Introduction
Fracture healing is a physiological process of
repair that proceeds in stages, each characterized by a different
predominant tissue in the fracture gap. Chondrogenesis as well as
osteogenesis in fracture healing and is downregulated in the later
phase of osteogenesis, as osteoblasts mature to osteocytes. In
addition, neurotrophins and their receptors may be of importance to
osteoblasts and endothelial cells during fracture healing. Cell
viability, osteoblast differentiation, and gene expression are
altered in human osteoblasts from hypertrophic fracture non-unions.
For fracture non-unions that do not heal after appropriate surgical
intervention, the question arises as to what extent systemic
cellular dysfunction is a pathogenic factor.
Mesenchymal stem cells (MSCs) are multipotent
stromal cells that can differentiate into a variety of unique
mesenchymal cell types, such as osteoblasts, chondrocytes and
adipocytes (1). Human MSCs
(hMSCs) have been reported to be a population of self-renewing
multipotent cells that may have clinical therapeutic potential
(2,3). They can differentiate into several
lineages, including osteoblasts, in response to stimulation by
multiple environmental factors (4). Skeletal development and homeostasis
are dependent on the activity of osteoblasts derived from MSCs. In
addition, bone development is delicately regulated by homeostasis,
which is maintained by the balance between osteoblasts and
adipocytes (5). MSCs have emerged
as key regulators of various biological and pathological processes,
and disruption of this differentiation balance leads to various
bone-related metabolic diseases.
MicroRNAs (miRNAs or miRs) are an abundant class of
small (18–25 nucleotides) non-coding single-stranded RNAs found in
diverse organisms. miRNAs have emerged as important
post-transcriptional regulators of gene expression (6–8).
They negatively regulate the translation of specific mRNAs by base
pairing with partially or fully complementary sequences in target
mRNAs (6–9). Although the biological functions of
the majority of miRNAs are not yet fully understood, they may be
essential in the regulation of various biological processes,
including cell proliferation (10,11), apoptosis (12), cell differentiation (13) and tumor formation (14,15). An increasing number of miRNAs have
been found to positively regulate osteoblast differentiation and
bone formation by targeting negative regulators of osteogenesis or
negatively by targeting important osteogenic factors (16–19). The effect of miRNAs has also been
investigated in the osteogenesis of hMSCs. A recent study
demonstrated that miRNAs target the principal transcription factors
and signaling molecules involved in osteoblast differentiation of
MSCs and osteoblast functions (20). miRNAs may thus represent novel
therapeutic targets for pharmacological control of bone cell
functions and enhancing bone formation.
In the present study, it was investigated whether
microRNA-153 (miR-153) was involved in the osteoblastic
differentiation of hMSCs. The expression of miRNAs during
osteogenic differentiation of hMSCs was assessed and miR-153 was
found to be significantly downregulated. Furthermore,
bioinformatics analysis was conducted using PicTar, TargetScan and
microRNA.org to predict the miRNAs that bind to the
bone morphogenetic protein receptor type II (BMPR2) 3′-untranslated
region (3′-UTR). Among the candidates, miR-153 was selected for
further investigation. The regulation of BMPR2 by miR-153 was
verified by the construction of deletion mutants, a luciferase
reporter assay, reverse transcription-quantitative polymerase chain
reaction (RT-qPCR), and western blot analysis. The potential role
of miR-153 in osteoblast differentiation was also investigated
using alkaline phosphatase (ALP) staining and Alizarin Red staining
(ARS).
Materials and methods
hMSC isolation and culture
Nine bone marrow samples from young subjects (<30
years old) and seven bone marrow samples from older subjects
(>60 years old) with slight or severe osteoporosis were
obtained, from which hMSCs were isolated and identified by cell
surface markers, as previously described (21). hMSCs were cultured in α-Minimum
Essential Medium (α-MEM; HyClone, Salt Lake City, UT, USA),
supplemented with 10% fetal bovine serum (FBS; Gibco Invitrogen,
Carlsbad, CA, USA) and 1% penicillin and streptomycin (Gibco
Invitrogen). The cells were incubated at 37°C in a 95% humidity
incubator with 5% CO2. The study was approved by the
Ethics Committee of Henan Hospital of Traditional Chinese Medicine.
Informed consent was signed by the participants.
Osteogenic differentiation
hMSCs were plated at a cell density of
1×105 cells in 12-well plates. At 80% confluence, medium
was replaced with osteoblast-specific induction medium containing
high-glucose Dulbecco's modified Eagle's medium (DMEM), 10% FBS, 5
mM β-glycerophosphate, 100 nM dexamethasone, and 50 µg/ml
ascorbic acid (all purchased from Sigma-Aldrich, St. Louis, MO,
USA) to induce differentiation. hMSCs were cultured in induction
medium for 15 days. The induction medium was changed every 3
days.
RNA extraction RT-qPCR
Total RNA was isolated with TRIzol reagent according
to the manufacturer's instructions (Invitrogen Life Technologies,
Carlsbad, CA, USA). The purity and concentration of total RNA were
determined in an Ultraviolet Spectrophotometer (Eppendorf, Hamburg,
German). cDNA was synthesized from 1 µg total RNA using a
Reverse Transcription kit (ReverTra Dash; Toyobo, Tokyo, Japan)
according to the manufacturer's instructions. qPCR was conducted
using an ABI 7500 Real-Time PCR system (Applied Biosystems,
Carlsbad, CA, USA). The reaction (20 µl) contained the cDNA
synthesized earlier, forward and reverse primers, and SYBR-Green
PCR MasterMix (Applied Biosystems). The amplification conditions
were 50°C for 2 min, 95°C for 10 min, followed by 40 cycles of
denaturation at 95°C for 15 sec, and annealing and extension at
60°C for 1 min. Relative expression of mRNA or microRNA was
evaluated using the 2−ΔΔCt method and normalized to the
expression of β-actin or U6, respectively. The primers for the
genes are listed in Table I.
 | Table IPrimer sequences for reverse
transcription-quantitative polymerase chain reaction. |
Table I
Primer sequences for reverse
transcription-quantitative polymerase chain reaction.
