Introduction
Renal interstitial fibrosis (RIF) is a pathological
process that is common to the majority of chronic kidney diseases,
resulting in functional deterioration which is largely independent
of the initial renal injury (1).
RIF is characterized by abnormal interstitial extracellular matrix
(ECM) deposition (2). While a
number of detailed studies on the mechanistic basis for RIF have
been published in recent years, the etiology of this condition has
yet to be fully clarified.
MicroRNAs (miRNAs/miRs) are short non-coding RNAs,
which can modulate gene expression by altering target mRNA
stability and translation to control key physiological and
pathological processes (3,4).
Indeed, miRNA dysregulation has been linked to an array of
different disease processes (5).
Within the kidneys, specific miRNAs regulate development,
homeostasis and normal physiology, while also regulating the onset
of a range of renal diseases (6). As such, further studies elucidating
the miRNAs that regulate the incidence of RIF have the potential to
highlight novel approaches with which to prevent or treat this
debilitating condition.
In the present study, a miRNA array-based approach
was utilized to identify miRNAs that were differentially expressed
between fibrotic and normal kidney tissues using a rat model of
mercury chloride (HgCl2)-induced RIF (7,8).
This analysis identified miR-101 (also termed miR-101a,
miR-101a-3p) as the miRNA that was the most significantly
downregulated in fibrotic renal tissue. In previous studies,
miR-101 has also been shown to be involved in fibrotic processes,
such as liver fibrosis (9,10), pulmonary fibrosis (11), cardiac fibrosis (12,13), bladder fibrosis (14) and cystic fibrosis (15). The role of this miRNA in the
context of RIF, however, remains to be clarified, as do the
molecular mechanisms underlying such a role. As such, the present
study further evaluated the ability of miR-101 to regulate RIF.
One of the key steps in the development of RIF is
renal tubular epithelial-to-mesenchymal transition (EMT), wherein
tubular epithelial cells adopt a mesenchymal-like phenotype and
lose their epithelial-like traits (16). The disruption of tubular EMT is
thus a viable approach for the treatment of RIF (17,18). Transforming growth factor-β1
(TGF-β1) is an essential regulator of this renal tubular EMT
process (19,20), signaling through its cognate
cell-surface type I and II receptors (TβR-I and TβR-II,
respectively) to induce appropriate downstream signaling pathway
activation. There is prior evidence to indicate that TβR-I is a
direct miR-101 target (21), and
as such, it was hypothesized that miR-101 may be able to regulate
renal tubular EMT by targeting TβR-I and thereby suppressing TGF-β1
signaling.
The present study thus examined the ability of
miR-101 to reverse TGF-β1-induced tubular EMT in human proximal
tubular epithelial (HK-2) cells. Through these analyses, it was
determined that TGF-β1 treatment led to the downregulation of
miR-101. In addition, miR-101 overexpression suppressed
TGF-β1-induced EMT, whereas miR-101 knockdown exerted the opposite
effect. At a mechanistic level, miR-101 suppressed TGF-β1 signaling
by inhibiting TβR-I expression, and the blocking of TβR-I signaling
ablated the effects of miR-101 inhibition on TGF-β1-induced EMT.
Overall, the results thus demonstrated that miR-101 inhibited
tubular EMT, at least in part by suppressing TβR-I expression.
Materials and methods
Rat model of HgCl2-induced
RIF
In total, 20 male Sprague-Dawley rats (4-5 weeks
old, weighing 120±10 g) were obtained from the Shanghai Laboratory
Animal Center, Chinese Academy of Sciences. All rats were housed in
a specific pathogen-free environment that was maintained at 22-24°C
with a relative humidity of 50-60% and a 12-h light/dark cycle. All
rats were provided with ad libitum access to food and water.
These animals were then randomly divided into the control (n=8) and
model (n=12) groups, with the animals in the latter group being
orally administered with HgCl2 (8 mg/kg; Shanghai
Tongren Pharmaceutical Co., Ltd.) once daily for nine weeks. These
experimental protocols were conducted in accordance with
internationally accepted laboratory principles and all animals
received humane care as well as free access to food and water. The
present study was approved by The Animal Research Ethics Committee
of Shanghai University of Traditional Chinese Medicine, Shanghai,
China. The humane endpoint for this study was a loss of 15% of the
starting body weight. The animals were anesthetized using 1%
pentobarbital sodium (50 mg/kg) by intraperitoneal (i.p.)
injection. The samples of blood (0.8 ml) were collected from the
vena cava. After collecting the blood and kidney, at the end of the
experimental procedure, the abdominal vasculature, including the
vena cava was cut to cause exsanguination under deep anesthesia.
Death was further confirmed by checking for the onset of rigor
mortis (22). The body weight
was measured at the time of sacrifice, and body weight loss was not
observed in any of the rats, with the body weight ranging from 331
to 492 g.
Cells, cell culture and treatment
HK-2 cells were obtained from the Institute of Basic
Medical Sciences, Chinese Academy of Medical Sciences, and were
grown in DMEM (Gibco; Thermo Fisher Scientific, Inc.) containing
10% fetal bovine serum (FBS, Gibco; Thermo Fisher Scientific, Inc.)
for 18 h. Cells were then transferred to serum-free medium and were
treated for 48 h with 5 ng/ml TGF-β1 to achieve EMT induction. In
appropriate experiments, cells were transfected with 20 nM of
miR-101 mimic (Guangzhou RiboBio Co., Ltd.; the mature sequence of
hsa-miR-101 mimic was UAC AGU ACU GUG AUA ACU GAA) or 50 nM of
miR-101 inhibitor (Guangzhou RiboBio Co., Ltd.) or appropriate
controls using a ribioFECT CP Transfection kit (Guangzhou RiboBio
Co., Ltd.) along with 5 ng/ml TGF-β1 for 48 h. In order to assess
how TβR-I affects these results, in appropriate experiments, cells
were treated with 10 µM of SB-431542 (Tocris Bioscience) to
specifically inhibit TβR-I for 12 h, and the cells were then
transfected with miR-101 inhibitor or appropriate controls using a
ribioFECT CP Transfection kit (Guangzhou RiboBio Co., Ltd.) along
with 5 ng/ml TGF-β1 or SB-431542 for a further 48 h.
Hydroxyproline measurements
Renal hydroxyproline levels were measured based upon
HCl hydrolysis as per the method described in the study by Jamall
et al (23). Briefly, 100
mg of renal tissue was homogenized in 2.5 ml of ice-cold
ddH2O, after which a BCA kit (Thermo Fisher Scientific,
Inc.) was used to quantify the protein levels in these samples.
Subsequently, 6 M HCl were used to hydrolyze 2 ml of these
homogenates for 18 h at 105°C, and the resultant hydrolysates were
filtered using 3-mm filter paper prior to drying at 40°C. Samples
were then incubated with Ehrlich's solution [25% (w/v)
p-dimethylaminobenzaldehyde and 27.3% (v/v) perchloric acid in
isopropanol] at 50°C for 1.5 h, followed by assessment at 558 nm
using a Tecan Infinite M200 Pro plate reader (Tecan Life Sciences),
with protein concentrations being used to normalize the resultant
values.
Histological analysis
The kidney samples were fixed using 10% formalin,
paraffin-embedded, and cut into 5-µM-thick sections. The
sections were then subjected to Masson's trichrome staining using a
modified Masson's Trichrome stain kit (Solarbio Science &
Technology Co., Ltd.) according to the manufacturer's instructions.
Briefly, the sections were stained in Harris hematoxylin for 5 min,
in Ponceau acid fuchsin staining solution for 10 min, and in
Aniline Blue solution for 5 min at room temperature. Hematoxylin
and eosin (H&E) staining was performed using a H&E staining
kit (Yeasen Biotech Co., Ltd.) according to the manufacturer's
instructions. Briefly, the sections were stained in hematoxylin for
5 min and in eosin solution for 1 min at room temperature. Images
were obtained using an Olympus IX73 microscope (Olympus
Corporation). ImageJ software for Windows V 1.52v (NIH) was used to
quantify the Masson's trichrome positive staining area.
miRNA microarray
Renal miRNA profiles in these animals (two samples
in each group) were evaluated using an Agilent Rat microRNA
Microarray 16.0 (Agilent Technologies. Inc.). Total renal tissue
RNA was extracted using a mirVana™ miRNA Isolation kit (Ambion;
Thermo Fisher Scientific, Inc.), after which an Agilent Bioanalyzer
2100 was used to assess the RNA quality based upon the RNA
integrity number (RIN) statistic. The miRNAs in these samples were
then labeled and hybridized with a miRNA Complete Labeling and Hyb
kit (Agilent Technologies, Inc.) based on the provided directions,
and the Agilent Microarray Scanner and Feature Extraction software
(v10.7) was used to analyze these slides under default settings.
