Introduction
Bladder cancer (BC) is one of the most common human
cancer types worldwide that results in a large number of
cancer-associated fatalities each year (1,2).
Rapid tumour growth and metastasis are the primary reasons for the
high mortality associated with patients with BC (2,3).
Therefore, exploring the regulatory mechanisms underlying the
proliferation and migration of BC cells may aid the development of
novel therapeutic strategies for this disease (2,3).
Long non-coding RNAs (lncRNAs) are a class of small
non-coding RNAs that consist of >200 nucleotides (4). By regulating downstream effectors,
including microRNAs (miRs) and proteins, lncRNAs serve important
roles in a variety of biological processes, such as cell
proliferation, survival, apoptosis, energy metabolism, migration,
invasion and tumourigenesis (5-7).
In the recent decade, a large number of lncRNAs have been reported
to function as oncogenes or tumour suppressors in various human
cancer types, including BC (8-10).
LncRNA metastasis associated lung adenocarcinoma transcript 1
(MALAT1) is frequently deregulated in some common cancer types and
has been revealed to serve as a tumour promotor or suppressor
(11-13). For instance, MALAT1 is upregulated
in osteosarcoma, and the knockdown of MALAT1 inhibits osteosarcoma
cell proliferation and migration and induces cell apoptosis
(11). In addition, MALAT1 has
been reported to promote renal cell carcinoma cell proliferation
and invasion, as well as inhibit cell apoptosis by modulating the
miR-205/enhancer of zeste homolog 2 (Ezh2) axis (14). Furthermore, a previous study
reported that MALAT1 promotes BC cell migration through the
induction of epithelial-to-mesenchymal transition (EMT) (15). In addition, Li et al
(16) identified that the high
expression of MALAT1 was associated with a poor prognosis and
promoted clinical progression and metastasis in BC. Xie et
al (17) demonstrated that
MALAT1 promoted BC cell invasion and inhibited cell apoptosis via
antagonizing miR-125b. However, it remains unclear whether other
miRs and downstream proteins are also involved in MALAT1-mediated
BC.
miRs are a type of non-coding small RNA that are
22-25 nucleotides in length. These RNAs directly bind to
complementary sites in the 3′-untranslated region (UTR) region of
target mRNAs and thus cause translation inhibition (18,19). By affecting the expression of
their target genes, miRs are also involved in various biological
processes (20,21). It is well-established that miR-34a
generally functions as a tumour suppressor in human cancer
(22). For instance, miR-34a
inhibits the proliferation and promotes the apoptosis of non-small
cell lung cancer (NSCLC) cells by targeting transforming growth
factor β receptor 2 (TGF-βR2) (22). In addition, miR-34a inhibits
osteosarcoma cell proliferation by the inhibition of ether à go-go
1 expression (23). In BC, Yu
et al (24) demonstrated
that miR-34a inhibited metastasis and angiogenesis by directly
targeting cluster of differentiation 44. Sun et al (25) reported that miR-34a inhibited BC
cell proliferation and invasion by targeting the orphan nuclear
receptor hepatocyte nuclear factor 4γ. However, to the best of our
knowledge the association between MALAT1 and miR-34a in BC has not
been previously reported.
The present study aimed to investigate the clinical
significance of MALAT1 expression in BC. In addition, the
regulatory role of MALAT1 in BC growth and metastasis in
vitro and the underlying mechanism were studied.
Materials and methods
Clinical tissues
The present study was approved by the Ethics
Committee of First People’s Hospital of Jining City (Jining,
China). BC and matched adjacent non-tumor tissues were collected
from a total of 56 patients with primary BC at the First People’s
Hospital of Jining City between April 2011 and March 2013. Written
informed consent was collected from all patients. These patients
with BC included 34 male and 22 female, and were between 42 to 78
years old. All tissues were pathologically confirmed by the
Department of Pathology at our hospital. No patients received
chemotherapy and radiotherapy prior to surgical resection.
Following surgical resection, the tissues were immediately frozen
and stored at −80°C until use.
Cell culture and transfection
BC cell lines (HT-1376, RT112, 253J and T24) and the
normal urinary tract epithelial cell line SV-HUC-1 were purchased
from the Chinese Academy of Sciences Cell Bank (Shanghai, China).
Cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM,
Thermo Fisher Scientific, Inc., Waltham, MA, USA) with 10% fetal
bovine serum (Thermo Fisher Scientific, Inc.) at 37°C in an
atmosphere containing 5% CO2.
For cell transfection, T24 cells were transfected
with 100 nM of negative control (NC) small interfering (si) RNA
(siNC group; forward sequence, 5′-UUC UCC GAA CGU GUC ACG UTT-3′;
and reverse sequence, 5′-ACG UGA CAC GUU CGG AGA ATT-3′), 100 nM of
MALAT1 siRNA (siMALAT1 group; forward sequence, 5′-GCA AAU GAA AGC
UAC CAA U-3′; and reverse sequence, 5′-AUU GGU AGC UUU CAU UUG
CTT-3′), 1 μg of pcDNA3.1 vector (Amspring, Changsha,
China), 100 nM of miR-NC mimics (miR-NC group; cat. no. HMC0002,
Sigma-Aldrich; Merck KGaA, Darmstadt, Germany), 100 nM of miR-34a
mimics (miR-34a group; cat. no. HMI0509, Sigma-Aldrich; Merck
KGaA), or co-transfected with siMALAT1 and NC inhibitor
(siMALAT1+anti-NC group), siMALAT1 and miR-34a inhibitor
(siMALAT1+anti-miR-34a group), siMALAT1 and 1 μg of blank
vector (siMALAT1+blank group; Amspring), siMALAT1 and CCND1 plasmid
(siMALAT1+CCND1; Amspring), miR-NC and blank vector (miR-NC+blank
group), miR-34a mimics and blank vector (miR-34a+blank group), and
miR-34a mimics and CCND1 plasmid (miR-34a+CCND1 group) using
Lipofectamine 2000 (Thermo Fisher Scientific, Inc.). At 48 h
post-transfection, the cells were harvested and reverse
transcription-quantitative polymerase chain reaction (RT-qPCR) was
conducted to determine the transfection efficiency.
RT-qPCR
Total RNA was isolated from the tissues and cell
lines using TRIzol reagent (Thermo Fisher Scientific, Inc.)
according to the manufacturer’s protocol. The total RNA (1
μg) was reverse transcribed into cDNA using the PrimeScript
RT Reagent Kit (Takara Bio, Inc., Otsu, Japan) according to the
manufacturer’s protocol. The miR-specific TaqMan miR assay kit
(Applied Biosystems; Thermo Fisher Scientific, Inc.) was used to
determine miR expression using the ABI 7500 Sequence Detection
System (Thermo Fisher Scientific, Inc.). U6 was used as the
internal control. For lncRNA and mRNA expression detection, PCR was
performed using the SYBR-Green qPCR mix (Toyobo Life Science,
Osaka, Japan) according to the manufacturer’s protocol.
Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was used as the
internal control. The PCR reaction condition was as follows: 95°C
for 5 min followed by 40 cycles of 95°C for 30 sec and 60°C for 30
sec. The relative expression levels were calculated using the
2-ΔΔCq method (26).
The primer sequences used were as follows: U6 forward, 5′-CTC GCT
TCG GCA GCA CAT ATA CT-3′ and reverse, 5′-CGC TTC ACG AAT TTG CGT
GT-3′; MALAT1 forward, 5′-GGT AAC GAT GGT GTC GAG GTC-3′ and
reverse, 5′-CCA GCA TTA CAG TTC TTG AAC ATG-3′; CCND1 forward,
5′-GCT GCG AAG TGG AAA CCA TC-3′ and reverse, 5′-CCT CCT TCT GCA
CAC ATT TGA A-3′; and GAPDH forward, 5′-GGA GCG AGA TCC CTC CAA
AAT-3′ and reverse, 5′-GGC TGT TGT CAT ACT TCT CAT GG-3′. The
primers for miR-34a were purchased from Guangzhou Fulengen Co.,
Ltd. (cat. no. HmiRQP0439; Guangzhou, China) and the sequences were
not supplied by the manufacturer.
Western blot analysis
Tissues and cells were lysed in cold
radioimmunoprecipitation assay buffer (Thermo Fisher Scientific,
Inc.). Protein concentration was examined using a BCA protein assay
kit (Beyotime Institute of Biotechnology, Haimen, China). Proteins
(50 μg per lane) were loaded onto 10% SDS-polyacrylamide
gels and were blotted onto poly-vinylidene difluoride membranes
(Thermo Fisher Scientific, Inc.). Membranes were blocked with 5%
non-fat dry milk in Tris-buffered saline with Tween-20 at room
temperature for 2 h. Following this, membranes were incubated at
4°C overnight with rabbit monoclonal anti-CCND1 primary antibody
(dilution 1:50; cat. no. ab134175; Abcam, Cambridge, MA, USA) and
rabbit polyclonal anti-GAPDH primary antibody (dilution 1:100; cat.
no. ab9485; Abcam). Following 3 wash steps with phosphate buffered
saline with Tween-20, the membranes were incubated with horseradish
peroxidase-conjugated goat anti-rabbit secondary antibody (dilution
1:5,000; cat. no. ab6721, Abcam) at room temperature for 1 h.
Subsequently, the proteins were visualized using an enhanced
chemiluminescent kit (ECL-plus, Thermo Fisher Scientific, Inc.) and
analysed using Image-Pro Plus software 6.0 (Media Cybernetics,
Inc., Rockville, MD, USA).
Cell Counting Kit-8 (CCK-8) assay
A CCK-8 assay was conducted to study cell
proliferation. Following 48 h of trans-fection, the cells (5,000
cells per well) were seeded onto 96-well plates and cell
proliferation was measured at 0, 24, 48 and 72 h using CCK-8
(Thermo Fisher Scientific, Inc.) according to the manufacturer’s
instructions. The absorbance was detected at an optical density of
450 nm using a spectrophotometer.
Cell migration assay
Transfected cells (5×105 cells/well) were
cultured in 6-well plates and were grown to 100% confluence. Cells
were scratched with sterile 200 μl pipette tips, and the
plates were washed twice with Dulbecco’s phosphate-buffered saline
to remove the detached cells. Following this, the cells were
incubated at 37°C in serum-free DMEM. During the following 24 h,
the cells migrated into the wounded area. Photographs were taken at
0 and 24 h using a light microscope (CKX41; Olympus Corporation,
Tokyo, Japan).
Luciferase reporter gene assay
To clarify the target association between miR-34a
and MALAT1, the MALAT1 sequences, containing the wild-type (WT) or
mutated (MT) miR-34a binding sites, were subcloned into the
pmiR-GLo luciferase reporter vector (Promega Corporation, Madison,
WI, USA). T24 cells were co-transfected with WT or MT MALAT1
reporter plasmids, together with the miR-34a mimic or miR-NC, using
Lipofectamine 2000. To clarify the target association between CCND1
and miR-34a, the CCND1 3′-UTR sequences, containing the WT or MT
miR-34a binding sites, were also subcloned into the pmiR-GLo
luciferase reporter vector. T24 cells were co-transfected with the
WT or MT CCND1 reporter plasmids, together with the miR-34a mimic
or miR-NC, using Lipofectamine 2000. At 48 h post-transfection, the
luciferase activity was conducted using the Dual-Luciferase
Reporter Assay System (Promega Corporation), according to the
manufacturer’s instructions. The ratio of Firefly luciferase
activity to Renilla luciferase activity was determined.
Statistical analysis
Data were presented as the mean ± standard deviation
and were analysed using SPSS 19.0 software (IBM Corp., Armonk, NY,
USA). A two-tailed Student’s t-test was used to compare the
difference between two groups. Multiple comparisons were performed
using one-way analysis of variance followed by Tukey’s post hoc
test. Kaplan-Meier analysis with a log-rank test was performed to
assess patient survival. The correlation between MALAT1 and miR-34a
expression was analysed using Spearman’s rank correlation.
P<0.05 was considered to indicate a statistically significant
difference.
Results
Upregulation of MALAT1 in BC tissues and
cell lines
MALAT1 expression was first examined in 56 BC
tissues and the matching adjacent non-tumour tissues using RT-qPCR.
