Introduction
Hepatocellular carcinoma (HCC) is a common malignant
tumor and the second most common cause of cancer-associated
mortality globally (1). The
overall survival rate is poor for patients with HCC, with a 5-year
survival rate of <10% (2).
Surgery is the most effective treatment for HCC, however ~80% of
patients with HCC are ineligible for surgery due to advanced
disease and 20-25% of patients will experience postoperative
recurrence within 5 years (2).
Chemotherapy serves an important role in the treatment of patients
with advanced HCC, however it has low efficacy primarily due to
apoptosis resistance (3).
However, the mechanism of apoptosis resistance remains unclear.
Cyclooxygenase-2 (COX-2) is a rate-limiting enzyme
that serves a role in the conversion of arachidonic acid to
prostaglandin H2 (4). COX-2 is
typically undetectable in healthy tissues under normal
physiological conditions, however, cytokines including tumor growth
factor, interleukin (IL)-1 and IL-6, hypoxia, Helicobacter
pylori infection and carcinogenic substances cells may trigger
the rapid and transient expression of COX-2 to modulate
inflammation and carcinogenesis (5). COX-2 is associated with a number of
tumor progression processes, including the stimulation of tumor
cell growth, the inhibition of tumor cell apoptosis, the promotion
of tumor angiogenesis and the enhancement of tumor invasion and
metastasis (6). A number of
studies have demonstrated that COX-2 is overexpressed in HCC cells
and serves an important role in apoptosis resistance (7,8).
It has been reported that COX-2 overexpression inhibits tumor cell
apoptosis; Leng et al (9)
revealed that transfecting HCC cells with COX-2 promotes cell
growth and resistance to butyrate-induced apoptosis. In addition,
the COX-2 inhibitor celecoxib suppresses the carcinogen diethyl
nitrosamine, which induces HCC growth (10). However, the potential mechanisms
underlying COX-2-mediated apoptosis resistance have not yet been
fully elucidated.
To maintain a survival advantage in the tumor
microenvironment, tumor cells generate energy mainly via aerobic
glycolysis even under aerobic conditions-this is called the
‘Warburg effect’ (11,12). Pyruvate kinase (PK) is a
rate-limiting enzyme in the glycolysis pathway and has four
isoenzymes, including L, R, M1 and M2. M1 PK is mainly expressed in
the skeletal muscle and brain tissue, which have high energy and
oxygen consumption (13,14). A critical mediator of the Warburg
effect is the PK M2 isoform (PKM2), a tumor-specific isoform of PK
that catalyzes the synthesis of pyruvate and ATP using
phosphoenolpyruvate (PEP) and ADP as substrates (15). PKM2 is primarily expressed in
embryonic tissues and proliferative cells, and elevated PKM2
expression has been reported in several tumor types, including HCC
(16,17). One previous study confirmed that
HCC cells express higher level of PKM2 compared with normal
adjacent tissues or benign tumor types, while PKM2 knockout has
been reported to inhibit the growth of HCC cells (18). Dong et al (19) demonstrated that PKM2 was
overexpressed in cancer tissues compared with adjacent normal
tissues and PKM2 downregulation in HepG2 cells inhibited cell
growth via targeting hypoxia inducible factor-1α (HIF-1α) and
B-cell lymphoma-extra large expression, suggesting that PKM2 may
promote HCC cell proliferation by exerting a protein kinase
function. However, whether COX-2 mediated resistance to apoptosis
is associated with PKM2 upregulation is unclear.
A number of studies have demonstrated that COX-2
serves an important role in apoptosis resistance (20,21). However, whether the HIF-1α/PKM2
pathway is involved in COX-2-induced apoptosis resistance remains
to be elucidated. In the present study, the association between
COX-2, PKM2 and HIF-1α was assessed.
Materials and methods
Patients and tissue microarray
construction
The present study was approved by the Ethics
Committee of the First Affiliated Hospital of Anhui Medical
University and conformed to the 1975 Declaration of Helsinki. All
patients provided written informed consent. Primary HCC tissues and
paired adjacent normal liver tissues were obtained from the
archives of the Department of Pathology, the First Affiliated
Hospital of Anhui Medical University between March 2010 and June
2016 in 143 patients with HCC (105 male, 38 female; mean age 51,
range 35 to 78 years), embedded in paraffin and 10%
neutral-buffered formalin was used to fix the tissues at room
temperature for 12 h. Histological tumor differentiation was
determined using the Edmondson-Steiner scoring system (22) and tumor types were classified
using the World Health Organization classification system (23) and the International Anti-cancer
Coalition Tumor Node Metastasis classification system (24). Tissue sections were dyed using
hematoxylin staining solution (containing 0.2% hematoxylin) and
1-2% eosin at room temperature for 5 min. The sections were
reviewed for the identification of the target area by two
pathologists independently. Three to five representative 1-mm cores
were produced from each sample and then inserted into a new
recipient paraffin block in a grid pattern using a manual tissue
array (Boyikang Instruments, Beijing, China).
Reagents
Dulbecco’s modified Eagle’s medium (DMEM), fetal
bovine serum (FBS), basement membrane matrix and PBS were obtained
from Hyclone (GE Healthcare Life Sciences, Logan, UT, USA).
Antibodies against COX-2 (cat. no. ab15191), HIF-1α (cat. no.
ab8366) and PKM2 (cat. no. ab150377) were purchased from Abcam
(Cambridge, MA, USA). Si-COX-2, Si-PKM2 and Si-HIF-1α were
purchased from Shanghai GenePharma Co., Ltd. (Shanghai, China). The
Plus Western Blotting Detection System was obtained from Bio-Rad
Laboratories, Inc. (Hercules, CA, USA). Reagents for reverse
transcription-quantitative polymerase chain reaction (RT-qPCR)
analysis were as follows: The RT kit was purchased from Invitrogen
(Thermo Fisher Scientific, Inc., Waltham, MA, USA) and the
Thunderbird SYBR RT-PCR Mix was from Toyobo Life Science (Osaka,
Japan). Annexin V/propidium iodide (PI) apoptosis detection kit was
obtained from BD Biosciences (San Jose, CA, USA). The TUNEL system
was purchased from Roche Diagnostics (Basel, Switzerland). Cell
Counting Kit-8 (CCK-8) was obtained from BioTek Instruments, Inc.
(Winooski, VT, USA).
Cell culture
The HepG2 cell line is originated from
hepatoblastoma, however it was frequently used as a HCC cell
(23). The HepG2 cell line was
purchased from the Shanghai Cell Bank (Chinese Academy of Sciences,
Shanghai, China) and cultured in DMEM supplemented with 10% (v/v)
heat-inactivated FBS at 37°C in a humidified atmosphere containing
5% CO2.
Immunohistochemical (IHC) analysis
The expression of COX-2, HIF-1α and PKM2 in HCC
samples from 143 patients was detected using IHC. Following
deparaffinization and antigen retrieval, sections were blocked with
5% goat serum (OriGene Technologies, Inc., Beijing, China) at 37°C
for 30 min. All sections were incubated at 4°C overnight with
rabbit antibodies against COX-2 and PKM2 (all 1:100), and mouse
monoclonal antibody against HIF-1α (1:100). Sections
(4-µm-thick) were then incubated with horseradish peroxidase
(HRP)-labeled goat anti-rabbit immunoglobulin G (IgG) (H+L) or goat
anti-mouse IgG (H+L) secondary antibodies (1:200, cat. no.
BS13278/BS12478; Bioworld Technology Inc., St. Louis Park, MN, USA)
at 37°C for 30 min. The secondary antibodies were diluted by PBS
phosphate buffer (0.01 mol/l, pH: 7.4-7.6). Then the sections were
stained using a DAB detection kit (cat. no. ZLI-9018; OriGene
Technologies, Inc.) at room temperature for 30 sec and hematoxylin
staining solution (containing 0.2% hematoxylin) at room temperature
for 5 min. Cytoplasm or karyon staining was considered to indicate
a positive expression of COX-2, HIF-1α or PKM2. Staining intensity
was graded as follows: 0, no staining; 1, weak intensity; 2,
moderate intensity; and 3, strong intensity. The degree of positive
staining was determined as follows: 0, <24%; 1, 25-49%; 2,
50-74%; and 3, ≥75%). Patients with HCC were classified into two
groups according to the total score (staining intensity + positive
staining); the negative expression group (total score 0-2) and the
positive expression group (total score 3-6). IHC results were
quantitatively analyzed using a biological image analysis system
that consisted of an Olympus CX31 light microscope (magnification,
×200; Olympus Corporation, Tokyo, Japan), Nikon Digital Camera DXM
1200F (Nikon Corporation, Tokyo, Japan), Image-ProPlus6.0 (Media
Cybernetics, Inc., Rockville, MD, USA) and JEOA 801D morphological
biological image analysis software version 6.0 (Zhejiang Jieda
Technology, Co., Ltd., Zhejiang, China).
siRNA transfection
HepG2 cells were seeded into 6-well plates
(1×105/ml) and cultured until the adherent cells reached
70% confluence. COX-2 (forward, 5′-GGA ACG UUG UGA AUA ACA UTT-3′
and reverse, 5′-AUG UUA UUC ACA ACG UUC CTT-3′), HIF-1α (forward,
5′-GCC GCU CAA UUU AUG AAU ATT-3′ and reverse, 5′-UAU UCA UAA AUU
GAG CGG CTT-3′) and PKM2 (forward, 5′-GGC UGG ACU ACA AGA ACA
UTT-3′ and reverse, 5′-AUG UUC UUG UAG UCC AGC CTT-3′) siRNAs were
designed and synthesized by Shanghai GenePharma Co., Ltd. siRNA was
transfected into the cells at a concentration of 20 nM using
Lipofectamine® 2000 (Thermo Fisher Scientific, Inc.) at
37°C for 24 h. Cells in the negative control group were treated
with Lipofectamine® 2000. Cells were incubated with
si-COX-2, si-HIF-1α or si-PKM2 at 37°C for 8 h and then cultured in
DMEM at 37°C for a further 16 h.