The Quantile algorithm was used to normalize the resultant raw data
with the Gene Spring Software v11.0 (Agilent Technologies, Inc.).
Shanghai Biotechnology Corporation conducted these microarray
analyses.
miRNA microarray analysis
Linear models and empirical Bayes methods were used
to detect differentially expressed miRNAs (24), with the thresholds for
differential expression being the following: P<0.05, false
discovery rate (FDR) <0.05 and fold change (FC) >1.5.
TargetScan (http://www.targetscan.org) was used to identify
putative miRNA target genes, with the resultant miRNA-gene network
being constructed based upon the associations between these miRNAs
and genes in the Sanger microRNA database (http://www.mirbase.org) and on the strength of their
interactions. This network was constructed as an adjacency matrix
A=[ai,j], where ai,j represents the
relational weights of gene 'I' and microRNA 'j'. In the final
network, squares and circles were used to represent miRNAs and
mRNAs, respectively, with single edges being used to represent
associations between miRNAs and target genes. Degree values were
used to represent centrality within this network, indicating the
contribution of one miRNA to the surrounding genes such that key
miRNAs have larger degree values (25).
Reverse transcription-quantitative PCR
(RT-qPCR)
A Qiagen miRNeasy Mini kit (Qiagen, Inc.) was used
to extract mRNA and miRNAs from the samples. Relative gene
expression was analyzed using the 2−ΔΔCq method
(26), with U6 and β-actin being
used to normalize miRNA and mRNA expression, respectively. A
SYBR-Green Real-Time PCR kit (Takara Bio, Inc.) was used for qPCR
with the following primers: Human E-cadherin forward, 5′-AAG ACA
AAG AAG GCA AGG TT-3′ and reverse, 5′-AAG AGA GTG TAT GTG GCA
AT-3′; human α-smooth muscle actin (α-SMA) forward, 5′-GGA CAT CAA
GGA GAA ACT GT-3′ and reverse, 5′-CCA TCA GGC AAC TCG TAA CT-3′;
human TβR-I forward, 5′-TGT GAA GCC TTG AGA GTA AT-3′ and reverse,
5′-TGT TGA CTG AGT TGC GAT AA-3′; and β-actin forward, 5′-CAC GAT
GGA GGG GCC GGA CTC ATC -3′ and reverse, 5′-TAA AGA CCT CTA TGC CAA
CAC AGT -3′. Primers for U6 and miRNA were from GeneCopoeia, Inc.
The cycling conditions were as follows: 95°C for 30 sec, followed
by 40 cycles of 95°C for 5 sec, and 60°C for 10 sec.
Western blot analysis
RIPA buffer supplemented with a complete mini
protease inhibitor cocktail and a PhosSTOP phosphatase inhibitor
cocktail (Roche Diagnostics) was used to lyse the HK-2 cell and
renal tissue samples, which were then spun for 15 min at 13,000 × g
at 4°C. A BCA assay (Thermo Fisher Scientific, Inc.) was then used
to assess the protein quantities in the collected supernatants, and
equal protein amounts (30 µg) were separated via 10%
SDS-PAGE prior to transfer onto nitrocellulose membranes. Following
a 1-h blocking step using 5% non-fat milk for 1 h at room
temperature, these blots were probed overnight at 4°C with primary
anti-α-SMA (1:1,000, ab7817), anti-E-cadherin (1:1,000 ab40772),
anti-Smad3 (1:1,000 ab40854), anti-p-Smad3 (1:500 ab52903),
anti-TβR-I (1:1,000 ab31013) and anti-GAPDH antibodies (1:5,000
ab8245) (Abcam). The blots were then washed, incubated with
secondary HRP-conjugated goat anti-mouse antibody (1:5,000,
ab97023, Abcam) or goat anti-rabbit antibody (1:5,000, ab205718,
Abcam) for 1 h at room temperature, and protein bands were then
visualized using an ECL reagent (Thermo Fisher Scientific, Inc.),
with a Tanon 5200 detection system (Tanon Science & Technology
Co., Ltd.) being used for imaging. Protein band densitometry was
quantified using ImageJ software for Windows V 1.52v (NIH) and
normalized to GAPDH.
Immunofluorescence staining
E-cadherin and α-SMA expression was assessed via
immunofluorescence in HK-2 cells by plating these cells in 96-well
plates, washing them with PBS twice, and then fixing them using 4%
paraformaldehyde. Subsequently, 5% BSA in PBS was used to block the
cells for 30 min at room temperature, after which they were
incubated with primary anti-α-SMA (1:200, ab7817; Abcam) and
anti-E-cadherin (1:50, ab40772; Abcam) antibodies at room
temperature for 1 h. Cy3-conjugated goat anti-rabbit secondary
antibody (1:200, A10520; Thermo Fisher Scientific, Inc.) and
FITC-conjugated goat anti-mouse secondary antibody (1:200, F2761;
Thermo Fisher Scientific, Inc.) were then incubated with these
cells for 1 h at 37°C, after which Hoechst 33258 (Thermo Fisher
Scientific, Inc.) was used for nuclear staining at room temperature
for 5 min, and a Thermo Fisher Scientific ArrayScan HCS reader was
used for image acquisition, with the Thermo HCS Studio™ 2.0 Cell
Analysis Program was used for subsequent data analysis.
Statistical analysis
Data are presented as the mean ± SEM. GraphPad Prism
7 was used for all statistical analysis. Data were compared using
Student's t-tests and one-way ANOVA with Tukey's post hoc analysis,
as appropriate. P<0.05 was the significance threshold, and all
experiments were repeated three or more times.
Results
HgCl2 treatment induces renal
inflammation and interstitial fibrosis
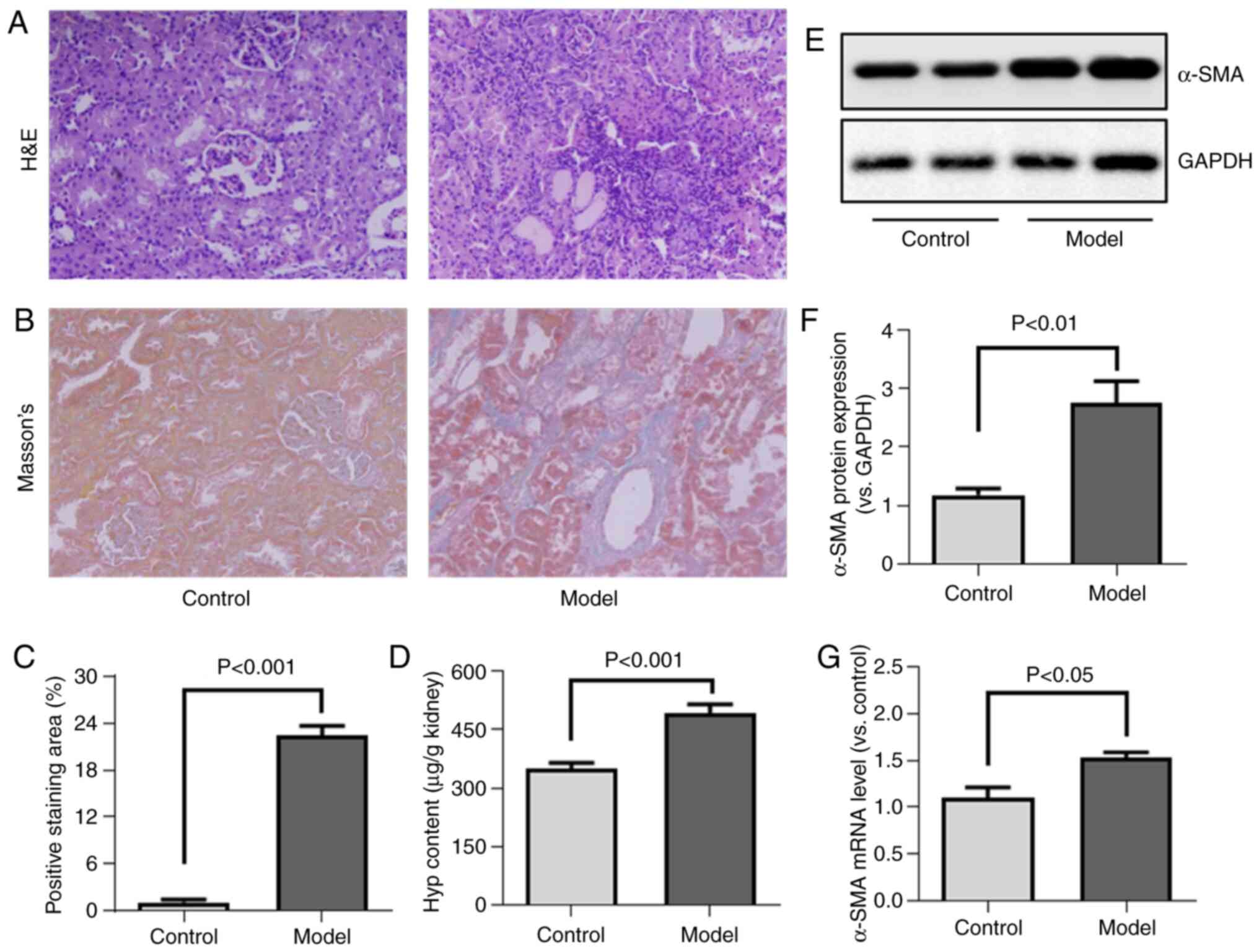
The present study first analyzed the H&E-stained
kidney sections from the rats, revealing that HgCl2
treatment was associated with renal tubular deformation and
mononuclear cell infiltration. This was in sharp contrast to the
normal glomerul'ar and tubular architecture observed in the control
animals (Fig. 1A).
