Introduction
Claudin-7 (CLDN7), a critical member of the claudin
family, was first cloned and identified in mice by Morita et
al in 1999 (1). Its encoding
gene maps to human chromosome 17p13.1 (2). The CLDN7 protein structure
comprises four transmembrane domains, two extracellular loops and a
cytoplasmic tail region (3).
CLDN7 exhibits a specific expression pattern in human tissues. It
is highly expressed in epithelial tissues such as the
gastrointestinal tract, mammary glands, lungs and kidneys, while
showing very low expression in neural and hematopoietic tissues,
suggesting its close involvement in epithelial tissue
differentiation and barrier maintenance (4). Abnormal CLDN7 expression is
strongly linked to tumorigenesis, proliferation and metastasis in
cancers including colorectal cancer (CRC), lung cancer and breast
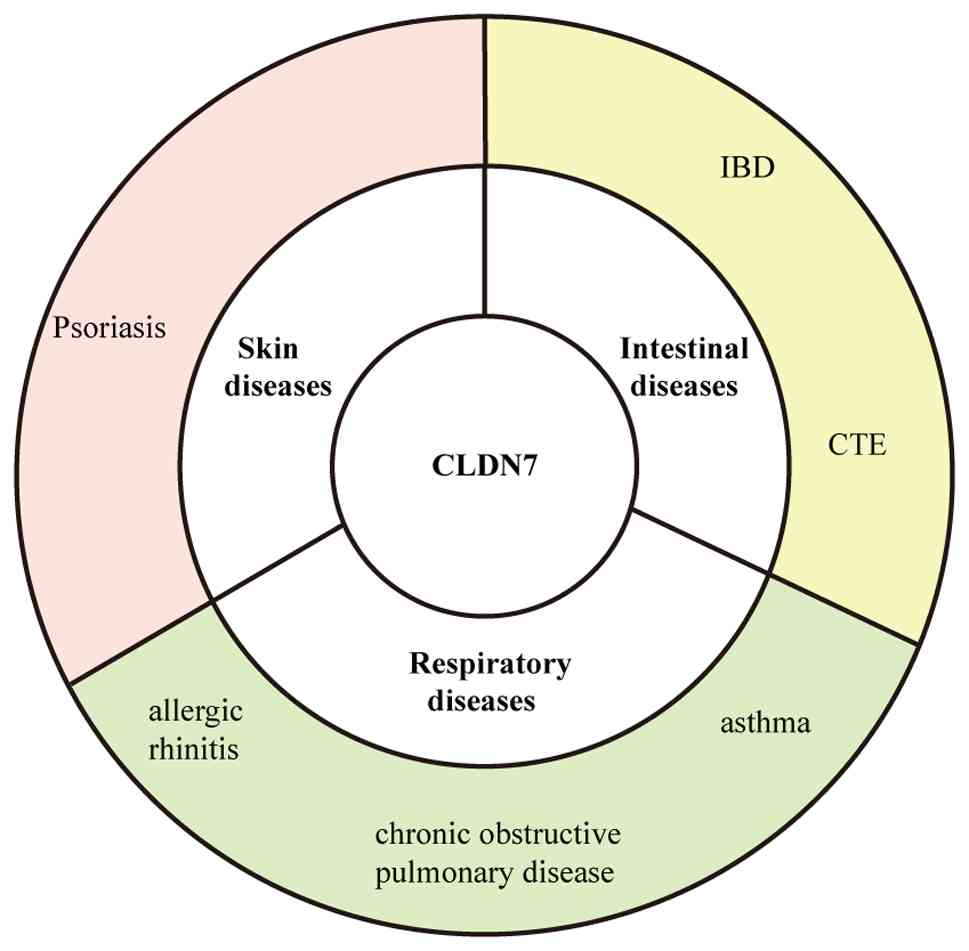
cancer (5-7). Additionally, CLDN7 plays a crucial
role in several non-neoplastic diseases, such as respiratory
disorders, intestinal diseases and skin conditions. For instance,
CLDN7 is essential for intestinal epithelial cell function and
self-renewal and its absence leads to spontaneous colitis (8,9).
Reduced CLDN7 expression levels may trigger excessive keratinocytes
proliferation and impaired barrier function, contributing to
psoriasis development (10,11). CLDN7 is widely distributed in the
respiratory barrier and is involved in respiratory diseases
including allergic rhinitis, chronic obstructive pulmonary disease
and asthma (12-14). However, there is currently a lack
of comprehensive reviews on the role and mechanisms of CLDN7 in
both neoplastic and non-neoplastic diseases. The present review
first introduced the structure, expression, regulation,
physiological functions and stability of CLDN7. Subsequently, its
expression, mechanisms of action and clinical significance in
tumors were outlined. Moreover, the role of CLDN7 in respiratory,
intestinal and skin diseases were discussed. The present review
provided a theoretical basis for developing CLDN7-targeted
therapies for both neoplastic and non-neoplastic diseases.
Methods
To comprehensively review research progress on
CLDN7, a search strategy combining Medical Subject Headings (MeSH)
and free-text keywords centered on 'CLDN7' or 'claudin-7' was
utilized. To enhance comprehensiveness, search terms also included
synonyms and narrower terms related to its functions (e.g., 'tight
junction', 'cell adhesion') and associated diseases (e.g.,
'cancer', 'carcinoma', 'colitis'). Online databases searched
included PubMed (https://pubmed.ncbi.nlm.nih.gov/), Web of Science
(https://www.webof-science.com/), and
Google Scholar (https://scholar.google.com/), with searches conducted
up to October 31, 2025; the Human Protein Atlas (HPA and Consensus
datasets, version 25.0), the iPTMnet database (https://research.bioinformatics.udel.edu/iptmnet/,
accessed September 15, 2025) and the Gene Expression Profiling
Interactive Analysis (GEPIA) database (http://gepia.cancer-pku.cn/, accessed August 28,
2025). The literature search included all relevant publications
from each database's inception up to October 31, 2025. Inclusion
criteria primarily encompassed English-language original research
articles (both preclinical and clinical), reviews and meta-analyses
explicitly examining the expression, function, or regulatory
mechanisms of CLDN7 in human diseases, particularly cancers and
inflammatory diseases.
Structure
The CLDN7 gene maps to chromosome 17p13.1,
comprising four exons and three introns. Transcription generates
three alternatively spliced mRNA variants encoding two distinct
protein subtypes (variants 1 and 2 encode the same subtype)
(https://atlasgeneticsoncology.org).
