Introduction
Oxidative stress is involved in the pathogenesis of
lifestyle-associated diseases, including atherosclerosis,
hypertension, diabetes mellitus, ischemic diseases and malignancies
(1). Oxidative stress results
from an imbalance in pro-oxidant/antioxidant homeostasis that leads
to the generation of toxic reactive oxygen species (ROS), such as
hydrogen peroxide, organic hydroperoxides, nitric oxide, superoxide
and hydroxyl radicals (2).
Excessive free radicals in the body attack proteins, lipids, DNA
and other biological macromolecules, leading to damaged cell
structures, interference with normal metabolic activity, disease
and accelerated aging (3). Thus,
reducing oxidative stress is both indispensable and significant in
improving the quality of life and treatment options for patients
with oxidative stress-associated diseases. Due to the potential
health hazards of synthetic antioxidants, such as dibutyl
hydroxyanisole, which is carcinogenic in animal models, searching
for efficient natural antioxidants from plants with low toxicity is
of utmost importance in the development of safe antioxidants.
The Kelch-like ECH-associated protein 1
(Keap1)/nuclear factor erythroid 2-related factor 2 (Nrf2)
signaling pathway is essential for cytoprotection against oxidative
stress. Nrf2 is a potent transcriptional activator that serves a
central role in the expression of numerous cytoprotective genes in
response to oxidative stress (4).
BTB and CNC homolog 1 (Bach1) is a transcription factor that
functions as an Nrf2 repressor, competing with Nrf2 to downregulate
the expression of antioxidant enzymes, and thus serving an
important role in the regulation of the body’s oxidant/antioxidant
imbalance (5). The cellular
oxidative stress induced by H2O2 shares the
biological and morphological characteristics of apoptosis (6). However, since oxidative damage
varies clinically and is not permanent, the effects of oxidative
stress cannot be solely explained by apoptosis. Under stress
conditions, such as nutrient deprivation, oxidative stress and/or
metabolic stress, autophagy produces metabolic substrates to meet
the bioenergetic needs of the cell, thus preventing cell death
(7). Therefore, it is necessary
to evaluate the protective efficacy of an antioxidant from the
perspective of autophagy. In auditory cells, oxidative stress
induces autophagy through molecular crosstalk among p62, Keap1 and
Nrf2 to provide protection against necrosis by ATP depletion
(8). This suggests that autophagy
may work together with the Keap1/Nrf2 pathway to resist oxidative
stress.
Flavonoids are commonly found in medicinal plants,
vegetables and fruits, and are known to significantly reduce the
incidence of cardiovascular disease.
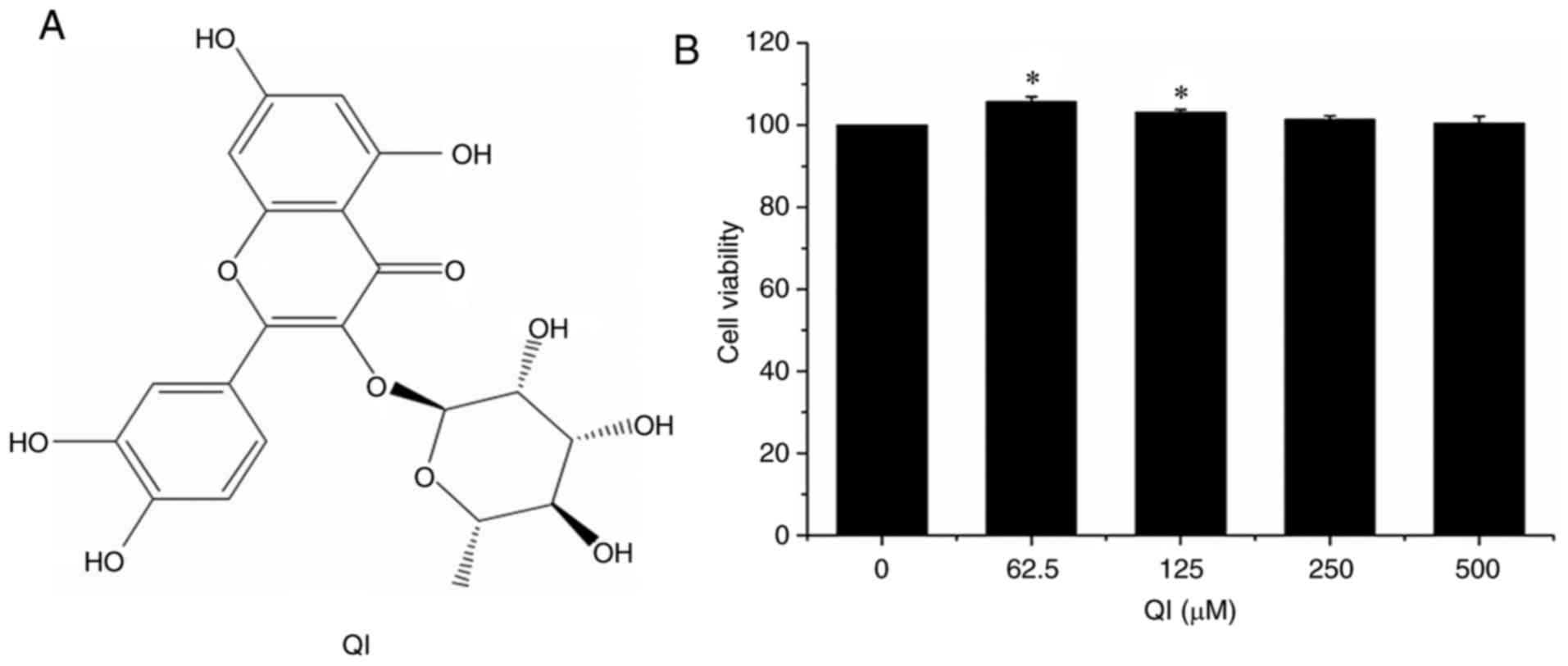
Quercetin-3-O-α-L-rhamnopyranoside (QI; molecular weight, 448;
Fig. 1A) is a natural polyphenol
that belongs to the flavonoid family and is produced in the leaves
of Lindera aggregata (Sims) Kosterm. QI has a wide range of
pharmacological activities, including antioxidant (9), antiviral (10), antidepressant (11), diabetic resistant (12), liver protectant (13) and cardiovascular protectant
activities (14). It has been
reported that QI reduces the apoptosis of endothelial progenitor
cells caused by oxidized low-density lipoprotein, and promotes
autophagy via extracellular signal-regulated kinase activation
(15). However, the underlying
mechanism of the action of QI requires further investigation.
In the present study, the aim was to examine whether
QI exerted a protective effect on human umbilical vein endothelial
cells (HUVECs) and if it activated the Nrf2 pathway by inducing
autophagy. The results revealed that the antioxidant effect of QI
occurred at the gene level. In vivo, QI significantly
improved the activity of superoxide dismutase (SOD) and glutathione
(GSH), and decreased malondialdehyde (MDA) levels in the serum and
heart tissue of aging rats. These results suggest that QI may be
used in the early treatment of cardiovascular disease, or serve as
a health supplement to alleviate oxidative stress.
Materials and methods
Reagents
Antibodies against light chain 3B (LC3B; cat. no.
3868), Caspase-9 (cat. no. 9502), Caspase-3 (cat. no. 9662),
poly(ADP-ribose) polymerase (PARP; cat. no. 9532), autophagy
related 5 (Atg5; cat. no. 12994), Atg13 (cat. no. 13468), Bach1
(cat. no. 4578) and β-actin (cat. no. 3700) were purchased from
Cell Signaling Technology, Inc. (Beverly, MA, USA), while
antibodies against Nrf2 (cat. no. R1312-8), heme oxygenase-1 (HO-1;
cat. no. ET1604-45), and Histone H3 (cat. no. ET1601-14) were
obtained from HuaBio (Hangzhou, Zhejiang, China). The secondary
antibodies used in the present study included goat anti-rabbit
horseradish peroxidase (HRP)-conjugated immunoglobulin (Ig)-G and
goat anti-mouse HRP-conjugated IgG (cat. nos. BL003A and BL001A;
Biosharp, Shanghai, China), as well as FITC goat anti-rabbit IgG
(HA1004, HuaBio, Hangzhou, Zhejiang, China). Control (CTL), Nrf2
and HO-1 small interfering (si)-RNAs (cat. nos. sc-37007, sc-37030
and sc-35554, respectively) were purchased from Santa Cruz
Biotechnology, Inc. (Dallas, TX, USA), while
Lipofectamine® 2000 transfection reagent and dimethyl
sulfoxide (DMSO) were purchased from Thermo Fisher Scientific, Inc.
(Waltham, MA, USA). 3-Methyladenine (3-MA), chloroquine (CQ) and
N-acetyl-L-cysteine (NAC) were purchased from Sigma-Aldrich
(Merck KGaA, Darmstadt, Germany), H2O2 was
obtained from Tianjin Yongda Chemical Reagent Co., Ltd. (Tianjin,
China), while
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl-2H-tetrazolium bromide
(MTT) and Triton X-100 were from Solarbio Science & Technology
Co., Ltd. (Beijing, China). The following kits were used in this
study: ROS determination kit, total SOD assay kit (hydroxylamine
method), MDA kit and reduced GSH assay kit, all purchased from
Jiancheng Bioengineering Institute (Nanjing, China); BCA protein
concentration determination kit and enhanced chemiluminescence
(ECL) system, both from Beyotime Institute of Biotechnology
(Shanghai, China); and Annexin V-FITC apoptosis detection kit, DNA
Purification kit, and Nuclear and Cytoplasmic Protein Extraction
kit, which were obtained from KeyGen Biotech Co., Ltd. (Nanjing,
China).
QI preparation
Young leaves of Lindera aggregate (Sims)
Kosterm. were collected from the Tiantaishan district (Zhejiang,
China) and authenticated by Professor Jingkui Tian in Zhejiang
University (Hangzhou, China), where the voucher specimens were
stored and evaluated during our previous study (no. LA201301-08)
(16). Separation, purification
and structural analysis of QI were performed as described in a
previous study (16). Briefly,
dried plant material (10 kg) was extracted using 70% ethanol under
reflux for 2 h, and the process was repeated three times. All the
extracts were combined and concentrated under a vacuum. The
combined extracts were diluted with H2O and
chromatographed over an AB-8 resin column, prior to eluting with
NaOH (1 BV; pH 9.0), H2O (3 BV) and 50% ethanol (3 BV).