Collagen deposition is a key hallmark of tissue
fibrosis, and can be evaluated via Masson's trichrome staining and
by measuring hydroxyproline content. Masson's staining indicated
that HgCl2 treatment resulted in increased collagen
deposition (Fig. 1B and C). The
renal hydroxyproline content was similarly elevated in the
HgCl2-treated rats compared with the controls (Fig. 1D).
α-SMA is a marker of ECM-secreting myofibroblasts,
and it was found that HgCl2 treatment significantly
enhanced α-SMA expression at the protein and mRNA level (Fig. 1E-G), thus confirming that this
treatment was linked to an enhanced renal myofibroblast
activation.
Identification of key differentially
expressed miRNAs associated with HgCl2-induced RIF
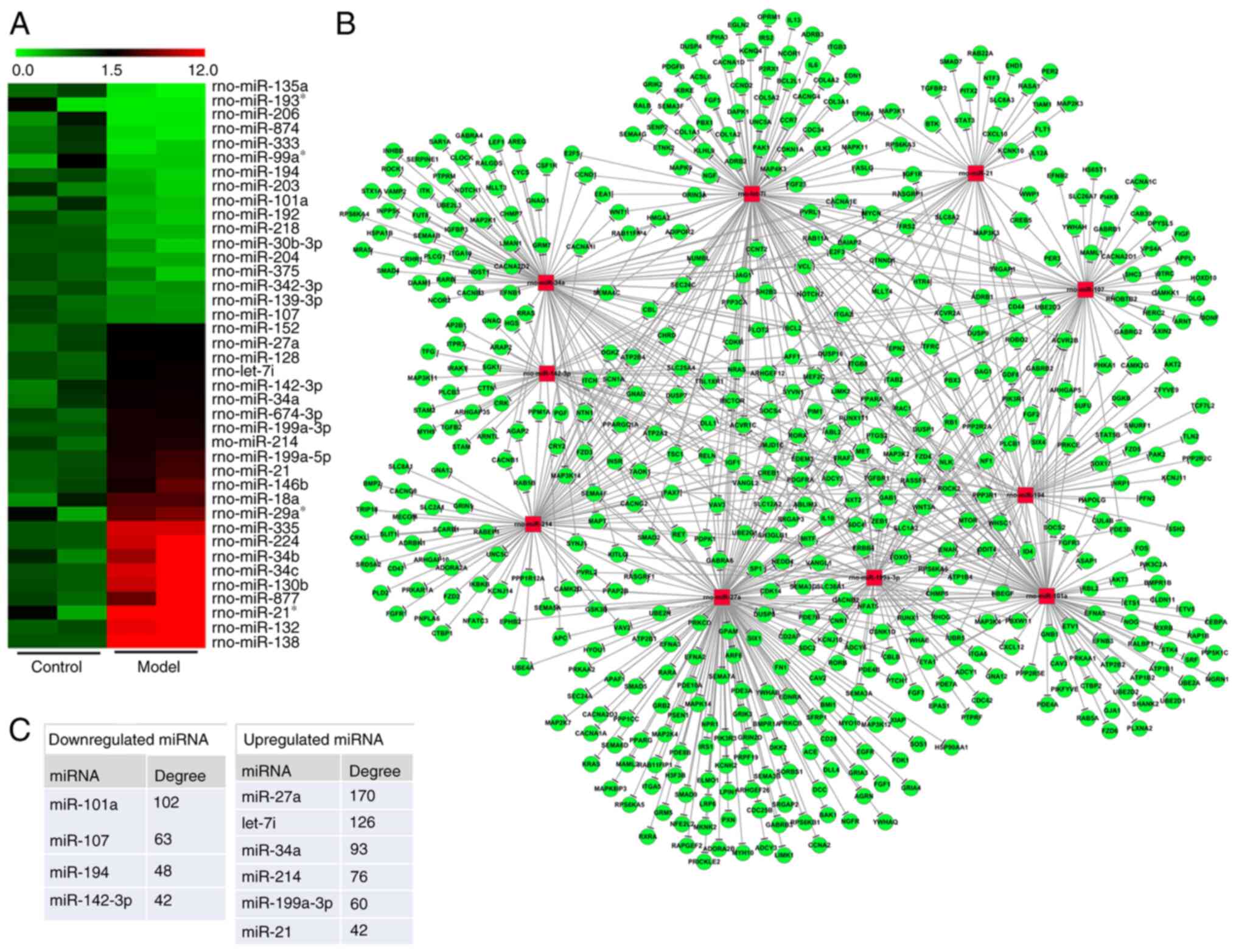
Using an array-based approach, 40 miRNAs that were
differentially expressed between the control and RIF model animals
were identified, of which 17 and 23 were downregulated and
upregulated, respectively (Fig.
2A). TargetScan was used to predict target genes for these
miRNAs and a miRNA-gene network based upon predicted interactions
among these miRNAs and genes was then constructed using the Sanger
miRNA database (Fig. 2B). The
degree metric was used to determine centrality within this network,
reflecting the degree to which a given miRNA contributes to the
regulation of the surrounding genes, with key miRNAs having a
larger degree value (25). The
top 10 most central miRNAs within this network included four
downregulated miRNAs (miR-101a, miR-107, miR-194 and miR-142-3p,
with respective degree values of 102, 63, 48 and 43) and six
upregulated miRNAs (miR-27a, let-7i, miR-34a, miR-214, miR-199a-3p
and miR-21, with respective degree values of 170, 126, 93, 76, 60
and 42) (Fig. 2C).
