Introduction
Lung cancer accounts for the highest proportion
(~25%) of all cancer-associated mortalities worldwide, and has a
mortality rate of >15% (1). Of
all lung cancer cases, >85% are non-small cell lung cancer
(NSCLC), which is associated with a high mortality rate (2,3).
At present, the most effective curative method for NSCLC is
surgical resection; however, surgical treatment has been
successfully applied to only a few patients, as NSCLC tends to be
diagnosed at advanced stages. Therefore, early diagnosis with
predictive biomarkers and therapeutic applications with prognostic
biomarkers are vital for improving the survival rate of patients
with NSCLC.
MicroRNAs (miRNAs or miRs) are a group of non-coding
RNAs (18-22 nucleotides in length) that play a role in the
post-translational regulation of cell proliferation,
differentiation and death, as well as other physiological and
pathological processes (4-7).
The role of miRNAs as tumor-inducers or -suppressors depends on
their targets in numerous types of cancer. As previously reported,
the dysregulation of miRNAs promotes tumor occurrence and
development (8). Accumulating
evidence supports the involvement of miR-340 in a variety of cancer
types (9-11). miR-340 modulates the growth,
migration and invasion of numerous types of cancer cells (12-15). miR-340 has been shown to suppress
the migration, invasion and metastasis of breast cancer cells via
the Wnt signaling pathway (16)
or by downregulating Rho kinase-1 (17,18); thus, miR-340 may be a key tumor
suppressor in the diagnosis and treatment of breast cancer. These
findings suggest that miR-340 may act as a suppressor of tumor
progression. A previous study however, reported the association
between miR-340 dysregulation and the development of NSCLC
(19). miR-340 functions as a
tumor suppressor by regulating the expression of cyclin-dependent
kinase-4 in NSCLC cells, inhibiting cell growth (20). In addition, it has been
demonstrated that miR-340 inhibits tumor cell proliferation and
induces apoptosis by targeting a variety of negative regulators of
p27 in NSCLC (21).
RABs, small G proteins of the Ras superfamily, serve
as regulators of vesicular transport in the exocytic and endocytic
pathways in eukaryotic cells (22). It has been demonstrated that RABs
play significant roles in endocytosis, cell secretion, growth and
signal transduction (23).
miR-30b/c has been shown to directly downregulate the expression of
RAB18 and to inhibit the proliferation of NSCLC cells, indicating
that miR-30b/c may serve as a tumor suppressor gene in the
pathogenesis of NSCLC (24). An
elevated expression of RAB27A has been shown to be associated with
NSCLC and resistance to conventional chemotherapeutic agents
(25); however, the link between
RAB27B and miR-340 in NSCLC remains unclear. Thus, the present
study aimed to investigate the function of RAB27B in association
with miR-340 in NSCLC.
The findings of the aforementioned studies indicate
that miR-340 may be a potential target in the treatment of NSCLC;
however, the molecular mechanisms of miR-340 in modulating the
expression of RAB27B in the occurrence and development of NSCLC
remain unclear. Thus, the present study aimed to investigate
miR-340 as a novel tumor suppressor in NSCLC.
Materials and methods
Cell culture
The human lung cancer cell lines A549, PC9 and
normal bronchial epithelial cells (BEAS-2B; American Type Culture
Collection) were cultured in Dulbecco's modified Eagle's medium
(DMEM; Gibco; Thermo Fisher Scientific, Inc.) containing 10% fetal
bovine serum (Gibco; Thermo Fisher Scientific, Inc.), 100 U/ml
penicillin and 100 g/ml streptomycin (Shanghai Xinyu Biotechnology
Pharmaceutical Co. Ltd.) at 37°C in 5% CO2.
Tissue collection
In total, 22 pairs of NSCLC and adjacent non-tumor
tissues were obtained from patients with NSCLC who were treated at
The Affiliated Hospital of Southwest Medical University between
2015 and 2018; samples were stored at −80°C. Written informed
consent was obtained from all patients. The present study was
approved by the Ethics Committee of the Affiliated Hospital of
Southwest Medical University.
Immunohistochemistry (IHC)
All tissues were immersed in 10% formalin and
embedded in paraffin. The expression of RAB family proteins,
including RAB27A, RAB27B, RAB21, RAB11A and RAB9A in the tissue
samples was detected by IHC. Upon deparaffinization with xylene and
rehydration with ethanol, the sections were washed with PBS twice
and immersed in H2O2 solution to block
endogenous peroxidase activity. Following washing with PBS, 10%
goat serum was added for 1 h at room temperature to block
nonspecific reactions. The tissues were then incubated overnight at
4°C with anti-RAB27A (dilution 1:200; cat. no. sc-74586; Santa Cruz
Biotechnology, Inc.), anti-RAB27B (dilution 1:200; cat. no.
DF12060; Affinity Biosciences), anti-RAB21 (dilution 1:200; cat.
no. sc-81917; Santa Cruz Biotechnology, Inc.), anti-RAB11A
(dilution 1:200; cat. no. sc-166912; Santa Cruz Biotechnology,
Inc.) and anti-RAB9A (dilution 1:200; cat. no. sc-71950; Santa Cruz
Biotechnology, Inc.) antibodies. Subsequently, the sections were
washed with PBS twice and incubated with a horseradish peroxidase
(HRP)-conjugated secondary antibody (dilution 1:200; cat. no.
sc-2347; Santa Cruz Biotechnology, Inc.) at 37°C for 30 min.
Finally, the sections were incubated with 3,3′-diaminobenzidine to
detect proteins. The samples were visualized under a light
microscope (Nikon Corp.).
Cell transfection
The BEAS-2B, A549 and PC9 cells were seeded into
6-well plates at a density of 5×104 cells/well and
transfected with miR-340 mimics, miR-340 inhibitors or mock miRNA
(Invitrogen; Thermo Fisher Scientific, Inc.) using
Lipofectamine® 2000 (Invitrogen; Thermo Fisher
Scientific, Inc.) according to the manufacturer's protocols. Total
protein was isolated from the transfected cells with protein
extraction kit (KeyGen Biotech) according to the manufacturer's
instructions at 4°C, then total protein was quantified using a BCA
assay (KeyGen Biotech) according to the manufacturer's instructions
at 4°C. Total RNA was extracted from the transfected cells using
TRIzol reagent (Invitrogen; Thermo Fisher Scientific, Inc.) and
quantified using NanoDrop@ One/One C Micro-UV-Vis
Spectrophotometer (Thermo Fisher Scientific, Inc.). Total protein
and RNA was used for further analysis. The sequences of miR-340
mimics and miR-340 inhibitors were as follows: miR-340 mimics,
UUAUAAAGCAAUGAGACUGAUU and miR-340 inhibitors,
AAUCAGUCUCAUUGAUUUAUAA.
Cell Counting kit-8 (CCK-8) assay
The proliferation of the transfected BEAS-2B, A549
and PC9 cells was assessed using a CCK-8 assay (Sigma-Aldrich;
Merck KGaA). Briefly, the cells were seeded into 24-well plates and
cultured for 24 h, and then transfected with miR-340 mimics,
miR-340 inhibitors or mock miRNA. Subsequently, 100 µl
transfected cells were added to 96-well plates at a density of
2×104 cells/well, and 10 µl CCK-8 solution were
added to each well, followed by 2 h of incubation at 37°C. Viable
cells were counted with BioTek Epoch microplate absorbance reader
(Bio-Tek Instruments, Inc.) at 450 nm.