Subsequently, the 50% ethanol elution was concentrated and
chromatographed on a reverse phase silica gel column (sample
amount, 1 g; intermediates, 40 ml column volume; ethanol
concentration, 14.25%; elution, 3 BV/h). Finally, an ethanol-water
system was used for re-crystallization, and the solution was then
set aside for 24 h at room temperature to obtain the QI.
Cells and culture conditions
Human umbilical vein endothelial cells (HUVECs) were
obtained from the American Type Culture Collection (Manassas, VA,
USA). Cells were cultured in Endothelial Cell Medium (ECM;
ScienCell Research Laboratories, Inc., San Diego, CA, USA)
supplemented with 5% fetal bovine serum (FBS) and 1%
penicillin/streptomycin solution (100 IU/ml penicillin and 100
mg/ml streptomycin). The cells were maintained in a 37°C humidified
incubator with 5% CO2.
MTT assay to determine cytotoxicity, cell
viability and the effect of the autophagy inhibitor 3-MA
The cytotoxicity of QI was evaluated by an MTT
assay. Briefly, cells were seeded in 96-well plates at
3×104 cells/well in a final volume of 100 µl.
After 16 h, fresh FBS-free medium with various concentrations of QI
(0, 62.5, 125, 250 and 500 µM) was added to each well.
Following 24 h of incubation, 20 µl MTT (5 mg/ml) per well
was added, and the cells were incubated for a further 2 h.
Subsequently, DMSO was used to dissolve the formazan crystals, and
a microplate reader was used to measure the optical density at 570
nm. Untreated cells were set to 100% viability in order to
determine the cytotoxicity of QI.
The antioxidant properties of QI on cell viability
were also assessed using MTT assay. Briefly, cells or transfected
cells were treated with DMSO, H2O2 only or
H2O2 (1,400 µM) and QI (62.5, 125, 250
and 500 µM) combined for 3 h. As described earlier, the
medium was removed and replaced by 20 µl MTT (5 mg/ml).
After 2 h, the supernatants were removed from the wells, and MTT
dye was solubilized in DMSO at 200 µl/well. The absorbance
at 570 nm was determined on a microplate reader.
HUVECs were seeded into 96-well plates at
3×104 cells/well in a final volume of 100 µl.
When completely adherent, cells were treated with 25 µM 3-MA
(an autophagy inhibitor) for 1 h at 37°C, followed by QI treatment
as aforementioned. Following 3 h, the medium was replaced with
fresh FBS-free ECM, and 20 µl MTT (5 mg/ml) was added to
each well for a further 2 h incubation. The crystals were then
dissolved in DMSO and the optical density at 570 nm was detected
using a microplate reader following high-speed vibration for 3 min.
The cell viability was calculated by considering untreated cells as
100% viable.
Plasmid transfection and observation of
apoptosis morphology
Cells were transfected with plasmids encoding red
fluorescent protein (RFP; Addgene, Inc., Cambridge, MA, USA) using
the VigeneFection (VGF) plasmid transfection reagent (Vigene
Biosciences, Inc., Rockville, MD, USA), according to the
manufacturer’s protocol. Briefly, cells were treated with 0.5
µg plasmid DNA and 2 µl VGF in each well of a 24-well
plate. Then HUVECs were seeded in confocal dishes following 24 h of
stable transfection. The expression of RFP in HUVECs was confirmed
by fluorescence detection. When 80-90% confluence was reached,
treatments were initiated. Confocal images were obtained with an
Olympus FV1000 confocal microscope (Olympus Corp., Tokyo,
Japan).
Transient transfection with siRNA
Cells were seeded in 6-well plates at
5×105 cells/well and then transfected with the CTL siRNA
(17), Nrf2 siRNA (18) or HO-1 siRNA (19) using Lipofectamine 2000™. Briefly,
3 µg of siRNA was added to each well with 6 µl
Lipofectamine 2000™. Treatments were initiated following 24 h
transfection. The expression of the Nrf2 and HO-1 proteins was
confirmed by western blot analysis.
SOD, GSH, and MDA assay in HUVECs
HUVECs were cultured at a density of
2×105 per well in 6-well plates and cultured overnight
and then treated with H2O2 (1,400 µM)
and QI (62.5, 125, 250 and 500 µM) for 3 h. Then cells were
collected and assay kits (Jiancheng Bioengineering Institute) were
used to measure the activity of MDA, GSH and SOD.
Cell apoptosis analysis
The role of QI in protecting HUVECs from
H2O2-induced apoptosis was assessed by flow
cytometry using an Annexin V-FITC apoptosis detection kit. Briefly,
cells were harvested by non-enzymatic cell dissociation and
centrifuged (500 × g for 5 min at room temperature) to remove the
medium. The cells were then washed twice with phosphate-buffered
saline (PBS) prior to re-suspending in 500 µl binding
buffer. Next, the cells were stained with 5 µl Annexin
V-FITC and 5 µl propidium iodide (PI). After 15-min
incubation without exposure to light, the Annexin V and PI
emissions were detected in the FL1-H and FL2-H channels,
respectively, with an excitation wavelength of 488 nm and emission
wavelength of 530 nm on a FACS-Vantage flow cytometer (Cytomics FC
500; Beckman Coulter, Inc., Brea, CA, USA).
DNA ladder assay
HUVECs were seeded in 6-well plates at a
concentration of 7.0×105 cells/well. At 16 h after
planting, the cells were treated with H2O2
with or without QI. Following the treatment, DNA was isolated with
a DNA Purification kit and electrophoretically analyzed on 1.5%
agarose gel.
ROS assay
Cells were seeded into 6-well plates at
5×105 cells/well and treated with
H2O2 and QI at a range of concentrations for
3 h. The medium was removed and replaced with 1 ml PBS containing 1
µM CM-H2DCFDA (Jiancheng Bioengineering
Institute), and cells were then incubated in a 5% CO2
incubator at 37°C for 30 min in the dark. Next, the cells were
collected in 15 ml centrifuge tubes. Subsequent to washing with PBS
three times, the ROS level for each treatment group was determined
using a flow cytometer (Cytomics FC 500) and analyzed using FlowJo
software (version 7.6; FlowJo LLC, Ashland, OR, USA).
Nuclear transfer of Nrf2
Cells treated with QI and
H2O2, or pretreated with 3-MA (25 µM)
were washed with cold PBS and fixed with methanol (pre-chilled at
−20°C) for 5 min at room temperature. Next, cells were
permeabilized with 0.5% Triton X-100 in PBS for 10 min and then
blocked with 1% bovine serum albumin (BSA) in PBS for 30 min.
Following washing with PBS, the cells were incubated overnight with
an anti-Nrf2 antibody in 1% BSA at 4°C, followed by incubation with
FITC goat anti-rabbit IgG in 1% BSA for 2 h at room temperature.
Nuclei were stained with DAPI in Vectashield mounting medium
(Vector Laboratories, Inc., Burlingame, CA, USA). Confocal images
were obtained using the 63x oil immersion lens on a confocal
microscope (LSM 780; Carl Zeiss AG, Oberkochen, Germany).
Western blot analysis
Drug-treated cells were collected by scraping,
followed by centrifugation (500 × g for 5 min at 4°C). Nuclear and
cytoplasmic proteins were extracted according to the protocol of
the Nuclear and Cytoplasmic Protein Extraction kit. The extraction
of total protein was performed as follows: Cells were washed once
with PBS and then lysed in a lysis buffer consisting of 1% sodium
dodecyl sulfate (SDS), 10 mM ethylene diamine tetra acetic acid
(EDTA) and 50 mM Tris-HCl (pH 8.1) in the presence of a 1% protease
inhibitor mixture (Sigma-Aldrich; Merck KGaA). Lysates were
sonicated (15 sec) to shear genomic DNA and then centrifuged at
13,000 × g for 10 min at 4°C. The protein concentrations in the
supernatants were determined using the Micro BCA Protein Assay kit.
Equal protein amounts were resolved on an SDS-polyacrylamide gel
and transferred to a polyvinylidene fluoride membrane (Bio-Rad
Laboratories, Inc., Hercules, CA, USA). The membrane was then
blocked with 5% non-fat milk in Tris-buffered saline containing
0.1% Tween-20 (TBST) for 30 min at room temperature, and then
incubated with the LC3B (1:1,000), Capase-9 (1:500), Caspase-3
(1:500), PARP (1:1,000), Atg13 (1:500), Atg5 (1:500), Nrf2 (1:500),
HO-1 (1:500), Bach-1 (1:500), Histone H3 (1:1,000) and β-Actin
(1:1,000) antibodies in TBST at 4°C overnight. The membrane was
then washed three times with TBST for a total of 30 min, followed
by incubation with goat secondary antibodies against rabbit or
mouse IgG conjugated to HRP (1:5,000 dilution) for 2 h at room
temperature. Subsequent to washing three times with TBST for a
total of 30 min, ECL was used to develop the images and the
immunoblots.
Detection of autophagic flux and the
effect of ROS on autophagy
HUVECs were seeded into 6-well-plates, and then
separately treated with CQ (10 µM) or QI (250 µM), or
co-treated with CQ and QI for 3 h. Cells were subsequently lysed to
determine LC3B and β-actin levels by western blot analysis.
Furthermore, cells that were separately treated with QI (0, 62.5,
125 and 250 µM) and NAC (5 mM) only or NAC pretreatment for
1 h with QI (62.5, 125 and 250 µM) were also lysed to
determine LC3B and β-actin levels by western blot analysis.
Fluorescent quantitative polymerase chain
reaction (qPCR)
Briefly, HUVECs were seeded into 96-well plates at
3×104 cells/well. Following treatment of cells with QI,
the medium was removed and cells were washed with 50 µl cold
PBS. For the extraction of cellular RNAs, the ExCellenCT Lysis kit
(Applied Biological Materials, Inc., Richmond, BC, Canada) was used
according to the manufacturer’s protocol. A total of 1 µl
protease and 50 µl lysis solution were added to each well,
mixed and incubated for 10 min in an incubator. To end the
reaction, 1 µl protease inhibitor and 5 µl stop
solution were added and incubated for 20 min at room temperature.
Next, the RNA concentration was measured with a microplate reader.