miR-101 is downregulated in fibrotic
kidney tissue and TGF-β1-treated HK-2 cells
Given that miR-101 was among the most significantly
downregulated miRNAs in the miRNA-mRNA network (Fig. 2C), the present study then
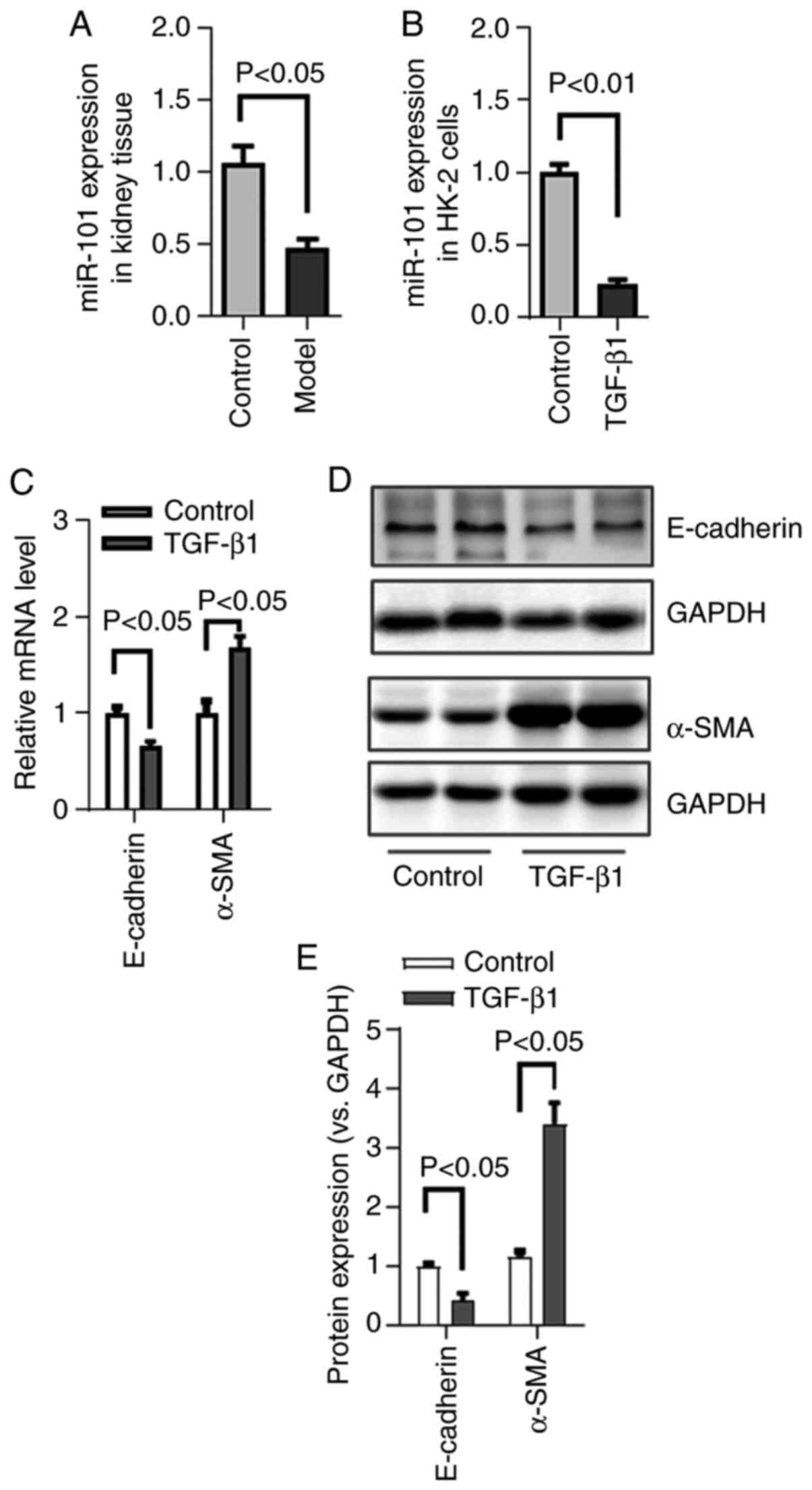
explored the role of this miRNA in the context of RIF. First it was
confirmed that miR-101 downregulation was evident in the
HgCl2-treated fibrotic kidney tissue by RT-qPCR, in line
with the microarray findings (Fig.
3A).
Renal tubular EMT is a key driver of the development
of RIF (27). The present study
therefore explored this process in vitro by treating HK-2
cells with TGF-β1. Consistent with the successful EMT induction,
TGF-β1 treatment resulted in both the loss of epithelial E-cadherin
expression and the upregulation of mesenchymal α-SMA expression in
HK-2 cells, as confirmed by western blot analysis (Fig. 3C-E). Additionally, miR-101
expression was assessed in these cells by RT-qPCR, and it was
confirmed that this miRNA was downregulated in the context of
TGF-β1-induced EMT (Fig.
3B).
miR-101 inhibits TGF-β1-induced EMT in
HK-2 cells
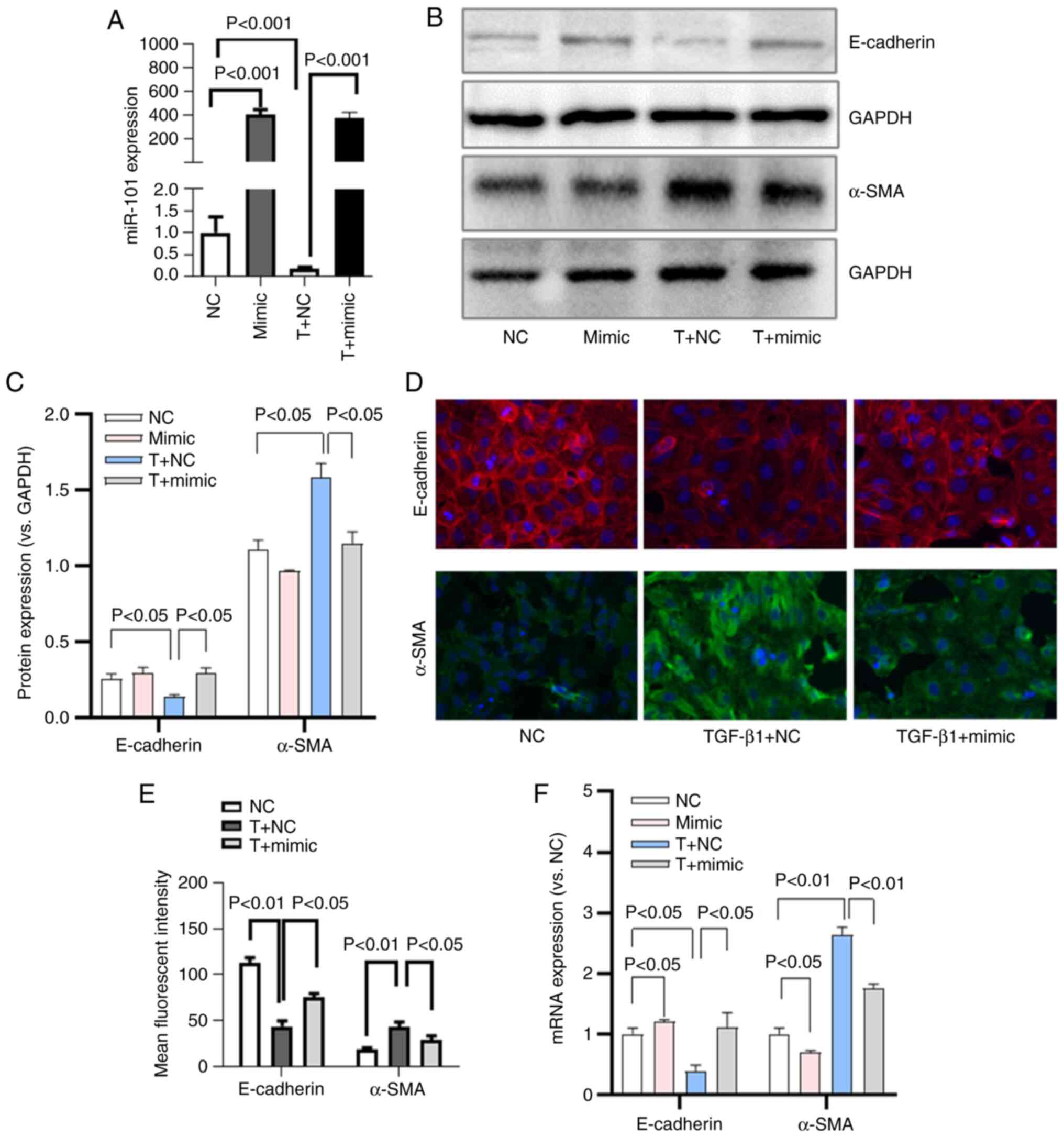
HK-2 cells were then transfected with a miR-101
mimic or a corresponding control construct, after which these cells
were treated with TGF-β1. As was expected, miR-101 mimic
transfection was associated with a significant increase in miR-101
expression, whereas TGF-β1 treatment decreased miR-101 expression
(Fig. 4A).
When the effects of miR-101 on TGF-β1-induced EMT
were evaluated in these HK-2 cells, it was found that miR-101 mimic
transfection resulted in an increased E-cadherin expression at the
protein (Fig. 4B-E) and mRNA
(Fig. 4F) level, along with a
decreased α-SMA protein (Fig.