Amino acid composition, molecular formula, molecular weight and
isoelectric point of the two CLDN7 subtypes were analyzed using
ProtParam online software (http://au.expasy.org/tools/). CLDN7 subtype 1 is a
classic sequence of 211 amino acids, primarily containing 25 Ala,
23 Gly and 21 Leu, residues. Its molecular formula is
C1007H1588N256O279S21,
with a molecular weight of 22,418.49 Da, a theoretical isoelectric
point (pI) of 8.91, an estimated in vitro half-life of 30 h
and an instability index of 49.60. CLDN7 subtype 2 comprises 145
amino acids and possesses a shorter C-terminal region, lacking
amino acids 159-211 compared with subtype 1. Its composition
includes 19 Ala, 19 Leu and 17 Gly residues. Its molecular formula
is
C671H1094N172O186S19,
with a molecular weight of 15,156.25 Da, theoretical pI of 8.93 and
instability index of 57.57. All claudin family members are
quadrimembrane proteins of 20-34 kDa with highly conserved
topology: An intracellular NH2-terminus, a longer
intracellular COOH-terminus, two extracellular loops (larger ECL1,
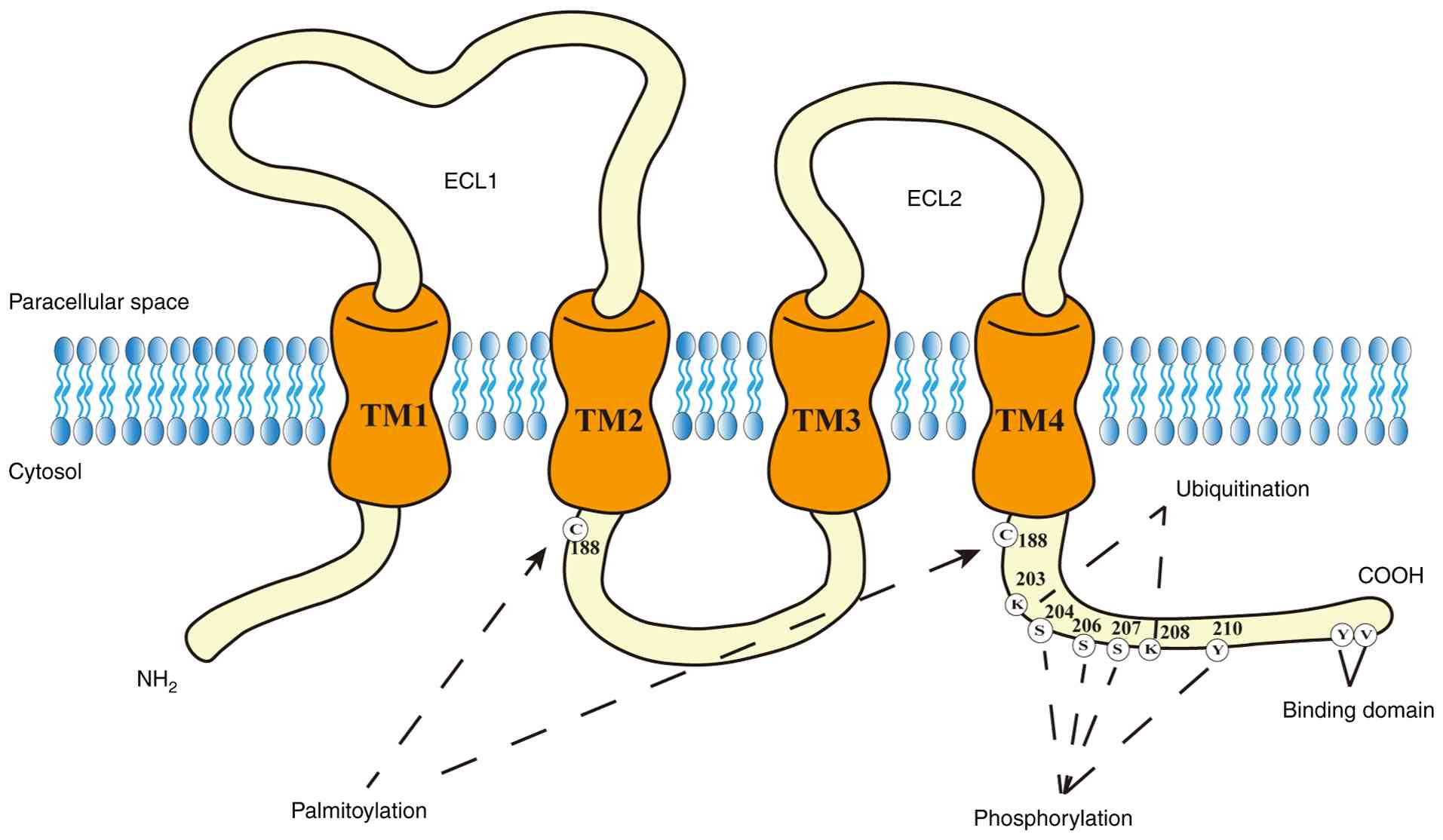
smaller ECL2) and a short intracellular loop (15). CLDN7 is a complete membrane
protein with four hydrophobic transmembrane domains, two
extracellular loops (ECL1 and ECL2), an amino terminus and a
carboxyl terminus (3). The
schematic structure of CLDN7 is illustrated in Fig. 1. ECL1 contains conserved cysteine
residues forming disulfide bonds and multiple positively charged
lysine and arginine residues. These confer high permeability to
anions (e.g., Cl−), establishing CLDN7 as a key protein
constituent of the anion channels (16). CLDN2, CLDN15 and CLDN22 are
primarily involved in cation channel formation (17). Several claudins (including CLDN1,
CLDN3, CLDN5, CLDN11 and CLDN19) possess an uncharged first
extracellular loop and thus lack ion selectivity, functioning
primarily to seal intercellular connections (18). Studies indicate that some
claudins form functional pores only through specific interactions
with other claudin molecules. For instance, the combination of
CLDN16 and CLDN19 constitutes a cation channel, whereas the pairing
of CLDN4 and CLDN8 forms an anion channel (19,20). The C-terminal tails of claudins
exhibit diversity in sequence and length and contain PDZ
domain-binding motifs (18).
These motifs anchor CLDN7 to scaffolding proteins such as ZO-1,
thereby stabilizing tight junctions (TJs) (21).
Expression and regulation of CLDN7
According to the HPA and Consensus datasets, CLDN7
shows high expression in most tissues, with tissue-specific
enhancement observed in the esophagus and intestines. Beyond the
gastrointestinal tract, CLDN7 expression is detected in epithelial
tissues including the breast, lung and prostate, but is very low or
nearly absent in muscle and neural tissues (https://www.proteinatlas.org/). Abnormal changes in
CLDN7 expression frequently occur during tumorigenesis. For
example, in CRC, CLDN7 mRNA and protein expression levels are
markedly lower than those in adjacent normal mucosa (22,23). In breast cancer, analysis of
numerous clinical samples indicates that ~50% of primary tumor
tissues show loss of CLDN7 expression (24). In ovarian cancer, CLDN7 mRNA is
highly upregulated in all four major subtypes (serous, mucinous,
clear cell and endometrioid) compared with normal ovarian tissues
(25). In non-neoplastic
diseases such as inflammatory bowel disease (IBD), reduced CLDN7
expression in the gut results in impaired barrier function
(26). Psoriasis patients
exhibit elevated expression of CLDN7 in keratinocytes, which
correlates with alterations in skin barrier function (27).
The expression of CLDN7 is regulated by multiple
factors, including primarily epigenetic mechanisms, transcription
factors, post-translational modifications and signaling pathways
(Fig. 2).