| Name | Forward
(5′-3′) | Reverse
(5′-3′) |
|---|
| ALP |
GACAAGAAGCCCTTCACTGC |
AGACTGCGCCTGGTAGTTGT |
| OC |
TGCTTGTGACGAGCTATCAG |
GAGGACAGGGAGGATCAAGT |
| COL1A1 |
GCAACAGTCGCTTCACCTACA |
CAATGTCCAAGGGAGCCACAT |
| BMPR2 |
CATTCGCTCAAGCAGTTTAGTGGAC |
GGTTCTGAAGCATTTCCTGGGC |
| U6 |
CGCTTCACGAATTTGCGTGTCAT |
GCTTCGGCAGCACATATACTAAAAT |
| β-actin |
AGATGTGGATCAGCAAGCAG |
GCGCAAGTTAGGTTTTGTCA |
Western blot analysis
Cells were lysed in radioimmunoprecipitation assay
buffer with protease inhibitors (Roche Diagnostics, Indianapolis,
IN, USA). Protein concentrations were determined using a Pierce BCA
Protein Assay kit (Thermo Fisher Scientific, Inc., Logan, UT, USA).
Equal quantities of protein were loaded and separated on 10%
SDS-PAGE, and then transferred to a polyvinylidene difluoride
membrane (Millipore, Billerica, MA, USA). Non-specific protein
interactions were blocked by incubation with 3% fat-free milk in
Tris-buffered saline buffer (containing 150 mM NaCl and 50 mM
Tris-HCl, pH 7.6) at 4°C for 1 h. The membranes were incubated with
the following primary antibodies: Anti-BMPR2 antibody (1:1,000;
Cell Signaling Technology, Beverly, MA, USA), anti-ALP antibody
(1:100) or anti-glyceraldehyde 3-phosphate dehydrogenase (GAPDH;
1:2,000) antibody (both from Santa Cruz Biotechnology, Inc., Santa
Cruz, CA, USA) at 4°C overnight. Unbound antibody was removed by
washing in TBS with Tween-20 (TBST) three times (10 min/wash). The
membranes were incubated with horseradish p eroxide-conjugated
secondary antibodies (Zhongshan, Beijing, China) at 25°C for 1 h,
and followed by washing with TBST buffer three times (10 min/wash).
The blots were developed with chemiluminescent ECL reagent
(Millipore). GAPDH was used as an internal control.
ALP staining
The osteoblast phenotype was evaluated by
determining ALP activity. ALP staining was performed using an ALP
staining kit (Institute of Hematology, Chinese Academy of Medical
Sciences, Beijing, China) followed by the accessory procedure.
Cells seeded in 24-well plates were transfected with miR-153
inhibitor or mimics and the respective negative control
(GenePharma, Inc., Shanghai, China) using Lipofectamine™ 2000
(Invitrogen). Prior to staining, the transfected cells were fixed
in 10% paraformaldehyde for 10 min at 25°C. After washing with
phosphate-buffered saline (PBS), t he cells were stained usi ng 300
µg/ml BCIP/NBT solution (Thermo Fisher Scientific, Inc.) for
20 min at 25°C. ALP-positive cells were stained blue/purple.
ARS mineralization assay
ARS was performed to detect calcification during
late induction. Cells seeded in 24-well plates were transfected
with miR-153 inhibitor or mimics and the respective negative
control using Lipofectamine™2000. After cells had been fixed in 5%
paraformaldehyde for 10 min, samples were evaluated by ARS
staining. Briefly, cells were stained with 2% Alizarin Red (pH 7.2)
for 15 min and then washed twice with PBS. Orange and red bodies
were identified as calcium nodules.
Dual luciferase reporter genes
construct
A 94-bp fragment of the BMPR2 3′-UTR containing the
predicted binding site for miR-153 was amplified by PCR from human
genomic DNA. The sequence for the mutation of the miR-153 binding
sites was introduced using the fast mutation kit (NEB, Ipswich, MB,
Canada) according to the manufacturer's instructions. The
constructed plasmids were termed BMPR2-WT (BMPR2-wild-type) and
BMPR2-Mut (BMPR2-mutation), respectively. The fragment was purified
and inserted into a pRL-TK vector (Promega Corporation, Madison,
WI, USA) between XhoI and NotI cleavage sites.
Dual luciferase reporter assay
Luciferase assays were performed according to a
previously published protocol. Cells were co-transfected with the
wild-type BMPR2 3′-UTR (WT) or (Mut) and miR-153 mimic or the
control mimic using Lipofectamine™2000 reagent according to the
manufacturer's instructions. After transfection (48 h), cells were
collected and luciferase activity was assayed for Renilla
and Firefly luciferase activity using the Dual-Luciferase Reporter
assay system (Promega Corportation), and each experiment was
repeated in triplicate.
Statistical analysis
All data are presented as the mean ± standard
deviation (n≥3). Differences between groups were analyzed via
Student's t-test using SPSS 16.0 (SPSS Inc., Chicago, IL, USA).
P<0.05 was considered to indicate a statistically significant
difference.
Results
miR-153 decreases during osteogenic
differentiation
Initially the changes in miR-153 expression during
osteogenic differentiation were examined using RT-qPCR. hMSCs were
used as a cell model and osteogenic differentiation was induced.