RT-qPCR
Following transfection, COX-2, HIF-1α and PKM2 mRNA
expression was assessed using RT-qPCR. Briefly, total RNA was
extracted using a TRIzol® kit (Thermo Fisher Scientific
Inc.) according to the manufacturer’s protocol. The total RNA
concentration was assessed by measuring absorbance at 260 and 280
nm using a NanoDrop spectrophotometer (TU-1810; Thermo Fisher
Scientific Inc.). RT-qPCR was performed using a two-step reaction
using TransStart® All-in-One First-Strand cDNA Synthesis
SuperMix and TransStart® Top Green qPCR SuperMix kit
(Beijing Transgen Biotech Co., Ltd., Beijing, China). Following
normalization to β-actin expression, the relative gene expression
was determined using a comparative standard curve. PCR primers
specific for human genes were provided by Shanghai GenePharma Co.,
Ltd. and were as follows: β-actin forward, 5′-CTC TTC CAG CCT TCC
TTC CT-3′ and reverse, 5′-AGC ACT GTG TTG GCG TAC AG-3′; COX-2,
forward 5′-TAA AAA CCC CAT AAC CCC GCC-3′ and reverse 5′-TTG GGC
TTT TCT CCT TTG GTT-3′; HIF-1α, forward 5′-CCA CCT CTG GAC TTG CCT
TT-3′ and reverse 5′-ACT TAT CTT TTT CTT GTC GTT CGC-3′; PKM2,
forward 5′-ACT CGG GCT GAA GGC AGT GA-3′ and reverse 5′-TGT GGG GTC
GCT GGT AAT GG-3′. Following denaturation at 95°C for 3 min,
amplification was performed for 40 cycles of 95°C for 10 sec, 57°C
for 30 sec and 65°C for 10 sec, with a single fluorescence
measurement. All reactions were performed in triplicate. The
expression levels were calculated using the 2−∆∆Cq
method (25).
Western blotting
To measure the expression of COX-2, HIF-1α and PKM2
proteins, western blotting was performed following transfection
with si-COX-2, si-HIF-1α or si-PKM2. Briefly, total protein was
extracted from HepG2 cells using radioimmunoprecipitation assay
buffer (Beyotime Institute of Biotechnology, Haimen, China) at 4°C
for 30 min. Protein concentration was determined using a Lowry
Protein assay (Thermo Fisher Scientific Inc.). Approximately 20
µg proteins were loaded in each lane and subjected to
electrophoresis using 10% SDS-PAGE, and were transferred onto
polyvinylidene difluoride membranes (EMD Millipore, Billerica, MA,
USA) at 110 V for 60 min. The membrane was blocked for 2 h at room
temperature in blocking solution (5% non-fat milk in Tris-buffered
saline with 0.05% Tween-20) and incubated with primary antibodies
(1:500) at 4°C overnight. Following three washes in tris-buffered
saline with 0.05% Tween-20, the membrane was incubated with
HRP-conjugated mouse anti-rabbit IgG (cat. no. sc-2357) or
HRP-conjugated goat anti-mouse IgG (cat. no. sc-2031) secondary
antibodies (Santa Cruz Biotechnology Inc., Dallas, TX, USA) at a
dilution of 1:10,000 for 60 min at room temperature. The following
antibodies were used: COX-2 (Abcam; cat. no. ab15191), HIF-1α
(Abcam; cat. no. ab8366), PKM2 (Abcam; cat. no. ab150377) and
β-actin (Abcam; cat. no. ab8226). The membranes were incubated in a
chromogenic substrate (Thermo Fisher Scientific Inc.) 2 min in room
temperature to develop protein bands. Results were quantified by
densitometric analysis using ImageJ software (V1.48u; National
Institutes of Health, Bethesda, MD, USA).
Flow cytometry
At 24 h following transfection, non-adherent cells
were removed by gentle washing, the remaining cells were collected
in a 15 ml sterile tube and centrifuged at 400 × g at room
temperature for 5 min. Cells were washed twice with cold PBS and
resuspended in 400 µl binding buffer at a concentration of
1×105 cells/ml. The cell suspensions were then mixed
with 5 µl Annexin V-fluorescein isothiocyanate solution and
10 µl PI, incubated for 15 min at 4°C in the dark and
analyzed using flow cytometry (BD Biosciences, Franklin Lakes, NJ,
USA) within 1 h. Flow cytometric analysis was performed using
FlowJo7.6.1 software (FlowJo LLC, Ashland, OR, USA). A total of
10,000 cells from each sample were analyzed and the experiment was
repeated at least three times.
TUNEL assay
HepG2 cells (1×105) were seeded into six
well plates. Following transfection, cover slips were washed twice
with PBS and then fixed in 4% paraformaldehyde solution at room
temperature for 25 min. Apoptotic cells were detected using a TUNEL
assay (TUNEL System kit; Roche Diagnostics) according to the
manufacturer’s protocol. The results were quantitatively analyzed
using a biological image analysis system, which consisted of an
Olympus CX31 light microscope (magnification, ×400), Nikon Digital
Camera DXM 1200F and ACT-1 version 2.63 software (Nikon
Corporation).
Cell proliferation assay
HepG2 cells were seeded into 96-well plates at a
density of 2×104 cells/well for 24 h to allow for cell
adherence. Transfection with specific siRNA was performed as
described above. Subsequently, 10 µl CCK-8 reagent (Shanghai
BestBio Beibo Bio, Shanghai, China) was added to each well at 37°C
for 4 h. The optical density was measured using a scanning
multi-well spectrophotometer (BioTek Instruments, Inc., Winooski,
VT, USA) at 490 nm.
Statistical analysis
At least three independent experiments were
performed for all assays. Statistical analysis was performed using
SPSS version 17.0 software (SPSS Inc., Chicago, IL, USA). The
results are expressed as the mean ± standard error of the mean.
Differences between two or multiple groups were compared using a
independent t-test or a one-way analysis of variance and a post-hoc
test (Bonferroni’s). Associations between protein expression and
clinicopathological parameters were assessed using Spearman’s
correlation analysis. P<0.05 was considered to indicate a
statistically significant difference.
Results
Expression of PKM2 protein is positively
associated with COX-2 expression and worse clinicopathological
characteristics in patients with HCC
To investigate the function of COX-2 in patients
with HCC, formalin-fixed paraffin-embedded HCC specimens were
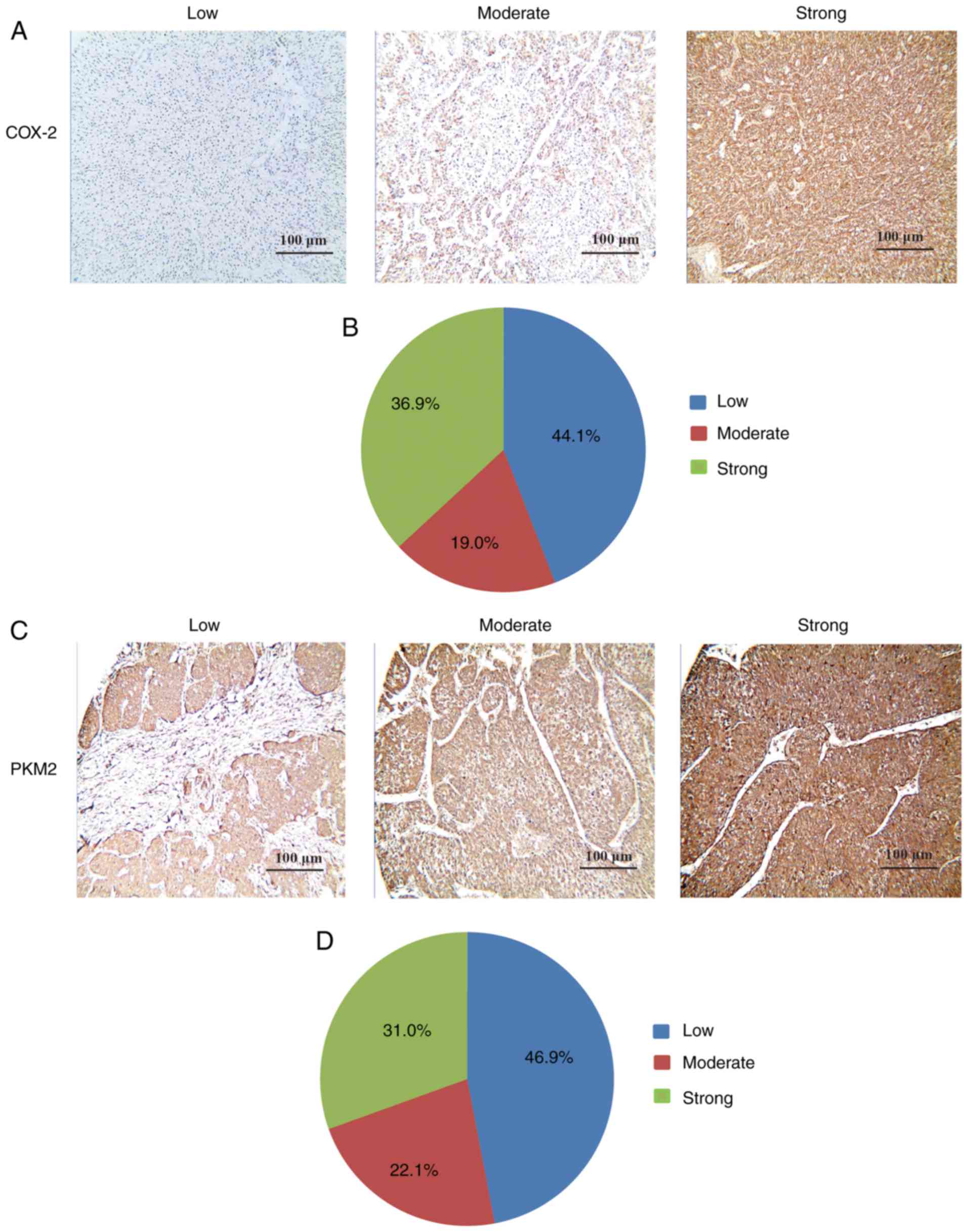
analyzed using IHC staining. It was demonstrated that COX-2 was
primarily expressed in the cytoplasm of tumor cells in a diffused
pattern (Fig. 1A). Furthermore,
55.9% (80/143) of tumor tissues had a high COX-2 expression
(moderate or strong) compared with adjacent liver tissues (Fig. 1B). The association between
clinicopathological characteristics and COX-2 expression is
presented in Table I. The
histological grade of HCC cells was significantly positively
associated with COX-2 expression; 81.2% of poorly-differentiated
tumor tissues had a high expression of COX-2, while only 37 and
66.7% of the well- and moderately-differentiated tumor cells had a
high COX-2 expression (P<0.001). However, no association was
observed between the expression of COX-2 and other
clinicopathological characteristics, including tumor size and
clinical stage (P>0.05).
