Introduction
Collagen VI is an important component of the
extracellular matrix (ECM). It forms a microfibrillar network that
is usually found to be in close association with the cell and
surrounding basement membrane. Collagen VI is also found in the
interstitial space of numerous tissues, including muscles, tendons,
skin, cartilage and intervertebral discs (1). Collagen VI is encoded by six
different genes, namely collagen α-1 (VI) chain (COL6A1),
COL6A2, COL6A3, COL6A4, COL6A5 and
COL6A6. Collagen VI-related myopathy (COL6-RD) is caused by
pathogenic variants in three collagen VI genes: COL6A1,
COL6A2 and COL6A3. These mutations can result in two
main types of muscle disorders that can range from severe forms of
Ullrich congenital muscular dystrophy (UCMD) to milder forms of
Bethlem myopathy (BM) (2).
A child with typical UCMD cannot walk independently,
or can only do so for short periods of time. These patients also
present with changes to their skin and joints, including variable
proximal contractures and distal hyperlaxity, congenital hip
dislocations, follicular hyperkeratosis over some of the extensor
surfaces of the limbs and soft velvety skin on their palms and
soles of their hands and feet. In addition, there is usually
abnormal wound healing, which can result in the formation of keloid
scars. BM is a relatively mild disorder, and can be characterized
by some proximal muscle weakness and contractures, with the onset
of these symptoms occurring within the first two decades of life
(2). These two major phenotypes
are not distinct and an intermediate phenotype, described as mild
UCMD with severe BM, has been defined by number of previous studies
(2,3). BM is transmitted as an autosomal
dominant or autosomal recessive inherited disorder and is usually
associated with mutations in the COL6 genes (4,5).
The prevalence of BM in the northern region of England in 2007 was
0.77/10,000 (6).
The distribution of the mutations in the COL6
genes is somewhat uniform, and lacks mutation hot spots. The most
common mutations in COL6-related myopathy are splicing and missense
mutations (3,7). As a consequence of this notable
heterogeneity at the clinical and molecular levels in the
COL6 genes, it appears to be difficult to establish a
correlation between the phenotype and the genotype of this disease.
In a previous study, dominant splice-site mutations that cause exon
skipping were observed in cases classed as moderate to progressive
(8).
Two common splice sites, the canonical splice donor
GT and acceptor AG, are present in 98.71% of all mammalian genes
(9). Mutations in these
canonical dinucleotides always result in splicing errors when
pre-mRNA transcripts are transformed into mature mRNAs (10). It was previously reported that
sequence variations surrounding certain canonical splice sites may
not result in splicing errors because these exon and intron
elements are only weakly conserved, and their alterations do not
always disrupt processes related to splicing (11). Therefore, it is essential to
determine the presence of aberrant mRNAs in order to predict the
clinical phenotypes of diseases. RNA-sequencing uses
high-throughput sequencing technology to rapidly and
comprehensively obtain the transcriptional status of biological
samples at a specific time (12). In the present study,
RNA-sequencing was successfully used to analyze the influence of
splice mutations.
The current study reported three patients from a
family who presented with typical clinical features of BM. A novel
heterozygous COL6A2 mutation (c.736-1G>C) was identified
in this family. The mechanism of this mutation has not been
previously reported in BM, to the best of our knowledge. Adiagnosis
of BM was made by observation of the clinical characteristics and
MRI features of the patients. Bioinformatics, in silico
analysis, RNA-sequencing, reverse transcription (RT)-PCR and
immunocytochemistry were performed to confirm the pathogenicity and
mechanism of the mutation. The present findings may help improve
the accurate molecular diagnosis of the disorder, and illustrated a
previously unrecognized category of BM.
Materials and methods
Patients and their families
This study was from January 2015 to August 2018. The
clinical data of the affected father (proband) and two sons were
collected at The First Affiliated Hospital of Guangxi Medical
University (Nanning, China). Three male patients age ranged from 40
to 14 years. The median age was 24 years. The family had nine
members, the parents, two sisters and a brother of the proband, his
wife and two sons. Information such as symptoms at onset and
creatine kinase levels were determined by means of a commercially
available Creatine Kinase reagent kit (cat. no. 23-666-208; Thermo
Fisher Scientific, Inc.) via rate-assay spectrophotometry according
to the manufacturer's instructions, as well as ultrasonic
cardiogram results, were obtained. Muscle MRI was performed using
the thighs of the three patients and a normal control (the
grandfather). The MRI results were assessed by two independent and
experienced radiologists and a neurologist. The right gastrocnemius
was biopsied in the father. Routine periodic acid-Schiff (PAS)
staining was performed on muscle cryostat sections to investigate
the presence of normal and abnormal glycogen levels.
Biological materials
All the biological materials, including peripheral
blood samples obtained from the cubital vein, skin biopsies, muscle
biopsies as well as fibroblasts for cell culture, were collected
after the appropriate informed consent was obtained. The use of all
appropriate human tissues was approved by The Medical Ethics
Committee of the First Affiliated Hospital of Guangxi Medical
University (Nanning, China; approval no. 2017-KY-E-154) and the
experimental protocols were performed in accordance with the
institutional guidelines and regulations. All nine family members
donated venous blood samples Genomic DNA was extracted with Qiagen
FlexiGene DNA kit (cat. no. 51206; Qiagen China Co., Ltd.)
according to the manufacturer's protocol. A muscle biopsy was
obtained from the proband and skin biopsies were obtained from the
inner side of the thigh in the proband and a son. Normal skin and
muscle tissues were collected from two male patients (median age 24
years) receiving skin or muscle resections for non-muscular
disease-related reasons, including circumcisions. Fibroblasts were
cultured from the skin biopsies and were used as primary
cultures.
DNA analysis
To systematically search for the disease-causing
genes, a myopathy panel based on next-generation sequencing was
performed on the proband. The myopathy panel (Table SI) was a kit designed by the
Zhongguancun Huakang Gene Institute (http://www.kangso.net/) and produced using the Agilent
SureSelect Target Enrichment technique (13).
High-throughput single-end sequencing was performed
on an Illumina NextSeq500 platform (Illumina, Inc.). In order to
construct the exome library, 4 µg genomic DNA from
peripheral blood samples of each patient was sheared into fragments
of 150-250 bp in length by sonication (30 sec on/30 sec off, 0°C
for 5 min) and subsequently hybridized for enrichment, using the
manufacturer's standard protocol (SureSelect Target Enrichment
System Target Sequence Enrichment kit, cat. no. 5190-5931;
Agilent). Indexing primers with 8 bp index A01-H12 were used to
amplify the captured library, and PCR was performed to amplify the
SureSelect-enriched DNA library. The amplified captured library was
purified using AMPure XP magnetic beads. To evaluate the DNA
quality and quantity, it was necessary to use the 2100 Bioanalyzer
instrument (Agilent) and a High Sensitivity DNA kit (Agilent)
according to the manufacturer's protocols. The final concentration
of the final library was >1.52×10−6 mmol/l. After
obtaining the human reference genome from the University of
California, Santa Cruz database (build 37.1, version hg19,
http://genome.ucsc.edu/), sequence alignment was
performed using the Burrows-Wheeler Alignment tool (14). Picard (http://sourceforge.net/projects/picard/) was then
employed to mark duplicates resulting from PCR amplification. All
candidate mutations were subsequently filtered against several
programs, including the single nucleotide polymorphism database
(http://www.ncbi.nlm.nih.gov/projects/SNP/snp_summary.cgi),
the international HapMap Project (http://hapmap.ncbi.nlm.nih.gov/) and the 1,000 genomes
project (2012 April release, http://www.1000genomes.org/) in order to remove loci
polymorphisms. Polymorphism phenotyping version 2 (PolyPhen-2) was
used for sorting intolerant from tolerant and mutation tasters to
predict whether an amino acid substitution could affect the
functions of the protein. Next, Sanger sequencing was used to
validate whether the identified potential disease-causing variant
occurred in the individual family members (the proband's parents,
brother, two sisters, two sons and his wife).
In silico analysis
To evaluate the probability of splice site mutations
being able to disrupt the normal splicing of COL6A2, two
online bioinformatics tools were used for prediction studies: Human
Splicer Finder (http://www.umd.be/HSF/) and Alternative Splice Site
Predictor (http://wangcomputing.com/assp/index.html). ExPASy
(https://www.expasy.org/)was used to predict their
impact on the protein that they would produce.
RNA-sequencing analysis
The effect of the splicing mutation on COL6A2
mRNA was determined by RNA-sequencing analysis. Tissue from the
muscle of the proband and a non-muscle disease control were
selected to extract total RNA for RNA-sequencing analysis.
