Introduction
Extracellular vesicles (EVs) are small particles
(30–1000 nm) secreted by various types of cells; they are enclosed
by a phospholipid bilayer and contain DNA, RNA and protein
(1). EVs can be released into
cell culture medium (CCM), and they are also found abundantly and
naturally in body fluids. They may therefore serve as biomarkers
for the development of superior, sensitive and minimally invasive
diagnostic alternatives in 'precision medicine' (2,3).
Due to the small size and heterogeneity of vesicles, EV detection
and classification is challenging. Different types of vesicles have
been identified, such as exosomes, microvesicles, apoptotic bodies,
secreted proteins and retrovirus-like vesicles (1,4).
In the present study, the widely used term 'exosomes' was used to
refer to EVs (exosomes and other types of cell-derived vesicles) in
general.
The commonly used protocol for isolation of exosomes
is ultracentrifugation (UC); the final step of which is
centrifugation at 100,000 × g at least for 70 min to pellet the
small vesicles that correspond to exosomes (5). In addition, sucrose density
gradients, ultrafiltration (6),
high performance liquid chromatography-based protocols (7) and immunoaffinity-capture methods
(8), singly or combined with the
application of UC, can provide high enrichment and purity of
exosomes (9). In recent years,
easy-to-use precipitation solutions, such as ExoQuick and Total
Exosomes Isolation Reagent (TEI), have been utilized to precipitate
particles in liquid. The procedure is convenient and time-saving
with no need for expensive equipment or technical challenge
(10,11). However, the 'salting out' methods,
unable to resolve particle heterogeneity, are not specific for
exosomes or other EVs and may easily lead to the isolation of
non-exosomal particles (12).
In the downstream analysis of exosomal content, a
number of alternative exosomal RNA (exoRNA) extraction methods have
been used, including phenol-based techniques (TRIzol) and combined
phenol and pure column-based techniques [miRNeasy and HiPure Liquid
RNA/miRNA kit (HLR)] (13).
Recently, commercial kits [SeraMir™ Exosomes RNA Amplification kit
(SeraMir), Total Exosomes RNA and Protein Isolation kit (TER)] have
been designed specifically for the isolation of RNA and protein
from a single enriched exosome preparation. The exoRNeasy
Serum/Plasma kits use a membrane-based affinity binding step to
isolate exoRNA directly from serum or plasma (14).
Given the highly attractive research value of
exoRNA, the availability of convenient methods to extract the
exoRNA with high quality, substantial yield, purity and appropriate
size distribution needs to be confirmed. Recently, comparative
studies on the impact of isolation methods for either exosomes
(15,16) or exoRNA (13,17) on downstream RNA yield, quality and
profiles have been conducted. However, in practice, the methods
used for extraction of both exosomes and exoRNA should be
considered in order to extract exoRNA of high-quality. In addition,
a large range of methods including several new commercial kits
designed specifically for exoRNA extraction (SeraMir, TEI and
exoRNeasy) should be utilized and compared. Therefore, in this
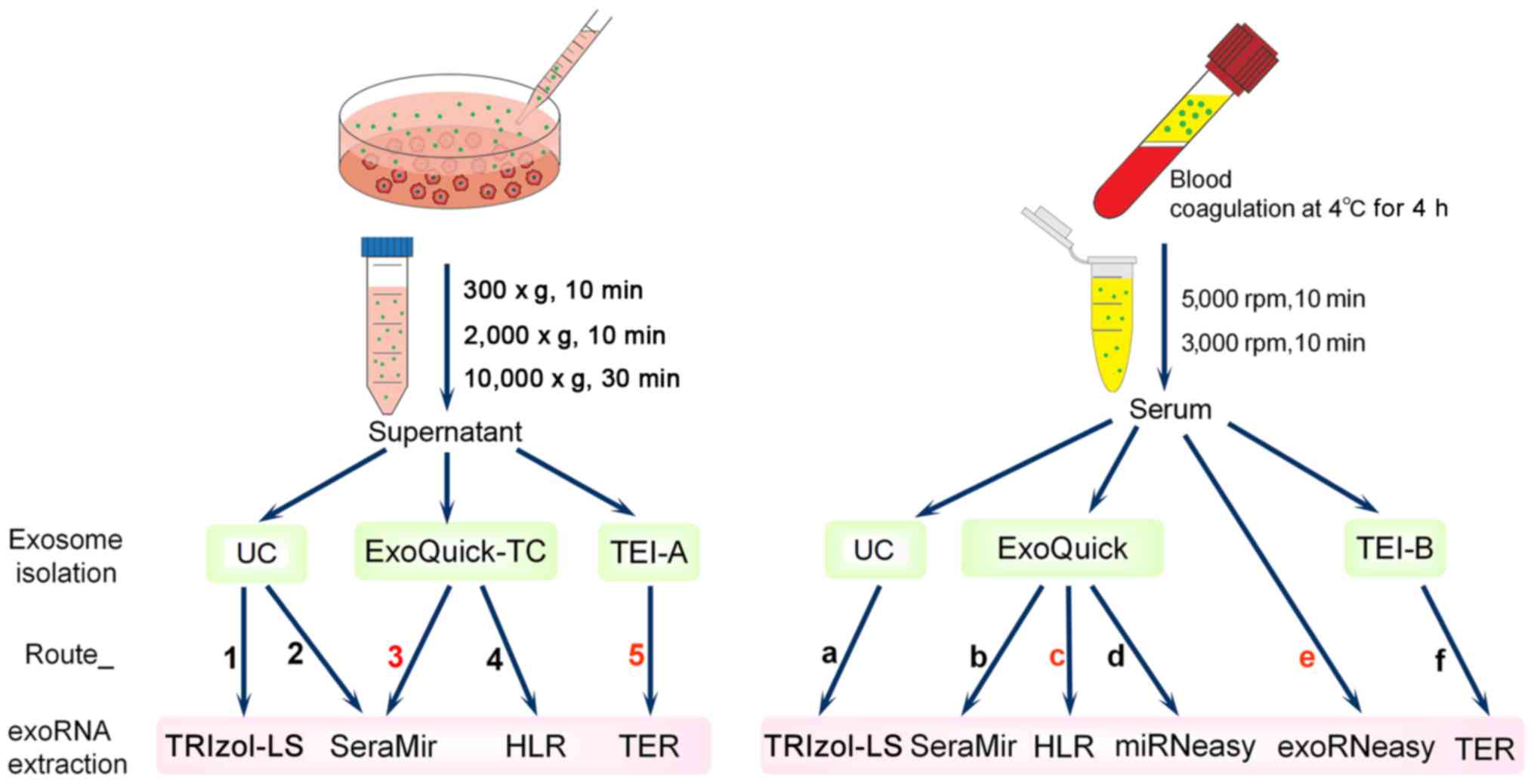
study (Fig. 1), nanoparticle
tracking analysis (NTA) and protein analysis were used to measure
and characterize CCM or serum-derived exosomes isolated using UC
and two commercially available kits (ExoQuick and TEI).
Subsequently, exoRNA was isolated by a combination of the above
methods for exosome isolation and six exoRNA isolation methods
(TRIzol-LS, SeraMir, TEI, HLR, miRNeasy and exoRNeasy). Given that
there were too many methods for each combination to be tested, we
determined the combination patterns (Route_1 to Route_5 for CCM and
Route_a to Route_f for serum) (Fig.
1) according to the recommendations of the relevant
manufacturers or other published studies (11,17–20). The quantity and quality of exoRNA
were determined using NanoDrop and Bioanalyzer 2100. By the use of
qPCR and high-throughput sequencing analysis, we provide evidence
that different combinations of isolation methods for exosomes and
exoRNA can affect exoRNA profiling.