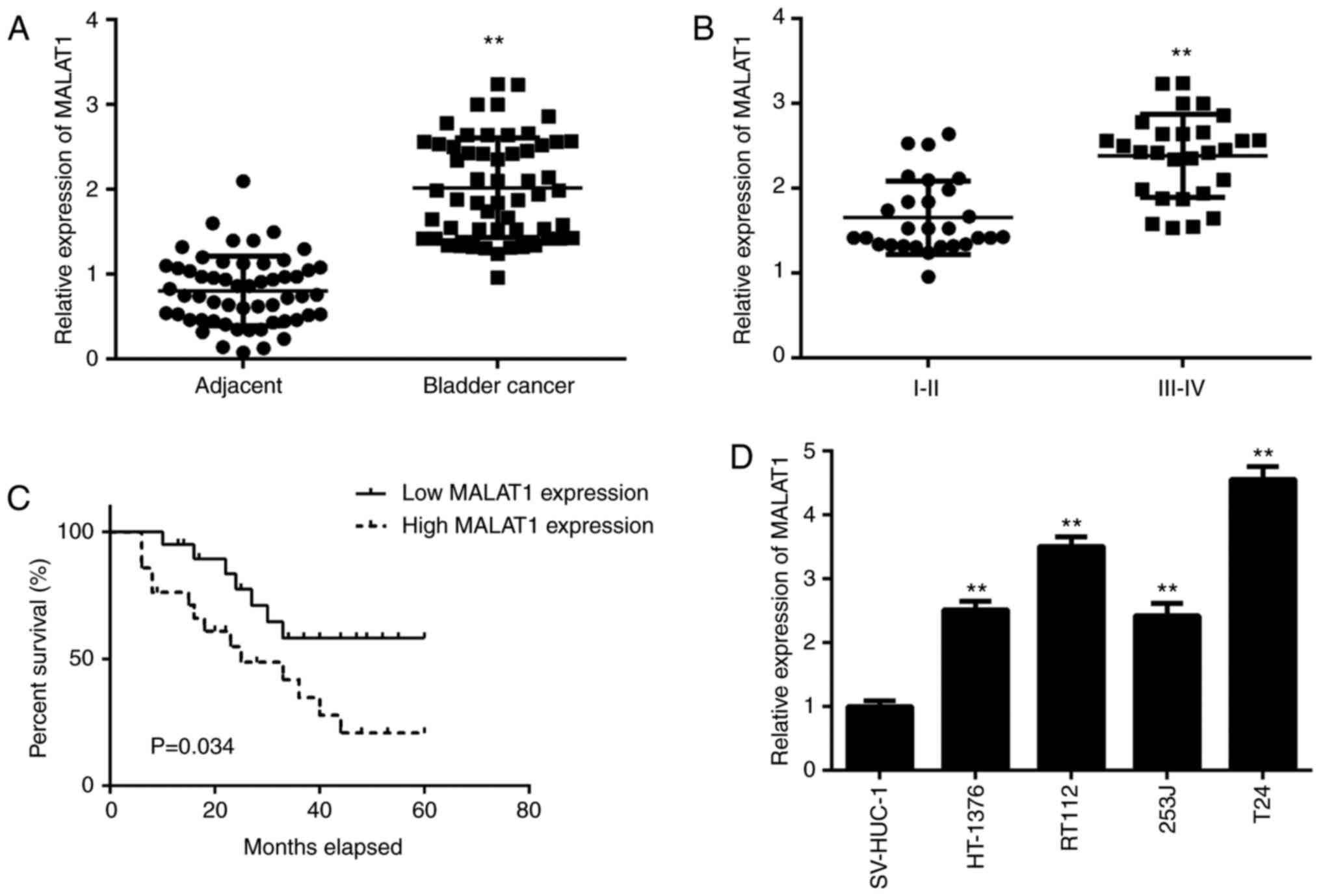
As indicated in Fig. 1A, the
expression levels of MALAT1 were significantly increased in the BC
tissues compared with the adjacent non-tumour tissues. Furthermore,
the expression levels of MALAT1 were significantly increased in the
BC tissues at stage III-IV compared with that in the tissues at
stage I-II (Fig. 1B). These
patients with BC were subsequently divided into high- and
low-MALAT1 expression groups based on the mean expression value in
the BC tissues. As indicated in Table
I, high expression of MALAT1 was significantly associated with
advanced Tumour, Node, Metastases (TNM) stage and positive lymph
node metastasis. Furthermore, patients with high MALAT1 expression
exhibited a shorter survival time when compared with those with a
low MALAT1 expression (Fig. 1C).
The expression of MALAT1 in the BC cell lines (HT-1376, RT112, 253J
and T24) and the normal urinary tract epithelial cell line SV-HUC-1
were also assessed. MALAT1 was also significantly upregulated in
the BC cell lines compared with the SV-HUC-1 cells (Fig. 1D).
 | Table IAssociation between MALAT1 expression
and clinicopathological characteristics in bladder cancer. |
Table I
Association between MALAT1 expression
and clinicopathological characteristics in bladder cancer.
|
Characteristics | Cases (n=56) | MALAT1 expression
| P-value |
|---|
| High (n=29) | Low (n=27) |
|---|
| Age | | | | 0.783 |
| <55 | 21 | 10 | 11 | |
| ≥55 | 35 | 19 | 16 | |
| Sex | | | | 0.790 |
| Male | 34 | 17 | 17 | |
| Female | 22 | 12 | 10 | |
| Lymph node
metastasis | | | | 0.013 |
| Negative | 36 | 14 | 22 | |
| Positive | 20 | 15 | 5 | |
| TNM stage | | | | 0.007 |
| I-II | 28 | 9 | 19 | |
| III-IV | 28 | 20 | 8 | |
Inhibition of MALAT1 reduces the
proliferation and migration of BC cells
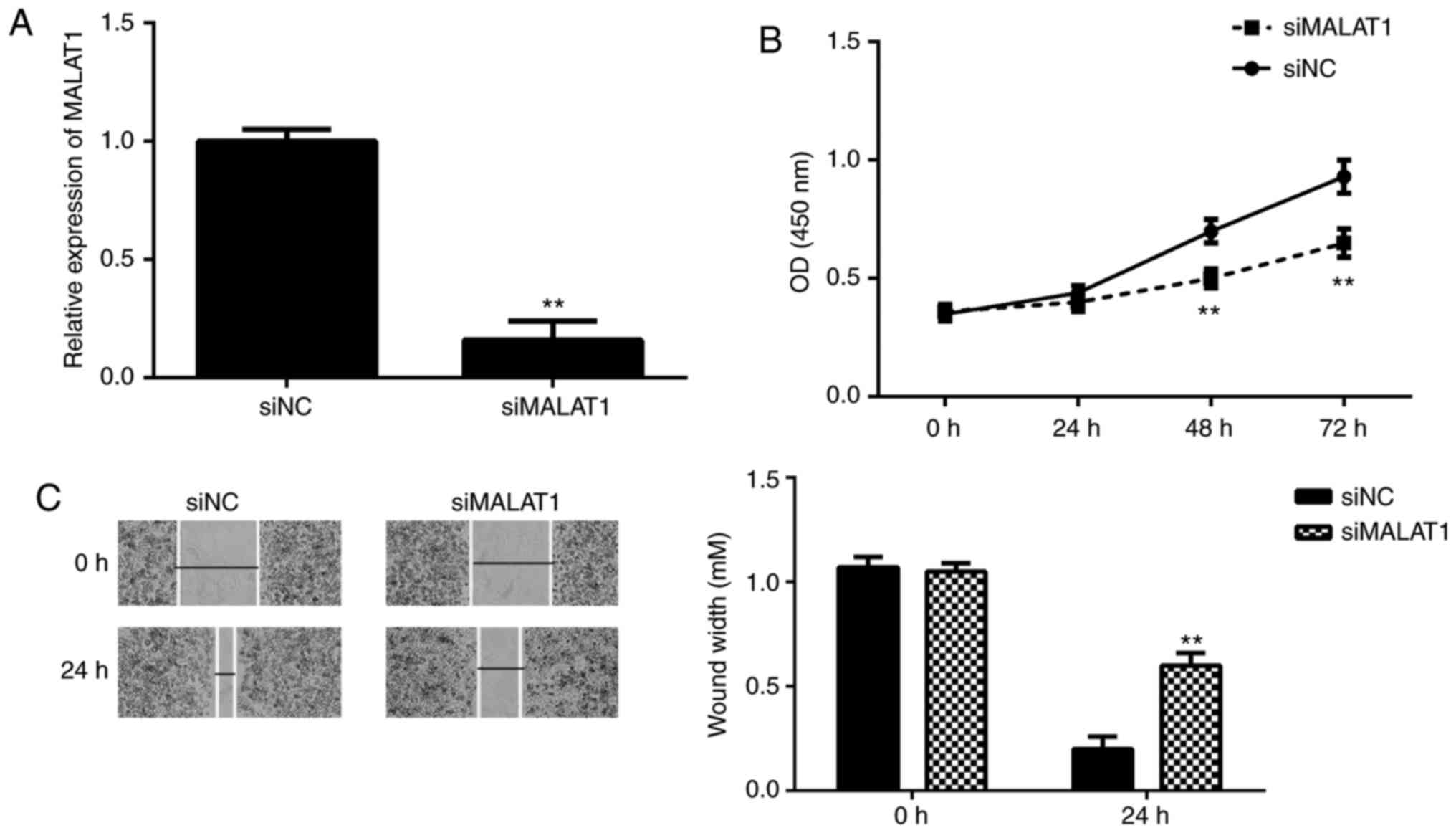
The function of MALAT1 in BC was studied using T24
cells. MALAT1-specific siRNA was transfected into T24 cells to
inhibit its expression. As indicated in Fig. 2A, MALAT1 was significantly
downregulated in the siMALAT1 group compared with the siNegative
Control (siNC) group. These data confirmed that the trans-fection
efficiency was successful. Data from CKK-8 and wound healing assays