Construction of the sequencing libraries followed the standard
protocols of the TruSeq™ RNA Sample Preparation guide (Illumina,
Inc.). TruSeq® Stranded mRNA Library Prep kit (Illumina,
Inc.) was used to prepare the sequencing libraries. The poly-A
containing mRNA molecules are purified from total RNA using poly-T
oligo-attached magnetic beads. During the elution of the poly-A
RNA, the RNA is also fragmented to 120-200-bp and primed for cDNA
synthesis. First-strand cDNA was then synthesized with SuperScript
II Reverse Transcriptase (Invitrogen; Thermo Fisher Scientific,
Inc.) using random primers. The temperature protocol was follows:
25°C For 10 min, 42°C for 15 min and 70°C for 15 min. AMPureXP
beads (Beckman Coulter, Inc.) were then used to isolate
double-stranded cDNA, which was synthesized using Second Strand
Master mix (Invitrogen; Thermo Fisher Scientific, Inc.) according
to the manufacturer's instructions. The adapters were then ligated
to the A-tailing fragment. Subsequently, 12 cycles of PCR were
performed to enrich those DNA fragments that had adapter molecules
at both ends following the program: 98°C For 30 sec; 15 cycles of
98°C for 10 sec, 60°C for 30 sec, 72°C for 30 sec and 72°C for 5
min. This technique also amplified the quantity of DNA in the
library. Purified libraries were further quantified using a
Qubit2.0 fluorometer (Invitrogen; Thermo Fisher Scientific, Inc.)
and then validated using an Agilent 2100 bioanalyzer (Agilent
Technologies, Inc.). Clusters were then generated using cBot with
the libraries, which were further diluted to 10 pM and were
subsequently sequenced using the Illumina HiSeq x Ten system
(Illumina, Inc.) for 150 cycles. Shanghai Sangong Biotech
Corporation (https://www.sangon.com/) was employed
to construct the libraries, and Illumina sequencing was performed
for the high-quality reads, which were kept for sequence analysis
after passing through Illumina quality filters. The sequencing data
sets are stored at http://bioinformatics.jnu.edu.cn/software/sequencing_datasets/
and Gene Expression Omnibus database (accession number, GSE42006)
for reference. Integrative Genomics Viewer (version 2.8.0) was used
to display the genomic data.
Skin fibroblast cultures
To extract total RNA for transcript analysis, skin
biopsies from two patients (proband and his elder son) and normal
controls were immersed into Dulbecco's modified Eagle's medium
(cat. no. 22390; Gibco; Thermo Fisher Scientific, Inc.)
supplemented with 20% heat-inactivated fetal bovine serum (cat. no.
A11-043; PAA Laboratories GmbH; GE Healthcare) and
penicillin-streptomycin (cat. no. 15070-063; Gibco; Thermo Fisher
Scientific, Inc.). Fibroblasts from skin biopsies were cultured in
Dulbecco's modified Eagle's medium containing 20% fetal bovine
serum (Sigma-Aldrich; Merck KGaA) and penicillin-streptomycin
(Sigma-Aldrich; Merck KGaA) at 37°C in an atmosphere of 95% air and
5% CO2 (15).
Transcript analysis
In order to validate the results of RNA-sequencing
analysis and the effect of the splice site mutation, transcript
analysis was performed using RT-PCR with RNA from fibroblasts
derived from two patients (proband and his elder son) and two
healthy individuals as controls. TRIzol® (0.5 ml;
Invitrogen; Thermo Fisher Scientific, Inc.) was used to extract
total RNA using the standard manufacturer's instructions.
In order to reveal the COL6A2 cryptic splice
acceptor, RT-PCR was performed with a forward primer within the 3′
of exon 3 and a reverse primer within exon 5. The primer pairs were
as follows: Forward, 5′-CTC TAC CGC AAC GAC TAC GC-3′ and reverse,
5′-TTC CAG GCA GCT CAC CTT-3′. RT-PCR, revealing COL6A2 exon
skipping, was performed using a forward primer within exon 3 and a
reverse primer within exon 6. The primer pairs were as follows:
Forward, 5′-CCC AAC CAG AAC CTG AAG GA-3′ and reverse, 5′-TCG GCT
CCA AAT TCA CCC T-3′. PCR was conducted with Premix Taq™ DNA
Polymerase (Takara Biotechnology Co., Ltd.) under the following
conditions: Initial denaturation at 95°C for 4 min, followed by 36
cycles at 95°C for 30 sec, 56°C for 30 sec and 72°C for 40 sec. PCR
products were separated on 2% agarose gels and extracted using a
gel purification kit (Favorgen Biotech Corp.) according to the
manufacturer's protocols. The eluted products were sequenced using
an ABI 3730 XL Genetic Analyser with BigDye™ Terminator Cycle