4B-E) and mRNA (Fig. 4F)
expression, consistent with the suppression of EMT in these
cells.
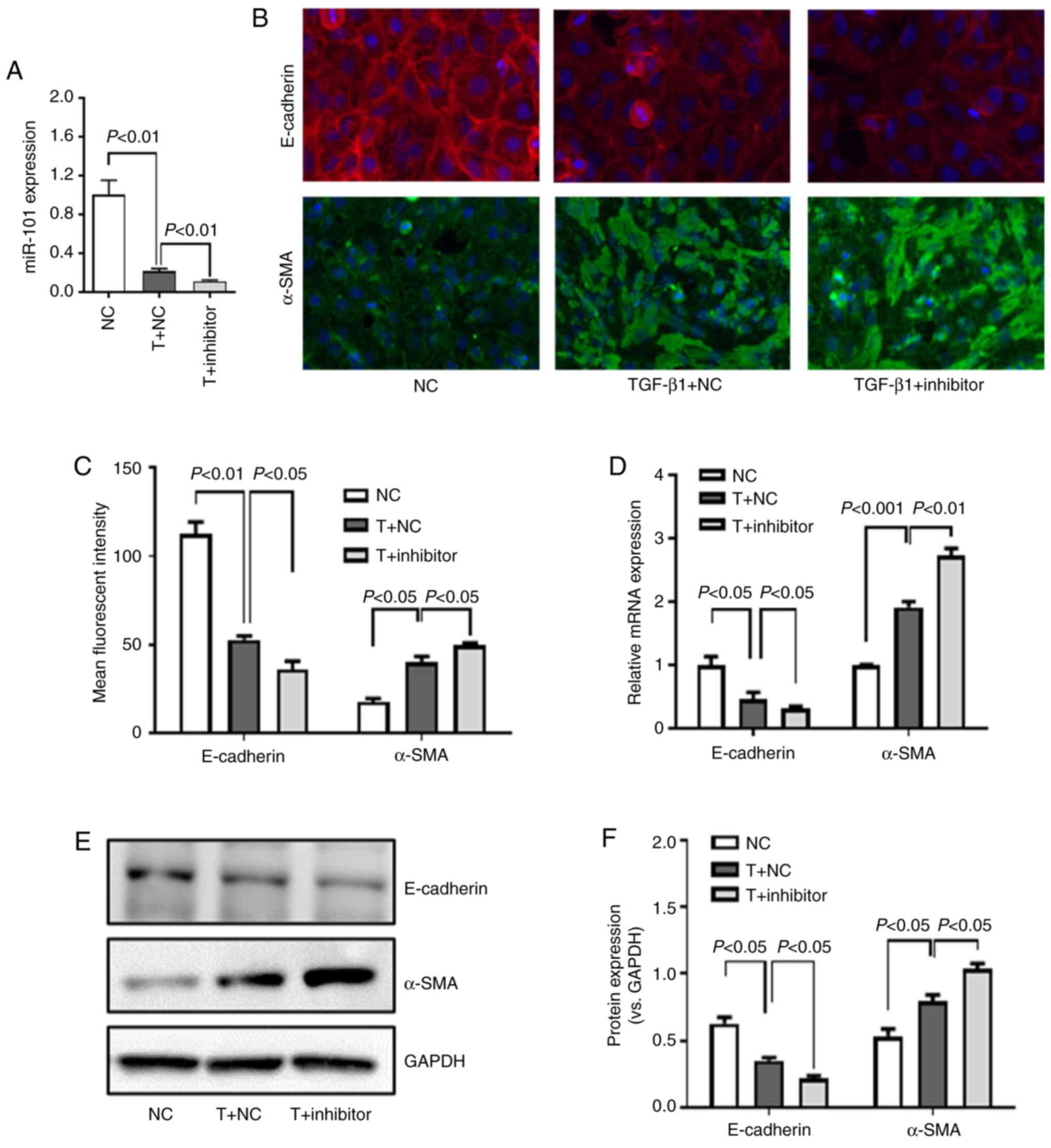
miR-101 knockdown promotes TGF-β1-induced
EMT in HK-2 cells
The present study then assessed the effects of
miR-101 downregulation on TGF-β1-induced EMT in HK-2 cells by
transfecting the cells with a miR-101 inhibitor prior to TGF-β1
treatment, resulting in a significant knockdown of miR-101
expression (Fig. 5A). This
inhibition of miR-101 was associated with a significant decrease in
E-cadherin protein (Fig. 5B, C, E
and F) and mRNA (Fig. 5D)
expression, while α-SMA protein (Fig. 5, C, E and F) and mRNA (Fig. 5D) expression increased
significantly in these cells. These results thus revealed that
miR-101 downregulation enhanced TGF-β1-induced EMT in these HK-2
cells.
Effect of miR-101 on TβR-I and Smad3
expression in HK-2 cells
Previous studies have identified TβR-I as a miR-101
target gene (21,28). The present study thus assessed
the effects of miR-101 on TβR-I expression in the TGF-β1-treated
HK-2 cells. This approach revealed that TGF-β1 treatment enhanced
TβR-I expression, whereas miR-101 mimic transfection inhibited its
expression (Fig. 6A and B). By
contrast, transfection with miR-101 inhibitor was associated with
an increased TβR-I expression in the TGF-β1-treated HK-2 cells
(Fig. 6D and E).
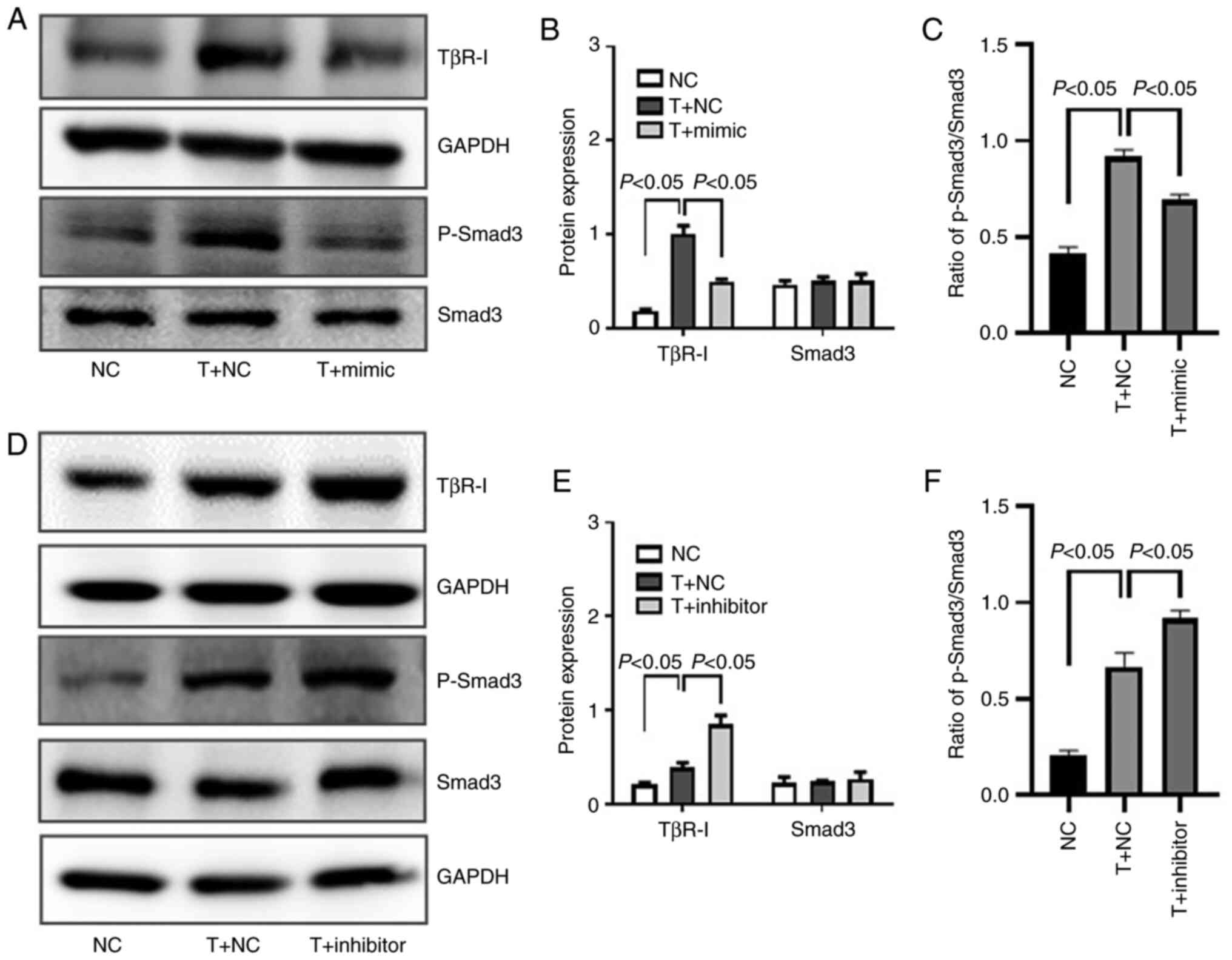 | Figure 6Effect of miR-101 on TβR-I and Smad3
in HK-2 cells. (A) TβR-I, Smad3 and p-Smad3 levels were assessed by
western blot analysis in HK-2 cells following miR-101 mimic
transfection. (B) Densitometric quantification of TβR-I and Smad3,
with GAPDH being used for normalization. (C) Ratio of p-Smad3 vs.