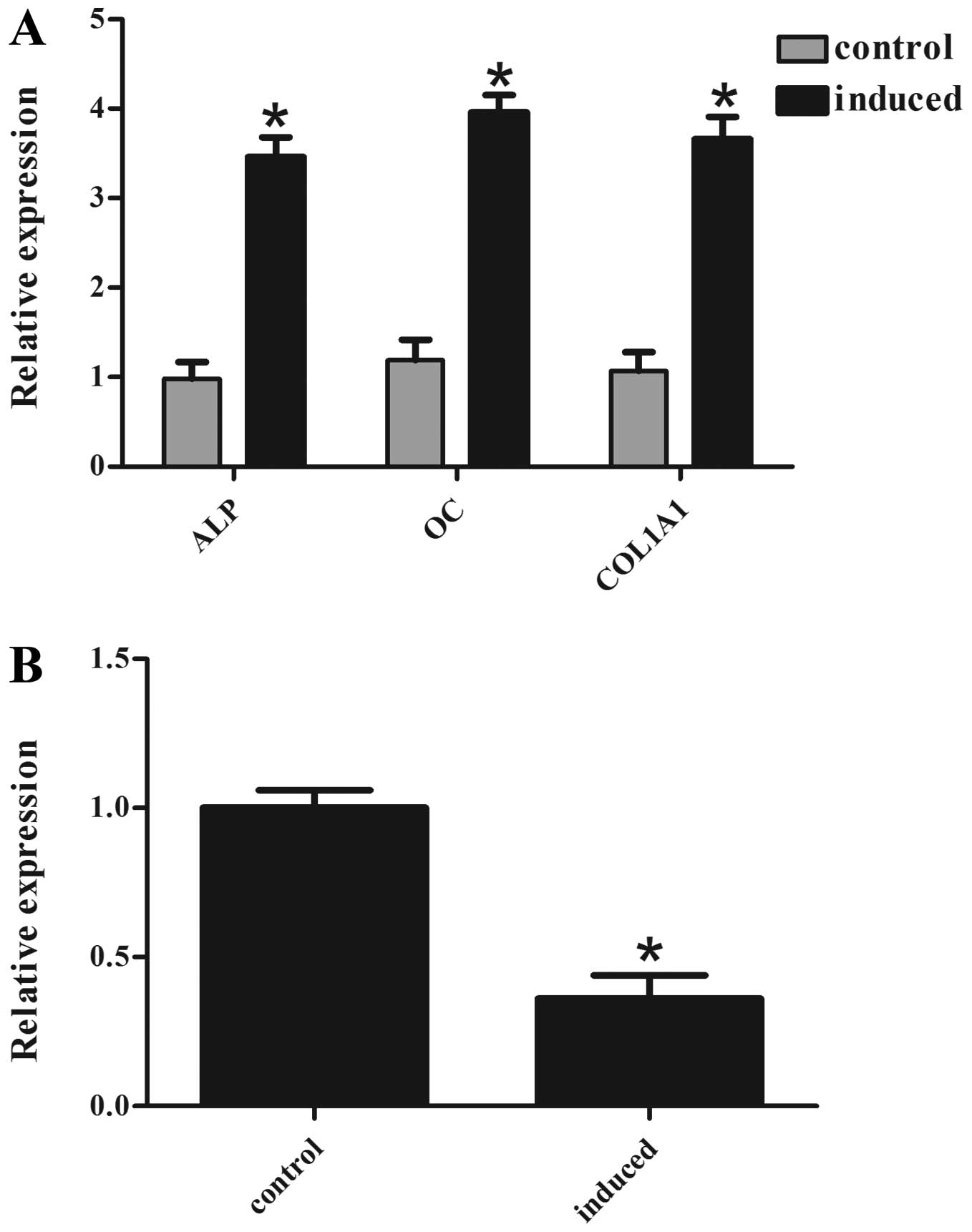
Several osteogenic factors, such as ALP, osteocalcin (OC) and
collagen type I (COL1A1), were used as phenotypic marker genes of
osteogenic differentiation. There was a marked increase in ALP, OC
and COL1A1 mRNA levels in the induced cells, suggesting successful
induction of osteogenic differentiation (Fig. 1A). It was also identified that the
miR-153 levels were significantly lower in the induced cells
compared with the non-induced cells (Fig. 1B). These data suggest that miR-153
may negatively regulate osteogenic differentiation.
miR-153 suppresses osteogenic
differentiation of hMSCs
To further investigate the role of miR-153 in
osteogenic differentiation, hMSCs were transfected with miR-153
inhibitor or mimics and the respective negative control; and then
the capacity for osteogenesis was examined. The effect of the
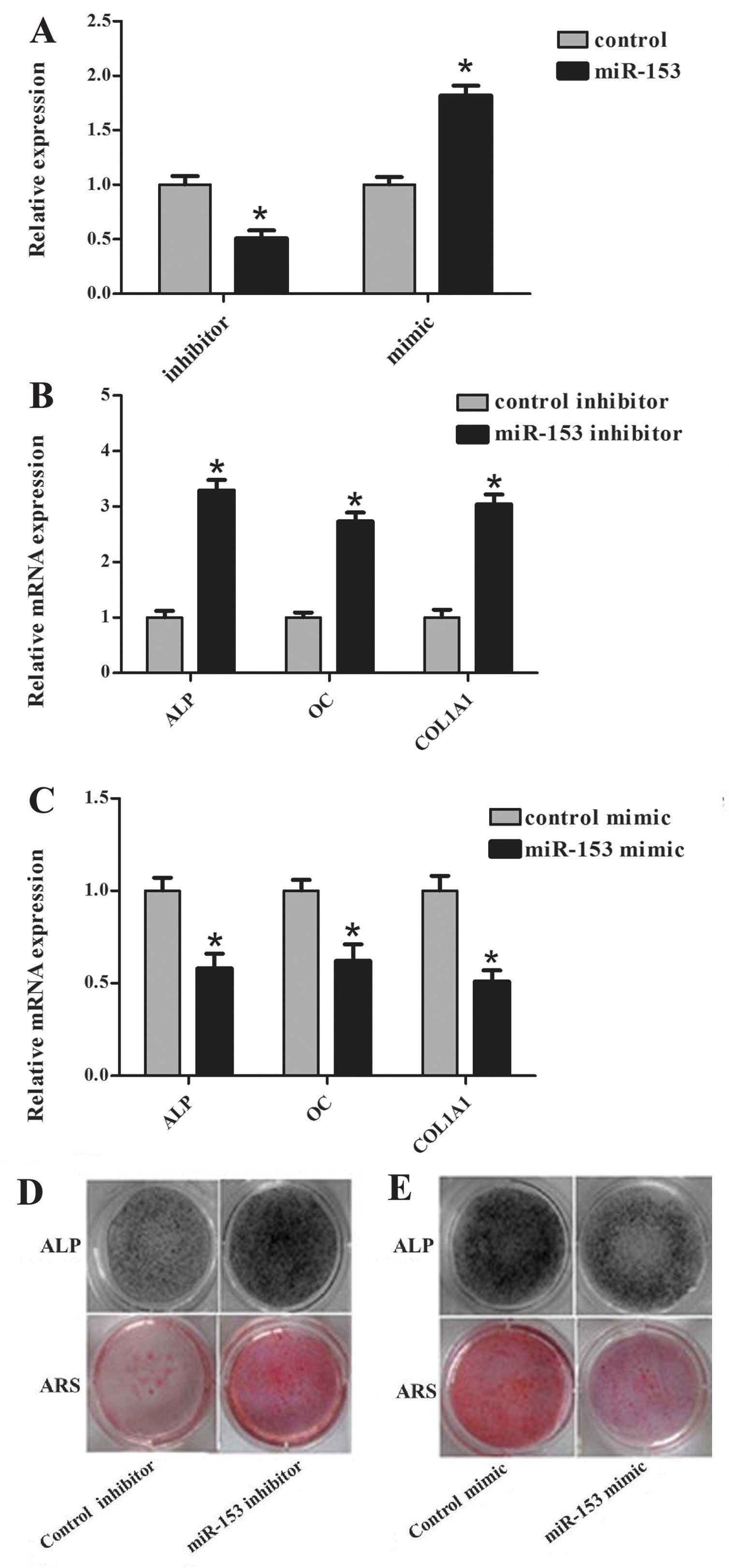
miR-153 mimic/inhibitor was determined by RT-qPCR. The expression
of miR-153 was decreased after transfection of miR-153 inhibitor,
and increased after transfection of miR-153 mimics (Fig. 2A). In addition, all of the
osteogenic differentiation markers examined were significantly
increased following transfection of the miR-153 inhibitors
(Fig. 2B). By contrast, hMSCs
transfected with miR-153 mimics showed decreased expression of ALP,
OC and COL1A1 compared with those transfected with the negative
