Introduction
Cutaneous squamous cell carcinoma (CSCC) is one of
the most common skin malignancies, accounting for 25-35% of all
skin cancer types (1). The extent
of tumor infiltration and metastasis greatly influences the
clinical stage and prognosis of the disease; and although numerous
treatment options are available for CSCC, including surgical
excision, radiotherapy, photodynamic therapy and topical drug
treatment, the prognosis of invasive and metastatic CSCC remains
relatively poor and is associated with substantial levels of
mortality (2). Furthermore, the
incidence of CSCC continues to increase, which is largely due to
the increased prevalence of risk factors including older
populations, immunosuppression, chronic sun exposure and
sensitivity to sunlight or ultraviolet radiation (1,3).
Circular RNAs (circRNAs) are a newly identified
group of non-coding RNAs that lack 5'-caps and 3'-tails (4), which leaves them resistant to
exonuclease- or ribonuclease-medi-ated degradation and permits
their stable expression in numerous types of organisms. A number of
studies have reported that circRNA expression levels are
significantly increased in various types of tumor, where they serve
as important molecules for tumor metastasis and recur-rence
(5). circRNA may competitively
bind to microRNA (miRNA) response elements to inhibit miRNA
expression or function, which ultimately affects target genes
(6,7). Fos-related antigen 2 (FOSL2) is a
member of the activator protein 1 (AP-1) transcription factor
family, which includes the various isoforms of Fos and Jun
(8-10). Previous studies have demonstrated
that FOSL2 is abnormally expressed in numerous different types of
tumor, where it serves important functions in cell adhesion,
migration, invasion, metastasis and proliferation (11-12).
The aim of the present study was to investigate the
function of circRNA_001937 in CSCC. In the present study,
differ-ential circRNA expression profiles of CSCC were analyzed
using the Arraystar Human circRNAs chip and verified by RT-qPCR. In
addition, the effects of circRNA on cell behavior, in particular
its regulation of the miRNA-mRNA axis, were also investigated.
Materials and methods
Patient samples
The present study was approved by the Ethics
Committee of the First Hospital of Jilin University (Jilin, China)
and written informed consent was obtained from all patients. Three
pairs of CSCC tissues and corresponding adjacent tissues were
collected from the Department of Plastic Surgery at the First
Hospital of Jilin University between September 2015 and November
2018. The clinicopathological features are shown in detail in
Table SI. All specimens were
confirmed by clinical and pathological diagnosis.
Cell lines
The human CSCC cell lines A431 and SCL-1, and the
human immortal keratinocyte cell line HaCaT, were purchased from
Guangzhou Genio Biotechnology Co., Ltd.
Reagents
circRNA_001937, miRNA-597-3p and FOSL2 mRNA primer
sequences were purchased from Kangcheng Co., Ltd. Anti-FOSL2
primary antibody (1:500; cat. no. H00116173-B01P) and anti-GAPDH
primary antibody (1:1,000; cat. no. R2655) were purchased from
Sigma-Aldrich (Merck KGaA). RPMI-1640 medium, fetal bovine serum
(FBS), crystal violet, Annexin V fluorescein isothiocyanate
(FITC)/propidium iodide (PI) Cell Apoptosis Detection kit (cat. no.
sc-4252 AK), MTT assays, Transwell plates, Matrigel and dimethyl
sulfoxide were purchased from Santa Cruz Biotechnology, Inc.
Profiling of circRNA expression
The Arraystar Human circRNAs chip (Arraystar Inc.)
was used to analyze the expres-sion of circRNAs in the CSCC tissues
and corresponding adjacent tissues. Total RNA was extracted from
the samples using an RNeasy Mini kit (cat. no. 74104; Qiagen GmbH),
and the RNA was analyzed on the circRNAs chips. The expression
levels were analyzed and quantified by Kangcheng Co., Ltd.
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA was extracted from A431 and SCL-1 cells
using TRIzol® reagent (Thermo Fisher Scientific, Inc.).
Total RNA was reverse transcribed into cDNA at room temperature
using TaqMan™ reverse transcription reagents (cat. no. 4304134;
Thermo Fisher Scientific, Inc.). qPCR was subsequently performed
using the iScript™ cDNA Synthesis kit (cat. no. 1708890; Bio-Rad
Laboratories, Inc.). The following thermocycling conditions were
used for the qPCR: 40 cycles at 94°C for 15 sec; 20 cycles at 55°C
for 30 sec; and 20 cycles at 70°C for 30 sec. The following primers
were used: circRNA_001937 forward, 5'-TGA AGA ACA GCT CTC TGG
CTG-3' and reverse, 5'-GCC CAC TTA ATC AGG GTC AGG T-3';
miRNA-597-3p forward, 5'-CGG AAT TCA TCT CAA GCC AAC-3' and
reverse, 5'-CGG GAT CCC TTC ATT CAA GGT CAA TG-3'; FOSL2 forward,
5'-GAG AGG AAC AAG CTG GCT GC-3' and reverse, 5'-GCT TCT CCT TCT
CCT TCT GC-3'; U6 (control for miRNA) forward, 5'-TTT AGG GCT TCG
ATA CT-3' and reverse, 5'-TCT GCT GCA GCA CA-3'; and GAPDH (control
for circRNA and mRNA) forward, 5'-GGT CCT GTT GTT TA-3' and
reverse, 5'-TGC TCA TTC CCT C-3'. Expression levels were quantified
using the 2-Δ∆Cq method (13) and the relative expression levels
of target RNAs were normalized to the loading control U6.
Cell transfection
Small interfering RNA (siRNA/si) targeting
circRNA_001937 (si-circRNA_001937) and the nega-tive control
(si-NC) labeled with green fluorescent protein, miRNA-597-3p mimic,
miRNA-597-3p inhibitor and the NC were synthesized by Kangcheng
Co., Ltd. The sequence of si-circRNA_001937 was 5'-GGC AGC ACA TGT
CAG GC-3' and the sequence of si-NC was 5'-TCT TTA GGG GTG TGC GTA
GG-3'. The quantity of siRNA transfected was 50 nM. A431 and SCL-1
cells (1x105/well) were transfected using
Lipofectamine® 2000 reagent (Thermo Fisher Scientific,
Inc.) according to the manufacturer's protocol. Cells were
trans-fected for 48 h prior to subsequent experimentation.
