Introduction
Congenital heart disease (CHD), characterized by the
developmental abnormality of the heart and intrathoracic great
vessels, is the most common birth defect in humans worldwide,
occurring in approximately 1% of live births, and is the major
non-infectious cause of infant morbidity and mortality, accounting
for approximately 30% of neonatal deaths resulting from
developmental malformations (1).
According to the specific anatomic lesions, CHD is categorized into
at least 21 clinical types, encompassing atrial septal defect,
ventricular septal defect, atrioventricular septal defect,
tetraology of Fallot, patent ductus arteriosus, transposition of
the great arteries, right ventricular outflow tract obstruction,
aortic stenosis, pulmonary atresia, coronary artery deformation,
tricuspid atresia, Ebstein’s anomaly of the tricuspid valve, double
outlet right ventricle, hypoplastic left heart syndrome,
interrupted aortic arch and total anomalous pulmonary venous
connection (1). If unrepaired,
these cardiovascular deformations may contribute to poor exercise
tolerance, degraded life quality, delayed fetal brain development,
infective endocarditis, metabolic disorders, pulmonary
hypertension, congestive heart failure, thromboembolic stroke,
arrhythmias and even sudden cardiac death (2–13).
Despite its high prevalence and important clinical significance,
the etiology for CHD in an overwhelming majority of patients
remains unclear.
In mammals, the heart is the first organ to form
during embryogenesis (14).
Cardiogenesis is a complex and dynamic biological process that
requires the orchestration of cardiac cell commitment,
differentiation, proliferation and migration, and both
environmental and genetic risk factors may perturb this exquisite
temporal and spatial cooperation, leading to a wide variety of CHD
(15–22). A growing body of evidence
underscores the key role of cardiac transcription factors in
embryonic cardiovascular morphogenesis, and a long list of
mutations in the genes coding for cardiac transcription factors,
including the NK and GATA families, have been associated with CHD
(23–41). However, CHD is a genetically
heterogeneous disease and the genetic defects responsible for CHD
in the majority of patients remain unknown.
Previous studies have indicated that the cardiac
transcription factor, PITX2c, a member of the bicoid-like
homeodomain family of transcription factors, is essential for
normal cardiovascular development (42–49). The PITX2c gene is
predominantly expressed in the embryonic and adult hearts, playing
a crucial role in the embryogenesis of the left atrium, cardiac
conduction system and pulmonary venous myocardium (50). In mice, targeted deletion of
PITX2c has been shown to lead to embryonic lethality due to
distinct types of CHD, including atrial isomerism, double-outlet
right ventricle, atrial septal defect, ventricular septal defect,
transposition of the great arteries, and abnormal aortic arch, as
well as incomplete closure of the body wall (42). In humans, PITX2c mutations have
been implicated in congenital atrial septal defect, ventricular
septal defect, double outlet of the right ventricle and atrial
fibrillation (51–54). These findings justify screening
PITX2c in other cohorts of patients with CHD.
Materials and methods
Study subjects
A cohort of 170 unrelated neonates with CHD was
recruited from the Chinese Han population. The available relatives
of the mutation carriers were also included. The patients were
evaluated by individual and familial histories, review of the
medical records, complete physical examination, 12-lead
electrocardiogram and 2-dimensional transthoracic echocardiography
with a color flow Doppler. Transesophageal echocardiography and
cardiac catheterization were performed in some patients. Most
patients underwent cardiac surgery or catheter-based repair. The
patients with known chromosomal abnormalities or syndromic
cardiovascular defects were excluded from the study. Clinical
investigations were carried out by cardiologists who had no
knowledge of the genotype.
A total of 200 unrelated, ethnically matched healthy
individuals randomly enlisted from the individuals undergoing
routine physical examinations were used as the control subjects.
According to the reviews of medical histories and analyses of the
echocardiographic records, the control individuals had no CHD. The
ethnic origin of a participant was ascertained by a combination of
self-reported ethnicity and a personal questionnaire asking
questions regarding birthplace, language, religion and
ancestry.
Peripheral venous blood specimens from patients with
CHD and control individuals were prepared. The study protocol was
reviewed and approved by the local institutional ethics committee
and written informed consent was obtained from the parents or
guardians of the participants prior to enrollment in the study.
Genetic analysis of human PITX2c
Genomic DNA was extracted from the blood lymphocytes
of each participant using the Wizard Genomic DNA Purification kit
(Promega Corp., Madison, WI, USA). The PITX2c gene was
sequenced initially in 170 unrelated neonates with CHD, and the
genotyping of PITX2c was subsequently performed in the
available relatives of the mutation carriers and the 200 unrelated
control individuals. The referential genomic DNA sequence of
PITX2c was derived from GenBank (Accession no. NC_000004),
which was at the National Center for Biotechnical Information
(NCBI; http://www.ncbi.nlm.nih.gov/).
The primer pairs used to amplify all the coding
exons and exon-intron boundaries of PITX2c by polymerase
chain reaction (PCR) were designed as previously described
(53). PCR was performed using
HotStar Taq DNA Polymerase (Qiagen GmbH, Hilden, Germany) on a
Veriti Thermal Cycler (Applied Biosystems, Foster, CA, USA), with
standard conditions and concentrations of reagents. Amplified
products were analyzed on 1% agarose gels stained with ethidium
bromide and purified using the QIAquick Gel Extraction kit (Qiagen
GmbH). Both strands of each PCR product were sequenced with a
BigDye® Terminator v3.1 Cycle Sequencing kit under an
ABI PRISM 3130 XL DNA Analyzer (both from Applied Biosystems). The
sequencing primers were the same as those used for the
above-mentioned specific region amplification. The DNA sequences
were viewed and analyzed with DNA Sequencing Analysis
Software® v5.1 (Applied Biosystems). The variant was
validated by re-sequencing an independent PCR-generated amplicon
from the same subject. Additionally, for an identified sequence
variant, the Exome Variant Server (EVS; http://evs.gs.washington.edu/EVS) and the NCBI single
nucleotide polymorphism (SNP; http://www.ncbi.nlm.nih.gov/SNP) online databases were
queried to confirm its novelty.