Reverse transcription was conducted in an RNase-free environment
using 5X All-In-One RT MasterMix (Applied Biological Materials,
Inc.) according to the manufacturer’s protocol. The reverse
transcription system contained 2 µl 5X All-In-One RT
MasterMix, 13 µl nuclease-free water and 5 µl total
RNA (2 µg). The reverse transcription conditions were as
follows: 25°C for 15 min, 42°C for 30 min, 85°C for 5 min, and then
held at 4°C. Subsequent to reverse transcription, EvaGreen 2X qPCR
MasterMix (Applied Biological Materials, Inc.) was applied in a
reaction system containing 0.3 µl forward primer (10
µM), 0.3 µl reverse primer (10 µM), 0.6
µl cDNA (10 ng/µl), 5 µl EvaGreen and 3.8
µl H2O. The qPCR reaction conditions (25°C for 15
min, 42°C for 30 min, 85°C for 5 min, and then held at 4°C) were
similar to those of reverse transcription. The primers used in the
present study were as follows: β-actin forward, 5′-CGG GAA ATC GTG
CGT GAC AT-3′, and reverse, 5′-GAA CTT TGG GGG ATG CTC GC-3′; Nrf2
forward, 5′-GAT TCT GAC TCC GGC ATT TC-3′, and reverse, 5′-TCC CCA
GAA GAA TGT ACT GG-3′; and HO-1 forward, 5′-GCC CTG CCC TTC AGC
AT-3′, and reverse, 5′-CTG CAT TTG AGG CTG A GC C-3′. The relative
quantification 2−ΔΔCq method (20) was used to evaluate quantitative
variation between treatments.
In vivo study
In the present study, 50 specific-pathogen-free
female rats of the Sprague-Dawley strain (>18 weeks; 500-550 g;
JOINN Laboratories, Suzhou, China) were housed in groups under a
12-h light/dark cycle, at a constant temperature of 24°C and
humidity of 40%; all animals were provided with sterilized food and
water ad libitum. Following 7 days of acclimatization, the
animals were randomly distributed into five groups, each containing
10 animals of similar average body weight. Animals in the five
groups received the following treatments: i) Vehicle (20%
polyethylene glycol 2000 and 10% polyethylene glycol 400 in
saline); ii) 4.0 mg/kg breviscapine, a Chinese medicine used to
treat cardiovascular and cerebrovascular diseases (21), serving as a positive control; iii)
QI at a dose of 4.0 mg/kg; iv) QI at a dose of 8.0 mg/kg; and v) QI
at a dose of 16.0 mg/kg. All treatments were administered by tail
vein injection once daily for 2 weeks. All experimental procedures
using live animals were conducted in accordance with protocols
approved by the Ethics Review Committee for the Use of Animal
Subjects of Zhejiang University (Zhejiang, China).
Preparation of homogenates
Animals underwent a 12-h fast from food prior to
sacrifice, and then hearts were collected. Tissue homogenates were
prepared by grinding 0.1 g of the tissue in 1 ml saline. Following
centrifugation at 1,500 × g for 15 min at 4°C, the supernatant was
removed and kept at −20°C until further use.
Preparation of sera and evaluation of
biochemical parameters
Blood was collected by orbital puncture from the
rats, following anesthesia by ethyl ether. The blood samples were
then allowed to settle for 30 min prior to centrifugation at 2,500
× g for 10 min at 4°C. Sera and heart tissue homogenates were used
for detecting biochemical parameters associated with oxidative
stress, including SOD, GSH and MDA, according to the protocols of
the corresponding kits.
Statistical analysis
Data are presented as the mean ± standard deviation.
One-way analysis of variance and least significant difference tests
were used for statistical analysis in all experiments. Differences
were considered to be statistically significant at P<0.05.
Statistical analyses were performed using the SPSS statistical
software (version 17.0; SPSS, Inc., Chicago, IL, USA).
Results
QI promotes the growth of HUVECs
The MTT cell proliferation assay was used to assess
QI toxicity. QI promoted the growth of HUVECs at all the
investigated concentrations (62.5-500 µM), and significantly
accelerated cell proliferation at the concentrations of 62.5 and
125 µM (Fig. 1B).
Treatment with 62.5, 125, 250 and 500 µM QI for 24 h
increased cell viability by 5.75, 3.14, 1.44 and 0.49%,
respectively, compared to the control (Fig. 1B). These findings suggest that, at
concentrations ranging between 62.5 and 500 µM, QI does not
damage HUVECs.
QI protects HUVECs from damage induced by
H2O2
The study next investigated the effect of QI on the
growth of HUVECs following H2O2-induced
oxidative damage. The HUVECs ultimately demonstrated apoptosis-like
changes in their morphology, including plasma rounding and cell
shrinkage, as a result of oxidative injury in the
H2O2-treated group (Fig. 2A). By contrast, fewer
apoptotic-like cells were observed in the cell groups co-treated
with various concentrations of QI and H2O2
(Fig. 2B). Co-treatment with
62.5, 125, 250 or 500 µM QI resulted in significant
dose-dependent protection against cell death induced by 1,400
µM H2O2 (Fig. 2C).
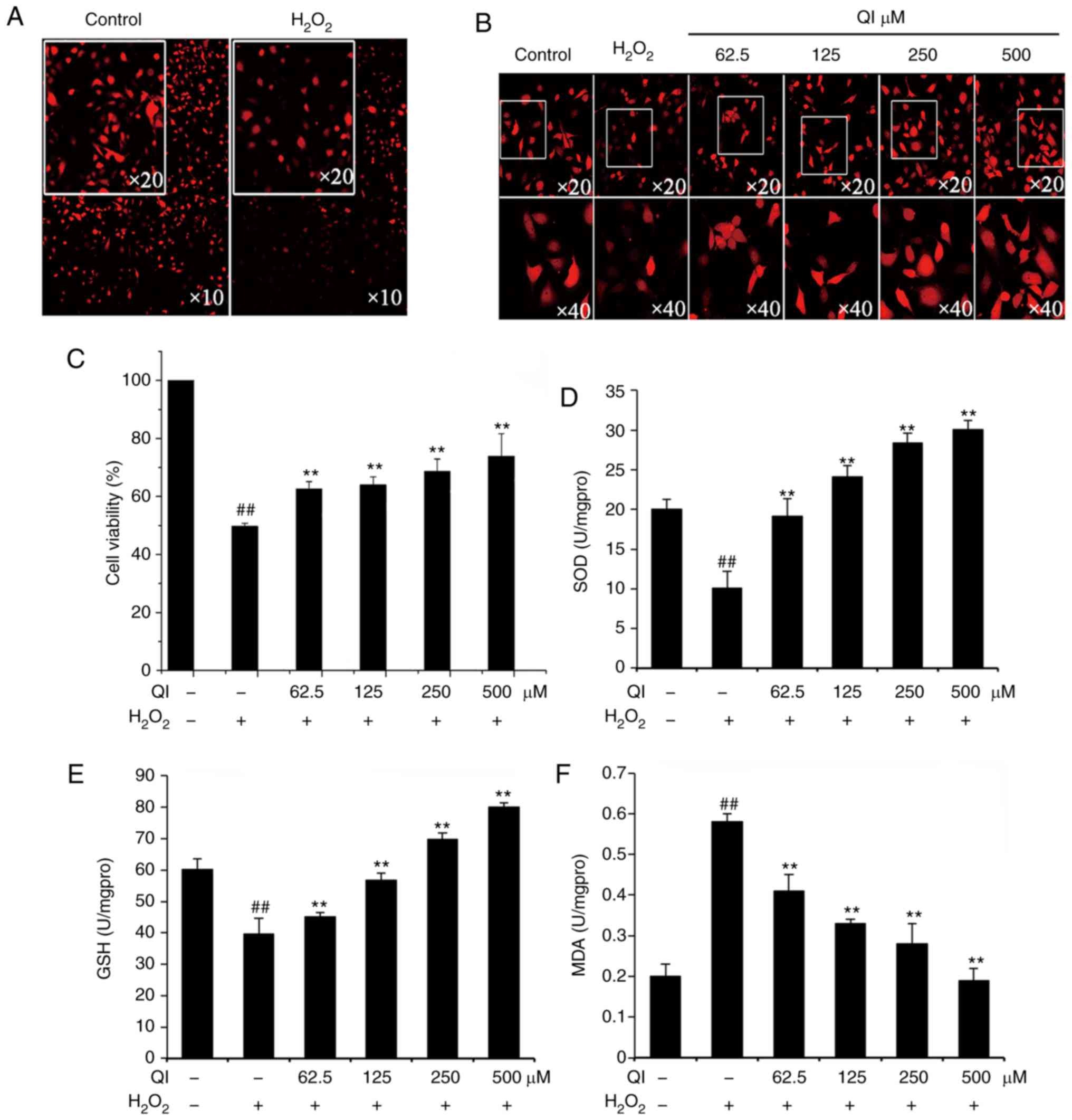 | Figure 2QI protects HUVECs from
H2O2-induced apoptosis by increasing SOD
activity and GSH content. Confocal images examining the morphology
of cells treated with (A) H2O2 alone and (B)
H2O2 in combination with QI, demonstrating
the protective effect of QI against
H2O2-induced oxidative stress injury in red
fluorescent protein-stained HUVECs (magnification, ×10, ×20 or
×40). (C) Effect of QI on the viability of HUVECs following
H2O2-induced damage was measured by an MTT
assay. Data are expressed as the percentage of viable cells
(treated vs. control cells). (D) SOD and (E) GSH intracellular
activities of HUVECs were improved by QI treatment, while (F) MDA
activity was reduced. Cells were treated with
H2O2 and QI for 3 h at 37°C, and then the
SOD, GSH and MDA activities were determined by the corresponding
assay kits. All assays were performed in triplicate, and data are
expressed as the mean ± standard deviation. ##P<0.01
vs. control; **P<0.01 vs. H2O2
treatment. QI, Quercetin-3-O-α-L-rhamnopyranoside; HUVECs, human
umbilical vein endothelial cells; SOD, superoxide dismutase; GSH,
glutathione; MDA, malondialdehyde. |
QI reduces
H2O2-induced increases in MDA level, and
improves intracellular SOD and GSH activities in HUVECs
Incubation of HUVECs with H2O2
for 3 h significantly increased MDA levels, and decreased SOD and
GSH activities as compared with those in the control group.
However, co-treatment of HUVECs with QI significantly reduced the
H2O2-induced MDA levels and restored the SOD
and GSH activities compared with the group treated with
H2O2 alone, in a concentration-dependent
manner (Fig. 2D-F). In comparison
with the H2O2-treated group, cells treated
with 62.5, 125, 250 and 500 µM QI displayed increased SOD
levels by 89.10, 138.61, 181.19 and 198.02%, respectively. A
similar improvement was observed in GSH activity following QI
treatment in HUVECs, with a significant increase detected (Fig. 2E). Compared with the
H2O2-only group, the cell groups treated with
62.5, 125, 250 and 500 µM QI displayed a markedly higher
content of GSH by 13.89, 43.18, 76.26 and 102.02%, respectively. By
contrast, MDA was reduced by 29.31, 43.10, 51.72 and 67.24% in the
62.5, 125, 250 and 500 µM QI-treated cells, respectively, as
compared with the H2O2 treatment alone.