further indicated that the knockdown of MALAT1 significantly
suppressed the proliferation and migration of T24 cells (Fig. 2B and C).
miR-34a is a target of MALAT1 in BC
cells
A bioinformatic analysis was conducted to predict
the potential MALAT1-miR interactions. As indicated in Fig. 3A, miR-34a had a potential binding
site in MALAT1. Consistent with this prediction, the inhibition of
MALAT1 expression significantly increased the expression of miR-34a
in T24 cells (Fig. 3B). To
further confirm this prediction, a luciferase reporter gene assay
was performed using the T24 cells, and the data revealed that
transfection with the miR-34a mimics significantly decreased the
luciferase activity of the MALAT1-WT luciferase reporter gene
plasmid, but had no significant effect on the luciferase activity
of the MALAT1-MT luciferase reporter gene plasmid in the T24 cells
(Fig. 3C). These findings
demonstrated that miR-34a and MALAT1 were associated. Following
this, it was indicated that the expression levels of miR-34a were
significantly reduced in the BC tissues and cell lines compared
with the adjacent non-tumour tissues and SV-HUC-1 cells,
respectively (Fig. 3D and E).
Furthermore, an inverse correlation between the expression of
miR-34a and MALAT1 in the BC tissues was observed (Fig. 3F).
miR-34a is involved in MALAT1-mediated BC
cell proliferation and migration
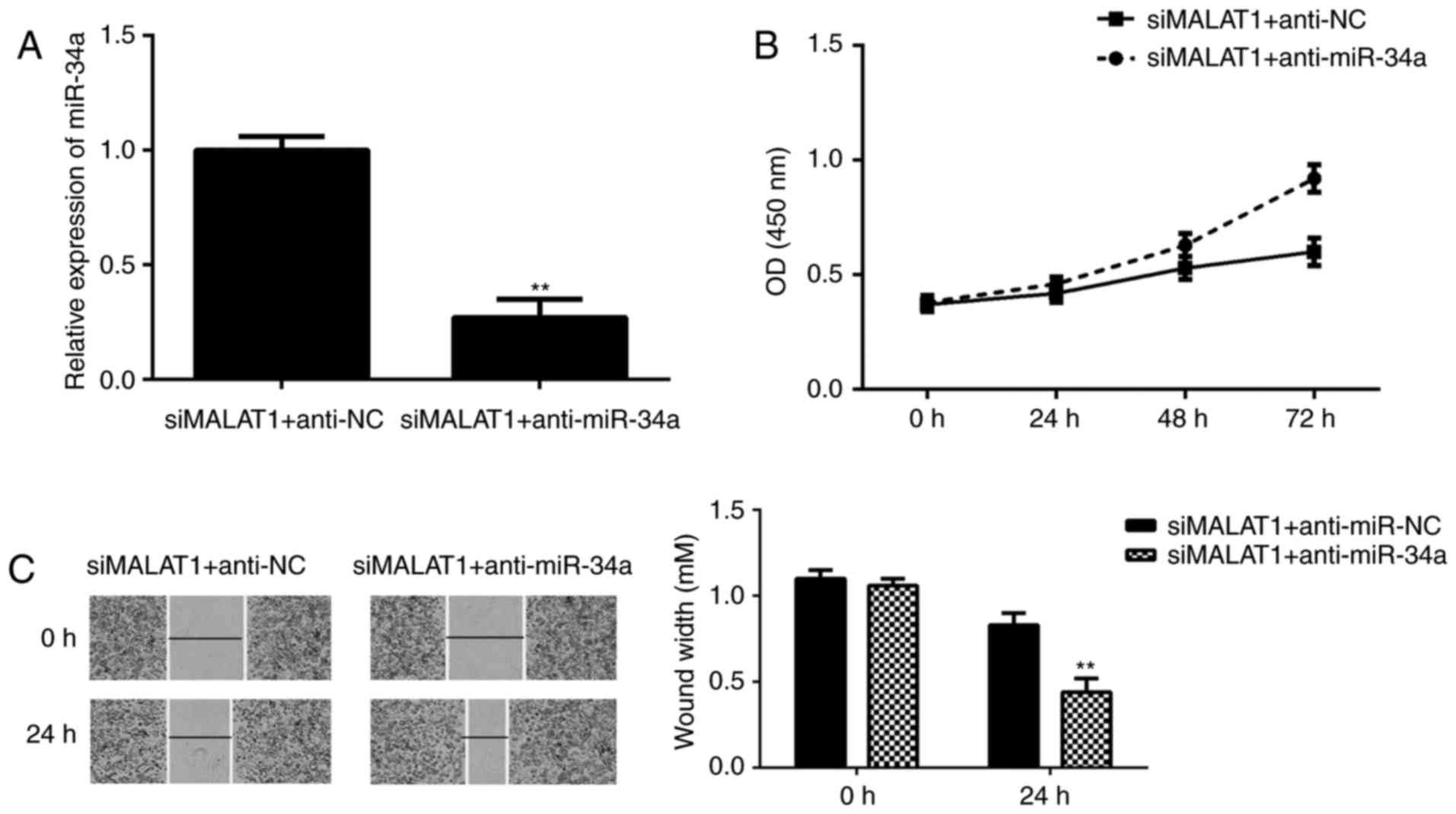
It was speculated that miR-34a may be involved in
MALAT1-mediated BC cell proliferation and migration. To clarify
this speculation, siMALAT1-transfected T24 cells were transfected
with miR-34a inhibitor or NC inhibitor. The qPCR data indicated
that miR-34a was significantly downregulated in the
siMALAT1+anti-miR-34a group compared with the siMALAT1+anti-NC
group (Fig. 4A). These data
confirmed that the transfection efficiency was successful. Further
study revealed that the proliferation and migration of T24 cells
were significantly increased in the siMALAT1+anti-miR-34a group
compared with the siMALAT1+anti-NC group (Fig. 4B and C), which suggested that
knockdown of miR-34a impaired the suppressive effects on the T24
cell proliferation and migration induced by MALAT1
downregulation.