Alignment of multiple PITX2c protein
sequences among species
Multiple PITX2c protein sequences across various
species were aligned using the online program, MUSCLE, version 3.6
(http://www.ncbi.nlm.nih.gov/homologene?cmd=Retrieve&dopt=MultipleAlignment&list_uids=55454).
Prediction of the pathogenic potential of
a PITX2c sequence variation
The disease-causing potential of a PITX2c
sequence variation was predicted by MutationTaster (http://www.mutationtaster.org), which automatically
yielded a probability for the variation to be either a pathogenic
mutation or a benign polymorphism. Notably, the P-value used here
is the probability of the correct prediction rather than the
probability of error as used in t-test statistics (i.e., a value
close to 1 indicates high accuracy of the prediction). Besides,
another online program PolyPhen-2 (http://genetics.bwh.harvard.edu/pph2) was also
utilized to evaluate the causative likeliness of a variant.
Expression plasmids and site-directed
mutagenesis
The recombinant expression plasmid PITX2c-pcDNA4,
which was constructed by Strungaru et al (55), was a gift from Professor Georges
Christé, from Physiopathologie des troubles du rythme cardiaque,
Faculté de Pharmacie de Lyon, Université Lyon 1, Lyon, France. The
atrial natriuretic factor (ANF)-luciferase reporter plasmid, which
contains the 2600-bp 5′-flanking region of the ANF gene,
namely ANF(-2600)-Luc, was kindly provided by Dr Ichiro Shiojima,
from the Department of Cardiovascular Science and Medicine, Chiba
University Graduate School of Medicine (Chiba, Japan). Each of the
identified variations was introduced into wild-type PITX2c
using a QuickChange II XL Site-Directed Mutagenesis kit
(Stratagene, La Jolla, CA, USA) with a complementary pair of
primers. The mutants were sequenced to confirm the desired
mutations and to exclude any other sequence variations.
Luciferase reporter gene assay
Chinese hamster ovary (CHO) cells were cultured in
Dulbecco’s modified Eagle’s medium supplemented with 10% fetal calf
serum, as well as 100 U/ml penicillin and 100 g/ml streptomycin.
The ANF(-2600)-Luc reporter construct and an internal control
reporter plasmid, pGL4.75 (hRluc/CMV; Promega), were used in
transient transfection assays to explore the transactivational
activity of the PITX2c mutant. The CHO cells were transfected with
2 μg of the wild-type PITX2c-pcDNA4 or mutant PITX2c-pcDNA4 (R91Q
or T129S) or the empty vector pcDNA4, 2.0 μg of ANF(-2600)-Luc
reporter construct, and 0.04 μg of pGL4.75 control reporter vector
using Lipofectamine 2000 Transfection Reagent (Invitrogen,
Carlsbad, CA, USA). For co-transfection experiments, 1 μg of
wild-type PITX2c-pcDNA4, 1 μg of mutant PITX2c-pcDNA4 (R91Q or
T129S), 2.0 μg of ANF(-2600)-Luc, and 0.04 μg of pGL4.75 were used.
The transfected cells were incubated for 24 h, then lysed and
assayed for reporter activities. Firefly luciferase and
Renilla luciferase activities were measured with the
Dual-Glo luciferase assay system (Promega). The activity of the
ANF promoter was presented as fold activation of Firefly
luciferase relative to Renilla luciferase. Three independent
experiments were conducted at minimum for wild-type or mutant
PITX2c.
Statistical analysis
Experimental data are expressed as the means ±
standard deviations. Continuous variables were tested for normality
of distribution, and the Student’s unpaired t-test was used for the
comparison of numeric variables between 2 groups. A comparison of
the categorical variables between 2 groups was performed using
Pearson’s χ2 test or Fisher’s exact test where
appropriate. A two-tailed P-value <0.05 was considered to
indicate a statistically significant difference.
Results
Baseline characteristics of the study
population
A cohort of 170 unrelated neonates with CHD was
enrolled and clinically evaluated in contrast to a total of 200
unrelated, ethnically-matched healthy individuals used as the
controls. All the participants had no established environmental
risk factors for CHD, such as maternal illness and drug use in the
first trimester of pregnancy, parental smoking, and long-term
exposure to toxicants and ionizing radiation. The baseline clinical
characteristics of the 170 unrelated CHD patients are summarized in
Table I.
 | Table IBaseline clinical characteristics of
the 170 unrelated neonates with congenital heart disease. |
Table I
Baseline clinical characteristics of
the 170 unrelated neonates with congenital heart disease.
| Parameter | No. or
quantity | Percentage or
range |
|---|
| Male | 89 | 52.4 |
| Age (days) | 12.6±8.5 | 1–26 |
| Birth weight
(kg) | 3.1±0.8 | 1.6–5.5 |
| Positive family
history | 51 | 30 |
| Distribution of
different types of CHD | | |
| Isolated CHD | 84 | 49.4 |
| VSD | 21 | 12.4 |
| PDA | 14 | 8.2 |
| ASD | 10 | 5.9 |
| PS | 9 | 5.3 |
| TGA | 7 | 4.1 |
| AVSD | 6 | 3.5 |
| COA | 5 | 2.9 |
| DORV | 5 | 2.9 |
| PTA | 3 | 1.8 |
| TAPVC | 2 | 1.2 |
| HLHS | 1 | 0.6 |
| PA | 1 | 0.6 |
| Complex CHD | 86 | 50.6 |
| TGA + VSD | 27 | 15.9 |
| TOF | 15 | 8.8 |
| VSD + PDA | 12 | 7.1 |
| ASD + VSD | 9 | 5.3 |
| PDA + TGA | 5 | 2.9 |
| VSD + DORV | 5 | 2.9 |
| ASD + TGA | 5 | 2.9 |
| ASD + PDA | 4 | 2.4 |
| ASD + VSD +
DORV | 2 | 1.2 |
| IAA + VSD | 1 | 0.6 |
| ASD + VSD +
PDA | 1 | 0.6 |
| Treatment | | |
| Surgical
repair | 110 | 64.7 |
| Follow-up | 60 | 35.3 |
PITX2c mutation
All the exons and splice junction sites of the
PITX2c gene was sequenced in the 170 unrelated neonates with
CHD, and 2 heterozygous sequence variations in PITX2c weree
identified in 2 out of the 170 patients, with a mutational
prevalence of approximately 1.18% based on the patient population.