Materials and methods
Study design and participant consent
This study was approved by the Ethics Committee of
Nanfang Hospital, Guangzhou, China, and written informed consent
was obtained from all participants. The sample collection and
treatment were carried out in accordance with the approved
guidelines. The experiment was repeated three times using three
completely independent sets of samples (three independent CCM or
serum samples prepared at different times). Each sample was divided
into 100 ml CCM and 500 µl serum for each extraction method.
A flowchart of the study design is shown in Fig. 1. The human lung cancer cell line
A549 (ATCC, Manassas, VA, USA) was cultured in serum-free RPMI-1640
medium and 2% Exo-FBS™ exosome-depleted fetal bovine serum (System
Biosciences, Mountain View, CA, USA) for 48 h and the CCM was
collected and centrifuged at 300 × g for 10 min, then at 2,000 × g
for 10 min, and finally at 10,000 × g for 30 min to remove dead
cells, cell debris and large particles (shedding vesicles and
apoptotic bodies). Blood samples were obtained from three healthy
donors (mean age, 28 years; gender, one female and two males) at
the Department of Laboratory Medicine, Nanfang Hospital, Southern
Medical University, Guangzhou, China. Blood in containers without
anticoagulant or coagulant was kept at 4°C for 4 h to ensure serum
separation. Serum samples were centrifuged at 5,000 rpm for 10 min,
and then at 3,000 rpm for 10 min and stored at −80°C before
use.
Methods for exosome isolation from CCM or serum
included: UC, ExoQuick (System Biosciences) and TEI (Invitrogen,
Life Technology, Carlsbad, CA, USA). RNA was extracted from
exosomes using four different methods: TRIzol-LS (Ambion, Life
Technology, Carlsbad, CA, USA), SeraMir (System Biosciences), TER
(Invitrogen), HLR (Magen, Guangzhou, China) and miRNeasy (Qiagen,
Hilden, Germany). ExoRNeasy (Qiagen) is able to purify exosomal RNA
from serum directly.
Exosome isolation
The UC method was used as previously described
(5). The supernatant was
ultracentrifuged using a W32Ti rotor (L-80XP; Beckman Coulter,
Brea, CA, USA) at 110,000 × g for 70 min to pellet the exosomes.
The pellet was washed in phosphate-buffered saline (PBS) to
eliminate contaminating proteins, and centrifuged again at 110,000
× g for 70 min. The PBS was removed and the exosomes re-suspended
in 100 µl PBS or nuclease-free water. The nanomaterial
exosome isolation methods tested comprised four kits (ExoQuick-TC,
TEI for CCM, ExoQuick, TEI for serum), which were used according to
the manufacturer's instructions. All centrifugation steps were
performed at 4°C.
Nanoparticle tracking analysis (NTA)
Vesicle suspensions with concentrations between
1×107/ml and 1×109/ml were examined using a
Nanosight NS300 (NanoSight Ltd., Amesbury, UK) equipped with a 405
nm laser to determine the size and quantity of particles isolated.
A video of 60-sec duration was taken with a frame rate of 30
frames/sec, and particle movement was analyzed using NTA software
(version 2.3; NanoSight Ltd.).
Transmission electron microscopy
(TEM)
A 20–40 µl solution of exosomes was placed on
a copper mesh and post-negatively stained with 2% phosphotungstic
acid solution for 10 min. The sample was then dried for 2 min under
incandescent light. The copper mesh was observed and photographed
under a transmission electron microscope (H-7650 Hitachi
microscope; Hitachi, Tokyo, Japan).
Western blot analysis
The exosome supernatant was denatured in 5X sodium
dodecyl sulfonate (SDS) buffer and subjected to western blot
analysis (10% SDS-polyacrylamide gel electrophoresis; 50 µg
protein/lane) using rabbit polyclonal antibody CD63 (sc-15363) in
CD9 (sc-13118; both from Santa Cruz Biotechnology, Inc., Dallas,
TX, USA), TSG101 (T5701; Sigma-Aldrich, Dorset, UK) and calnexin
(BS1438; Bioworld Technology, St. Louis Park, MN, USA). The
proteins were visualized on the Bio-Rad ChemiDoc XRS Imager system
(Bio-Rad Laboratories, Berkeley, CA, USA).
ExoRNA isolation and RNA analyses
TRIzol-LS reagent was designed to isolate
high-quality total RNA from liquid samples, and 250 µl of
exosome solution was lysed in 750 µl TRIzol-LS.
Subsequently, 200 µl of chloroform was used for phase
separation and 100% isopropanol for RNA precipitation. Finally, RNA
was eluted in 30 µl RNase-free water after being washed
twice in 75% ethanol. TER, HLR, miRNeasy, SeraMir and exoRNeasy
kits were used according to each manufacturer's total RNA isolation
procedure.
The RNA concentration was assessed using a NanoDrop
2000 spectrophotometer (Thermo Scientific, Waltham, MA, USA). The
RNA yield and size distribution were analyzed using an Agilent 2100
Bioanalyzer with an RNA 6000 Pico kit (Agilent Technologies, Foster
City, CA, USA).
RNA profiling analysis
Aliquots containing 100 ng of three exoRNA samples
from CCM isolated by different methods were used for RNA library
preparation, following the instructions for the NEBNext Multiplex
Small RNA library preparation kit (New England Biolabs, Ipswich,
MA, USA). The PCR amplified cDNA construct (from 140–160 bp) was
purified using a QIAquick PCR Purification kit (Qiagen). The
purified cDNA was directly sequenced using an Illumina MiSeq 2000
platform (Illumina, San Diego, CA, USA). The miRNA contained in
serum exosomes was analyzed using an All-in-One™ miRNA First-Strand
cDNA synthesis kit and miRNA qPCR kit (GeneCopoeia, Rockville, MD,
USA).
Statistical analysis
One-way or two-way analysis of variance (ANOVA) was
used to investigate the differences between groups. Differences in
paired samples were compared using the two-tailed Student's t-test.
Correlations between variables were assessed by Pearson's
correlation. Linear regression was applied to determine the linear
regression equation. SPSS 15.0 was used for statistical analyses
(SPSS, Inc., Chicago, IL, USA). P<0.05 was considered to be
statistically significant.