Smad3. (D) TβR-I, Smad3 and p-Smad3 levels were assessed by western
blot analysis in HK-2 cells following miR-101 inhibitor
transfection. (E) Densitometric quantification of TβR-I and Smad3,
with GAPDH being used for normalization. (F) Ratio of p-Smad3 vs.
Smad3. NC, negative control; T, TGF-β1; mimic, miR-101 mimic;
inhibitor, miR-101 inhibitor; α-SMA, α-smooth muscle actin; TGF-β1,
transforming growth factor-β1. |
The TGF-β1/Smad pathway serves to facilitate
canonical TGF-β1 signaling within cells, with Smad3 phosphorylation
being a key step in this pathway (29). The present study confirmed, by
western blot analysis, that miR-101 mimic transfection was
associated with a decreased Smad3 phosphorylation (Fig. 6A and C), whereas the inhibition
of miR-101 exerted the opposite effect in TGF-β1-treated HK-2 cells
(Fig. 6D and F).
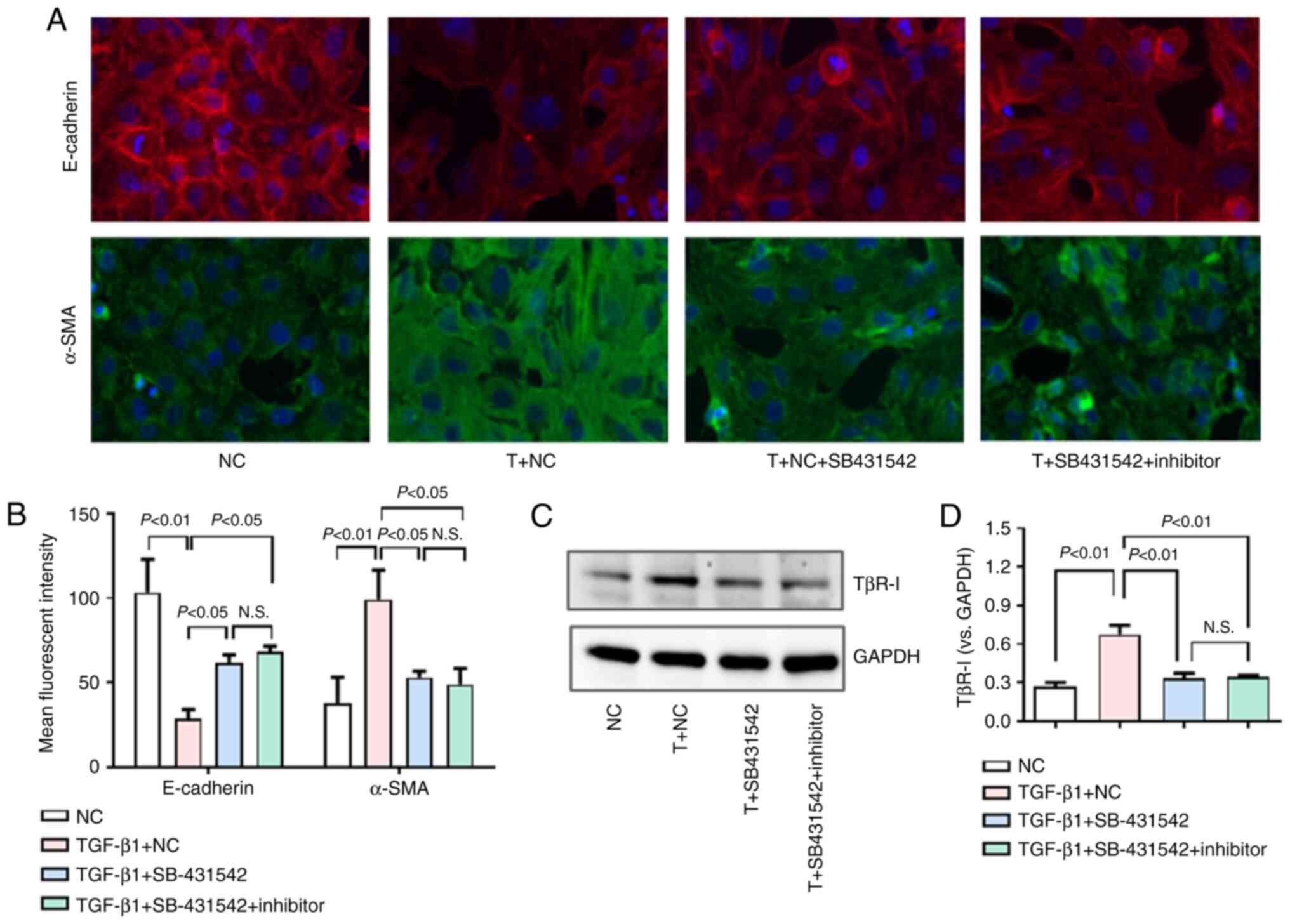
TβR-I inhibition impairs TGF-β1-induced
EMT and the effect of miR-101 inhibition on HK-2 cells
In an effort to more fully assess whether miR-101
targets TβR-I to suppress the EMT process in HK-2 cells, the
present study then assessed whether the effect of miR-101 inhibitor
transfection on EMT was disrupted when the cells were treated with
the potent TβR-I inhibitor, SB-431542 (30). As was expected, SB-431542
markedly impaired TGF-β1-induced EMT in the HK-2 cells, enhancing
E-cadherin expression and suppressing α-SMA expression (Fig. 7A and B). When the cells were
treated with SB-431542, this ablated the effects of miR-101
inhibitor transfection on EMT-related phenotypes (Fig. 7A and B), indicating that miR-101
targets TβR-I to inhibit TGF-β1-induced EMT. When TβR-I protein
expression was assessed, it was confirmed that SB-431542 treatment
markedly decreased TβR-I protein expression, whereas miR-101
inhibitor treatment did not further affect the TβR-I protein levels
in these SB-431542-treated HK-2 cells (Fig. 7C and D).
Discussion
A number of studies to date have highlighted the
roles played by miRNAs in the context of both pathological and
physiological processes (5).
Specific miRNAs have been found to regulate renal development,
homeostasis and the pathology of RIF as well as other diseases
(6,31). While the general etiology of RIF
has been thoroughly studied, the specific regulatory roles of
individual miRNAs in this disease context remain to be fully
elucidated. By further studying the ability of these miRNAs to
influence RIF progression, it may be possible to identify novel
approaches to preventing or treating this condition.