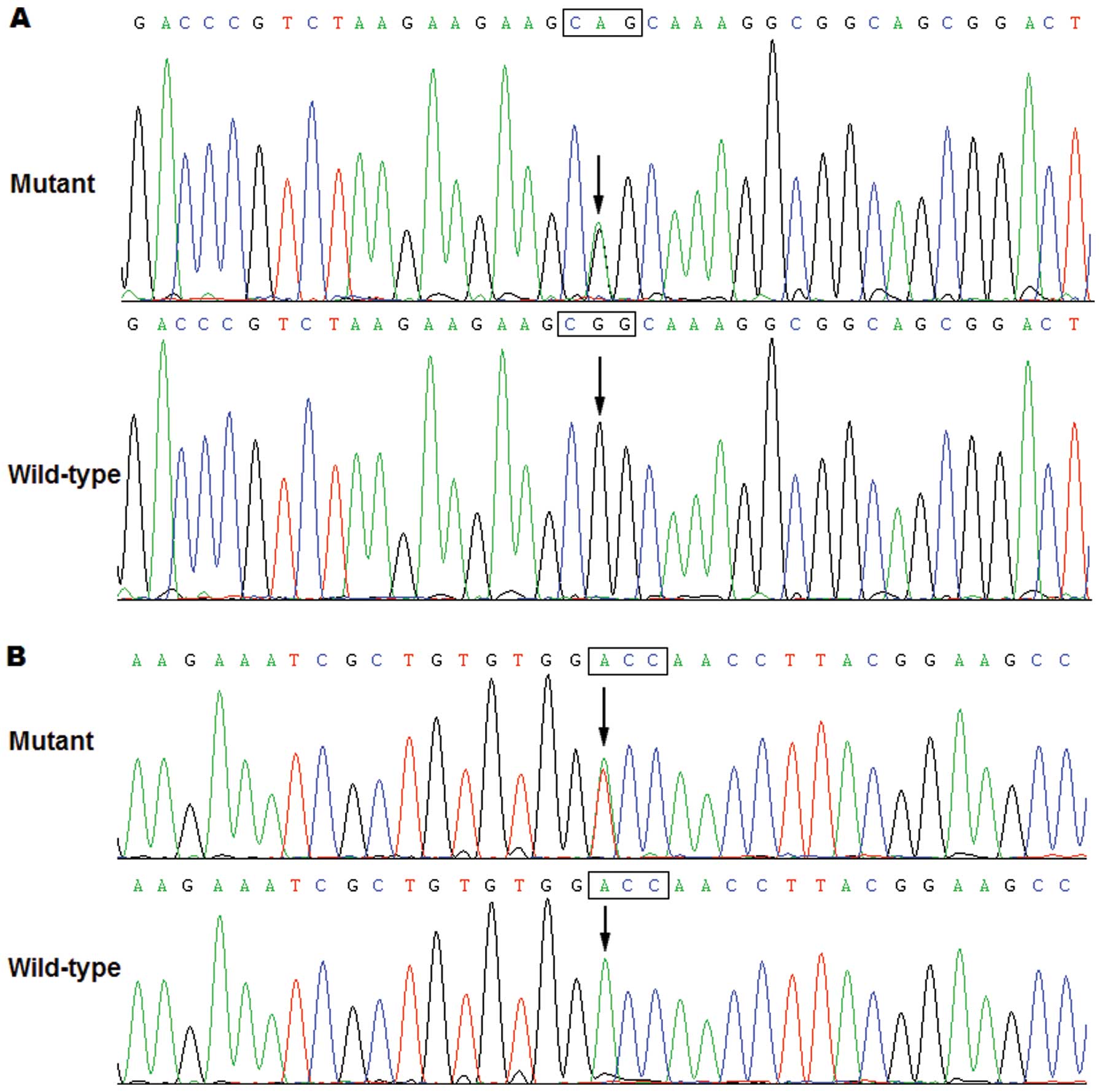
Specifically, a substitution of adenine for guanine at the second
nucleotide of codon 91 of the PITX2c gene (c.272G>A),
predicting the transition of arginine into glutamine at amino acid
91 (p.R91Q), was identified in a neonate with transition of great
arteries and ventricular septal defect. A change of adenine into
thymine at the first nucleotide of codon 129 of the PITX2c
gene (c.385A>T), equivalent to the transversion of threonine
into serine at amino acid 129 (p.T129S), was detected in another
newborn with transition of great arteries and ventricular septal
defect. The sequence electropherograms showing the identified
heterozygous PITX2c variations compared with the
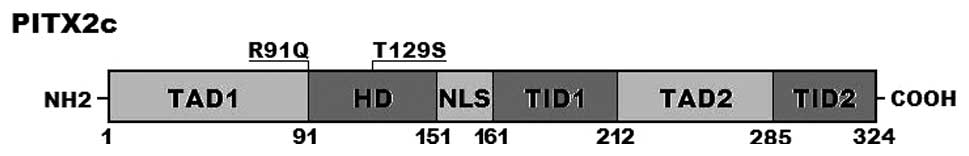
corresponding control sequences are shown in Fig. 1. A schematic diagram of
PITX2c showing the structural domains (56,57) and the locations of the detected
mutations is presented in Fig. 2.
