Introduction
The course of the eukaryotic cell cycle is regulated
by genes and accompanied by periodic fluctuations in the expression
levels of a number of genes (1).
Maintaining cell cycle homeostasis requires coordinated
interactions between proteins such as tubulin (2), actin (3) and cellular structures. Cell cycle
dysregulation leads to genomic instability which leads to cell
death, diseases and malignancies, such as breast cancer (4), esophageal cancer (5), and head and neck squamous cell
carcinoma (6,7). Schizosaccharomyces pombe
(S. pombe) is a eukaryotic organism with well-defined cell
shapes and highly active homologous recombination mechanisms,
making it a high-quality model organism in biology (8,9). The
growth of fission yeast cells is susceptible to environmental
changes; in conditions such as high temperatures or nutritional
deficiencies, cells trigger a stress response that regulates cell
cycle checkpoints to delay or arrest the mitotic process,
disrupting cell cycle homeostasis (10). For example, when fission yeast
cells are cultured at a suitable temperature (such as 25˚C),
protein function in the cells is normal; however, when cultured at
higher temperatures (such as 37˚C), protein function during mitosis
is disturbed and cell cycle homeostasis is destroyed (11,12).
Almost all aspects of eukaryotic gene expression and
its regulation involve ATP-dependent RNA helicases. The DEAD box
protein family of RNA helicases is a key regulator of RNA splicing,
mRNA export and ribosome biosynthesis. Mutations and dysregulation
of the DEAD box protein family have been associated with a variety
of diseases, such as cancer and neurological disorders (13). The ATP-dependent RNA helicase Fal1
of the S. pombe DEAD box protein family is involved in the
constitution of the exon junction complex subunit that recognizes
and degrades mis-spliced mRNAs in the nucleus during the cell
cycle. The fal1 gene deletion results in the formation of
ascus with more or fewer than four spores and abnormal mitosis
(14,15). Ste13 has ATP-dependent histone
chaperone activity that regulates the involvement of histone
chaperones in chromatin assembly and de-assembly. The ste13
mutant cannot enter the G0 phase of the cell cycle or
initiate sexual reproduction to produce spores (16). Dbp2 has RNA helicase activity and
binds ATP and mRNA; it is involved in the composition of the
ribonucleoprotein complex and regulates ribosome biogenesis.
Deletion of the dbp2 gene results in reduced localization of
proteins to chromatin and mitotic cell cycle abnormalities
(15). Translation initiation RNA
helicase, Ded1, has translation initiation factor activity. In
response to stress, Ded1 inhibits the activity of the Cdc25 protein
in mitosis and activates the mitotic G2/M checkpoint;
however, under high temperature conditions, ded1 mutant
cells are unable to enter S phase and mitotic processes (11,17).
Prp19 complex WD repeat protein Prp5, which is involved in the
composition of the Prp19 complex, regulates DNA repair,
recombination and spore formation, but at high temperatures,
prp5 mutations result in abnormal cell morphology, defective
cell cycle proteins and cell cycle arrest at mitosis or the
G2/M border (18,19).
The uap56 gene encodes the TREX complex subunit,
ATP-dependent RNA Uap56, which is involved in the mRNA metabolic
process by catalyzing the attachment of RNA fragments through the
spliceosome. Under high temperature conditions, the mitotic cell
cycle of the uap56Δ strain is abnormal (15).
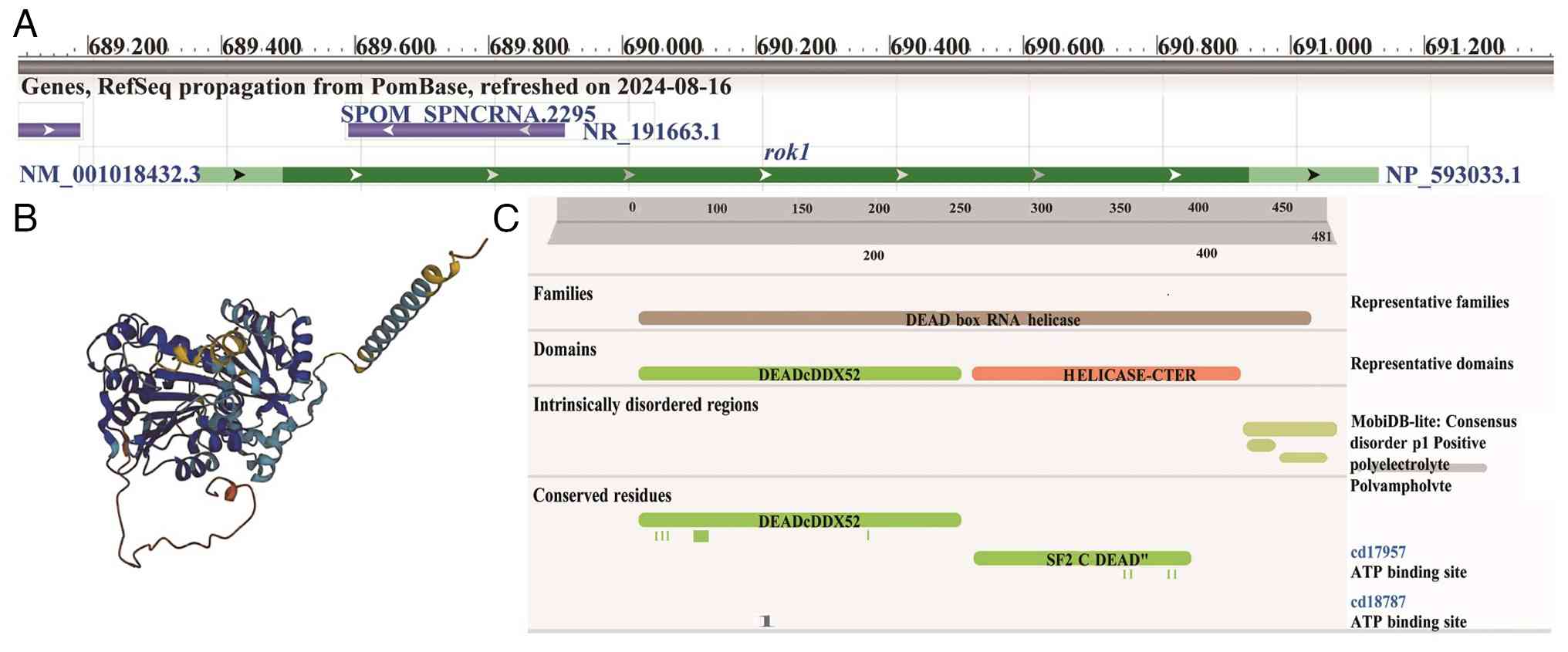
The rok1 gene encodes the ATP-dependent RNA
helicase Rok1 in S. pombe, which is a member of the DEAD box
protein family (Fig. 1). In cells,
the Rok1 protein is localized in the nucleolus and is involved in
regulating the maturation of small subunit ribosomal RNA (rRNA) and
thus the process of ribosome biogenesis (20). In budding yeast, the Rok1 protein
binds to ATP, stabilizes the binding of its cofactor Rrp5 to the
40S ribosome (21) and is involved
in rRNA processing and control of cell cycle progression in budding
yeast (22). In fission yeast,
rok1 gene deletion results in decreased cell growth under
high-temperature conditions and in media with galactose, maltose,
sucrose or xylose as the carbon source, increased growth in media
with lysine, proline or serine as the nitrogen source and increased
susceptibility to bleomycin, brefeldin A, cycloheximide, diamide,
formamide and sodium dodecyl sulfate. Abnormal chromosome
segregation during meiosis was also observed in rok1∆ cells
(23). In addition, the
rok1 gene is closely associated with human diseases, and
Rok1 and Cd168 accelerate the progression of androgen-independent
(AI) prostate cancer and accelerate the invasion and metastasis of
AI cells in the endothelial layer of human bone marrow (24).
To the best of our knowledge, the effects of fission
yeast rok1 gene deletion on cell mitosis under normal and
stress temperature conditions and its molecular mechanisms have not
been reported. In the present study, the effects of rok1
gene deletion on cell growth, sporulation and spindle, actin,
kinetochores and centromere protein dynamics in mitosis were
investigated using the fission yeast as a research model.
RNA-sequencing (RNA-Seq) and bioinformatics analyses also revealed
key genes and pathways of the abnormal growth of the rok1Δ
strain, providing a scientific basis for further revealing the role
of Rok1 protein in cell division.
Materials and methods
Experimental strains
The strains used in the present study are listed in
Table I (25,26).
All strains constructed in this experiment were engineered with
fluorescent labels via spore production, as later described.
 | Table IStrains and genotypes used in the
present study. |
Table I
Strains and genotypes used in the
present study.
| Strain | Genotype | Source |
|---|
| PT286 | h-
wt | Lab of
Trana |
| PT287 | h+
wt | Lab of
Trana |
| PT2514 | h-
Mis12-GFP:Leu/mC-Atb2:HygR | Lab of
Trana |
| PT3850 | h+
Pact1-LifeAct-mGFP:LEU1 | Lab of
Trana |
| YL20 | h-
Sid4-GFP:NatMX/mC-Atb:HygR | Ding Zhang
(25) |
| YL24 | h-
mC-Atb2:HygR/Cut11-GFP :HygR | Ding Zhang
(25) |
| YL26 | hb GFP-Atb2:HygR/Hht1-RFP:HygR | Yu et al
(26) |
| 2125-A | h+
rok1∆:kanR | Lab of
Trana |
| 2125-B | h-
rok1Δ:kanR | Lab of
Trana |
| 2125-1 | hb
rok1Δ:KanR/Mis12-GFP:Leu/mC-Atb2:HygR | Present study |
| 2125-2 | hb
rok1Δ:KanR/Pact1-LifeAct-mGFP:LEU1 | Present study |
| 2125-3 | hb
rok1Δ:KanR/Sid4-GFP:NatMX/mC-Atb:HygR | Present study |
| 2125-4 | hb
rok1Δ:KanR/mC-Atb2:HygR/Cut11-GFP :HygR | Present study |
| 2125-5 | hb
rok1Δ:KanR/GFP-Atb2:HygR/Hht1-RFP:HygR | Present study |
Culture media and main reagents
YE5S Liquid Medium was prepared by adding 2.5 g
Yeast Extract (Thermo Fisher Scientific, Inc.), 0.01125 g Amino
Acids (Ade/Leu/Ura/His/Lys) (Merck KGaA), 15 g Dextrose (Shanghai
Aladdin Biochemical Technology Co., Ltd.) and 500 ml ultrapure
Water. YE5S Solid Medium was prepared by adding 8.5 g agar
(BioFroxx; neoFroxx GmbH) to the liquid medium. G418, HygR or NatMx
Resistance Selection Medium were prepared by adding 0.3 mg/ml G418
(Guangzhou Saigou Biotech Co., Ltd.), 0.3 mg/ml HygR (Shanghai
Macklin Biochemical Co., Ltd.) or 0.1 mg/ml NatMx (Beijing Solarbio
Science & Technology Co., Ltd.) to YE5S solid medium.
Leu-deficient medium was prepared by removing Leu from YE5S solid
medium. 1X snail enzyme solution was prepared by mixing 100X snail
enzyme buffer with 10 g/ml snail enzyme (Shanghai Yuanye
Bio-Technology Co., Ltd.) and 990 µl ultrapure water. The 100X
Snail Enzyme Buffer contains 1 mol/l sorbitol (Shanghai Yuanye
Bio-Technology Co., Ltd.), 100 mmol/l EDTA (Shanghai Macklin
Biochemical Co., Ltd.) and 14 mmol/l β-mercaptoethanol (Merck
KGaA). All media and reagents were prepared at room temperature and
sterilized prior to use.
Fluorescent protein labeling
construction
The wild-type strain with fluorescent marker
(PT2514, PT3850, YL20, YL24, YL26) and the opposite mating-type
rok1Δ strain (2125-A or 2125-B) (Table I) were inoculated onto YE5S solid
medium and were activated at 25˚C for 3 days. The activated
h+ and h- strains were then
mixed on EMM-N nitrogen-deficient medium for sporulation (27) and incubated at 25˚C for 2 days.
Colonies were picked and examined using an OLYMPUS BX51 microscope
(Olympus Corporation) to monitor sporulation. Upon confirmation of
spore formation, the cells were treated with 1 ml prepared 1X snail
enzyme working solution (28,29)
to release the spore suspension. The suspension was spread onto
YE5S solid medium and incubated at 25˚C for 2 days to allow spore
germination and single colony formation. Colonies were subsequently
transferred using the replica plating method onto
nutrient-deficient or antibiotic-containing media (30). Positive clones obtained were
expanded through serial passages, stored with 30% glycerol as
cryoprotectant in an 80˚C ultra-low temperature freezer (Thermo
Fisher Scientific, Inc.) for future use.
Live cell imaging
Live cell imaging of the experimental strains was
performed using a TCS-SP8 laser confocal microscope (Leica
Microsystems GmbH) at 25 and 37˚C. The parameters were set as
follows: Green fluorescence received in the wavelength range of
493-545 nm, red fluorescence received in the wavelength range of
584-736 nm, pixels of 512x512 µm, seven optical slices with a pitch
of 6.02 µm, exposure time of 400 msec, shooting interval of 2 min
and total shooting time of 120 min.
Outlier detection
Measurements of the dynamics of cellular mitotic
spindle, actin, kinetochore, and centrosome proteins were analyzed
using ImageJ (version 1.51s; National Institutes of Health).
Potential outliers were identified and excluded based on the
Z-score method. The Z-score was calculated as Z=(χ-µ)/σ, where χ is
the measured value, µ is the mean of measurements and σ is the
population standard deviation. Data points with Z>2 were
considered outliers (31). To
ensure data consistency across all metrics including microtubule,
actin, kinetochore and centrosomal protein data, each initially
collected from 20 cells (No. 1-No. 20), if a cell was identified as
an outlier in any metric, all measurement data from that cell were
excluded.
For example, using actin ring data from rok1Δ
strains cultured at 25˚C, six metrics were analyzed: Actin ring
length, total time from assembly to complete disappearance,
assembly time, contraction time, total contraction rate from
assembly to complete disappearance and contraction rate during the
contraction phase. After Z-score normalization of measurements from
the 20 cells, three outliers (Z>2) were identified: The
contraction rate of the No. 4 cell and both the total duration and
assembly time of the actin ring of the No. 14 cell. Consequently,
all six parameters from the No. 4 and the No. 14 cells were
excluded. Following this procedure, data from 5 cells were excluded
for each metric, ultimately retaining a uniform set of 15 cells per
metric for subsequent analysis.
Statistical analysis
Data analysis was performed using SPSS Statistics 26
(International Business Machines Corporation). The Shapiro-Wilk
test assessed data normality, and the unpaired Student's t-test
analyzed differences between wild-type and rokI∆ strains.
Differentially expressed genes were identified using an adjusted
P<0.05, log2 fold change >0(32) as screening criteria. RT-qPCR data
were analyzed using the 2-∆∆Cq relative quantification
method (33) to describe the
transcriptional expression changes of target genes in the
rok1Δ strain relative to the wild-type strain. P<0.05 was
considered to indicate a statistically significant difference.
RNA-Seq
The PT287 and 2125 strains were cultured at 25˚C and
37˚C until the cells entered the logarithmic growth phase, and then
the organisms were collected and frozen. Total RNA was extracted
using the Yeast total RNA kit (Omega Bio-Tek, Inc.) according to
the manufacturer's standard protocol, and sent to Beijing Novogene
technology Co., Ltd. for quality testing and RNA-Seq. Reference
genes were compared using Hisat2 v2.0.5 software (34); gene expression was quantified using
Feature Counts and Stringtie (1.3.3b) software (35); and differential significance of
gene expression of samples was analyzed using DESeq2 (1.20.0) and
EdgeR (3.22.5) software (36).
Differential genes were processed by Gene Ontology (GO) function
enrichment (37), Kyoto
Encyclopedia of Genes and Genomes (KEGG) (38) pathway enrichment analysis was
performed using Cluster Profile software (version 3.8.1) (39).