MTT assay
Following transfection, A431 and SCL-1 cells in the
logarithmic growth phase were seeded at a density of
1x105/well into 96-well culture plates. Following 0, 12,
24 or 48 h of culture, 20 µl MTT solution was added/well for 6 h to
determine cell proliferation. Following the MTT incubation, cells
were washed with phosphate buffered saline (PBS) and the purple
formazan crystals were dissolved using 100 µl dimethyl
sulfoxide/well. Proliferation was subsequently analyzed at a
wavelength of 490 nm using a microtiter plate reader.
Colony formation assay
Following transfection, A431 and SCL-1 cells
(1x104/well) in the logarithmic growth phase were plated
into 6-well culture plates and cultured for 2 weeks at 37°C.
Following incubation, the cells were washed twice with PBS, fixed
with 2% paraformaldehyde at 37°C for 30 min and subsequently
stained with 0.5% crystal violet at 37°C for 15 min. Colonies were
visualized using a Nikon electron microscope (magnification, x100;
Nikon Corporation) and analyzed using ImageJ version 1.47 software
(National Institutes of Health).
Matrigel invasion assay
Following transfection, A431 and SCL-1 cells
(1x105/well) in the logarithmic growth phase were plated
in the upper chambers of Transwell plates with Matrigel and
fibronectin was also added to the upper chamber. RPMI-1640 medium
supplemented with 20% FBS was plated in the lower chambers.
Following incubation at 37°C for 24 h, the lower chamber cells were
fixed with 2% parafor-maldehyde at 37°C for 30 min and stained with
0.5% crystal violet at 37°C for 15 min. Stained A431 and SCL-1
cells were visualized using a confocal microscope (magnification,
x100) and ImageJ version 1.47 software was used to quantify the
number of invasive cells.
Flow cytometric analysis of
apoptosis
Cell apoptosis was performed using the Annexin
V-FITC/PI apoptosis detection kit, according to the manufacturer's
protocol. Following trans-fection, A431 and SCL-1 cells in the
logarithmic growth phase were seeded into 96-well culture plates
and were subsequently incubated with Annexin V-FITC and PI solution
at 37°C for 20 min in the dark. Apoptotic cells were visualized
using a BD FACSCalibur flow cytometer (BD Biosciences).
Dual-luciferase reporter assay
A431 and SCL-1 cells in the logarithmic growth phase
were digested and seeded into 24-well culture plates. Subsequently,
the A431 and SCL-1 cells were co-transfected with miRNA-597-3p
mimics or the miRNA-597-3p NC and the wild-type (wt) or mutated
(mut) 3'-untranslated region (UTR) of circRNA_001937 and FOSL2.
Following 36 h of transfection, the luciferase activity was
performed by a Dual-Luciferase Reporter Assay system (cat. no.
E1910; Promega Corporation). Relative luciferase activity was
normalized to the Renilla luciferase internal control.
Western blotting
A431 and SCL-1 cells of each group were lysed with
RIPA buffer (Sigma-Aldrich; Merck KGaA). Protein quantification was
carried out using a BCA protein assay kit (Promega Corporation). A
total of 50 µg protein/lane was separated by 10% SDS-PAGE and
subsequently transferred to polyvinylidene difluoride membranes.
The membranes were then blocked for 1 h at room temperature with
non-fat dry milk in TBST (Bio-Rad Laboratories, Inc.). The
membranes were incubated with the anti-FOSL2 primary antibody
(1:500; cat. no. H00116173-B01P; Gibco; Thermo Fisher Scientific,
Inc.) and anti-β-actin (1:1,000; cat. no. R2655; Sigma-Aldrich;
Merck KGaA) at 37°C overnight. Then, membranes were washed with
PBS-Tween-20 buffer and subsequently incu-bated with a horseradish
peroxidase-conjugated anti-rabbit antibody (1:1,000; cat. no.
G-21234; Invitrogen; Thermo Fisher Scientific, Inc.) for 2 h at
4°C. Protein bands were visualized using the Pierce ECL Western
blotting substrate (Pierce; Thermo Fisher Scientific, Inc.).
Statistical analysis
Statistical analysis was performed using SPSS 17.0
software (SPSS, Inc.) and the data were presented as the mean ±
standard deviation. Statistical differences between two groups were
determined using an unpaired Student's t-test or an χ2
test, whereas statistical differences between >2 groups were
analyzed using a one-way analysis of variance, followed by Tukey's
multiple comparison test. P<0.05 was considered to indicate a
statistically significant difference.