The mutation was neither observed in 400 control chromosomes nor
reported in the EVS and NCBI SNP databases, which were consulted
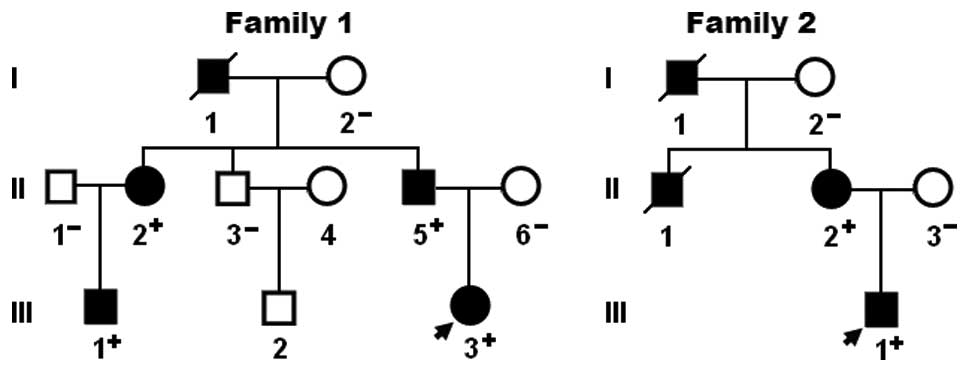
again on January 12, 2014. A genetic scan of the available family
members of the mutation carriers revealed that in each family the
mutation was present in all affected family members alive, but
absent in the unaffected family members examined. An analysis of
the pedigrees revealed that in each family, the mutation
co-segregated with CHD transmitted as an autosomal dominant trait
with complete penetrance. Atrial fibrillation was confirmed by the
early electrocardiograms in patients I-1 and II-1 from family 1.
The pedigree structures of the families are shown in Fig. 3. The phenotypic characteristics
and the results of genetic screening of the affected family members
are listed in Table II.
 | Table IIPhenotypic characteristics and status
of the PITX2c mutations in the affected family members. |
Table II
Phenotypic characteristics and status
of the PITX2c mutations in the affected family members.
| Subject
Information | Phenotype | Genotype (PITX2c
mutation) |
|---|
|
|
|---|
| Identity | Gender | Agea | CHD | AF |
|---|
| Family 1 | | | | | R91Q |
| I-1 | M | 52b | VSD | + | NA |
| II-1 | F | 31 | VSD | + | +/− |
| II-5 | M | 26 | VSD | − | +/− |
| III-1 | M | 2 | VSD | − | +/− |
| III-3 | F | 0 | TGA, VSD | − | +/− |
| Family 2 | | | | | T129S |
| I-1 | M | 49b | VSD | − | NA |
| II-1 | M | 0b | TGA, VSD | − | NA |
| II-2 | F | 25 | VSD | − | +/− |
| III-1 | M | 0 | TGA, VSD | − | +/− |
Alignment of multiple PITX2c protein
sequences
A cross-species alignment of multiple PITX2c protein
sequences displayed that the affected amino acids were completely
conserved evolutionarily (Fig.
4), indicating that the amino acids are functionally
important.
Causative potential of the PITX2c
variations
The PITX2c sequence variations of c.272G>A
and c.385A>T were both predicted to be disease-causing by
MutationTaster, with the same P-value of 1.000. No SNPs in the
altered regions were found in the MutationTaster database. In
addition, these 2 amino acid substitutions (p.R91Q and p.T129S)
were also predicted to be possibly damaging by PolyPhen-2, with the
same scores of 0.995 (sensitivity, 0.68; specificity, 0.97) for
p.R91Q and p.T129S.
Functional defect associated with PITX2c
mutations
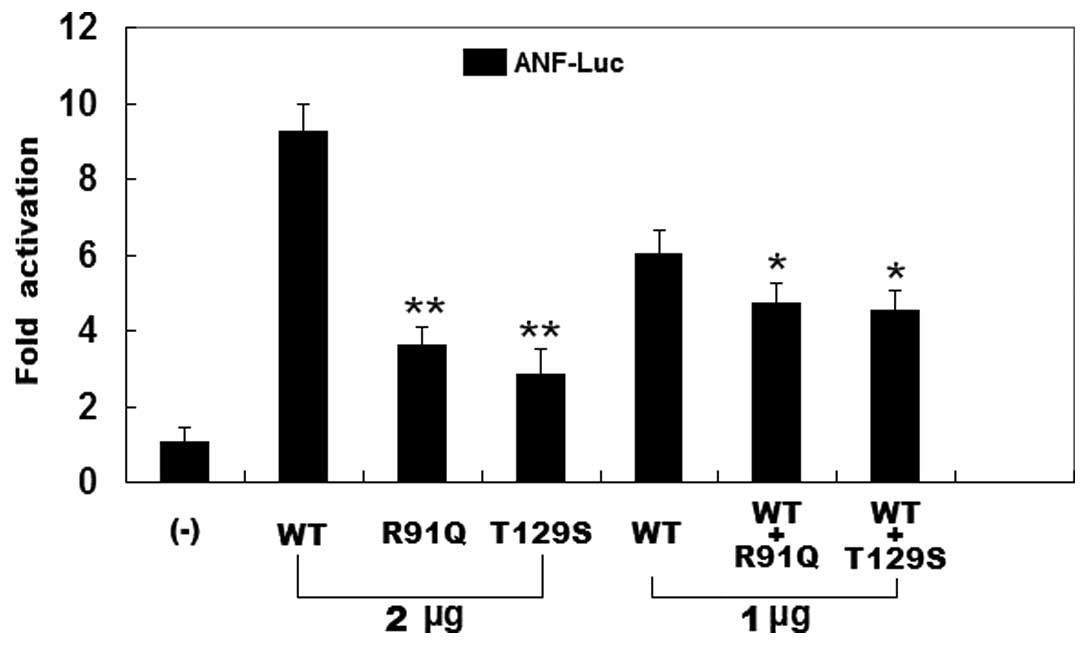
As shown in Fig.