Reverse transcription-quantitative PCR
(RT-qPCR)
PT287 and 2125 strains were cultured at 25 and 37˚C
until the cells reached the logarithmic growth phase, then
collected and frozen. Total RNA was extracted using the Yeast Total
RNA Kit and reverse-transcribed into cDNA using the RevertAid First
Strand cDNA Synthesis Kit (Thermo Fisher Scientific, Inc.) at 42˚C
for 60 min. The qPCR conditions were as follows: 95˚C
pre-denaturation for 30 sec, followed by 39 cycles of 95˚C
denaturation for 5 sec and 60˚C annealing/extension for 30 sec. The
melting curve analysis stage consisted of 95˚C for 10 sec, 65˚C for
5 sec and 95˚C for 5 sec. The act1 gene was used as the
internal control, and qPCR was performed using the CFX96 Real-Time
PCR Detection System (Bio-Rad Laboratories, Inc.) to analyze gene
expression levels. The primer sequences used are listed in Table II.
 | Table IIPrimer sequences used for
quantitative PCR. |
Table II
Primer sequences used for
quantitative PCR.
| Gene | Forward primer
(5'→3') | Reverse primer
(5'→3') |
|---|
| psc3 |
CGTTCAGCCTCAAGAACGGA |
AATTGCGATACGAGCTAGGC |
| psm1 |
ATCAACGCTGAGTTACGCCA |
CTAACTGCGCAGAGTCTCGT |
| myo51 |
TCGGTAGGCTCGGAATGTTG |
AAGGCGACTCTGATTGACCG |
| blt1 |
TGACGGTTCATCCATCGTTT |
ACTGGGCGTTTTCGTCTTCT |
| act1 |
CCCAAATCCAACCGTGAGAAG |
CCAGAGTCCAAGACGATACCAGTG |
Results
Changes in cell growth and spore
production in the rok1Δ strain
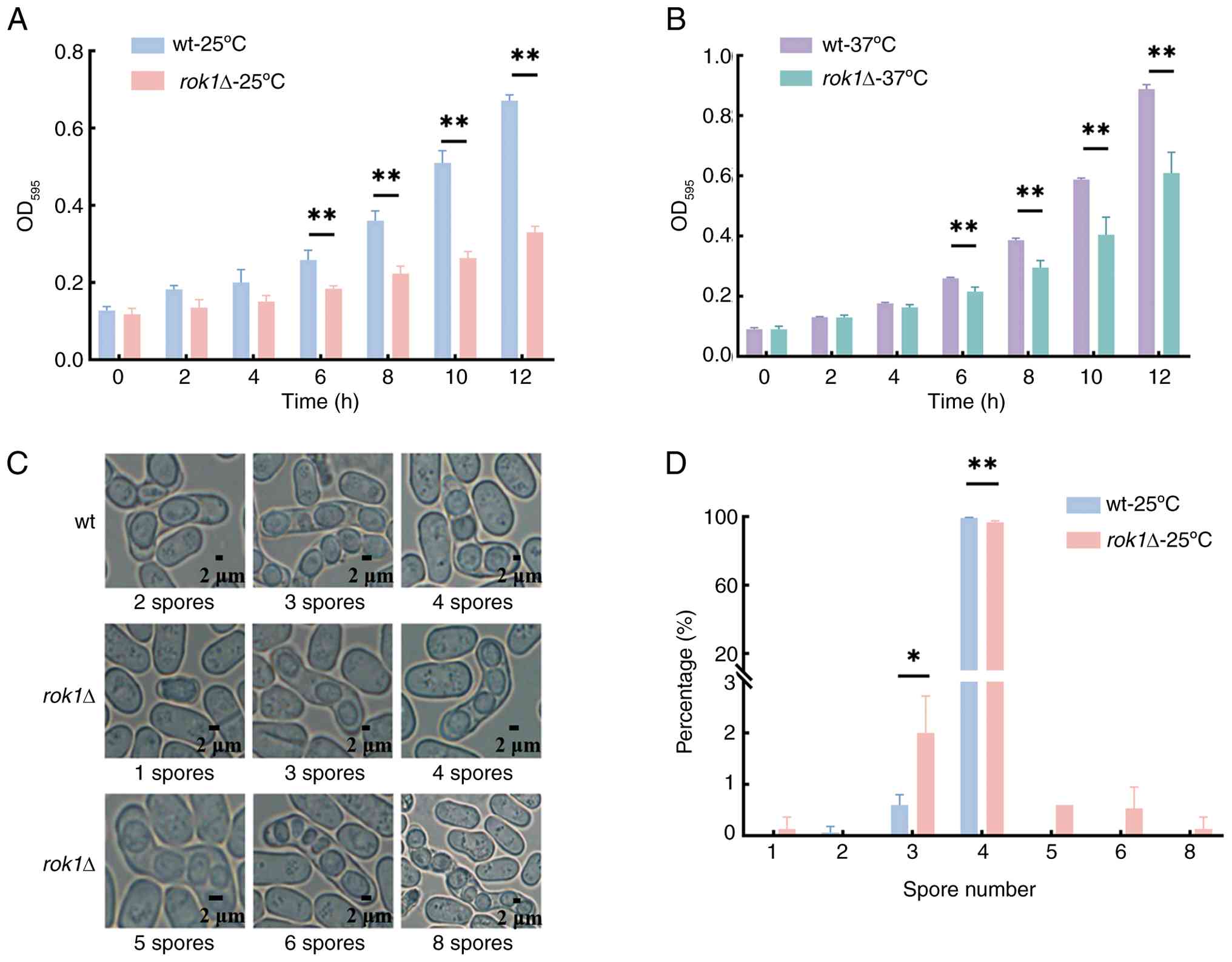
The growth rates of the wild-type strain and the
rok1Δ strain were compared and the results showed that there
was no significant difference in growth rates between the two over
the period of 0-4 h at 25˚C and 37˚C (all P>0.05). After 6 h,
the wild-type strain entered the logarithmic growth phase and the
growth rate of the rok1Δ strain significantly decreased. At
25˚C, optical density (OD)595 at 12 h was 0.68±0.00 and
0.33±0.02 for the wild-type and rok1Δ strains, respectively.
At 37˚C, OD595 at 12 h was 0.89±0.01 and 0.61±0.07 for
the wild-type and rok1Δ strains, respectively (Fig. 2A and B), which were significantly different
(P<0.01). The aforementioned results indicated that rok1
gene deletion significantly decreased the cell growth rate.
The results of spore number production showed that
99.33±0.31% of the wild-type strains produced 4 spores, whereas the
percentage of the rok1Δ strain producing 4 spores was
96.60±0.92%, which was a significant difference. In addition, there
were 2.00±0.72% of the rok1Δ strain producing 3 spores
(Fig. 2C and D). The results showed that the
rok1Δ strain produced an abnormal number of spores compared
to the wild-type strain, with a significant decrease in the number
of spores. It is noteworthy that the high temperature of 37˚C
significantly impacts spore production. Previous studies have
reported that at 37˚C, gene-deleted fission yeast either fail to
produce spores or exhibit a low probability of spore formation
(40). Therefore, spore production
was investigated exclusively at 25˚C.
A significant decrease in the growth rate of the
rok1Δ strain was found by growth rate measurements;
therefore, the mitosis-related protein dynamics of the rok1Δ
strain was investigated.
Changes of spindle during mitosis of
rok1Δ strain under different temperature conditions
During mitosis, microtubule proteins assemble to
form the spindle, which is responsible for the accurate segregation
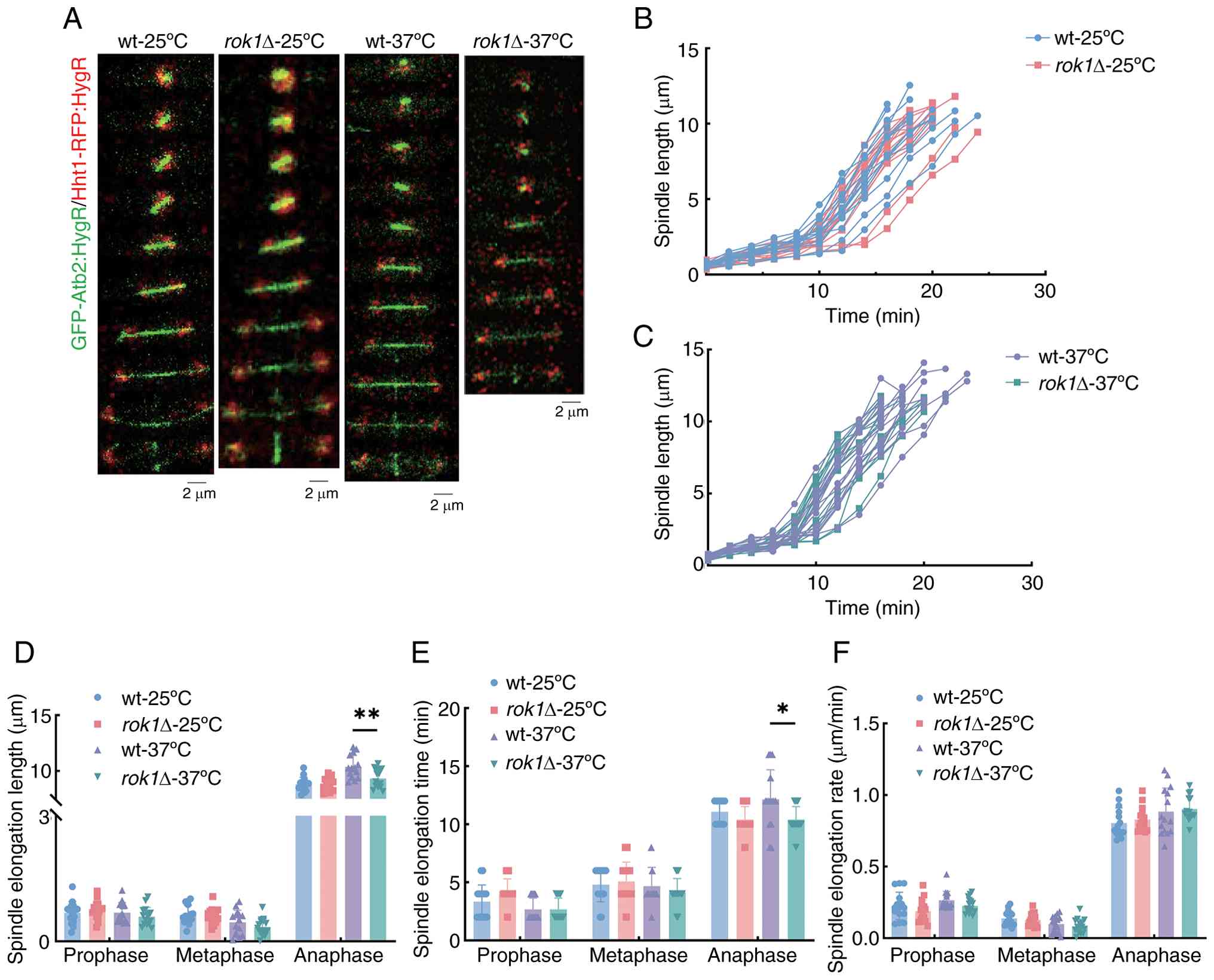
of chromosomes during cell division (41). Live-cell imaging of wild-type and
rok1Δ cells with green fluorescent protein (GFP)-Atb2-tagged
microtubule proteins was used to assess spindle properties. The
analysis of spindle dynamics at 25˚C showed that the spindle
elongation lengths of the wild-type strains at the prophase,
metaphase and anaphase were 0.68±0.27, 0.63±0.23 and 8.82±0.62 µm
and for the rok1Δ strain they were 0.75±0.22, 0.61±0.19 and
8.57±0.80 µm, respectively (Fig.
3A-D). The rok1Δ strain exhibited no significant changes
in spindle elongation length compared to the wild-type strain in
all phases (all P>0.05). Further analysis revealed (Fig. 3E and F) that the spindle elongation times of
the wild-type strains at prophase, metaphase and anaphase were
3.33±1.45, 4.80±1.47 and 11.07±1.03 min and for the rok1Δ
strain they were 4.27±1.83, 5.07±1.67 and 10.40±1.12 min,
respectively. The spindle elongation rates of the wild-type and
rok1Δ strains at the prophase, metaphase and anaphase were
0.22±0.10, 0.14±0.05 and 0.80±0.10 µm/min and 0.19±0.08, 0.13±0.04
and 0.83±0.09 µm/min, respectively. There was no significant
difference in the elongation time and elongation rate of the
spindle in all periods between the wild-type and rok1Δ
strains (all P>0.05). These results indicate that the deletion
of the rok1 gene does not affect spindle dynamics at
25˚C.
The spindle elongation lengths at 37˚C showed that
compared with the wild-type there was a significant decrease in the
spindle elongation length of the rok1Δ strains at anaphase
(10.38±1.02 vs. 9.33±0.83 µm, respectively); however, there were no
significant changes in spindle elongation length at prophase or
metaphase. Further analysis revealed that this trend was consistent
for the spindle elongation time with wild-type strains displaying
significantly longer elongation times compare with rok1Δ
strains at anaphase (12.13±2.56 vs. 10.40±1.12 min, respectively),
with no significant changes observed at prophase and metaphase
(P>0.05) (Fig. 3E). In
addition, there was no significant difference in spindle elongation
rates between the wild-type and rok1Δ strains at prophase
(0.26±0.06 vs. 0.23±0.05 µm/min, respectively), metaphase
(0.10±0.06 vs. 0.08±0.05 µm/min, respectively) and anaphase
(0.88±0.17 vs. 0.90±0.09 µm/min), respectively (both P>0.05)
(Fig. 3F).
These results indicate that the rok1 gene
deletion decelerated the spindle elongation process at 37˚C.
Comparison of 25˚C and 37˚C revealed that a high temperature stress
of 37˚C had an inhibitory effect on the spindle elongation
process.
Changes of actin during mitosis of
rok1Δ strain under different temperature conditions
During mitosis, actin filaments co-assemble with
myosin II to form an actomyosin ring that help the cell complete
cytoplasmic division (42).