Results
Among the three exosome isolation methods
(UC, ExoQuick and TEI), UC shows the lowest yield and recovery, but
the highest protein purity
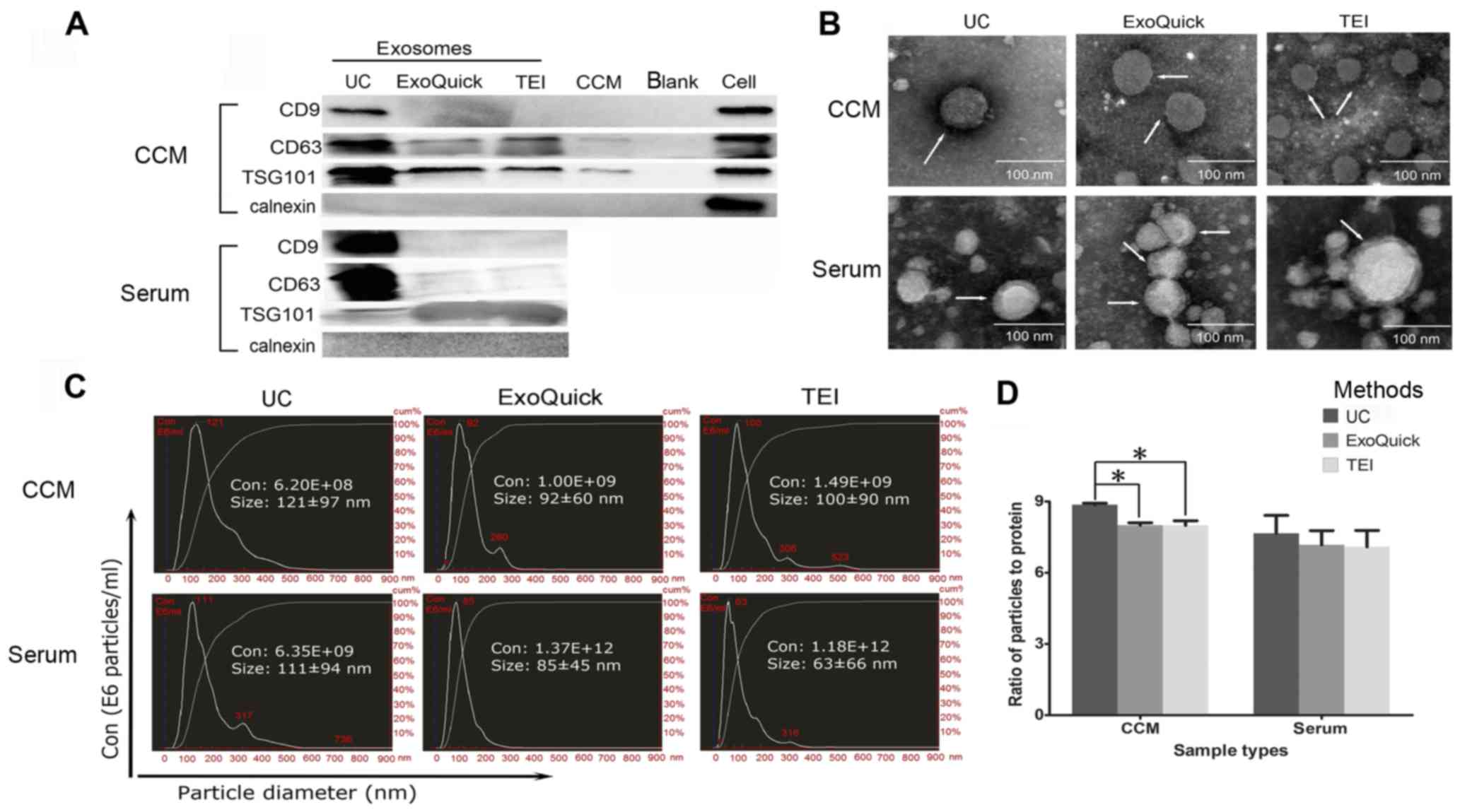
To identify exosomes, the vesicles isolated from
A549 CCM and serum were investigated by western blot analysis
(Fig. 2A), TEM (Fig. 2B) and NTA (Fig. 2C). The existence of exosomal
protein markers (CD9, CD63 or TSG101) and non-existence of
non-exosomal markers (calnexin), the lipid bilayer structure, and
the size of particles (50–200 nm) were used to demonstrate the
presence of exosomes.
The recovery rate of exosomes generated by the three
methods was compared using NTA (Table
I). Either in CCM or serum, the two commercial kits (ExoQuick
and TEI) showed higher exosomal recovery than UC. In CCM, TEI
produced a higher yield of exosomes than ExoQuick. However, in
serum, no difference was found between the two commercial kits.
 | Table ITotal number and diameter of particles
isolated from CCM and serum by the different methods. |
Table I
Total number and diameter of particles
isolated from CCM and serum by the different methods.
| Samples | N | Concentration
(particles/ml)
| Size (mode ± SD)
|
|---|
| Ultra | ExoQuick | TEI | Ultra | ExoQuick | TEI |
|---|
| CCM | 1 | 6.20E+08 | 1.00E+09 | 1.49E+09 | 121±97 | 92±60 | 100±90 |
| 2 | 6.33E+07 | 4.65E+08 | 5.52E+08 | 165±131 | 132±87 | 173±96 |
| 3 | 9.17E+06 | 5.51E+07 | 9.33E+07 | 136±74 | 104±66 | 92±201 |
| Serum | 1 | 6.35E+09 | 1.37E+12 | 1.18E+12 | 111±94 | 85±45 | 63±66 |
| 2 | 2.22E+09 | 1.48E+11 | 4.72E+11 | 161±109 | 133±75 | 106±96 |
| 3 | 1.23E+09 | 6.94E+10 | 4.56E+10 | 102±60 | 108±58 | 251±239 |
Exosomes are commonly quantified using protein
concentration and particle numbers. The ratio of particle number to
protein concentration has been used to assess co-isolation of
protein contaminants and exosome purity (12). Overall, significant differences in
this ratio were found among the three exosomes isolation methods
(UC, ExoQuick and TEI) in CCM (P=0.005) (Fig. 2D). Exosomes isolated by UC showed
a higher ratio than those from ExoQuick (UC vs. ExoQuick: 8.86±0.10
vs. 8.00±0.18, P=0.024) and TEI (UC vs. TEI: 8.86±0.11 vs.
7.99±0.34, P=0.049) in CCM (Fig.
2D), indicating the lowest level of protein contaminants and
highest purity. No differences were found between ExoQuick and TEI.
Although the differences were not significant (P>0.05), the same
trend for UC to have a higher ratio was also observed in serum (UC
vs. ExoQuick vs. TEI: 7.66±1.30 vs. 7.17±1.03 vs. 7.09±1.19)
(Fig. 2D). Interestingly, the
finding that exosomal markers were the most highly enriched in UC
exosome preparations (Fig. 2A)
also demonstrated a higher purity than that obtained with ExoQuick
and TEI.