Herein, a previously described
HgCl2-induced rat model of RIF was employed (8,32). By analyzing renal tissues from
these animals, 17 and 23 miRNAs were identified that were
downregulated and upregulated, respectively, in fibrotic kidney
tissue samples relative to the control kidney samples. Through
bioinformatics analyses, the 10 most critical of these miRNAs were
then identified, including four that were downregulated (miR-101,
miR-107, miR-194 and miR-142-3p) and six that were upregulated
(miR-27a, let-7i, miR-34a, miR-214, miR-199a-3p and miR-21) in
fibrotic renal tissue. The majority of these miRNAs have previously
been linked to renal diseases, such as RIF. For example, Hou et
al (33) found miR-27a to
suppress peroxisome proliferator-activated receptor-γ signaling and
to thereby promote RIF, whereas miR-34a has been shown to regulate
Klotho expression in tubular epithelial cells, thereby controlling
RIF (34), while also inducing
the apoptotic death of these cells (35). There is also evidence to indicate
that miR-214 is upregulated in the context of renal injury, and the
knockdown of this miRNA is sufficient to attenuate unilateral
ureteral obstruction (UUO)-induced RIF (36). miR-21 is among the most
well-characterized miRNAs associated with fibrosis in a range of
tissue types (37-39). In healthy renal tissue, minimal
miR-21 expression is observed, whereas it is significantly
upregulated in the context of RIF, wherein it can target PTEN and
peroxisome proliferator-activated receptor-α to promote fibrotic
progression (38). These
findings thus confirm that HgCl2 induces a model of RIF similar to
that induced by a UUO-based approach, while also confirming the
reliability of our miRNA array findings.
Using bioinformatics analyses, the present study
identified miR-101 as the most downregulated miRNA in the rat model
of HgCl2-induced RIF. This miRNA has previously been
reported to play roles in other fibrotic processes, including liver
fibrosis (9,10), pulmonary fibrosis (11), cardiac fibrosis (12,13), bladder fibrosis (14) and cystic fibrosis (15); however, its importance in the
context of RIF has not been well-characterized. As such, the
present study explored its role in this pathological setting. Renal
tubular EMT is a key step in RIF progression (40), and is characterized by tubular
cells undergoing a shift from an epithelial-like to a
mesenchymal-like phenotype, whereupon these cells are able to
produce high levels of ECM components and to thereby drive RIF
pathogenesis (8). TGF-β1 is the
most well-studied inducer of EMT (41), which is also downregulated in
HgCl2-induced RIF (8,32), and as such, this was utilized in
the present study to induce this process in HK-2 cells. In line
with the findings in vivo, a reduced miR-101 expression was
observed in the cells following TGF-β1 treatment. When miR-101 was
overexpressed in these same cells, it was found that this reversed
TGF-β1-induced EMT, whereas miR-101 inhibition exerted the opposite
effect. Taken together, these findings highlight miR-101 as an
inhibitor of tubular EMT.
The TGF-β1/Smad pathway is an essential mediator of
EMT progression that is initiated upon the binding of TGF-β1 to the
cell surface TβR-II molecule, in turn resulting in TβR-I
activation, Smad2/3 phosphorylation and nuclear translocation and
altered gene expression. Previous research has demonstrated that
miR-101 can suppress fibrosis owing to its ability to target TβR-I
and to thereby suppress TGF-β1 signaling (21). In line with this finding, the
present study determined that miR-101 was able to inhibit EMT
progression via downregulating TβR-I, as miR-101 overexpression
disrupted TGF-β1-induced TβR-I mRNA and protein expression. The
TGF-β1/Smad pathway is an essential mediator of canonical TGF-β1
signaling (29), with Smad3
phosphorylation being a key component of this process.
Consistently, it was determined that miR-101 was able to inhibit
TGF-β1-induced Smad3 phosphorylation in HK-2 cells, suggesting that
this miRNA can suppress TGF-β1/Smad3 signaling. Taken together,
these findings demonstrate that miR-101 can control TβR-I
expression so as to suppress TGF-β1 signaling.
To confirm these results, the cells were
additionally treated with the potent TβR-I inhibitor, SB-431542. As
was expected, SB-431542 suppressed TGF-β1 induced EMT in HK-2
cells. When cells were treated with this TβR-I inhibitor, the
ability of miR-101 inhibition to impact EMT phenotypes was ablated,
thus confirming that miR-101 ablates TGF-β1-induced EMT by directly
targeting TβR-I. Recently, Zhao et al (42) also found that miR-101 inhibited
acute kidney injury-chronic kidney disease transition by inhibiting
the EMT process. The results of the present study revealed that
miR-101 expression was downregulated in a rat renal fibrosis model,
confirming that miR-101 may be important in different rodent renal
diseases model. In an in vitro study, Zhao et al
(42) found that miR-101
overexpression using miR-101 mimic inhibited EMT. Apart from the
gain-of function experiment, in the present study, it was further
confirmed that the downregulation of miR-101 using miR-101
inhibitor promoted the process of EMT. Taken together, the results
further verified the key role of miR-101 in kidney diseases, and
suggested that it may be a potential target for the treatment of
renal fibrosis.
In conclusion, the findings of the present study
highlight miR-101 as a key miRNA associated with
HgCl2-induced RIF. It was found that this miRNA can
suppress renal tubular EMT by targeting TβR-I and thereby
inhibiting TGF-β1 signaling, thus indicating that miR-101 may be a
viable target for treating RIF, although further research is
required needed to confirm this hypothesis.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
PL, CL and QW conceived the study and established
the initial design of the study. QW, YT and HX performed the
experiments and analyzed the data. QW prepared the manuscript. All
authors read and approved the final manuscript. QW and PL confirm
the authenticity of all the raw data.
Ethics approval and consent to
participate
The present study was approved by the Animal
Research Ethics Committee of Shanghai University of Traditional
Chinese Medicine, Shanghai, China.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Acknowledgments
Not applicable.
Funding
The present study was supported by the National Natural Science
Foundation of China (grant nos. 81573810 and 30901943), the China
Postdoctoral Science Foundation (grant no. 2015T80445) and the
National Science and Technology Major Project 'Key New Drug
Creation and Manufacturing Program' of China (grant no.
2019ZX09201001).
Abbreviations:
|
α-SMA
|
α-smooth muscle actin
|
|
ANOVA
|
one-way analysis of variance
|
|
DMEM
|
Dulbecco's modified Eagle's medium
|
|
ECM
|
extracellular matrix
|
|
EMT
|
epithelial-to-mesenchymal
transition
|
|
FBS
|
fetal bovine serum
|
|
FDR
|
false discovery rate
|
|
HgCl2
|
mercury chloride
|
|
miRNAs/miRs
|
microRNAs
|
|
RIF
|
renal interstitial fibrosis
|
|
TβR-I
|
TGF-β1 type I receptor
|
|
TβR-II
|
TGF-β1 type II receptor
|
|
TGF-β1
|
transforming growth factor-β
|
References
|
1
|
Farris AB and Colvin RB: Renal
interstitial fibrosis: Mechanisms and evaluation. Curr Opin Nephrol
Hypertens. 21:289–300. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Strutz F and Zeisberg M: Renal fibroblasts
and myofibroblasts in chronic kidney disease. J Am Soc Nephrol.
17:2992–2998. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
He L and Hannon GJ: MicroRNAs: Small RNAs
with a big role in gene regulation. Nat Rev Genet. 5:522–531. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Fabian MR, Sonenberg N and Filipowicz W:
Regulation of mRNA translation and stability by microRNAs. Annu Rev
Biochem. 79:351–379. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Kloosterman WP and Plasterk RH: The
diverse functions of microRNAs in animal development and disease.
Dev Cell. 11:441–450. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Chung AC, Yu X and Lan HY: MicroRNA and
nephropathy: Emerging concepts. Int J Nephrol Renovasc Dis.
6:169–179. 2013.PubMed/NCBI
|
|
7
|
Yuan JL, Tao YY, Wang QL, Shen L and Liu
CH: Fuzheng Huayu Formula () prevents rat renal interstitial
fibrosis induced by HgCl2 via antioxidative stress and
down-regulation of nuclear factor-kappa B activity. Chin J Integr
Med. 23:598–604. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Wang QL, Tao YY, Yuan JL, Shen L and Liu
CH: Salvianolic acid B prevents epithelial-to-mesenchymal
transition through the TGF-beta1 signal transduction pathway in
vivo and in vitro. BMC Cell Biol. 11:312010. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Lei Y, Wang QL, Shen L, Tao YY and Liu CH:
MicroRNA-101 suppresses liver fibrosis by downregulating
PI3K/Akt/mTOR signaling pathway. Clin Res Hepatol Gastroenterol.