Sequencing kit (Applied Biosystems; Thermo Fisher Scientific,
Inc.). The COL6A2 sequence of the patients was compared with
that observed in the control and reference sequences of
COL6A2.
Immunohistochemistry of muscle biopsies
and immunocytochemistry of skin fibroblasts
The morphology was observed under a microscope via
hematoxylin and eosin (H&E) staining. Fresh specimens were
fixed in 10% neutral buffered formalin and further processed into
paraffin-embedded blocks, sectioned at 6 -µm thickness,
dehydrated in graded ethanol (70, 80, 90, 95 and 100%) solutions
and processed according to the H&E staining protocol (16). The stained tissues were observed
using a biomicroscopy imaging system microscope (cat. no. DP73;
Olympus Corporation). Samples derived from skin fibroblast cultures
and muscle biopsies were subsequently prepared for
immunohistochemistry to evaluate collagen VI protein expression.
Briefly, sections were incubated at 4°C overnight with the primary
antibody which was a monoclonal antibody raised in rabbits (clone
#EPR17072) that targeted collagen VI (cat. no. ab182744; 1:100;
Abcam), and then incubated at room temperature for 30 min with
Supervision™ Universal (Anti-Mouse/Rabbit) Detection reagent (HRP)
(cat. no. D-3004; Thermo Fisher Scientific, Inc.) conjugated to
peroxidase in Tris-HCI buffer containing carrier protein and
anti-microbial agent. The experiment was performed according to the
manufacturer's protocol.
Results
Three patients in a family display
typical features of BM
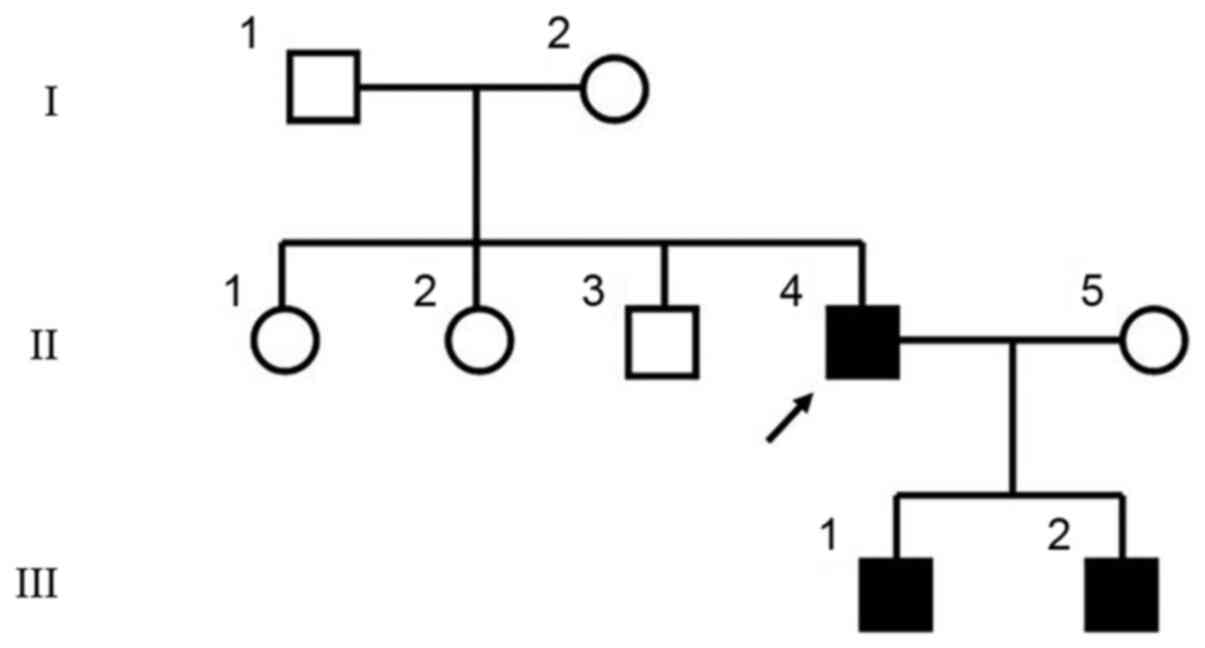
The three patients in this study were from three
generations of a Chinese family (Fig. 1). The proband (II-4) was 40 years
old. From 4 years of age, he experienced difficulty getting up from
the floor and climbing stairs; at 8 years of age, he began toe
walking and needed to use a rail to climb stairs. He underwent
Achilles tendon release aged 8 years. He had mild weakness in the
shoulder girdle and upper limb muscles, and moderate weakness in
the axial and hip girdle muscles. At the age of 40, he was able to
walk unassisted but with a marked Trendelen burg gait. Physical
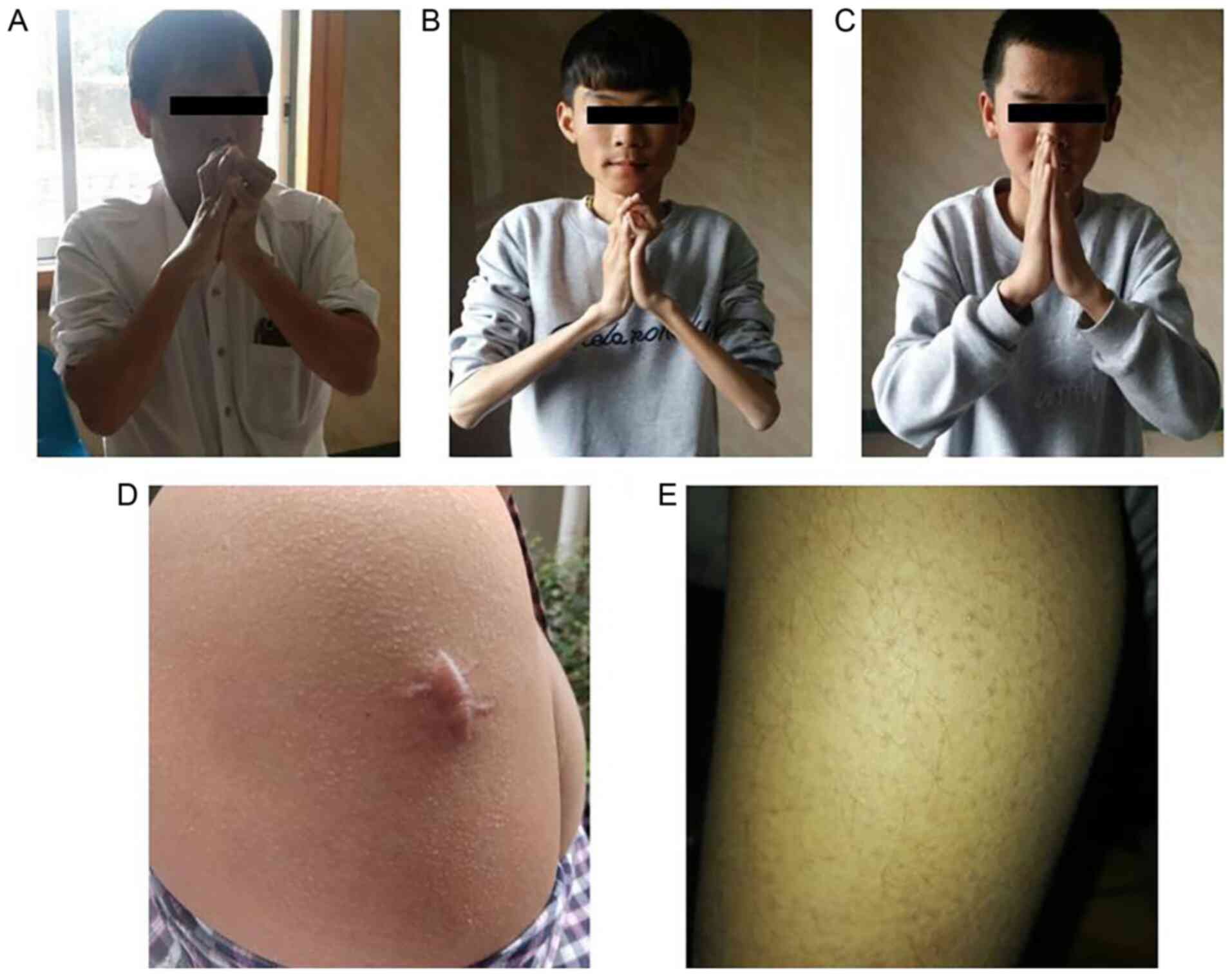
examination revealed proximal muscle weakness and amyotrophy. Joint
contractures of the shoulders, hips, knees and feet were observed
(Fig. 2A). Skin examination
revealed that he had keratosis pilaris along the extensor surfaces
of his arms and legs.
Two sons (III-1 and III-2) of the proband shared
similar clinical symptoms. The father was the most severe case, the
younger brother the mildest. Examination in our hospital occurred
when the older boy was 18 years old. He could not get up from the
floor. The younger 14-year-old brother experienced difficulty when
getting up from the floor and climbing stairs, and he walked on his
toes (Table I). Both underwent
Achilles tendon release 10 years of age.
 | Table IClinical data and biochemical results
of three patients in the family. |
Table I
Clinical data and biochemical results
of three patients in the family.
| Variable | Patient
|
|---|
| II:4 | III:1 | III:2 |
|---|
| Age, years | 40 | 18 | 14 |
| Approximate age at
which muscle weakness was diagnosed, years | 4 | 4 | 5 |
| Capable of climbing
stairs | No | No | No |
| Capable of walking
100 m | Yes | Yes | Yes |
| Capable of jumping
up and down for 1 min | No | No | No |
| Capable of standing
for 10 min | No | No | No |
| Skin
appearance | Some scarring | Keratosis
pilaris | Normal |
| Rigidity of
spine | No | No | No |
| Plasma creatine
kinase levels, U/l | 576 | 747 | 759 |
| Ultrasonic
cardiogram | Normal | Normal | Normal |
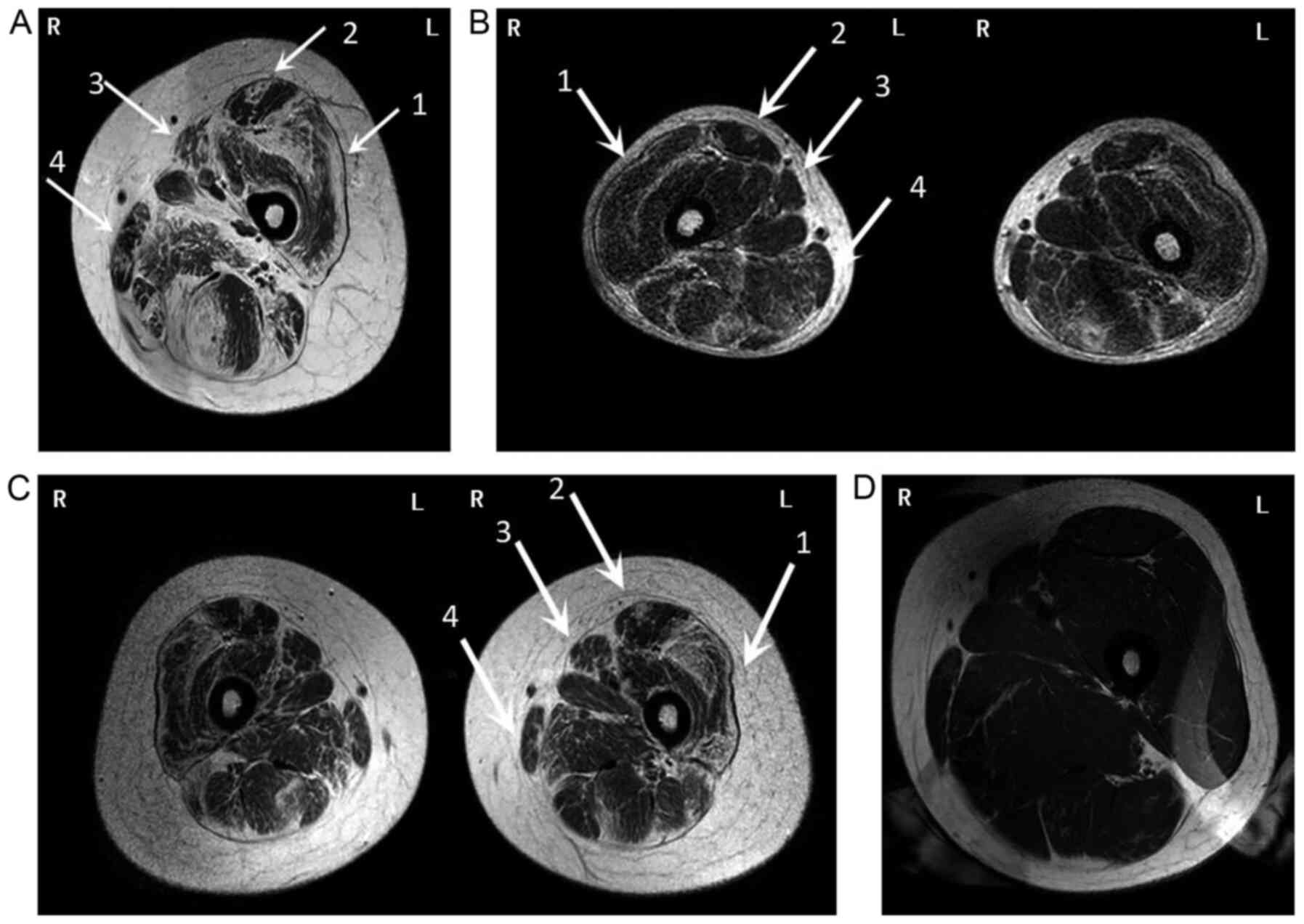
To further demonstrate the patients' pattern of
muscle involvement and to determine the potential relevance of
these unknown genetic variants, muscle MRI for the lower
extremities was recommended. This revealed a distinct pattern of
fatty infiltration in both lower extremities, and the anterior
group muscles were more impaired than the rear groups in the thigh
(Fig. 3A-C). This is consistent
with a pattern of symptoms termed 'central shadow' in the rectus
femoris and a pattern further described as 'outside-in' in the
vastus lateralis muscle. Fatty infiltration of the thigh muscles
varied greatly among the three patients. The father (Fig. 3A) was the most severe case, while
the youngest son (Fig. 3C)
exhibited the mildest symptoms.