 | Table IAssociation between COX-2 expression
level in hepatocellular cancer and clinicopathological factors. |
Table I
Association between COX-2 expression
level in hepatocellular cancer and clinicopathological factors.
|
Characteristics | Samples (n) | COX-2
| Positive rate
(%) | χ2 | P-value |
|---|
| Low | High |
|---|
| Sex | | | | | 2.708 | 0.100 |
| Male | 105 | 34 | 71 | 67.6 | | |
| Female | 38 | 18 | 20 | 52.6 | | |
| Age (years) | | | | | 0.001 | 0.973 |
| <60 | 101 | 46 | 55 | 54.5 | | |
| ≥60 | 42 | 19 | 23 | 54.8 | | |
| Tumor size
(cm) | | | | | 2.292 | 0.130 |
| <5 | 52 | 13 | 39 | 75.0 | | |
| ≥5 | 91 | 34 | 57 | 62.6 | | |
| Hepatitis | | | | | 0.008 | 0.931 |
| Positive | 122 | 36 | 86 | 70.5 | | |
| Negative | 21 | 6 | 15 | 71.4 | | |
| Liver
cirrhosis | | | | | 0.133 | 0.706 |
| Positive | 123 | 42 | 81 | 65.9 | | |
| Negative | 20 | 6 | 14 | 70.0 | | |
| Tumor number | | | | | 0.037 | 0.848 |
| Single | 24 | 6 | 18 | 75.0 | | |
| Multiple | 119 | 32 | 87 | 73.1 | | |
| Portal vein
invasion | | | | | 0.590 | 0.443 |
| Present | 60 | 18 | 42 | 70.0 | | |
| Absent | 83 | 30 | 53 | 63.9 | | |
| AFP | | | | | 0.277 | 0.599 |
| High | 137 | 36 | 101 | 73.7 | | |
| Normal | 6 | 1 | 5 | 83.3 | | |
| Clinical stage | | | | | 2.197 | 0.533 |
| I | 14 | 6 | 8 | 57.1 | | |
| II | 100 | 25 | 75 | 75.0 | | |
| III | 25 | 8 | 17 | 68.0 | | |
| IV | 4 | 1 | 3 | 75.0 | | |
| Histological
grade | | | | | 18.663 | <0.001 |
|
Well-differentiated | 54 | 34 | 20 | 37.0 | | |
|
Moderately-differentiated | 57 | 19 | 38 | 66.7 | | |
|
Poorly-differentiated | 32 | 6 | 26 | 81.2 | | |
IHC was used to investigate the association between
PKM2 and COX-2 in HCC cells. PKM2 was mainly expressed in the
cytoplasm of tumor tissues (Fig.
1C). Furthermore, PKM2 expression was upregulated in 53.1%
(76/143) of tumor tissues compared with adjacent liver tissues
(Fig. 1D). The association
between clinicopathological characteristics and PKM2 expression is
presented in Table II. The
histological grade of HCC cells was significantly positively
associated with PKM2 expression; 75.0% of poorly-differentiated
tumor cells had a high expression of PKM2, while only 29.6 and
63.1% of well- and moderately-differentiated tumor cells had a high
expression of PKM2 proteins (P<0.001). To assess whether PKM2
expression is associated with COX-2 activation, Spearman’s
correlation analysis was performed. As illustrated in Table III, 92.5% of COX-2 positive
tumor cells had a significantly high PKM2 expression (P<0.001),
while only 3.2% of COX-2 negative tumor cells highly expressed
PKM2. Altogether, these results suggest that COX-2 activation is
associated with a high PKM2 expression and worse
clinico-pathological characteristics in HCC patients.
 | Table IIAssociation between PKM2 expression
level in hepatocellular cancer and clinicopathological factors. |
Table II
Association between PKM2 expression
level in hepatocellular cancer and clinicopathological factors.
|
Characteristics | Samples (n) | PKM2
| Positive rate
(%) | χ2 | P-value |
|---|
| Low | High |
|---|
| Sex | | | | | 3.122 | 0.077 |
| Male | 105 | 28 | 77 | 73.3 | | |
| Female | 38 | 16 | 22 | 57.9 | | |
| Age (years) | | | | | 0.098 | 0.754 |
| <60 | 101 | 51 | 50 | 49.5 | | |
| ≥60 | 42 | 20 | 22 | 52.3 | | |
| Tumor size
(cm) | | | | | 0.949 | 0.330 |
| <5 | 52 | 11 | 41 | 78.8 | | |
| ≥5 | 91 | 26 | 65 | 71.4 | | |
| Hepatitis | | | | | 0.096 | 0.756 |
| Positive | 122 | 33 | 89 | 72.9 | | |
| Negative | 21 | 5 | 16 | 76.1 | | |
| Liver
cirrhosis | | | | | 0.284 | 0.594 |
| Positive | 123 | 38 | 85 | 69.1 | | |
| Negative | 20 | 5 | 15 | 75.0 | | |
| Tumor number | | | | | 0.163 | 0.686 |
| Single | 24 | 7 | 17 | 70.8 | | |
| Multiple | 119 | 30 | 89 | 74.8 | | |
| Portal vein
invasion | | | | | 0.817 | 0.366 |
| Present | 60 | 16 | 44 | 73.3 | | |
| Absent | 83 | 28 | 55 | 66.3 | | |
| AFP | | | | | 0.371 | 0.542 |
| High | 137 | 31 | 106 | 77.4 | | |
| Normal | 6 | 2 | 4 | 66.7 | | |
| Clinical stage | | | | | 4.235 | 0.237 |
| I | 14 | 6 | 8 | 57.1 | | |
| II | 100 | 22 | 78 | 78.0 | | |
| III | 25 | 6 | 19 | 76.0 | | |
| IV | 4 | 0 | 4 | 100.0 | | |
| Histological
grade | | | | | 20.425 | <0.001 |
|
Well-differentiated | 54 | 38 | 16 | 29.6 | | |
|
Moderately-differentiated | 57 | 21 | 36 | 63.1 | | |
|
Poorly-differentiated | 32 | 8 | 24 | 75.0 | | |
 | Table IIICorrelation between COX-2 and
PKM2. |
Table III
Correlation between COX-2 and
PKM2.