Luciferase reporter assays
Potential targets of miR-340 were predicted using
TargetScan (www.targetscan.org/vert_71/) and further confirmed by
MicroRNA.org30 (www.microrna.org), and miRBase (http://www.mirbase.org/). To further confirm whether
RAB27B is a direct target of miR-340, luciferase reporter
experiments were performed. The cells plated into 24-well plates at
40% confluence were co-transfected for 24 h with miR-340 mimics,
miR-340 inhibitors or mock miRNA and luciferase reporter plasmids
[pGL3 RAB27B 3′-untranslated region (UTR) wild-type or mutant;
Genepharm, Inc.] using Lipofectamine 2000 according to the
manufacturer's instructions. Luciferase activity was measured 24 h
later with the Dual-Luciferase® Assay (Promega Corp.).
Lightswitch Renilla activity was normalized to pGL3 to
assess transfection efficiency. Each assay was conducted in
triplicate.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
Total RNA was isolated from NSCLC tissues, non-tumor
tissues and cell lines using TRIzol reagent for RT-qPCR analysis.
miRNA or mRNA expression was evaluated by two-step RT-qPCR. cDNA
was prepared using A-MLV reverse transcriptase (Invitrogen; Thermo
Fisher Scientific, Inc.); qPCR with SYBR-Green dye was performed
according to the manufacturer's protocols (Takara Inc.). For qPCR,
cDNA (0.5 µl), forward primer (0.5 µl), reverse
primer (0.5 µl), deoxyribonucleotide triphosphate mixture (2
µl, 2.5 mM), DNA polymerase (0.5 µl, 10 U/µl),
5X buffer (6 µl) and double distilled H2O (15
µl) were subjected to 32 cycles at 94°C for 5 min, 95°C for
15 sec and 56°C for 35 sec. Each sample was evaluated in duplicate,
and the mean quantification cycle (Cq) was calculated.
The following forward and reverse primers were used:
5′-GCGCTAGTTTCCTGT-3′ and 5′-GTGCAGGGTCCGAGGT-3′ (miR-340);
5′-TGCGGGACAAGAGCGGTTCCG-3′ and 5′-GCCAGTTCCCGAGCTTGCCGTT-3′
(RAB27B); and 5′-CAATGACCCCTTCATTGACC-3′ and
5′-GACAAGCTTCCCGTTCTCAG-3′ (GAPDH). PCR amplification of miRNA was
conducted using miRNA-specific forward primers and the universal
poly(T) adaptor reverse primer with U6 as an internal control. The
forward sequence of Homo sapiens miR-340 was
5′-GCTTATAAAGCAATGAGACTGATT-3′. U6 was used as an internal control
(forward, 5′-CTCGCTTCGGCAGCACA-3′ and reverse,
5′-AACGCTTCACGAATTTGCGT-3′). The results were quantified using the
2-ΔΔCq method as previously described (26).
Transwell assay
The effects of miR-340 on A549 cell invasion and
migration were analyzed using Transwell assays (Costar, Aiyan
Biotechnology Co. Ltd.) with or without Matrigel (Clontech,
Laboratories, Inc.), respectively. The A549 cells transfected for
24 h were resuspended in the upper 24-well chambers at a density of
1×105 cells/well. Serum-free DMEM and 10% FBS-DMEM were
added to the upper and lower chambers, respectively. After 24 h,
cells on the upper membrane were collected using cotton swabs,
while cells in the lower chamber were fixed with 100% methanol for
15 min and stained with 0.05% crystal violet (Beyotime
Biotechnology Company) for 20 min at 37°C. Invasive and migrated
cells were observed under a inverted Fluorescence Microscope IX53
(Olympus). Each assay was conducted in triplicate.
Western blot analysis
Total proteins were extracted from tissues and
transfected cells using lysis buffer, and the protein concentration
was determined using a BCA protein assay kit. Proteins (20
µg) were subjected to 10% SDS-PAGE and transferred onto
polyvinylidene difluoride membranes. The membranes were blocked
with 2% bovine serum albumin and cultured with primary antibodies
against RAB27A (dilution 1:600; cat. no. sc-74586; Santa Cruz
Biotechnology, Inc.), RAB27B (dilution 1:800; cat. no. DF12060;
Affinity Biosciences), RAB21 (dilution 1:1,000; cat. no. sc-81917;
Santa Cruz Biotechnology, Inc.), RAB11A (dilution 1:1,200; cat. no.
sc-166912; Santa Cruz Biotechnology, Inc.) and RAB9A (dilution
1:1,000; cat. no. sc-71950; Santa Cruz Biotechnology, Inc.) for 60
min at 37°C. Following 3 washes with TBST, the membranes were
incubated with goat anti-rabbit IgG-HRP secondary antibody
(dilution 1:1,000; cat. no. sc-2004; Santa Cruz Biotechnology,
Inc.) at room temperature for 30 min and then washed as
aforementioned. Subsequently, specific binding was detected with
the chemiluminescence (GE Healthcare Life Sciences). The detection
of the chemiluminescent signal was performed in the gel
documentation system ImageQuant LAS 4000 Mini (GE Healthcare Life
Sciences). The intensity of the bands corresponding to the target
proteins was analyzed using ImageJ 1.8.0 (National Institutes of
Health).
Statistical analysis
Data were analyzed with SPSS 19.0 (IBM Corp.,
Armonk, NY, USA) and expressed as the means ± standard error of the
mean. Differences between groups were compared by one-way analysis
of variance followed by a Tukey's post-hoc test. P<0.05 was
considered to indicate a statistically significant difference. Each
experiment was performed at least 3 times.