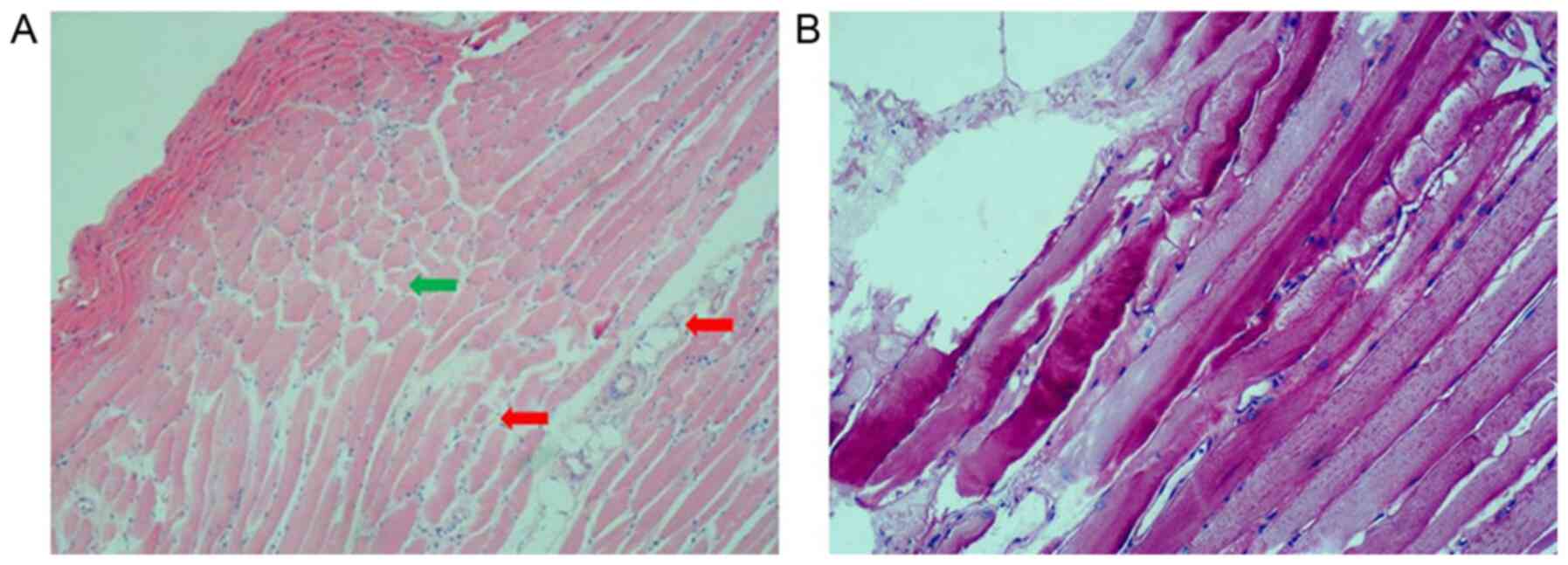
H&E staining of the gastrocnemius muscle of the
proband showed mild dystrophic features, such as muscle
degeneration and necrotic fibers. In addition, connective tissue
hyperplasia was observed in the muscle fibers and these were
unequal in size and had certain structural distortion (Fig. 4A). Abnormalities of glycogen
deposition were not observed when tissue sections of muscle were
stained with PAS staining (Fig.
4B).
Identification of a heterozygous splicing
mutation in the COL6A2 gene
To make a genetic diagnosis, 2.84 million 75-bp
single-end sequence reads were generated from the proband. Of
these, 2.83 million reads (99.93%) aligned to the human reference
genome. Of the 169 selected genes (Table SI), a heterozygous splicing
mutation (c.736-1G>C) in the COL6A2 gene was identified
in the proband and Sanger sequencing validated that the same
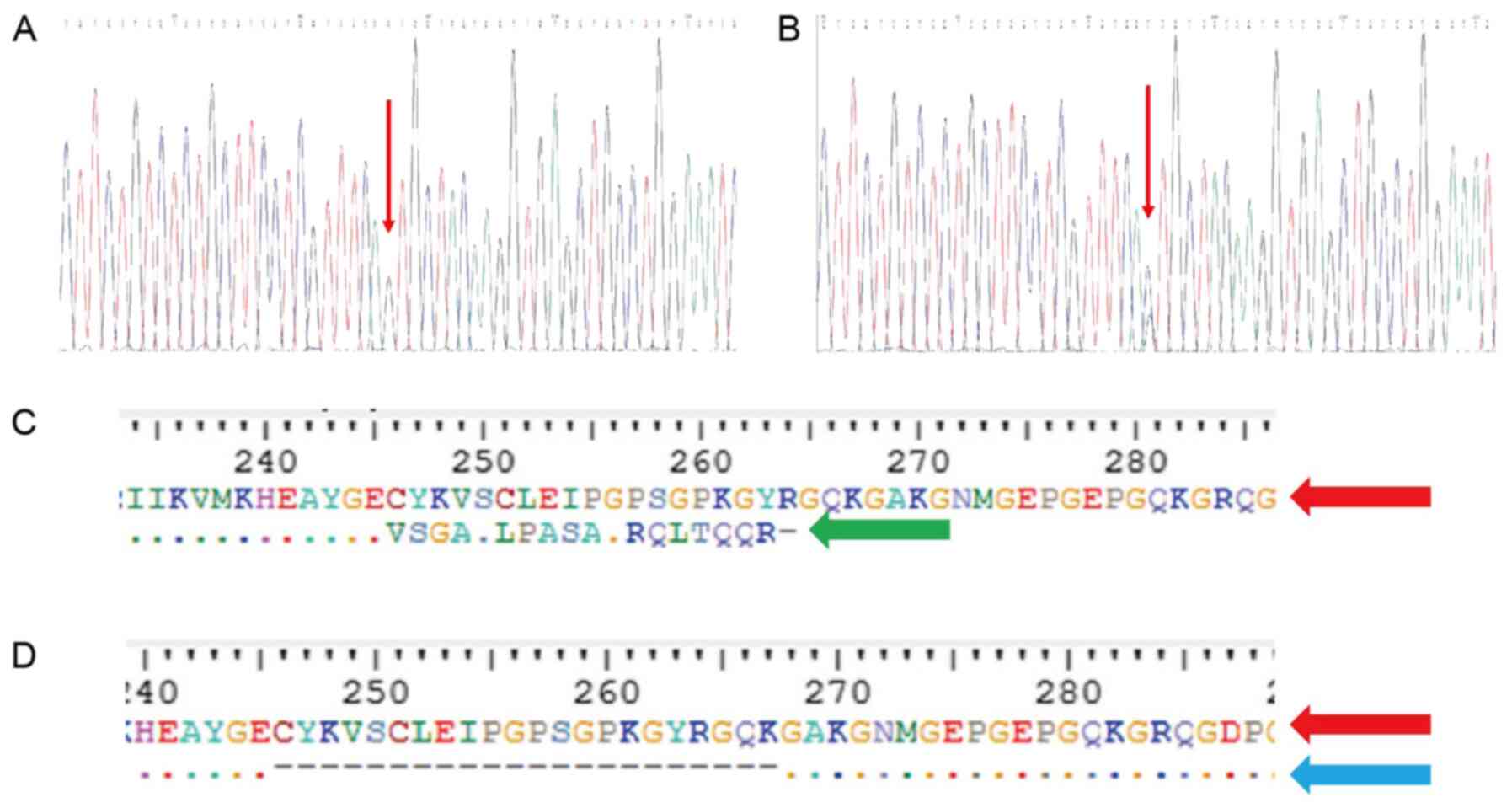
mutation existed in the two sons (Fig. 5A), while the spouse (II-5),
proband's siblings (II-1, 2 and 3), father (I-1) and mother (I-2)
were normal (Fig. 5B). The
members who had the mutation in the COL6A2 gene had a
similar phenotype, while those without this mutation did not have
muscle weakness or motor disability. This splice acceptor site
mutation observed in intron 4 was co-segregated within the
family.