| PKM2 | COX-2
| r | P-value |
|---|
| − | + |
|---|
| Negative | 61 | 6 | 0.889 | <0.001 |
| Positive | 2 | 74 | | |
Reduced COX-2 expression promotes
apoptosis and downregulates proliferation in HepG2 cells
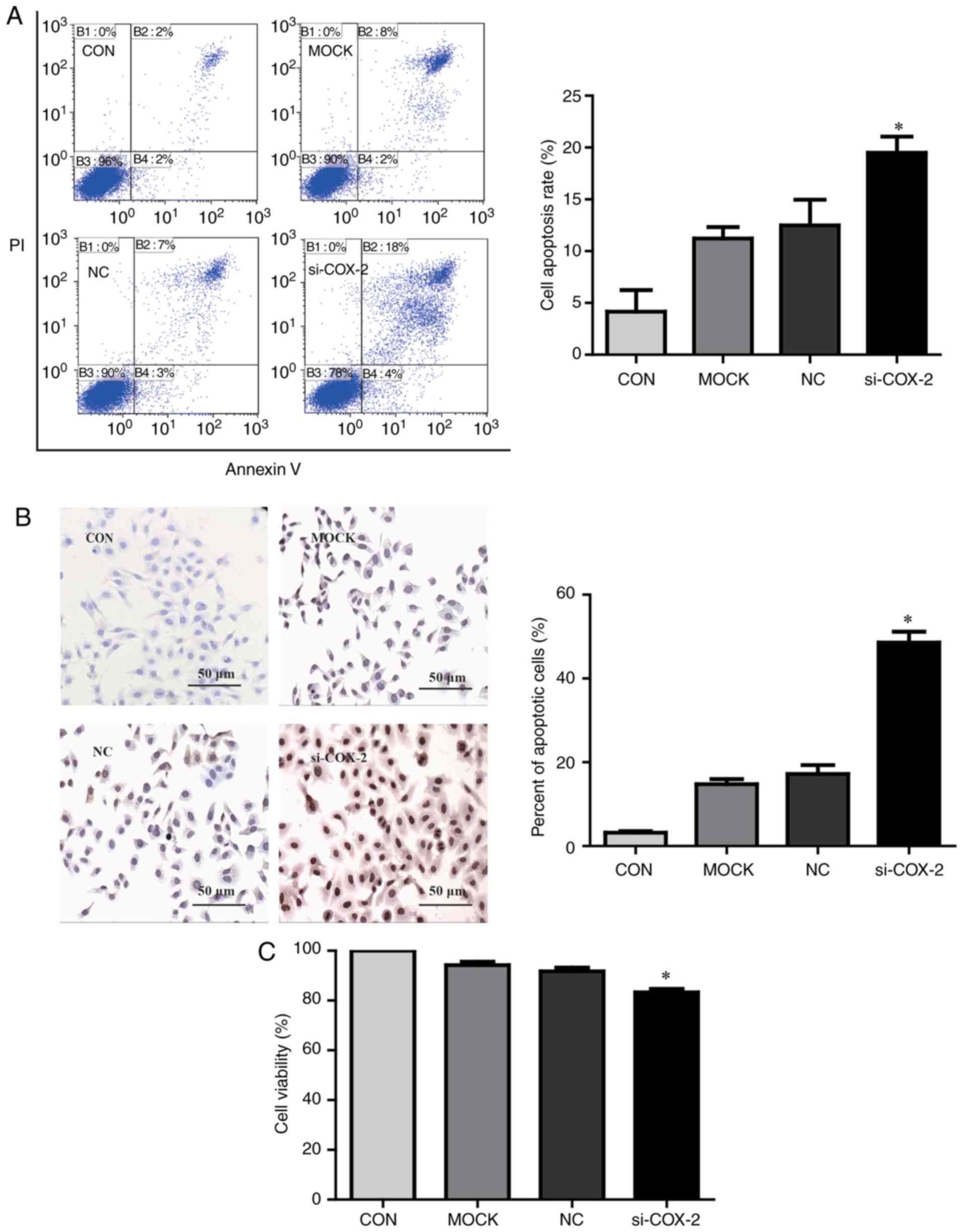
To investigate whether COX-2 influences apoptosis in
HCC cells, HepG2 cells were transfected with COX-2 siRNA for 24 h.
As presented in Fig. 2A,
apoptosis was significantly promoted in COX-2 siRNA-transfected
HepG2 cells compared with untransfected cells (P<0.05). The
results of a TUNEL assay revealed that COX-2 knockdown
significantly enhanced the apoptosis in HepG2 cells compared with
paired controls (P<0.05; Fig.
2B). Furthermore, a CCK-8 assay revealed that COX-2 knockdown
significantly reduced cell viability compared with untransfected
HepG2 cells (P<0.05; Fig.
2C).
PKM2 knockdown increases apoptosis in HCC
cells and downregulates proliferation
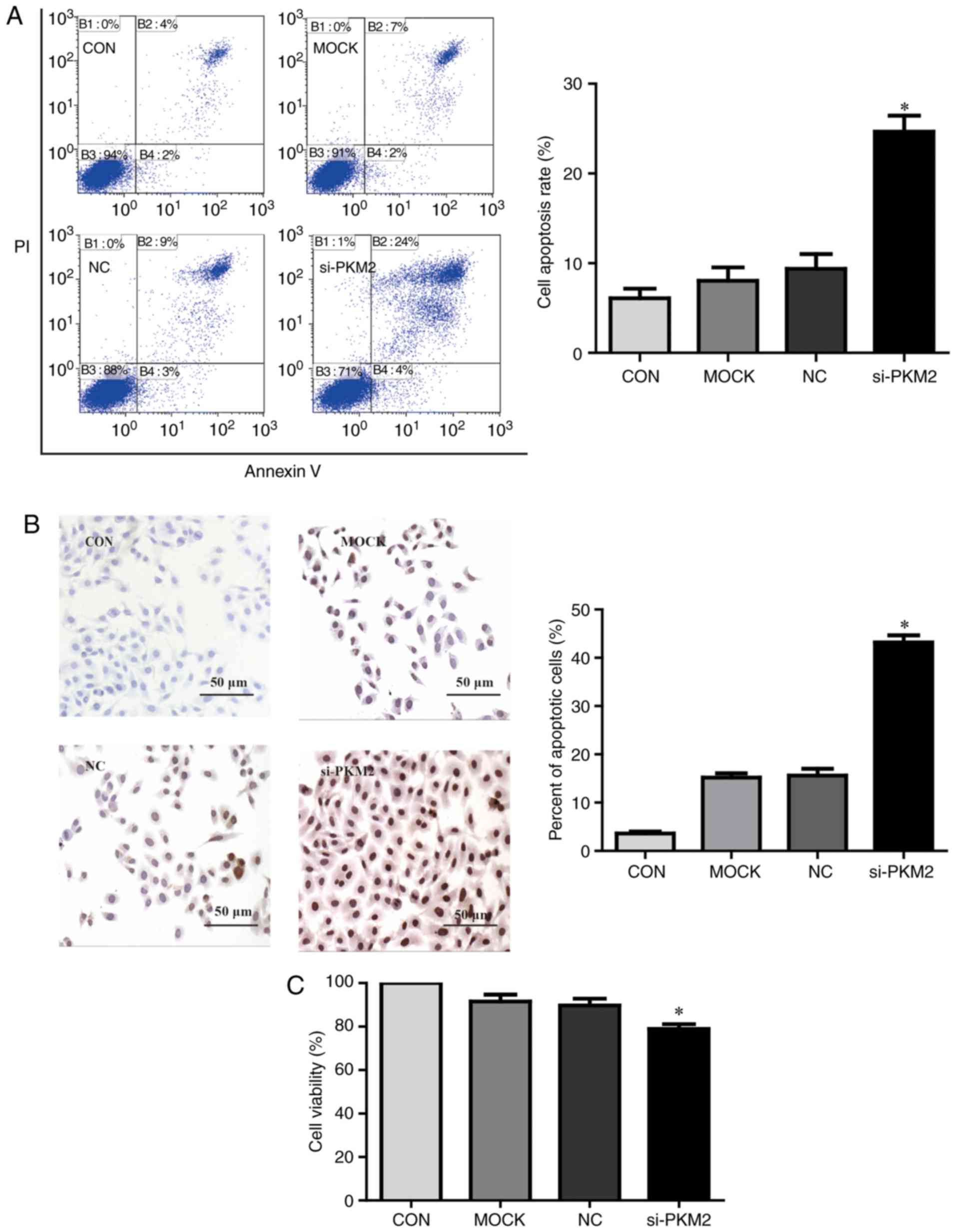
To investigate whether PKM2 affects apoptosis in HCC
cells, HepG2 cells were transfected with PKM2 siRNA. Annexin V/PI
double-staining and TUNEL assays were performed to assess
apoptosis. As presented in Fig.
3A, PKM2 knockdown resulted in a significant increase in
apoptosis compared with untransfected HepG2 cells (P<0.05). The
results of a TUNEL assay further confirmed that PKM2 knockdown in
HepG2 cells significantly increased apoptosis compared with
untransfected control cells (P<0.05; Fig. 3B). Furthermore, a CCK-8 assay
revealed that PKM2 knockdown significantly reduced cell viability
compared with untransfected HepG2 cells (P<0.05; Fig. 3C).
COX-2 regulates PKM2 expression in HCC
cells
To investigate the association between COX-2 and
PKM2 in HCC cells in vitro, HepG2 cells were transfected
with COX-2 siRNA, RT-qPCR was performed and the results revealed
that the expression levels of COX-2 (P<0.05; Fig. 4A) and PKM2 (P<0.05; Fig. 4B) mRNA were significantly reduced
compared with untransfected cells. Western blotting also confirmed
that COX-2 knockdown significantly reduced PKM2 expression at the
protein level (P<0.05; Fig.
4C). Subsequently, RT-qPCR revealed that PKM2 mRNA was
significantly downregulated in HepG2 cells following PKM2 siRNA
transfection (P<0.001; Fig.
4D); however, COX-2 mRNA expression levels were unaffected
(P>0.05; Fig. 4E). Western
blotting also revealed that PKM2 siRNA was able to effectively
downregulate PKM2 expression levels (P<0.05) but had no impact
on COX-2 expression levels (P>0.05; Fig. 4F). These results suggest that
COX-2 is able to modulate PKM2 expression in HCC cells.