Results
miR-340 expression is downregulated in
NSCLC tissues and cell lines
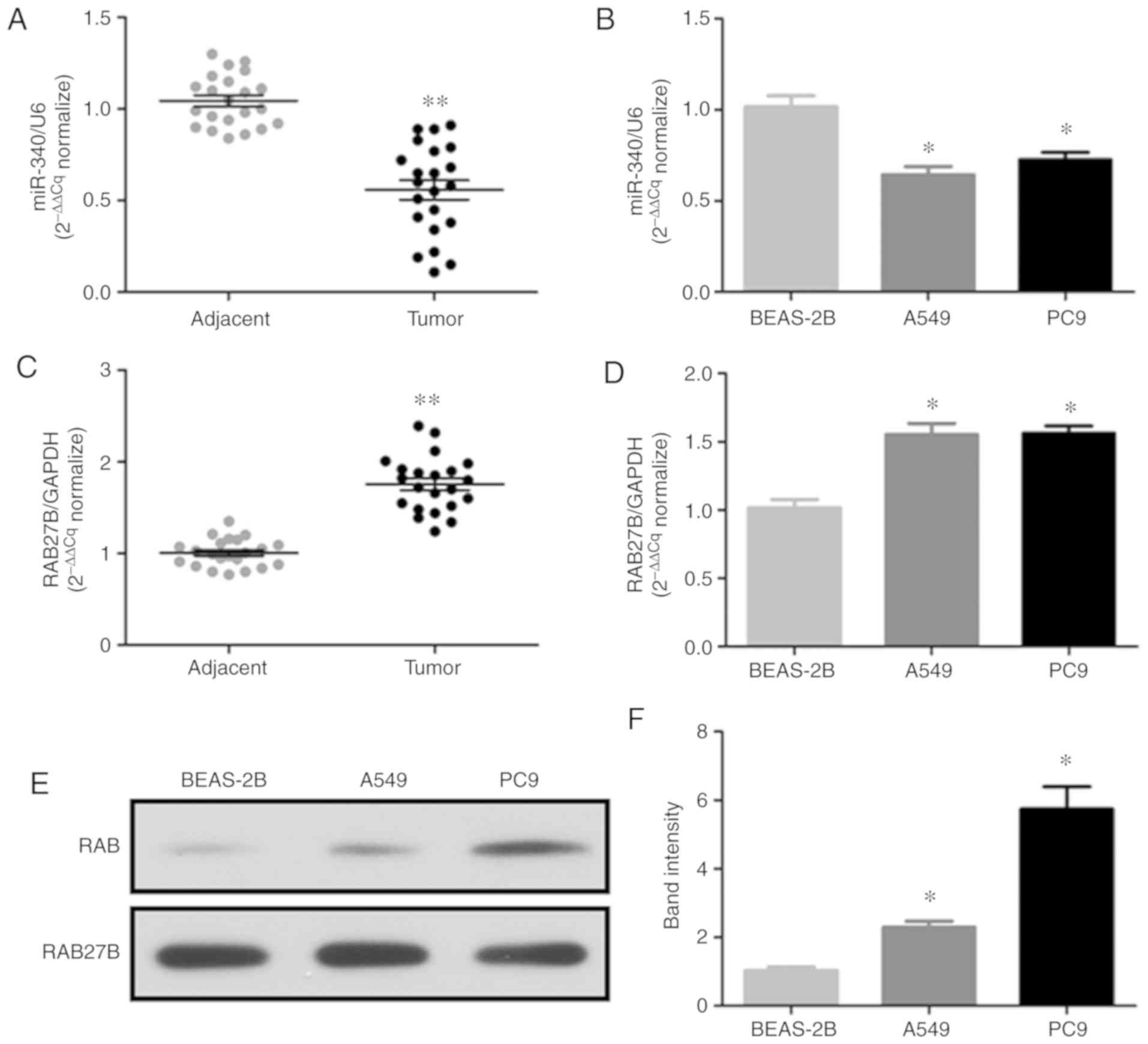
The present study analyzed the expression levels of
miR-340 and RAB27B in 22 pairs of NSCLC tissues, adjacent tissues
and nornal BEAS-2B cells or NSCLC cell lines (A549 and PC9) by
RT-qPCR; the protein expression of RAB27B in the NSCLC cell lines
was determined by western blot analysis. The results revealed that
miR-340 expression was significantly decreased in NSCLC tissues
(Fig. 1A) and cell lines
(Fig. 1B). On the contrary, the
mRNA expression levels of RAB27B were significantly increased in
NSCLC tissues (Fig. 1C) and cell
lines (Fig. 1D), while RAB27B
protein expression was markedly upregulated in the NSCLC cell lines
(Fig. 1E and F). Thus, the
downregulation of miR-340 may be associated with the upregulation
of RAB27B in NSCLC.
Analysis of the RAB family of proteins in
NSCLC tissues
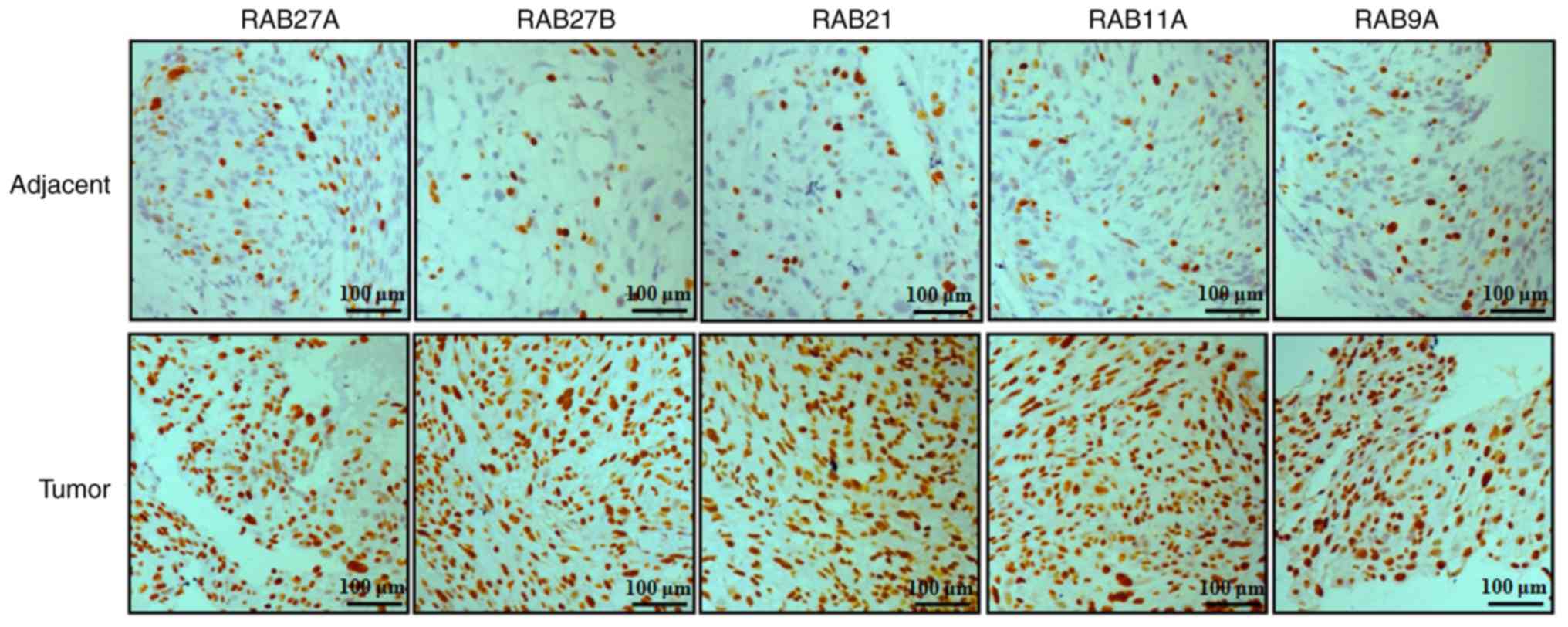
In order to examine the expression of RAB family
proteins in NSCLC tissues, the expression levels of 5 RAB proteins,
including RAB27A, RAB27B, RAB21, RAB11A and RAB9A, were evaluated
by immunohistochemical analysis in NSCLC and adjacent tissues
(Fig. 2). The results indicated
that the protein expression levels of RAB27A, RAB27B, RAB21, RAB11A
and RAB9A were notably increased in NSCLC tissues compared with
non-tumor tissues (Fig. 2).