QI inhibits
H2O2-induced apoptosis in HUVECs
Staining with Annexin V-FITC/PI and flow cytometry
were performed to identify the apoptotic cells. The results
revealed that cells exposed to H2O2 alone
exhibited a higher apoptotic rate (46.7±3.3%) as compared with the
control cells (1.3±0.1%). A two-dimensional diagram also revealed
that QI treatment at concentrations of 62.5, 125, 250 and 500
µM decreased the proportion of apoptotic cells to 30.3±1.8,
26.9±1.7, 19.3±1.4 and 18.4±0.8%, respectively, as compared with
that observed in cells that suffered H2O2
injury (Fig. 3A and B).
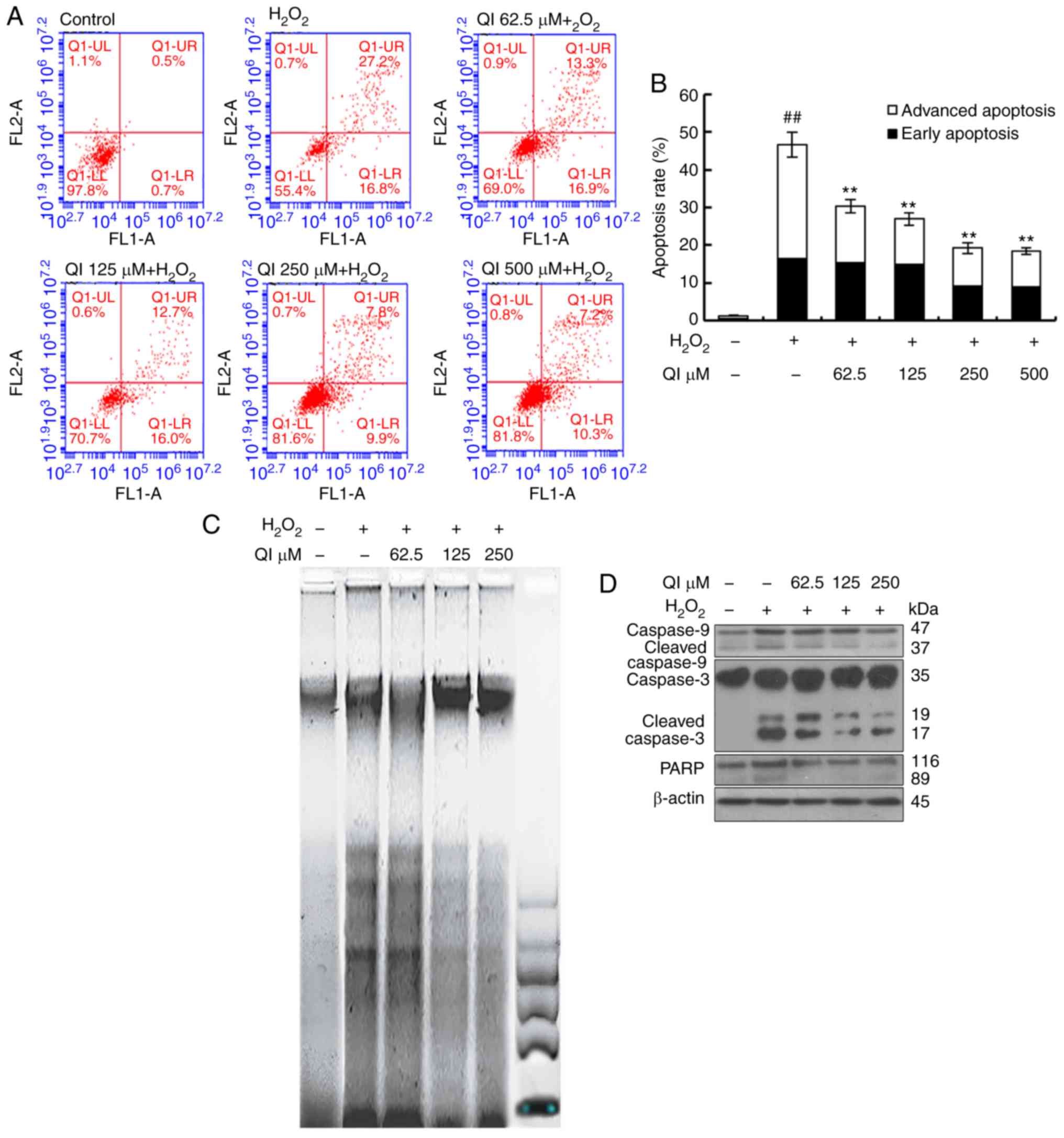 | Figure 3Inhibitory effect of QI on cell
apoptosis that was induced by H2O2. (A)
Representative dot-plots of living, early apoptotic and late
apoptotic/necrotic cells, as determined by flow cytometry with
Annexin V and PI double staining. HUVECs were incubated in
drug-free medium, or medium containing H2O2
and QI for 3 h. The horizontal axis represents the Annexin V
intensity, and the vertical axis represents the PI staining. (B)
Apoptosis rate analysis. All assays were performed in triplicate,
and data are expressed as the mean ± standard deviation.
##P<0.01 vs. control; **P<0.01 vs.
H2O2 treatment. (C) QI inhibited
H2O2-induced DNA damage in HUVECs. Fragmented
DNA was extracted and analyzed on 1.5% agarose electrophoresis
gels. (D) Western blot analysis of Caspase-9, cleaved Caspase-9,
Caspase-3, cleaved Caspase-3 and PARP protein expression levels in
HUVECs following H2O2 treatment with or
without QI. HUVECs, human umbilical vein endothelial cells; QI,
Quercetin-3-O-α-L-rhamnopyranoside; PARP, poly(ADP-ribose)
polymerase. |
The present study further investigated whether
H2O2 caused DNA fragmentation, which is
another hallmark of apoptosis, in HUVECs. Agarose gel
electrophoresis revealed that HUVECs treated with
H2O2 exhibited DNA fragmentation, as
indicated by the typical DNA laddering pattern, whereas QI-treated
cells demonstrated significantly reduced DNA laddering (Fig. 3C). Additionally, treatment with QI
resulted in a dose-dependent decrease in cleaved Caspase-3 and PARP
levels, as well as activation of Caspase-9, compared with the
H2O2 treatment alone (Fig. 3D). Collectively, these results
indicate that QI protects against cell death by inhibiting
H2O2-induced apoptosis.
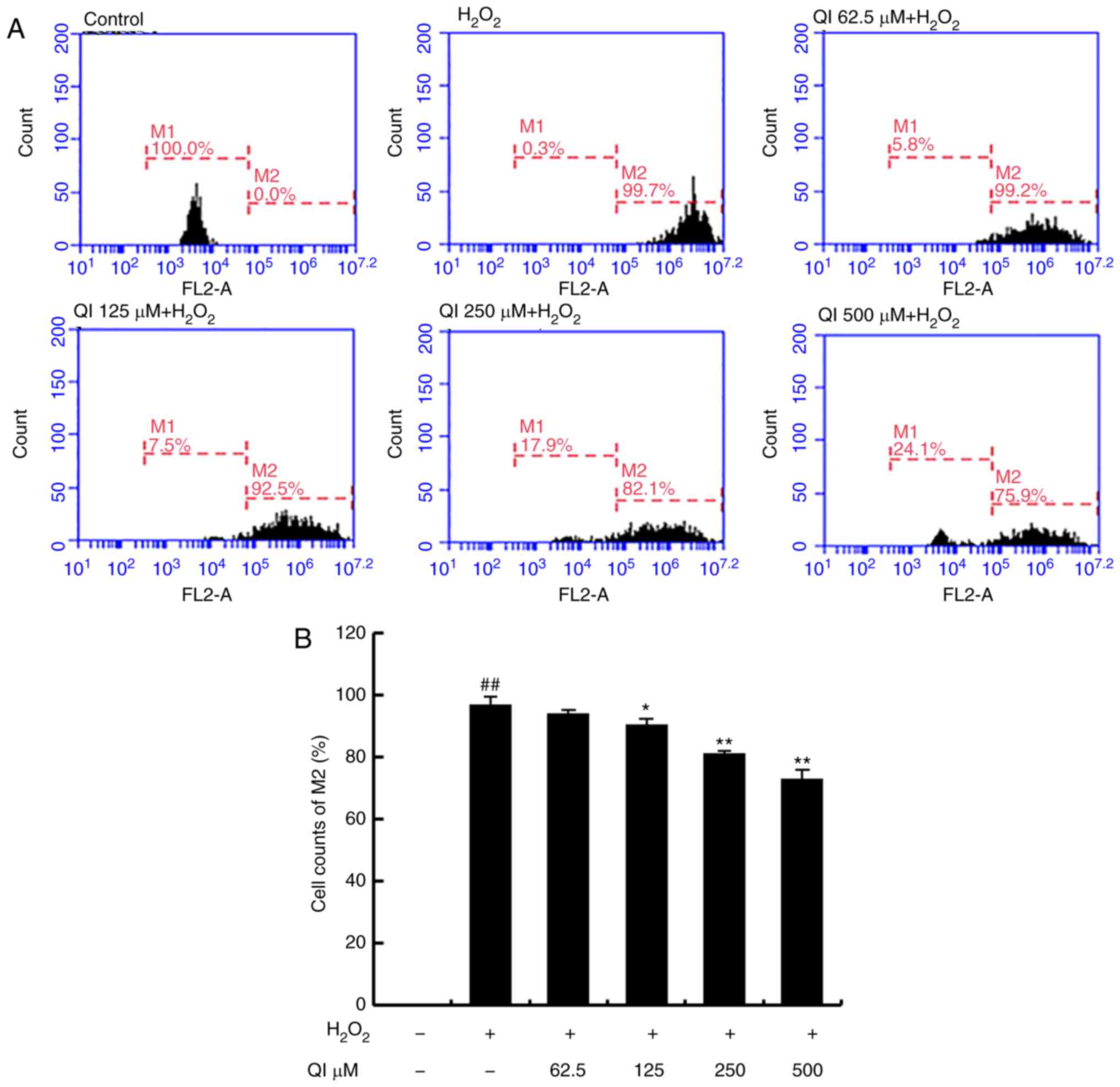
QI inhibits the production of ROS
triggered by H2O2
Oxidative stress inhibition was further corroborated
by the fact that QI mediated decreases in
H2O2-induced ROS production in the HUVECs.