CCND1 is a target gene of miR-34a in BC
cells
Bioinformatic analysis data demonstrated that CCND1
was a target gene of miR-34a (Fig.
5A). The luciferase reporter gene assay was conducted to
confirm this prediction, and the data indicated that transfection
with the miR-34a mimics significantly downregulated the luciferase
activity of the CCND1-WT luciferase reporter gene plasmid, but had
no significant effect on the luciferase activity of the CCND1-MT
luciferase reporter gene plasmid (Fig. 5B). These findings demonstrated
that miR-34a was a target gene of miR-34a. In addition, it was
revealed that the inhibition of MALAT1 expression caused a
reduction in CCND1 expression in T24 cells, which was abolished by
the knockdown of miR-34a (Fig. 5C and
D). These findings suggested that MALAT1 may positively
regulate the expression of CCND1 via directly targeting miR-34a in
T24 cells. Following this, it was investigated whether CCND1 was
involved in miR-34a-mediated BC growth. T24 cells were
co-transfected with miR-NC and blank vector (miR-NC+blank), miR-34a
mimics and blank vector (miR-34a+blank), and miR-34a mimics and
CCND1 plasmid (miR-34a+CCND1). CKK-8 assay data demonstrated that
the proliferation of T24 cells was significantly reduced in the
miR-34a+blank group when compared with the miR-NC+blank group;
however, proliferation was restored in the miR-34a+CCND1 group
(Fig. 5E). These findings
suggested that miR-34a inhibits BC cell proliferation, and
upregulation of CCND1 impaired the suppressive effects of miR-34a
on T24 cell proliferation.
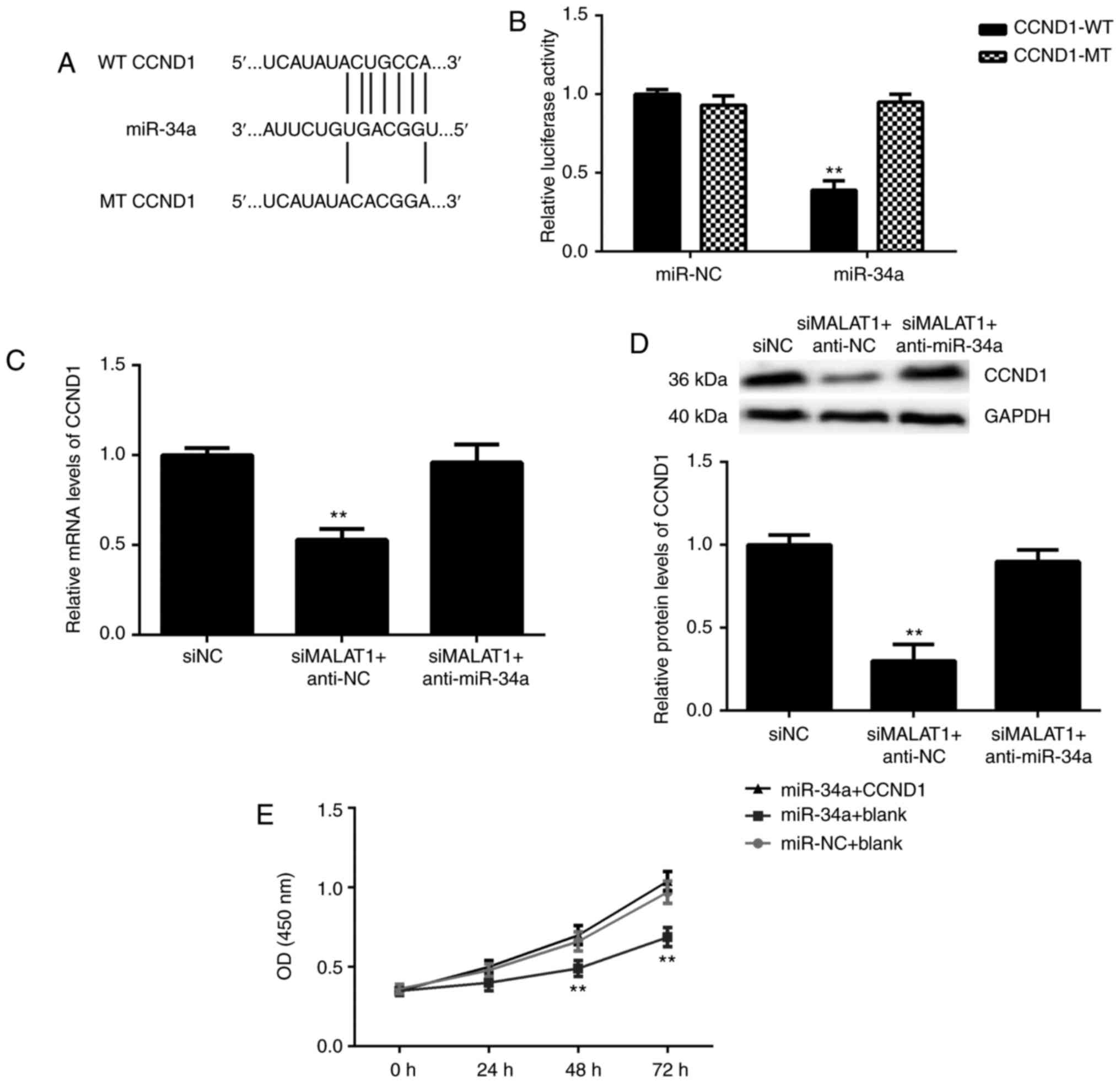 | Figure 5CCND1 is a target gene of miR-34a in
bladder cancer cells. (A) miR-34a had a potential binding site in
CCND1. (B) Transfection with the miR-34a mimics significantly
downregulated the luciferase activity of the CCND1-WT luciferase