miR-340 targets RAB27B and directly
decreases its expression
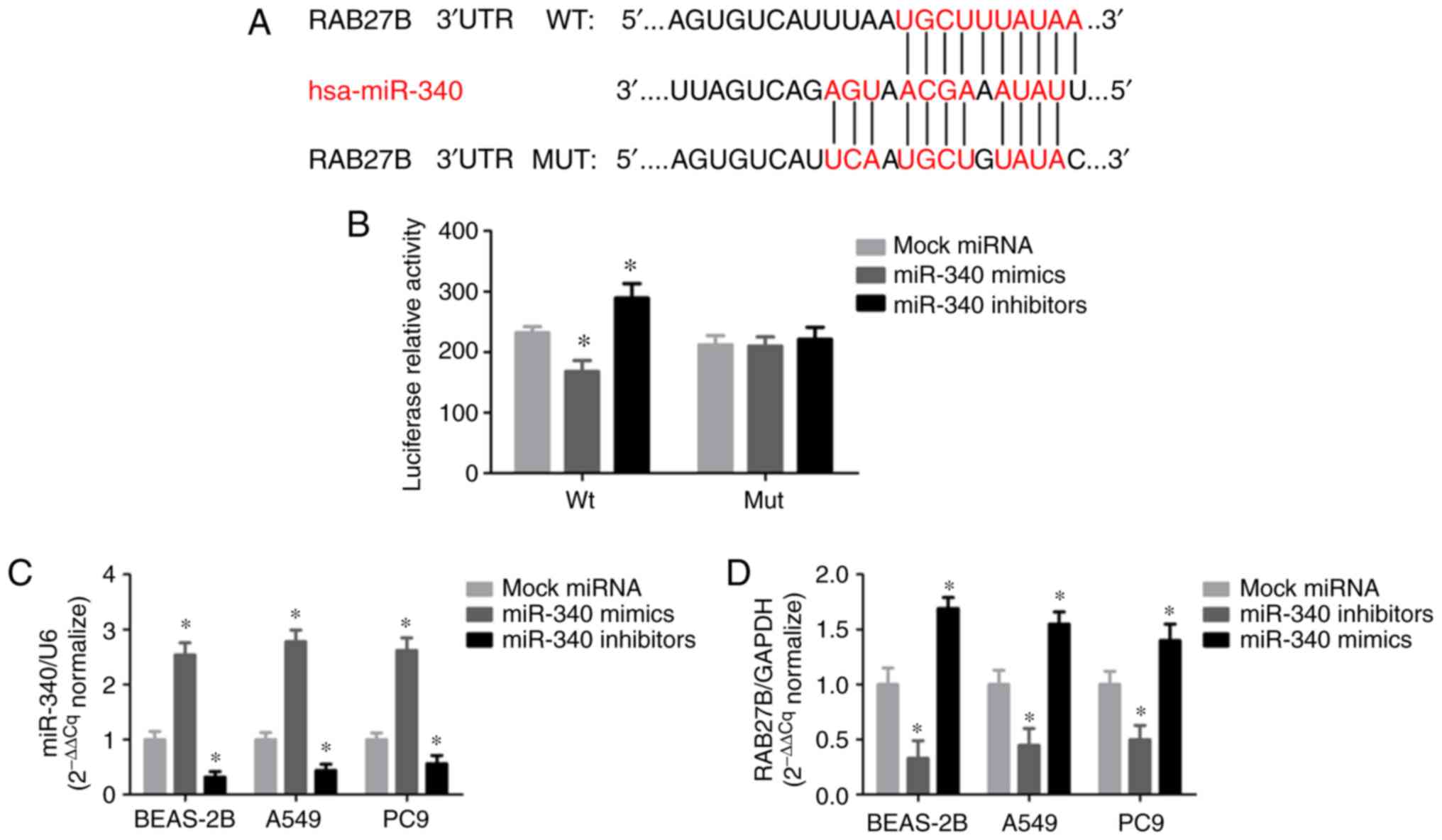
The present study revealed that RAB27B may be a
target of miR-340 using predictive tools, including TargetScan,
miRanda and miRBase. Of note, a putative binding site of miR-340
was identified in the 3′-UTR of RAB27B (Fig. 3A). The luciferase activity of the
RAB27B wild-type 3′-UTR reporter was markedly suppressed by
transfection with miR-340 mimics compared with the negative
control, while the mutant luciferase reporter was significantly
activated by transfection with miR-340 inhibitors (Fig. 3B). miR-340 expression was
significantly increased in all cell lines transfected with miR-340
mimics, whereas it was decreased in NSCLC cells transfected with
miR-340 inhibitors (Fig. 3C).
RAB27B mRNA expression was notably decreased following transfection
with miR-340 mimics; however, RAB27B expression in the A549 cells
was upregulated in response to the silencing of miR-340 with
miR-340 inhibitors (Fig. 3D).
These results indicate that miR-340 regulates the
post-transcriptional expression of RAB27B by directly binding to
its 3′-UTR.
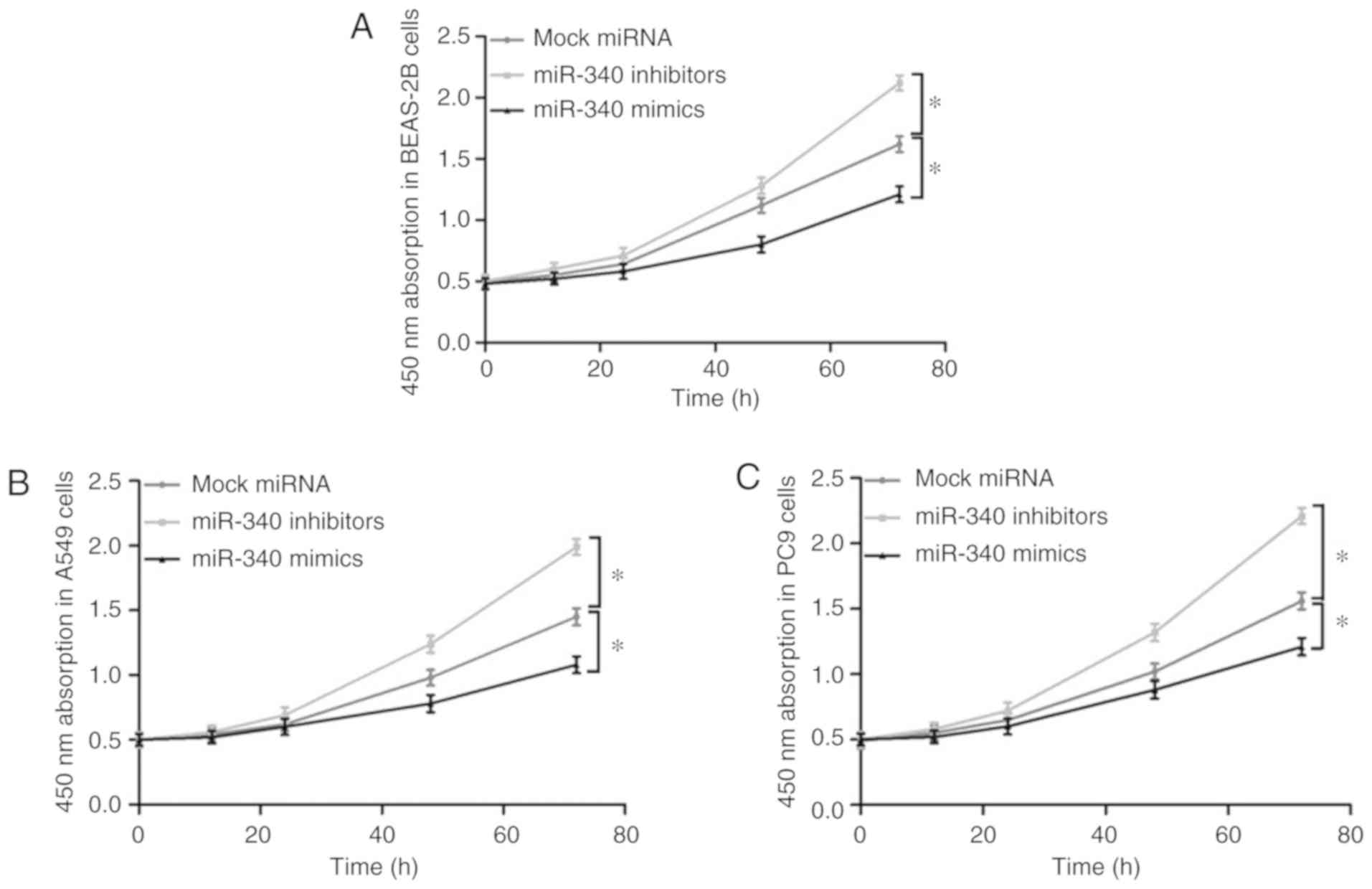
Effects of miR-340 overexpression on the
proliferation of NSCLC cell lines
The effects of miR-340 overexpression or knockdown
on the proliferation of the BEAS-2B, A549 and PC9 cells were
evaluated by a CKK-8 assay (Fig.