The rate of ROS-positive cells was decreased from 94.2±1.1% in the
62.5 µM QI-treated group to 73.2±2.7% in the 500 µM
QI-treated group, as compared with the ROS rate of 97.1±2.3% in the
H2O2 treatment alone group (Fig. 4A and B). These results indicate
that QI provides resistance to ROS in a dose-dependent manner.
QI protects HUVECs from
H2O2-induced cytotoxicity through activation
of the Keap1/Nrf2 pathway and inhibition of Bach1
In light of the aforementioned findings, the
pathways involved in QI-mediated protection against oxidative
stress were examined in HUVECs by immunofluorescence and western
blot assays. Nrf2 is essential for cytoprotection against oxidative
stress (22). The present study
revealed that nuclear Nrf2 levels were increased subsequent to QI
treatment, as indicated by both the immunofluorescence and western
blot assay results; Histone H3 was used as a nuclear internal
reference (Fig. 5A–C).
Furthermore, Bach1 nuclear expression was decreased in QI-treated
cells, but was increased in the cytoplasm, indicating that QI
promoted the transfer of Bach1 from the nucleus to the cytoplasm,
thus releasing its inhibitory effect on Nrf2 (Fig. 5C). The expression of HO-1 protein,
which acts downstream of Nrf2, was also increased following QI
treatment (Fig. 5D), suggesting
that the antioxidant effect of QI was regulated by the Nrf2
antioxidant pathway.
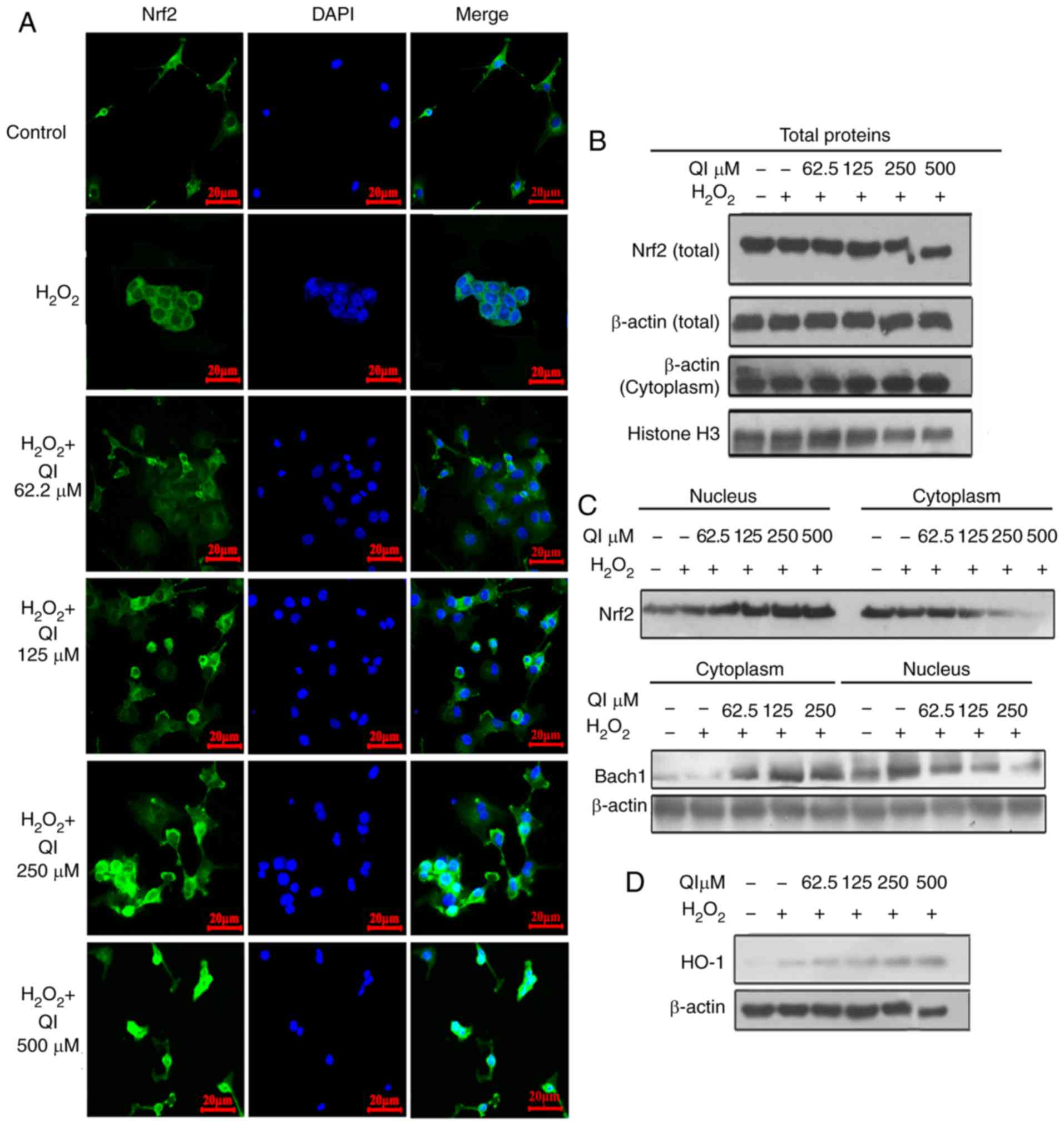 | Figure 5QI promotes Nrf2 nuclear transfer,
increased the expression of the antioxidative protein HO-1 and
inhibits Bach1 competition. (A) Immunohistochemical analysis of
Nrf2 in human umbilical vein endothelial cells (blue, DAPI; green,
Nrf2). Confocal images were obtained using the 63X oil immersion
lens of a confocal microscope (scale bar, 20 µm). Western
blot analysis results of (B) Nrf2 and Histone H3 total proteins,
(C) Nrf2 and Bach1 protein levels in the nucleus and cytoplasm, and
(D) HO-1 levels (an Nrf2 downstream protein). QI,
Quercetin-3-O-α-L-rhamnopyranoside; Nrf2, nuclear factor erythroid
2-related factor 2; Bach1, BTB and CNC homolog 1; HO-1, heme
oxygenase-1. |
QI upregulates the expression levels of
the Nrf2 and HO-1 genes
qPCR was used to further investigate the
antioxidative mechanism of QI in vitro. The data
demonstrated that the expression levels of Nrf2 and HO-1 genes
increased following QI treatment (Fig. 6A). Taken together, these results
revealed that QI affects both the protein levels of Nrf2 and HO-1,
as well as their transcriptional levels.
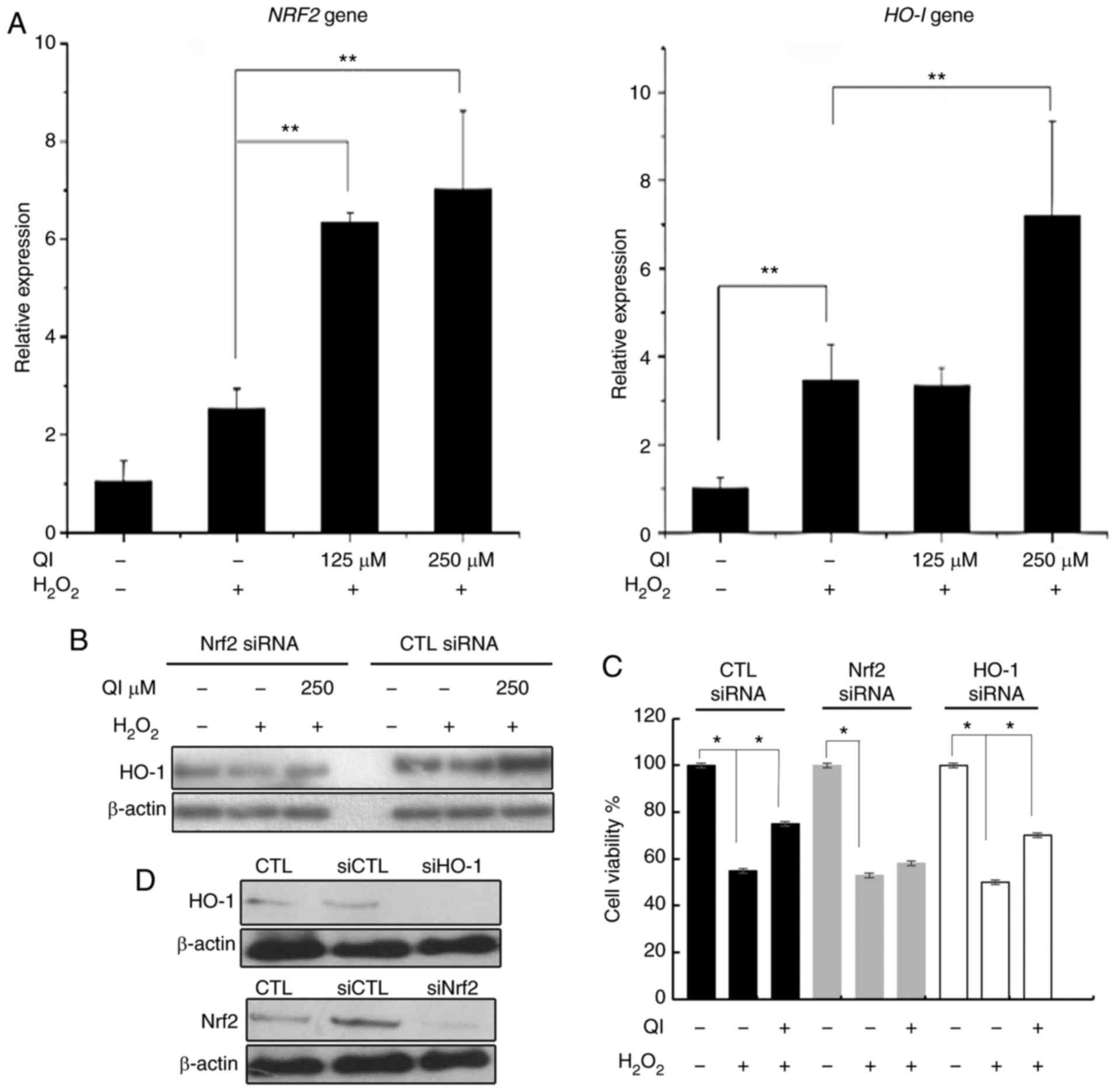 | Figure 6QI upregulates the expression of
antioxidant genes Nrf2 and HO-1, while knockdown of Nrf2 by siRNA
transfection suppresses the QI-induced antioxidant effect in
HUVECs. (A) Quantitative polymerase chain reaction analysis of Nrf2
and HO-1 gene levels. (B) Western blot analysis of HO-1 protein
expression in cells transfected with CTL and Nrf2 siRNA, followed
by QI and H2O2 treatment. (C) Effect of QI on
the viability of HUVECs with H2O2-induced
damage was measured by an MTT assay following CTL, Nrf2 or HO-1
siRNA transfection (expressed as the percentage of viable cells;
treated vs. control cells). (D) Western blot analysis was also
performed to confirm that Nrf2 and HO-1 siRNA transfection resulted
in successful knockdown. All assays were performed in triplicate,
and data represent the mean ± standard deviation.