Upon comparison of exoRNA extraction,
Route_3 and Route_e show the highest quantity and recovery among
the five extraction methods for CCM and the six methods for serum,
respectively
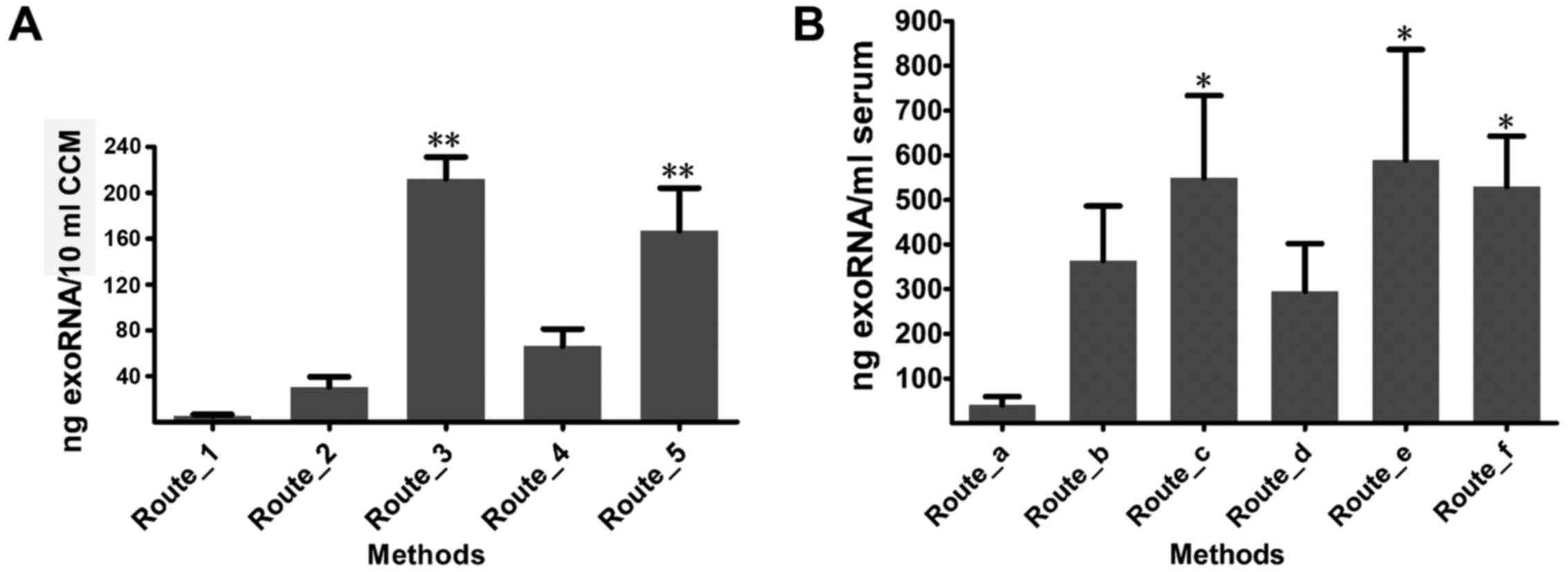
To evaluate the influence of isolation methods on
the exoRNA yield of CCM, five combinations of extraction methods
for exosomes and exoRNA (Route_1 to Route_5) were compared
(Fig. 3A). The experiment was
repeated three times using three CCM samples. Among these methods,
Route_3 showed the highest extraction efficiency (212.13±19.14 ng
RNA/10 ml CCM), and was followed by Route_5 with a concentration of
167.06±37.02 ng RNA/10 ml CCM. In addition, the RNA concentrations
obtained using Route_3 and Route_5 were significantly higher than
those from the other three routes [Route_1 (5.48±1.25), Route_2
(30.71±8.84) and Route_4 (66.18±15.05)] (P<0.01). Overall, when
a high exoRNA yield from CCM is needed, Route_3 and Route_5 are the
recommended methods.
With respect to exoRNA isolation from serum, six
combinations of methods (Route_a to Route_f) were used. The
experiments were repeated three times using samples from three
different donors (Fig. 3B) which
may explain the variation in exoRNA quality. Overall, Route_e
recovered the most exoRNA among all the methods (589.20±247.26 ng
RNA/ml serum), and was followed by Route_c (549.00±184.66 ng RNA/ml
serum) and Route_f (530.00±112.69 ng RNA/ml serum). Route_c,
Route_e and Route_f extracted a higher yield of exoRNA than Route_a
(40.45±19.21 ng/ml serum) (P<0.05). Therefore, on the basis of
exoRNA yield, Route_e, followed by Route_c and Route_f could be
recommended for serum exoRNA isolation.
The exoRNA obtained using Route_4 and
Route_5 (for CCM), or Route_e (for serum) shows the highest yield
and the most uniform size of small RNA
Total exoRNA recovery from CCM and serum was
characterized using an Agilent 2100 Bioanalyzer (Fig. 4). It was shown that the majority
of exoRNA from CCM was small (<200 nt), but there were
variations in RNA quantity and quality among the different methods.
Route_1, Route_2 and Route_3 presented not only the main peak
around 100 nt but also some longer RNA species including 18S and
28S rRNA. However, Route_4 and Route_5 showed the best quality of
small RNA which was free of 18S RNA contamination; therefore,
Route_4 and Route_5 may be more suitable for small exoRNA
research.
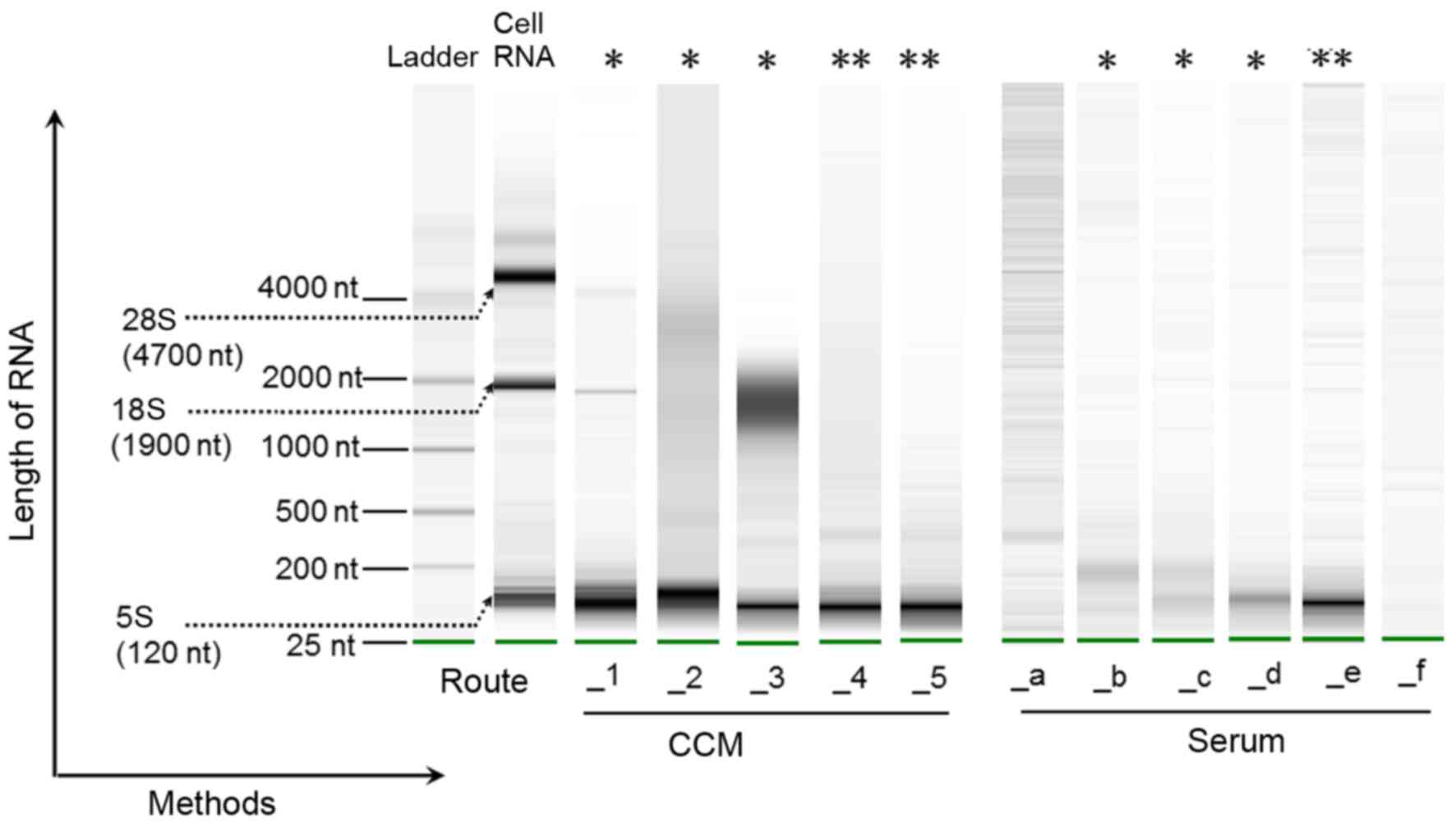 | Figure 4Bioanalyzer analysis of total exosomal
RNA (ExoRNA) by Agilent RNA Pico chip. The experiment was repeated
three times and the trend was the same, thus one of the results is
shown. The RNA 6000 ladder standard (in the first lane) contains
six RNA fragments ranging in size from 0.2 to 6 kb. Representative
bands of cellular RNA (in the second lane) showed 5S (120 nt), 18S
(1,900 nt) and 28S rRNA (4,700 nt). In bands of exoRNA from cell
culture medium (CCM), almost all of the samples showed an obvious
band in the small RNA area. Among the five combination methods,
Route_4 and Route_5 (labeled by **) showed a narrow size
distribution pattern of small RNA around 100 nt. Some longer RNA
species, including 18S and 28S ribosomal RNA, were found in bands
obtained using Route_1, Route_2 and Route_3 (labeled by *). For
exoRNA from serum, Route_e (labeled by **) had the most obvious
band in the position of small RNA, and was followed by Route_b,
Route_c and Route_d (labeled by *). No visible bands could be found
in samples from Route_a and Route_f. |
For serum exoRNA, 18S and 28S ribosomal RNA were
absent from the samples produced by all methods (Fig. 4). Overall, Route_e (exoRNeasy)
resulted in the highest yield and the most uniform size of small
RNA, indicating the highest extraction efficiency of small exoRNA
from serum among the six combinations of methods (Route_a to
Route_f).