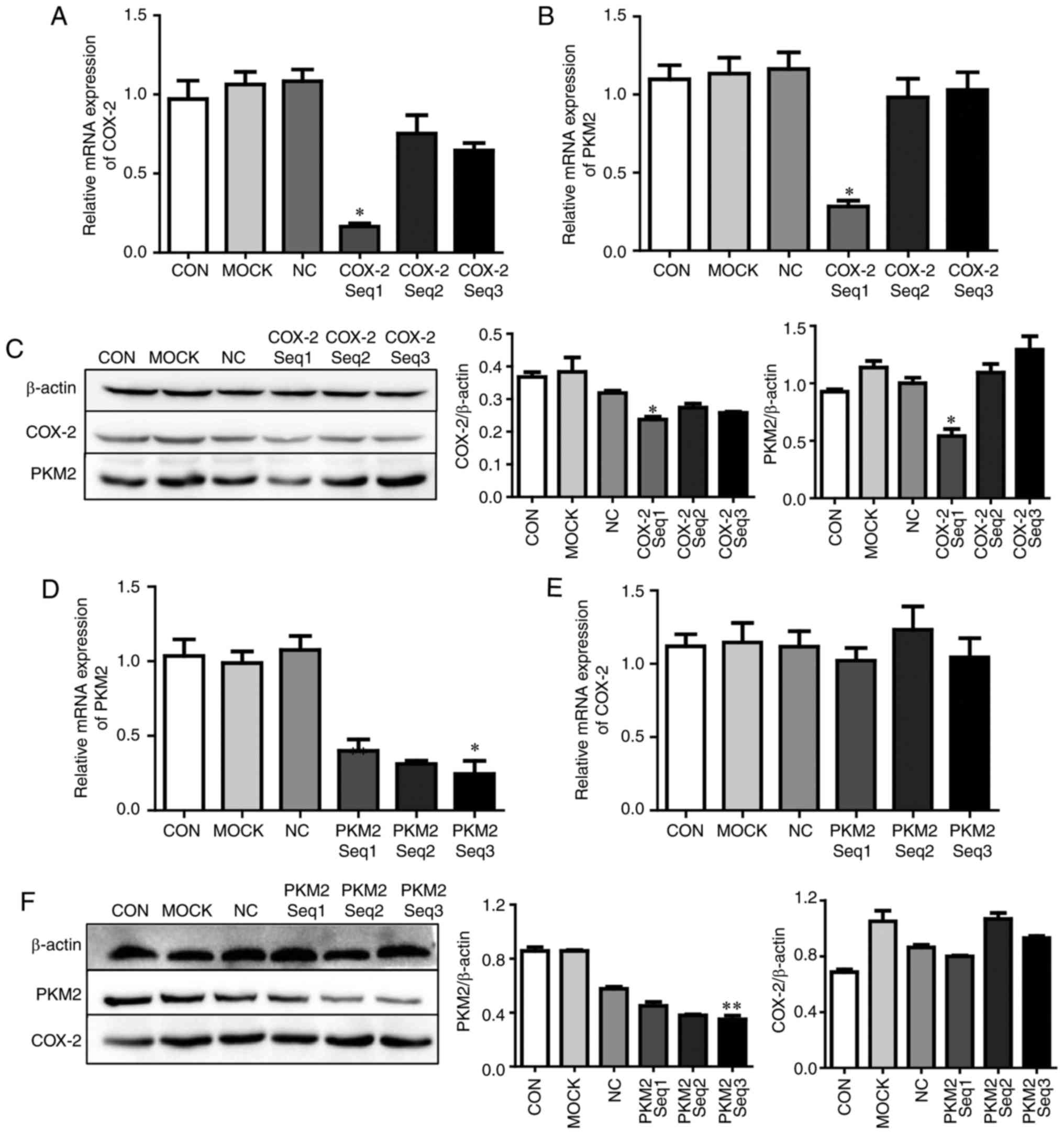 | Figure 4COX-2 regulates PKM2 expression in
HepG2 cells in vitro. Firstly, HepG2 cells were transfected
with COX-2 siRNA for 24 h. (A) COX-2 and (B) PKM2 mRNA was measured
using RT-qPCR. (C) COX-2 and PKM2 protein expression was measured
by western blotting. Secondly, HepG2 cells were transfected with
PKM2 siRNA for 24 h, and (D) PKM2 and (E) COX-2 mRNA was also
measured using RT-qPCR, and (F) COX-2 and PKM2 protein expression
was measured using western blotting. *P<0.05 and
**P<0.01 vs. CON. COX-2, cyclooxygenase-2; PKM2,
pyruvate kinase M2 isoform; siRNA, small interfering RNA; CON,
untreated hepatocellular carcinoma cells; MOCK, hepatocellular
carcinoma cells treated with Lipofectamine® 2000 alone;
NC, hepatocellular carcinoma cells treated with negative control
siRNA; reverse transcription-quantitative polymerase chain
reaction. |
HIF-1α is associated with the
COX-2/PKM2-mediated regulation of cell apoptosis
To investigate whether HIF-1α is associated with
modulation of the COX-2/PKM2 pathway, HIF-1α expression in HCC
tissues was measured using IHC. HIF-1α was mainly expressed in the
cytoplasm of tumor cells and was upregulated in 46.2% (66/143) of
HCC specimens compared with adjacent liver tissues (Fig. 5A). The associations between
HIF-1α, COX-2 and PKM2 were assessed and the results revealed that
54.3% of COX-2-positive tissues and 60.5% of PKM2-positive tissues
expressed HIF-1α, whereas only 35.5% and 29.9% of COX-2 and
PKM2-negative HCC tissues, respectively, expressed HIF-1α
(P<0.001; Table IV).
Furthermore, HIF-1α protein expression was positively correlated
with COX-2 (r=0.187, P=0.025) and PKM2 (r=0.307, P<0.001)
expression, suggesting that HIF-1α may be involved in the
modulation of the COX-2/PKM2 pathway.
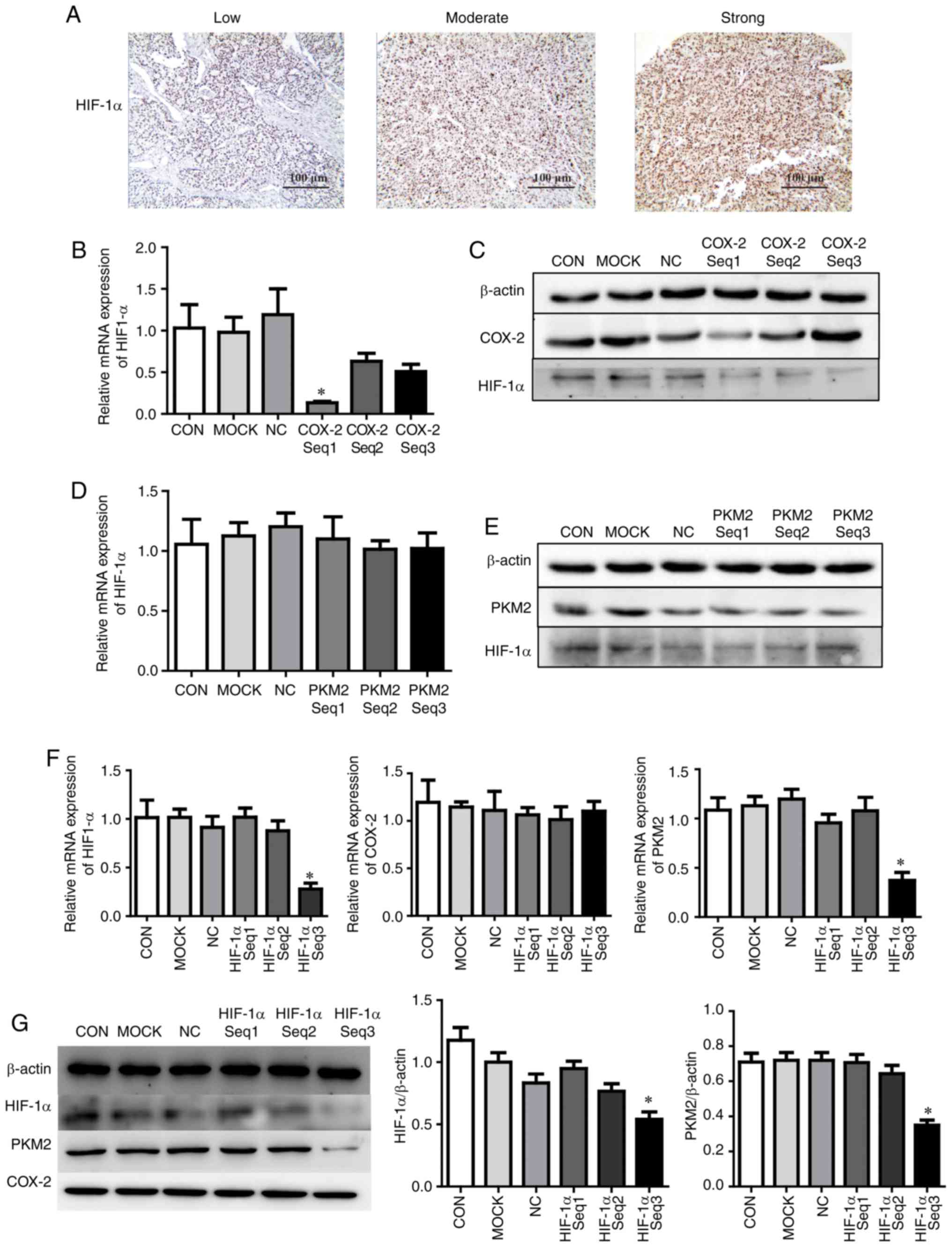 | Figure 5HIF-1α expression is associated with
COX-2/PKM2-mediated regulation of cell apoptosis. (A)
Immunohistochemical analysis of HIF-1α expression in
paraffin-embedded hepatocellular carcinoma tissue samples (scale
bar=100 µm). HepG2 cells were transfected with COX-2 siRNA
for 24 h, and the expression of HIF-1α (B) mRNA and (C) protein was
measured using RT-qPCR and western blotting, respectively. HepG2
cells were transfected with PKM2 siRNA for 24 h and the expression
of HIF-1α (D) mRNA and (E) protein was measured using RT-qPCR and
western blotting, respectively. HepG2 cells were transfected with
HIF-1α siRNA for 24 h and the expression of HIF-1α (F) mRNA and (G)
protein was measured using RT-qPCR and western blotting,
respectively. *P<0.05 vs. CON. HIF-1α,
hypoxia-inducible factor 1-α; COX-2, cyclooxygenase-2; PKM2,
pyruvate kinase M2 isoform; siRNA, small interfering RNA; CON,
untreated hepatocellular carcinoma cells; MOCK, hepatocellular
carcinoma cells treated with Lipofectamine® 2000 alone;
NC, hepatocellular carcinoma cells treated with negative control
siRNA; reverse transcription-quantitative polymerase chain
reaction. |
 | Table IVCorrelation between HIF-1α and
COX-2/PKM2. |
Table IV
Correlation between HIF-1α and
COX-2/PKM2.