4). It was observed that transfection with miR-340 mimics
significantly suppressed the proliferative ability of these cell
lines; however, the cell proliferative ability was markedly
promoted following the knockdown of miR-340 with inhibitors.
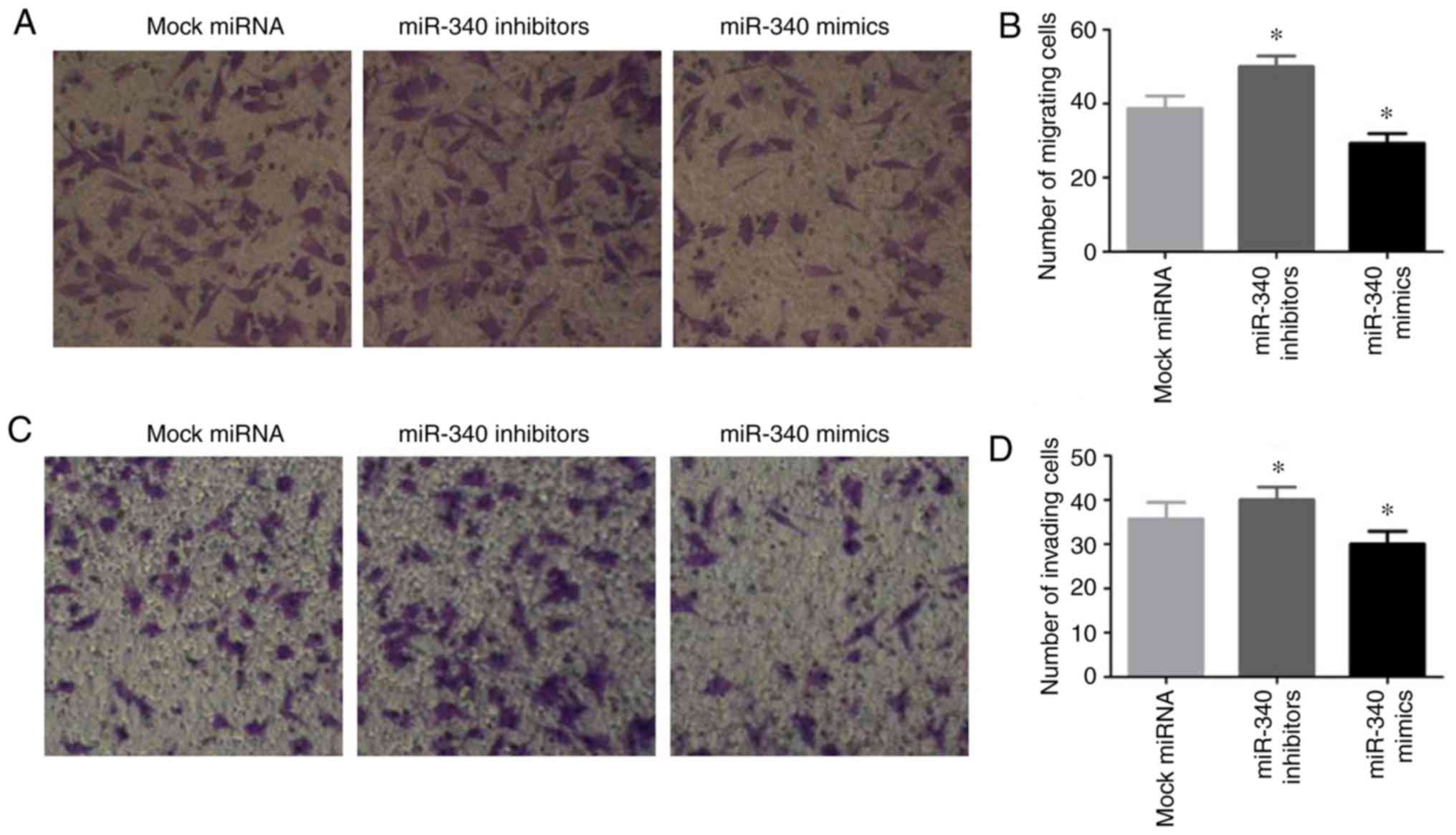
Effects of miR-340 on the migratory and
invasive abilities of NSCLC cells
To enhance our understanding of the biological
functions of miR-340 in NSCLC cells, the A549 cells were
transfected with miR-340 mimics or inhibitors, and the migratory
and invasive abilities of the cells were investigated by a
Transwell assay. The results revealed that transfection with
miR-340 mimics notably inhibited A549 cell migration, while
transfection with miR-340 inhibitors exerted opposite effects
(Fig. 5A and B). Furthermore,
transfection with miR-340 mimics suppressed cell invasion, and this
was promoted by transfection with miR-340 inhibitors (Fig. 5C and D).
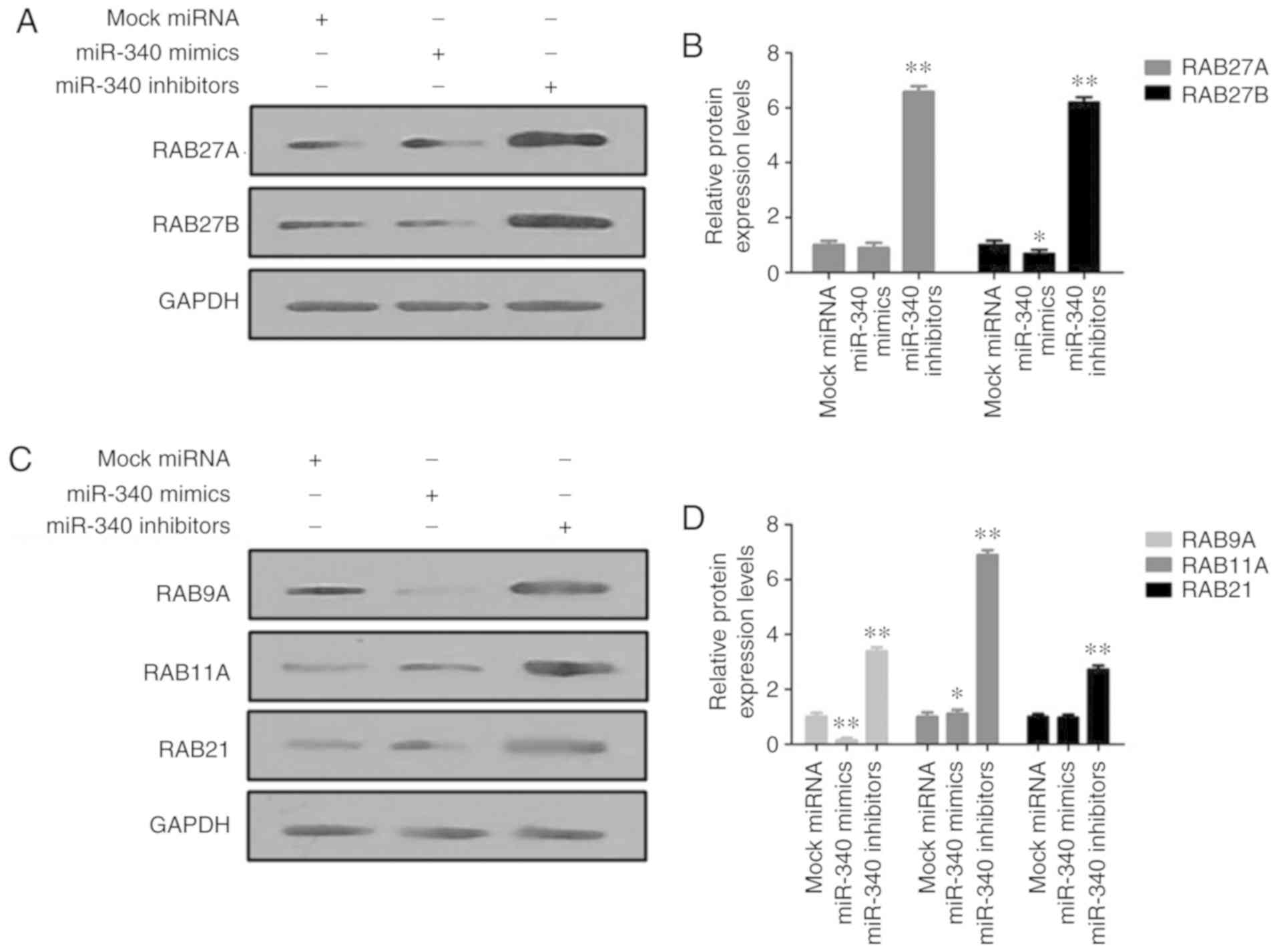
In vitro analysis of the effects of
miR-340 on RAB27 proteins
The protein expression levels of RAB27A and RAB27B
were markedly decreased in the miR-340 mimic-transfected A549
cells, but were increased upon transfection with miR-340 inhibitors
(Fig. 6A and B). In addition,
transfection with miR-340 mimics significantly suppressed the
expression levels of RAB9A, RAB11A and RAB21 in A549 cells, while
miR-340 inhibitors exerted opposite effects (Fig. 6C and D).