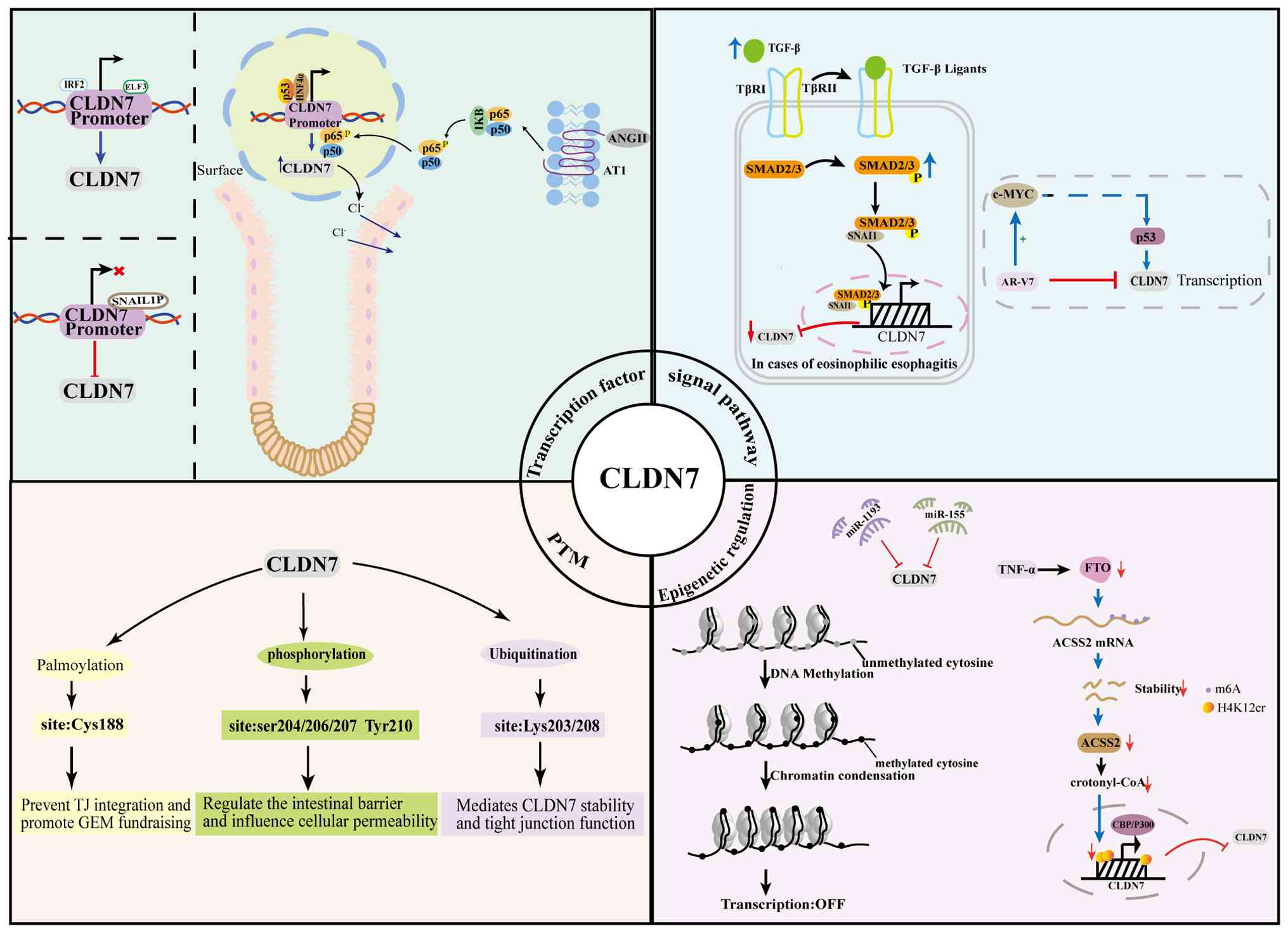 | Figure 2Schematic diagram of CLDN7 expression
and regulatory mechanism. CLDN7, claudin-7; PTM, post-translational
modification; ELF3, E74-like factor 3; IRF2, interferon regulatory
factor 2; HNF4α, hepatocyte nuclear factor 4α; p65, nuclear
factor-kappa B subunit p65; p50, nuclear factor-kappa B1; p53,
tumor protein p53; SNAIL1P, SNAIL family transcriptional repressor;
IκB, inhibitor of nuclear factor-kappa B; ANGⅡ, angiotensin II;
AT1, angiotensin II type 1 receptor; TGF-β, transforming growth
factor-beta; TβR Ⅰ and Ⅱ, transforming growth factor-beta receptor
type I and type II; SMAD2/3, SMAD family member 2 and SMAD family
member 3; c-MYC, MYC proto-oncogene; AR-V7, androgen receptor
variant 7; ACSS2, acyl-CoA synthase short-chain family member 2;
crotonyl-CoA, crotonyl coenzyme A; FTO, fat mass and
obesity-associated protein; CBP/P300, CREB-binding
protein/E1A-binding protein p300; m6A, N6-methyladenosine; H4K12cr,
histone H4 lysine 12 crotonylation; miR-155, microRNA 155;
miR-1193, microRNA 1193. |
Epigenetic regulation
Epigenetic regulation of CLDN7 expression primarily
involves microRNA (miRNA)-mediated mechanisms, DNA methylation and
histone modifications.
microRNAs (miRNAs/miRs)
miRNAs are small endogenous RNA molecules, typically
18-25 nucleotides in length. They mediate gene silencing by binding
to the 3'-untranslated region (3'-UTR) of target mRNAs, leading to
either RNA cleavage or translational repression (28). Studies have demonstrated that
miRNAs regulate CLDN7 expression.
miR-155: In eosinophilic esophagitis (EoE), miR-155
expression is markedly elevated and directly targets the 3'-UTR of
CLDN7. This reduces CLDN7 mRNA and protein expression, ultimately
impairing epithelial barrier function (29).
miR-1193: In cervical cancer, miR-1193 is markedly
downregulated, negatively regulating CLDN7 expression through
direct binding to its 3'-UTR. This suppresses the proliferation,
invasion and migration of cervical cancer cells (30).
DNA methylation
Promoter methylation induces transcriptional
silencing by preventing transcription factors from binding to their
target sites. Hypermethylation of CpG islands in the CLDN7 promoter
by DNMT3A/DNMT3B and DNMT1 causes chromatin condensation and
transcriptional repression. This disruption of the epithelial
barrier promotes tumor progression (22,31,32). Hypermethylation-induced silencing
of CLDN7 and subsequent reduced expression play a critical role in
tumor development (32).
Hypermethylation of CLDN7 occurs at the invasive front of
esophageal squamous cell carcinoma, potentially leading to CLDN7
heterogeneity and reduced expression (33). Additionally, hypermethylation of
the CLDN7 promoter region results in reduced mRNA expression,
positively correlating with deeper tumor invasion, higher
histological grade and advanced clinical stage. It is also
associated with metastasis and poor prognosis in cancers including
CRC and clear cell renal cell carcinoma (32,34).
Histone modification
Research has revealed that in IBD, TNF-α signaling
downregulates the demethylase fat mass and obesity-associated
protein, enhancing N6-methyladenosine modification and suppressing
AR-V7 androgen receptor variant 7 expression. This deficiency
impairs crotonyl-CoA synthesis, leading to reduced
CBP/p300-catalyzed histone mark H4K12cr modification at the CLDN7
promoter. Ultimately, this suppresses CLDN7 transcription by
altering the chromatin state (35).