Results
circRNA_001937 expression levels are
significantly increased in CSCC
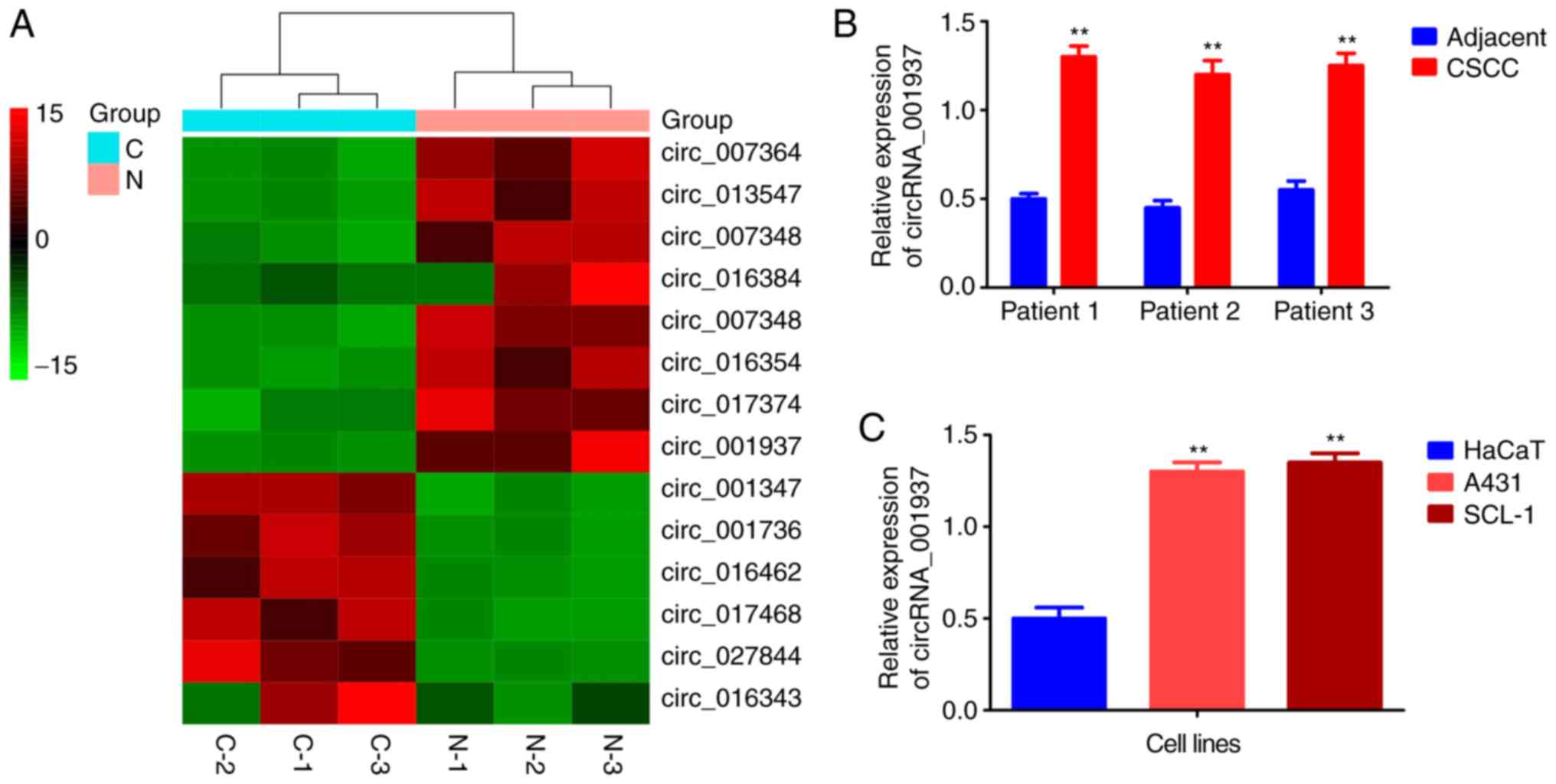
The cluster heat map demonstrated that
circRNA_001937 expression levels were increased by 14.58-fold in
CSCC tissues (Fig. 1A).
Similarly, results from the RT-qPCR also reported that
circRNA_001937 levels were significantly increased in CSCC tissues
(P<0.01; Fig. 1B), and A431
and SCL-1 cells (P<0.01; Fig.
1C) compared with the corresponding adjacent tissues and HaCaT
cells.
si-circRNA_001937 transfection
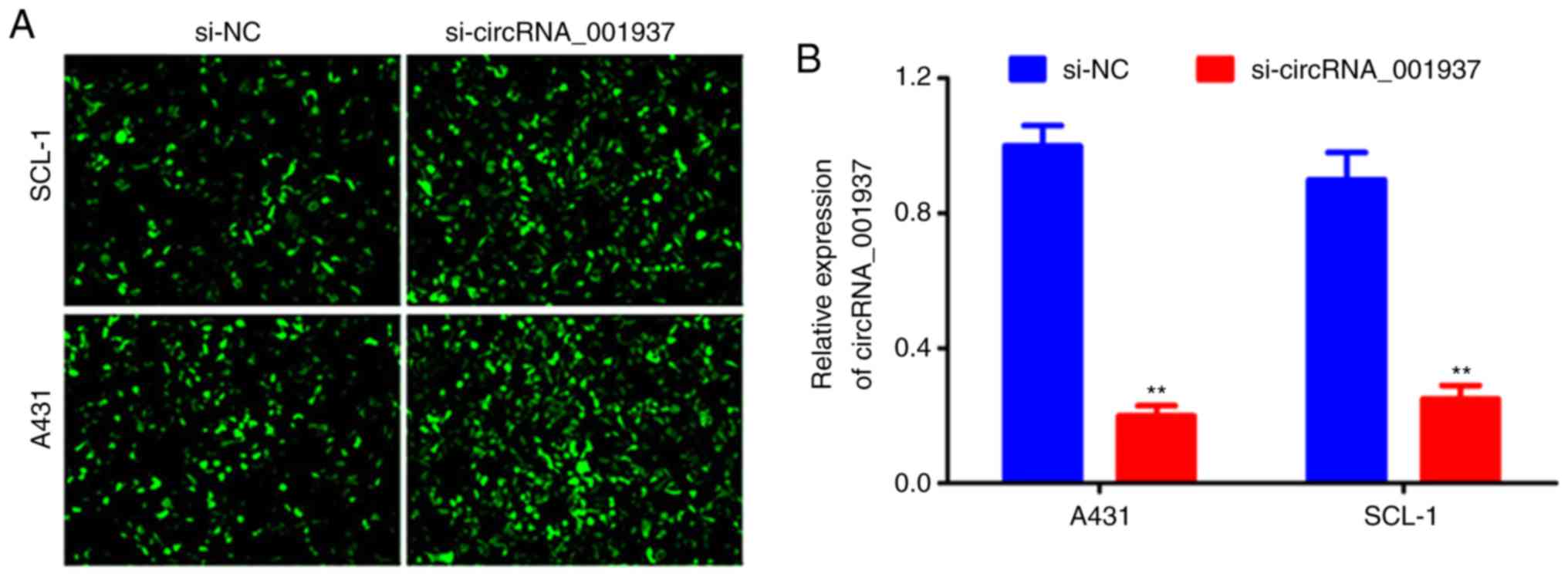
Following cell transfection with si-circRNA_001937
or si-NC, bright green fluorescence was observed in the si-NC and
si-circRNA_001937 groups (Fig.
2A). The expression levels of circRNA_001937 in the
si-circRNA_001937 group were significantly decreased compared with
the si-NC group (P<0.01; Fig.