5, the same amounts of wild-type PITX2c (2 μg), R91Q-mutant
PITX2c (2 μg) and T129S-mutant PITX2c (2 μg) activated the
ANF promoter by approximately a 9-, 4- and 3-fold increase,
respectively, when compared with the empty plasmid. When the same
amount of wild-type PITX2c (1 μg) was transfected in combination
with mutant PITX2c (1 μg of R91Q-mutant or 1 μg of T129S-mutant),
the induced activation of the ANF promoter was increased by
approximately 5-fold compared with the empty plasmid. These results
indicate that the PITX2c mutants are associated with significantly
reduced activation activity compared with their wild-type
counterpart.
Discussion
The human PITX2c gene maps to chromosome
4q25, coding for a protein of 324 amino acids (58). PITX2c is predominantly expressed
in the developing and adult heart and is required for normal
cardiovascular development (59).
In the present study, 2 novel heterozygous mutations of PITX2c,
p.R91Q and p.T129S, were identified in 2 newborns with CHD. The
mutant alleles were absent in the 400 reference chromosomes from an
ethnically matched control population. Cross-species alignment of
PITX2c protein sequences revealed that the altered amino acids were
completely conserved evolutionarily. These 2 variations were
predicted to be pathogenic by both MutationTaster and PolyPhen-2,
and functional analysis demonstrated that the mutants were
associated with a significantly reduced transcriptional activity.
Therefore, it is likely that functionally compromised PITX2c
predisposes to CHD in these mutation carriers.
PITX2 is a member of the paired-like homeobox
transcription factor family. To date, 4 distinct PITX2 transcripts,
generated by differential mRNA splicing and alternative promoter
usage, have been reported, of which PITX2a, PITX2b and PITX2c
differ only in their amino-termini and have been identified in
humans, mice, chicks, zebrafish and Xenopus, whereas the 4th
isoform, PITX2d, which lacks the whole amino-terminal domain and
most homeodomains, has only been identified in humans. The unique
amino-termini of PITX2a, PITX2b and PITX2c may have an effect on
their transcriptional activity in a cell-type and
promoter-dependent manner. The homeodomain may recognize and bind
to specific DNA sequences (5′-TAATCC-3′), which is responsible for
DNA binding and interaction with other transcription factors
(60). The PITX2c mutations of
p.R91Q and p.T129S identified in the present study are located in
the homeodomain, and thus they may be expected to exert an effect
on the transcriptional activity of PITX2c by perturbing its DNA
binding.
PITX2c is an upstream regulator of multiple target
genes expressed in the heart during embryogenesis, including the
ANF gene (61). Therefore,
the functional characteristics of a PITX2c mutation can be
investigated by the assay of the transcriptional activity of the
ANF promoter in cells expressing PITX2c mutant in contrast
to its wild-type counterpart. In this study, the functional effect
of 2 novel PITX2c mutations identified in patients with CHD was
characterized by transcriptional activity analysis and the results
demonstrated that the mutants were associated with a significantly
decreased transcriptional activity on the downstream gene,
ANF, suggesting that PITX2c loss-of-function mutations are
potentially an alternative pathological mechanism of CHD.
The fact that dysfunctional PITX2c confers enhanced
susceptibility to CHD has been substantiated in animal models. In
mice, PITX2c is expressed specifically in the trabecular and
septal myocardium with a strong expression bias in the myocardium
associated with endocardial cushions of the atrioventricular canal
and outflow tract, which are crucial for cardiac septation
(62), and the targeted
disruption of the PITX2c gene has been shown to result in
embryonic lethality due to cardiovascular defects, including atrial
isomerism, ventricular septal defect, double-outlet right
ventricle, atrial septal defect and abnormal aortic arch (42). In Xenopus, the knockdown of
PITX2c by the use of chemically modified antisense oligonucleotides
has ben shown to lead to aberrant cardiac morphology, of which the
most commonly observed cardiac deformity was a failure of rightward
migration of the outflow tract, occurring in 23% of embryos
injected with the PITX2c antisense oligonucleotides. Other cardiac
deformations caused by PITX2c-targeted mRNA interference included
anomalies of atrial septation, extracellular matrix restriction,
relative atrial-ventricular chamber positioning and restriction of
ventricular development (43).
These experimental findings highlight an exquisite sensitivity of
the developing cardiovascular system to the level of PITX2c.
Notably, mutant PITX2c has been causally linked to
lone or familial atrial fibrillation (53,54). In this study, 2 novel PITX2c
mutations were identified in 2 families with ventricular septal
defect, of which 2 family members also had atrial fibrillation, and
2 family members also had transition of the great arteries.
Different genetic backgrounds and epigenetic modifiers may account
for the pronounced phenotypic heterogeneity among these mutation
carriers.
In conclusion, the current study associates PITX2c
loss-of-function mutations with transition of the great arteries
and ventricular septal defect in humans, which provides additional
evidence supporting that the fact that PITX2c plays an important
role in cardiovascular development.
Acknowledgements
The authors are thankful to the participants for
their dedication to the study. This study was supported by grants
from the National Natural Science Foundation of China (81270161 and
81271927) and the Science and Technology Foundation of the Medical
College of Shanghai Jiao Tong University (13XJ10070).
References
|
1
|
Roger VL, Go AS, Lloyd-Jones DM, Benjamin
EJ, Berry JD, Borden WB, Bravata DM, Dai S, Ford ES, Fox CS,
Fullerton HJ, Gillespie C, Hailpern SM, Heit JA, Howard VJ, Kissela
BM, Kittner SJ, Lackland DT, Lichtman JH, Lisabeth LD, Makuc DM,
Marcus GM, Marelli A, Matchar DB, Moy CS, Mozaffarian D, Mussolino
ME, Nichol G, Paynter NP, Soliman EZ, Sorlie PD, Sotoodehnia N,
Turan TN, Virani SS, Wong ND, Woo D and Turner MB; American Heart
Association Statistics Committee and Stroke Statistics
Subcommittee. Heart disease and stroke statistics-2012 update: a
report from the American Heart Association. Circulation.