reporter gene plasmid, but had no significant effect on the
luciferase activity of the CCND1-MT luciferase reporter gene
plasmid. **P<0.01 vs. miR-NC. (C and D) Knockdown of
MALAT1 caused a reduction in the mRNA and protein expression levels
of CCND1 in T24 cells, which was abolished by the knockdown of
miR-34a. **P<0.01 vs. siNC. (E) T24 cells were
co-transfected with miR-NC and blank vector (miR-NC+blank), miR-34a
mimics and blank vector (miR-34a+blank), and miR-34a mimics and
CCND1 plasmid (miR-34a+CCND1). The Cell counting Kit-8 assay was
then conducted to study cell proliferation. **P<0.01
vs. miR-NC+blank. MALAT1, metastasis associated lung adenocarcinoma
transcript 1; miR, microRNA; WT, wild-type; MUT, mutant; NC,
negative control; si, small interfering RNA; CCND1, cyclin D1. |
CCND1 acts as a downstream effector of
MALAT1 in BC cells
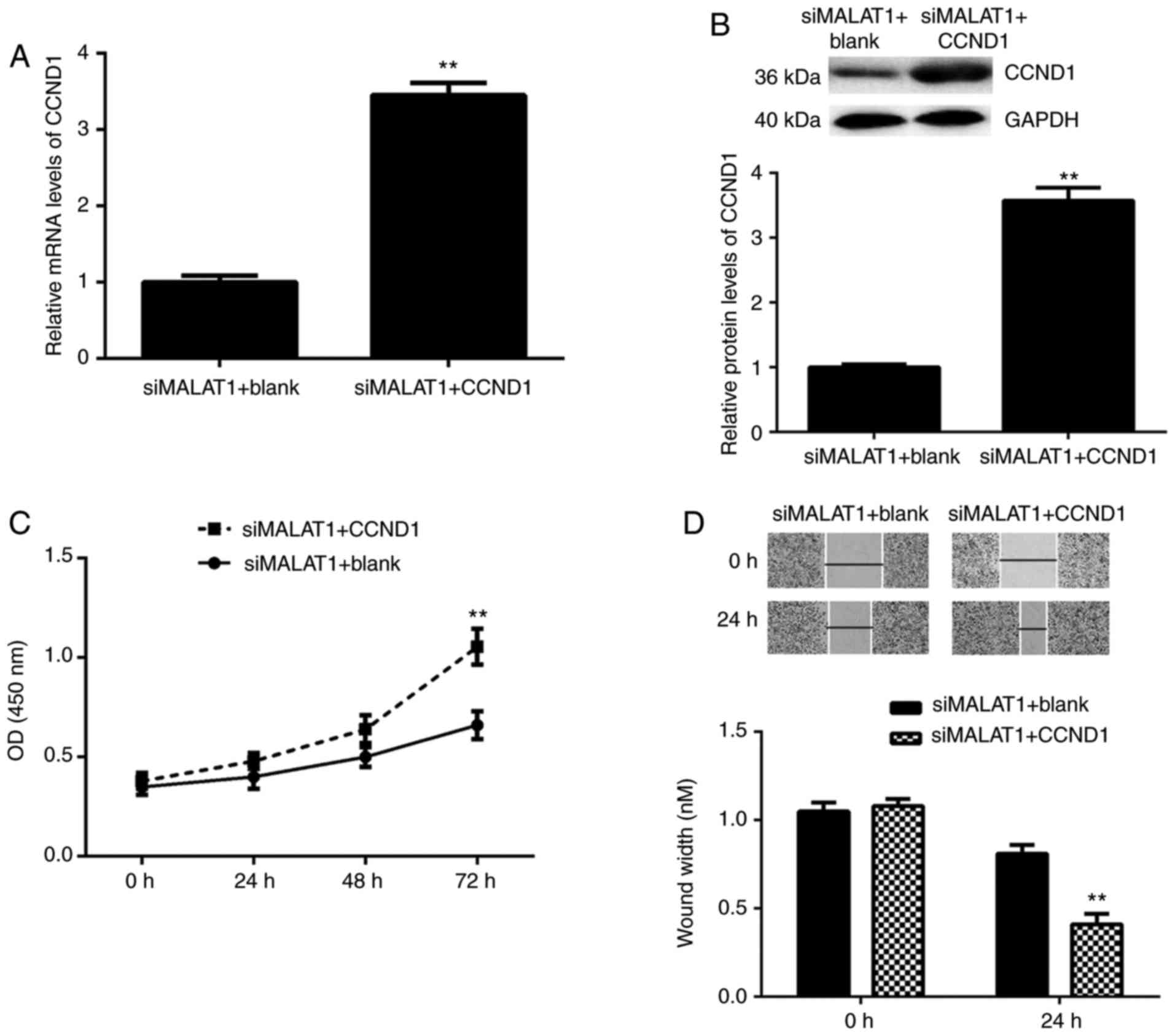
Based on the above findings, it was speculated that
CCND1 may be a downstream effector of MALAT1 in BC cells. To
clarify this speculation, siMALAT1-transfected T24 cells were
transfected with a CCND1 expression plasmid or a blank vector. As
indicated in Fig. 6A and B, CCND1
mRNA and protein expression levels were significantly upregulated
in the siMALAT1+CCND1 group compared with the siMALAT1+blank group.
These data confirmed that the transfection efficiency was
successful. Further investigation identified that cell
proliferation and migration were also significantly increased in
the siMALAT1+CCND1 group compared with the siMALAT1+blank (Fig. 6C and D), which suggested that
CCND1 upregulation may impair the inhibitory effects of MALAT1
inhibition on T24 cell proliferation and migration. Therefore, the
results suggested that CCND1 acted as a downstream effector of
MALAT1 in BC cells.