2B).
Silencing circRNA_001937 expression
inhibits CSCC prolif- eration, and induces apoptosis
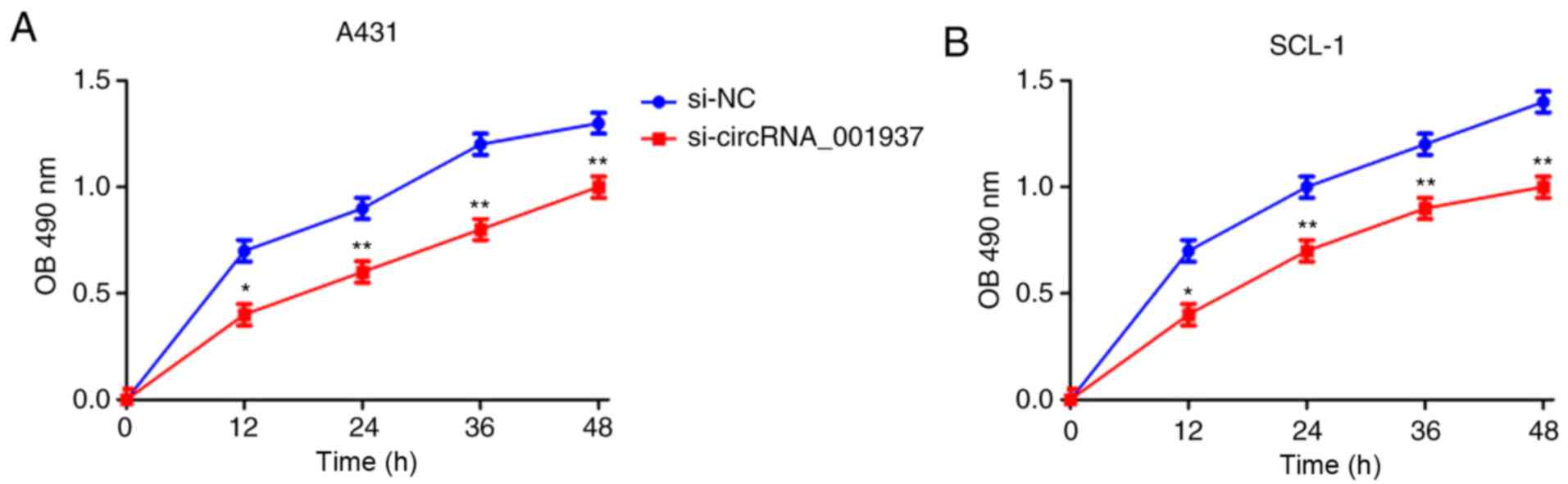
Subsequently, the effect of circRNA_001937 on cell
behavior was investigated. The proliferative rate (P<0.05) were
significantly reduced in the si-circRNA_001937 group compared with
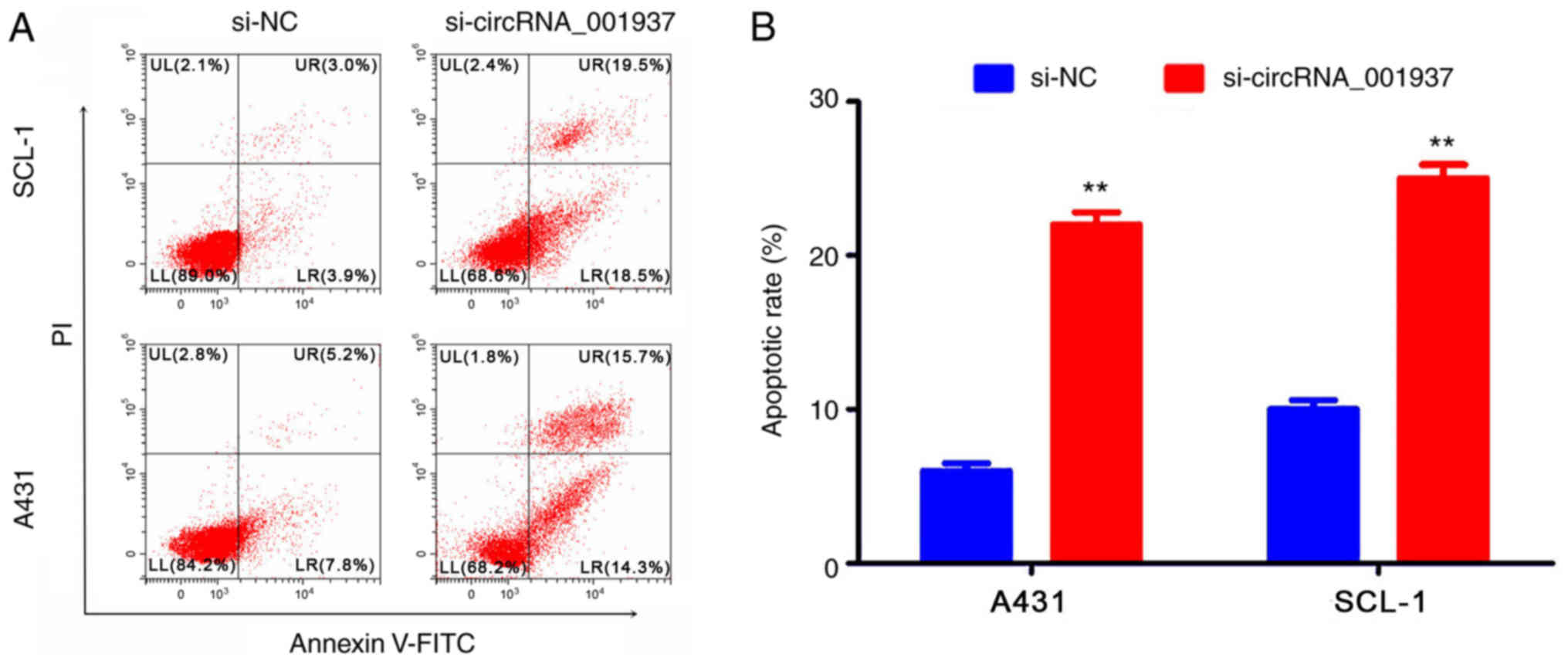
the si-NC group (Fig. 3). In
addition, flow cytometric analysis identified that the apoptotic
rate was significantly increased in the si-circRNA_001937 group
compared with the si-NC group (P<0.01; Fig. 4).