125:e2–e220. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Bang JS, Jo S, Kim GB, Kwon BS, Bae EJ,
Noh CI and Choi JY: The mental health and quality of life of adult
patients with congenital heart disease. Int J Cardiol. 170:49–53.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Idorn L, Jensen AS, Juul K, Overgaard D,
Nielsen NP, Sørensen K, Reimers JI and Søndergaard L: Quality of
life and cognitive function in Fontan patients, a population-based
study. Int J Cardiol. 168:3230–3235. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Kröönström LA, Johansson L, Zetterström
AK, Dellborg M, Eriksson P and Cider A: Muscle function in adults
with congenital heart disease. Int J Cardiol. 170:358–363.
2014.
|
|
5
|
Lu JC, Cotts TB and Dorfman AL: Diastolic
function and patient-reported quality of life for adolescents and
adults with repaired tetralogy of Fallot: a tissue Doppler study.
Pediatr Cardiol. 33:618–624. 2012. View Article : Google Scholar
|
|
6
|
Broberg CS, Van Woerkom RC, Swallow E,
Dimopoulos K, Diller GP, Allada G and Gatzoulis MA: Lung function
and gas exchange in Eisenmenger syndrome and their impact on
exercise capacity and survival. Int J Cardiol. 171:73–77. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Donofrio MT, Duplessis AJ and
Limperopoulos C: Impact of congenital heart disease on fetal brain
development and injury. Curr Opin Pediatr. 23:502–511. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Rushani D, Kaufman JS, Ionescu-Ittu R,
Mackie AS, Pilote L, Therrien J and Marelli AJ: Infective
endocarditis in children with congenital heart disease: cumulative
incidence and predictors. Circulation. 128:1412–1419. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Passarella G, Trifirò G, Gasparetto M,
Moreolo GS and Milanesi O: Disorders in glucidic metabolism and
congenital heart diseases: detection and prevention. Pediatr
Cardiol. 34:931–937. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Martínez-Quintana E, Rodríguez-González F
and Nieto-Lago V: Subclinical hypothyroidism in grown-up congenital
heart disease patients. Pediatr Cardiol. 34:912–917.
2013.PubMed/NCBI
|
|
11
|
Zomer AC, Vaartjes I, van der Velde ET, de
Jong HM, Konings TC, Wagenaar LJ, Heesen WF, Eerens F, Baur LH,
Grobbee DE and Mulder BJ: Heart failure admissions in adults with
congenital heart disease; risk factors and prognosis. Int J
Cardiol. 168:2487–2493. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Ueda A, Adachi I, McCarthy KP, Li W, Ho SY
and Uemura H: Substrates of atrial arrhythmias: histological
insights from patients with congenital heart disease. Int J
Cardiol. 168:2481–2486. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Perry JC: Sudden cardiac death and
malignant arrhythmias: the scope of the problem in adult congenital
heart patients. Pediatr Cardiol. 33:484–490. 2012. View Article : Google Scholar
|
|
14
|
Olson EN: Gene regulatory networks in the
evolution and development of the heart. Science. 313:1922–1927.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Lee LJ and Lupo PJ: Maternal smoking
during pregnancy and the risk of congenital heart defects in
offspring: a systematic review and metaanalysis. Pediatr Cardiol.
34:398–407. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Ackerman C, Locke AE, Feingold E, Reshey
B, Espana K, Thusberg J, Mooney S, Bean LJ, Dooley KJ, Cua CL,
Reeves RH, Sherman SL and Maslen CL: An excess of deleterious
variants in VEGF-A pathway genes in Down-syndrome-associated
atrioventricular septal defects. Am J Hum Genet. 91:646–659. 2012.
View Article : Google Scholar
|
|
17
|
Tan HL, Glen E, Töpf A, Hall D, O’Sullivan
JJ, Sneddon L, Wren C, Avery P, Lewis RJ, ten Dijke P, Arthur HM,
Goodship JA and Keavney BD: Nonsynonymous variants in the SMAD6
gene predispose to congenital cardiovascular malformation. Hum
Mutat. 33:720–727. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Soemedi R, Wilson IJ, Bentham J, Darlay R,
Töpf A, Zelenika D, Cosgrove C, Setchfield K, Thornborough C,
Granados-Riveron J, Blue GM, Breckpot J, Hellens S, Zwolinkski S,
Glen E, Mamasoula C, Rahman TJ, Hall D, Rauch A, Devriendt K,
Gewillig M, O’Sullivan J, Winlaw DS, Bu’Lock F, Brook JD,
Bhattacharya S, Lathrop M, Santibanez-Koref M, Cordell HJ, Goodship
JA and Keavney BD: Contribution of global rare copy-number variants
to the risk of sporadic congenital heart disease. Am J Hum Genet.
91:489–501. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Sanchez-Castro M, Gordon CT, Petit F, Nord
AS, Callier P, Andrieux J, Guérin P, Pichon O, David A, Abadie V,
Bonnet D, Visel A, Pennacchio LA, Amiel J, Lyonnet S and Le Caignec
C: Congenital heart defects in patients with deletions upstream of
SOX9. Hum Mutat. 34:1628–1631. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wu M, Li Y, He X, Shao X, Yang F, Zhao M,
Wu C, Zhang C and Zhou L: Mutational and functional analysis of the
BVES gene coding region in Chinese patients with non-syndromic
tetralogy of Fallot. Int J Mol Med. 31:899–903. 2013.PubMed/NCBI
|
|
21
|
Aoki Y, Niihori T, Banjo T, Okamoto N,
Mizuno S, Kurosawa K, Ogata T, Takada F, Yano M, Ando T, Hoshika T,
Barnett C, Ohashi H, Kawame H, Hasegawa T, Okutani T, Nagashima T,
Hasegawa S, Funayama R, Nagashima T, Nakayama K, Inoue S, Watanabe
Y, Ogura T and Matsubara Y: Gain-of-function mutations in RIT1
cause Noonan syndrome, a RAS/MAPK pathway syndrome. Am J Hum Genet.