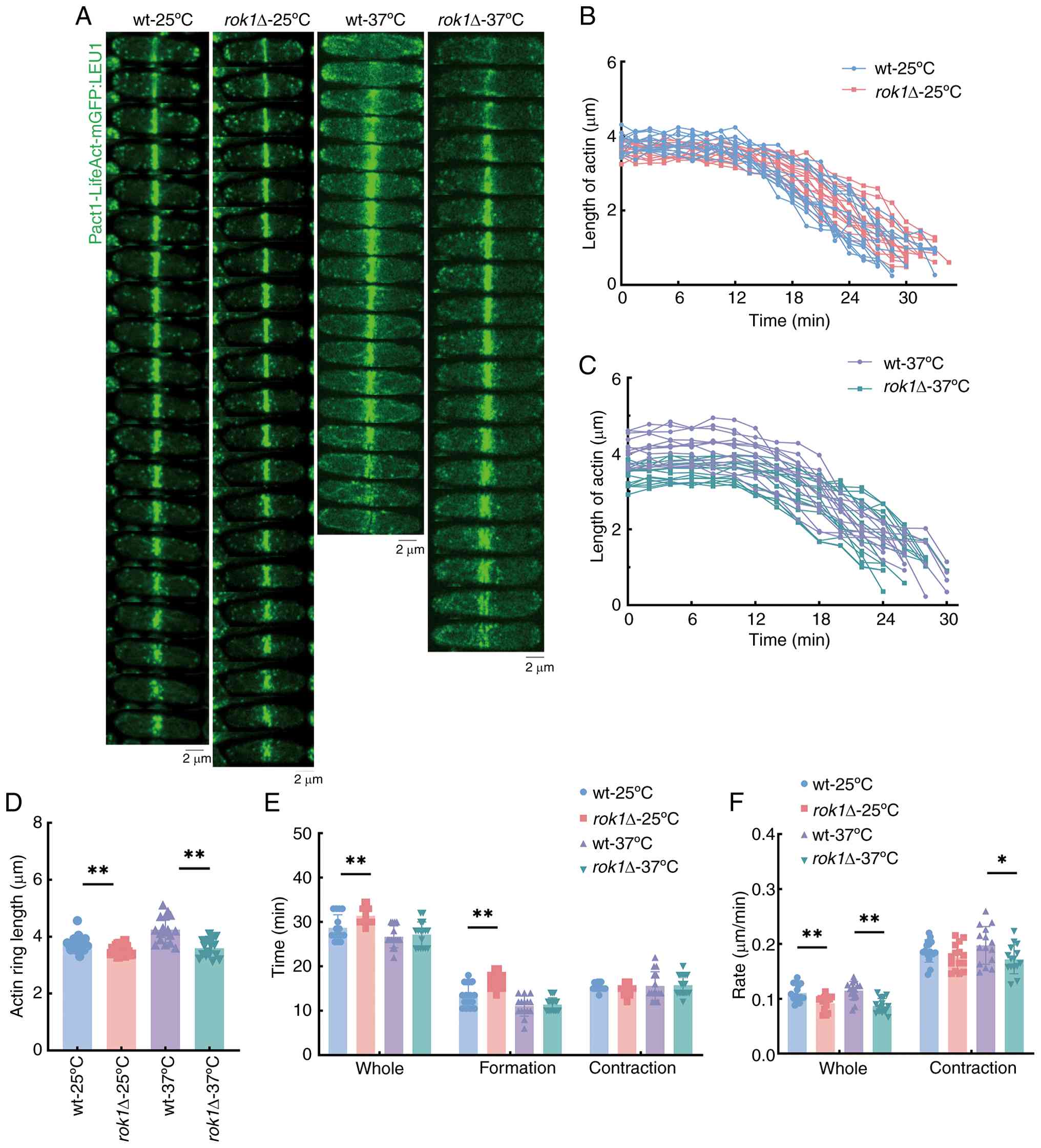
Live-cell imaging was performed of wild-type and rok1Δ cells
with Pact1-LifeAct-mGFP-tagged actin. Analysis of actin rings at
25˚C indicated that the actin ring length was significantly
decreased (3.75±0.17 vs. 3.52±0.15 µm), total time from assembly to
complete disappearance was significantly increased (28.70±2.84 vs.
31.40±1.29 min) and total contraction rates were significantly
decreased (0.11±0.02 vs. 0.09±0.01 µm/min), in the rok1Δ
strain compared with the wild-type stain (Fig. 4A-F). Further analysis results
indicated that the assembly times of the actin rings in the
wild-type and rok1Δ strains were 13.31±2.46 and 16.70±1.78
min, respectively, which was a significant difference (Fig. 4E). Entering the contraction phase,
the actin rings of the wild-type and rok1Δ strains
contracted for 15.39±0.85 and 14.70±1.92 min, respectively, and the
contraction rates were 0.19±0.02 and 0.18±0.02 µm/min, respectively
(Fig. 4F); there were no
significant differences in contraction time and contraction rate
(both P>0.05). These results indicate that rok1 gene
deletion prolongs the actin ring assembly process at 25˚C.
At 37˚C the contraction lengths were 4.09±0.38 vs.
3.50±0.26 µm and the total contraction rate of the actin ring was
0.12±0.02 vs. 0.09±0.01 µm/min for the wild-type and rok1Δ
strains, respectively. Both actin ring length and total contraction
rate were significantly decreased in the rok1Δ strain
compared with the wild-type. However, the actin ring assembly time
was 11.07±2.25 and 11.20±1.47 min for the wild-type and
rok1Δ strains, respectively, which was not a significant
difference (Fig. 4A-F). Entering
the contraction phase, the contraction time and rate of the actin
ring for the wild-type and rok1Δ strains were 15.60±3.22 vs.
15.33±1.95 min and 0.20±0.03 vs. 0.17±0.03 µm/min, respectively.
The actin ring contraction rate of the rok1Δ strain was
significantly decreased compared with the wild-type, and there was
no significant difference in the contraction time (Fig. 4E and F). These results indicate that
rok1 gene deletion affected the contraction process of actin
rings at 37˚C. Comparison of results at 25˚C and 37˚C revealed that
rok1 gene deletion both decreased the initiation length and
contraction length of the actin ring and decreased the contraction
rate. Unlike at 37˚C, rok1 gene deletion prolonged the actin
ring assembly process at 25˚C.
Changes of kinetochores during mitosis
of rok1Δ strain at different temperature conditions
The kinetochores interact with spindle microtubules
to assist in the correct segregation of chromosomes during mitosis.
Live-cell imaging of wild-type and rok1Δ cells with
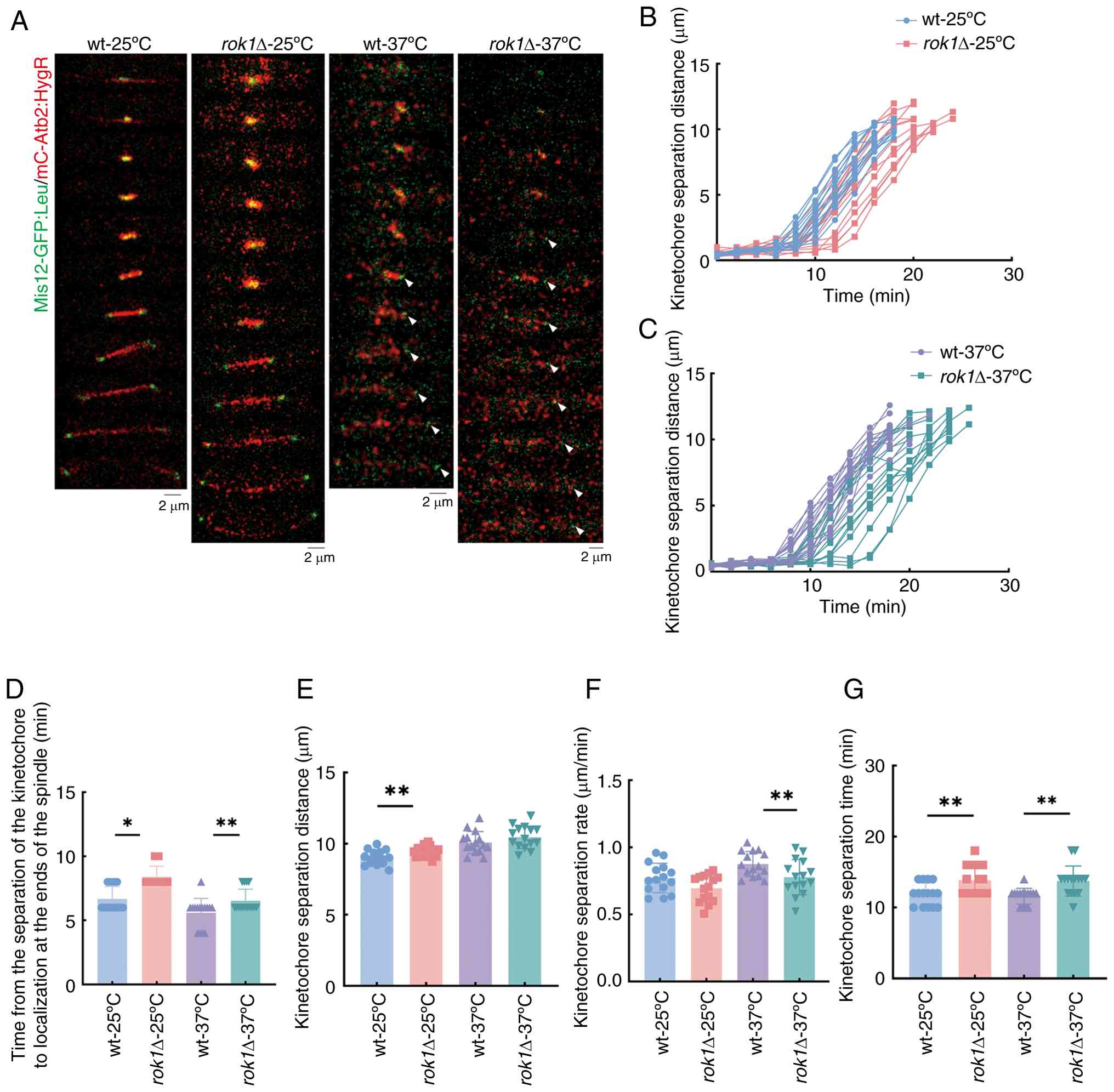
Mis12-GFP-tagged kinetochore proteins. The analysis of kinetochore
dynamics at 25˚C showed that the time from the separation to
localization at the spindle poles of kinetochores in the wild-type
and rok1Δ strains was 6.67±0.07 vs. 8.40±0.83 min,
respectively, which was a significant difference (Fig. 5A-D). Further analysis showed that
the separation distance (9.44±0.78 vs. 10.06±0.67 µm) and
separation time (11.87±1.60 vs. 13.87±2.07 min) were significantly
increased in rok1Δ strains compared with wild-type, but
there was no significant change in separation rates (0.81±0.11 vs.
0.74±0.10 µm/min) of kinetochores (Fig. 5E-G). These results indicate that
rok1 gene deletion prolonged the separation distance and
separation time of kinetochores as well as the time for
kinetochores to reach the poles of the spindle at 25˚C.
The time from the separation to localization at the
spindle poles of kinetochores in the wild-type and rok1Δ
strains was 5.60±1.12 and 6.63±0.92 min, respectively, at 37˚C,
which was a significant difference (Fig. 5A-D). Further analysis indicated
that the kinetochores separation distances of wild-type and
rok1Δ strains were 10.08±0.78 and 10.45±0.08 µm,
respectively, which was not significant difference (Fig. 5E). The separation time was
significantly increased (11.60±1.12 vs. 13.73±2.12 min; Fig. 5F) and the rate of kinetochores was
significantly decreased (0.88±0.09 vs. 0.78±0.13 µm/min; Fig. 5G) in rok1Δ strains compared
with wild-type. These results indicate that rok1 gene
deletion delayed the kinetochore separation process at 37˚C. At
25˚C and 37˚C, rok1 gene deletion prolonged the kinetochore
segregation time, the time from the separation to localization at
the spindle poles of kinetochores and thus delayed the kinetochore
segregation process. The inhibition of the kinetochore segregation
process was more effective with temperature stress at 37˚C.
Changes of centrosomes during mitosis
of rok1Δ strain at different temperatures
During mitosis, centrosomes assist in the completion
of microtubule assembly and influence the organization and function
of the spindle. Live-cell imaging of wild-type and rok1Δ
cells with Sid4-GFP-tagged centromere proteins was used to
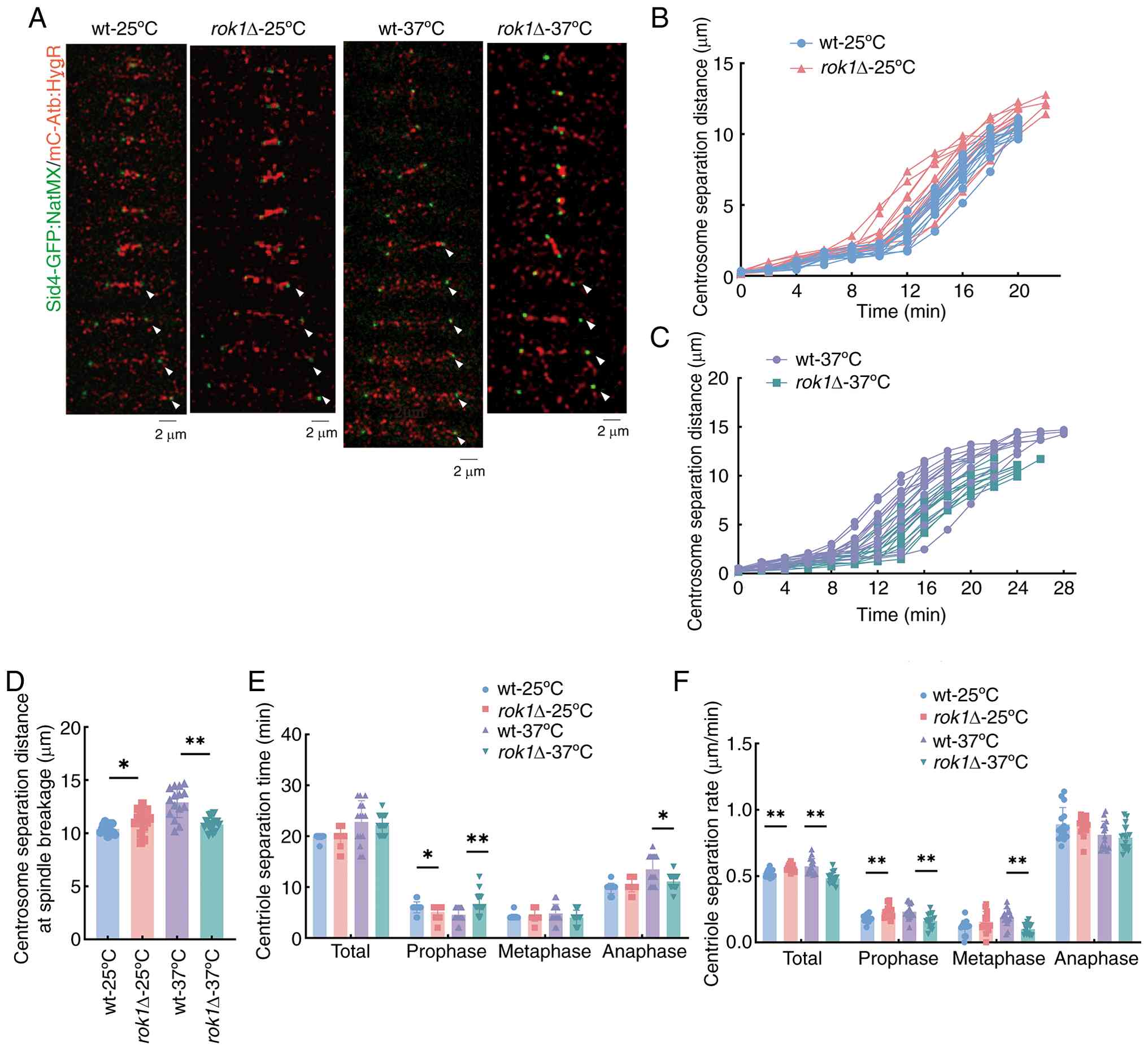
investigate changes in the centrosomes. The analysis of centrosome
dynamics at 25˚C indicated that the total separation time was
unchanged (19.87±0.52 vs. 19.87±1.92 min); however, the final
separation distances (10.40±0.43 vs. 11.17±1.06 µm) and the total
centrosome separation rates (0.52±0.03 vs. 0.56±0.02 µm/min) were
significantly increased in rok1Δ strains compared with
wild-type strains (Fig. 6A-F).
Further analysis of the centrosome separation process revealed that
the centrosome separation time of the wild-type and rok1Δ
strains at the prophase were 6.00±1.07 and 5.07±1.28 min, at
metaphase they were 4.13±0.52 and 4.40±1.12 min and at anaphase
they were 9.73±1.03 and 10.40±1.35 min, respectively (Fig. 6E). The centrosome separation rates
at prophase were 0.18±0.03 and 0.23±0.05 µm/min, at metaphase they
were 0.11±0.06 and 0.16±0.08 µm/min and at anaphase the rates were
0.89±0.13 and 0.88±0.07 µm/min in the wild-type and rok1Δ
strains, respectively. The centrosome separation time of
rok1Δ strains at prophase was significantly decreased and
the separation rate at prophase was significantly increased when
compared with the wild-type. These results indicated that at 25˚C,
rok1 gene deletion resulted in increased centromere
separation distance and increased separation rate.
When the experiments were conducted at 37˚C, the
total separation time was not significantly altered (22.80±4.20 vs.
22.53±1.77 min); on the other hand, there was a significant
decrease in both the final separation distance (12.90±1.44 vs.