circRNA_001937 functions as an miRNA
sponge for miRNA-597-3p
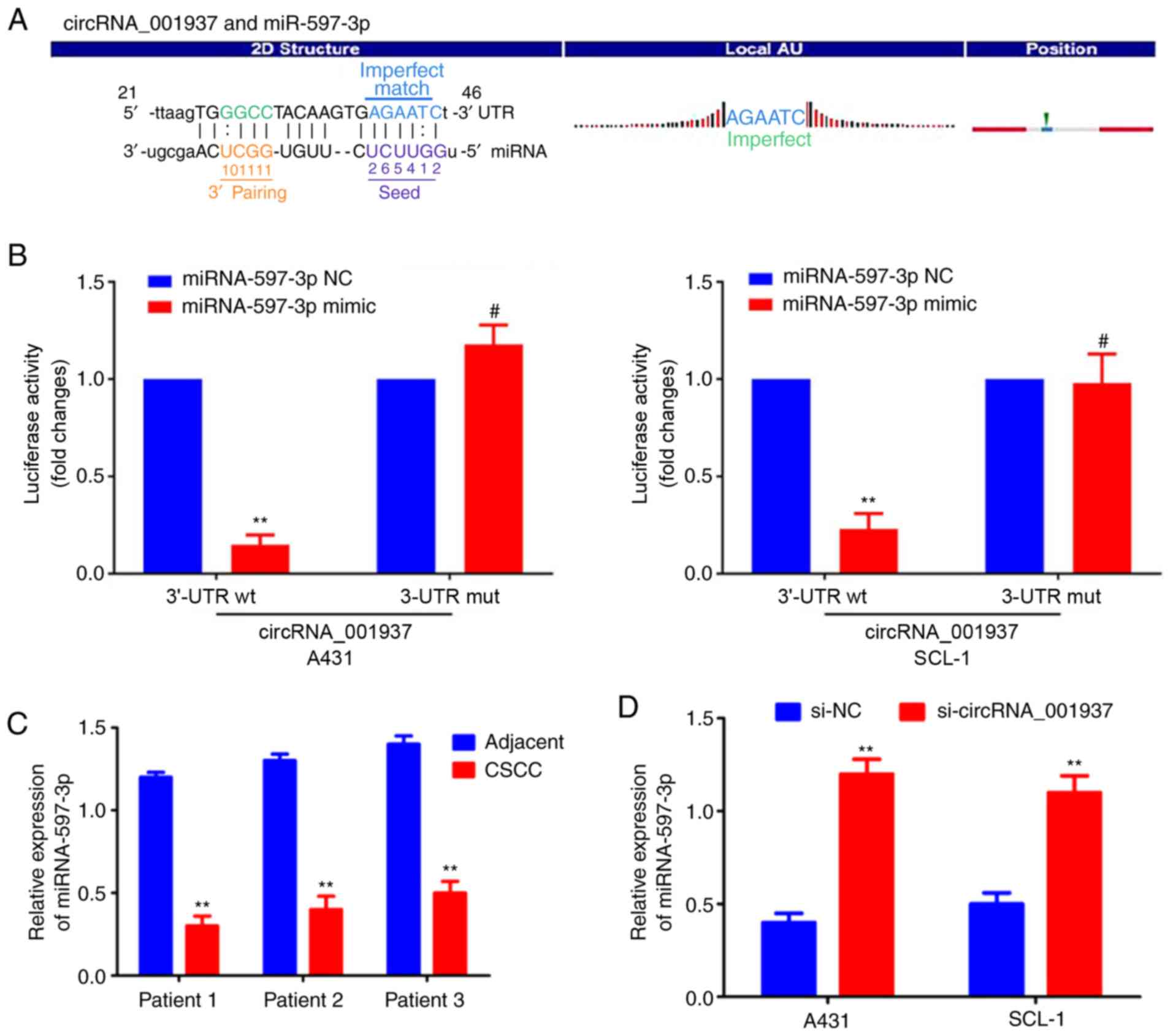
The specific binding sequences between
circRNA_001937 and miRNA-597-3p are presented in Fig. 5A. Dual-luciferase reporter assays
were used to demonstrate that the relative luciferase activity in
the circRNA_001937-3'-UTR-Wt and miRNA-597-3p mimic co-transfection
group was significantly decreased compared with the groups
co-transfected with miRNA-597-3p NC or circRNA_001937-3'-UTR-Mut
(P<0.01; Fig. 5B).
miRNA-597-3p expression levels were also significantly decreased in
CSCC tissues compared with corresponding adjacent tissues
(P<0.01; Fig. 5C); however,
these miRNA-597-3p expression levels were significantly increased
following transfection with si-circRNA_001937 compared with the
si-NC group (P<0.01; Fig.
5D).
FOSL2 is a direct target of
miRNA-597-3p
The specific binding sequences between FOSL2 3'-UTR
and miRNA-597-3p were predicted using TargetScan 7.0 software
(Fig. 6A). Dual-luciferase
reporter assay results demonstrated that the relative luciferase
activity in cells co-transfected with FOSL2-3'-UTR-Wt and
miRNA-597-3p mimic was significantly decreased compared with the
miRNA-597-3p NC or FOSL2-3'-UTR-Mut groups (P<0.01; Fig. 6B). In addition, FOSL2 mRNA
expression levels were significantly increased in CSCC tissues
compared with the adjacent noncancerous tissues (P<0.01;
Fig. 6C), whereas FOSL2
expression levels were significantly decreased in the miRNA-597-3p
mimic group, and significantly increased in the miRNA-597-3p
inhibitor group, compared with the miRNA-597-3p NC group
(P<0.01; Fig. 6D).