93:173–180. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Chang SW, Mislankar M, Misra C, Huang N,
Dajusta DG, Harrison SM, McBride KL, Baker LA and Garg V: Genetic
abnormalities in FOXP1 are associated with congenital heart
defects. Hum Mutat. 34:1226–1230. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Schott JJ, Benson DW, Basson CT, Pease W,
Silberbach GM, Moak JP, Maron BJ, Seidman CE and Seidman JG:
Congenital heart disease caused by mutations in the transcription
factor NKX2-5. Science. 281:108–111. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Wang J, Xin YF, Liu XY, Liu ZM, Wang XZ
and Yang YQ: A novel NKX2-5 mutation in familial ventricular septal
defect. Int J Mol Med. 27:369–375. 2011.PubMed/NCBI
|
|
25
|
Xie WH, Chang C, Xu YJ, Li RG, Qu XK, Fang
WY, Liu X and Yang YQ: Prevalence and spectrum of Nkx2.5 mutations
associated with idiopathic atrial fibrillation. Clinics (Sao
Paulo). 68:777–784. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Huang RT, Xue S, Xu YJ, Zhou M and Yang
YQ: A novel NKX2.5 loss-of-function mutation responsible for
familial atrial fibrillation. Int J Mol Med. 31:1119–1126.
2013.PubMed/NCBI
|
|
27
|
Garg V, Kathiriya IS, Barnes R,
Schluterman MK, King IN, Butler CA, Rothrock CR, Eapen RS,
Hirayama-Yamada K, Joo K, Matsuoka R, Cohen JC and Srivastava D:
GATA4 mutations cause human congenital heart defects and reveal an
interaction with TBX5. Nature. 424:443–447. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Wang J, Fang M, Liu XY, Xin YF, Liu ZM,
Chen XZ, Wang XZ, Fang WY, Liu X and Yang YQ: A novel GATA4
mutation responsible for congenital ventricular septal defects. Int
J Mol Med. 28:557–564. 2011.PubMed/NCBI
|
|
29
|
Liu XY, Wang J, Zheng JH, Bai K, Liu ZM,
Wang XZ, Liu X, Fang WY and Yang YQ: Involvement of a novel GATA4
mutation in atrial septal defects. Int J Mol Med. 28:17–23.
2011.PubMed/NCBI
|
|
30
|
Yang YQ, Gharibeh L, Li RG, Xin YF, Wang
J, Liu ZM, Qiu XB, Xu YJ, Xu L, Qu XK, Liu X, Fang WY, Huang RT,
Xue S and Nemer G: GATA4 loss-of-function mutations underlie
familial tetralogy of fallot. Hum Mutat. 34:1662–1671. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Yang YQ, Wang J, Wang XH, Wang Q, Tan HW,
Zhang M, Shen FF, Jiang JQ, Fang WY and Liu X: Mutational spectrum
of the GATA5 gene associated with familial atrial fibrillation. Int
J Cardiol. 157:305–307. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Jiang JQ, Li RG, Wang J, Liu XY, Xu YJ,
Fang WY, Chen XZ, Zhang W, Wang XZ and Yang YQ: Prevalence and
spectrum of GATA5 mutations associated with congenital heart
disease. Int J Cardiol. 165:570–573. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Wei D, Bao H, Liu XY, Zhou N, Wang Q, Li
RG, Xu YJ and Yang YQ: GATA5 loss-of-function mutations underlie
tetralogy of fallot. Int J Med Sci. 10:34–42. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Wang XH, Huang CX, Wang Q, Li RG, Xu YJ,
Liu X, Fang WY and Yang YQ: A novel GATA5 loss-of-function mutation
underlies lone atrial fibrillation. Int J Mol Med. 31:43–50.
2013.PubMed/NCBI
|
|
35
|
Zheng GF, Wei D, Zhao H, Zhou N, Yang YQ
and Liu XY: A novel GATA6 mutation associated with congenital
ventricular septal defect. Int J Mol Med. 29:1065–1071.
2012.PubMed/NCBI
|
|
36
|
Wang J, Luo XJ, Xin YF, Liu Y, Liu ZM,
Wang Q, Li RG, Fang WY, Wang XZ and Yang YQ: Novel GATA6 mutations
associated with congenital ventricular septal defect or tetralogy
of fallot. DNA Cell Biol. 31:1610–1617. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Yang YQ, Wang XH, Tan HW, Jiang WF, Fang
WY and Liu X: Prevalence and spectrum of GATA6 mutations associated
with familial atrial fibrillation. Int J Cardiol. 155:494–496.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Li J, Liu WD, Yang ZL and Yang YQ: Novel
GATA6 loss-of-function mutation responsible for familial atrial
fibrillation. Int J Mol Med. 30:783–790. 2012.PubMed/NCBI
|
|
39
|
Huang RT, Xue S, Xu YJ and Yang YQ:
Somatic mutations in the GATA6 gene underlie sporadic tetralogy of
Fallot. Int J Mol Med. 31:51–58. 2013.PubMed/NCBI
|
|
40
|
Bruneau BG: The developmental genetics of
congenital heart disease. Nature. 451:943–948. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
McCulley DJ and Black BL: Transcription
factor pathways and congenital heart disease. Curr Top Dev Biol.
100:253–277. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Liu C, Liu W, Palie J, Lu MF, Brown NA and
Martin JF: Pitx2c patterns anterior myocardium and aortic arch
vessels and is required for local cell movement into
atrioventricular cushions. Development. 129:5081–5091.