Splicing mutations can result in aberrant
splicing
In silico analysis
In order to determine the effects of the mutation on
the splicing of COL6A2 mRNA, in silico and in
vitro analyses, which involved RNA-sequencing analysis and
RT-PCR, was performed.
Human Splicer Finder and Alternative Splice Site
Predictor, which are two freely available online bioinformatics
tools, were used to predict the effect of the c.736-1G>C
variation on COL6A2 mRNA splicing. Human Splicer Finder
shown that wild-type sequencing existed a normal splice acceptor
sites of 3′ end; while the normal acceptor sites disappeared and
replaced by a new one in the mutant-type. The same result was found
by Alternative Splice Site Predictor. Wild-type sequencing existed
a splice acceptor sites of 3′ end; while the acceptor sites
disappeared in the mutant-type. These tools suggested that this
mutation would probably result in the elimination of any wild-type
splice acceptor sites that would create a new cryptic splice
acceptor site. Thus, a 129 bp sequence at the 3′ end of intron 4
was retained (data not shown).
Prediction of influence on the
protein
To understand the influence of the two abnormal
splicing variants, ExPASy was used to predict their impact on the
protein that they would produce. The new cryptic splice acceptor
site obtained suggested that the loss of the wild-type splice site
could result in a frame-shift in amino acid 246, which would
eventually lead to a premature stop codon in amino acid 264
(p.Cys246fs264*) of the protein. The skipping of exon 5 appeared to
result in the deletion of 21 amino acids, namely from 246 to 267,
in the protein (p.Cys246_Lys267del) (Fig. 5C and D). The truncated protein
lacked the triple-helical domain (THD) region.
RNA-sequencing analysis
Integrative Genomics Viewer was used to display the
genomic data. In addition to the normal base sequence, two abnormal
splicing variants adjacent to the mutation site were identified in
the COL6A2 gene. One variant had a 141-bp sequence retained
at the 3′ end of intron 4, causing a new cryptic splice acceptor
site. Another variant was due to exon 5 skipping. The percentage of
presence of the normal, 3′ end of intron 4 retention and exon 5
skipping variants was 50, 36 and 14%, respectively (Table II). The coverage (reads) of the
patient was lower than compared with that of the control.
Differential gene expression analysis revealed that the expression
of COL6A2 in the patient was ~5-fold higher compared with that of
the normal control.
 | Table IICoverage of splicing variants in
proband and normal control. |
Table II
Coverage of splicing variants in
proband and normal control.
| Splicing
variant | Patient
| Normal control
|
|---|
| Reads | Percentage, % | Reads | Percentage, % |
|---|
| Wild-type
transcripts sequence | 18 | 50 | 4,846 | 99.5 |
| Intron 4
retention | 13 | 36 | 18 | 0.4 |
| Exon 5
skipping | 5 | 14 | 7 | 0.1 |
RT-PCR
To validate the results demonstrated by
RNA-sequencing analysis, RNA derived from cultured skin fibroblasts
from the proband and his elder son, and two healthy controls were
used to perform RT-PCR experiments. The difference between the
wild-type transcripts and the mutant transcripts in patients was
that the wild-type transcripts were longer (which was caused by a
cryptic splice acceptor site activation) or shorter (which was
caused by exon skipping) compared with the mutant transcripts when
identical primers were used to amplify the PCR products that
flanked the region of mutation. Therefore, this method can serve as
a practical tool to distinguish the two transcripts by when the
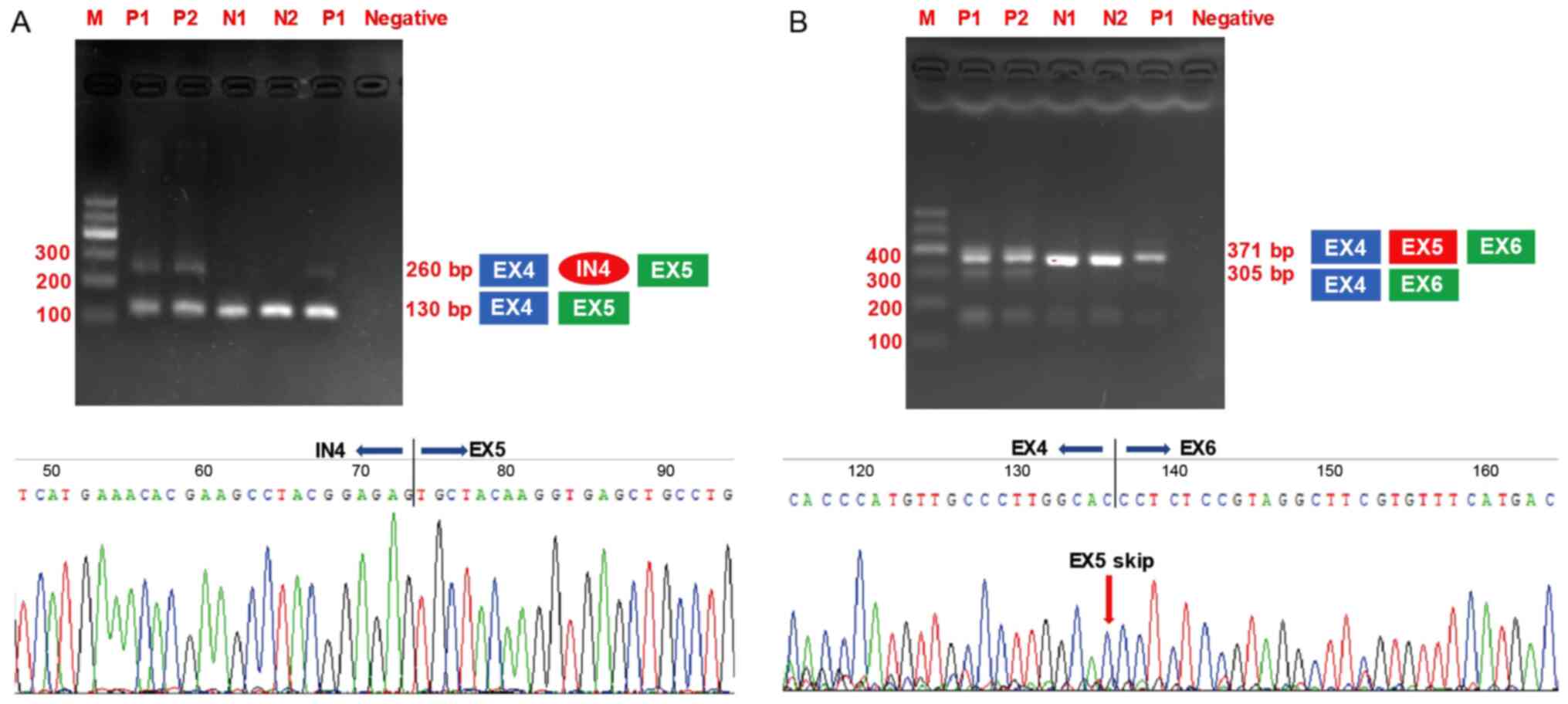
samples are subjected to gel electrophoresis. The amplification of
exon 3 to exon 5 in COL6A2 generated a 130-bp product for
all the cDNA samples from the controls and patients. Notably, an
additional larger product of 260 bp was only detected in the cDNA
samples obtained from the patients (Fig. 6A). The RT-PCR products were
subjected to gel extraction and Sanger sequencing, which showed
that a 129-bp sequence from the 3′ end of intron 4 was retained
between exon 4 and exon 5. This result was consistent with the
in silico analysis. RT-PCR amplification of exon 3 to exon 6
also produced a 371-bp product when using cDNA samples from either
the control individuals or the patients as templates. However, a
smaller product of 305 bp was only detected in the patients'
samples (Fig. 6B). Gel
extraction and Sanger sequencing also revealed that exon 4 directly
spliced the in-frame to exon 6 and hence skiped exon 5.