*P<0.05 and **P<0.01, as indicated. QI,
Quercetin-3-O-α-L-rhamnopyranoside; HUVECs, human umbilical vein
endothelial cells; Nrf2, nuclear factor erythroid 2-related factor
2; HO-1, heme oxygenase-1; CTL, control; siRNA, small interfering
RNA. |
Nrf2 inhibition suppresses QI-induced
antioxidant expression and cytoprotection
As mentioned earlier, nuclear localization of Nrf2
was found to be increased, while Nrf2 gene expression was
upregulated by QI under oxidative stress conditions (Figs. 5 and 6A). Therefore, the study further
examined whether reducing Nrf2 expression using siRNA transfection
affects antioxidants. Nrf2 is known to activate the transcription
of numerous cytoprotective enzymes, such as HO-1. The current study
results revealed that Nrf2 siRNA transfection suppressed the effect
of QI on increased HO-1 expression under oxidative stress
conditions (Fig. 6B).
Next, the study examined whether Nrf2 siRNA or HO-1
siRNA were able to suppress the QI-induced cytoprotection in HUVECs
by assessing the cell viability using an MTT assay following siRNA
transfection. QI significantly reduced the
H2O2-induced cytotoxicity of HUVECs
transfected with CTL siRNA, with a cell viability of 55.1% observed
in the H2O2 group and 75.2% in the
H2O2+QI group (P<0.05; Fig. 6C). By contrast, QI treatment did
not markedly reduce the H2O2-induced
cytotoxicity of HUVECs transfected with Nrf2 siRNA
(H2O2, 53.2%; H2O2+QI,
58.3%; P>0.05). However, in cells transfected with HO-1 siRNA,
QI was able to reduce the H2O2-induced
cytotoxicity, with a cell viability of 50.3 and 70.2% in the
H2O2 and H2O2+QI
groups, respectively (P<0.05; Fig.
6C). Fig. 6D presents the
blots confirming that the transfection of Nrf2 siRNA and HO-1 siRNA
resulted in successful knockdown. These results suggest that
QI-induced Nrf2 nuclear transcription is critical for
cytoprotection against oxidative stress.
QI promotes Nrf2 transcription and
inhibits HUVEC damage by inducing autophagy
Autophagy detection was also performed in the
present study. The expression levels of Atg5, Atg13 and the
autophagic marker LC3B-II were increased following QI treatment
(Fig. 7A and B). When combined
with autophagy flux detection, treatment with both QI and CQ
resulted in an evident improvement in LC3B-II levels as compared
with that observed in cells with QI or CQ treatment alone (Fig. 7C), suggesting that QI promotes
autophagy. To further assess whether ROS or QI induced autophagy,
the ROS inhibitor NAC was used. The results found that addition of
NAC had no effect on the QI-induced increase of LC3B-II levels
(Fig. 7D), further demonstrating
that QI can induce autophagy. Notably, the transfer of Nrf2 from
the cytoplasm to the nucleus was inhibited by 3-MA, an autophagy
inhibitor (Fig. 7E). To further
test the hypothesis that autophagy is involved, the cell viability
was measured following 3-MA pre-treatment. It was observed that the
protective effect of QI was diminished by the addition of 3-MA
(Fig. 7F), indicating that QI
serves its antioxidant role in HUVECs by inducing autophagy.
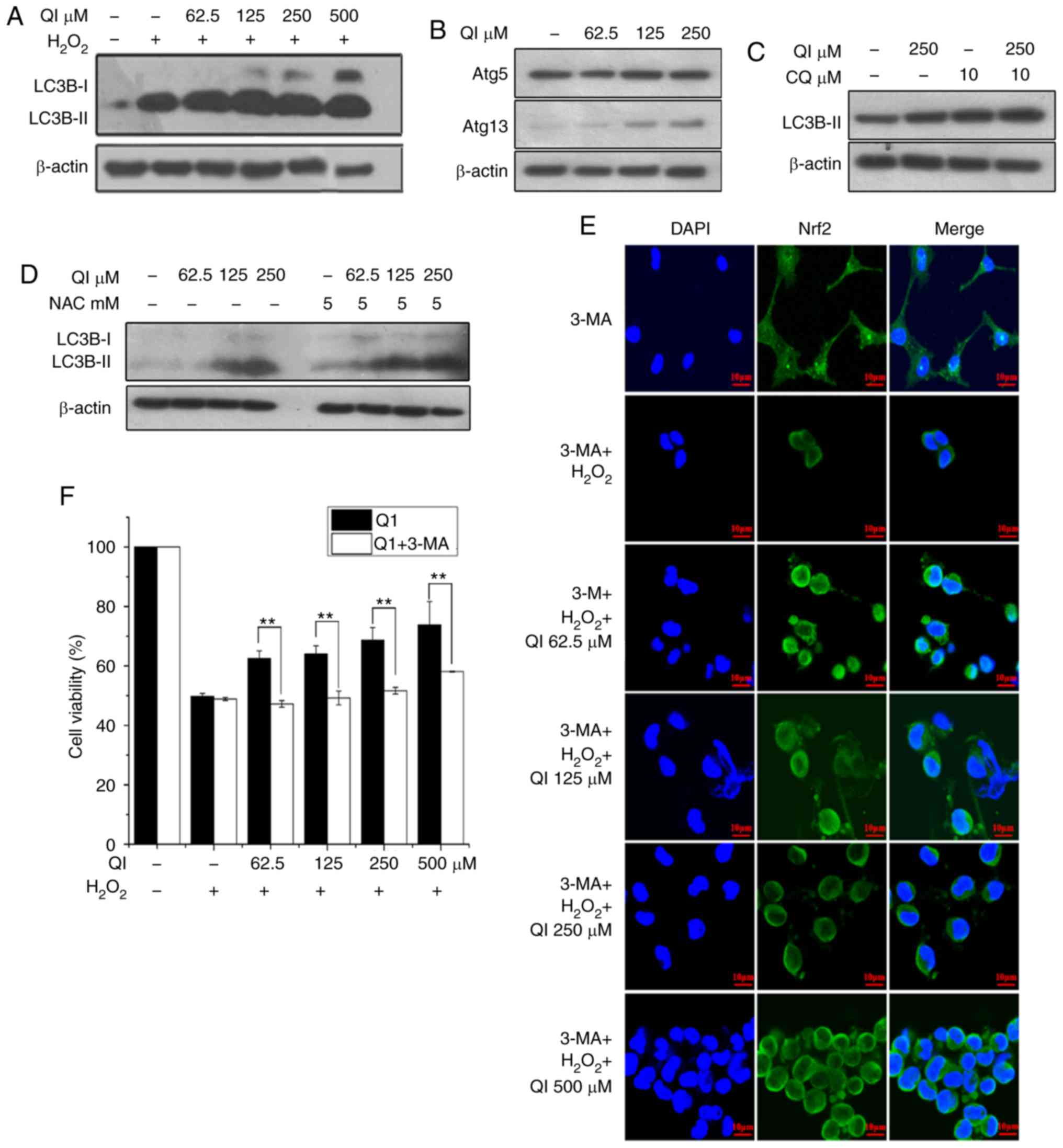 | Figure 7QI protects human umbilical vein
endothelial cells from H2O2-induced
cytotoxicity by evoking autophagy-induced Nrf2 nuclear
translocation. (A) LC3B-I/LC3B-II and (B) Atg5/Atg13 protein levels
examined by western blot analysis to determine the effect of QI on
autophagy. LC3B-II levels were also examined by western blot
analysis in (C) cells that were separately treated with CQ (10
µM) or QI (250 µM), or cotreated with both for 3 h,
or (D) cells that were treated with QI (62.5, 125 and 250
µM) alone or with NAC (5 mM) pre-treatment for 1 h. (E)
Confocal images obtained by immunohistochemical assay (blue, DAPI;
green, Nrf2; scale bar, 10 µm), demonstrating the Nrf2
nuclear transfer following pretreatment with 3-MA (an autophagy
inhibitor). (F) Cell viability determined by MTT assay in cells
pretreated with 3-MA, followed by QI treatment. All assays were
performed in triplicate, and data represent the mean ± standard
deviation. **P<0.01, as indicated. QI,
Quercetin-3-O-α-L-rhamnopyranoside; Nrf2, nuclear factor erythroid
2-related factor 2; Atg, autophagy related; 3-MA, 3-methyladenine;
CQ, chloroquine; NAC, N-acetyl-L-cysteine. |
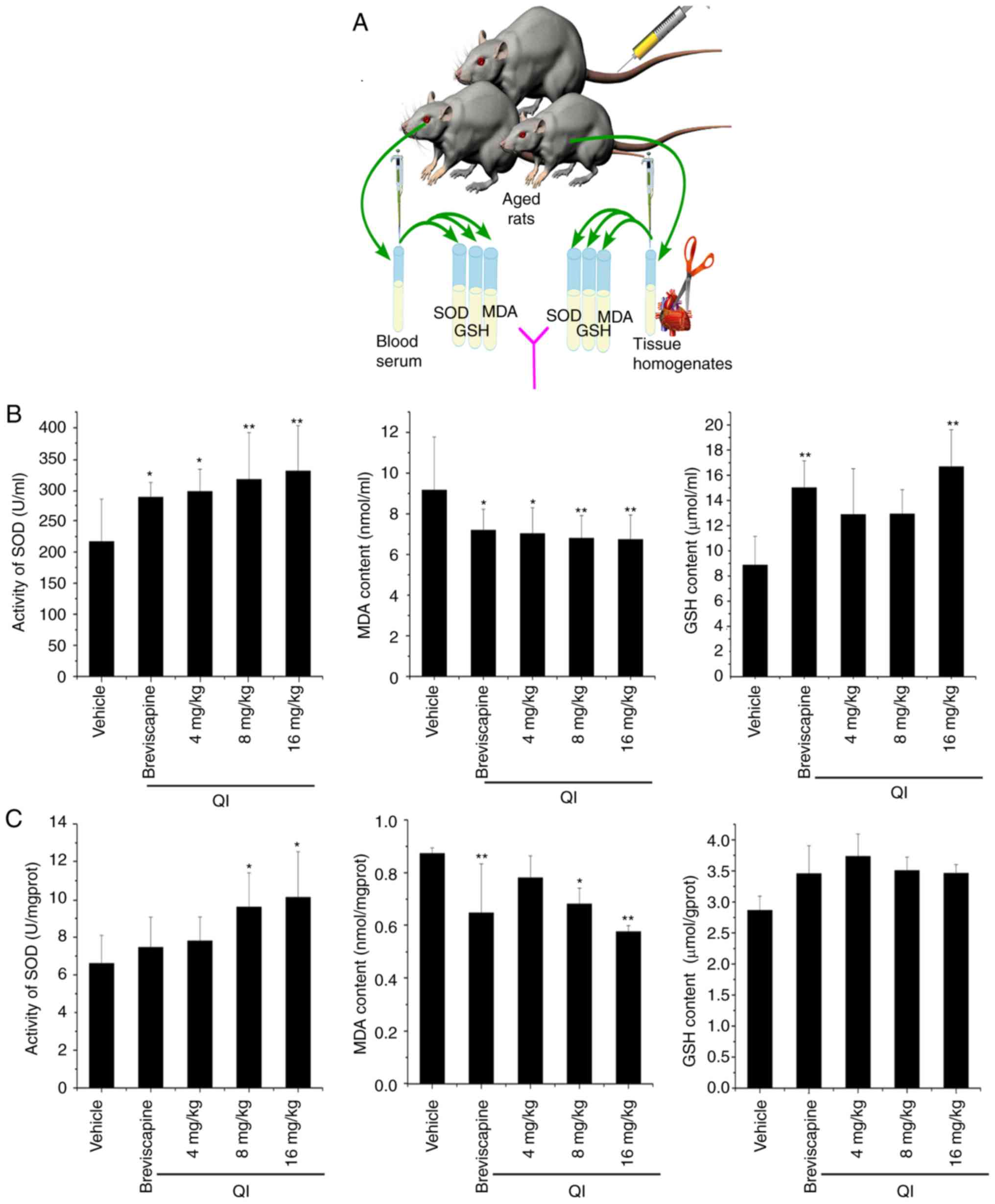
QI improves the activities of SOD and
GSH, and decreases the content of MDA in the serum and heart tissue
of rats
Next, rats were used to assess the antioxidant
ability of QI in vivo(Fig.