control (Fig. 2C). At 6 day of
differentiation, ALP staining (Fig.
2D upper) was more intense in the cells transfected with the
miR-153 inhibitor when compared with the cells transfected with
control inhibitor. Furthermore, the staining of the cells
transfected with miR-153 mimic was less intense than those
transfected with the control mimic (Fig. 2E upper). ARS (Fig. 2D and E lower) on day 15 showed the
same tendency at the matrix mineralization level. These results
indicate that miR-153 could suppress osteogenic
differentiation.
BMPR2 is a direct target of miR-153
It is well known that miRNAs act by suppressing the
expression of their target genes. Among the predicted candidates,
BMPR2 was identified, which is a kinase receptor of BMPs with an
established role in osteogenesis (22,23). To understand the molecular
mechanisms underlying the effects of BMPR2, bioinformatics analyses
using miRNA target analysis tools PicTar (http://www.pictar.org/cgi-bin/PicTar_vertebrate.cgi),
TargetScan (http://www.targetscan.org/), and microRNA.org (http://www.microrna.org/microrna/home.do) were
performed to predict the putative miRNAs that bind to the BMPR2
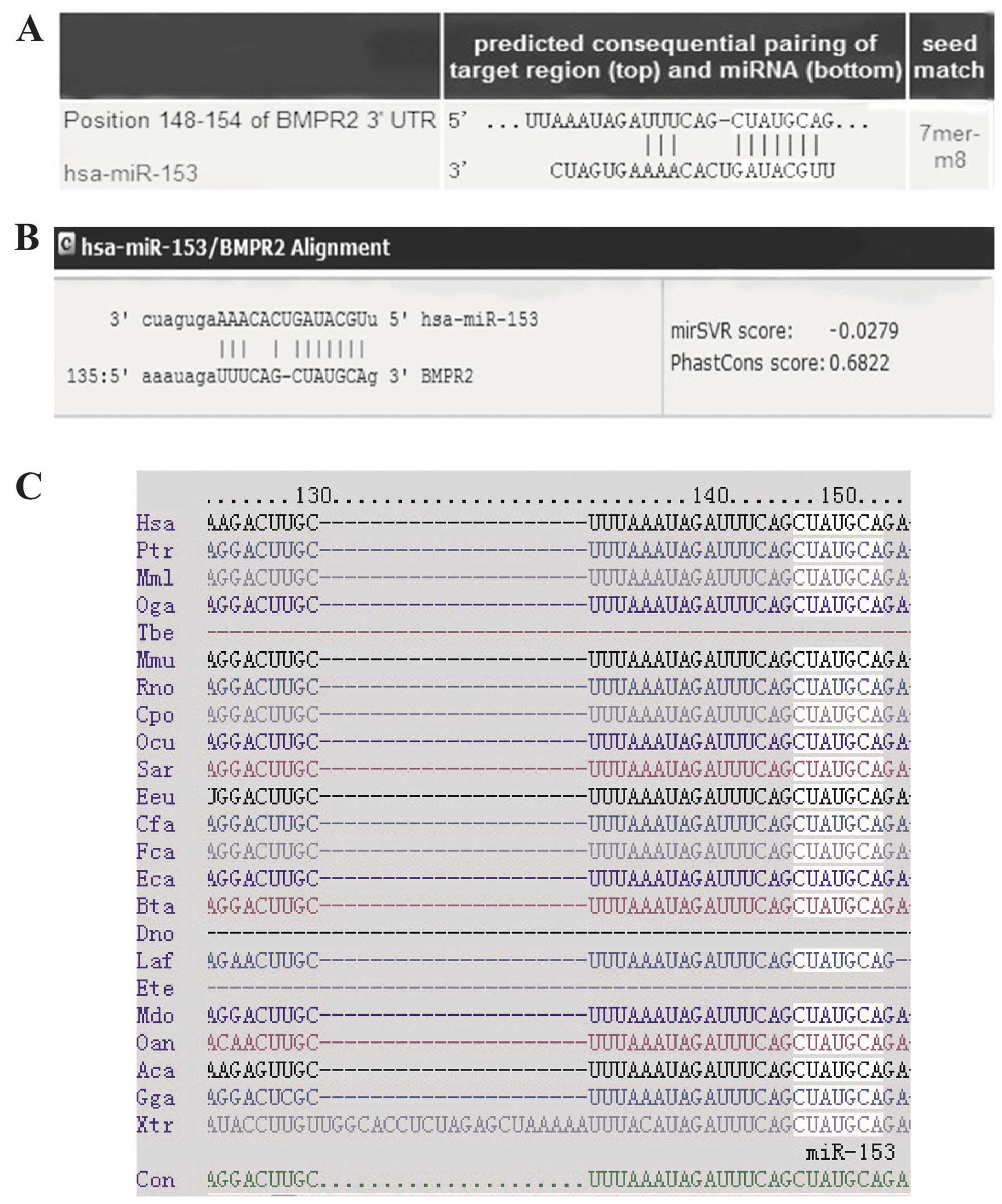
3′-UTR. According to the analysis, the three programs all predicted
that the binding sequence in the 3′-UTR of BMPR2 was a match for
the miR-153 seed (Fig. 3A and B).