Collagen VI is absent in fibroblasts but
is expressed normally in muscle
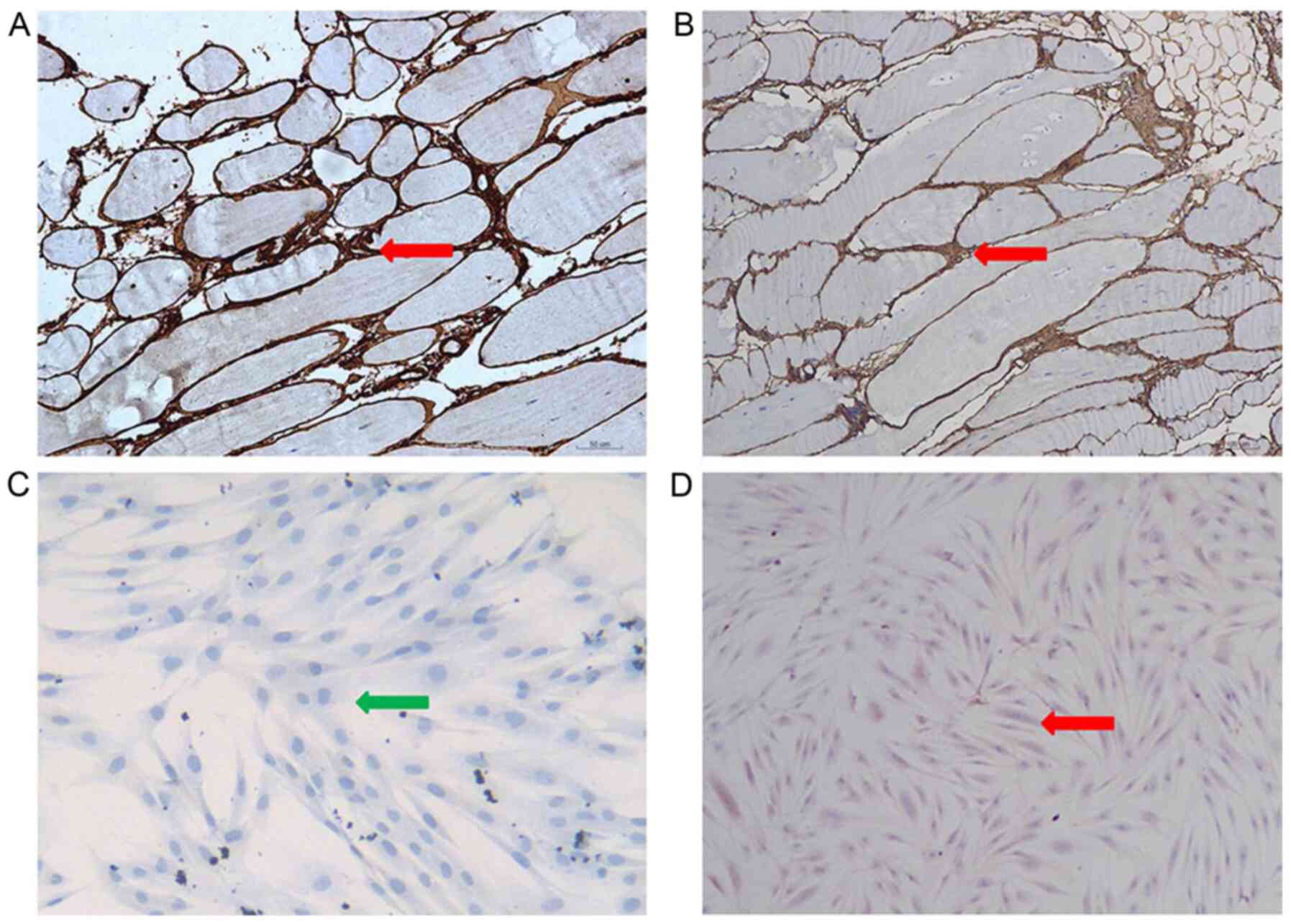
To further investigate the consequences of the
c.736-1G>C mutation on the protein product, immunohistochemical
analysis was performed using a rabbit monoclonal antibody against
collagen VI (cat. no. EPR17072). Collagen VI immunohistochemistry
indicated that this protein was absent in the skin fibroblasts
derived from patients but showed a normal pattern in the
fibroblasts derived from the controls. In contrast, a normal
pattern was found in the patient muscle fibers, attesting to the
integrity of the basement membrane (Fig. 7C and D). Fig. 7C and D do not have the same
background with stain. The difference between the two figures may
be due to non-specific factors, for example the degree of
pathological staining.
Discussion
BM is a well-defined collagen VI disorder. BM has
been associated with heterozygous, dominant and recessive mutations
in one of the COL6A1, COL6A2 or COL6A3 genes,
and is either inherited from an affected parent or occurs de
novo (5). The present study
was able to identify a heterozygous mutation (c.736-1G>C) in a
splice-site of the COL6A2 gene within a Chinese family. The
pathogenicity of this mutation has not been reported previously, to
the best of our knowledge. Three patients in the family shared
similar classical BM presentation and disease progression,
presenting with an early onset, slow progression and a relatively
mild disease. The severity of the clinical symptoms had a tendency
to worsen with advancing age.
The majority of patients with COL6-RD exhibit a
pathogenomic pattern in muscle MRI scans, and this is particularly
noticed in patients with certain degree of muscle weakness
(17). Patients with BM can
exhibit patterns that were termed 'central shadow' in the rectus
femoris and 'outside-in' in the vastus lateralis muscle in previous
reports (2). In the present
study, the presentation of muscle MRI scans were similar to the
previous study; fatty infiltration of the thigh muscles varied
greatly among the three patients studied. The degree of fatty
infiltration observed in the MRI scans was associated with the
severity of the clinical symptoms. Thigh muscle MRI scans can also
be used to assess the progression of the disease in patients with
BM (18).
To confirm the clinical findings, a myopathy panel
was performed on the proband. A heterozygous mutation
(c.736-1G>C) was identified in a splice-site of the
COL6A2 gene. This mutation has not been previously reported
in previous literature, to the best of our knowledge. The splice
acceptor site mutations at positions 1 and 2 are considered
pathogenic In silico analysis predicted that c.736-1G>C,
which is a splice site mutation, likely damages the splice acceptor
site. This evidence supports the likely pathogenicity of the novel
splice-site mutation.
To identify the aberrant splice variant caused by
the splice-site mutation, RNA-sequencing was used to analyze the
muscle obtained from the proband. The results revealed a 129-bp
retention of the 3′ end of intron 4 and skipping of exon 5.
Finally, RT-PCR was performed to validate the results of
RNA-sequencing. The splice site mutation (c.736-1G>C) destroyed
the normal splice acceptor site, which led to two main
consequences. Firstly, the activation of a new cryptic splice
acceptor site in intron 4 caused a frameshift and ultimately
resulted in premature termination of the protein. Secondly, there
was a skipped sequence of 66 bp on exon 5, which introduced a
deletion comprising of 22 amino acids. However, this resulted in no
alteration of the open reading frame of COL6A2. As reported in a
previous study, the mutant mRNA, exon skipping or the retention of
the intron in the mRNA was detected, which was similar to the
mutant mRNA in our study. It formed was degraded by the nonsense
mediated decay (NMD) pathway (19). Even if the present protein
escaped the NMD pathway, the truncated protein lacked the
triple-helical domain (THD) region. Hence, these variants may
result in aberrant splicing and affect the structure of collagen
VI.
Differential gene expression analysis revealed that
the expression of COL6A2 in the patient was ~5-fold higher compared
with that of the normal control. The low level of COL6A2 mRNA most
likely resulted from NMD in one allele. In a previous report,
nonsense-mediated decay was found in patients with splicing
mutations. Lucarini et al (20) reported a patient with UCMD and a
homozygous A to G transition at position -10 of intron 12. Northern
blot hybridization reported that the COL6A2 mRNA level in the
patient was substantially reduced compared with that in the normal
control. The data suggested that the majority of the COL6A2 mRNA in
the patient's fibroblasts was degraded, most likely through
nonsense-mediated mRNA decay. This possibility was further
investigated. The patient's fibroblasts were treated with
cycloheximide to inhibit RNA degradation. Next, RT-PCR analysis of
total RNA prepared from the patient's fibroblasts indicated that
aberrant splicing had occurred. Zhang et al (21) reported the existence of three
patients from two families with UCMD resulting from recessive
mutations of the COL6A2 gene. A splice acceptor site
mutation in intron 17 was found in two of the patients, causing NMD
in patients and in their unaffected parents. Fibroblasts from the
patients expressed low levels of COL6A2 mRNA. In the parents, the
COL6A2 mRNA levels were reduced to 57-73% of those of the control.
In the present study, the splicing mutation led to a truncated
protein. To avoid the accumulation of the truncated protein, NMD
may decrease the level of aberrant mRNA. This might be the reason
why the patients in the present study had a mild phenotype.
In order to identify the consequence of the mutation
at the protein level in the current study, immunohistochemical
analyses of the patients' muscles and fibroblasts were performed.
Collagen VI was expressed differently between these two samples.