Discussion
It is well known that one miRNA can target numerous
genes. The function of miR-340 in the occurrence of various types
of cancer has been widely studied in recent years. As previously
reported, miR-340 restricts the development of breast cancer cells
by targeting numerous oncogenes, including c-Met, zinc finger
E-box-binding homeobox 1 and enhancer of zeste homolog 2 (27-29). Furthermore, miR-340 has been shown
to inhibit the progression of glioma by targeting cyclin-dependent
kinase 6, cyclin-D1/D2 and tissue plasminogen activator (30,31). In addition, miR-340 has been shown
to play a suppressive role in lung, colorectal (32), ovarian (33) and prostate cancers (34), laryngeal squamous cell carcinoma
(35) and osteosarcoma (36); however, the role of miR-340 in
NSCLC remains unknown. The present study reported that miR-340 was
downregulated in NSCLC, whereas miR-340 overexpression suppressed
the growth and migration of NSCLC cells. This suggests that miR-340
may play a tumor-suppressing role; however, a previous study
revealed that miR-340 played an oncogenic role in gastric cancer by
targeting cyclin G2 (37). The
biological function of miR-340 may depend on its target gene, which
is mainly modulated by miR-340.
Various studies have demonstrated that the RAB
family of proteins play important roles in NSCLC. The
downregulation of RAB27A has been shown to suppress the
proliferative, migratory and invasive abilities of NSCLC cells
in vitro, and to inhibit the growth of xenograft tumors in
mice, indicating that RAB27A may be a potential therapeutic target
in NSCLC (38). miR-451 regulates
the survival of NSCLC cells via the down-regulation of RAB14,
suggesting that targeting the interaction between miR-451 and RAB14
may be a novel therapeutic target in NSCLC (39). miR-30b/c directly targets and
downregulates the expression of RAB18 to inhibit the proliferation
of NSCLC cells. RAB11A protein has been shown to be overexpressed
in NSCLC tissues, and to be associated with advanced tumor, node
and metastasis stage, positive nodal status and poor patient
prognosis. In addition, the overexpression of RAB11A has been shown
to promote the proliferation, invasion and migration of NSCLC cells
via the upregulation of cyclin D1 and cyclin E, and the
downregulation of p27 (40).
RAB27, a member of the small GTPase family (41), has been reported to be associated
with various human cancers. For instance, RAB27A has been
identified as an inducer of melanoma growth (42), while the inhibition of RAB27A in
melanoma cell lines has been shown to suppress primary tumor growth
and lung metastasis (43). The
nuclear factor-κB-mediated RAB27A expression facilitates cytokine
secretion to promote the stemness of colon cancer cells (44). RAB27A expression has been
associated with tumor grade and the unfavorable prognosis of glioma
(45). These studies indicate
RAB27A as a potential tumor-promoting protein in human cancers,
whereas others have suggested that RAB27A serves as a tumor
suppressor. RAB27A upregulation has been shown to be associated
with the favorable prognosis of patients with colorectal cancer
(46). Furthermore, RAB27B
facilitates the invasion and migration of estrogen
receptor-positive breast cancer cells (47); the downregulated expression RAB27A
and RAB27B has been detected in the late stages of prostate cancer
(48).
Few studies have reported the association between
RAB and miR-340 in NSCLC; thus, TargetScan software was employed in
the present study, which predicted RAB11A, RAB27B and RAB43 as
potential targets of miR-340. RAB27B was proposed to be the most
likely target of miR-340 of the aforementioned RAB proteins. In
addition, we also conducted a luciferase reporter experiment to
confirm whether RAB27B is a direct target of miR-340. The present
study identified RAB27B as a novel target gene of miR-340, and
observed that miR-340 was downregulated, while RAB27B was
upregulated in NSCLC tissues and cell lines. miR-340 overexpression
inhibited the proliferation and invasion of NSCLC cells; these
effects were reversed by the knockdown of RAB27B. The mechanisms
underlying the effects of miR-340 comprise the targeting and
inhibition of RAB27B by miR-340. Furthermore, the miR-340/RAB27B
axis may be involved in the occurrence and progression of
NSCLC.
Previous reports have demonstrated that RAB27A and
RAB27B are the major components involved in vesicle fusion and
trafficking, and exosome secretion, playing important roles in
tumor progression and metastasis. The increased expression of
RAB27B has been observed in hepatocellular carcinoma (49), colorectal cancer (50) and breast cancer (51), indicating that RAB27B may be a
valuable predictor of metastasis and prognosis, or a potential
therapeutic target for the treatment of various types of cancer.
The upregulated expression of RAB27B may be an unfavorable
prognostic factor in patients with squamous cell carcinoma of the
lungs (52). Additionally, the
upregulated expression of RAB27B has been shown to be associated
with the malignant features of lung adenocarcinoma (LUAD); RAB27B
was identified as a potential indicator of metastasis and the
prognosis of LUAD (53).
In conclusion, miR-340 plays a critical role in
NSCLC, and its overexpression restricts the growth and invasion of
NSCLC cells by downregulating RAB27B. The results of the present
study may provide novel insight into the molecular mechanisms
underlying the occurrence and development of NSCLC, However, the
potential regulatory mechanisms of miR-340 targeting RAB27B require
further investigation.
Acknowledgments
Not applicable.
Funding
This study was supported by grants from the large
data system platform for laboratory medicine consultation oriented
to precision medicine (2017TJPT0003).