| HIF-1α | COX-2
| r | P-value | PKM2
| r | P-value |
|---|
| − | + | − | + |
|---|
| Negative | 40 | 37 | 0.187 | 0.025 | 47 | 30 | 0.307 | <0.001 |
| Positive | 22 | 44 | | | 20 | 46 | | |
HepG2 cells were transfected with COX-2 siRNA and
RT-qPCR analysis (P<0.05; Fig.
5B) and western blotting (Fig.
5C) demonstrated that HIF-1α was significantly reduced compared
with untransfected cells. HepG2 cells were next transfected with
PKM2 siRNA and RT-qPCR (P>0.05; Fig. 5D) and western blotting (Fig. 5E) revealed that PKM2 knockdown had
no effect on HIF-1α expression. Finally, HepG2 cells were
transfected with HIF-1α siRNA and RT-qPCR (Fig. 5F) and western blotting (P<0.05;
Fig. 5G) revealed that HIF-1α
knockdown did not influence COX-2 expression compared with
untransfected cells, whereas PKM2 expression was significantly
reduced.
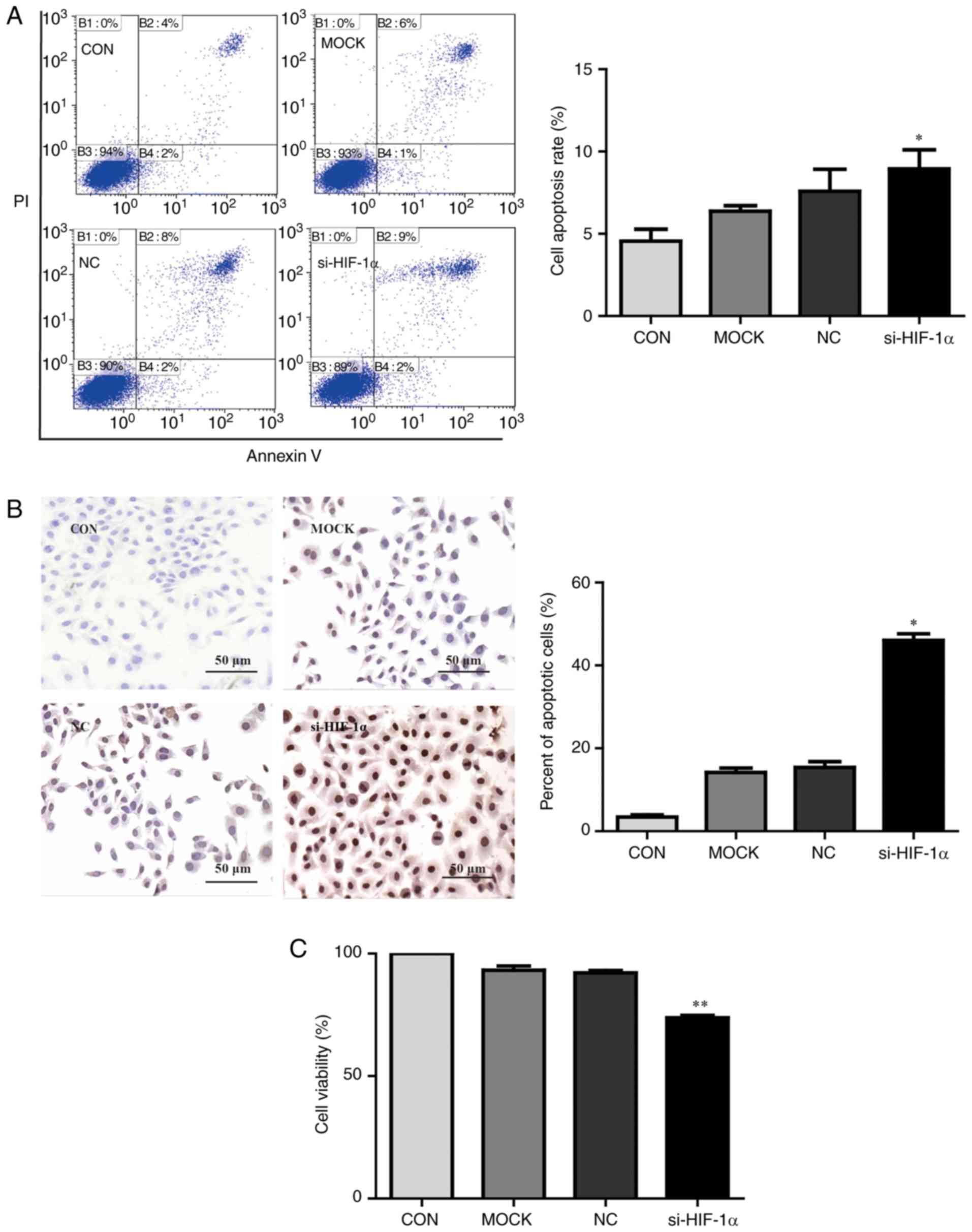
The impact of HIF-1α on apoptosis in HepG2 cells was
assessed using an Annexin V/PI double-staining assay and TUNEL
assay. As presented in Fig. 6A,
HIF-1α knockdown significantly increased apoptosis compared with
untransfected cells (P<0.05). TUNEL assays further confirmed
that HIF-1α knockdown in HepG2 cells significantly increased
apoptosis compared with untransfected control cells (P<0.05;
Fig. 6B). Furthermore, HepG2
cells transfected with HIF-1α siRNA significantly reduced cell
viability compared with untransfected cells (P<0.01; Fig. 6C). Altogether, these results
suggest that COX-2 is able to regulate the expression of PKM2 in
HCC cells via modulating HIF-1α.
Discussion
Standard metabolic processes are adapted in cancer
cells compared with normal cells in order to maintain their rapid
proliferation and progression, and PKM2 serves an important role in
cancer cell metabolism (18,19). It has previously been reported
that COX-2 is overexpressed in a number of tumor types, including
HCC (26). However, whether COX-2
overexpression affects tumor cells metabolism via regulating PKM2
is currently unclear. In the present study, it was demonstrated
that COX-2 and PKM2 were significantly upregulated in HCC tissues
compared with normal liver tissues (P<0.05), and that their
expression was positively associated with worse
clinico-pathological characteristics. Furthermore, it was
demonstrated that COX-2 knockdown significantly reduced the
expression of PKM2 and HIF-1α in HepG2 cells, whereas apoptosis was
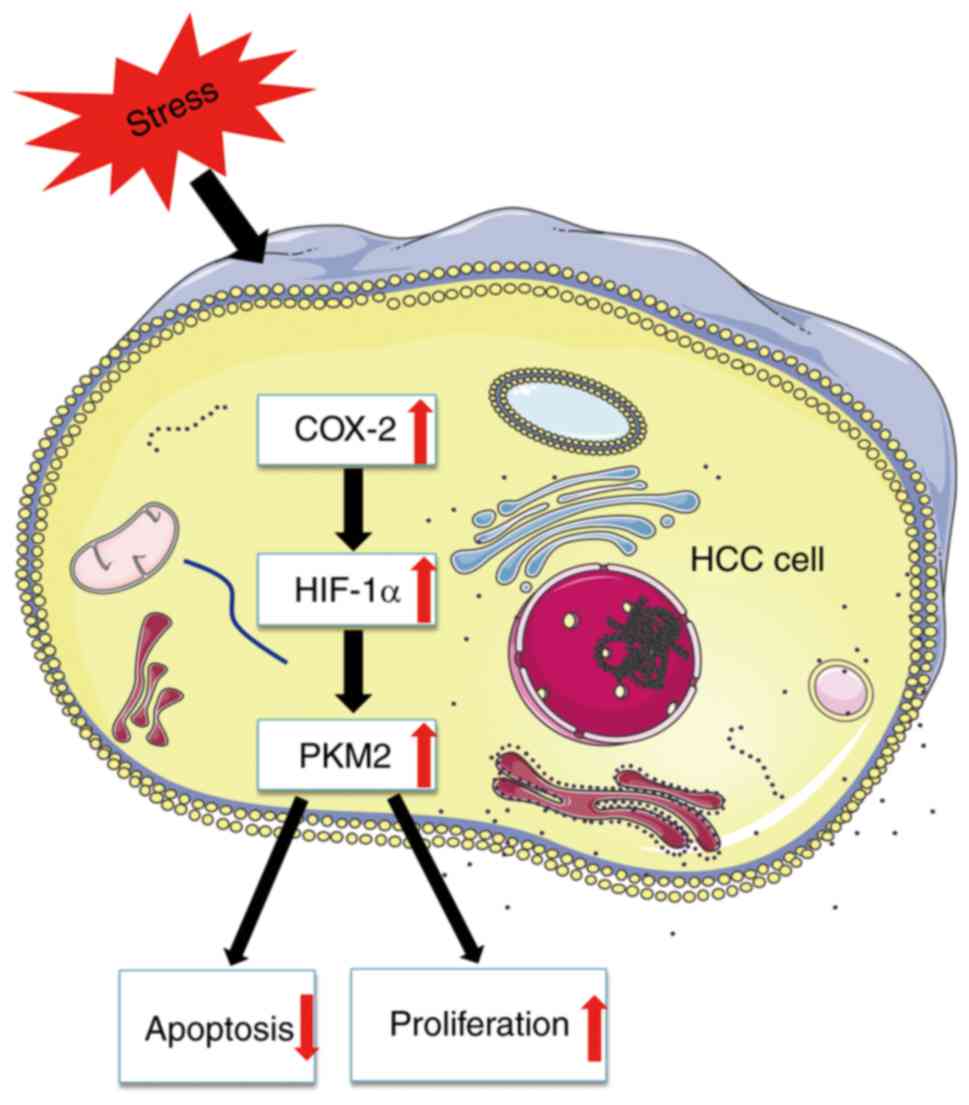
increased. The results of the present study suggest a novel method
of metabolism regulation via the COX-2-mediated activation of the
HIF-1α/PKM2 pathway in HCC (Fig.