The miR-153 target site in the 3′-UTR of BMPR2 is highly conserved
among vertebrates (Fig. 3C).
These data indicate that the translation of BMPR2 protein may be
regulated by miR-153.
miR-153 inhibits BMPR2 expression
To confirm the targeting of BMPR2 by miR-153, a
luciferase reporter assay was performed. The pRL-TK vector contains
Renilla luciferase and firefly luciferase genes. Firefly
luciferase was used as an internal control and Renilla
luciferase was linked with 3′-UTR sequences as a reporter. The
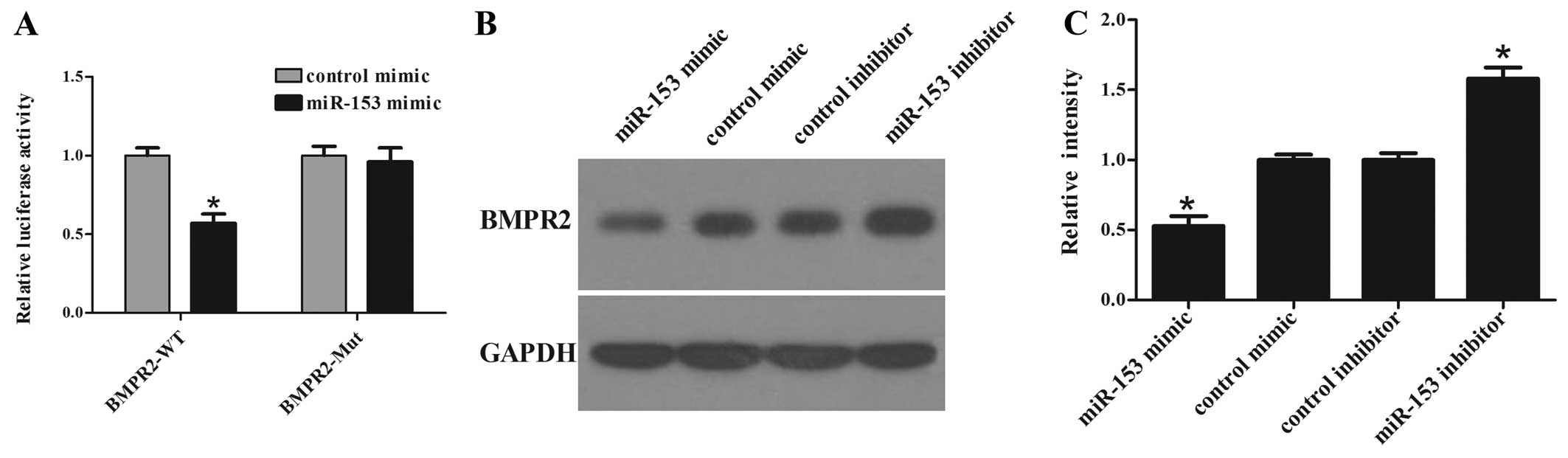
luciferase activity assay (Fig.
4A) showed that the luciferase activity in BMPR2-WT transfected
cells was significantly decreased in miR-153 mimic co-transfected
cells compared with that in control co-transfected cells. In
addition, site-directed mutagenesis of the seed region abolished
the inhibitory effect of miR-153 mimics and Mut luciferase activity
did not change. These results indicate that miR-153 significantly
suppressed the activity of WT but not Mut BMPR2 3′-UTR in hMSCs.
Overexpression of miR-153 significantly suppressed the protein
level of BMPR2, while inhibition of miR-153 elevated the protein
level of BMPR2 in hMSCs (Fig. 4B and
C). These results reveal that miR-153 could negatively regulate
BMPR2 expression through a partially complementary binding site in
the 3′-UTR of BMPR2.
miR-153 suppresses osteogenic
differentiation by targeting BMPR2
Since miR-153 could suppress osteogenic
differentiation and inhibit BMPR2 expression, it was further
explored whether inhibition of BMPR2 by siRNA could also have a
similar effect as miR-153 overexpression on osteogenic
differentiation of cells. The effect of BMPR2 siRNA was confirmed
by RT-qPCR (Fig. 5A) and western
blot analysis (Fig. 5B and C). As
expected, cells transfected with BMPR2 siRNA showed a similar
effect to cells transfected with miR-153 mimic (Figs. 2B and 5D). Furthermore, co-transfection of
miR-153 mimic with BMPR2 overexpression plasmid attenuated the
effect of miR-153 on the osteogenic differentiation of cells
(Fig. 5E). These results indicate
that miR-153 regulates osteogenic differentiation of cells through
BMPR2.