8A). Our preliminary experimental results demonstrated that QI
has no toxic or side effects on rats when administered at a dose of
0-100 mg/kg once daily for 2 weeks by tail vein injection (data not
shown). In addition, the pre-experimental SOD vitality test results
revealed that the lowest effective dose was 4 mg/kg (data not
shown); therefore, the concentrations of 4, 8 and 16 mg/kg were
selected for subsequent experiments. QI treatment displayed a
dose-dependent reduction in MDA levels and increase in SOD
activity. This effect reached a maximum level at the highest tested
dosage (16 mg/kg) in the serum and heart tissue of rats (Fig. 8B and C). In the serum, QI at doses
of 4, 8 and 16 mg/kg significantly enhanced GSH content, whereas
only the 16 mg/kg dose was able to increase the GSH content in the
heart tissue. These results indicate that QI achieves an
antioxidant capacity in aging rats primarily through improving SOD,
scavenging MDA and moderately increasing GSH.
Discussion
Endothelial dysfunction has been implicated in the
initiation and propagation of vascular diseases, including
atherosclerosis, hypertension, cardiac hypertrophy and congestive
heart failure. The oxidative stress induced by ROS, such as
superoxide, H2O2 and peroxynitrite, is a key
cause of endothelial cell dysfunction (23). The main chemical constituents in
the leaves of Lindera aggregata (Sims) Kosterm. are
flavonoids, which are known to have a favorable effect on the
prevention and treatment of cardiovascular disease. In the present
study, the antioxidant activity of QI derived from the leaves of
Lindera aggregata (Sims) Kosterm. was investigated in
vitro and in vivo. The results revealed that in QI
treatment, autophagy serves an important role in regulating the
Nrf2 pathway to protect HUVECs from damage caused by oxidative
stress.
Oxidative metabolites are involved in the
functional inactivation of endothelial cells by increasing cell
permeability and as potent inducers of endothelial cell death. The
MDA level reflects the extent of cell damage induced by oxidative
stress (24). An oxidative stress
model was established in the present study, according to the
results of preliminary experiments, which identified that a
concentration of 1,400 µM H2O2
resulted in a cell viability of ~50%. The antioxidant enzyme SOD
and reducing compound GSH are considered to augment the antioxidant
defenses in endothelial cells. SOD scavenges superoxide radicals by
converting them to hydrogen peroxide, which is then converted to
water by catalase and GSH peroxidase (25). The results of the present study
revealed that QI alleviated the H2O2-induced
decrease in SOD activity and GSH content in HUVECs, suggesting that
the cytoprotective effect of QI is likely associated with the
restoration of endogenous antioxidation and the decrease of lipid
peroxidation. This effect was also verified in aging rats, which
were considered as a suitable model in the current study since the
amount and activity of antioxidant enzymes in the body decrease
with age, causing an increase in reactive oxygen and thus leading
to a decline in physical function and various diseases. Aging rats
undergoing QI treatment displayed improved SOD activity and greater
GSH levels in the sera and heart tissues, as compared with the
vehicle control. The MDA concentration was also reduced in
QI-treated cells and rats, confirming that QI improves the ability
of endogenous antioxidation.
Excessive production of ROS in cells can directly
and indirectly cause mitochondrial dysfunction, apoptosis and cell
death (26). Nrf2 target genes
are involved in the elimination of ROS (27). Therefore, Nrf2 is a potent
transcriptional activator that serves a central role in the
expression of several cytoprotective genes in response to oxidative
stress (4). It was recently
reported that the autophagy pathway maintains the integrity of the
Keap1/Nrf2 pathway for normal liver function by governing Keap1
turnover and that Nrf2 accumulation is the dominant cause of liver
damage in autophagy-deficient mice (28). Additionally, autophagy is an
effective protector of the inner ear against oxidative stress
(8). To elucidate the underlying
mechanism of these antioxidant effects, the expression levels of
relevant proteins were measured in
H2O2-damaged HUVECs following QI treatment in
the present study. Treatment with QI inhibited the activation of
Caspase-9, cleaved Caspase-3, and cleaved PARP, demonstrating that
QI protects HUVECs from damage resulting from
H2O2-mediated apoptosis. Immunofluorescence
imaging and western blot analysis demonstrated the transfer of Nrf2
protein from the cytoplasm to the nucleus following QI treatment.
Furthermore, transfection of cells with Nrf2 siRNA resulted in
attenuation of the cytoprotective effect of QI. It is, thus,
speculated that Nrf2 regulation is the critical cause of the
antioxidant properties of QI.
Atg5 is E3-like activity for Atg8s-lipidaton and is
required for autophagosome formation, while Atg13 is a member of
the Atg1-Atg13-Atg17-Atg29 complex and is critical for correct
localization of ULK1 to the pre-autophagosome and stability of ULK1
protein (29). Following QI
treatment in the current study, Atg5, Atg13 and LC3B-II levels were
upregulated, suggesting that QI promotes autophagy. This was
further confirmed using 3-MA to study whether autophagy was
involved in the protective mechanism. It was observed that
pretreatment with 3-MA attenuated the protective effect of QI and
inhibited the nuclear transfer of Nrf2, suggesting that QI protects
HUVECs and activates the Nrf2 pathway by inducing autophagy.
Further analysis demonstrated that the antioxidant effect of QI
also occurs at the transcriptional level.
In conclusion, the current study provided evidence
that QI elicits significant autophagy by Nrf2 activation and
Nrf2-dependent induction of major cellular antioxidant enzymes, and
effectively attenuates H2O2-induced oxidative
stress in HUVECs (Fig. 9). These
observations demonstrate the importance of the QI/autophagy/Nrf2
axis in the cellular response to oxidative stress-induced cell
death and suggest that this axis may be a potential therapeutic
target for oxidative stress-induced diseases.
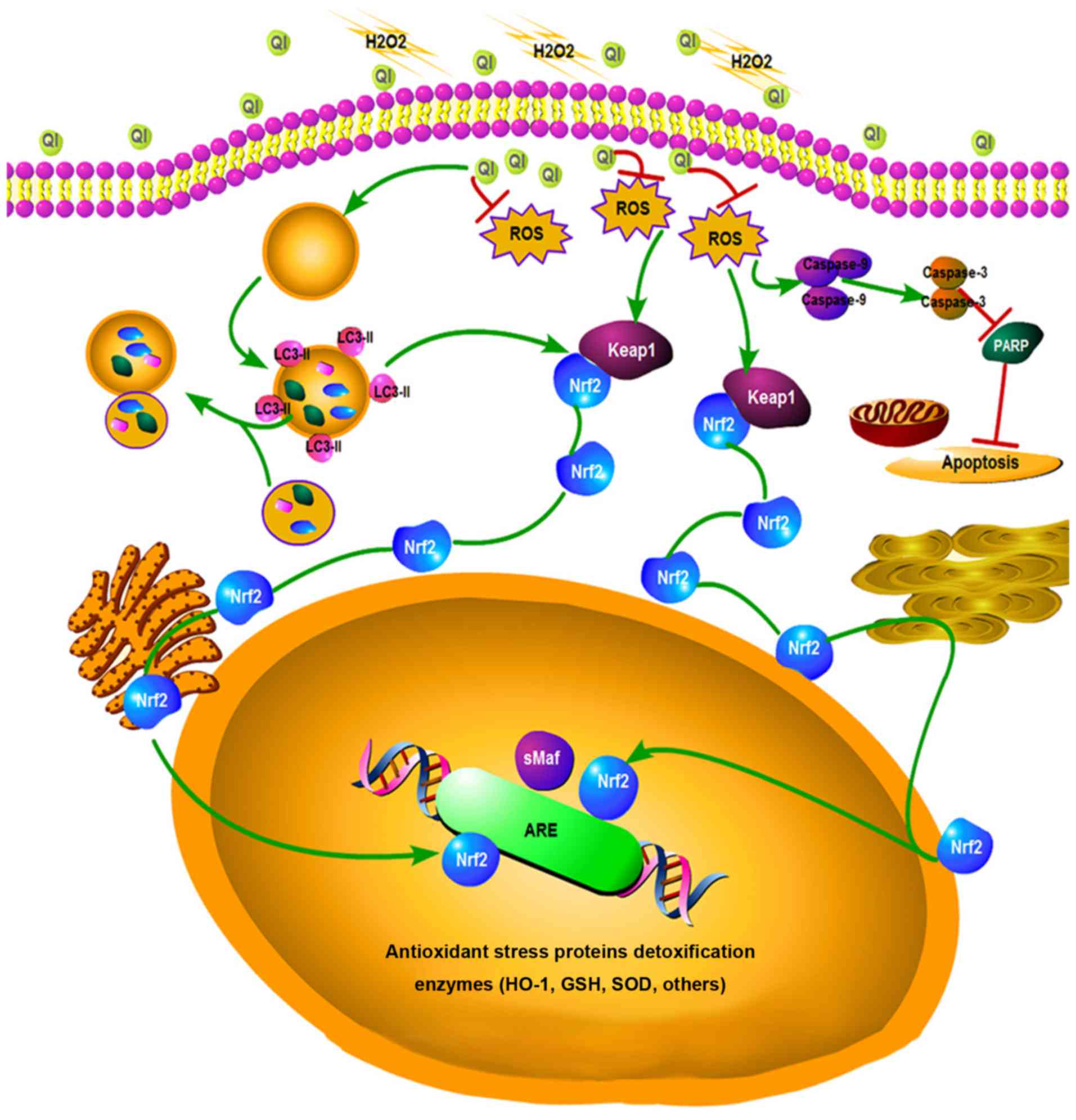 | Figure 9Diagram of the whole pathway. QI
elicits significant autophagy for Nrf2 activation and
Nrf2-dependent induction of major cellular antioxidant enzymes, and
effectively attenuates H2O2-induced oxidative
stress in human umbilical vein endothelial cells. QI,
Quercetin-3-O-α-L-rhamnopyranoside; Nrf2, nuclear factor erythroid
2-related factor 2; ROS, reactive oxygen species; LC, light chain;
Keap1, Kelch-like ECH-associated protein 1; PARP, poly(ADP-ribose)
polymerase; ARE, antioxidant response element; HO-1, heme
oxygenase-1; GSH, glutathione; SOD, superoxide dismutase. |
Acknowledgments
Not applicable.