7).
COX-2 is an inducible enzyme that serves an
important function in multiple pathophysiological processes,
including inflammation, atherosclerosis, tissue injury,
angiogenesis and tumorigenesis (14,27). COX-2 is chronically overexpressed
in numerous premalignant, malignant and metastatic human cancer
types (20,21). One previous study reported that
COX-2 regulates multiple cellular processes, including survival,
proliferation and apoptosis resistance in cancer (28). In the present study, COX-2
overexpression was observed in liver tumor tissues and was
positively associated with poorly differentiated histological
grades. COX-2 knockdown significantly reduced cell proliferation
and increased HepG2 cell apoptosis (P<0.05). Furthermore, it was
revealed that COX-2 expression was positively associated with PKM2
expression, while PKM2 was downregulated and apoptosis was
increased in HepG2 cells transfected with COX-2 siRNA. These
results suggest that PKM2 may serve a function in COX-2 induced
apoptosis resistance in HCC.
The primary function of the PK enzyme is to regulate
the final rate-limiting step of glycolysis (29), which catalyzes the transfer of a
phosphate group from PEP to ADP, yielding one molecule of pyruvate
and one of ATP (30,31). Due to its overexpression in tumor
tissues, PKM2 has been widely studied and has been reported to
serve an essential role in tumor progression (32,33). A number of studies have indicated
that PKM2 regulates apoptosis and proliferation (34,35); however, the precise molecular
mechanisms responsible remain elusive (36). In the present study, PKM2
expression was detected in paraffin-embedded HCC specimens using
IHC staining. It was revealed that PKM2 is upregulated in HCC
tissues compared with normal hepatic tissues and is associated with
poor differentiation. PKM2 function was also investigated using
in vitro knockdown assays. The results demonstrated that
PKM2 knockdown resulted in impaired cell viability and augmented
tumor cell apoptosis in vitro. These data suggest that
targeting PKM2 proteins may have potential as a therapeutic
strategy for the treatment of HCC.
PKM2 expression varies among different types of
cancer, suggesting that the function of PKM2 in tumorigenesis
depends on the signaling context (37). Therefore, a mechanistic insight
into the expression of PKM2 and its molecular pathways in HCC may
provide important data for developing novel HCC therapies (38,39). HIF-1 is a transcriptional factor
that comprises two subunits: HIF-1α and HIF-1β (40). HIF-1α mediates systemic hypoxia by
activating the transcription of multiple genes including
erythropoietin, glycolytic enzymes and vascular endothelial growth
factor (41,42). HIF-1α is known to regulate aerobic
glycolysis in a number of cancer types and promotes tumor growth by
regulating PKM2 expression (21,43). To determine whether PKM2
expression is modulated by HIF-1α in HCC, HIF-1α was silenced in
HepG2 cells via siRNA transfection. The results revealed that
si-HIF-1α effectively blocked the expression of PKM2 in HepG2
cells, inhibiting cell growth and promoting apoptosis. Similarly, a
significant reduction in HIF-1α expression was observed following
COX-2 knockdown (P<0.05). These data clearly demonstrate that
the HIF-1α/PKM2 pathway may contribute to COX-2-induced apoptosis
resistance in HCC.
In conclusion, the present study demonstrates that
COX-2 knockdown inhibits HepG2 cell viability and induces apoptosis
in vitro, and this may be mediated by the HIF-1α/PKM2
signaling pathway. These results suggest that the COX-2/HIF-1α/PKM2
pathway may facilitate apoptosis resistance in HCC cells,
suggesting that COX-2-induced apoptosis resistance in HCC may
result from metabolic changes associated with the HIF-1α/PKM2
pathway.
Acknowledgments
The authors would like to thank Professor Wei Wei
and Mrs. Weiqin Zhang for their technical assistance.
Funding
The present study was supported by the National
Natural Science Foundation of China (grant nos. 81572430 and
81402040).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors’ contributions
GS and JL designed the experiments and wrote the
paper. QW and DL performed the biological experiments with
assistance from LF, YuhL and YuL. HY and QW contributed to the
immunohistochemical analysis. GS and HW draft the manuscript. GS,
HW and JL analyzed data and edited the paper. All authors read and
approved the final manuscript.
Ethics approval and consent to
participate
The present study was approved by the Ethics
Committee of the First Affiliated Hospital of Anhui Medical
University and conformed to the 1975 Declaration of Helsinki. All
patients provided written informed consent.
Patient consent for publication
All patients consented to the publication of their
data.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Borzio M, Dionigi E, Parisi G, Raguzzi I
and Sacco R: Management of hepatocellular carcinoma in the elderly.
World J Hepatol. 7:1521–1529. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Mazzoccoli G, Tarquini R, Valoriani A,
Oben J, Vinciguerra M and Marra F: Management strategies for
hepatocellular carcinoma: Old certainties and new realities. Clin
Exp Med. 16:243–256. 2016. View Article : Google Scholar
|
|
3
|
Hoffmann EK and Lambert IH: Ion channels
and transporters in the development of drug resistance in cancer
cells. Philos Trans R Soc Lond B Biol Sci. 369:201301092014.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Ciccoli R, Sahi S, Singh S, Prakash H,
Zafiriou MP, Ishdorj G, Kock JL and Nigam S: Oxygenation by COX-2
(cyclo-oxygenase-2) of 3-HETE (3-hydroxyeicosatetraenoic acid), a
fungal mimetic of arachidonic acid, produces a cascade of novel
bioactive 3-hydroxyeicosanoids. Biochem J. 390:737–747. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Chandrashekar N, Selvamani A, Subramanian
R, Pandi A and Thiruvengadam D: Baicalein inhibits pulmonary
carcinogenesis-associated inflammation and interferes with COX-2,
MMP-2 and MMP-9 expressions in-vivo. Toxicol Appl Pharmacol.
261:10–21. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Esbona K, Inman D, Saha S, Jeffery J,
Schedin P, Wilke L and Keely P: COX-2 modulates mammary tumor
progression in response to collagen density. Breast Cancer Res.
18:352016. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Chen G, Li X, Yang J, Li J, Wang X, He J
and Huang Z: Prognostic significance of cyclooxygenase-2 expression
in patients with hepatocellular carcinoma: A meta-analysis. Arch
Med Sci. 12:1110–1117. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Zhao QT, Yue SQ, Cui Z, Wang Q, Cui X,
Zhai HH, Zhang LH and Dou KF: Potential involvement of the
cyclooxygenase-2 pathway in hepatocellular carcinoma-associated
angiogenesis. Life Sci. 80:484–492. 2007. View Article : Google Scholar
|
|
9
|
Leng J, Han C, Demetris AJ, Michalopoulos
GK and Wu T: Cyclooxygenase-2 promotes hepatocellular carcinoma
cell growth through Akt activation: Evidence for Akt inhibition in
celecoxib-induced apoptosis. Hepatology. 38:756–768. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Galant LW, de Mattos AA, Menti E, Valiatti
FB, de Mattos AZ, Porawski M, Hartmann A and Rhoden CR: The effect
of celecoxib on the development of diethylnitrosamine-induced liver
tumors in rats. Ann Hepatol. 12:425–433. 2013.PubMed/NCBI
|
|
11
|
Sui W, Zhang Y, Wang Z, Wang Z, Jia Q, Wu
L and Zhang W: Antitumor effect of a selective COX-2 inhibitor,
celecoxib, may be attributed to angiogenesis inhibition through
modulating the PTEN/PI3K/Akt/HIF-1 pathway in an H22
murine hepatocarcinoma model. Oncol Rep. 31:2252–2260. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Choe J, Yoon Y, Kim J and Jung YJ:
Positive feedback effect of PGE2 on cyclooxygenase-2 expression is
mediated by inhibition of Akt phosphorylation in human follicular
dendritic cell-like cells. Mol Immunol. 87:60–66. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Al-Harbi NO, Imam F, Al-Harbi MM, Ansari
MA, Zoheir KM, Korashy HM, Sayed-Ahmed MM, Attia SM, Shabanah OA
and Ahmad SF: Dexamethasone attenuates LPS-induced acute lung
injury through inhibition of NF-κB, COX-2, and pro-inflammatory
mediators. Immunol Invest. 45:349–369. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Merk H, Zhang S, Lehr T, Muller C, Ulrich
M, Bibb JA, Adams RH, Bracher F, Zahler S, Vollmar AM and Liebl J:
Inhibition of endothelial Cdk5 reduces tumor growth by promoting
non-productive angiogenesis. Oncotarget. 7:6088–6104. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Wang ZS, Chen LZ, Zhou HP, Liu XH and Chen
FH: Diarylpentadienone derivatives (curcumin analogues): Synthesis
and anti-inflammatory activity. Bioorg Med Chem Lett. 27:1803–1807.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Hacker HJ, Steinberg P and Bannasch P:
Pyruvate kinase isoenzyme shift from L-type to M2-type is a late
event in hepatocarcinogenesis induced in rats by a
choline-deficient/DL-ethionine-supplemented diet. Carcinogenesis.