10.82±0.63 µm) and the total centrosome separation rates (0.57±0.06
vs. 0.48±0.04 µm/min) in the rok1Δ strains compared with
wild-type. Further analysis of the centrosome separation process
revealed that the centrosome separation time of the wild-type and
rok1Δ strains at different phases were as follows: Prophase,
4.53±1.41 and 7.07±2.12 min; metaphase, 4.80±1.82 and 3.87±1.60
min; and anaphase, 13.47±2.88 and 11.60±1.12 min, in wild-type and
rok1Δ strains, respectively. The centrosome separation rates
were as follows: Prophase, 0.23±0.06 and 0.16±0.05 µm/min;
metaphase, 0.19±0.06 and 0.10±0.04 µm/min; and anaphase, 0.81±0.10
and 0.79±0.10 µm/min, in wild-type and rok1Δ strains,
respectively. The centrosome separation time at prophase in the
rok1Δ strain was significantly increased, but the separation
time at anaphase was significantly decreased compared with the
wild-type. Furthermore, the separation rate of the rok1Δ
strain was significantly decreased at prophase and metaphase
compared with the wild-type. These results indicated that at 37˚C,
rok1 gene deletion shortened the centrosome separation
distance and decreased the separation rate. Comparison of results
at 25 and 37˚C revealed that 37˚C high temperature stress reversed
the effect of rok1 gene deletion on the centrosome
segregation process.
Sequencing quality analysis
Combining the three replicates, the total number of
bases in the high-quality analyzed data for the wild-type-25˚C
strain, the wild-type-37˚C strain, the rok1Δ-25˚C strain and
the rok1Δ-37˚C strain were 6.80 Giga bases (G), 6.67 G, 6.88
G and 6.74 G, respectively. The Qphred20 (percentage of
bases with a phred score >20, where Phred=-10·log10(e) and e
represents the sequencing error rate) (43) value of all four groups of samples
was >98.00%, the Qphred30 (percentage of bases with a
Phred score >30) value was >94%, the GC content was between
41-43% and the sequencing error rate was <0.03%. These results
suggest that the sequencing data were accurate and reliable and
could be analyzed and studied subsequently.
Highly expressed gene analysis
Gene expression levels were quantitatively analyzed,
with FPKM <1 as low or no gene expression, FPKM >1 as genes
expressed, FPKM >60 as genes highly expressed and FPKM >5,000
as genes very highly expressed (35,44).
The results indicated that there were 4,950, 7,757, 4,997 and 7,488
genes expressed in the wild-type-25˚C strain, the wild-type-37˚C
strain, the rok1Δ-25˚C strain and the rok1Δ-37˚C
strain, respectively, which accounted for 39.00, 61.11, 39.37 and
59.00% of the total genes capable of expression in fission yeast.
In addition, there were 1,966, 2,181, 1,811 and 2,176 highly
expressed genes, accounting for 15.17, 17.18, 14.27 and 17.14% of
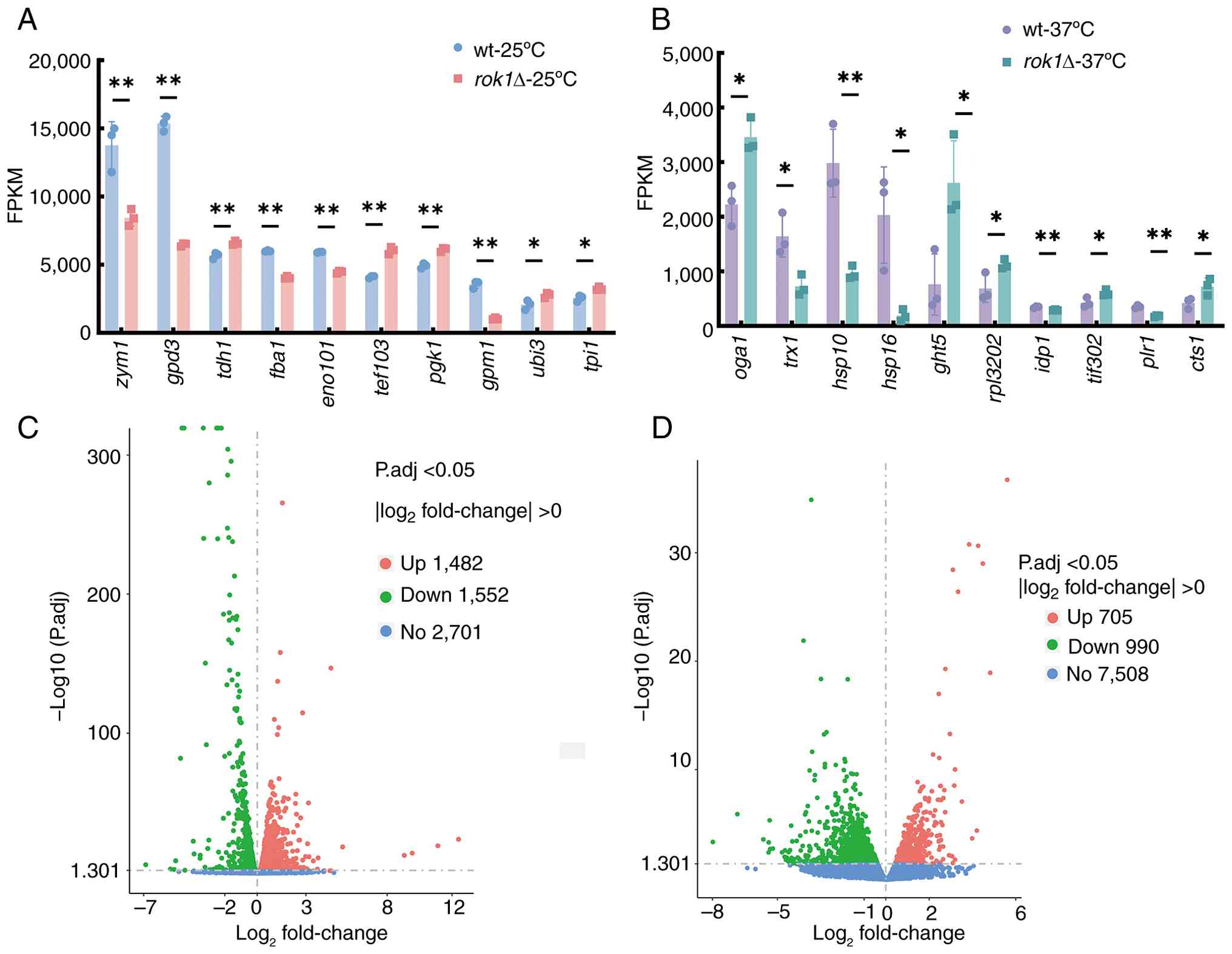
the total genes, respectively. From these highly expressed genes,
representative genes exhibiting statistically significant
expression differences between the rok1∆ strain and the
wild-type strain were further selected for display (Fig. 7A and B; Tables
III and IV).
 | Table IIIHighly expressed genes in the wt-25˚C
and rok1Δ-25˚C strains. |
Table III
Highly expressed genes in the wt-25˚C
and rok1Δ-25˚C strains.
| Gene | FPKM (wt) | FPKM
(rok1Δ) | Description |
|---|
| zym1 | 13,760.91 | 8,448.89 | Metallothionein,
Zym1 |
| gpd3 | 15,353.45 | 6,483.99 | Glyceraldehyde
3-phosphate dehydrogenase, Gpd3 |
| tdh1 | 5,679.17 | 6,599.78 |
Glyceraldehyde-3-phosphate dehydrogenase,
Tdh1 |
| fba1 | 6,012.45 | 4,059.26 |
Fructose-bisphosphate aldolase, Fba1 |
| eno101 | 5,927.17 | 4,461.72 | Enolase |
| tef103 | 4,107.47 | 6,054.25 | Translation
elongation factor EF-1 α, Ef1a-c |
| pgk1 | 4,919.32 | 6,107.86 | Phosphoglycerate
kinase, Pgk1 |
| gpm1 | 3,555.49 | 1,041.72 | BPG-dependent
phosphoglycerate mutase (PGAM), Gpm1 |
| ubi3 | 2,068.30 | 2,770.01 | Ribosomal-ubiquitin
fusion protein, Ubi3 |
| tpi1 | 2,552.69 | 3,237.18 | Triosephosphate
isomerase |
 | Table IVHighly expressed genes in the wt-37˚C
and rok1Δ-37˚C strains. |
Table IV
Highly expressed genes in the wt-37˚C
and rok1Δ-37˚C strains.
| Gene | FPKM (wt) | FPKM
(rok1Δ) | Description |
|---|
| oga1 | 2,229.10 | 3,462.16 | Ribosome
preservation factor, Stm1 homolog, Oga1 |
| trx1 | 1,644.48 | 726.54 | Cytosolic
thioredoxin, Trx1 |
| hsp10 | 2,983.73 | 964.69 | Mitochondrial heat
shock protein, Hsp10 |
| hsp16 | 2,031.58 | 187.98 | Heat shock protein,
Hsp16 |
| ght5 | 762.67 | 2,620.96 | Plasma membrane
high-affinity proton symporter, Ght5 |
| rpl3202 | 687.85 | 1,122.77 | 60S ribosomal
protein L32 |
| idp1 | 346.51 | 292.29 | Isocitrate
dehydrogenase, Idp1 |
| tif302 | 427.34 | 604.58 | Translation
initiation factor eIF3b (p84) |
| plr1 | 351.44 | 180.39 | Pyridoxal
reductase, Plr1 |
| cts1 | 424.50 | 722.51 | CTP synthase,
Cts1 |
Among the genes expressed to a very high level in
both wild-type and rok1∆-25˚C strains, the FPKM value of the
zym1 gene decreased by 1.629-fold compared to the wild-type
strain (P<0.01). The FPKM values of the tdh1 and
pgk1 genes increased by 1.1621- and 1.5047-fold,
respectively (both P<0.01). Among the genes that were expressed
at very high levels in both wild-type and rok1∆-37˚C
strains, the FPKM value of the rpl3202 gene increased
1.6323-fold in the wild-type strain (P<0.05). The FPKM values of
trx1 and plr1 genes were decreased by 0.4418-fold
(P<0.05) and 0.5132-fold (P<0.01), respectively.
Analysis of differentially expressed
genes
The differentially expressed genes of the wild-type
strain and the rok1Δ-25˚C and rok1Δ-37˚C strains were
analyzed. The results indicated that there were 3,034 and 1,695
differentially expressed genes in the rok1Δ-25˚C and
rok1Δ-37˚C strains, respectively, compared with the
wild-type strains, which included 1,482 and 705 upregulated genes
and 1,552 and 990 downregulated genes, respectively (Fig. 7C and D).
Compared with the wild-type strain, in the
rok1Δ-25˚C strain, mei2, map2, map3 and
psc3 gene expression were upregulated by 1.2866-, 1.8764-,
2.7593- and 1.3973-fold, respectively (Table V). In the rok1Δ-37˚C strain,
gpa2, rgs1, myo51 and blt1 gene
expression was upregulated by 1.9694-, 2.1606-, 0.7366- and
1.0046-fold, respectively (Table
VI). In the rok1Δ-25˚C strain, ddx27 and
pas1 gene expression were downregulated by 1.7320- and
1.6148-fold, respectively (Table
VII). In the rok1Δ-37˚C strain, rho5,
wos2, apc14, and pas1 genes were downregulated
by 1.4299-fold, 1.1209-fold, 0.7467-fold, and 1.3259-fold,
respectively. arp1, dil1, cmk1 and mfr1
gene expression were downregulated by 0.9087-, 1.6737-, 1.3417- and
1.1414-fold, respectively (Table
VIII). Notably, although arp1, dli1, cmk1,
and mfr1 genes were all significantly downregulated, the
downregulation factor for arp1 was 0.9087-fold, lower than
that of the other genes (>1). This result indicates that while
the deletion of the rok1 gene broadly suppresses the
transcription of these genes, its inhibitory effect on arp1 is
relatively weaker.
 | Table VUpregulated differentially expressed
genes in the wild-type-25˚C and rok1Δ-25˚C strains. |
Table V
Upregulated differentially expressed
genes in the wild-type-25˚C and rok1Δ-25˚C strains.
| Gene |
log2FoldChange | Adjusted
P-value | Length, bp | Description |
|---|
| hsp90 | 1.5155 |
3.25x10-266 | 2,600 | Hsp90
chaperone |
| ght2 | 1.2251 |
6.59x10-138 | 2,245 | Hexose
transmembrane transporter, Ght2 |
| map3 | 2.7593 |
3.18x10-115 | 2,888 | Pheromone M-factor
receptor, Map3 |
| bip1 | 1.0174 |
1.60x10-110 | 2,805 | ER heat shock
protein, BiP |
| mei2 | 1.2866 |
1.24x10-104 | 4,033 | RNA-binding protein
involved in meiosis, Mei2 |
| map2 | 1.8764 |
1.91x10-53 | 3,726 | P-factor pheromone
Map2 |
| tef101 | 0.7861 |
5.12x10-53 | 1,569 | Translation
elongation factor EF-1 α, Ef1a-a |
| wtf25 | 1.5509 |
1.45x10-51 | 2,271 | Wtf element |
| psc3 | 1.3973 |
3.73x10-40 | 3,989 | Mitotic cohesin
complex, non-SMC subunit, Psc3 |
| ctt1 | 0.6184 |
1.83x10-39 | 2,212 | Catalase |
 | Table VIUpregulated differentially expressed
genes in the wild-type-37˚C and rok1Δ-37˚C strains. |
Table VI
Upregulated differentially expressed
genes in the wild-type-37˚C and rok1Δ-37˚C strains.
| Gene |
log2FoldChange | Adjusted
P-value | Length, bp | Description |
|---|
| ppk1 | 2.4226 |
6.88x10-15 | 3,072 | Serine/threonine
protein kinase, Ppk1 |
| rgs1 | 2.1606 |
1.98x10-9 | 1,446 | Regulator of
G-protein signaling, Rgs1 |
| put4 | 3.1716 |
3.38x10-8 | 1,659 | Plasma membrane
proline transmembrane transporter, Put4 |
| snoR54b | 2.5113 |
7.20x10-7 | 96 | Small nucleolar
RNA, snR54b |
| uck2 | 1.6349 |
9.23x10-7 | 663 | Uracil
phosphoribosyltransferase, Uck2 |
| gpa2 | 1.9694 |
6.84x10-6 | 1,065 | Heterotrimeric G
protein α-2 subunit, Gpa2 |
| myo51 | 0.7366 |
1.52x10-2 | 4,416 | Myosin type V |
| fib1 | 0.7617 |
1.74x10-2 | 918 | Fibrillarin, rRNA
and histone methyltransferase |
| blt1 | 1.0046 |
3.53x10-2 | 2,103 | Ubiquitin
domain-like protein, Blt1 |
| tif11 | 0.6131 |
3.54x10-2 | 417 | Translation
initiation factor eIF1A |
 | Table VIIDownregulated differentially
expressed genes in the wild-type-25˚C and rok1Δ-25˚C
strain. |
Table VII
Downregulated differentially
expressed genes in the wild-type-25˚C and rok1Δ-25˚C
strain.
| Gene |
log2FoldChange | Adjusted
P-value | Length, bp | Description |
|---|
| bgl2 | -1.8472 |
7.99x10-305 | 1,654 | Glucan
β-glucosidase, Bgl2 |
| mae2 | -1.6447 |
3.56x10-296 | 2,080 | Malic enzyme,
malate dehydrogenase, Mae2 |
| ddx27 | -1.7320 |
5.34x10-200 | 2,423 | ATP-dependent RNA
helicase, Ddx27/Drs1 |
| prz1 | -1.7525 |
2.89x10-187 | 3,442 | Calcineurin
responsive transcription factor, Prz1 |
| gal7 | -2.1082 |
3.39x10-186 | 1,566 |
Galactose-1-phosphate uridylyltransferase,
Gal7 |
| pas1 | -1.6148 |
1.99x10-165 | 3,109 | Cyclin, Pas1 |
| sgf73 | -3.2303 |
5.61x10-151 | 1,416 | SAGA complex
subunit, Sgf73 |
| per1 | -1.5861 |
5.54x10-139 | 2,963 | Plasma membrane
amino acid permease, Per1 |
| shd1 | -0.7636 |
1.83x10-47 | 4,782 | Cytoskeletal
protein binding protein Sla1 family, Shd1 |
| mbx1 | -1.2067 |
1.88x10-45 | 3,183 | MADS-box
transcription factor, Mbx1 |
 | Table VIIIDownregulated differentially
expressed genes in the wild-type-37˚C and rok1Δ-37˚C
strain. |
Table VIII
Downregulated differentially
expressed genes in the wild-type-37˚C and rok1Δ-37˚C
strain.