Availability of data and materials
All data generated or analyzed during this study are
included in this published article or are available from the
corresponding author on reasonable request.
Authors' contributions
XZ and JL conceived and designed the experiments.
XZ, GT, JQ and PH performed these experiments and analyzed the
data. XZ and JL drafted and revised the manuscript. All authors
have read and approved the final manuscript.
Ethics approval and consent to
participate
The present study was approved by the Ethics
Committee of the Affiliated Hospital of Southwest Medical
University. Written informed consent was obtained from all
patients.
Patient consent for publication
Not applicable.
Competing interests
All the authors declare that there are no any
competing interests.
References
|
1
|
Siegel R, Ma J, Zou Z and Jemal A: Cancer
statistics, 2014. CA Cancer J Clin. 64:9–29. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Ferlay J, Soerjomataram I, Dikshit R, Eser
S, Mathers C, Rebelo M, Parkin DM, Forman D and Bray F: Cancer
incidence and mortality worldwide: Sources methods and major
patterns in GLOBOCAN 2012. Int J Cancer. 136:E359–E386. 2015.
View Article : Google Scholar
|
|
3
|
Rossi A, Maione P, Sacco PC, Sgambato A,
Casaluce F, Ferrara ML, Palazzolo G, Ciardiello F and Gridelli C:
ALK inhibitors and advanced non-small cell lung cancer (Review).
Int J Oncol. 45:499–508. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Raitoharju E, Seppala I, Oksala N,
Lyytikainen LP, Raitakari O, Viikari J, Ala-Korpela M, Soininen P,
Kangas AJ, Waldenberger M, et al: Blood microRNA profile associates
with the levels of serum lipids and metabolites associated with
glucose metabolism and insulin resistance and pinpoints pathways
underlying metabolic syndrome: The cardiovascular risk in young
finns study. Mol Cell Endocrinol. 391:41–49. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Jiang F, Yu Q, Chu Y, Zhu X, Lu W, Liu Q
and Wang Q: MicroRNA-98-5p inhibits proliferation and metastasis in
non-small cell lung cancer by targeting TGFBR1. Int J Oncol.
54:128–138. 2019.
|
|
6
|
Tang Q, Li M, Chen L, Bi F and Xia H:
miR-200b/c targets the expression of RhoE and inhibits the
proliferation and invasion of non-small cell lung cancer cells. Int
J Oncol. 53:1732–1742. 2018.PubMed/NCBI
|
|
7
|
Lujambio A and Lowe SW: The microcosmos of
cancer. Nature. 482:347–355. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Othman N and Nagoor NH: Overexpression of
miR-361-5p plays an oncogenic role in human lung adenocarcinoma
through the regulation of SMAD2. Int J Oncol. 54:306–314. 2019.
|
|
9
|
Rongxin S, Pengfei L, Li S, Xiaochen J and
Yihe H: MicroRNA-340-5p suppresses osteosarcoma development by
down-regulating the Wnt/β-catenin signaling pathway via targeting
the STAT3 gene. Eur Rev Med Pharmacol Sci. 23:982–991.
2019.PubMed/NCBI
|
|
10
|
Shi Z, Li Y, Qian X, Hu Y, Liu J, Zhang S
and Zhang J: miR-340 inhibits triple-negative breast cancer
progression by reversing EZH2 mediated miRNAs dysregulated
expressions. J Cancer. 8:3037–3048. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Wei P, Qiao B, Li Q, Han X, Zhang H, Huo Q
and Sun J: microRNA-340 suppresses tumorigenic potential of
prostate cancer cells by targeting high-mobility group
nucleosome-binding domain 5. DNA Cell Biol. 35:33–43. 2016.
View Article : Google Scholar
|
|
12
|
Huang D, Qiu S, Ge R, He L, Li M, Li Y and
Peng Y: miR-340 suppresses glioblastoma multiforme. Oncotarget.
6:9257–9270. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Huang T, Zhou Y, Zhang J, Wong CC, Li W,
Kwan JSH, Yang R, Chan AKY, Dong Y, Wu F, et al: SRGAP1, a crucial
target of miR-340 and miR-124, functions as a potential oncogene in
gastric tumorigenesis. Oncogene. 37:1159–1174. 2018. View Article : Google Scholar :
|
|
14
|
Xie L, Chen Z, Liu H, Guan L, Wang Z and
Li W: Effects of miR-340 on hepatocellular carcinoma by targeting
the DcR3 gene. Dig Liver Dis. 50:291–296. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Zhao P, Ma W, Hu Z, Zhang Y, Zhang S and
Wang Y: Up-regulation of miR-340-5p promotes progression of thyroid
cancer by inhibiting BMP4. J Endocrinol Invest. 41:1165–1172. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Chen CP, Sun ZL, Lu X, Wu WX, Guo WL, Lu
JJ, Han C, Huang JQ and Fang Y: miR-340 suppresses cell migration
and invasion by targeting MYO10 in breast cancer. Oncol Rep.
35:709–716. 2016. View Article : Google Scholar
|
|
17
|
Mohammadi-Yeganeh S, Paryan M, Arefian E,
Vasei M, Ghanbarian H, Mahdian R, Karimipoor M and Soleimani M:
MicroRNA-340 inhibits the migration, invasion, and metastasis of
breast cancer cells by targeting Wnt pathway. Tumour Biol.
37:8993–9000. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Maskey N, Li D, Xu H, Song H, Wu C, Hua K,
Song J and Fang L: MicroRNA-340 inhibits invasion and metastasis by
down-regulating ROCK1 in breast cancer cells. Oncol Lett.
14:2261–2267. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Zhang Z, Wang Y, Zhang W, Li J, Liu W and
Lu W: Long non-coding RNA SNHG14 exerts oncogenic functions in
non-small cell lung cancer through acting as a miR-340 sponge.
Biosci Rep. 39:pii: BSR20180941. 2019.
|
|
20
|
Qin Y, Zhou X, Huang C, Li L, Liu H, Liang
N, Chen Y, Ma D, Han Z, Xu X, et al: Lower miR-340 expression
predicts poor prognosis of non-small cell lung cancer and promotes
cell proliferation by targeting CDK4. Gene. 675:278–284. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Fernandez S, Risolino M, Mandia N, Talotta
F, Soini Y, Incoronato M, Condorelli G, Banfi S and Verde P:
miR-340 inhibits tumor cell proliferation and induces apoptosis by
targeting multiple negative regulators of p27 in non-small cell
lung cancer. Oncogene. 34:3240–3250. 2015. View Article : Google Scholar
|
|
22
|
Chia WJ and Tang BL: Emerging roles for
Rab family GTPases in human cancer. Biochim Biophys Acta.
1795:110–116. 2009.PubMed/NCBI
|
|
23
|
Bobrie A, Krumeich S, Reyal F, Recchi C,
Moita LF, Seabra MC, Ostrowski M and Théry C: Rab27a supports
exosome-dependent and -independent mechanisms that modify the
tumour microenvironment and can promote tumour progression. Cancer
Res. 72:4920–4930. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Zhong K, Chen K, Han L and Li B:
MicroRNA-30b/c inhibits non-small cell lung cancer cell
proliferation by targeting Rab18. BMC Cancer. 14:7032014.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Li W, Mu D, Tian F, Hu Y, Jiang T, Han Y,
Chen J, Han G and Li X: Exosomes derived from Rab27a-overexpressing
tumor cells elicit efficient induction of antitumor immunity. Mol
Med Rep. 8:1876–1882. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
27
|
Rezaei Z, Sebzari A, Kordi-Tamandani DM
and Dastjerdi K: Involvement of the dysregulation of miR-23b-3p,
miR-195-5p, miR-656-5p and miR-340-5p in trastuzumab resistance of
HER2-positive breast cancer cells and system biology approach to
predict their targets involved in resistance. DNA Cell Biol.