Transcription factors
E74-like Factor 3 (ELF3): ELF3, also known as ESE-1
or ESX, is a transcription factor belonging to the E Twenty-Six
(ETS) family, encoded by the ELF3 gene on chromosome 1q32.1. It is
highly expressed in epithelial cells (36). ELF3 directly activates CLDN7
transcription by recognizing the Ets-binding site located 150 bp
upstream of the CLDN7 promoter, thereby maintaining TJs and the
epithelial phenotype (37).
Hepatocyte Nuclear Factor 4α (HNF4α): HNF4α is a
prototypical nuclear receptor transcription factor. Its expression
increases markedly along with p53 during colonic epithelial
differentiation. Together, these factors synergistically activate
CLDN7 transcription. Subsequently, CLDN7 protein localizes at the
apical TJs to form a Cl− selective channel, shifting ion
transport from basal secretion to apical absorption, thus
maintaining electrolyte homeostasis (38).
p65: p65 (RelA) is a key subunit of the nuclear
factor κB transcription factor family. In colonic epithelium, Ang
II activates the IKK signaling pathway via the AT1 receptor,
promoting nuclear translocation of p65. This translocation enables
p65 to bind the κB site in the CLDN7 promoter and recruit the
p300/CBP coactivator, forming an enhancement complex. This complex
upregulates CLDN7 expression and enhances the paracellular
permeability of TJs to Cl− (39).
Interferon regulatory factor 2 (IRF2): IRF2 is a
member of the IRF transcription factor family, which activates
transcription of multiple direct target genes and regulates immune
responses and immune cell development (40,41). In oral squamous cell carcinoma
(OSCC), IRF2 binds the promoter region of CLDN7 and induces CLDN7
expression, thereby inhibiting proliferation, invasion and
migration of OSCC cells (42).
SNAIL: SNAIL1 (Snail) and SNAIL2 (Slug) belong to
the Snail family of zinc-finger transcriptional repressors
(43). In esophageal squamous
cell carcinoma and breast cancer cell lines, Snail and its homolog
SNAILP suppress CLDN7 transcription by binding to the E-box of its
promoter (44,45). Additionally, Slug may regulate
CLDN7 expression in squamous cell carcinoma and adenocarcinoma of
the lung by binding to the E-box on the CLDN7 promoter; however,
further validation is required (46).
Post-translational modifications
(PTMs)
PTM refers to covalent, enzymatic, or non-enzymatic
addition of specific chemical groups to amino acid side chains.
Such modifications include phosphorylation, palmitoylation,
ubiquitination, glycosylation and acetylation, profoundly affecting
protein conformation, stability, trafficking and function (47). PTM sites of CLDN7 are shown in
Fig. 1.
Palmitoylation
Palmitoylation occurs at Cys188, preventing CLDN7
incorporation into TJs. Instead, it redirects CLDN7 to
glycolipid-enriched membrane microdomains, influencing membrane
microdomain localization and downstream signaling (48).
Phosphorylation
Phosphorylation is the most extensively studied PTM
of CLDN7. Phosphorylation at Ser206 mediated by WNK4 reduces CLDN7
integration into TJs and enhances paracellular permeability to
Cl− (49).
Conversely, phosphorylation at Ser204/S207 and Tyr210 promotes
CLDN7 assembly at TJs, strengthening epithelial barrier function
(50,51).
Ubiquitination
Research on CLDN7 ubiquitination remains relatively
limited. According to annotations from the iPTMnet database,
ubiquitination of CLDN7 occurs primarily at lysine 203 and lysine
208 (https://research.bioinformatics.udel.edu/iptmnet/,
accessed September 15, 2025). Existing studies suggest that this
modification process may be dynamically regulated by E3 ubiquitin
ligases (such as LNX1) and deubiquitinating enzymes (such as
USP28), thereby influencing CLDN7 protein stability and
tight-junction (TJ) function (52,53).
Signaling pathway
In hepatocellular carcinoma, the androgen receptor
variant 7 (AR-V7) collaborates with c-MYC to counteract downstream
transcriptional activation of CLDN7 by p53. This markedly
suppresses CLDN7 expression, disrupting epithelial barrier
integrity and accelerating c-MYC-driven tumor progression (54). Activation of the transforming
growth factor-β (TGF-β) signaling pathway results in SMAD2/3
phosphorylation. Phosphorylated SMAD2/3 forms a transcriptional
repression complex with SNAL1, which directly binds to the CLDN7
promoter. This interaction inhibits CLDN7 transcription, leading to
reduced expression and impaired epithelial barrier integrity
(55).
Stability of CLDN7
Multiple factors influence CLDN7 stability,
including its oligomerization properties, degradation processes and
interactions with other proteins. Purified CLDN7 forms multiple
oligomeric structures (monomers, dimers, trimers and higher-order
oligomers) in detergents. These oligomers may contribute to CLDN7
stability and function within the cell membrane (56). CLDN7 degradation also affects its
stability. CLDN7 undergoes degradation not only by MMP7 hydrolysis
(57), but also through the
ubiquitin-proteasome pathway mediated by E3 ubiquitin ligases
(58,59). Additionally, CLDN7 protein
degradation involves lysosomal pathway: Lysosomes serve as major
intracellular degradation sites, encapsulating and degrading
proteins through endocytosis and autophagy (60). Epithelial cell adhesion molecule
(EpCAM) interacts with CLDN7, stabilizing it on epithelial cells
surfaces (61,62). Further research demonstrates that
EpCAM and trophoblast cell surface antigen 2 (Trop2) share
regulatory functions for CLDN7, allowing partial compensation.
Dual-knockout mice lacking both proteins exhibit severe impairment
in CLDN7 expression and localization (63). The absence of EpCAM leads to
CLDN7 loss in mouse intestinal epithelial cells, causing TJ
defects. Interestingly, this deletion does not reduce CLDN7 mRNA
levels, suggesting that EpCAM regulates CLDN7 expression via
post-transcriptional mechanisms (64). Moreover, EpCAM protects CLDN7
from endocytosis and lysosomal degradation (61,62). These findings indicate that EpCAM
stabilizes CLDN7 proteins at the cell surface and that the direct
interaction between these two proteins is essential for CLDN7
stability.
The physiological functions of CLDN7
Barrier function
CLDN7 localizes not only to apical regions of
intestinal epithelial cells but also strongly distributes along
basolateral membranes in the intestine (65). Intestinal epithelial integrity is
essential for nutrient absorption and defense against pathogens.
CLDN7 forms a physical barrier in intestinal epithelial cells,
isolating harmful substances (such as bacteria, toxins and
antigens) from the internal environment (34). Intestinal-specific CLDN7 knockout
mice exhibit severe intestinal defects, including mucosal
ulceration, epithelial cell shedding, inflammation and elevated
matrix metalloproteinase expression (8,66). In normal intestine, integrin α2
forms a stable protein complex with CLDN7 and CLDN1. This complex
is disrupted in CLDN7-deficient mice. Furthermore, the expression
and localization of integrin α2 are also altered, highlighting
CLDN7's role in cell-matrix interactions (66). Additionally, CLDN7 maintains
intestinal homeostasis and prevents IBD and colorectal cancer (CRC)
progression. In intestinal-specific CLDN7 knockout mice, CLDN7
deficiency promotes colitis and subsequent CRC progression by
disrupting TJ integrity and amplifying inflammation (8).