| Gene |
log2FoldChange | Adjusted
P-value | Length, bp | Description |
|---|
| rho5 | -1.4299 |
1.39x10-6 | 603 | Rho family GTPase,
Rho5 |
| mbx1 | -1.8473 |
1.56x10-6 | 1,374 | DNA-binding
transcription factor, MADS-box, Mbx1 |
| adg1 | -1.8871 |
1.73x10-6 | 501 |
Schizosaccharomyces pombe specific
protein, Adg1 |
| vip1 | -1.2490 |
1.50x10-3 | 774 | RNA-binding
protein, Vip1 |
| wos2 | -1.1209 |
1.54x10-3 | 561 | p23 homolog, Hsp90
co-chaperone, Wos2 |
| mug125 | -1.0961 |
1.56x10-3 | 858 |
Schizosaccharomyces pombe specific
protein, Mug125 |
| pas1 | -1.3259 |
9.58x10-3 | 1,236 | Cyclin, Pas1 |
| apc14 | -0.7467 |
3.10x10-2 | 324 | Anaphase-promoting
complex subunit, Apc14 |
| arp1 | -0.9087 |
3.44x10-2 | 1,140 | Dynactin complex
subunit, centractin family actin-like protein, Arp1 |
| dli1 | -1.6737 |
4.40x10-2 | 1,083 | Meiotic dynein
intermediate light chain, Dli1 |
| cmk1 | -1.3417 |
4.60x10-5 | 1,008 |
Calcium/calmodulin-dependent protein
kinase, Cmk1 |
| mfr1 | -1.1414 |
8.59x10-4 | 1,266 | Meiotic APC
activator, Mfr1 |
Differential gene expression
validation
Analysis of differentially expressed genes from the
transcriptomic data revealed that psc3 and psm1 were
key genes at 25˚C, whereas myo51 and blt1 were key
genes at 37˚C. To validate the expression changes of these genes,
specific primers were designed using act1 as the reference
gene and verified for specificity through the NCBI Primer-BLAST
tool. The results demonstrated that in the S. pombe (taxid:
4896) genome, the primers for psc3, psm1, myo51, blt1 and
act1 all specifically matched unique target sequences,
confirming the validity of the primer design and their suitability
for RT-qPCR experiments.
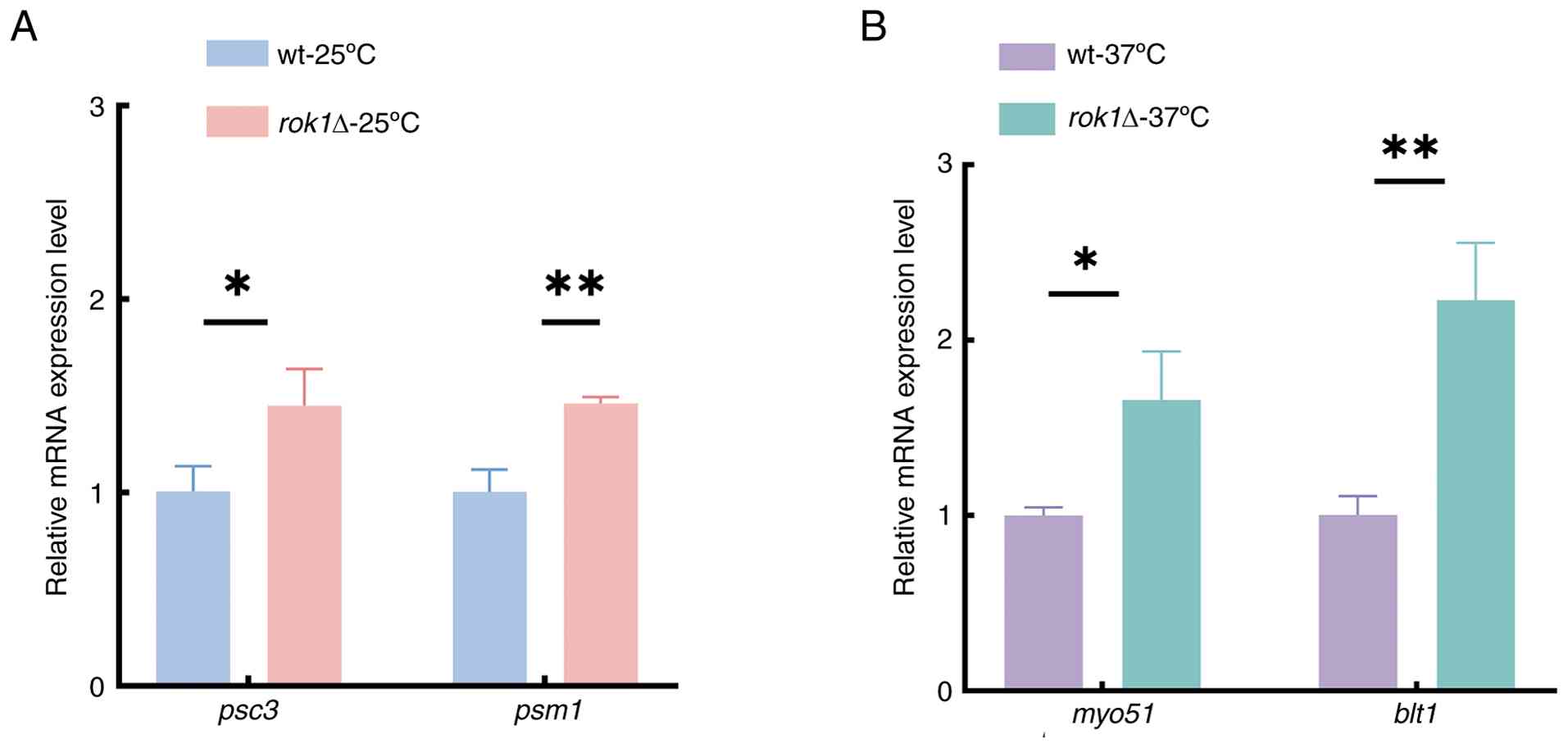
The RT-qPCR results showed that the expression
levels of psc3 and psm1 were significantly
upregulated at 25˚C (Fig. 8A); in
addition, myo51 and blt1 were significantly
upregulated at 37˚C (Fig. 8B) with
rok1 deletion compared with the wild-type. These findings
were consistent with the RNA-Seq data, confirming the reliability
of the transcriptomic results. These results suggest that
rok1 regulate the expression of these key genes, thereby
influencing the dynamics of mitotic progression.
Differential gene GO functional
enrichment analysis
GO enrichment analysis of differentially expressed
genes was performed in wild-type and rok1Δ strains, and the
10 categories with the most significant up- and downregulated
differentially expressed genes were selected and plotted as bar
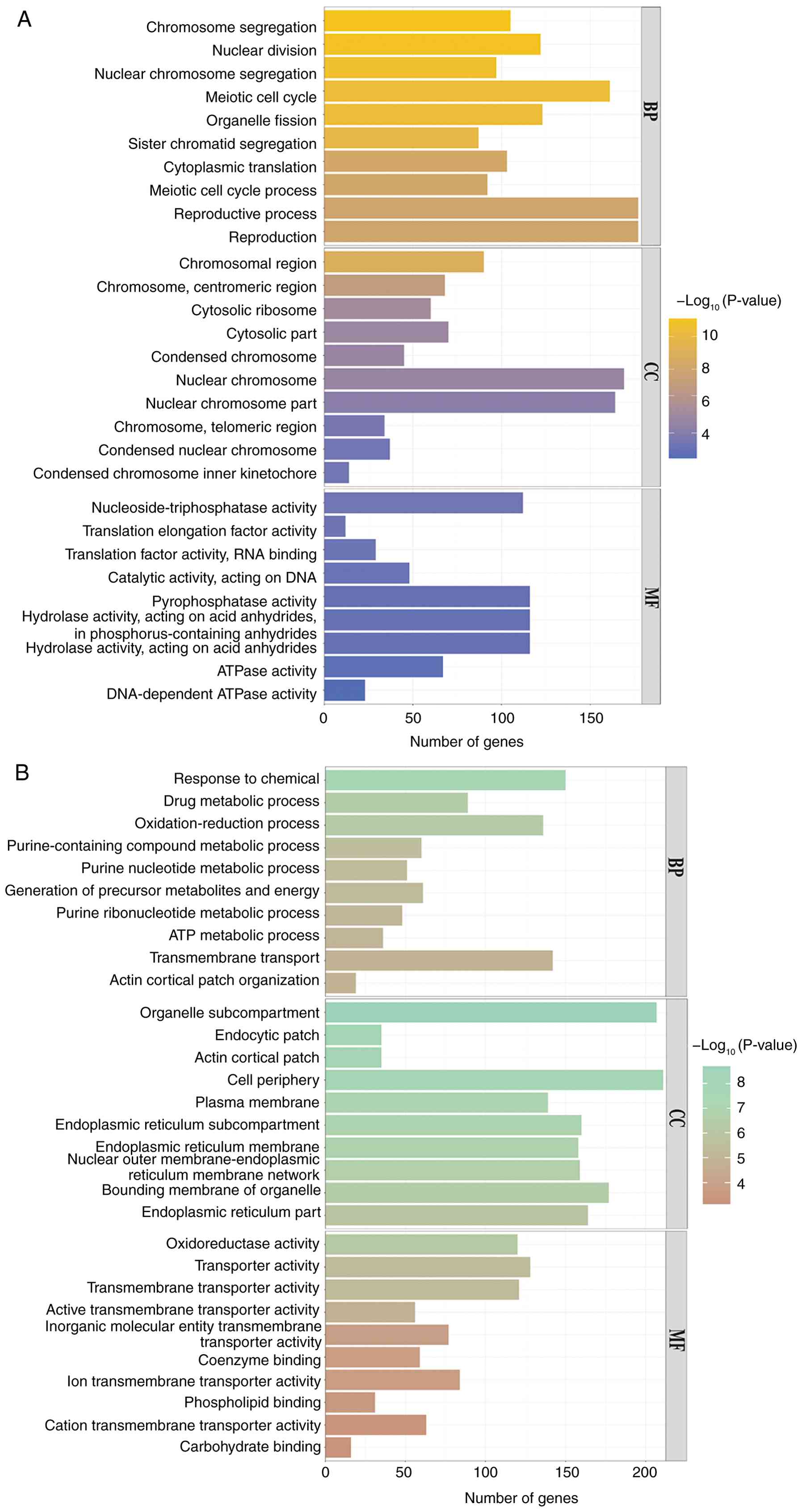
graphs. The analysis revealed that at 25˚C, the rok1Δ strain
was enriched for differential genes up to 303 GO branches (P≤0.05),
including 230 biological processes, 41 cellular components and 32
molecular functions, compared with the wild-type strain. Among the
biological processes, upregulated genes were enriched in ‘sister
chromatid segregation’ and ‘reproductive process’. In cellular
components, upregulated genes were enriched in ‘chromosomes,
centromeric region’ and ‘condensed chromosome inner kinetochore’
regions. In molecular function, upregulated genes were enriched in
‘nucleoside-triphosphatase activity’ and ‘translation elongation
factor activity’ (Fig. 9A). In
addition, downregulated differential genes were enriched in
‘purine-containing compound metabolic process’ and ‘actin cortical
patch organization’, ‘actin cortical patch’ and ‘plasma membrane’
regions in cellular components, with ‘transmembrane transporter
activity’ and ‘oxidoreductase activities’ in molecular functions
(Fig. 9B).
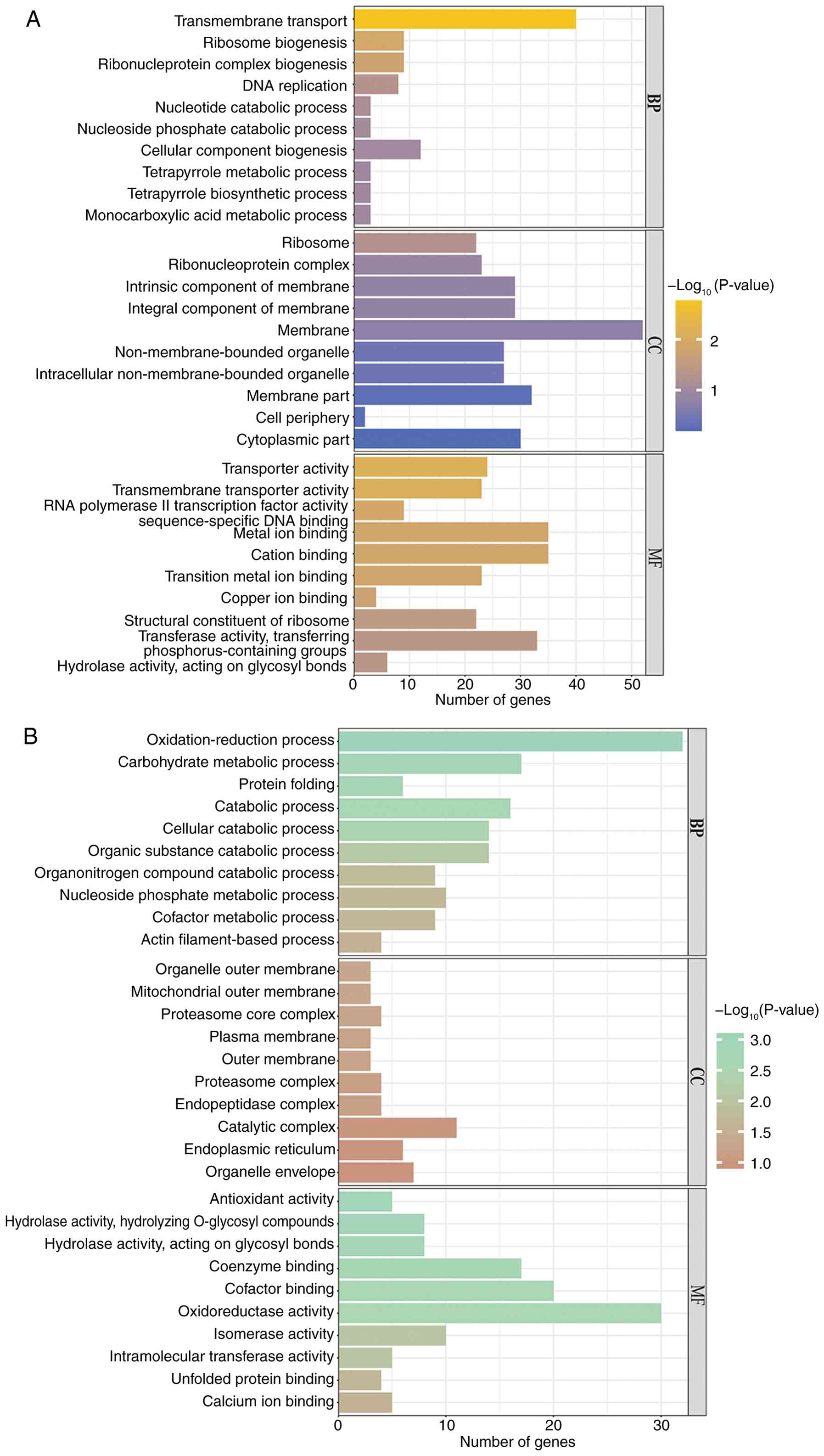
At 37˚C, the differential genes of the rok1Δ
strain were enriched in 28 GO branches (P≤0.05), including 3
biological processes and 25 cellular components. compared with the
wild-type strain. upregulated genes were enriched in ‘transmembrane
transport’ and ‘ribosome biogenesis’ during biological processes;
in cellular components, upregulated genes were enriched in the
‘ribosome’; in molecular function, upregulated genes were enriched
in ‘transmembrane transporter activity’ and ‘RNA polymerase II
transcription factor activity, sequence-specific DNA binding’
(Fig. 10A). In addition,
downregulated differential genes were enriched in ‘protein folding
‘and ‘actin filament-based’ processes, ‘mitochondrial outer
membrane’ and ‘proteasome core complex’ regions in cellular
components and ‘hydrolase activity, acting on glycosyl bonds’ in
molecular functions (Fig. 10B).