CCND1 is upregulated in BC
Finally, it was observed that the expression of
CCND1 was significantly increased in the BC tissues and cell lines
compared with the adjacent non-tumour tissues and the normal
urinary tract epithelial cell line SV-HUC-1 (Fig. 7). Therefore, the findings
suggested that CCND1 was upregulated in BC.
Discussion
The regulatory mechanism of MALAT1 underlying BC
growth and metastasis has not been fully elucidated. The present
study indicated that MALAT1 was significantly upregulated in the BC
tissues and cell lines compared with that in adjacent non-tumour
tissues and the normal urinary tract epithelial cell line SV-HUC-1.
Furthermore, the expression levels of MALAT1 were significantly
higher in the BC tissues at stage III-IV when compared with the
tissues at stage I-II. Notably, the knockdown of MALAT1
significantly inhibited BC cell proliferation and migration by
targeting miR-34a. In addition, the expression of miR-34a was
significantly decreased in the BC tissues and cell lines compared
with the adjacent non-tumour tissues and the SV-HUC-1 cells.
Furthermore, the expression of miR-34a was inversely correlated
with the expression of MALAT1 in the BC tissues. CCND1 was then
identified as a target gene of miR-34a, and its expression was
negatively mediated by miR-34a in the BC cells. Notably, the
upregulation of CCND1 impaired the effect of MALAT1 inhibition on
BC cell proliferation and migration. In addition, the expression of
CCND1 was significantly increased in the BC tissues and cell
lines.
An increasing number of lncRNAs have been
demonstrated to function as key regulators in various types of
human cancer, including BC. For instance, lncRNA activated by TGF-β
promotes BC cell proliferation, migration and invasion by
suppressing the expression of miR-126 (8). LncRNA growth arrest-specific 5
promotes BC cell apoptosis by inhibiting the transcription of Ezh2
(27). The findings in the
present study revealed that MALAT1 was significantly upregulated in
BC tissues and cell lines, and its upregulation was significantly
associated with advanced TNM stage, lymph node metastasis and a
shorter survival time in patients with BC. Consistent with these
data, Li et al (16) also
reported that the high expression of MALAT1 promoted BC progression
and metastasis and was associated with poor prognosis. In the
present study, it was revealed that the knockdown of MALAT1
decreased BC cell proliferation and migration. Similarly, Jiao
et al (28) reported that
the overexpression of MALAT1 promoted the proliferation, migration
and invasion of bladder transitional cell carcinoma cells.
LncRNAs typically function by regulating miR
expression and, thus, affect downstream target genes (12,29). For instance, the lncRNA X-inactive
specific transcript promotes TGF-β-induced EMT in NSCLC cells by
regulating the miR-367/141-zinc finger E-box binding homeobox 2
axis (30). LncRNA taurine
up-regulated 1 promotes oral squamous cell carcinoma progression by
upregulating formin like 2 by sequestering miR-219 (31). In the present study, knockdown of
MALAT1 increased miR-34a expression in T24 cells, and it was
confirmed that MALAT1 directly targeted miR-34a by using a
luciferase reporter gene assay. Furthermore, miR-34a was
downregulated in BC, and its downregulation was inversely
correlated with the increased expression of MALAT1 in the BC
tissues. These findings suggested that the decreased expression of
miR-34a may be due to the increased expression of MALAT1 in BC. The
present findings suggested that the suppressive effects of MALAT1
downregulation on the proliferation and migration of the T24 cells
was through the upregulation of miR-34a.
Bioinformatic analysis was performed to study the
potential target gene of miR-34a in BC cells, and CCND1 was
selected. The result of the luciferase reporter gene assay
indicated that CCND1 was a direct target gene of miR-34a in T24
cells. Notably, several previous studies also confirmed targeting
associations in other cancer types. For instance, Ye et al
(32) indicated that the
inhibition of miR-34a promoted the proliferation and migration of
laryngeal carcinoma cells by targeting CCND1. In addition, miR-34a
overexpression induced significant G1 cell-cycle arrest in NSCLC
cells via the inhibition of CCND1 (33). In the present study, the
inhibition of MALAT1 downregulated the expression of CCND1 in the
T24 cells, which was impaired by the silencing of miR-34a,
suggesting that MALAT1 impacted CCND1 expression via miR-34a.
Furthermore, CCND1 overexpression eliminated the inhibitory effects
on T24 cell proliferation and migration induced by MALAT1
downregulation, suggesting that CCND1 was a downstream effector of
MALAT1 in BC cells.
To the best of our knowledge, the present study
indicated for the first time that MALAT1, which is upregulated in
BC, promotes tumour cell proliferation and migration via modulating
the miR-34a/CCND1 axis. Thus, the findings suggest that MALAT1 may
be a potential therapeutic target for BC treatment.
Acknowledgments
Not applicable.
Funding
No funding was received.
Availability of data and materials
All data generated or analyzed during this study are
included in this published article.
Authors’ contributions
YL collected clinical tissues and wrote the
manuscript. YL, SG and QD performed the experiments. QZ designed
the study and revised the manuscript.
Ethics approval and consent to
participate
The present study was been approved by the Ethics
Committee of First People’s Hospital of Jining City (Jining,
China). All patients provided written informed consent.
Patient consent for publication
All patients provided written informed consent.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2015. CA Cancer J Clin. 65:5–29. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Skeldon SC and Larry Goldenberg S: Bladder
cancer: A portal into mens health. Urol Oncol. 33:40–44. 2015.
View Article : Google Scholar
|
|
3
|
Sathe A and Nawroth R: Targeting the
PI3K/AKT/mTOR pathway in bladder cancer. Methods Mol Biol.
1655:335–350. 2018. View Article : Google Scholar
|
|
4
|
Zhou Y, Meng X, Chen S, Li W, Li D, Singer
R and Gu W: IMP1 regulates UCA1-mediated cell invasion through
facilitating UCA1 decay and decreasing the sponge effect of UCA1
for miR-122-5p. Breast Cancer Res. 20:322018. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Smolle MA and Pichler M: The role of long
non-coding RNAs in osteosarcoma. Noncoding RNA. 4:E72018.PubMed/NCBI
|
|
6
|
Ruan X: Long noncoding RNA central of
glucose homeostasis. J Cell Biochem. 117:1061–1065. 2016.