Stem cell fate regulation
CLDN7 regulates small intestinal stem cell (ISC)
functions, influencing ISC survival, self-renewal and epithelial
differentiation (9,67). The balance between stem cell
homeostasis and cell fate determination in the intestine depends
upon interactions between Wnt and Notch signaling pathways
(68,69). CLDN7 precisely regulates the
Wnt/β-catenin signaling pathway through direct interaction with
β-catenin. This ensures moderate activation, maintaining stable
expansion and stemness of Lgr5+/Olfm4+
intestinal stem cells. Concurrently, CLDN7 synergizes with Notch
signaling, suppressing excessive differentiation toward the
secretory cell lineage and promoting steady-state renewal and
differentiation of absorptive epithelial cells (9). A recent study revealed that CLDN7
regulates colonic stem cell behavior by modulating the Notch and
Hippo signaling pathways. Activation of Notch signaling or
inhibition of Hippo signaling rescues defects caused by CLDN7
deficiency (70). Under CLDN7
deficiency, β-catenin stability decreases, becoming susceptible to
degradation. This weakens Wnt signaling pathway activity, leading
to stem cell depletion and differentiation toward secretory
lineages (9). Exogenous
activation of Wnt signaling or restoration of CLDN7 expression
effectively reverses this pathological process, restores intestinal
epithelial homeostasis and promote regenerative repair after injury
(67).
Renal ion transport
CLDN7 is crucial for renal ion transport and
localizes primarily to the distal tubules, collecting ducts and the
thick ascending limb of Henle's loop in the kidney (71). Renal phenotypes indicate that
CLDN7 is essential for maintaining salt homeostasis in the distal
nephron. CLDN7 forms non-selective paracellular channels that
facilitate Cl− and Na+ reabsorption in the
collecting duct. When overexpressed, CLDN7 reduces Cl−
conductance and increases Na+ conductance, indicating it
promotes NaCl reabsorption through charge-selective channels
(72). Negatively charged amino
acids (D38, E53) within the first extracellular domain (ED1) of
CLDN7 are critical for the paracellular Cl− barrier.
Mutating these residues to positive charges markedly increases
Cl− permeability without affecting Na+
permeability or TJ ultrastructure. Thus, ED1 determines charge
selectivity, while ED2 has minimal impact (16). CLDN7 knockout mice die shortly
after birth, partly due to severe renal salt depletion and
dehydration (73). In the renal
collecting duct, CLDN7 is phosphorylated by WNK4 kinase, with
Ser206 of the COOH-terminus region being the phosphorylation site
(49). WNK4-mediated
phosphorylation of CLDN7 regulates paracellular chloride
permeability, maintaining salt balance. Gain-of-function mutations
in WNK4 lead to CLDN7 hyperphosphorylation, increased chloride
leakage, impaired sodium chloride reabsorption and abnormal
potassium handling (74). The
WNK4-CLDN7 phosphorylation mechanism is a key regulator of
salt-sensitive hypertension (75).
CLDN7 and diseases
CLDN7 and tumors
Expression of CLDN7 in tumors
CLDN7 expression is widely dysregulated in numerous
malignant tumors, including lung, colorectal, ovarian, breast,
gastric, esophageal and prostate cancers and is closely associated
with tumor progression and metastasis (34,76,77). Expression profiles of CLDN7
across different cancer types are systematically summarized in
Table I. Research indicates that
abnormal CLDN7 expression and disrupted localization affect
tumorigenesis through dual mechanisms. On one hand, compromised
epithelial barrier integrity occurs. On the other hand, it
interferes with fundamental cellular regulatory networks
(proliferation, differentiation, apoptosis), disrupting homeostasis
and potentially inducing cellular transformation and tumor
development (78). Notably, loss
of CLDN7 expression often correlates with impaired cell adhesion
and malignant progression (79).
However, some studies suggest that elevated CLDN7 expression may
enhance tumor cell migration, invasion and metastasis (80). These findings indicate that CLDN7
may have bidirectional roles in tumorigenesis, acting either as a
tumor suppressor or a pro-oncogenic factor.
 | Table IThe changes in the expression pattern
of CLDN7 in tissues. |
Table I
The changes in the expression pattern
of CLDN7 in tissues.
| Cancer | Methodology | Sample size | Positive rate,
% | CLDN7 expression
level | (Refs.) |
|---|
| Colorectal
cancer | IHC | 11 | 28 | Downregulation | (82) |
| Lung cancer | RT-qPCR, WB | - | - | Downregulation | (5,78) |
| Ovarian cancer | IHC, WB | 29 | 66 | Upregulation | (25) |
| Breast cancer | RT-qPCR | 65 | 29.23 | Downregulation | (83) |
| Nasopharyngeal
carcinoma | IHC | 18 | 55.5 | Upregulation | (84) |
| Oral squamous cell
carcinoma | IHC | 66 | 93.3 | Upregulation | (34) |
| Liver cancer | IHC | 67 | 91.04 | Upregulation | (85) |
| Gastric cancer | IHC | 134 | 29.9 | Upregulation | (86) |
| Prostate
cancer | IHC | 141 | - | Upregulation | (87) |
| Thyroid cancer | IHC | 107 | 94 | Downregulation | (88) |
| Triple-negative
breast cancer | IHC | 222 | 32.4 | Upregulation | (89) |
| Cervical
adenocarcinoma | IHC | 55 | - | Upregulation | (90,91) |
| Pancreatic
cancer | IHC | 141 | - | Downregulation | (92) |
| Endometrial
cancer | IHC | 31 | 54.8 | Downregulation | (93) |
| Esophageal
adenocarcinoma | IHC | 227 | 70 | Upregulation | (94) |
| Esophageal squamous
cell carcinoma | IHC,WB | 40 | 30 | Downregulation | (51,95) |
| Lung
adenocarcinoma | IHC | 86 | - | Upregulation | (96) |
| Cervical
adenocarcinoma | IHC | 55 | - | Upregulation | (90) |
| Superficial
(non-muscle-invasive) urothelial bladder carcinoma | IHC | 111 | 22 | Downregulation | (97) |
| Adenoid cystic
carcinoma of the salivary gland | IHC | 50 | 38 | Downregulation | (98) |
| Human throat
cancer | IHC | 80 | 22.5 | Downregulation | (99) |
Regulation of tumor phenotypes by
CLDN7
Tumor proliferation
CLDN7 exhibits variable expression levels across
different tumors, influencing cancer cell proliferation (81,82). Furthermore, CLDN7 regulates tumor
cell migration and invasion through barrier function and signaling
pathways (7,8). Regulatory mechanisms of CLDN7 in
cancer are illustrated in Fig.
3.