Funding
The present study was supported by Zhejiang
Provincial Science and Technology Planning Project (grant no.
2016C04005), the Jiangsu Provincial Natural Science Foundation of
China (grant no. BK20161269) and the National Science Foundation of
China (grant no. 81473182).
Availability of data and materials
The datasets used and/or analyzed during the
current study are available from the corresponding author on
reasonable request.
Authors’ contributions
HH and LZ designed the study. HH, BX and AA
performed the in vitro experiments. HH, XY, HL and MG
conducted the in vivo experiments. HH and AA analyzed the
data and wrote the manuscript. All authors read and approved the
final manuscript.
Ethics approval and consent to
participate
All experimental procedures using live animals were
conducted in accordance with protocols approved by the Ethics
Review Committee for the Use of Animal Subjects of Zhejiang
University (Zhejiang, China).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Finkel T and Holbrook NJ: Oxidants,
oxidative stress and biology of ageing. Nature. 408:239–247. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Rahman T, Hosen I, Islam MM and Shekhar
HU: Oxidative stress and human health. Adv Biosci Biotechnol.
3:997–1019. 2012. View Article : Google Scholar
|
|
3
|
Birben E, Sahiner UM, Sackesen C, Erzurum
S and Kalayci O: Oxidative stress and antioxidant defence. World
Allergy Organ J. 5:9–19. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Motohashi H and Yamamoto M: Nrf2-Keap1
defines a physiologically important stress response mechanism.
Trends Mol Med. 10:549–557. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Dhakshinamoorthy S, Jain AK, Bloom DA and
Jaiswal AK: Bach1 competes with Nrf2 leading to negative regulation
of the antioxidant response element (ARE)-mediated NAD(P)H: Quinone
oxidoreductase 1 gene expression and induction in response to
antioxidants. J Biol Chem. 280:16891–16900. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Chen S, Tang Y, Qian Y, Chen R, Zhang L,
Wo L and Chai H: Allicin prevents
H2O2-induced apoptosis of HUVECs by
inhibiting an oxidative stress pathway. BMC Complement Altern Med.
14:3212014. View Article : Google Scholar
|
|
7
|
Maiuri MC, Zalckvar E, Kimchi A and
Kroemer G: Self-eating and self-killing: Crosstalk between
autophagy and apoptosis. Nat Rev Mol Cell Biol. 8:741–752. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Hayashi K, Dan K, Goto F, Tshuchihashi N,
Nomura Y, Fujioka M, Kanzaki S and Ogawa K: The autophagy pathway
maintained signaling crosstalk with the Keap1-Nrf2 system through
p62 in auditory cells under oxidative stress. Cell Signal.
27:382–393. 2015. View Article : Google Scholar
|
|
9
|
Kim JA, Jung YS, Kim MY, Yang SY, Lee S
and Kim YH: Protective effect of components isolated from
Linderaerythrocarpa against oxidative stress-induced apoptosis of
H9c2 cardiomyocytes. Phytother Res. 25:1612–1617. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Yu YB, Miyashiro H, Nakamura N, Hattori M
and Park JC: Effects of triterpenoids and flavonoids isolated from
Alnus firma on HIV-1 viral enzymes. Arch Pharm Res. 7:820–826.
2007. View Article : Google Scholar
|
|
11
|
Boppana K, Dubey PK, Jagarlapudi SA,
Vadivelan S and Rambabu G: Knowledge based identification of MAO-B
selective inhibitors using pharmacophore and structure based
virtual screening models. Eur J Med Chem. 44:3584–3590. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Jang DS, Kim JM, Kim J, Yoo JL, Kim YS and
Kim JS: Effects of compounds isolated from the fruits of Rumex
japonicas on the protein glycation. Chem Biodivers. 5:2718–2723.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Choi B, Bae JY, Kim DS, Li J, Kim JL, Lee
YJ and Kang YH: Dietary compound quercitrindampents VEGF induction
and PPAR gamma activation in oxidized LDL-exposed murine
macrophages: Association with scavenger receptor CD36. J Agric Food
Chem. 58:1333–1341. 2010. View Article : Google Scholar
|
|
14
|
Hubert DJ, Dawe A, Florence NT, Gilbert
KD, Angele TN, Buonocore D, Finzi PV, Vidari G, Bonaventure NT,
Marzatico F and Paul MF: In vitro hepatoprotective and antioxidant
activities of crude extract and isolated compounds from
Ficusgnaphalocarpa. Inflammonpharmacology. 19:35–43. 2011.
View Article : Google Scholar
|
|
15
|
Zhi K, Li M, Bai J, Wu Y, Zhou S, Zhang X
and Qu L: Quercitrin treatment protects endothelial progenitor
cells from oxida-tive damage via inducing autophagy through
extracellular signal-regulated kinase. Angiogenesis. 19:311–324.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Xu C, Yang B, Zhu W, Li X, Tian J and
Zhang L: Characterisation of polyphenol constituents of Linderae
aggregata leaves using HPLC fingerprint analysis and their
antioxidant activities. Food Chem. 186:83–89. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ning L, Wentworth L, Chen H and Weber SM:
Down-regulation of Notch1 signaling inhibits tumor growth in human
hepatocellular carcinoma. Am J Transl Res. 1:358–366.
2009.PubMed/NCBI
|
|
18
|
Choi HI, Kim HJ, Park JS, Kim IJ, Bae EH,
Ma SK and Kim SW: PGC-1α attenuates hydrogen peroxide-induced
apoptotic cell death by upregulating Nrf-2 via GSK3β inactivation
mediated by activated p38 in HK-2 cells. Sci Rep. 7:4319–4331.
2017. View Article : Google Scholar
|
|
19
|
Li W, Ma F, Zhang L, Huang Y, Li X, Zhang
A, Hou C and Zhu Y and Zhu Y: S-Propargyl-cysteine exerts a novel
protective effect on methionine and choline deficient diet-induced
fatty liver via Akt/Nrf2/HO-1 pathway. Oxid Med Cell Longev.
2016:46908572016. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
21
|
Li XL, Li YQ, Yan WM, Li HY, Xu H, Zheng
XX, Guo DW and Tang LK: A study of the cardioprotective effect of
breviscapine during hypoxia of cardiomyocytes in vitro and during
myocardial infarction in vivo. Planta Medica. 70:1039–1044. 2004.
View Article : Google Scholar
|
|
22
|
Gao Y, Chu S, Shao Q, Zhang M, Xia C, Wang
Y, Li Y, Lou Y, Huang H and Chen N: Antioxidant activities of
ginsenoside Rg1 against cisplatin-induced hepatic injury through
Nrf2 signaling pathway in mice. Free Radic Res Commun. 51:1–13.
2017. View Article : Google Scholar
|
|
23
|
Hadi H, Carr CS and Suwadi JA: Endothelial
dysfunction: Cardiovascular risk factors, therapy, and outcome.
Vasc Health Risk Manag. 1:183–198. 2005.
|
|
24
|
Lum H and Roebuck KA: Oxidant stress and
endothelial cell dysfunction. Am J Physiol Cell Physiol.
280:C719–C741. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zhai L, Zhang P, Sun RY, Liu XY, Liu WG
and Guo XL: Cytoprotective effects of CSTMP, a novel stilbene
derivative, against H2O2-induced oxidative
stress in human endothelial cells. Pharmacol Rep. 63:1469–1480.
2011. View Article : Google Scholar
|
|
26
|
Kir HM, Dillioglugil MO, Tugay M,
Eraldemir C and Özdoğan HK: Effects of vitamins E, A and D on MDA,
GSH, NO levels and SOD activities in 5/6 nephrectomized rats. Am J
Nephrol. 25:441–446. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Sugawara T and Chan PH: Reactive oxygen
radicals and pathogenesis of neuronal death after cerebral
ischemia. Antioxid Redox Signal. 5:597–607. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Yates MS, Tran QT, Dolan PM, Osburn WO,
Shin SN, McCulloch CC, Silkworth JB, Taguchi K, Yamamoto M,
Williams CR, et al: Genetic versus chemoprotective activation of
Nrf2 signaling: Overlapping yet distinct gene expression profiles
between Keap1 knockout and triterpenoid-treated mice.
Carcinogenesis. 30:1024–1031. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Shibutani ST and Yoshimori T: A current
perspective of autophagosome biogenesis. Cell Res. 24:58–68. 2014.
View Article : Google Scholar :
|