19:99–107. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Christofk HR, Vander Heiden MG, Wu N,
Asara JM and Cantley LC: Pyruvate kinase M2 is a
phosphotyrosine-binding protein. Nature. 452:181–186. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Jung Y, Jang YJ, Kang MH, Park YS, Oh SJ,
Lee DC, Xie Z, Yoo HS, Park KC and Yeom YI: Metabolic signature
genes associated with susceptibility to pyruvate kinase, muscle
type 2 gene ablation in cancer cells. Mol Cells. 35:335–341. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Dong T, Yan Y, Chai H, Chen S, Xiong X,
Sun D, Yu Y, Deng L and Cheng F: Pyruvate kinase M2 affects liver
cancer cell behavior through up-regulation of HIF-1α and Bcl-xL in
culture. Biomed Pharmacother. 69:277–284. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Han D, Wei W, Chen X, Zhang Y, Wang Y,
Zhang J, Wang X, Yu T, Hu Q, Liu N and You Y: NF-κB/RelA-PKM2
mediates inhibition of glycolysis by fenofibrate in glioblastoma
cells. Oncotarget. 6:26119–26128. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Huang M, Wang L, Chen J, Bai M, Zhou C,
Liu S and Lin Q: Regulation of COX-2 expression and
epithelial-to-mesenchymal transition by hypoxia-inducible
factor-1alpha is associated with poor prognosis in hepatocellular
carcinoma patients post TACE surgery. Int J Oncol. 48:2144–2154.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zhou L, Rui JA, Zhou WX, Wang SB, Chen SG
and Qu Q: Edmondson-Steiner grade: A crucial predictor of
recurrence and survival in hepatocellular carcinoma without
microvascular invasio. Pathol Res Pract. 213:824–830. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Calderaro J, Couchy G, Imbeaud S, Amaddeo
G, Letouze E, Blanc JF, Laurent C, Hajji Y, Azoulay D, Bioulac-Sage
P, et al: Histological subtypes of hepatocellular carcinoma are
related to gene mutations and molecular tumour classification. J
Hepatol. 67:727–738. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Faria SC, Szklaruk J, Kaseb AO, Hassabo HM
and Elsayes KM: TNM/Okuda/Barcelona/UNOS/CLIP international
multidisciplinary classification of hepatocellular carcinoma:
Concepts, perspectives, and radiologic implications. Abdom Imaging.
39:1070–1087. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
26
|
Yang HJ, Jiang JH, Yang YT, Yang XD, Guo
Z, Qi YP, Zeng FH, Zhang KL, Chen NZ, Xiang BD and Li LQ:
Cyclooxygenase-2 expression is associated with initiation of
hepatocellular carcinoma, while prostaglandin receptor-1 expression
predicts survival. World J Gastroenterol. 22:8798–8805. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Kim KN, Ko YJ, Yang HM, Ham YM, Roh SW,
Jeon YJ, Ahn G, Kang MC, Yoon WJ, Kim D and Oda T:
Anti-inflammatory effect of essential oil and its constituents from
fingered citron (Citrus medica L. var sarcodactylis) through
blocking JNK, ERK and NF-κB signaling pathways in LPS-activated RAW
2647 cells. Food Chem Toxicol. 57:126–131. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Christofk HR, Vander Heiden MG, Harris MH,
Ramanathan A, Gerszten RE, Wei R, Fleming MD, Schreiber SL and
Cantley LC: The M2 splice isoform of pyruvate kinase is important
for cancer metabolism and tumour growth. Nature. 452:230–233. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Lou Y, Wang C, Zheng W, Tang Q, Chen Y,
Zhang X, Guo X and Wang J: Salvianolic acid B inhibits
IL-1β-induced inflammatory cytokine production in human
osteoarthritis chondrocytes and has a protective effect in a mouse
osteoarthritis model. Int Immunopharmacol. 46:31–37. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Bu LJ, Yu HQ, Fan LL, Li XQ, Wang F, Liu
JT, Zhong F, Zhang CJ, Wei W, Wang H and Sun GP: Melatonin, a novel
selective ATF-6 inhibitor, induces human hepatoma cell apoptosis
through COX-2 downregulation. World J Gastroenterol. 23:986–998.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Zhang JX, Xing JG, Wang LL, Jiang HL, Guo
SL and Liu R: Luteolin Inhibits Fibrillary β-Amyloid1-40-induced
inflammation in a human blood-brain barrier model by suppressing
the p38 MAPK-Mediated NF-κB signaling pathways. Molecules.
22:2017.
|
|
32
|
Ghosh S and Hayden MS: New regulators of
NF-κB in inflammation. Nat Rev Immunol. 8:837–848. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Li Z, Yang P and Li Z: The multifaceted
regulation and functions of PKM2 in tumor progression. Biochim
Biophys Acta. 1846:285–296. 2014.PubMed/NCBI
|
|
34
|
Hamaguchi Y, Mori A, Fujimoto Y, Ito T,
Iida T, Yagi S, Okajima H, Kaido T and Uemoto S: Longer warm
ischemia can accelerate tumor growth through the induction of
HIF-1α and the IL-6-JAK-STAT3 signaling pathway in a rat
hepatocellular carcinoma model. J Hepatobiliary Pancreat Sci.
23:771–779. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Yang CM, Chen YW, Chi PL, Lin CC and Hsiao
LD: Resveratrol inhibits BK-induced COX-2 transcription by
suppressing acetylation of AP-1 and NF-κB in human rheumatoid
arthritis synovial fibroblasts. Biochem Pharmacol. 132:77–91. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Radunz S, Treckmann J, Baba HA, Best J,
Muller S, Theysohn JM, Paul A and Benko T: Long-term outcome after
liver transplantation for hepatocellular carcinoma following
Yttrium-90 radioembolization bridging treatment. Ann Transplant.
22:215–221. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Yang W, Xia Y, Cao Y, Zheng Y, Bu W, Zhang
L, You MJ, Koh MY, Cote G, Aldape K, et al: EGFR-induced and
PKCepsilon monoubiquitylation-dependent NF-κB activation
upregulates PKM2 expression and promotes tumorigenesis. Mol Cell.
48:771–784. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Xie W, Wang H, He Y, Li D, Gong L and
Zhang Y: CDK5 and its activator P35 in normal pituitary and in
pituitary adenomas: relationship to VEGF expression. Int J Biol
Sci. 10:192–199. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Magierowski M, Magierowska K,
Hubalewska-Mazgaj M, Sliwowski Z, Pajdo R, Ginter G, Kwiecien S and
Brzozowski T: Exogenous and endogenous hydrogen sulfide protects
gastric mucosa against the formation and time-dependent development
of ischemia/reperfusion-induced acute lesions progressing into
deeper ulcerations. Molecules. 22:pii: E2952017. View Article : Google Scholar
|
|
40
|
Wos J, Brys M, Lewy-Trenda I, Stasikowska
O, Papiez P, Papierz W and Starska K: Analysis of HIF-1α and COX-2
expression in tumor stroma and correlation with the degree of
neoplasm invasiveness in laryngeal cancer-preliminary study.
Otolaryngol Pol. 65:102–108. 2011.In Polish.
|
|
41
|
Tao T, Li G, Dong Q, Liu D, Liu C, Han D,
Huang Y, Chen S, Xu B and Chen M: Loss of SNAIL inhibits cellular
growth and metabolism through the miR-128-mediated
RPS6KB1/HIF-1α/PKM2 signaling pathway in prostate cancer cells.
Tumour Biol. 35:8543–8550. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Salvolini E, Buldreghini E, Lucarini G,
Vignini A, Giulietti A, Lenzi A, Mazzanti L, Di Primio R and
Balercia G: Interleukin-1β, cyclooxygenase-2, and hypoxia-inducible
factor-1α in astheno-zoospermia. Histochem Cell Biol. 142:569–575.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Zhao CX, Luo CL and Wu XH: Hypoxia
promotes 786-O cells invasiveness and resistance to sorafenib via
HIF-2α/COX-2. Med Oncol. 32:4192015. View Article : Google Scholar
|