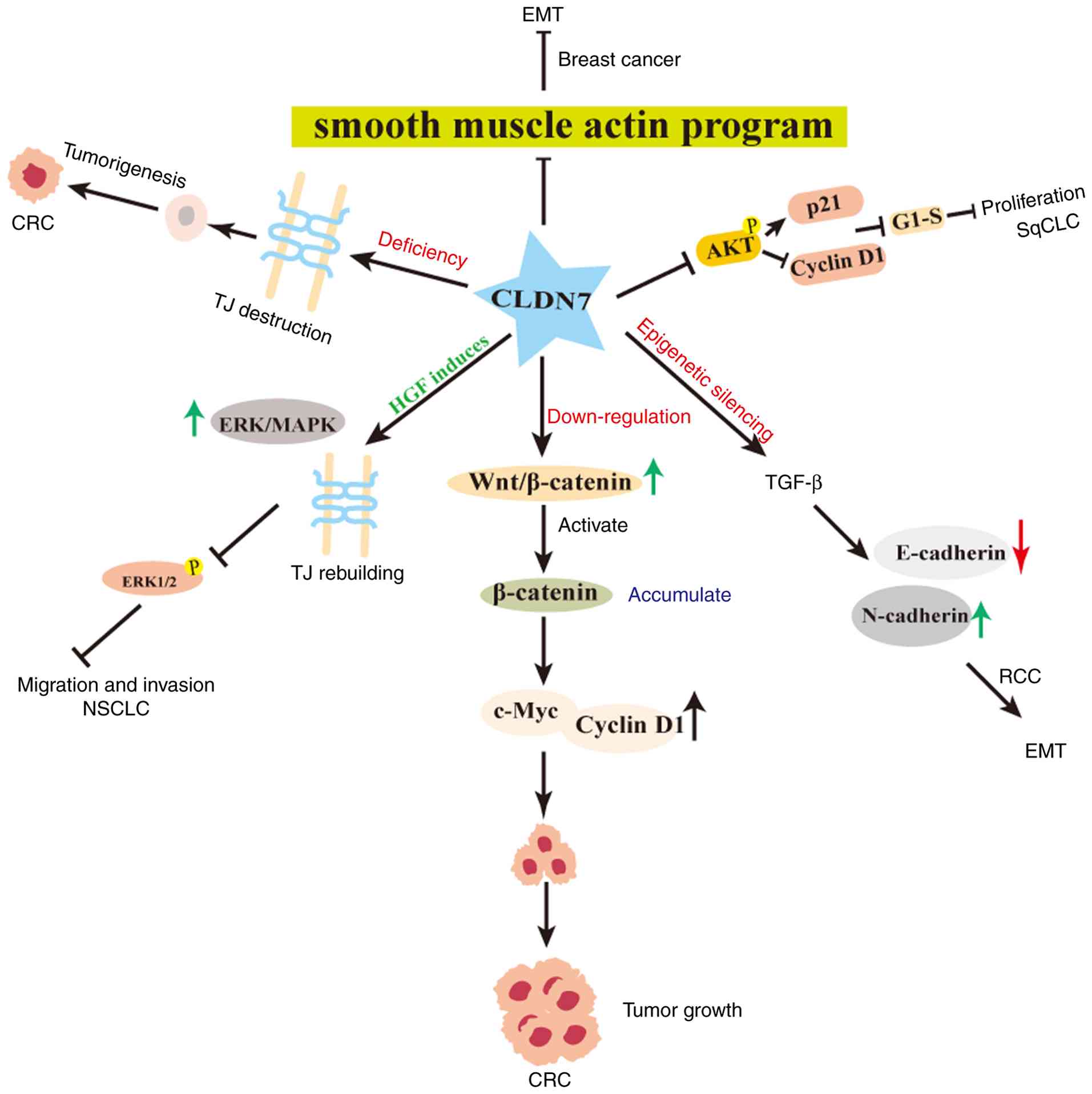 | Figure 3Regulatory mechanisms of CLDN7 in
cancer. CLDN7, claudin-7; EMT, epithelial-mesenchymal transition;
AKT, AKT serine/threonine kinase; p21, cyclin-dependent kinase
inhibitor 1A; Cyclin D1, G1/Specific Cyclin-D1; Wnt/β-catenin,
wingless/integrated/β-catenin; c-Myc, MYC proto-oncogene; TGF-β,
transforming growth factor-beta; SqCLC, squamous-cell lung cancer;
RCC, renal cell carcinoma; CRC, colorectal cancer; NSCLC,
non-small-cell lung cancer. |
Cell proliferation is closely associated with the
cell cycle and CLDN7 inhibit tumor cell cycle progression. Its
tumor-suppressive role in CRC depends on p53. In p53 wild-type
cells, CLDN7 induces G0/G1 phase arrest and
apoptosis (6). In squamous lung
cancer cells, CLDN7 accelerates the interaction between PDK1 and
AKT, inhibits sustained AKT phosphorylation, upregulates p21,
decreases Cyclin D1 levels and suppresses the G1-S
transition, thus inhibiting cell proliferation (5). Through immunofluorescence
co-localization and immunoprecipitation, Lu et al
demonstrated that CLDN7 co-localizes with integrin β1, forming a
protein complex in human lung cancer cells, contributing to its
anti-proliferative effects (83). Moreover, in CRC tissues, CLDN7
knockout mouse models and CRC cell lines have shown that its loss
suppresses tumor growth, migration and apoptosis via a
SOX-9-mediated Wnt/β-catenin pathway (7). Therefore, CLDN7 can suppress tumor
proliferation by regulating the cell cycle and modulating signaling
pathways.
Tumor migration and invasion
CLDN7 regulates tumor migration and invasion mainly
by controlling barrier function and epithelial-mesenchymal
transition (EMT) (76,84). CLDN7 also modulates signaling
pathways (such as Wnt/β-catenin), suppressing tumor metastasis
(81).
The intestinal epithelial barrier protects against
infection and injury. Chronic inflammation is a critical factor in
tumorigenesis (85). In normal
epithelium, CLDN7 localizes at the apical TJs, forming a
high-resistance barrier that prevents tumor cells from penetrating
the basement membrane and infiltrating into the stroma (86). In an AOM/DSS-induced
colitis-carcinoma model, CLDN7 deficiency disrupts TJ integrity,
increases mucosal permeability and disrupts linear E-cadherin
staining (8). Additionally,
CLDN7 knockout mice exhibit severe enteritis, disrupted epithelium,
necrotic glands and markedly loosened basement membrane junctions
(87).
EMT promotes tumor dissemination by causing loss of
epithelial markers and gain of mesenchymal markers. EMT disrupts
intercellular adhesion, detaching epithelial cells from the basal
layer, thus enhancing motility (76). CLDN7 is an epithelial marker in
multiple tumors and its expression regulates EMT, influencing tumor
metastasis (32,88). In CRC, reduced CLDN7 expression
correlates with lower histological grade, facilitating invasion and
metastasis through EMT regulation (76). Moreover, APC loss or β-catenin
mutation activates Wnt signaling, downregulating CLDN7 and
E-cadherin, while upregulating Snail/ZEB1/Vimentin, disrupting TJs
and initiating EMT (89). In
renal cell carcinoma, epigenetic silencing of CLDN7 promotes
TGF-β-driven EMT (E-cadherin↓, N-cadherin↑) and restoring CLDN7
expression can reverse EMT, suppressing local invasion and distant
metastasis (32). In salivary
adenoid cystic carcinoma (SACC), CLDN7 knockdown enhances
metastasis, decreases E-cadherin expression and increases
N-cadherin and vimentin expression (81). Furthermore, West et al
(84) used mouse models, human
organoids and multi-omics data to demonstrate that CLDN7 blocks EMT
by inhibiting SMA-actin, thereby reducing breast cancer invasion
and metastasis.