FOSL2 is activated by the
circRNA_001937/miRNA-597-3p axis
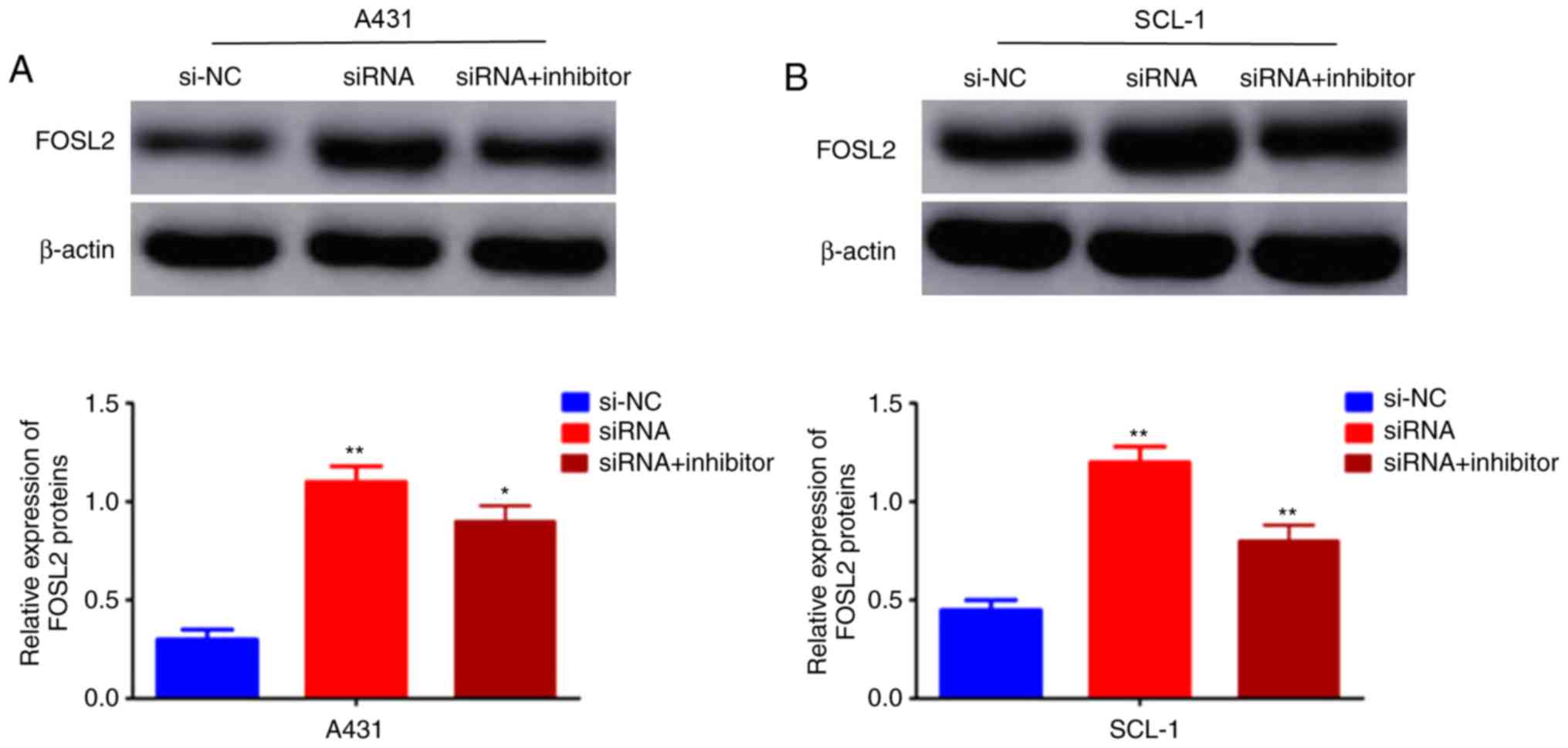
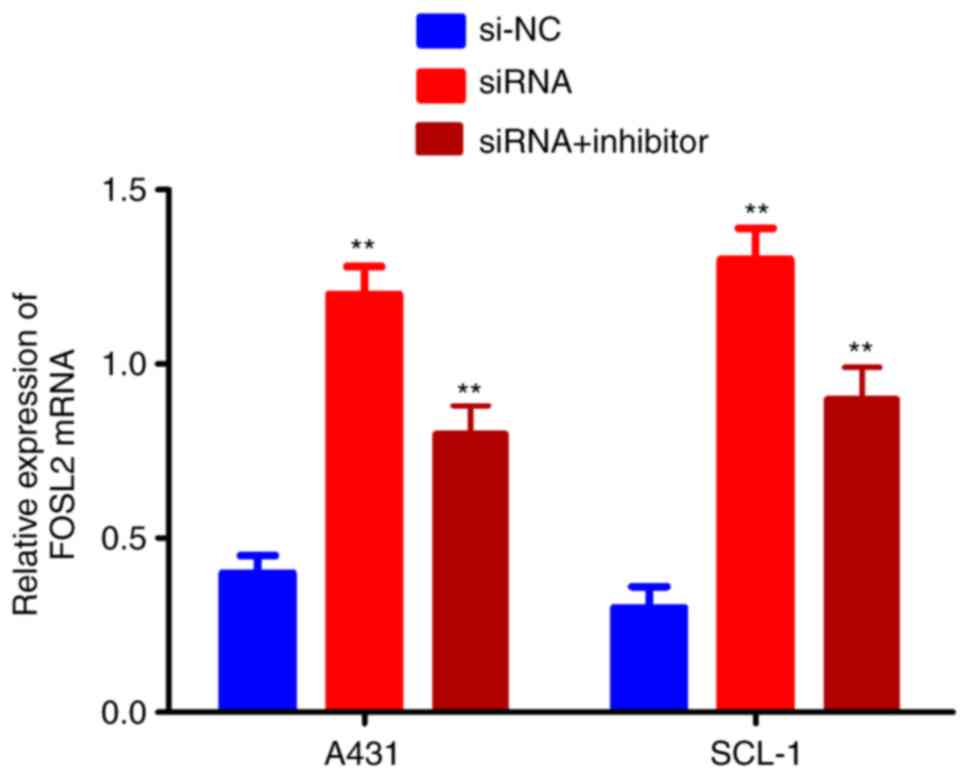
The effects of the circRNA_001937/miRNA-597-3p axis
on FOSL2 mRNA and protein expression levels was further
investigated using RT-qPCR and western blotting, respectively. The
expression levels of FOSL2 protein (P<0.01; Fig. 7A and B) and mRNA (P<0.01;
Fig. 8) were significantly
increased in the si-circRNA_001937 group compared with the si-NC
group; and were substantially decreased in the si-circRNA_001937
and miRNA-597-3p inhibitor co-transfection group compared with the
si-circRNA_001937 group.