This finding is consistent with those of previous studies. In
patients with BM, muscle biopsies immunostained with various
anti-collagen VI antibodies tended to be positive and therefore
could not be used for diagnostic purposes (22). This phenomenon also occurs in
other muscular dystrophies. For example, in patients with
limb-girdle muscular dystrophy1B, lamin A/C labeling of muscle
tissue is a common event (23).
The cause of this phenomenon is unclear. It can be hypothesized
that the protein expression is normal but structure and function
are impaired. Further research is needed to confirm this
hypothesis. However, further tests, such as western blotting, on
the samples obtained from the patients, were not possible in our
study, due to the limited quantity of sample available.
COL6A2 encodes the α2 chain of type VI
collagen and comprises a signaling peptide domain and three
different von Willebrand factor type A domains. These are typically
protein binding domains that have been shown to bind ECM proteins
as well as several collagen units (1). In the patients with BM in the
present study, the splice-site mutation (c.736-1G>C) was located
within the α2 THD region. The majority of mutations in the THD
region reported previously were mostly associated with UCMD
(24), and their phenotypes were
different from those of the current patients. It is conceivable
that non-penetrance and incomplete expression could be the causes
for this difference. Therefore, the correlation between phenotype
and genotype is not clear in patients with COL6-RD. Zhang et
al (25) reported a mutation
(c.736-1G>A) at the same position in a patient with UCMD, who
acquired ambulation after a delay of 16 months. Subsequently, he
lost ambulation at the age of 8 years old. Hip dislocation,
torticollis, extended talipes and skin changes were also present in
this patient.
In contrast, the patients in the current study
presented only with mild symptoms. The proband patient was 40 years
old and still walking independently. It was hypothesized that the
fact that certain transcripts 'escape' exon 5 skipping (by
retaining intron 4 instead) contributes to reducing the load of the
dominant-negative protein isoform, whereas the c.736-1G>A
mutation primarily produces the exon 5-skipped variant, which could
explain the reportedly severe phenotype. Finally, it should be
noted that similar types of mutations or mutations within the same
domains of the protein do not necessarily result in comparable
phenotypic severity of the disease, since these have been
identified in patients with differing clinical severity by the
present group and other researchers (26).
The interpretation of numerous splicing variants of
unknown biological and clinical significance found in general
genetic screenings represents a major challenge when attempting to
assess the molecular diagnosis of genetic diseases (12). A fraction of splicing mutations
may be deleterious because they may eventually affect mRNA splicing
(27). The majority of online
splice prediction programs tend to offer tools that predict
putative 5′-splice donor and 3′-splice acceptor sites (including
several cryptic splice sites). Additionally, these will give
probability scores or potential alterations of the splicing,
including Splice-Port, ESEfinder, BDGP Spice Site Prediction, Human
Splicing Finder and Alternative Splice Site Predictor (28). However, these programs do not
offer direct predictions of functional consequences of splice-site
mutations with regards to exon skipping, exon deletion or intron
retention (10). Researchers
often have to make their own predictions based on the calculated
probability scores. Some splice-site mutations can result in both
exon skipping and partial exon deletion, which is occasionally due
to activation of certain cryptic splice sites within the exon
(29). Therefore, it appears to
be unwise to predict the eventual functional consequences of these
alterations in the genomic sequence. Therefore, assays for
detecting mutant mRNA splicing isoforms are needed and can
determine the mechanism of mutation at a splice site.
According to the literature data, RT-PCR analysis
(29) of patient mRNA and
transfection of mini-gene constructs (30) expressing the mutated exons in
cell lines are two of the most widely used methods for validating
mutant mRNA splicing isoforms. RT-PCR and mini-gene assays can
probe the specific variant in question but do not reveal the
relative abundance of each of the different isoforms. In addition,
mini-gene construct expression only reveals the molecular
consequence of a splice-site mutation in vitro, while
alterations in vivo cannot be detected by mini-gene assays
(31). RT-PCR can identify any
mutant mRNAs directly, but is sophisticated for certain genes, such
as the dystrophin gene, which contain several alternative
promoters, markedly large introns and cryptic exons (32).
In the current study, RNA-sequencing was
successfully used to analyze the influence of splicing mutations.
This approach can directly identify the sequences of aberrant mRNAs
and the relative abundance of each isoform of the protein (12). This appears to be much simpler
and faster than RT-PCR and mini-gene assays. However, a limitation
of high-throughput sequencing is the existence of sequencing errors
(33). In the present study,
discrepancies between the RNA-sequencing and RT-PCR results were
observed. Specific RT-PCR was needed to validate the results of
RNA-sequencing. RNA-sequencing holds expanded promise for the
splicing mutation pathogenesis in various diseases, including
myopathy, developmental disorders and neurodegenerative disorders.
RNA-sequencing can be beneficial due to its diagnostic, prognostic
and therapeutic applicability (34). Our future studies will focus on
processing additional cases of BM in order to evaluate this method,
so that the specific impact of sequence variants on splicing can be
assessed.
Muscle biopsies are invasive operations (35). In our study, limited muscle
sample was obtained from the proband. The total RNA extracted from
the muscle was consumed by the RNA-sequencing and the rest of the
sample was used hematoxylin and eosin staining and immunochemistry.
Unfortunately, there was not enough RNA for RT-PCR to validate the
results of RNA-sequencing. BM is a disease involving both muscle
and skin. COL6 is abundantly expressed by skin fibroblasts, and
human skin biopsies and in vitro primary skin fibroblast
cultures have been widely used to characterize COL6 defects in BM
(5,15). Notably, fibroblast are relatively
easy to culture from the skin biopsies (36), therefore the present study
performed reverse transcription PCR using RNA from fibroblasts
derived from two patients and two healthy individuals as controls.
Our future studies will extract more total RNA for the RT-PCR
validation, aiming to compare the expression of the mRNA in muscle
and skin fibroblast.
In conclusion, the present study described a
BM-affected family resulting from a novel splice-site mutation
within intron 4 of the COL6A2 gene. Using a number of
bioinformatics and experimental methods, the hypothesis that the
c.736-1G>C mutation caused the BM phenotype in this family was
supported. These findings extend our present knowledge of the
possible mutations of the COL6A2 gene associated with BM.
The splicing mutation c.736-1G>C in the COL6A2 gene can
produce two different mRNA splice variants: One retaining intron 4
and one skipping exon 5 (in-frame), which possibly causes aberrant
splicing and leads to premature termination and truncation of
translation. Ultimately, this would affect the quaternary structure
of collagen VI. RNA-sequencing can be a powerful tool to determine
the underlying mechanism of a disease-causing mutation at a splice
site.
Supplementary Data
Availability of data and materials
The datasets generated and/or analyzed during the
current study are available in the Genome Sequence Archive
repository (https://bigd.big.ac.cn/gsa-human/). The results of
Human Splicer Finder and Alternative Splice Site Predictor. Human
Splicer Finder additional data are available upon request.
Authors' contributions
DL and JZ summarized the clinical information,
analyzed the genetic test results and drafted the manuscript. YX
helped to collect the clinical information and analyze the genetic
test results. YD assessed the pathological biopsy results and
performed the pathological biopsies. JZ collected the additional
clinical information. YS performed the MRI. All authors read and
approved the final manuscript.
Ethics approval and consent to
participate
All procedures performed in this study involving
human participants were conducted in accordance with the ethical
standards of the institutional and national research committees and
was approved by The Medical Ethics Committee of the First
Affiliated Hospital of Guangxi Medical University (Nanning, China;
approval no. 2017-KY-E-154). Written informed consent was obtained
from each participant included in the study or a parent.
Patient consent for publication
Consent for publication was also obtained from all
participants included in this study or a parent.
Competing interests
The authors declare that they have no competing
interests.
Acknowledgments
The authors would like to thank Dr Dev Sooranna,
Imperial College London, for editing the manuscript.
Abbreviations:
|
BM
|
Bethlem myopathy
|
|
COL6-RD
|
collagen type VI-related
dystrophies
|
|
ECM
|
extracellular matrix
|
|
MRI
|
magnetic resonance imaging
|
|
NMD
|
non-sense mediated decay
|
|
PolyPhen-2
|
Polymorphism phenotyping version 2
|
|
RT-PCR
|
reverse transcription-polymerase chain
reaction
|
|
THD
|
triple-helical domains
|
|
UCMD
|
Ullrich congenital muscular
dystrophy
|
References
|
1
|
Cescon M, Gattazzo F, Chen PW and Bonaldo
P: Collagen VI at a glance. J Cell Sci. 128:3525–3531. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Bushby KMD, Collins J and Hicks D:
Collagen type VI myopathies. Adv Exp Med Biol. 802:185–199. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Kim SY, Kim WJ, Kim H, Choi SA, Lee JS,
Cho A, Jang SS, Lim BC, Kim KJ, Kim JI, et al: Collagen VI-related
myopathy: Expanding the clinical and genetic spectrum. Muscle
Nerve. 58:381–388. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Foley AR, Hu Y, Zou Y, Columbus A,
Shoffner J, Dunn DM, Weiss RB and Bönnemann CG: Autosomal recessive
inheritance of classic Bethlem myopathy. Neuromuscular Disord.