In the canonical Wnt/β-catenin pathway, β-catenin
accumulates in the cytoplasm, enters the nucleus and promotes tumor
progression by activating downstream genes (90). CLDN7 silencing activates
Wnt/β-catenin signaling, promoting proliferation and metastasis in
SACC and CRC. In SACC, nuclear β-catenin accumulation accompanies
EMT, whereas in CRC, β-catenin drives transcription of c-Myc and
Cyclin D1, promoting CRC cell proliferation and metastasis
(7,84). Additionally, hepatocyte growth
factor induction enables CLDN7 to re-establish TJs and suppress
ERK/MAPK signaling in human lung cancer cells, markedly reducing
ERK1/2 phosphorylation and thereby inhibiting non-small cell lung
cancer cell migration and invasion (77).
Clinical significance of CLDN7 in
tumors
Analysis using the GEPIA database indicates that
CLDN7 expression correlates with overall survival (OS) and
disease-free survival (DFS) (Fig.
4). In low-grade glioma (LGG) and glioblastoma multiforme, high
CLDN7 expression is associated with poorer OS. In clear cell renal
cell carcinoma and uveal melanoma, low CLDN7 expression predicts
poorer OS. High CLDN7 expression correlates with reduced DFS in
adrenal cortical carcinoma and LGG, whereas the opposite occurs in
clear cell renal cell carcinoma. Histopathological analysis of
pancreatic and nasopharyngeal carcinoma tissues shows a
statistically significant association between reduced CLDN7
expression and lower survival rates (91). In gastric cancer, high CLDN7
expression correlates with shorter survival (92). Overexpression of CLDN7 in breast
cancer is associated with poorer DFS (88). In colon cancer, low CLDN7
expression and positive perineural infiltration are independent
predictors of poor DFS (93).
Thus, CLDN7 may serve as a prognostic biomarker in various
cancers.
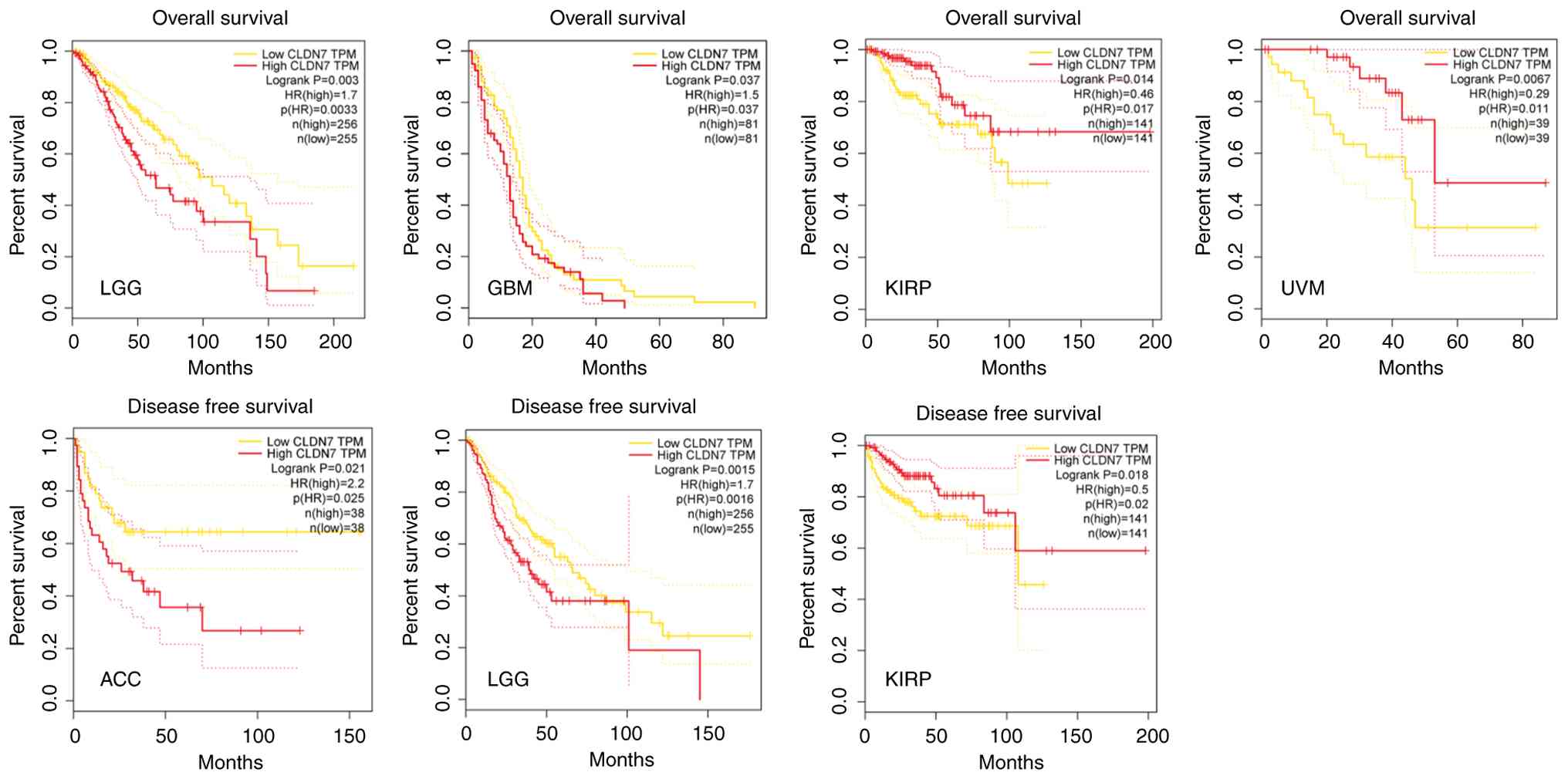 | Figure 4Correlation between CLDN7 levels and
OS or DFS in different tumor tissues. Data were retrieved from the
Gene Expression Profiling Interactive Analysis (GEPIA) public
database (http://gepia.cancer-pku.cn). CLDN7,
claudin-7; OS, overall survival; DFS, disease-free survival; TPM,
transcripts per million; HR, hazard ratio; LGG, low-grade glioma;
GBM, glioblastoma multiforme; KIRP, clear cell renal cell
carcinoma; UVM, uveal melanoma; ACC, adrenal cortical
carcinoma. |
CLDN7 downregulation is closely associated with
aggressive phenotypes and poor prognosis across multiple tumor
types. Knockdown of CLDN7 markedly increases ERK1/2
phosphorylation, promoting cell proliferation and invasion, whereas
restoration of CLDN7 expression markedly attenuates these malignant
phenotypes (83).
