wang J: Identification and validation of an
immune-related gene-based prognostic index for bladder cancer. Am J
Transl Res. 12:5188–5204. 2020.PubMed/NCBI
|
|
49
|
Xiang X: Identification, expression and
prognostic significance of FCER1G gene related to CD4+ T lymphocyte
infiltration in Diffuse Large B-cell Lymphoma. CNKI: Sichuan
University, 2023.
|
|
50
|
Li B, Cai Z, Zhang Y, Chen R, Tang S, Kong
F, Li W, Ding L, Chen L and Xu H: Biomarkers associated with
papillary thyroid carcinoma and Hashimoto's thyroiditis:
Bioinformatic analysis and experimental validation. Int
Immunopharmacol. 143(113532)2024.PubMed/NCBI View Article : Google Scholar
|
|
51
|
Xiang X, Gao LM, Zhang Y, Tang Y, Zhao S,
Liu W, Ye Y and Zhang W: Identification of FCER1G related to
Activated Memory CD4(+) T Cells Infiltration by Gene Co-expression
Network and Construction of a Risk Prediction Module in Diffuse
Large B-Cell Lymphoma. Front Genet. 13(849422)2022.PubMed/NCBI View Article : Google Scholar
|
|
52
|
Li J, Shi H, Yuan Z, Wu Z, Li H, Liu Y and
Lu M and Lu M: The role of SPI1-TYROBP-FCER1G network in
oncogenesis and prognosis of osteosarcoma, and its association with
immune infiltration. BMC Cancer. 22(108)2022.PubMed/NCBI View Article : Google Scholar
|
|
53
|
Qiu X, Zhang JH, Xu Y, Cao YX, Zhang RT,
Hu LN and Zhou JH: Identification of FCER1G as a key gene in
multiple myeloma based on weighted gene co-expression network
analysis. Hematology. 28(2210904)2023.PubMed/NCBI View Article : Google Scholar
|
|
54
|
Bigley AB, Spade S, Agha NH, Biswas S,
Tang S, Malik MH, Dai L, Masoumi S, Patiño-Escobar B, Hale M, et
al: FcεRIγ-negative NK cells persist in vivo and enhance efficacy
of therapeutic monoclonal antibodies in multiple myeloma. Blood
Adv. 5:3021–3031. 2021.PubMed/NCBI View Article : Google Scholar
|
|
55
|
Shukla M and Sarkar RR: Differential
cellular communication in tumor immune microenvironment during
early and advanced stages of lung adenocarcinoma. Mol Genet
Genomics. 299(100)2024.PubMed/NCBI View Article : Google Scholar
|
|
56
|
A Self-reactive Innate-like T-cell program
was identified in cancer immunity. Cancer Discov.
12(Of11)2022.PubMed/NCBI View Article : Google Scholar
|
|
57
|
Morrish E and Ruland J: Cytotoxic
FCER1G(+) innate-like T cells: New potential for tumour
immunotherapy. Signal Transduct Target Ther. 7(204)2022.PubMed/NCBI View Article : Google Scholar
|
|
58
|
Gunderson AJ, Kaneda MM, Tsujikawa T,
Nguyen AV, Affara NI, Ruffell B, Gorjestani S, Liudahl SM, Truitt
M, Olson P, et al: Bruton tyrosine kinase-dependent immune cell
cross-talk drives pancreas cancer. Cancer Discov. 6:270–285.
2016.PubMed/NCBI View Article : Google Scholar
|
|
59
|
Andreu P, Johansson M, Affara NI, Pucci F,
Tan T, Junankar S, Korets L, Lam J, Tawfik D, DeNardo DG, et al:
FcRgamma activation regulates inflammation-associated squamous
carcinogenesis. Cancer Cell. 17:121–134. 2010.PubMed/NCBI View Article : Google Scholar
|
|
60
|
Kong D, Zhang Y, Jiang L, Long N, Wang C
and Qiu M: Comprehensive analysis reveals the tumor suppressor role
of macrophage signature gene FCER1G in hepatocellular carcinoma.
Sci Rep. 15(3995)2025.PubMed/NCBI View Article : Google Scholar
|
|
61
|
Clynes R, Takechi Y, Moroi Y, Houghton A
and Ravetch JV: Fc receptors are required in passive and active
immunity to melanoma. Proc Natl Acad Sci USA. 95:652–656.