19:813–817. 2009. View Article : Google Scholar
|
|
5
|
Gualandi F, Urciuolo A, Martoni E,
Sabatelli P, Squarzoni S, Bovolenta M, Messina S, Mercuri E,
Franchella A, Ferlini A, et al: Autosomal recessive Bethlem
myopathy. Neurology. 73:1883–1891. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Norwood FLM, Harling C, Chinnery PF, Eagle
M, Bushby K and Straub V: Prevalence of genetic muscle disease in
Northern England: In-depth analysis of a muscle clinic population.
Brain. 132:3175–3186. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Lee JH, Shin HY, Park HJ, Kim SH, Kim SM
and Choi YC: Clinical, pathologic, and genetic features of collagen
VI-related myopathy in Korea. J Clin Neurol. 13:331–339. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Briñas L, Richard P, Quijano-Roy S,
Gartioux C, Ledeuil C, Lacène E, Makri S, Ferreiro A, Maugenre S,
Topaloglu H, et al: Early onset collagen VI myopathies: Genetic and
clinical correlations. Ann Neurol. 68:511–520. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Burset M, Seledtsov IA and Solovyev VV:
Analysis of canonical and non-canonical splice sites in mammalian
genomes. Nucleic Acids Res. 28:4364–4375. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Ohno K, Takeda J and Masuda A: Rules and
tools to predict the splicing effects of exonic and intronic
mutations. Wiley Interdiscip Rev RNA. 9:2018. View Article : Google Scholar
|
|
11
|
Demir K, Kattan WE, Zou M, Durmaz E,
BinEssa H, Nalbantoğlu Ö, Al-Rijjal RA, Meyer B, Özkan B and Shi Y:
Novel CYP27B1 gene mutations in patients with vitamin D-dependent
rickets type 1A. PLoS One. 10:e01313762015. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Ozsolak F and Milos PM: RNA sequencing:
Advances, challenges and opportunities. Nat Rev Genet. 12:87–98.
2011. View
Article : Google Scholar :
|
|
13
|
Chen R, Im H and Snyder M: Whole-exome
enrichment with the agilent sureselect human all exon platform.
Cold Spring Harb Protoc. 2015:626–633. 2015.PubMed/NCBI
|
|
14
|
Li H and Durbin R: Fast and accurate short
read alignment with Burrows-Wheeler transform. Bioinformatics.
25:1754–1760. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Martoni E, Urciuolo A, Sabatelli P, Fabris
M, Bovolenta M, Neri M, Grumati P, D'Amico A, Pane M, Mercuri E, et
al: Identification and characterization of novel collagen VI
non-canonical splicing mutations causing ullrich congenital
muscular dystrophy. Hum Mutat. 30:E662–E672. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Wang C, Yue F and Kuang S: Muscle
histology characterization using H&E staining and muscle fiber
type classification using immunofluorescence staining. Bio Protoc.
7:e22792017. View Article : Google Scholar :
|
|
17
|
Fu J, Zheng YM, Jin SQ, Yi JF, Liu XJ, Lyn
H, Wang ZX, Zhang W, Xiao JX and Yuan Y: 'Target' and 'Sandwich'
signs in thigh muscles have high diagnostic values for collagen
VI-related myopathies. Chin Med J (Engl). 129:1811–1816. 2016.
View Article : Google Scholar
|
|
18
|
Mercuri E, Lampe A, Allsop J, Knight R,
Pane M, Kinali M, Bonnemann C, Flanigan K, Lapini I, Bushby K, et
al: Muscle MRI in Ullrich congenital muscular dystrophy and Bethlem
myopathy. Neuromuscul Disord. 15:303–310. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Vanegas OC, Zhang RZ, Sabatelli P,
Lattanzi G, Bencivenga P, Giusti B, Columbaro M, Chu ML, Merlini L
and Pepe G: Novel COL6A1 splicing mutation in a family affected by
mild Bethlem myopathy. Muscle Nerve. 25:513–519. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Lucarini L, Giusti B, Zhang RZ, Pan TC,
Jimenez-Mallebrera C, Mercuri E, Muntoni F, Pepe G and Chu ML: A
homozygous COL6A2 intron mutation causes in-frame triple-helical
deletion and nonsense-mediated mRNA decay in a patient with Ullrich
congenital muscular dystrophy. Hum Genet. 117:460–466. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Zhang RZ, Sabatelli P, Pan TC, Squarzoni
S, Mattioli E, Bertini E, Pepe G and Chu ML: Effects on collagen VI
mRNA stability and microfibrillar assembly of three COL6A2
mutations in two families with Ullrich congenital muscular
dystrophy. J Biol Chem. 277:43557–43564. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Jimenez-Mallebrera C, Maioli MA, Kim J,
Brown SC, Feng L, Lampe AK, Bushby K, Hicks D, Flanigan KM,
Bonnemann C, et al: A comparative analysis of collagen VI
production in muscle, skin and fibroblasts from 14 Ullrich
congenital muscular dystrophy patients with dominant and recessive
COL6A mutations. Neuromuscul Disord. 16:571–582. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Norwood F, de Visser M, Eymard B,
Lochmüller H and Bushby K; EFNS Guideline Task Force: EFNS
guideline on diagnosis and management of limb girdle muscular
dystrophies. Eur J Neurol. 14:1305–1312. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Lampe AK and Bushby KMD: Collagen VI
related muscle disorders. J Med Genet. 42:673–685. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zhang YZ, Zhao DH, Yang HP, Liu AJ, Chang
XZ, Hong DJ, Bonnemann C, Yuan Y, Wu XR and Xiong H: Novel collagen
VI mutations identified in Chinese patients with Ullrich congenital
muscular dystrophy. World J Pediatr. 10:126–132. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Yang HP, Zhang YZ, Ding J, Jiao H, Lü JL
and Xiong H: Clinical and mutation analyses of a Chinese family
with Bethlem myopathy. Zhonghua Yi Xue Za Zhi. 92:2820–2824.
2012.In Chinese.
|
|
27
|
Vaz-Drago R, Custódio N and Carmo-Fonseca
M: Deep intronic mutations and human disease. Hum Genet.
136:1093–1111. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Tang R, Prosser DO and Love DR: Evaluation
of bioinformatic programmes for the analysis of variants within
splice site consensus regions. Adv Bioinformatics.
2016:56140582016. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
BinEssa HA, Zou M, Al-Enezi AF, Alomrani
B, Al-Faham MSA, Al-Rijjal RA, Meyer BF and Shi Y: Functional
analysis of 22 splice-site mutations in the PHEX, the causative
gene in X-linked dominant hypophosphatemic rickets. Bone.
125:186–193. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Steffensen AY, Dandanell M, Jønson L,
Ejlertsen B, Gerdes AM, Nielsen FC and Hansen TV: Functional
characterization of BRCA1 gene variants by mini-gene splicing
assay. Eur J Hum Genet. 22:1362–1368. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Wang Y, Sun Y, Liu M, Zhang X and Jiang T:
Functional characterization of argininosuccinate lyase gene
variants by mini-gene splicing assay. Front Genet. 10:4362019.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Zaum AK, Stüve B, Gehrig A, Kölbel H,
Schara U, Kress W and Rost S: Deep intronic variants introduce DMD
pseudoexon in patient with muscular dystrophy. Neuromuscul Disord.
27:631–634. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Lou DI, Hussmann JA, Mcbee RM, Acevedo A,
Andino R, Press WH and Sawyer SL: High-throughput DNA sequencing
errors are reduced by orders of magnitude using circle sequencing.
Proc Natl Acad Sci USA. 110:19872–19877. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Lee H, Huang AY, Wang LK, Yoon AJ,
Renteria G, Eskin A, Signer RH, Dorrani N, Nieves-Rodriguez S, Wan
J, et al: Diagnostic utility of transcriptome sequencing for rare
Mendelian diseases. Genet Med. 22:490–499. 2020. View Article : Google Scholar :
|
|
35
|
Alieva M, van Rheenen J and Broekman MLD:
Potential impact of invasive surgical procedures on primary tumor
growth and metastasis. Clin Exp Metastas. 35:319–331. 2018.
View Article : Google Scholar
|
|
36
|
Barker SE, Grosse SM, Siapati EK, Kritz A,
Kinnon C, Thrasher AJ and Hart SL: Immunotherapy for neuroblastoma
using syngeneic fibroblasts transfected with IL-2 and IL-12. Brit J
Cancer. 97:210–217. 2007. View Article : Google Scholar : PubMed/NCBI
|