The GO enrichment results indicate that the ribosome biogenesis
process of the rok1Δ strain was affected, the mitotic
cytoskeleton was abnormal and the normal cell growth and
reproduction process was disrupted.
Differential gene KEGG enrichment
analysis
KEGG enrichment analysis was performed on the
differentially expressed genes of the wild-type strain and the
rok1Δ strain, and the top 10 pathways of selected
upregulated genes and downregulated genes were plotted as bar
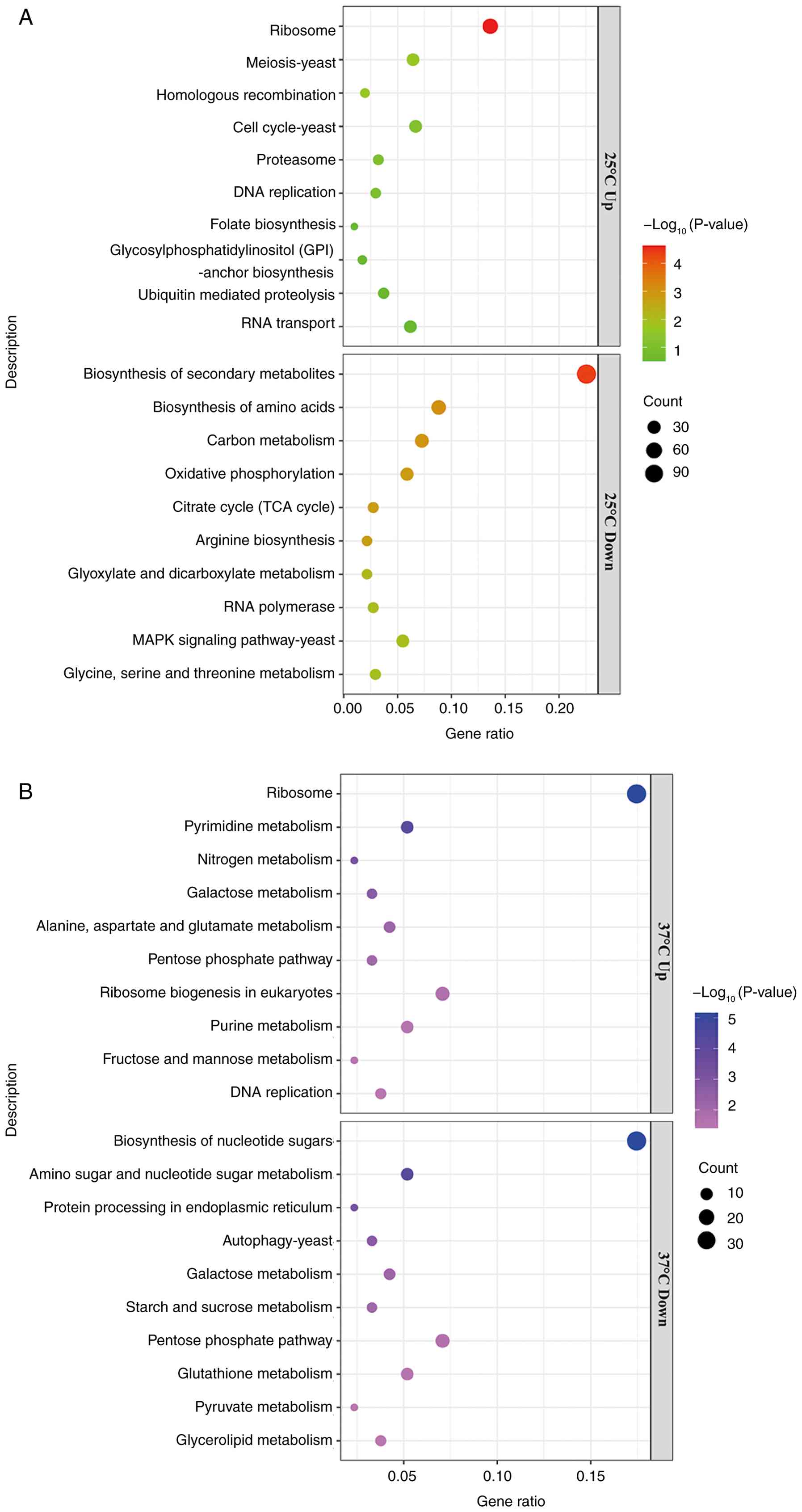
graphs (Fig. 11A and B). The results indicated that 1,634
differential genes were enriched in 80 pathways in the
rok1Δ-25˚C strain. The upregulated genes were enriched in
pathways such as ‘ribosome’, ‘cell cycle-yeast’ and ‘proteasome’,
whereas the downregulated genes were enriched in pathways such as
‘MAPK signaling pathway-yeast’ and ‘citrate cycle (TCA cycle)’. The
rok1Δ-37˚C strain was enriched for 787 differential genes in
89 pathways. Upregulated genes were enriched in pathways such as
‘ribosome’, ‘ribosome biogenesis in eukaryotes’ and ‘DNA
replication’, whereas downregulated genes were mainly enriched in
pathways such as ‘autophagy-yeast’ and ‘protein processing in
endoplasmic reticulum’. At 25˚C, the ‘cell cycle-yeast’ pathway was
enriched for 27 upregulated genes (Fig. 11A). The genes that were
upregulated in this pathway included spo4, rad17,
psm1, mis4 and mad1. At 37˚C, the
‘autophagy-yeast’ pathway was enriched for 18 downregulated genes
(Fig. 11B). The major genes with
downregulated expression in this pathway included pas1,
atg17, atg13, atg15 and arc1.
Discussion
In transcriptomic analysis, highly expressed genes
indicate the core functions and active metabolic pathways of cells
under specific conditions, serving as a crucial entry point and a
central analytical step for uncovering cellular functions and
adaptive mechanisms. Among the genes expressed to a very high level
in both wild-type and rok1Δ-25˚C strains was the zym1
gene. This encodes the metallothionein Zym1, and deletion of the
zym1 gene results in aberrant segregation of meiotic
chromosomes and aberrant spore formation (14,23).
The FPKM values of zym1 were decreased in the
rok1Δ-25˚C strain, which is in agreement with the spore
production anomaly in the present study. The tdh1 gene
encodes the glyceraldehyde-3-phosphate dehydrogenase Tdh1, which
physically binds to the response regulator Mcs4 and the
stress-responsive MAPKKK and is involved in the positive regulation
of the MAPK cascade response. Through phosphorylation, the MAPK
pathway delivers model factors that regulate biological growth and
division-related signaling (45).
The pgk1 gene encodes the phosphoglycerate kinase Pgk1,
which can be phosphorylated by the cell cycle protein-dependent
kinase CDK1, and thus participates in cellular glycolysis as well
as gluconeogenesis, providing energy for cell division, growth, and
protein synthesis (46). FPKM
values of tdh1 and pgk1 were upregulated in the
rok1Δ-25˚C strain, which suggested that phosphorylation of
the MAPK signaling pathway and glycolytic processes was being
compensated.
Among the genes that were expressed at very high
levels in both wild-type and rok1Δ-37˚C strains was the
trx1 gene, which encodes the cytosolic thioredoxin Trx1,
whose deletion results in abnormal DNA replication checkpoints in
mitosis and spore formation (23,47).
The plr1 gene encodes the pyridoxal reductase Plr1, which is
involved in the process of pyridoxal biosynthesis, whereas
plr1 gene deletion results in abnormal sporulation (23). The FPKM values of trx1 and
plr1 were downregulated in the rok1Δ-37˚C strain,
which indicated abnormal cell cycle progression and sporulation,
which was consistent with the abnormalities of cell division in the
rok1Δ-37˚C strain. The rpl3202 gene encodes the 60S
ribosomal protein L32, which participates in ribosome composition
and cytoplasmic translation processes as a cytosolic large
ribosomal subunit component and directs mitotic protein synthesis
(48). The FPKM value of
rpl3202 was increased in the rok1Δ-37˚C strain, which
suggested that ribosome composition and translation processes had
been compensated.
Differential gene analysis can directly reveal the
molecular mechanisms, key regulatory pathways, and potential
functional targets underlying phenotypic differences by deciphering
the dynamic changes in gene expression under different conditions
(49). In this study, we conducted
further analysis of differentially expressed genes between the
wild-type and rok1Δ strains. The results showed that among
the genes upregulated in the rok1Δ-25˚C strain, the
mei2 gene encodes the RNA-binding protein involved in
meiosis Mei2, which switches the cell from a mitotic cell cycle to
a meiotic cell cycle by dephosphorylation, allowing the cell to
stably express meiosis-specific mRNAs (50). The map2 gene encodes the
P-factor pheromone Map2 and map3 encodes the pheromone
M-factor receptor Map3. Both Map2 and Map3 are involved in the
pheromone-responsive MAPK cascade; map2 is involved in the positive
regulation of cell fusion coupling through signaling (51-53).
The psc3 gene encodes the STAG protein subunit Psc3 of the
mitotic cohesin complex, which assembles with Rad21, Psm1 and Psm3
into a functional complex. This complex maintains the connection
between sister chromatids through physical interactions,
participates in establishing and sustaining sister chromatid
cohesion, and ensures accurate chromosome segregation (54). The psc3 gene is consistent
with the lagging of centromeres observed in this study at 25˚C,
suggesting that the Rok1 protein participates in regulating the
formation and maintenance of cohesion between sister
chromatids.
Among the genes upregulated in the
rok1Δ-37˚C strain (Table
VI), the gpa2 gene encodes a protein that is a
structural component of the mitotic spindle pole body, and Gpa2 is
involved in the regulation of mitotic progression (55). The rgs1 gene encodes the
regulator of G-protein signaling Rgs1, and rgs1 gene
deletion affects the pheromone-responsive MAPK cascade response
process and results in aberrant protein localization in the actin
fusion focus (56). The
myo51 gene encodes a type V myosin whose motor domain and
tail domain interact with actin filaments, effectively mediating
the integration of nodes into the actin filament network, thereby
regulating the actin ring assembly and cytokinesis (57-59).
Myo51 can participate in CAR assembly and cytokinesis process
together with myosin II. Studies have shown that at 29˚C, cells
lacking myo51 can maintain normal morphology and growth rate,
whereas overexpression of myo51 leads to elongated cells and
failure to form functional septa, exhibiting phenotypes similar to
those observed in the present study; this indicates that Myo51
plays a non-essential role in cytokinesis (60). Further research has revealed that
Myo51 primarily plays a critical role in the assembly phase of the
contractile ring but cannot independently drive ring contraction
during the constriction phase (61). The present findings demonstrate a
slowed contraction rate of the actin ring, accompanied by
upregulated expression of myo51 according to transcriptomic data;
however, the specific molecular mechanisms underlying this
phenomenon require further investigation. The blt1 gene
encodes the ubiquitin domain-like protein Blt1, which possesses
cytoskeletal protein-membrane anchoring activity. It recruits the
Nod1-Gef2 complex to form a cell division node, assembles the
mitotic actomyosin contractile ring, and participates in
cytokinesis (62,63). In summary, the upregulation of
myo51 and blt1 may be a key factor underlying the
impaired actin ring assembly observed in the rok1Δ strain at
37˚C in this study. These findings suggest that the Rok1 protein
may influence actin polymerization, thereby participating in the
regulation of actin ring assembly, contraction, and
cytokinesis.
Among the genes downregulated in the
rok1Δ-25˚C strain, the ddx27 gene encodes the
ATP-dependent RNA helicase Ddx27/Drs1, which regulates rRNA
processing during ribosome biogenesis, and deletion of the
ddx27 gene results in abnormal mitotic cell cycle (15). The pas1 gene encodes the
cyclin Pas1, which as a partner protein of the cell cycle
protein-dependent protein kinase Pho85/PhoA-like Pef1, is involved
in the negative regulation of sister chromatid cohesion together
with Pef1(64). The downregulation
of the pas1 gene correlates with the observed lagging of
centromeres at 25˚C in this study, suggesting that the Rok1 protein
participates in regulating the formation and maintenance of
cohesion between sister chromatids.
Among the genes downregulated in the
rok1Δ-37˚C strain, the rho5 gene encodes the Rho
family GTPase Rho5, which is localized to the terminal end of
interphase cells and the intermediate region of mitotic cells and
participates in the regulation of the actin cytoskeletal
organization and the synthesis of the cell wall (65). The wos2 gene encodes the p23
homolog, the Hsp90 co-chaperone Wos2, which is involved in the
regulation of protein-containing complex assembly, and deletion of
the wos2 gene results in cell lysis and abnormal mitotic
cell cycle (15). The
anaphase-promoting complex subunit Apc14 is involved in
anaphase-promoting complex-dependent catabolic process and mitotic
sister chromatid segregation, and apc14 gene deletion
results in reduced mitotic checkpoint complex binding (66,67).
The downregulation of apc14 may account for the delayed
separation of centromeres during mitosis in the rok1Δ strain
at 37˚C in this study. This suggests that the Rok1 protein may
influence the ability of centromeres to migrate along the
spindle.
The arp1 gene encodes the dynactin complex
subunit, the centractin family actin-like protein Arp1. In S.
pombe, Arp1, Mug5 and Jnm1 constitute core components of the
dynactin complex, with Arp1 localizing to dynein anchoring sites at
the cell cortex. Arp1, Mug5, Jnm1 and the dynactin
microtubule-binding subunit Ssm4, which interacts with Mug5,
participate in dynein anchoring and regulation of microtubule
contraction. Deletion of the arp1 gene results in reduced
microtubule depolymerization rates (68). The dli1 gene encodes the
meiotic dynein intermediate light chain Dli1, which facilitates the
increased expression of the dynein complex core molecule Dhc1. This
enhancement promotes the binding of dynein to both the spindle pole
body (SPB) and microtubules (MTs), thereby facilitating MT
elongation (69). The mfr1
gene encodes the meiotic APC activator Mfr1; Mfr1 binds to the
meiosis-specific protein Mes1, and its expression can rescue the
entry defect at meiosis II in mes1Δ cells (70). Additionally, Mfr1 mediates the
rapid degradation of Cdc13 cyclin at the end of meiosis II,
ensuring proper exit from meiosis. Mfr1 null mutants complete
meiosis II but maintain high levels of Cdc13 and Cdc2 kinase
activity, resulting in delayed exit from cell division (71). The cmk1 gene encodes the
Ca2+/calmodulin-dependent kinase Cmk1; Cmk1
phosphorylates the M-phase inducer Cdc25 phosphatase, regulating
Cdk1 activity, and levels are closely associated with spindle
dynamics: During metaphase when Cdk1 activity is high, the
microtubule site clamping complex subunit Mde4 is phosphorylated at
its Cdk1 phosphorylation sites and localizes to kinetochores;
during anaphase, decreased Cdk1 activity leads to Mde4
dephosphorylation and translocation to the spindle, promoting
spindle elongation and maintaining its integrity (72). Furthermore, cmk1 gene
expression was downregulated in rok1Δ cells at 37˚C,
accompanied by shortened spindle elongation length and reduced
centromere separation distance. However, RNA-Seq revealed no
differential expression of centromere-related genes, with changes
only detected in genes involved in the microtubule assembly
pathway, such as cmk1. These results indicate that although
prominent phenotypic alterations were observed at the centromere
level, the underlying cause likely does not originate from aberrant
expression of centromere-associated proteins. The findings suggest
that Rok1 more plausibly influences centromere phenotypic variation
indirectly by modulating the microtubule assembly process, in which
Cmk1 appears to serve a representative role.