Isolation methods influence small RNA
constituent ratio, miRNA content and amount
To examine the effect of the isolation methods on
the exoRNA profile of CCM, RNA sequencing analysis was performed
using three exoRNA samples (100 ng exoRNA/sample) isolated from the
same CCM by three commonly used and representative routes: Route_1
isolated by traditional UC and TRIzol-LS methods, Route_3 isolated
by kits from the Systems Biosciences (SBI) Company (ExoQuick and
SeraMir), and Route_5 isolated by kits from Life Technology (TEI
and TER). Among the three samples, we obtained 12.56 million raw
reads from Route_1, 9.73 million reads from Route_3 and 12.61
million reads from Route_5. After trimming low-quality reads,
5,994,511 (51.21%) clean reads from Route_1, 8,030,999 (87.80%)
from Route_3 and 9,379,868 (83.86%) from Route_5 could be mapped to
known RNAs of the human genome. These mappable sequences were
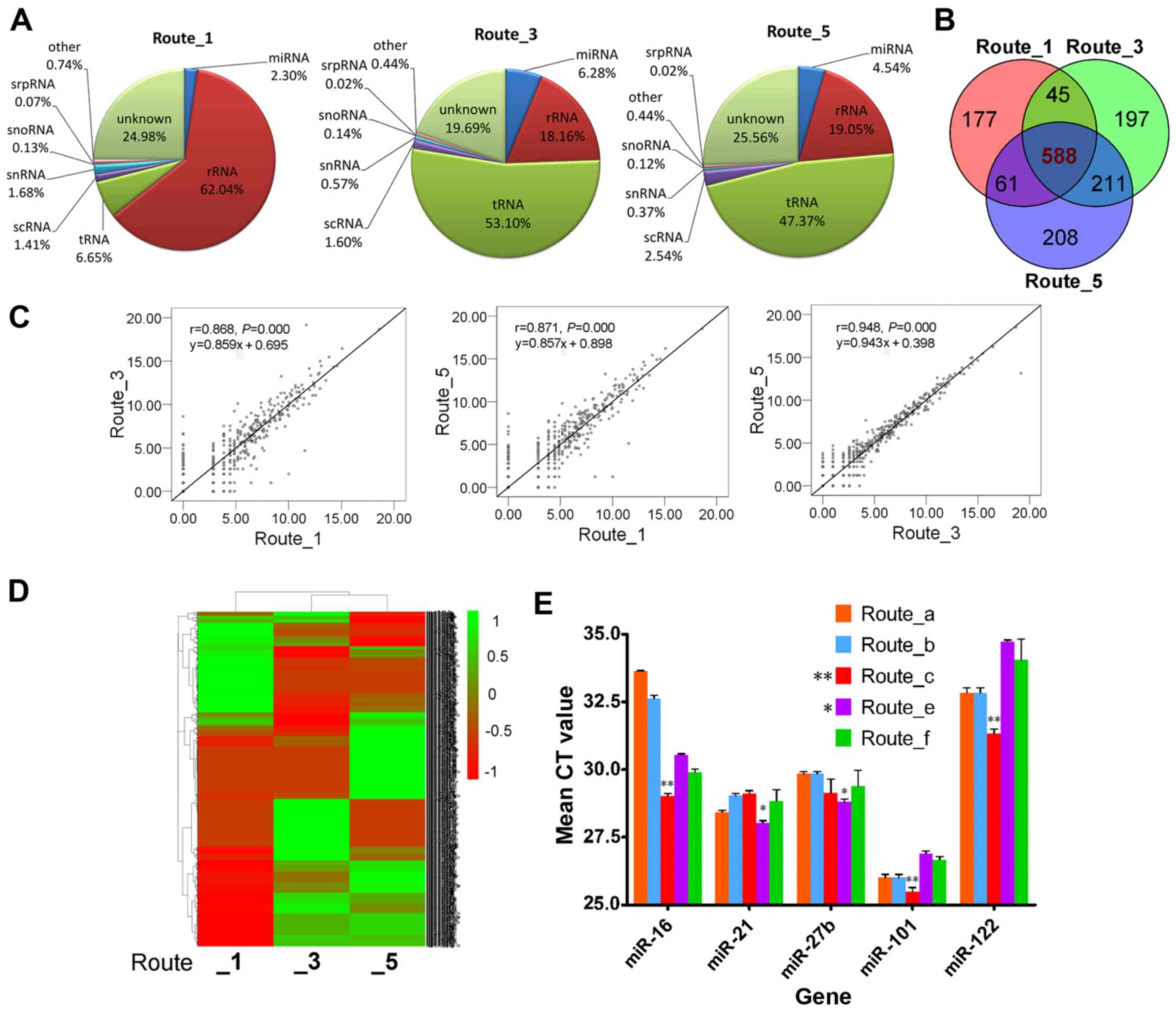
annotated to miRNA and other small non-coding RNAs. The percentages
of miRNA, rRNA, tRNA and other RNA are shown in a pie chart
(Fig. 5A): rRNA or tRNA was the
most abundant small RNA in the three samples. Route_1 samples
mainly contained rRNA (62.4% of all small RNA), but tRNA was the
main RNA in samples from Route_3 (53.1%) and Route_5 (47.37%). In
addition, Route_3 (6.28%) yielded a higher percentage of miRNA than
Route_1 (2.3%) and Route_5 (4.54%) (Fig. 5A).
We defined detectable miRNAs as those that had at
least one sequence per million mappable miRNA reads. Accordingly,
we detected 402 known hsa-miRNAs in samples from Route_1, 570 known
hsa-miRNAs in samples from Route_3 and 574 known hsa-miRNAs in
samples from Route_5. To demonstrate the methodological variations,
we investigated whether the miRNAs were unique to or common to the
different exoRNA preparation protocols. Among all detectable
miRNAs, 588 miRNAs were detected in samples obtained using all
three routes (Fig. 5B). In
addition to the miRNAs shared by three samples, or at least by two
samples, some miRNAs were also identified that were unique to each
isolation method. For example, miR-33b-3p, miR-500a-5p and
miR-1247-3p were only detected in exoRNA samples isolated by
Route_1, while miR-433-3p, miR-338-5p and miR-1262 were only found
with Route_3 and miR-219a-5p, miR-30c-1-3p and miR-616-5p only with
Route_5.