38:184–192. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Wu ZS, Wu Q, Wang CQ, Wang XN, Huang J,
Zhao JJ, Mao SS, Zhang GH, Xu XC and Zhang N: miR-340 inhibition of
breast cancer cell migration and invasion through targeting of
oncoprotein c-Met. Cancer. 117:2842–2852. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Hou LK, Yu Y, Xie YG, Wang J, Mao JF,
Zhang B, Wang X and Cao XC: miR-340 and ZEB1 negative feedback loop
regulates TGF-β-mediated breast cancer progression. Oncotarget.
7:26016–26026. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Li X, Gong X, Chen J, Zhang J, Sun J and
Guo M: miR-340 inhibits glioblastoma cell proliferation by
suppressing CDK6, cyclin-D1 and cyclin-D2. Biochem Biophys Res
Commun. 460:670–677. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Yamashita D, Kondo T, Ohue S, Takahashi H,
Ishikawa M, Matoba R, Suehiro S, Kohno S, Harada H, Tanaka J and
Ohnishi T: miR340 suppresses the stem-like cell function of
glioma-initiating cells by targeting tissue plasminogen activator.
Cancer Res. 75:1123–1133. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Takeyama H, Yamamoto H, Yamashita S, Wu X,
Takahashi H, Nishimura J, Haraguchi N, Miyake Y, Suzuki R, Murata
K, et al: Decreased miR-340 expression in bone marrow is associated
with liver metastasis of colorectal cancer. Mol Cancer Ther.
13:976–985. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Li P, Sun Y and Liu Q: MicroRNA-340
induces apoptosis and inhibits metastasis of ovarian cancer cells
by inactivation of NF-x03BA;B1. Cell Physiol Biochem. 38:1915–1927.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Huang K, Tang Y, He L and Dai Y:
MicroRNA-340 inhibits prostate cancer cell proliferation and
metastasis by targeting the MDM2-p53 pathway. Oncol Rep.
35:887–895. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Yu W, Zhang G, Lu B, Li J, Wu Z, Ma H,
Wang H and Lian R: miR-340 impedes the progression of laryngeal
squamous cell carcinoma by targeting EZH2. Gene. 577:193–201. 2016.
View Article : Google Scholar
|
|
36
|
Zhou X, Wei M and Wang W: MicroRNA-340
suppresses osteosarcoma tumor growth and metastasis by directly
targeting ROCK1. Biochem Biophys Res Commun. 437:653–658. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Yin G, Zhou H, Xue Y, Yao B and Zhao W:
MicroRNA-340 promotes the tumor growth of human gastric cancer by
inhibiting cyclin G2. Oncol Rep. 36:1111–1118. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Li X, Wang H, Ni Q, Tang Z, Ni J, Xu L,
Huang H, Ni S and Feng J: Effects of silencing Rab27a gene on
biological characteristics and chemosensitivity of non-small cell
lung cancer. Oncotarget. 8:94481–94492. 2017.PubMed/NCBI
|
|
39
|
Wang R, Wang ZX, Yang JS, Pan X, De W and
Chen LB: MicroRNA-451 functions as a tumor suppressor in human
non-small cell lung cancer by targeting ras-related protein 14
(RAB14). Oncogene. 30:2644–2658. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Dong Q, Fu L, Zhao Y, Du Y, Li Q, Qiu X
and Wang E: Rab11a promotes proliferation and invasion through
regulation of YAP in non-small cell lung cancer. Oncotarget.
8:27800–27811. 2017.PubMed/NCBI
|
|
41
|
Imai A, Yoshie S, Ishibashi K,
Haga-Tsujimura M, Nashida T, Shimomura H and Fukuda M: EPI64 pr
parotid acinar cells otein functions as a physiological
GTPase-activating protein for RAB27 protein and regulates amylase
release in rat. J Biol Chem. 286:33854–33862. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Akavia UD, Litvin O, Kim J, Sanchez-Garcia
F, Kotliar D, Causton HC, Pochanard P, Mozes E, Garraway LA and
Pe'er D: An integrated approach to uncover drivers of cancer. Cell.
143:1005–1017. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Peinado H, Alečković M, Lavotshkin S,
Matei I, Costa-Silva B, Moreno-Bueno G, Hergueta-Redondo M,
Williams C, García-Santos G, Ghajar C, et al: Melanoma exosomes
educate bone marrow progenitor cells toward a pro-metastatic
phenotype through MET. Nat Med. 18:883–891. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Feng F, Jiang Y, Lu H, Lu X, Wang S, Wang
L, Wei M, Lu W, Du Z, Ye Z, et al: Rab27A mediated by NF-κB
promotes the stemness of colon cancer cells via up-regulation of
cytokine secretion. Oncotarget. 7:63342–63351. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Wang H, Zhao Y, Zhang C, Li M, Jiang C and
Li Y: Rab27a was identified as a prognostic biomaker by mRNA
profiling, correlated with malignant progression and subtype
preference in gliomas. PLoS One. 9:e897822014. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Shi C, Yang X, Ni Y, Hou N, Xu L, Zhan F,
Zhu H, Xiong L and Chen P: High RAB27A expression indicates
favorable prognosis in CRC. Diagn Pathol. 10:682015. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Hendrix A, Maynard D, Pauwels P, Braems G,
Denys H, Van den Broecke R, Lambert J, Van Belle S, Cocquyt V,
Gespach C, et al: Effect of the secretory small GTPase RAB27B on
breast cancer growth, invasion, and metastasis. J Natl Cancer Inst.
102:866–880. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Worst TS, Meyer Y, Gottschalt M, Weis CA,
von Hardenberg J, Frank C, Steidler A, Michel MS and Erben P:
RAB27A, RAB27B and VPS36 are downregulated in advanced prostate
cancer and show functional relevance in prostate cancer cells. Int
J Oncol. 50:920–932. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Yang X, Ye X, Sun L, Gao F, Li Y, Ji X and
Wang X, Feng Y and Wang X: Downregulation of serum RAB27B confers
improved prognosis and is associated with hepatocellular carcinoma
progression through PI3K-AKT-P21 signaling. Oncotarget.
8:61118–61132. 2017.PubMed/NCBI
|
|
50
|
Bao J, Ni Y, Qin H, Xu L, Ge Z, Zhan F,
Zhu H, Zhao J, Zhou X, Tang X and Tang L: Rab27b is a potential
predictor for metastasis and prognosis in colorectal cancer.
Gastroenterol Res Pract. 2014:9131062014. View Article : Google Scholar
|
|
51
|
Hendrix A, Braems G, Bracke M, Seabra M,
Gahl W, De Wever O and Westbroek W: The secretory small GTPase
Rab27B as a marker for breast cancer progression. Oncotarget.
1:304–308. 2010.
|
|
52
|
Koh HM and Song DH: Prognostic role of
Rab27A and Rab27B expression in patients with non-small cell lung
carcinoma. Thorac Cancer. 10:143–149. 2019. View Article : Google Scholar
|
|
53
|
Zhang L, Fan W, Xu L, Mao Q, Chen Y, Mao
Y, Xu L and Wang J: Rab27b is a potential indicator for lymph node
metastasis and unfavorable prognosis in lung adenocarcinoma. Dis
Markers. 2018:72939622018. View Article : Google Scholar
|