Discussion
circRNAs serve as gene regulators in a variety of
physiological functions and pathological processes, of which their
functions in numerous types of cancer are currently being
investigated. For example, circ_0003159 was identified as a
potential biomarker for patients with gastric cancer. circ_001569
was identified to serve as a sponge for miR-145, which may prove
beneficial as increased miR-145 expression promotes proliferation
and invasion in colorectal cancer (14). Furthermore, circ_0043278 promoted
non-small cell lung cancer (NSCLC) proliferation and migration
through regulating miR-520 expression (15). Previous studies have also reported
that circ_0016788 regu-lated hepatocellular carcinoma (HCC)
tumorigenesis through the miR-486/cyclin-dependent kinase 4 pathway
(16-18) and circRNA-mitochondrial tRNA
translation optimization 1, whose expression was decreased in HCC,
and was observed to sequester miRNA-9 and suppress HCC
progression.
In the present study, the circRNAs chip and RT-qPCR
results demonstrated that circRNA_001937 expression levels were
increased in CSCC, and that silencing circRNA_001937 inhibited cell
proliferation and invasion and induced cell apoptosis.
circRNA_001937 is an exonic circRNA that is 2,850 nucleotides in
length, and is located on chromosome 16 (19). Huang et al(20) reported that circRNA_001937
expression was significantly increased in the peripheral blood
mononuclear cells of patients with tuberculosis, and that
circRNA_001937 was correlated with tuberculosis severity, with
expression levels successfully decreasing following treatment The
results obtained in the present study suggest that circRNA_001937
may be used as a potential diagnostic biomarker for CSCC. However,
the number of patients with CSCC used in the present study was
small, and large-scale clinical samples and adequate follow-up
studies are required for further verification. In addition,
dual-luciferase reporter assays confirmed that circRNA_001937
served as a miRNA sponge towards miRNA-597-3p, which subsequently
increased FOSL2 expres-sion. There are a limited number of studies
examining the miRNA-597-3p/FOSL2 pathway. miRNA-597-3p is located
on the 8p23.1 chromosome (21).
Xie et al(22) reported
that miR-597 targeting 14-3-3σ enhances cellular invasion and the
epithelial-mesenchymal transition in nasopharyngeal carcinoma
cells, whilst Zhang et al(23) revealed that a low expression of
miR-597 is correlated with tumor stage and a poor outcome in breast
cancer. FOSL2 is a member of the AP-1 transcription factor family
(24). Previous studies have
demonstrated that FOSL2 is abnormally expressed in numerous
different types of tumor. Wang et al(25) identified that FOSL2 may positively
regulate transforming growth factor-β1 signaling in NSCLC. Sun
et al(26) confirmed that
miR-143-3p inhibited the proliferation, migration and invasion of
osteosarcoma through targeting FOSL2. Additionally, FOSL2
expression may be regulated through a number of different
mechanisms. miRNA regulation is one method, including miRNA-30e
(27) and miR-143-3p (26), which have been revealed to
regulate the expression of FOSL2. These studies demonstrate that
miRNA-597-3p and FOSL2 participate in the carcinogenesis and
development of cancer. The present study demonstrated that
miRNA-597-3p expression was significantly decreased, and FOSL2
expression was significantly increased, in CSCC tissues. The FOSL2
gene was additionally observed to be directly targeted by
miRNA-597-3p, and FOSL2 expression levels were observed to be
increased by circRNA_001937 serving as a sponge for
miRNA-597-3p.
In conclusion, to the best of our knowledge, the
present study provides the first evidence that circRNA_001937
expression is significantly increased in CSCC, and that silencing
circRNA_001937 inhibits CSCC proliferation and invasion and induces
apoptosis. Silencing circRNA_001937 gene expression may inhibit
CSCC progression by preventing the sponging of the
miRNA-597-3p/FOSL2 pathway. These results suggest a novel,
potential therapeutic target for the treatment of patients with
CSCC. Large-scale, clinical and adequate follow-up studies are
required for further verification of these results.
Supplementary Data
Funding
No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
LG and QL designed the study and performed the
experiments. HJJ and DZ analyzed the data. All authors have read
and approved the final manuscript.
Ethics approval and consent to
participate
The present study was approved by the Ethics
Committee of the First Hospital of Jilin University (Jilin, China),
and written informed consent was obtained from all patients.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Acknowledgements
Not applicable.
References
|
1
|
Castelo B, Viñal D, Maseda R, Ostios L,
Sánchez D, García-Salvatierra B, Escámez MJ, Martínez-Santamaría L,
Del Río M, Mora-Rillo M, et al: Epidemiology and natural history of
cutaneous squamous cell carcinoma in recessive dystrophic
epidermolysis bullosa patients: 20 years' experience of a reference
centre in Spain. Clin Transl Oncol. 21:1573–1577. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Xiong Y, Dresser K and Cornejo KM:
Frequent TLE1 expression in cutaneous neoplasms. Am J
Dermatopathol. 41:1–6. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Ogata D and Tsuchida T: Systemic
immunotherapy for advanced cutaneous squamous cell carcinoma. Curr
Treat Options Oncol. 20:302019. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Ding L, Zhao Y, Dang S, Wang Y, Li X, Yu
X, Li Z, Wei J, Liu M and Li G: Circular RNA circ-DONSON
facilitates gastric cancer growth and invasion via NURF complex
dependent activation of transcription factor SOX4. Mol Cancer.