The 100 most abundant miRNAs made up 95.3–96.9% of
the detectable miRNA sequences, among which the top 20 abundant
miRNA are shown in Table II. The
top 20 abundant miRNAs obtained with the different isolation
methods were similar, but methodological variation was also evident
in these miRNAs. The rank of each miRNA obtained by the three
routes was different. To examine the variation in the miRNA content
resulting from methodological variability, a correlation analysis
was performed (using log2 transformed values after normalizing
reads to per million counts). Overall, correlations were found
among the three pairs of samples (P=0.000 for all) (Fig. 5C). The correlation (r) values were
0.868 between Route_1 and Route_3, 0.871 between Route_1 and
Route_5, and 0.948 between Route_3 and Route_5 (Fig. 5C), indicating that similar samples
were produced by the two commercial kits used in Route_3 and
Route_5. However, hierarchical clustering analysis (Fig. 5D) showed that significant
differences existed among the methods, although there was higher
similarity between Route_3 and Route_5.
 | Table IIThe 20 most abundant exosome miRNA of
three samples isolated by the different methods from CCM. |
Table II
The 20 most abundant exosome miRNA of
three samples isolated by the different methods from CCM.
| miRNA ID | Route_1
| Route_3
| Route_5
|
|---|
| TOP | Value | TOP | Value | TOP | Value |
|---|
| hsa-miR-21-5p | 1 | 396,340 | 1 | 419,405 | 1 | 37,2135 |
| hsa-miR-27b-3p | 2 | 34,483 | 2 | 89,643 | 2 | 75,594 |
| hsa-miR-30a-5p | 5 | 14,547 | 3 | 44,750 | 3 | 43,926 |
| hsa-miR-24-3p | 8 | 8,041 | 4 | 32,584 | 5 | 25,602 |
| hsa-miR-100-5p | 3 | 24,660 | 7 | 22,281 | 6 | 32,235 |
| hsa-let-7i-5p | 4 | 19,625 | 5 | 20,411 | 4 | 31,015 |
| hsa-miR-22-3p | 7 | 8,780 | 6 | 22,380 | 7 | 19,343 |
| hsa-miR-23a-3p | 16 | 3,984 | 8 | 18,174 | 9 | 14,045 |
| hsa-miR-27a-3p | 10 | 7,114 | 10 | 11,330 | 8 | 14,362 |
| hsa-let-7g-5p | 9 | 7,737 | 9 | 12,918 | 12 | 8,507 |
|
hsa-miR-378a-3p | 14 | 4,376 | 11 | 10,457 | 11 | 9,142 |
| hsa-miR-182-5p | 20 | 630 | 12 | 9,714 | 14 | 9,621 |
| hsa-miR-192-5p | 6 | 11,837 | 13 | 7,018 | 13 | 8,728 |
| hsa-miR-30d-5p | 17 | 3,274 | 15 | 5,890 | 10 | 9,057 |
| hsa-miR-29a-3p | 13 | 5,223 | 14 | 5,957 | 16 | 6,668 |
|
hsa-miR-151a-3p | 12 | 5,303 | 16 | 5,051 | 17 | 7,640 |
| hsa-miR-224-5p | 18 | 2,449 | 17 | 4,898 | 15 | 6,698 |
| hsa-miR-423-3p | 11 | 5,585 | 20 | 4,355 | 19 | 5,625 |
| hsa-miR-26a-5p | 15 | 4,079 | 18 | 3,302 | 18 | 5,763 |
| hsa-miR-25-3p | 19 | 1,594 | 19 | 4,708 | 20 | 3,926 |
Given the limited serum volume, we were unable to
extract sufficient RNA for sequencing analysis. Therefore, the
recovery of exosomal microRNA from serum was compared by qPCR. To
confirm the presence and amount of exosomal microRNA in serum, five
microRNAs (miR-16, miR-21, miR-27b, miR-101 and miR-122) earlier
reported to be present in the circulatory system or exosomes were
analyzed by qPCR (Fig. 5E).
Overall, the miRNA showed significant differences (P<0.05) among
the different routes. Specifically, the cycle threshold (CT) value
of miR-16 was highest for Route_a (33.64±0.03), indicating the
lowest copy number in the extracted sample. Route_c resulted in the
highest yield of miR-16 (27.05±0.53). In addition, the amounts of
miR-101 (25.49±0.27) and miR-122 (31.33±0.27) were also highest for
Route_c, indicating that Route_c provided high and stable
extraction efficiency for miRNA in serum. Route_e showed changeable
profiling, and produced the lowest level of miR-101 (26.91±0.15)
and miR-122 (34.73±0.09) but the highest level of miR-21
(28.8±0.19) and miR-27b (28.03±0.13), indicating selectivity of the
different miRNAs isolated by this method.
Discussion
In the present study, different traditional methods
and commercial kits were tested and compared to define the most
suitable isolation protocols for exosomes or exoRNA for use with
CCM and serum samples. The results were as follows. i) For exosome
isolation, two commercial kits (ExoQuick and TEI) showed higher
extraction efficiency than traditional UC, but UC samples had less
protein contamination than those from ExoQuick and TEI. ii) Route_3
and Route_5 showed high and stable exoRNA recovery from CCM. iii)
Route_e, followed by Route_c and Route_f, showed high exoRNA
recovery from serum; iv) high yield and narrow size distribution
pattern of small RNA presented in exoRNA isolated by Route_4 and
Route_5 from CCM, and Route_e from serum. v) RNA sequencing
analysis proved that isolation method can influence the small RNA
profile.
As a general rule, at least two different
technologies should be used to characterize individual EVs.
Therefore, three methods, western blot analysis, TEM and NTA, were
used to demonstrate the presence of exosomes in this study.
According to the minimal requirements for EV studies suggested by
Lötvall et al (21),
investigators should report the amount of several (three or more)
'exosome-enriched' and non-EV protein markers in any EV
preparation. Calnexin, the marker for non-EV components, was not
expressed in any of the exosome samples, indicating the absence of
contamination with intracellular proteins during exosome
purification. The cytosolic protein (TSG101) and the transmembrane
protein (CD63) were present in all exosome preparations. However,
the other transmembrane protein (CD9) was found only in exosomes
isolated by UC, but not in those obtained using the two kits. As
Lötvall et al recommend (21), the composition of EVs should
ideally be compared with that of the secreting cells. We found that
exosomes isolated by UC had a comparable (CD9) or even greater
(CD63 and TSG101) level of enrichment of the EV components than the
originating cells. The weak bands were found in CCM, indicating
that the supernatant 70 min post ultracentrifugation still
contained significant quantities of the remaining EVs (22).
NTA technology can rapidly calculate the total
number and the overall size distribution of vesicles. However, a
criticism of this technology, which is based on light scattering
and Brownian motion, is that it cannot differentiate adequately
among synthetic nanoparticles, large protein aggregates and
biologic vesicles (23). Although
protein quantification is the method most frequently used to
estimate the number of exosomes, it may overestimate this number by
detecting contaminating proteins that are not associated with
exosomes (24). On considering
the advantages and disadvantages of the two approaches, a
combination of both particle number and protein concentration
analysis was used to analyze the efficiency and purity of exosome
isolation. Currently, the ratio of particles to protein
concentration (particle number/microgram of protein) is accepted as
a good indicator of particle purity. Some precipitation protocols
produce highest high yield of particles, yet have low ratios of
particles to protein, potentially due to co-isolation of
contaminants (12,25).