1998.PubMed/NCBI View Article : Google Scholar
|
|
62
|
Domínguez-Soto A, de las Casas-Engel M,
Bragado R, Medina-Echeverz J, Aragoneses-Fenoll L, Martín-Gayo E,
van Rooijen N, Berraondo P, Toribio ML, Moro MA, et al: Intravenous
immunoglobulin promotes antitumor responses by modulating
macrophage polarization. J Immunol. 193:5181–5189. 2014.PubMed/NCBI View Article : Google Scholar
|
|
63
|
Duhan V, Gomez MR, Le TT, Rana S, Chen YE,
Balakrishnan D, Kelly G, Johnston RL, Krebs P and Khanna R:
FcRγ-dependent NK cell licensing through CD244 promotes antitumour
immunity. Cancer Immunol Res. 13:2075–2092. 2025.PubMed/NCBI View Article : Google Scholar
|
|
64
|
Chan RH, Chen PC, Yeh YM, Lin BW, Yang KD,
Shen MR and Lin PC: The expression quantitative trait loci in
immune response genes impact the characteristics and survival of
colorectal cancer. Diagnostics (Basel). 12(315)2022.PubMed/NCBI View Article : Google Scholar
|
|
65
|
Zhang M, Zhang Z, Garmestani K, Goldman
CK, Ravetch JV, Brechbiel MW, Carrasquillo JA and Waldmann TA:
Activating Fc receptors are required for antitumor efficacy of the
antibodies directed toward CD25 in a murine model of adult t-cell
leukemia. Cancer Res. 64:5825–5829. 2004.PubMed/NCBI View Article : Google Scholar
|
|
66
|
Zhang Z, Zhang M, Ravetch JV, Goldman C
and Waldmann TA: Effective therapy for a murine model of adult
T-cell leukemia with the humanized anti-CD2 monoclonal antibody,
MEDI-507. Blood. 102:284–288. 2003.PubMed/NCBI View Article : Google Scholar
|
|
67
|
Zhang M, Yao Z, Zhang Z, Garmestani K,
Goldman CK, Ravetch JV, Janik J, Brechbiel MW and Waldmann TA:
Effective therapy for a murine model of human anaplastic large-cell
lymphoma with the anti-CD30 monoclonal antibody, HeFi-1, does not
require activating Fc receptors. Blood. 108:705–710.
2006.PubMed/NCBI View Article : Google Scholar
|
|
68
|
Shemesh A, Su Y, Calabrese DR, Chen D,
Arakawa-Hoyt J, Roybal KT, Heath JR, Greenland JR and Lanier LL:
Diminished cell proliferation promotes natural killer cell
adaptive-like phenotype by limiting FcεRIγ expression. J Exp Med.
219(e20220551)2022.PubMed/NCBI View Article : Google Scholar
|















![Protein structure of FCER1G and FcεRI
schematic structure. (A) Protein structure of FCER1G
[source: GeneCards database (https://www.genecards.org/)]. In the figure,
blackish-brown represents the FcRγ subunit; bronze-gold represents
the FcεRIβ subunit; green represents the FcεRIα subunit; brown
represents the immunoglobulin heavy constant epsilon; and pink and
bluish-purple represent the immunoglobulin kappa region. (B)
Structural diagram of the FcεRI receptor. The FcεRIα chain binds
individual IgE molecules, thereby mediating for IgE recognition.
The FcεRIβ chain facilitates the maturation and trafficking of
FcεRIα and stabilizes the FcεRI complex on the cell surface. FcRγ
contains an ITAM motif; two molecules of FcRγ assemble with FcεRIα
and FcεRIβ chains to form a complete tetrameric high-affinity IgE
receptor. The figure was drawn with FigDraw2.0 (supplied by
Hangzhou Sifei Technology Co., Ltd.). FCER1G, Fc fragment of IgE
receptor 1G; FcεRI, Fc ε receptor I; ITAM, immunoreceptor
tyrosine-based activation motif.](/article_images/etm/31/5/etm-31-05-13141-g00.jpg)