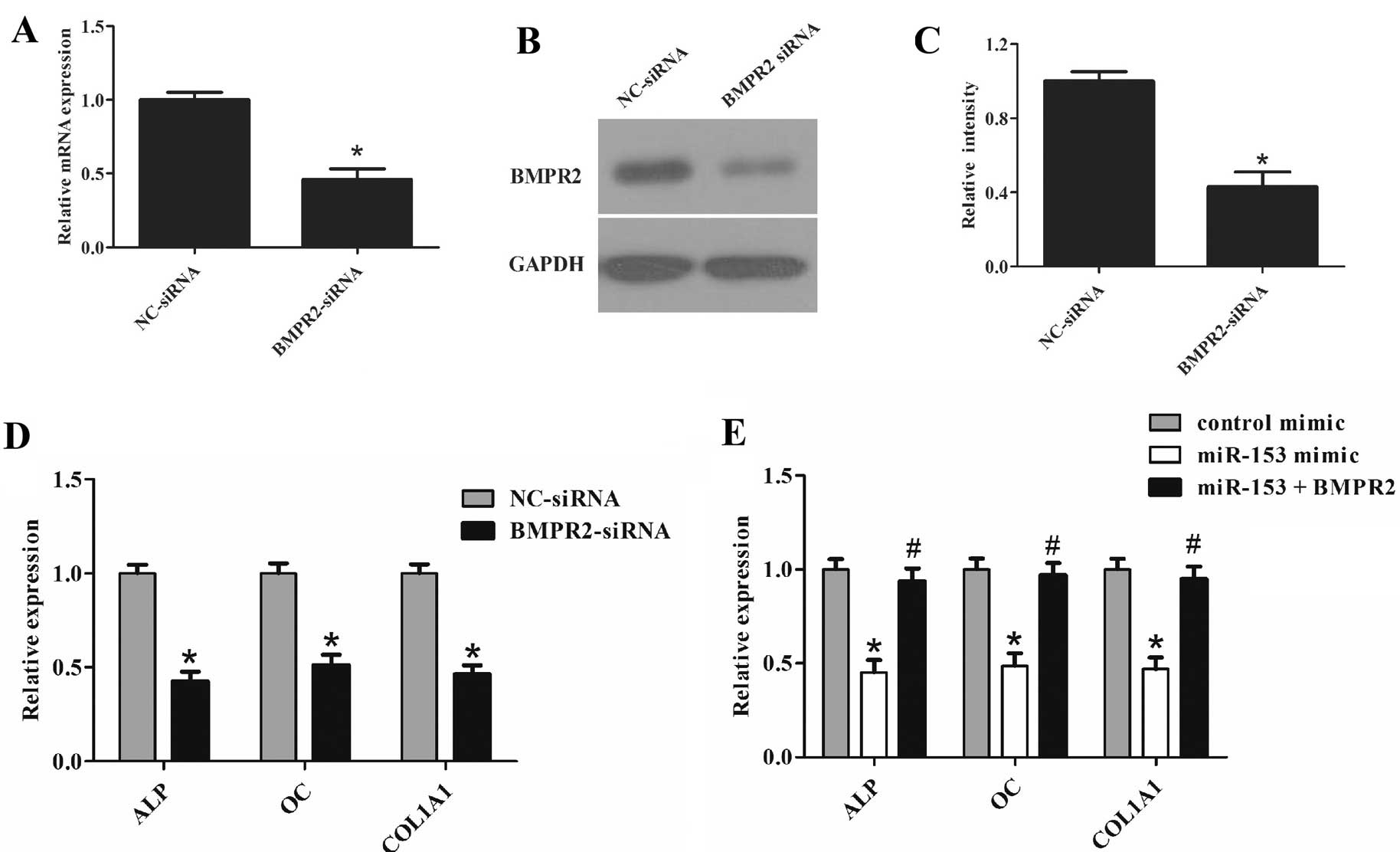 | Figure 5miR-153 suppressed osteogenic
differentiation by targeting BMPR2. (A) Cells were transfected with
BMPR2 siRNA or the control for 72 h. Expression of BMPR2 was
determined by RT-qPCR. (B and C) Cells were transfected with BMPR2
siRNA or the control for 72 h. Western blot analysis was used to
detect the protein level of BMPR2. (D) Cells were transfected with
BMPR2 siRNA or the control for 72 h. Expression of ALP, OC and
COL1A1 was determined by RT-qPCR. (E) Cells were co-transfected
with miR-153 mimic (100 nM)/BMPR2-pcDNA3 for 48 h. Expression of
ALP, OC and COL1A1 was determined by RT-qPCR. Glyceraldehyde
3-phosphate dehydrogenase was used as the internal control. The
data were obtained from three independent experiments.
*P<0.05, compared with the control;
#P<0.05, compared with miR-153 mimic-transfected
cells. miR, microRNA; BMPR2, bone morphogenetic protein receptor
type II; siRNA, small interfering RNA; ALP, alkaline phosphatase;
OC, osteocalcin; COL1A1, collagen type I; RT-qPCR, reverse
transcription-quantitative polymerase chain reaction. |
Discussion
Bone homeostasis involves a balance between bone
formation and bone absorption by osteoblasts and osteoclasts
respectively (24). A growing
body of evidence has revealed that miRNAs are critical in normal
biological processes and the pathogenesis of human diseases by
post-transcriptionally regulating gene expression (25). A number of miRNAs have been
emerging as important negative or positive regulators of
post-transcriptional gene expression and are considered critical
for osteogenesis (26). For
example, miR-34 was reported to inhibit osteoblast differentiation
by targeting Special AT-rich sequence-binding protein 2 (27). miR-100 suppressed osteoblast
differentiation by inhibiting BMPR2 (28). miR-133 and miR-135 functionally
inhibit osteoprogenitor differentiation by abolishing Runx2 and
SMAD family member 5 (Smad5) pathways, which contributed to bone
formation synergistically (29).
Sp7 has recently been identified as a key regulator of osteoblast
differentiation, and was reported to be regulated by miR-93,
miR-125b, miR-214 and miR-637 (30–33). miRNAs are endogenous small
noncoding RNAs that regulate the activities of target mRNAs and
cellular processes. Although no miRNA has been reported to be
important in the regulation of fracture healing, several miRNAs
control key elements in tissue repair processes, such as
inflammation, hypoxia response, angiogenesis, stem cell
differentiation, osteogenesis, and chondrogenesis. In this study,
miR-153 was identified as a suppressive regulator of osteogenic
differentiation. It was demonstrated that miR-153 was downregulated
during the osteogenic differentiation of cells. Overexpression of
miR-153 inhibited osteogenic differentiation, whereas inhibition of
miR-153 enhanced the osteogenic potential.