Our analysis of NTA showed that particle numbers
obtained using UC were lower than those of the two commercial kits
(ExoQuick and TEI), but the results of western blotting showed that
samples isolated by UC methods were more enriched for exosome
markers. This may be caused by the following: i) biologic vesicles
may be damaged by repeated ultracentrifugation steps, which causes
the low particle recovery of UC methods; ii) polymeric
precipitation (ExoQuick and TEI) shows higher particle recovery,
but the lack of a method to separate exosomes and high-density
protein aggregates may also result in contamination with free
proteins in the exo-pellet samples, especially in serum samples
with abundant albumin (26). In
addition, the ratio of particles to protein also suggests different
degrees of protein contamination in samples from the three methods.
The UC method provided the lowest exosome recovery rate, but a
higher ratio indicated higher protein purity of exosomes obtained
by UC than with ExoQuick and TEI. Therefore, UC is more suitable
for proteomic research, which requires higher protein purity of
exosomes. Notably, our results are consistent with the general
consensus but partially in opposition to the research of Caradec's
group (24). They found less
albumin contamination and better exosome enrichment in exosomes
isolated by ExoQuick rather than by UC. They based this conclusion
on the western blot result of LAMP2 but not on the exosome markers
(CD9, CD63 and TSG101) used in this study. The relative proportions
of different proteins (including exosome markers) seem to vary in
the different types of EV (21).
Therefore, subsets of EVs isolated by various methods may be
different. The amount of blood collected from patients may be
limited, and although UC showed higher protein purity of exosomes,
it is not always applicable to clinical samples given the large
volume of starting material required and the low extraction
efficiency of this method. A recent study reported that the qEV
column from Izon Science Company which extracted exosomes on the
basis of size exclusion chromatography, provided exosomes with both
superior purity and higher exosome recovery rate than UC combined
with density gradient purification, and therefore this may be an
alternative method for proteomic research using CCM or clinical
samples (12). Although ExoQuick
and TEI samples may be contaminated with proteins, in some exoRNA
research, when the analysis may be less influenced by proteins, the
two kits may be good choices because of their high extraction
efficiency.
In the RNA isolation step, we compared the RNA yield
and purity of the different combinations of traditional or
commercial extraction methods for exosomes and exoRNA to define the
most suitable exoRNA isolation protocol for CCM. Overall, Route_3
showed the highest exoRNA recovery from CCM and Route_1 showed the
lowest exoRNA recovery. Because Route_3 uses dispersive
electrophoresis strips not only in the small RNA area (<200 nt)
but also in the long RNA area (1000–2000 nt), it may be more
suitable for total RNA research rather than small RNA research.
Although the Route_3 methods (ExoQuick-TC and SeraMir) are often
recommended for use in combination, we found that either
ExoQuick-TC combined with other RNA isolation methods (Route_4) or
SeraMir combined with other exosome isolation methods (Route_2) was
also feasible. Despite the fact that the exoRNA recovery is
reduced, the long RNA banding weakens or even disappears after
either the exosome or the exoRNA extraction method is changed.
These changes would be more applicable for small RNA analysis.
Intriguingly, Route_5 showed high exoRNA recovery and also small
RNA bands with higher quality, indicating superior methods for CCM
small exoRNA research.
For serum exoRNA extraction, Route_e (ExoRNeasy)
showed high recovery yield and also the most apparent and narrow
well-size distribution pattern of small RNA. In addition,
researchers can isolate exoRNA by using this kit alone without a
specific exosome isolation process. The spin columns have the
capacity to handle up to a 4-ml sample volume, enabling use of
finite resources such as clinical samples or concentrated CCM.
Therefore, exoRNeasy is an improvement, and is a rapid method that
could be easily adapted to the clinical laboratory (14). Of note, although the miRNeasy (in
Route_d) and exoRNeasy kits came from the same company, exoRNeasy
is specifically designed for exoRNA isolation. Our finding that
Route_e showed a more obvious band of small RNA than Route_d
indicated that exoRNeasy was more suitable for exoRNA isolation
than miRNeasy. Moreover, Route_b, Route_c can be used as an
alternative method for extraction of exoRNA owing to the high RNA
recovery and visible bands in the small RNA area.
In the last step, the RNA cargo of the isolated
exosomes was analyzed. High sequencing analysis of CCM exoRNA
profiling revealed that the RNA content was similar, but some
differences existed among samples obtained using the various
isolation methods. Firstly, the primary data showed that more clean
reads were found in samples from commercial kits (Route_3 and
Route_5) than from the traditional Route_1. In addition, the final
sequencing results after mapping the data base revealed that higher
proportions of miRNA were found in exoRNA isolated by kit methods
than with UC. Therefore, Route_3 and Route_5 are also applicable to
the preparation of exoRNA for small RNA sequencing, and may have
more advantages for miRNA sequencing. Finally, most miRNAs were
common to the three methods and Pearson correlation analysis
demonstrated good correlations among them. However, a higher
correlation coefficient was found between the two kits (Route_3 and
Route_5), and we also discovered more similarity between the two
kit methods in constituent ratio analysis of small RNA and cluster
analysis of miRNA. These results may be explained by the similar
experimental principles used in Route_3 and Route_5.
With regard to the miRNA expression of exosomes in
serum, among the three methods tested Route_c had the highest level
of detected miRNA, indicating a good and stable extraction
efficiency of miRNA. Variations in miRNAs were also found in
samples obtained using the different routes. For example, Route_e
performed best for miR-21 and miR-27b but worse for miR-101 and
miR-122 expression. In contrast, Route_c performed worst for miR-21
but best for miR-101 and miR-122 expression. A conclusion from our
experiments is that different isolation methods have different
affinity and performance for specific miRNA molecules, which is in
good agreement with previous studies (13,15,27).
The major limitation of the study is that each
exosome isolation method was grouped together with a particular
exoRNA purification procedure, and not with the others. Although
this design is helpful in finding the optimum extraction mode, the
differences in the initial exosome isolation procedure could affect
the results of exoRNA isolation. Considering that some researchers
have decided on the exosome isolation method to be used in their
laboratories, they may be more concerned with the efficiency of
exoRNA extraction methods. Therefore, in further studies, the same
initial exosomes should be used separately to compare each exoRNA
isolation method.
A conclusion from this study is that different
isolation methods may account for the concentration, purity and
size discrepancies of exosomes or exoRNA, therefore it is important
to maintain consistency and use one isolation method for each
research application. This study also showed advantages and
disadvantages of each method and their application to different
types of research (Table III),
therefore providing a reference for use when choosing an isolation
method according to research design.
 | Table IIIConclusion for the characteristics of
each isolation method. |
Table III
Conclusion for the characteristics of
each isolation method.
| Route | Isolation
efficiency
| Advantage and
disadvantage | Recommended
use |
|---|
Exosomes
| exoRNA
|
|---|
| Con | Purity | Con | Purity |
|---|
| CCM |
| _1, _2 | L | H | L | M | High purity, low
extraction efficiency | Large volume
samples, proteomic research |
| _3 | H | L | H | L | High extraction
efficiency; protein contamination; exoRNA contain long RNA | Total exoRNA
analysis |
| _4 | H | L | M | H | High extraction
efficiency; protein contamination; high purity of small RNA | High sequencing or
other analysis of small RNA |
| _5 | H | L | H | H | High extraction
efficiency; protein contamination; high purity of small RNA | High sequencing or
other analysis of small RNA |
| Serum |
| _a | L | M | L | L | Low extraction
efficiency; no RNA band found in Agilent bioanalyzer analysis | Exosome isolation
of large volume samples, exosome-depleted FBS preparation |
| _b, _d | H | L | M | M | Protein
contamination | Alternative offer
for exoRNA analysis |
| _c | H | L | H | M | High level of
miRNA | Recommended methods
for miRNA qPCR analysis |
| _e | - | - | H | H | High level and
purity of small RNA, no need for exosome isolation process, handle
up to 4 ml serum | Small RNA
sequencing, miRNA qPCR analysis, recommended method easily used in
clinical laboratory |
| _f | H | M | H | L | No RNA band found
in Agilent bioanalyzer analysis | Exosome isolation
of small volume samples |
Acknowledgments
This study was funded by the National Natural
Science Foundation of China (grant no. 81371901), the Science and
Technology Program of Guangzhou (grant no. 1563000220).