View Article : Google Scholar
|
|
7
|
Wapinski O and Chang HY: Long noncoding
RNAs and human disease. Trends Cell Biol. 21:354–361. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Zhai X and Xu W: Long noncoding RNA ATB
promotes proliferation, migration and invasion in bladder cancer by
suppressing microRNA-126. Oncol Res. 26:1063–1072. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Xu R, Zhu X, Chen F, Huang C, Ai K, Wu H,
Zhang L and Zhao X: LncRNA XIST/miR-200c regulates the stemness
properties and tumourigenicity of human bladder cancer stem
cell-like cells. Cancer Cell Int. 18:412018. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Peng Z, Liu C and Wu M: New insights into
long noncoding RNAs and their roles in glioma. Mol Cancer.
17:612018. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Zhang Y, Dai Q, Zeng F and Liu H: MALAT1
Promotes the proliferation and metastasis of osteosarcoma cells by
activating the Rac1/JNK pathway via targeting miR-509. Oncol Res.
Apr 27–2018.Epub ahead of print. View Article : Google Scholar
|
|
12
|
Liu S, Song L, Zeng S and Zhang L:
MALAT1-miR-124-RBG2 axis is involved in growth and invasion of
HR-HPV-positive cervical cancer cells. Tumour Biol. 37:633–640.
2016. View Article : Google Scholar
|
|
13
|
Ji Q, Liu X, Fu X, Zhang L, Sui H, Zhou L,
Sun J, Cai J, Qin J, Ren J and Li Q: Resveratrol inhibits invasion
and metastasis of colorectal cancer cells via MALAT1 mediated
Wnt/beta-catenin signal pathway. PLoS One. 8:e787002013. View Article : Google Scholar
|
|
14
|
Hirata H, Hinoda Y, Shahryari V, Deng G,
Nakajima K, Tabatabai ZL, Ishii N and Dahiya R: Long noncoding RNA
MALAT1 promotes aggressive renal cell carcinoma through Ezh2 and
interacts with miR-205. Cancer Res. 75:1322–1331. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Ying L, Chen Q, Wang Y, Zhou Z, Huang Y
and Qiu F: Upregulated MALAT-1 contributes to bladder cancer cell
migration by inducing epithelial-to-mesenchymal transition. Mol
Biosyst. 8:2289–2294. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Li C, Cui Y, Liu LF, Ren WB, Li QQ, Zhou
X, Li YL, Li Y, Bai XY and Zu XB: High expression of long noncoding
RNA MALAT1 indicates a poor prognosis and promotes clinical
progression and metastasis in bladder cancer. Clin Genitourin
Cancer. 15:570–576. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Xie H, Liao X, Chen Z, Fang Y, He A, Zhong
Y, Gao Q, Xiao H, Li J, Huang W and Liu Y: LncRNA MALAT1 inhibits
apoptosis and promotes invasion by antagonizing miR-125b in bladder
cancer cells. J Cancer. 8:3803–3811. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Farazi TA, Hoell JI, Morozov P and Tuschl
T: MicroRNAs in human cancer. Adv Exp Med Biol. 774:1–20. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Garg D and Cohen SM: miRNAs and aging: A
genetic perspective. Ageing Res Rev. 17:3–8. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Boon RA and Vickers KC: Intercellular
transport of microRNAs. Arterioscler Thromb Vasc Biol. 33:186–192.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Hwang HW and Mendell JT: MicroRNAs in cell
proliferation, cell death, and tumorigenesis. Br J Cancer.
94:776–780. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Ma ZL, Hou PP, Li YL, Wang DT, Yuan TW,
Wei JL, Zhao BT, Lou JT, Zhao XT, Jin Y and Jin YX: MicroRNA-34a
inhibits the proliferation and promotes the apoptosis of non-small
cell lung cancer H1299 cell line by targeting TGFbetaR2. Tumour
Biol. 36:2481–2490. 2015. View Article : Google Scholar
|
|
23
|
Wu X, Zhong D, Gao Q, Zhai W, Ding Z and
Wu J: MicroRNA-34a inhibits human osteosarcoma proliferation by
downregulating ether a go-go 1 expression. Int J Med Sci.
10:676–682. 2013. View Article : Google Scholar :
|
|
24
|
Yu G, Yao W, Xiao W, Li H, Xu H and Lang
B: MicroRNA-34a functions as an anti-metastatic microRNA and
suppresses angiogenesis in bladder cancer by directly targeting
CD44. J Exp Clin Cancer Res. 33:7792014. View Article : Google Scholar
|
|
25
|
Sun H, Tian J, Xian W, Xie T and Yang X:
miR-34a inhibits proliferation and invasion of bladder cancer cells
by targeting orphan nuclear receptor HNF4G. Dis Markers.
2015:8792542015. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
27
|
Wang M, Guo C, Wang L, Luo G, Huang C, Li
Y, Liu D, Zeng F, Jiang G and Xiao X: Long noncoding RNA GAS5
promotes bladder cancer cells apoptosis through inhibiting EZH2
transcription. Cell Death Dis. 9:2382018. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Jiao D, Li Z, Zhu M, Wang Y, Wu G and Han
X: LncRNA MALAT1 promotes tumor growth and metastasis by targeting
miR-124/foxq1 in bladder transitional cell carcinoma (BTCC). Am J
Cancer Res. 8:748–760. 2018.PubMed/NCBI
|
|
29
|
Xie CH, Cao YM, Huang Y, Shi QW, Guo JH,
Fan ZW, Li JG, Chen BW and Wu BY: Long non-coding RNA TUG1
contributes to tumorigenesis of human osteosarcoma by sponging
miR-9-5p and regulating POU2F1 expression. Tumour Biol.
37:15031–15041. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Li C, Wan L, Liu Z, Xu G, Wang S, Su Z,
Zhang Y, Zhang C, Liu X, Lei Z and Zhang HT: Long non-coding RNA
XIST promotes TGF-beta-induced epithelial-mesenchymal transition by
regulating miR-367/141-ZEB2 axis in non-small-cell lung cancer.
Cancer Lett. 418:185–195. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Yan G, Wang X, Yang M, Lu L and Zhou Q:
Long non-coding RNA TUG1 promotes progression of oral squamous cell
carcinoma through upregulating FMNL2 by sponging miR-219. Am J
Cancer Res. 7:1899–1912. 2017.PubMed/NCBI
|
|
32
|
Ye J, Li L, Feng P, Wan J and Li J:
Downregulation of miR-34a contributes to the proliferation and
migration of laryngeal carcinoma cells by targeting cyclin D1.
Oncol Rep. 36:390–398. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Sun F, Fu H, Liu Q, Tie Y, Zhu J, Xing R,
Sun Z and Zheng X: Downregulation of CCND1 and CDK6 by miR-34a
induces cell cycle arrest. FEBS Lett. 582:1564–1568. 2008.
View Article : Google Scholar : PubMed/NCBI
|