It has been reported that miR-153 is important in a
number of types of cancer (34,37). However, its effects do not appear
to be consistent. miR-153 suppresses tumor growth in glioblastoma
(34), epithelial cancer
(35) and leukemia (36). Conversely, in prostate cancer,
miR-153 promotes cell proliferation via downregulation of the PTEN
tumor suppressor gene (37).
However, the role of miR-153 in the regulation of osteoblastic
differentiation remains poorly understood. In the present study, a
novel role of miR-153 in the osteogenesis of cells was identified.
To investigate the molecular mechanism by which miR-153 regulates
the osteogenic differentiation of cells, potential target genes
were analyzed that have an established function in promoting
osteogenesis using the target prediction tools, PicTar, TargetScan
and microRNA.org. The results revealed that BMPR2 may
be a possible target with a 9-nt non-consecutive match site
complementary to miR-153 in its 3′-UTR. It was demonstrated that
overexpression of miR-153 resulted in the suppression of BMPR2
expression, whereas functional inhibition of miR-153 led to
elevation of BMPR2, strongly suggesting that BMPR2 is regulated by
miR-153. In addition, the dual luciferase reporter assay also
identified BMPR2 as a direct target of miR-153. Supplementing BMPR2
could also partially reverse the suppressive effect of miR-153 on
osteoblastic differentiation. These data suggest that miR-153 may
negatively regulate osteogenic differentiation partially by
targeting BMPR2.
BMP signaling has been reported to be important in
the commitment of human adipose-derived mesenchymal stem cell
(hASC) osteogenic differentiation (38) and BMPs have important clinical
significance serving as strong osteoinductive factors for bone
tissue repair and regeneration (39). BMP signaling is initiated by the
binding of extracellular BMPR, resulting in BMPR2-mediated
activation of the BMPR1, which, in turn, causes the phosphorylation
and activation of intracellular Smad signaling molecules, and then
activates osteoblast-essential genes (23). These studies indicated that BMPR2
may be a crucial factor in the pathway of osteogenic
differentiation. BMP2 is key in skeletal development, repair and
regeneration. In the present study, it was demonstrated that
miR-153 negatively targets BMPR2 and suppresses osteogenic
differentiation.
In conclusion, the data indicate that miR-153 is a
novel regulator of BMPR2 and that it serves as a suppressor of
osteogenic differentiation. Thus, BMPR2 and miR-153 may be
potential therapeutic targets in the management of skeletal
diseases.
References
|
1
|
Bruder SP, Fink DJ and Caplan AI:
Mesenchymal stem cells in bone development, bone repair, and
skeletal regeneration therapy. J Cell Biochem. 56:283–294. 1994.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Locke M, Feisst V and Dunbar PR: Concise
review: human adipose-derived stem cells: separating promise from
clinical need. Stem Cells. 29:404–411. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Wang S, Qu X and Zhao RC: Mesenchymal stem
cells hold promise for regenerative medicine. Front Med. 5:372–378.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Gimble JM and Guilak F: Differentiation
potential of adipose derived adult stem (ADAS) cells. Curr Top Dev
Biol. 58:137–160. 2003. View Article : Google Scholar
|
|
5
|
Mathieu PS and Loboa EG: Cytoskeletal and
focal adhesion influences on mesenchymal stem cell shape,
mechanical properties, and differentiation down osteogenic,
adipogenic, and chondrogenic pathways. Tissue Eng Part B Rev.
18:436–444. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Eskildsen T, Taipaleenmäki H, Stenvang J,
Abdallah BM, Ditzel N, Nossent AY, Bak M, Kauppinen S and Kassem M:
MicroRNA-138 regulates osteogenic differentiation of human stromal
(mesenchymal) stem cells in vivo. Proc Natl Acad Sci USA.
108:6139–6144. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Dong J, Cui X, Jiang Z and Sun J:
MicroRNA-23a modulates tumor necrosis factor-alpha-induced
osteoblasts apoptosis by directly targeting Fas. J Cell Biochem.
114:2738–2745. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Hu R, Li H, Liu W, Yang L, Tan YF and Luo
XH: Targeting miRNAs in osteoblast differentiation and bone
formation. Expert Opin Ther Targets. 14:1109–1120. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Inose H, Ochi H, Kimura A, et al: A
microRNA regulatory mechanism of osteoblast differentiation. Proc
Natl Acad Sci USA. 106:20794–20799. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Laneve P, Di Marcotullio L, Gioia U, Fiori
ME, Ferretti E, Gulino A, Bozzoni I and Caffarelli E: The interplay
between microRNAs and the neurotrophin receptor tropomyosin-related
kinase C controls proliferation of human neuroblastoma cells. Proc
Natl Acad Sci USA. 104:7957–7962. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Wang X, Tang S, Le SY, Lu R, Rader JS,
Meyers C and Zheng ZM: Aberrant expression of oncogenic and
tumor-suppressive microRNAs in cervical cancer is required for
cancer cell growth. PLoS One. 3:e25572008. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Song G, Zhang Y and Wang L: MicroRNA-206
targets notch3, activates apoptosis, and inhibits tumor cell
migration and focus formation. J Biol Chem. 284:31921–31927. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Kawasaki H and Taira K: Hesl is a target
of microRNA-23 during retinoic-acid-induced neuronal
differentiation of NT2 cells. Nature. 423:838–842. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Sun T, Wang Q, Balk S, Brown M, Lee GS and
Kantoff P: The role of microRNA-221 and microRNA-222 in
androgen-independent prostate cancer cell lines. Cancer Res.