References
|
1
|
van der Pol E, Böing AN, Harrison P, Sturk
A and Nieuwland R: Classification, functions, and clinical
relevance of extracellular vesicles. Pharmacol Rev. 64:676–705.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
An T, Qin S, Xu Y, Tang Y, Huang Y, Situ
B, Inal JM and Zheng L: Exosomes serve as tumour markers for
personalized diagnostics owing to their important role in cancer
metastasis. J Extracell Vesicles. 4:275222015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Melo SA, Luecke LB, Kahlert C, Fernandez
AF, Gammon ST, Kaye J, LeBleu VS, Mittendorf EA, Weitz J, Rahbari
N, et al: Glypican-1 identifies cancer exosomes and detects early
pancreatic cancer. Nature. 523:177–182. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Cocucci E, Racchetti G and Meldolesi J:
Shedding microvesicles: Artefacts no more. Trends Cell Biol.
19:43–51. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Thery C, Amigorena S, Raposo G and Clayton
A: Isolation and characterization of exosomes from cell culture
supernatants and biological fluids. Curr Protoc Cell Biol. Chapter
3: Unit 3.22. 2006. View Article : Google Scholar
|
|
6
|
Musante L, Tataruch D, Gu D, Benito-Martin
A, Calzaferri G, Aherne S and Holthofer H: A simplified method to
recover urinary vesicles for clinical applications, and sample
banking. Sci Rep. 4:75322014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Lai RC, Arslan F, Lee MM, Sze NS, Choo A,
Chen TS, Salto-Tellez M, Timmers L, Lee CN, El Oakley RM, et al:
Exosome secreted by MSC reduces myocardial ischemia/reperfusion
injury. Stem Cell Res (Amst). 4:214–222. 2010. View Article : Google Scholar
|
|
8
|
Shao H, Chung J, Lee K, Balaj L, Min C,
Carter BS, Hochberg FH, Breakefield XO, Lee H and Weissleder R:
Chip-based analysis of exosomal mRNA mediating drug resistance in
glioblastoma. Nat Commun. 6:69992015. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Tauro BJ, Greening DW, Mathias RA, Ji H,
Mathivanan S, Scott AM and Simpson RJ: Comparison of
ultracentrifugation, density gradient separation, and
immunoaffinity capture methods for isolating human colon cancer
cell line LIM1863-derived exosomes. Methods. 56:293–304. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Zhu L, Qu XH, Sun YL, Qian YM and Zhao XH:
Novel method for extracting exosomes of hepatocellular carcinoma
cells. World J Gastroenterol. 20:6651–6657. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Schageman J, Zeringer E, Li M, Barta T,
Lea K, Gu J, Magdaleno S, Setterquist R and Vlassov AV: The
complete exosome workflow solution: From isolation to
characterization of RNA cargo. BioMed Res Int. 2013:2539572013.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Lobb RJ, Becker M, Wen SW, Wong CS,
Wiegmans AP, Leimgruber A and Möller A: Optimized exosome isolation
protocol for cell culture supernatant and human plasma. J Extracell
Vesicles. 4:270312015. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Eldh M, Lötvall J, Malmhäll C and Ekström
K: Importance of RNA isolation methods for analysis of exosomal
RNA: Evaluation of different methods. Mol Immunol. 50:278–286.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Enderle D, Spiel A, Coticchia CM, Berghoff
E, Mueller R, Schlumpberger M, Sprenger-Haussels M, Shaffer JM,
Lader E, Skog J, et al: Characterization of RNA from exosomes and
other extracellular vesicles isolated by a novel spin column-based
method. PLoS One. 10:e01361332015. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Rekker K, Saare M, Roost AM, Kubo AL,
Zarovni N, Chiesi A, Salumets A and Peters M: Comparison of serum
exosome isolation methods for microRNA profiling. Clin Biochem.
47:135–138. 2014. View Article : Google Scholar
|
|
16
|
Van Deun J, Mestdagh P, Sormunen R,
Cocquyt V, Vermaelen K, Vandesompele J, Bracke M, De Wever O and
Hendrix A: The impact of disparate isolation methods for
extracellular vesicles on downstream RNA profiling. J Extracell
Vesicles. 3:32014. View Article : Google Scholar
|
|
17
|
El-Khoury V, Pierson S, Kaoma T, Bernardin
F and Berchem G: Assessing cellular and circulating miRNA recovery:
The impact of the RNA isolation method and the quantity of input
material. Sci Rep. 6:195292016. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Li M, Rai AJ, DeCastro GJ, Zeringer E,
Barta T, Magdaleno S, Setterquist R and Vlassov AV: An optimized
procedure for exosome isolation and analysis using serum samples:
Application to cancer biomarker discovery. Methods. 87:26–30. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Quackenbush JF, Cassidy PB, Pfeffer LM,
Boucher KM, Hawkes JE, Pfeffer SR, Kopelovich L and Leachman SA:
Isolation of circulating microRNAs from microvesicles found in
human plasma. Methods Mol Biol. 1102:641–653. 2014. View Article : Google Scholar
|
|
20
|
Josson S, Gururajan M, Sung SY, Hu P, Shao
C, Zhau HE, Liu C, Lichterman J, Duan P, Li Q, et al: Stromal
fibroblast-derived miR-409 promotes epithelial-to-mesenchymal
transition and prostate tumorigenesis. Oncogene. 34:2690–2699.
2015. View Article : Google Scholar
|
|
21
|
Lötvall J, Hill AF, Hochberg F, Buzás EI,
Di Vizio D, Gardiner C, Gho YS, Kurochkin IV, Mathivanan S,
Quesenberry P, et al: Minimal experimental requirements for
definition of extracellular vesicles and their functions: A
position statement from the International Society for Extracellular
Vesicles. J Extracell Vesicles. 3:269132014. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Cvjetkovic A, Lötvall J and Lässer C: The
influence of rotor type and centrifugation time on the yield and
purity of extracellular vesicles. J Extracell Vesicles. Mar
25–2014.Epub ahead of print. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Gercel-Taylor C, Atay S, Tullis RH,
Kesimer M and Taylor DD: Nanoparticle analysis of circulating
cell-derived vesicles in ovarian cancer patients. Anal Biochem.
428:44–53. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Caradec J, Kharmate G, Hosseini-Beheshti
E, Adomat H, Gleave M and Guns E: Reproducibility and efficiency of
serum-derived exosome extraction methods. Clin Biochem.
47:1286–1292. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Webber J and Clayton A: How pure are your
vesicles? J Extracell Vesicles. Jan 10–2013.Epub ahead of print.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Lamparski HG, Metha-Damani A, Yao JY,
Patel S, Hsu DH, Ruegg C and Le Pecq JB: Production and
characterization of clinical grade exosomes derived from dendritic
cells. J Immunol Methods. 270:211–226. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Moret I, Sánchez-Izquierdo D, Iborra M,
Tortosa L, Navarro-Puche A, Nos P, Cervera J and Beltrán B:
Assessing an improved protocol for plasma microRNA extraction. PLoS
One. 8:e827532013. View Article : Google Scholar :
|