18:452019. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Li Q, Deng C, Zhu T, Ling J, Zhang H, Kong
L, Zhang S, Wang J and Chen X: Dynamics of physiological and miRNA
changes after long-term proliferation in somatic embryogenesis of
Picea balfouriana. Trees. 33:469–480. 2019. View Article : Google Scholar
|
|
6
|
Niwa Y, Yamada S, Sonohara F, Kurimoto K,
Hayashi M, Tashiro M, Iwata N, Kanda M, Tanaka C, Kobayashi D, et
al: Identification of a serum-based miRNA signature for response of
esophageal squamous cell carcinoma to neoadjuvant chemo-therapy. J
Transl Med. 17:12019. View Article : Google Scholar
|
|
7
|
Kogure A, Kosaka N and Ochiya T:
Cross-talk between cancer cells and their neighbors via miRNA in
extracellular vesicles: An emerging player in cancer metastasis. J
Biomed Sci. 26:72019. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
He J, Mai J, Li Y, Chen L, Xu H, Zhu X and
Pan Q: miR-597 inhibits breast cancer cell proliferation, migration
and invasion through FOSL2. Oncol Rep. 37:2672–2678. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Wrann CD, Eguchi J, Bozec A, Xu Z,
Mikkelsen T, Gimble J, Nave H, Wagner EF, Ong SE and Rosen ED:
FOSL2 promotes leptin gene expression in human and mouse
adipocytes. J Clin Invest. 122:1010–1021. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Jahangiri L, Sharpe M, Novikov N,
González-Rosa JM, Borikova A, Nevis K, Paffett-Lugassy N, Zhao L,
Adams M, Guner-Ataman B, et al: The AP-1 transcription factor
component Fosl2 potentiates the rate of myocardial differentiation
from the zebrafish second heart field. Development. 143:113–122.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Hu Y, Li J, Li S, Wang S, Gao G and Hou J:
Expression level of FOSL2 mRNA in blood leukocytes of type 2
diabetes mellitus patients in Uygurs in Xinjiang and its clinical
sinificance. J Jilin Univ. 42:545–550. 2016.In Chinese.
|
|
12
|
Li Z, Liu Y, Yan J, Zeng Q, Hu Y, Wang H,
Li H, Li J and Yu Z: Circular RNA hsa_circ_0056836 functions an
onco-genic gene in hepatocellular carcinoma through modulating
miR-766-3p/FOSL2 axis. Aging (Albany NY). 12:2485–2497. 2020.
View Article : Google Scholar
|
|
13
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expres-sion data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
14
|
Xie H, Ren X, Xin S, Lan X, Lu G, Lin Y,
Yang S, Zeng Z, Liao W, Ding YQ and Liang L: Emerging roles of
circRNA_001569 targeting miR-145 in the proliferation and invasion
of colorectal cancer. Oncotarget. 7:26680–26691. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Cui J, Li W, Liu G, Chen X, Gao X, Lu H
and Lin D: A novel circular RNA, hsa_circ_0043278, acts as a
potential biomarker and promotes non-small cell lung cancer cell
proliferation and migration by regulating miR-520. Artif Cells
Nanomed Biotechnol. 47:810–821. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Guan Z, Tan J, Gao W, Li X, Yang Y, Li X,
Li Y and Wang Q: Circular RNA hsa_circ_0016788 regulates
hepatocellular carci-noma tumorigenesis through miR-486/CDK4
pathway. J Cell Physiol. 234:500–508. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Han D, Li J, Wang H, Su X, Hou J, Gu Y,
Qian C, Lin Y, Liu X, Huang M, et al: Circular RNA circMTO1 acts as
the sponge of microRNA-9 to suppress hepatocellular carcinoma
progression. Hepatology. 66:1151–1164. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Zhou J, Zhang WW, Peng F, Sun JY, He ZY
and Wu SG: Downregulation of hsa_circ_0011946 suppresses the
migration and invasion of the breast cancer cell line MCF-7 by
targeting RFC3. Cancer Manag Res. 10:535–544. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Ojha R, Nandani R, Chatterjee N and
Prajapati VK: Emerging role of circular RNAs as potential
biomarkers for the diagnosis of human diseases. Adv Exp Med Biol.
1087:141–157. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Huang ZK, Yao FY, Xu JQ, Deng Z, Su RG,
Peng YP, Luo Q and Li JM: Microarray expression profile of circular
RNAs in peripheral blood mononuclear cells from active tuberculosis
patients. Cell Physiol Biochem. 45:1230–1240. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Bay A, Coskun E, Oztuzcu S, Ergun S,
Yilmaz F and Aktekin E: Plasma microRNA profiling of pediatric
patients with immune thrombocytopenic purpura. Blood Coagul
Fibrinolysis. 25:379–383. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Xie L, Jiang T, Cheng A, Zhang T, Huang P,
Li P, Wen G, Lei F, Huang Y, Tang X, et al: MiR-597 targeting
14-3-3σ enhances cellular invasion and EMT in nasopharyngeal
carcinoma cells. Curr Mol Pharmacol. 12:105–114. 2019. View Article : Google Scholar
|
|
23
|
Zhang XY, Liu DJ, Yuan RB, Zhang DH, Li
SR, Zhang SH and Zhang LY: Low expression of miR-597 is correlated
with tumor stage and poor outcome in breast cancer. Eur Rev Med
Pharmacol Sci. 22:456–460. 2018.PubMed/NCBI
|
|
24
|
Li S, Fang XD, Wang XY and Fei BY:
Fos-like antigen 2 (FOSL2) promotes metastasis in colon cancer. Exp
Cell Res. 373:57–61. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Wang J, Sun D, Wang Y, Ren F, Pang S, Wang
D and Xu S: FOSL2 positively regulates TGF-β1 signalling in
non-small cell lung cancer. PLoS One. 9:e1121502014. View Article : Google Scholar
|
|
26
|
Sun X, Dai G, Yu L, Hu Q, Chen J and Guo
W: miR-143-3p inhibits the proliferation, migration and invasion in
osteosar-coma by targeting FOSL2. Sci Rep. 8:6062018. View Article : Google Scholar
|
|
27
|
Ling L, Zhang SH, Zhi LD, Li H, Wen QK, Li
G and Zhang WJ: MicroRNA-30e promotes hepatocyte proliferation and
inhibits apoptosis in cecal ligation and puncture-induced sepsis
through the JAK/STAT signaling pathway by binding to FOSL2. Biomed
Pharmacother. 104:411–419. 2018. View Article : Google Scholar : PubMed/NCBI
|