KEGG, as a systematic pathway database, provides
standardized biological pathway annotations and enrichment analysis
for differentially expressed genes, serving as a core tool for
deciphering the molecular mechanisms underlying high-throughput
data (38). Cell cycle regulation
is a process required to ensure the maintenance of genome
integrity, and mitotic abnormalities caused by cell cycle
dysregulation trigger autophagy through various protein
interactions. Autophagy combines cell growth with cell division to
maintain the integrity of the nuclear and mitochondrial genomes.
Defective autophagy leads to defective cell growth and is
associated with abnormal mitosis (73). At 25˚C, in the cell cycle-yeast
pathway, the Spo4 protein has protein serine/threonine kinase
activity and is involved in the reorganization of specific spindle
pole bodies during meiosis, regulating ascospore formation; Spo4
protein deficiency results in abnormal prospore membrane formation
and chromosome segregation (23,74).
The rad17 gene encodes the replication factor C-related
checkpoint protein Rad17, which is localized in the nucleus by
binding to chromatin; Rad17 is involved in regulating mitotic DNA
replication checkpoint and DNA damage checkpoint signaling in
G2 phase (75). The
mad1 gene encodes the mitotic spindle checkpoint protein
Mad1, which recruits Cut7 proteins to the mismatched kinetochore in
chromosomes, promotes chromosome sliding on the spindle and is
involved in the mitotic spindle assembly checkpoint signaling
process (76). The psm1
gene encodes the ATPase subunit Psm1 of the mitotic/meiotic cohesin
complex, which participates in establishing sister chromatid
cohesion during mitosis. The Psm1-Psm3 head domain heterodimer,
Mis4 and the N-terminal helical domain of Rec8 bound to the
coiled-coil region of Psm3 collectively form the primary
chromatin-binding region of the cohesin complex. By recruiting Plo1
kinase, this complex promotes the phosphorylation of Rec8, thereby
ensuring the mono-orientation of sister kinetochores (77,78).
The cohesin loading factor (adherin) Mis4/Scc2, which has ATPase
activator activity, forms a tertiary complex with cohesins on DNA
and is involved in the regulation of mitotic sister chromatid
cohesion and chromosome segregation. Abnormalities in the Mis4
protein lead to abnormal cell cycle arrest and premature sister
chromatid segregation in mitosis (79,80).
Abnormal expression of cell cycle pathway genes suggested that the
cytokinesis process was affected in the rok1Δ-25˚C strain.
This was consistent with the abnormal results of spore formation
and mitosis in the rok1Δ-25˚C strain, and with the results
of GO enrichment analysis (Fig.
9).
At 37˚C, the autophagy-yeast pathway, the
pas1 gene encodes the cell cycle protein Pas1, which is
involved in the composition of the cyclin-dependent protein kinase
holoenzyme complex that negatively regulates the mitotic cell cycle
G1/S transition and sister chromatid cohesion (64). Atg17 participates in cellular
autophagosome assembly as an autophagy associated protein kinase
activator to remove its own damaged cellular structures; deletion
of Atg17 results in a decreased sporulation frequency and an
abnormal G1 to G0 transition (81,82).
Atg13 is co-localized with other Atg machinery proteins at the
phagocytic vesicle assembly site in the proximal endoplasmic
reticulum and is involved in cellular autophagosome assembly;
atg13 gene deletion results in abnormal spore formation and
autophagy processes (83). The
atg15 gene encodes the autophagy-associated
lysophospholipase Atg15, which is involved in the regulation of
microautophagy of the nucleus. Chromosome segregation and cellular
autophagy are abnormal in atg15Δ strains, and the frequency
of spore formation is decreased (23,83).
The arc1 gene encodes the Arp2/3 actin organizing complex WD
repeat subunit Sop2, which is involved in the composition of the
Arp2/3 protein complex to mediate actin polymerization; aberrant
Sop2 protein decreases protein localization to actomyosin
contractile ring during mitosis (84). Abnormal expression of the autophagy
pathway indicated impaired mitotic processes in the
rok1Δ-37˚C strain. This was consistent with the abnormal
results of actin formation and contraction and kinetochore
separation in the rok1Δ-37˚C strain, as well as with the
results of GO enrichment analysis. In addition, although the
ribosomal pathway was upregulated in both rok1Δ-25˚C and
rok1Δ-37˚C strains, few genes were upregulated in both.
Therefore, rok1 gene deletion mainly regulated the mitotic
process through up- and downregulation of different pathways at 25
and 37˚C.
Temperature is a critical factor influencing
protein synthesis and function in cells. In the present study, two
culture temperatures, 25 and 37˚C, were selected to investigate the
potential biological functions of the rok1 gene deletion
under heat stress to amplify the phenotypic effects results. This
experimental approach has been widely adopted in related studies;
for instance, Codlin et al (85) discovered that btn1-deficient cells
exhibited only mild proliferation defects under normal growth
conditions at 25˚C, whereas severe depolarization and cell lysis
were observed under heat stress at 37˚C. This revealed the role of
Btn1p in the F-actin-dependent endocytosis-polarized growth
coupling pathway. Similarly, Hoya et al (86) identified
functional overlap between exomer and GGA22 by analyzing the
synthetic growth defects of the gga22Δ cfr1Δ double
mutant under both low-temperature (22˚C) and high-temperature
(36˚C) stress conditions. It is important to note that phenotypic
variations in strains at different temperatures are a common
phenomenon and are generally not defined as temperature-sensitive
strains. Temperature-sensitive strains specifically refer to those
that exhibit complete growth arrest or total loss of protein
function at a restrictive temperature (87). Such strains show 100%
growth inhibition under non-permissive conditions; therefore, the
rok1Δ strain investigated here is not a
temperature-sensitive strain.
Notably, no widespread ribosome biosynthesis
dysfunction was observed in the rok1Δ strain in the present
study; mitotic progression remained normal and transcriptomic
analysis did not reveal a global decline in protein translation
levels. Instead, only specific alterations in the expression of
actin-related proteins were detected. Based on comprehensive review
of literature on the rok1 gene in fission yeast, to the best
of our knowledge, there is currently no evidence indicating that
rok1 directly affects protein translation through ribosome
biosynthesis. Therefore, the present study proposes a novel
mechanism: The decreased translation levels of actin-related
proteins represent a direct consequence of rok1 deletion,
rather than being secondary to ribosome biogenesis defects,
revealing a previously unrecognized role of the rok1 gene in
mitotic dynamics.
Currently, research on the rok1 gene in
fission yeast has primarily focused on its role in ribosome
biogenesis, while its mechanism in regulating mitotic dynamics
remains unclear. There is a particular lack of studies
investigating the effects of rok1 deletion on mitosis and its
molecular mechanisms under both normal and stress temperature
conditions. Using fission yeast as a model, the present study
demonstrated that rok1 deletion leads to slow cell growth,
abnormal spore numbers, shortened spindles and centromere
separation distance, delayed actin ring assembly and blocked
migration of the kinetochores on the spindle. Integrated RNA-Seq
and bioinformatics analyses suggest that the Rok1 protein regulates
actin polymerization, thereby participating in actin ring assembly,
contraction and cytokinesis, while potentially influencing
kinetochore mobility on the spindle or contributing to the
establishment and maintenance of sister chromatid cohesion.
Notably, this novel function of Rok1 appears independent of its
canonical RNA helicase activity and may instead be linked to
differential expression of genes such as myo51, blt1,
psm1 and psc3 in rok1Δ cells.
The present study preliminarily reveals potential
functional associations between rok1 and these
differentially expressed genes. However, the specific interaction
relationships and regulatory mechanisms have not yet been directly
validated at the protein level, which represents a major limitation
of the current research. To further clarify the roles of these
genes within the Rok1 regulatory network, subsequent plans involve
screening key target genes through overexpression or knockout
experiments combined with phenotypic analysis. Building upon this
foundation, in-depth studies on protein interactions and functional
validation will be conducted.
Acknowledgements
The authors would like to acknowledge Associate
Professor Phong Tran (Department of Cell and Developmental Biology,
University of Pennsylvania) for donating the yeast strains.
Funding
Funding: The present study was funded by the Sichuan Province
Science and Technology Support Project (grant nos. 2022NZZJ0003,
22ZYZFSF0009, 2022NSFSC0107 and 23ZHSF0082).
Availability of data and materials
The transcriptome data in the present study have
been deposited in the CNCB and NCBI databases under BioProject
accession numbers PRJCA051218, https://ngdc.cncb.ac.cn/bioproject/browse/PRJCA051218
and PRJNA1208695, https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1208695/.
All other data presented in the present study may be requested from
the corresponding author.
Authors' contributions
YH and XD conceived and designed the experiments of
the present study. JH, ML and JX performed the experiments and
analyzed the data. JH, ML, JX and XD drafted the manuscript and
revised it critically. All authors read approved the final version
of the manuscript. JH and ML confirm the authenticity of all the
raw data.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Cho RJ, Campbell MJ, Winzeler EA,
Steinmetz L, Conway A, Wodicka L, Wolfsberg TG, Gabrielian AE,
Landsman D, Lockhart DJ and Davis RW: A genome-wide transcriptional
analysis of the mitotic cell cycle. Mol Cell. 2:65–73.
1998.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Cook BD, Chang F, Flor-Parra I and
Al-Bassam J: Microtubule polymerase and processive plus-end
tracking functions originate from distinct features within TOG
domain arrays. Mol Biol Cell. 30:1490–1504. 2019.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Plastino J and Blanchoin L: Dynamic
stability of the actin ecosystem. J Cell Sci.
132(jcs219832)2018.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Nema R and Kumar A: BUB1, miR-495-3p, and
E2F1/E2F8 axis is associated with poor prognosis of breast cancer
patients and infiltration of Th2 cells in the tumor
microenvironment. Cancer Biomark.
42(18758592241310109)2025.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Chu A, Liu X, Liu S, Li M, Song R, Gan L,
Wang Y, Liu Z and Sun C: RNA-seq analysis reveals key genes
associated with downregulation of APE1 in esophageal squamous cell
carcinoma. Front Genet. 16(1549371)2025.PubMed/NCBI View Article : Google Scholar
|
|
6
|
McInerny CJ: Cell cycle regulated gene
expression in yeasts. Adv Genet. 73:51–85. 2011.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Wang L, Chen K, Weng S, Xu H, Ren Y, Cheng
Q, Luo P, Zhang J, Liu Z and Han X: PI3K pathway mutation predicts
an activated immune microenvironment and better immunotherapeutic
efficacy in head and neck squamous cell carcinoma. World J Surg
Oncol. 21(72)2023.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Vejrup-Hansen R, Mizuno K, Miyabe I, Fleck
O, Holmberg C, Murray JM, Carr AM and Nielsen O:
Schizosaccharomyces pombe Mms1 channels repair of perturbed
replication into Rhp51 independent homologous recombination. DNA
Repair (Amst). 10:283–295. 2011.PubMed/NCBI View Article : Google Scholar
|
|
9
|
MacQuarrie CD, Mangione MC, Carroll R,
James M, Gould KL and Sirotkin V: The S. pombe adaptor
protein Bbc1 regulates localization of Wsp1 and Vrp1 during
endocytic actin patch assembly. J Cell Sci.
132(jcs233502)2019.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Szkotnicki L, Crutchley JM, Zyla TR,
Bardes ESG and Lew DJ: The checkpoint kinase Hsl1p is activated by
Elm1p-dependent phosphorylation. Mol Biol Cell. 19:4675–4686.
2008.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Grallert B, Kearsey SE, Lenhard M, Carlson
CR, Nurse P, Boye E and Labib K: A fission yeast general
translation factor reveals links between protein synthesis and cell
cycle controls. J Cell Sci. 113:1447–1458. 2000.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Johnson CA, Brooker HR, Gyamfi I, O'Brien
J, Ashley B, Brazier JE, Dean A, Embling J, Grimsey E, Tomlinson
AC, et al: Temperature sensitive point mutations in fission yeast
tropomyosin have long range effects on the stability and function
of the actin-tropomyosin copolymer. Biochem Biophys Res Commun.
506:339–346. 2018.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Karbstein K: Attacking a DEAD problem: The
role of DEAD-box ATPases in ribosome assembly and beyond. Methods
Enzymol. 673:19–38. 2022.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Dudin O, Merlini L, Bendezú FO, Groux R,
Vincenzetti V and Martin SG: A systematic screen for morphological
abnormalities during fission yeast sexual reproduction identifies a
mechanism of actin aster formation for cell fusion. PLoS Genet.
13(e1006721)2017.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Hayles J, Wood V, Jeffery L, Hoe KL, Kim
DU, Park HO, Salas-Pino S, Heichinger C and Nurse P: A genome-wide
resource of cell cycle and cell shape genes of fission yeast. Open
Biol. 3(130053)2013.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Maekawa H, Nakagawa T, Uno Y, Kitamura K
and Shimoda C: The ste13+ gene encoding a putative RNA helicase is
essential for nitrogen starvation-induced G1 arrest and initiation
of sexual development in the fission yeast Schizosaccharomyces
pombe. Mol Gen Genet. 244:456–464. 1994.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Forbes KC, Humphrey T and Enoch T:
Suppressors of cdc25p overexpression identify two pathways that
influence the G2/M checkpoint in fission yeast. Genetics.
150:1361–1375. 1998.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Ren L, McLean JR, Hazbun TR, Fields S,
Vander Kooi C, Ohi MD and Gould KL: Systematic two-hybrid and
comparative proteomic analyses reveal novel yeast pre-mRNA splicing
factors connected to Prp19. PLoS One. 6(e16719)2011.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Potashkin J, Kim D, Fons M, Humphrey T and
Frendewey D: Cell-division-cycle defects associated with fission
yeast pre-mRNA splicing mutants. Curr Genet. 34:153–163.
1998.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Linder P and Jankowsky E: From unwinding
to clamping-the DEAD box RNA helicase family. Nat Rev Mol Cell
Biol. 12:505–516. 2011.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Khoshnevis S, Askenasy I, Johnson MC,
Dattolo MD, Young-Erdos CL, Stroupe ME and Karbstein K: The
DEAD-box protein Rok1 orchestrates 40S and 60S ribosome assembly by
promoting the release of Rrp5 from Pre-40S ribosomes to allow for
60S maturation. PLoS Biol. 14(e1002480)2016.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Jeon S, Lim S, Ha J and Kim J:
Identification of Psk2, Skp1, and Tub4 as trans-acting factors for
uORF-containing ROK1 mRNA in Saccharomyces cerevisiae. J Microbiol.
53:616–622. 2015.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Blyth J, Makrantoni V, Barton RE, Spanos
C, Rappsilber J and Marston AL: Genes important for
Schizosaccharomyces pombe meiosis identified through a
functional genomics screen. Genetics. 208:589–603. 2018.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Lin SL, Chang D and Ying SY: Hyaluronan
stimulates transformation of androgen-independent prostate cancer.
Carcinogenesis. 28:310–320. 2007.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Zhang D, Yu W, Liu M, Qing X, Ding X and
Hou Y: Effects of the mitochondrial fission gene dnm1 deletion on
mitosis and energy metabolism in Saccharomyces cerevisiae cells. J
Beijing Normal Univ. 60:331–343. 2024.
|
|
26
|
Yu W, Yuan R, Liu M, Liu K, Ding X and Hou
Y: Effects of rpl1001 gene deletion on cell division of fission
yeast and its molecular mechanism. Curr Issues Mol Biol.
46:2576–2597. 2024.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Furuya K and Niki H: Mating, spore
dissection, and selection of diploid cells in
Schizosaccharomyces japonicus. Cold Spring Harb Protoc.
2017(prot091843)2017.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Hu Z, Chen C, Zheng X, Yuan J, Zou R and
Xie C: Establishing gene expression and knockout methods in esteya
vermicola CBS115803. Mol Biotechnol. 66:2872–2881. 2024.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Lai CJS, Tan T, Zeng SL, Xu LR, Qi LW, Liu
EH and Li P: An enzymatic protocol for absolute quantification of
analogues: Application to specific protopanoxadiol-type
ginsenosides. Green Chem. 17:2580–2586. 2015.
|
|
30
|
Brown SD and Lorenz A: Single-step marker
switching in Schizosaccharomyces pombe using a lithium
acetate transformation protocol. Bio Protoc.