43:575–584. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Wang B, Yan Z, Li L, Wang Z and Liu H:
Effect of MiR-101 on rats with CCl4-induced liver fibrosis through
regulating Nrf2-ARE pathway. Panminerva Med. Jul 30–2019.Epub ahead
of print.
|
|
11
|
Huang C, Xiao X, Yang Y, Mishra A, Liang
Y, Zeng X, Yang X, Xu D, Blackburn MR, Henke CA and Liu L:
MicroRNA-101 attenuates pulmonary fibrosis by inhibiting fibroblast
proliferation and activation. J Biol Chem. 292:16420–16439. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Pan Z, Sun X, Shan H, Wang N, Wang J, Ren
J, Feng S, Xie L, Lu C, Yuan Y, et al: MicroRNA-101 inhibited
postinfarct cardiac fibrosis and improved left ventricular
compliance via the FBJ osteosarcoma oncogene/transforming growth
factor-β1 pathway. Circulation. 126:840–850. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Li X, Zhang S, Wa M, Liu Z and Hu S:
MicroRNA-101 protects against cardiac remodeling following
myocardial infarction via downregulation of runt-related
transcription factor 1. J Am Heart Assoc. 8:e0131122019. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wang N, Duan L, Ding J, Cao Q, Qian S,
Shen H and Qi J: MicroRNA-101 protects bladder of BOO from
hypoxia-induced fibrosis by attenuating TGF-β-smad2/3 signaling.
IUBMB Life. 71:235–243. 2019.
|
|
15
|
Viart V, Bergougnoux A, Bonini J, Varilh
J, Chiron R, Tabary O, Molinari N, Claustres M and Taulan-Cadars M:
Transcription factors and miRNAs that regulate fetal to adult CFTR
expression change are new targets for cystic fibrosis. Eur Respir
J. 45:116–128. 2015. View Article : Google Scholar
|
|
16
|
Liu Y: Cellular and molecular mechanisms
of renal fibrosis. Nat Rev Nephrol. 7:684–696. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Yang J and Liu Y: Blockage of tubular
epithelial to myofibroblast transition by hepatocyte growth factor
prevents renal interstitial fibrosis. J Am Soc Nephrol. 13:96–107.
2002. View Article : Google Scholar
|
|
18
|
Allison SJ: Fibrosis: Targeting EMT to
reverse renal fibrosis. Nat Rev Nephrol. 11:5652015. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Loeffler I and Wolf G: Transforming growth
factor-beta and the progression of renal disease. Nephrol Dial
Transplant. 29(Suppl 1): i37–i45. 2014. View Article : Google Scholar
|
|
20
|
Iwano M: EMT and TGF-beta in renal
fibrosis. Front Biosci (Schol Ed). 2:229–238. 2010. View Article : Google Scholar
|
|
21
|
Zhao X, Wang K, Liao Y, Zeng Q, Li Y, Hu
F, Liu Y, Meng K, Qian C, Zhang Q, et al: MicroRNA-101a inhibits
cardiac fibrosis induced by hypoxia via targeting TGFβRI on cardiac
fibroblasts. Cell Physiol Biochem. 35:213–226. 2015. View Article : Google Scholar
|
|
22
|
Close B, Banister K, Baumans V, Bernoth
EM, Bromage N, Bunyan J, Erhardt W, Flecknell P, Gregory N,
Hackbarth H, et al: Recommendations for euthanasia of experimental
animals: Part 1. DGXI of the European commission. Lab Anim.
30:293–316. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Jamall IS, Finelli VN and Que Hee SS: A
simple method to determine nanogram levels of 4-hydroxyproline in
biological tissues. Anal Biochem. 112:70–75. 1981. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Smyth GK: Linear models and empirical
bayes methods for assessing differential expression in microarray
experiments. Stat Appl Genet Mol Biol. 3:Article3. 2004. View Article : Google Scholar
|
|
25
|
Joung JG, Hwang KB, Nam JW, Kim SJ and
Zhang BT: Discovery of microRNA-mRNA modules via population-based
probabilistic learning. Bioinformatics. 23:1141–1147. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
27
|
Iwano M, Plieth D, Danoff TM, Xue C, Okada
H and Neilson EG: Evidence that fibroblasts derive from epithelium
during tissue fibrosis. J Clin Invest. 110:341–350. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Tu X, Zhang H and Zhang J, Zhao S, Zheng
X, Zhang Z, Zhu J, Chen J, Dong L, Zang Y and Zhang J: MicroRNA-101
suppresses liver fibrosis by targeting the TGFβ signalling pathway.
J Pathol. 234:46–59. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Meng XM, Chung AC and Lan HY: Role of the
TGF-β/BMP-7/Smad pathways in renal diseases. Clin Sci (Lond).
124:243–254. 2013. View Article : Google Scholar
|
|
30
|
Chaudhary NI, Roth GJ, Hilberg F,
Müller-Quernheim J, Prasse A, Zissel G, Schnapp A and Park JE:
Inhibition of PDGF, VEGF and FGF signalling attenuates fibrosis.
Eur Respir J. 29:976–985. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Rudnicki M, Perco PD, Haene B, Leierer J,
Heinzel A, Mühlberger I, Schweibert N, Sunzenauer J, Regele H,
Kronbichler A, et al: Renal microRNA- and RNA-profiles in
progressive chronic kidney disease. Eur J Clin Invest. 46:213–226.
2016. View Article : Google Scholar
|
|
32
|
Wang QL, Yuan JL, Tao YY, Zhang Y, Liu P
and Liu CH: Fuzheng Huayu recipe and vitamin E reverse renal
interstitial fibrosis through counteracting TGF-beta1-induced
epithelial-to-mesenchymal transition. J Ethnopharmacol.
127:631–640. 2010. View Article : Google Scholar
|
|
33
|
Hou X, Tian J, Geng J, Li X, Tang X, Zhang
J and Bai X: MicroRNA-27a promotes renal tubulointerstitial
fibrosis via suppressing PPARγ pathway in diabetic nephropathy.
Oncotarget. 7:47760–47776. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Liu Y, Bi X, Xiong J, Han W, Xiao T, Xu X,
Yang K, Liu C, Jiang W, He T, et al: MicroRNA-34a promotes renal
fibrosis by downregulation of klotho in tubular epithelial cells.
Mol Ther. 27:1051–1065. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Li H, Xu Y, Zhang Q, Xu H, Xu Y and Ling
K: Microvesicles containing miR-34a induce apoptosis of proximal
tubular epithelial cells and participate in renal interstitial
fibrosis. Exp Ther Med. 17:2310–2316. 2019.PubMed/NCBI
|
|
36
|
Denby L, Ramdas V, Lu R, Conway BR, Grant
JS, Dickinson B, Aurora AB, McClure JD, Kipgen D, Delles C, et al:
MicroRNA-214 antagonism protects against renal fibrosis. J Am Soc
Nephrol. 25:65–80. 2014. View Article : Google Scholar :
|
|
37
|
Zhang J, Jiao J, Cermelli S, Muir K, Jung
KH, Zou R, Rashid A, Gagea M, Zabludoff S, Kalluri R and Beretta L:
miR-21 inhibition reduces liver fibrosis and prevents tumor
development by inducing apoptosis of CD24+ progenitor
cells. Cancer Res. 75:1859–1867. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
McClelland AD, Herman-Edelstein M, Komers
R, Jha JC, Winbanks CE, Hagiwara S, Gregorevic P, Kantharidis P and
Cooper ME: miR-21 promotes renal fibrosis in diabetic nephropathy
by targeting PTEN and SMAD7. Clin Sci (Lond). 129:1237–1249. 2015.
View Article : Google Scholar
|
|
39
|
Yuan J, Chen H, Ge D, Xu Y, Xu H, Yang Y,
Gu M, Zhou Y, Zhu J, Ge T, et al: Mir-21 promotes cardiac fibrosis
after myocardial infarction via targeting Smad7. Cell Physiol
Biochem. 42:2207–2219. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Carew RM, Wang B and Kantharidis P: The
role of EMT in renal fibrosis. Cell Tissue Res. 347:103–116. 2012.
View Article : Google Scholar
|
|
41
|
Xu J, Lamouille S and Derynck R:
TGF-beta-induced epithelial to mesenchymal transition. Cell Res.
19:156–172. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Zhao JY, Wang XL, Yang YC, Zhang B and Wu
YB: Upregulated miR-101 inhibits acute kidney injury-chronic kidney
disease transition by regulating epithelial-mesenchymal transition.
Hum Exp Toxicol. 39:1628–1638. 2020. View Article : Google Scholar : PubMed/NCBI
|