69:3356–3363. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Kumar MS, Lu J, Mercer KL, Golub TR and
Jacks T: Impaired microRNA processing enhances cellular
transformation and tumorigenesis. Nat Genet. 39:673–677. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Dong S, Yang B, Guo H and Kang F:
MicroRNAs regulate osteogenesis and chondrogenesis. Biochem Biophys
Res Commun. 418:587–591. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Li H, Xie H, Liu W, et al: A novel
microRNA targeting HDAC5 regulates osteoblast differentiation in
mice and contributes to primary osteoporosis in humans. J Clin
Invest. 119:3666–3677. 2009. View
Article : Google Scholar : PubMed/NCBI
|
|
18
|
Guo J, Ren F, Wang Y, et al: miR-764-5p
promotes osteoblast differentiation through inhibition of
CHIP/STUB1 expression. J Bone Miner Res. 27:1607–1618. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Gao J, Yang T, Han J, Yan K, Qiu X, Zhou
Y, Fan Q and Ma B: MicroRNA expression during osteogenic
differentiation of human multipotent mesenchymal stromal cells from
bone marrow. J Cell Biochem. 112:1844–1856. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Taipaleenmäki H, Bjerre Hokland L, Chen L,
Kauppinen S and Kassem M: Mechanisms in endocrinology: micro-RNAs:
targets for enhancing osteoblast differentiation and bone
formation. Eur J Endocrinol. 166:359–371. 2012. View Article : Google Scholar
|
|
21
|
Zhang JF, Li G, Meng CL, Dong Q, Chan CY,
He ML, Leung PC, Zhang YO and Kung HF: Total favonoids of Herba
Epimedii improves osteogenesis and inhibits osteoclastogenesis of
human mesenchymal stem cells. Phytomedicine. 16:521–529. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Balint E, Lapointe D, Drissi H, van der
Meijden C, Young DW, van Wijnen AJ, Stein JL, Stein GS and Lian JB:
Phenotype discovery by gene expression profiling: Mapping of
biological processes linked to BMP-2-mediated osteoblast
differentiation. J Cell Biochem. 89:401–426. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Wang A, Ding X, Sheng S and Yao Z: Bone
morphogenetic protein receptor in the osteogenic differentiation of
rat bone marrow stromal cells. Yonsei Med J. 51:740–745. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Hu W, Ye Y, Zhang W, Wang J, Chen A and
Guo F: miR-142-3p promotes osteoblast differentiation by modulating
Wnt signaling. Mol Med Rep. 7:689–693. 2013.
|
|
25
|
Lamouille S, Subramanyam D, Blelloch R and
Derynck R: Regulation of epithelial-mesenchymal and
mesenchymal-epithelial transitions by microRNAs. Curr Opin Cell
Biol. 25:200–207. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Lian JB, Stein GS, van Wijnen AJ, Stein
JL, Hassan MQ, Gaur T and Zhang Y: MicroRNA control of bone
formation and homeostasis. Nat Rev Endocrinol. 8:212–227. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Wei J, Shi Y, Zheng L, Zhou B, Inose H,
Wang J, Guo XE, Grosschedl R and Karsenty G: miR-34s inhibit
osteoblast proliferation and differentiation in the mouse by
targeting SATB2. J Cell Biol. 197:509–521. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Zeng Y, Qu X, Li H, Huang S, Wang S, Xu Q,
Lin R, Han Q, Li J and Zhao RC: MicroRNA-100 regulates osteogenic
differentiation of human adipose-derived mesenchymal stem cells by
targeting BMPR2. FEBS Lett. 586:2375–2381. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Li Z, Hassan MQ, Volinia S, van Wijnen AJ,
Stein JL, Croce CM, Lian JB and Stein GS: A microRNA signature for
a BMP2-induced osteoblast lineage commitment program. Proc Natl
Acad Sci USA. 105:13906–13911. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Yang L, Cheng P, Chen C, He HB, Xie GQ,
Zhou HD, Xie H, Wu XP and Luo XH: miR-93/Sp7 function loop mediates
osteoblast mineralization. J Bone Miner Res. 27:1598–1606. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Goettsch C, Rauner M, Pacyna N, Hempel U,
Bornstein SR and Hofbauer LC: miR-125b regulates calcification of
vascular smooth muscle cells. Am J Pathol. 179:1594–1600. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Shi K, Lu J, Zhao Y, Wang L, Li J, Qi B,
Li H and Ma C: MicroRNA-214 suppresses osteogenic differentiation
of C2C12 myoblast cells by targeting Osterix. Bone. 55:487–494.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Zhang JF, Fu WM, He ML, et al: MiR-637
maintains the balance between adipocytes and osteoblasts by
directly targeting Osterix. Mol Biol Cell. 22:3955–3961. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Zhao S, Deng Y, Liu Y, Chen X, Yang G, Mu
Y, Zhang D, Kang J and Wu Z: MicroRNA-153 is tumor suppressive in
glioblastoma stem cells. Mol Biol Rep. 40:2789–2798. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Xu Q, Sun Q, Zhang J, Yu J, Chen W and
Zhang Z: Downregulation of miR-153 contributes to
epithelial-mesenchymal transition and tumor metastasis in human
epithelial cancer. Carcinogenesis. 34:539–549. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Liu L, Chen R, Huang S, Wu Y, Li G, Zhang
B, Liu Q, Yin D and Liang Y: miR-153 sensitized the K562 cells to
As2O3-induced apoptosis. Med Oncol. 29:243–247. 2012. View Article : Google Scholar
|
|
37
|
Wu Z, He B, He J and Mao X: Upregulation
of miR-153 promotes cell proliferation via downregulation of the
PTEN tumor suppressor gene in human prostate cancer. Prostate.
73:596–604. 2013. View Article : Google Scholar
|
|
38
|
Kang Q, Song WX, Luo Q, et al: A
comprehensive analysis of the dual roles of BMPs in regulating
adipogenic and osteogenic differentiation of mesenchymal progenitor
cells. Stem Cells Dev. 18:545–559. 2009. View Article : Google Scholar
|
|
39
|
Wang Q, Huang C, Xue M and Zhang X:
Expression of endogenous BMP-2 in periosteal progenitor cells is
essential for bone healing. Bone. 48:524–532. 2011. View Article : Google Scholar :
|