2002.PubMed/NCBI
|
|
43
|
Dagle JM, Sabel JL, Littig JL, Sutherland
LB, Kolker SJ and Weeks DL: Pitx2c attenuation results in cardiac
defects and abnormalities of intestinal orientation in developing
Xenopus laevis. Dev Biol. 262:268–281. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Bamforth SD, Bragança J, Farthing CR,
Schneider JE, Broadbent C, Michell AC, Clarke K, Neubauer S, Norris
D, Brown NA, Anderson RH and Bhattacharya S: Cited2 controls
left-right patterning and heart development through a Nodal-Pitx2c
pathway. Nat Genet. 36:1189–1196. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Li Q, Pan H, Guan L, Su D and Ma X: CITED2
mutation links congenital heart defects to dysregulation of the
cardiac gene VEGF and PITX2C expression. Biochem Biophys Res
Commun. 423:895–899. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Mommersteeg MT, Brown NA, Prall OW, de
Gier-de Vries C, Harvey RP, Moorman AF and Christoffels VM: Pitx2c
and Nkx2-5 are required for the formation and identity of the
pulmonary myocardium. Circ Res. 101:902–909. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Galli D, Domínguez JN, Zaffran S, Munk A,
Brown NA and Buckingham ME: Atrial myocardium derives from the
posterior region of the second heart field, which acquires
left-right identity as Pitx2c is expressed. Development.
135:1157–1167. 2008. View Article : Google Scholar
|
|
48
|
Lozano-Velasco E, Chinchilla A,
Martínez-Fernández S, Hernández-Torres F, Navarro F, Lyons GE,
Franco D and Aránega AE: Pitx2c modulates cardiac-specific
transcription factors networks in differentiating cardiomyocytes
from murine embryonic stem cells. Cells Tissues Organs.
194:349–362. 2011. View Article : Google Scholar
|
|
49
|
Liu C, Liu W, Lu MF, Brown NA and Martin
JF: Regulation of left-right asymmetry by thresholds of Pitx2c
activity. Development. 128:2039–2048. 2001.PubMed/NCBI
|
|
50
|
Clauss S and Kääb S: Is Pitx2 growing up?
Circ Cardiovasc Genet. 4:105–107. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Yuan F, Zhao L, Wang J, Zhang W, Li X, Qiu
XB, Li RG, Xu YJ, Xu L, Qu XK, Fang WY and Yang YQ: PITX2c
loss-of-function mutations responsible for congenital atrial septal
defects. Int J Med Sci. 10:1422–1429. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Wang J, Xin YF, Xu WJ, Liu ZM, Qiu XB, Qu
XK, Xu L, Li X and Yang YQ: Prevalence and spectrum of PITX2c
mutations associated with congenital heart disease. DNA Cell Biol.
32:708–716. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Zhou YM, Zheng PX, Yang YQ, Ge ZM and Kang
WQ: A novel PITX2c loss-of-function mutation underlies lone atrial
fibrillation. Int J Mol Med. 32:827–834. 2013.PubMed/NCBI
|
|
54
|
Yang YQ, Xu YJ, Li RG, Qu XK, Fang WY and
Liu X: Prevalence and spectrum of PITX2c mutations associated with
familial atrial fibrillation. Int J Cardiol. 168:2873–2876. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Strungaru MH, Footz T, Liu Y, Berry FB,
Belleau P, Semina EV, Raymond V and Walter MA: PITX2 is involved in
stress response in cultured human trabecular meshwork cells through
regulation of SLC13A3. Invest Ophthalmol Vis Sci. 52:7625–7633.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Footz T, Idrees F, Acharya M, Kozlowski K
and Walter MA: Analysis of mutations of the PITX2 transcription
factor found in patients with Axenfeld-Rieger syndrome. Invest
Ophthalmol Vis Sci. 50:2599–2606. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Acharya M, Lingenfelter DJ, Huang L, Gage
PJ and Walter MA: Human PRKC apoptosis WT1 regulator is a novel
PITX2-interacting protein that regulates PITX2 transcriptional
activity in ocular cells. J Biol Chem. 284:34829–34838. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Semina EV, Reiter R, Leysens NJ, Alward
WL, Small KW, Datson NA, Siegel-Bartelt J, Bierke-Nelson D, Bitoun
P, Zabel BU, Carey JC and Murray JC: Cloning and characterization
of a novel bicoid-related homeobox transcription factor gene, RIEG,
involved in Rieger syndrome. Nat Genet. 14:392–399. 1996.
View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Kirchhof P, Kahr PC, Kaese S, Piccini I,
Vokshi I, Scheld HH, Rotering H, Fortmueller L, Laakmann S,
Verheule S, Schotten U, Fabritz L and Brown NA: PITX2c is expressed
in the adult left atrium, and reducing Pitx2c expression promotes
atrial fibrillation inducibility and complex changes in gene
expression. Circ Cardiovasc Genet. 4:123–133. 2011. View Article : Google Scholar
|
|
60
|
Simard A, Di Giorgio L, Amen M, Westwood
A, Amendt BA and Ryan AK: The Pitx2c N-terminal domain is a
critical interaction domain required for asymmetric morphogenesis.
Dev Dyn. 238:2459–2470. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Ganga M, Espinoza HM, Cox CJ, Morton L,
Hjalt TA, Lee Y and Amendt BA: PITX2 isoform-specific regulation of
atrial natriuretic factor expression: synergism and repression with
Nkx2.5. J Biol Chem. 278:22437–22445. 2003. View Article : Google Scholar
|
|
62
|
Furtado MB, Biben C, Shiratori H, Hamada H
and Harvey RP: Characterization of Pitx2c expression in the mouse
heart using a reporter transgene. Dev Dyn. 240:195–203. 2011.
View Article : Google Scholar : PubMed/NCBI
|