6(e2075)2016.PubMed/NCBI View Article : Google Scholar
|
|
31
|
DeVore GR: Computing the Z score and
centiles for cross-sectional analysis: A practical approach. J
Ultrasound Med. 36:459–473. 2017.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Katz Y, Wang ET, Airoldi EM and Burge CB:
Analysis and design of RNA sequencing experiments for identifying
isoform regulation. Nat Methods. 7:1009–1015. 2010.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Hampton TH, Taub L, Ferreria-Fukutani K,
Stanton BA and MacKenzie TA: Analyzing qPCR data: Better practices
to facilitate rigor and reproducibility. Biochem Biophys Rep.
44(102356)2025.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Kim D, Paggi JM, Park C, Bennett C and
Salzberg SL: Graph-based genome alignment and genotyping with
HISAT2 and HISAT-genotype. Nat Biotechnol. 37:907–915.
2019.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Liao Y, Smyth GK and Shi W: featureCounts:
An efficient general purpose program for assigning sequence reads
to genomic features. Bioinformatics. 30:923–930. 2014.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Liu S, Wang Z, Zhu R, Wang F, Cheng Y and
Liu Y: Three differential expression analysis methods for RNA
sequencing: limma, EdgeR, DESeq2. J Vis Exp, 2021.
|
|
37
|
Ashburner M, Ball CA, Blake JA, Botstein
D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT,
et al: Gene ontology: Tool for the unification of biology. The gene
ontology consortium. Nat Genet. 25:25–29. 2000.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Ogata H, Goto S, Sato K, Fujibuchi W, Bono
H and Kanehisa M: KEGG: Kyoto encyclopedia of genes and genomes.
Nucleic Acids Res. 27:29–34. 1999.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Yu G, Wang LG, Han Y and He QY:
clusterProfiler: An R package for comparing biological themes among
gene clusters. OMICS. 16:284–287. 2012.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Brown SD, Audoynaud C and Lorenz A:
Intragenic meiotic recombination in Schizosaccharomyces
pombe is sensitive to environmental temperature changes.
Chromosome Res. 28:195–207. 2020.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Nunes V and Ferreira JG: From the
cytoskeleton to the nucleus: An integrated view on early spindle
assembly. Semin Cell Dev Biol. 117:42–51. 2021.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Rezig IM, Yaduma WG and McInerny CJ:
Processes controlling the contractile ring during cytokinesis in
fission yeast, including the role of ESCRT proteins. J Fungi
(Basel). 10(154)2024.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Goldstein LD, Cao Y, Pau G, Lawrence M, Wu
TD, Seshagiri S and Gentleman R: Prediction and quantification of
splice events from RNA-Seq data. PLoS One.
11(156132)2016.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Zhao Y, Li MC, Konaté MM, Chen L, Das B,
Karlovich C, Williams PM, Evrard YA, Doroshow JH and McShane LM:
TPM, FPKM, or normalized counts? A comparative study of
quantification measures for the analysis of RNA-seq data from the
NCI patient-derived models repository. J Transl Med.
19(269)2021.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Morigasaki S, Shimada K, Ikner A, Yanagida
M and Shiozaki K: Glycolytic enzyme GAPDH promotes peroxide stress
signaling through multistep phosphorelay to a MAPK cascade. Mol
Cell. 30:108–113. 2008.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Zhang Y and Barberis M: Exploring cell
cycle-mediated regulations of glycolysis in budding yeast. Front
Microbiol. 14(1270487)2023.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Boronat S, Domènech A, Carmona M,
García-Santamarina S, Bañó MC, Ayté J and Hidalgo E: Lack of a
peroxiredoxin suppresses the lethality of cells devoid of electron
donors by channelling electrons to oxidized ribonucleotide
reductase. PLoS Genet. 13(e1006858)2017.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Matsuyama A, Arai R, Yashiroda Y, Shirai
A, Kamata A, Sekido S, Kobayashi Y, Hashimoto A, Hamamoto M,
Hiraoka Y, et al: ORFeome cloning and global analysis of protein
localization in the fission yeast Schizosaccharomyces pombe.
Nat Biotechnol. 24:841–847. 2006.PubMed/NCBI View Article : Google Scholar
|
|
49
|
Xiang B, Chen ML, Gao ZQ, Mi T, Shi QL,
Dong JJ, Tian XM, Liu F and Wei GH: CCNB1 is a novel prognostic
biomarker and promotes proliferation, migration and invasion in
Wilms tumor. BMC Med Genomics. 16(189)2023.PubMed/NCBI View Article : Google Scholar
|
|
50
|
Otsubo Y, Yamashita A, Ohno H and Yamamoto
M: S. pombe TORC1 activates the ubiquitin-proteasomal
degradation of the meiotic regulator Mei2 in cooperation with Pat1
kinase. J Cell Sci. 127:2639–2646. 2014.PubMed/NCBI View Article : Google Scholar
|
|
51
|
Sasuga S, Abe R, Nikaido O, Kiyosaki S,
Sekiguchi H, Ikai A and Osada T: Interaction between pheromone and
its receptor of the fission yeast Schizosaccharomyces pombe
examined by a force spectroscopy study. J Biomed Biotechnol.
2012(804793)2012.PubMed/NCBI View Article : Google Scholar
|
|
52
|
Miyata M, Matsuoka M and Inada T:
Induction of sexual co-flocculation of heterothallic fission yeast
(Schizosaccharomyces pombe) cells by mating pheromones. J
Gen Appl Microbiol. 43:169–174. 1997.PubMed/NCBI View Article : Google Scholar
|
|
53
|
Imai Y and Yamamoto M: The fission yeast
mating pheromone P-factor: Its molecular structure, gene structure,
and ability to induce gene expression and G1 arrest in the mating
partner. Genes Dev. 8:328–338. 1994.PubMed/NCBI View Article : Google Scholar
|
|
54
|
Ilyushik E, Pryce DW, Walerych D, Riddell
T, Wakeman JA, McInerny CJ and McFarlane RJ: Psc3 cohesin of
Schizosaccharomyces pombe: Cell cycle analysis and
identification of three distinct isoforms. Biol Chem. 386:613–621.
2005.PubMed/NCBI View Article : Google Scholar
|
|
55
|
Pereira G and Schiebel E: The role of the
yeast spindle pole body and the mammalian centrosome in regulating
late mitotic events. Curr Opin Cell Biol. 13:762–769.
2001.PubMed/NCBI View Article : Google Scholar
|
|
56
|
Merlini L, Khalili B, Dudin O, Michon L,
Vincenzetti V and Martin SG: Inhibition of Ras activity coordinates
cell fusion with cell-cell contact during yeast mating. J Cell
Biol. 217:1467–1483. 2018.PubMed/NCBI View Article : Google Scholar
|
|
57
|
Tang Q, Billington N, Krementsova EB,
Bookwalter CS, Lord M and Trybus KM: A single-headed fission yeast
myosin V transports actin in a tropomyosin-dependent manner. J Cell
Biol. 214:167–179. 2016.PubMed/NCBI View Article : Google Scholar
|
|
58
|
Wang N, Lo Presti L, Zhu YH, Kang M, Wu Z,
Martin SG and Wu JQ: The novel proteins Rng8 and Rng9 regulate the
myosin-V Myo51 during fission yeast cytokinesis. J Cell Biol.
205:357–375. 2014.PubMed/NCBI View Article : Google Scholar
|
|
59
|
Tyree ZL, Bellingham-Johnstun K,
Martinez-Baird J and Laplante C: The myosin-V Myo51 and
alpha-actinin Ain1p cooperate during contractile ring assembly and
disassembly in fission yeast cytokinesis. J Fungi (Basel).
10(647)2024.PubMed/NCBI View Article : Google Scholar
|
|
60
|
Win TZ, Gachet Y, Mulvihill DP, May KM and
Hyams JS: Two type V myosins with non-overlapping functions in the
fission yeast Schizosaccharomyces pombe: Myo52 is concerned
with growth polarity and cytokinesis, Myo51 is a component of the
cytokinetic actin ring. J Cell Sci. 114:69–79. 2001.PubMed/NCBI View Article : Google Scholar
|
|
61
|
Laplante C, Berro J, Karatekin E,
Hernandez-Leyva A, Lee R and Pollard TD: Three myosins contribute
uniquely to the assembly and constriction of the fission yeast
cytokinetic contractile ring. Curr Biol. 25:1955–1965.
2015.PubMed/NCBI View Article : Google Scholar
|
|
62
|
Goss JW, Kim S, Bledsoe H and Pollard TD:
Characterization of the roles of Blt1p in fission yeast
cytokinesis. Mol Biol Cell. 25:1946–1957. 2014.PubMed/NCBI View Article : Google Scholar
|
|
63
|
Jourdain I, Brzezińska EA and Toda T:
Fission yeast Nod1 is a component of cortical nodes involved in
cell size control and division site placement. PLoS One.
8(e54142)2013.PubMed/NCBI View Article : Google Scholar
|
|
64
|
Birot A, Tormos-Pérez M, Vaur S, Feytout
A, Jaegy J, Alonso Gil D, Vazquez S, Ekwall K and Javerzat JP: The
CDK Pef1 and protein phosphatase 4 oppose each other for regulating
cohesin binding to fission yeast chromosomes. Elife.
9(e50556)2020.PubMed/NCBI View Article : Google Scholar
|
|
65
|
Nakano K, Arai R and Mabuchi I: Small
GTPase Rho5 is a functional homologue of Rho1, which controls cell
shape and septation in fission yeast. FEBS Lett. 579:5181–5186.
2005.PubMed/NCBI View Article : Google Scholar
|
|
66
|
Yoon HJ, Feoktistova A, Wolfe BA, Jennings
JL, Link AJ and Gould KL: Proteomics analysis identifies new
components of the fission and budding yeast anaphase-promoting
complexes. Curr Biol. 12:2048–2054. 2002.PubMed/NCBI View Article : Google Scholar
|
|
67
|
May KM, Paldi F and Hardwick KG: Fission
yeast Apc15 stabilizes MCC-Cdc20-APC/C complexes, ensuring
efficient Cdc20 ubiquitination and checkpoint arrest Curr. Biol.
27:1221–1228. 2017.PubMed/NCBI View Article : Google Scholar
|
|
68
|
Fujita I, Yamashita A and Yamamoto M:
Dynactin and Num1 cooperate to establish the cortical anchoring of
cytoplasmic dynein in S. pombe. J Cell Sci. 128:1555–1567.
2015.PubMed/NCBI View Article : Google Scholar
|
|
69
|
Scheffler K, Minnes R, Fraisier V,
Paoletti A and Tran PT: Microtubule minus end motors kinesin-14 and
dynein drive nuclear congression in parallel pathways. J Cell Biol.
209:47–58. 2015.PubMed/NCBI View Article : Google Scholar
|
|
70
|
Kimata Y, Kitamura K, Fenner N and Yamano
H: Mes1 controls the meiosis I to meiosis II transition by
distinctly regulating the anaphase-promoting complex/cyclosome
coactivators Fzr1/Mfr1 and Slp1 in fission yeast. Mol Biol Cell.
22:1486–1494. 2011.PubMed/NCBI View Article : Google Scholar
|
|
71
|
Blanco MA, Pelloquin L and Moreno S:
Fission yeast mfr1 activates APC and coordinates meiotic nuclear
division with sporulation. J Cell Sci. 114:2135–2143.
2001.PubMed/NCBI View Article : Google Scholar
|
|
72
|
Gregan J, Riedel CG, Pidoux AL, Katou Y,
Rumpf C, Schleiffer A, Kearsey SE, Shirahige K, Allshire RC and
Nasmyth K: The kinetochore proteins Pcs1 and Mde4 and
heterochromatin are required to prevent merotelic orientation. Curr
Biol. 17:1190–1200. 2007.PubMed/NCBI View Article : Google Scholar
|
|
73
|
Azzopardi M, Farrugia G and Balzan R:
Cell-cycle involvement in autophagy and apoptosis in yeast. Mech
Ageing Dev. 161:211–224. 2017.PubMed/NCBI View Article : Google Scholar
|
|
74
|
Ucisik-Akkaya E, Leatherwood JK and Neiman
AM: A genome-wide screen for sporulation-defective mutants in
Schizosaccharomyces pombe. G3 (Bethesda). 4:1173–1182.
2014.PubMed/NCBI View Article : Google Scholar
|
|
75
|
al-Khodairy F, Fotou E, Sheldrick KS,
Griffiths DJ, Lehmann AR and Carr AM: Identification and
characterization of new elements involved in checkpoint and
feedback controls in fission yeast. Mol Biol Cell. 5:147–160.
1994.PubMed/NCBI View Article : Google Scholar
|
|
76
|
Akera T, Goto Y, Sato M, Yamamoto M and
Watanabe Y: Mad1 promotes chromosome congression by anchoring a
kinesin motor to the kinetochore. Nat Cell Biol. 17:1124–1133.
2015.PubMed/NCBI View Article : Google Scholar
|
|
77
|
Tomonaga T, Nagao K, Kawasaki Y, Furuya K,
Murakami A, Morishita J, Yuasa T, Sutani T, Kearsey SE, Uhlmann F,
et al: Characterization of fission yeast cohesin: Essential
anaphase proteolysis of Rad21 phosphorylated in the S phase. Genes
Dev. 14:2757–2770. 2000.PubMed/NCBI View Article : Google Scholar
|
|
78
|
Liu Y, Min Y, Liu Y and Watanabe Y:
Phosphorylation of Rec8 cohesin complexes regulates
mono-orientation of kinetochores in meiosis I. Life Sci Alliance.
7(e202302556)2024.PubMed/NCBI View Article : Google Scholar
|
|
79
|
Kurokawa Y and Murayama Y: DNA binding by
the Mis4Scc2 loader promotes topological DNA entrapment
by the cohesin ring. Cell Rep. 33(108357)2020.PubMed/NCBI View Article : Google Scholar
|
|
80
|
Furuya K, Takahashi K and Yanagida M:
Faithful anaphase is ensured by Mis4, a sister chromatid cohesion
molecule required in S phase and not destroyed in G1 phase. Genes
Dev. 12:3408–3418. 1998.PubMed/NCBI View Article : Google Scholar
|
|
81
|
Zhao D, Liu XM, Yu ZQ, Sun LL, Xiong X,
Dong MQ and Du LL: Atg20- and Atg24-family proteins promote
organelle autophagy in fission yeast. J Cell Sci. 129:4289–4304.
2016.PubMed/NCBI View Article : Google Scholar
|
|
82
|
Zahedi Y, Durand-Dubief M and Ekwall K:
High-throughput flow cytometry combined with genetic analysis
brings new insights into the understanding of chromatin regulation
of cellular quiescence. Int J Mol Sci. 21(9022)2020.PubMed/NCBI View Article : Google Scholar
|
|
83
|
Mukaiyama H, Kajiwara S, Hosomi A,
Giga-Hama Y, Tanaka N, Nakamura T and Takegawa K:
Autophagy-deficient Schizosaccharomyces pombe mutants
undergo partial sporulation during nitrogen starvation.
Microbiology (Reading). 155:3816–3826. 2009.PubMed/NCBI View Article : Google Scholar
|
|
84
|
Balasubramanian MK, Feoktistova A,
McCollum D and Gould KL: Fission yeast Sop2p: A novel and
evolutionarily conserved protein that interacts with Arp3p and
modulates profilin function. EMBO J. 15:6426–6437. 1996.PubMed/NCBI
|
|
85
|
Codlin S, Haines RL and Mole SE: btn1
affects endocytosis, polarization of cholesterol-enriched membrane
domains, and polarized growth in Schizosaccharomyces pombe.
Traffic. 9:936–950. 2008.PubMed/NCBI View Article : Google Scholar
|