Clinicopathological analyses demonstrate markedly reduced CLDN7
expression in peritumoral tissues and metastatic lymph nodes of
esophageal cancer. Its loss correlates positively with deeper tumor
invasion, lymphatic vessel invasion and nodal metastasis,
suggesting that CLDN7 may predict lymph-node metastasis (33). At the invasive front of
esophageal squamous-cell carcinoma, CLDN7 is frequently
hypermethylated, causing heterogeneous downregulation. Its deletion
decreases E-cadherin expression, promoting tumor growth and
invasion, while abnormal localization contributes to transformation
and E-cadherin dysregulation (94). Similarly, loss of CLDN7 in oral
squamous-cell carcinoma correlates with high tumor grade, advanced
TNM stage, vascular invasion and regional lymph-node involvement
(95). Low CLDN7 expression in
breast and endometrial cancers correlates with higher histological
grade and distant metastasis, representing an independent
prognostic factor (96,97). In non-small-cell lung cancer,
ectopic CLDN7 expression enhances cisplatin-induced apoptosis via
caspase activation, thereby increasing drug sensitivity (98).
Overexpression of CLDN7 may promote malignant tumor
behavior (34). Microarray
analysis of esophageal adenocarcinoma reveals markedly higher CLDN7
transcription levels compared with adjacent normal mucosa,
suggesting that its activation may represent an early event in
tumorigenesis (95).
Palmitoylated CLDN7 forms complexes with MMP-family proteins and
CD147, potentially promoting tumor metastasis (48). Gastric cancer also exhibits
sustained CLDN7 upregulation, which progressively increases with
disease stage (99).
Adenocarcinomas of the prostate (100), pancreas (100), cervix (101) and ovary (25) frequently display CLDN7
overexpression, associated with poor prognosis. Conversely,
silencing CLDN7 via RNA interference markedly reduces invasive and
migratory capabilities of cancer cells (82,102). In ovarian cancer cells, high
CLDN7 expression is associated with platinum resistance, whereas
its knockdown restores cisplatin sensitivity (103).
Currently, research on CLDN7-targeted anti-cancer
drugs remains at the preclinical stage and no specific CLDN7 agents
have received clinical approval. However, several compounds
modulate CLDN7 expression or function to exert anti-tumor effects.
In CRC, the methylation inhibitor 5-aza-2'-deoxycytidine reverses
promoter methylation, markedly increasing CLDN7 expression
(31,104). Additionally, 2-deoxy-D-glucose
(2-DG) inhibits glycolysis, blocking tumor-derived CLDN7-mediated
metabolic reprogramming and immunosuppressive effects of
tumor-associated neutrophils, thus indirectly reducing the
tumor-promoting role of CLDN7 (82).
Role of CLDN7 in non-cancer diseases
CLDN7 also contributes to the development of
intestinal, skin and respiratory diseases (Fig. 5).
CLDN7 and intestinal diseases
IBD
IBD represents a group of disorders characterized by
intestinal epithelial barrier damage and chronic inflammation
(105). CLDN7 expression is
markedly reduced in IBD and patients exhibit persistent epithelial
hyperpermeability and microbial invasion (106). CLDN7 deficiency correlates with
impaired gut epithelial integrity, exacerbated intestinal
inflammation and increased matrix metalloproteinase expression
(57). Furthermore, CLDN7
deficiency selectively increases paracellular permeability to
small-molecule organic solutes, enhancing bacterial product
infiltration and exacerbating intestinal inflammation (107). Analysis of gut microbiota in
CLDN7-deficient mice demonstrates reduced microbial diversity,
intensifying DSS-induced inflammation (108). When therapeutic interventions
mitigate intestinal injury, CLDN7 expression is restored, promoting
intestinal epithelial recovery (109).
Congenital tufting enteropathy
(CTE)
CTE is a rare intestinal disorder characterized by
intractable neonatal diarrhea and intestinal epithelial tufting
(110). CTE primarily results
from loss-of-function mutations in the EpCAM gene. A subset of
cases involves mutations in SPINT2, which normally protects EpCAM
from matriptase-mediated degradation; SPINT2 mutations also result
in near-complete EpCAM loss (111,112). Studies show that
loss-of-function mutations in EpCAM or SPINT2 cause excessive
degradation of CLDN7, severely impairing or abolishing its
stability (111,112). Despite distinct genetic
origins, both mutations ultimately lead to significant CLDN7
dysfunction, disrupting TJ structure and intestinal epithelial
barrier function. This breakdown is a major contributor to severe
diarrhea characteristic of CTE.
CLDN7 and skin diseases
Psoriasis is an inflammatory skin disease
characterized by excessive keratinocyte proliferation, impaired
barrier function and marked inflammatory cell infiltration
(113). CLDN7 expression is
dysregulated in psoriasis. Kirschner et al (11) detected markedly reduced CLDN7
expression in early-stage psoriasis using immunofluorescence and
real-time PCR. Additionally, CLDN7 localization at the cell
membrane is diminished in the basal and granular layers, indicating
compromised barrier integrity. Another study demonstrated that
pro-inflammatory cytokines, such as IL-36γ, downregulate CLDN7
expression in keratinocytes, exacerbating epidermal barrier defects
and driving inflammatory responses (10). Conversely, elevated CLDN7
expression occurs in the granular epidermal layer of psoriasis
patients and inhibition of HMG-CoA reductase reduces this elevation
(27). Thus, CLDN7 likely
contributes to psoriasis pathogenesis.
CLDN7 and respiratory diseases
CLDN7 is broadly expressed in the respiratory
barrier and is implicated in respiratory diseases, including
allergic rhinitis (AR), chronic obstructive pulmonary disease
(COPD), and asthma.
AR
AR is a common condition affecting ~400 million
individuals worldwide (114).
Although AR is not life-threatening, it markedly disrupts daily
activities. Reports indicate that damage to the nasal epithelium is
crucial in AR pathogenesis (115,116). Nasal epithelium from AR
patients exhibits markedly lower CLDN7 transcript levels compared
with non-allergic controls. This reduction strongly associates with
secondhand smoke exposure, where oxidative stress disrupts the
CLDN7-mediated TJ barrier, increasing susceptibility to AR
(14).
COPD
Long-term exposure to tobacco smoke and particulate
matter (PM) disrupts TJs in airway epithelium, increasing barrier
permeability, mucus secretion and bacterial translocation in COPD
(12). Studies indicate markedly
reduced mRNA expression levels of TJ proteins (CLDN1, CLDN3, CLDN7,
CLDN15) in COPD, suggesting their potential as novel biomarkers for
early COPD diagnosis (117).
Asthma
Dysfunction of airway epithelial cells in asthma is
associated with impaired wound healing, compromised TJs and
excessive cell proliferation. These factors contribute to abnormal
airway responses to external pathogens (118-120). Asthmatic patients exhibit lower
plasma CLDN7 levels, positively correlating with lung function
(FEV1/FVC). Additionally, CLDN7 expression is suppressed by
titanium dioxide exposure, suggesting that PM exposure may induce
airway epithelial barrier dysfunction, inflammation and
hyperresponsiveness (13).
Exposure to diesel exhaust particles similarly alters CLDN4, CLDN5
and CLDN17 expression in mouse nasal passages and lungs (121). Such consistent regulation
across respiratory barriers offers potential therapeutic targets
for airway diseases.
In summary, CLDN7 markedly influences multiple
non-tumor diseases, including intestinal disorders, skin conditions
and respiratory diseases. Additionally, CLDN7 plays a critical role
in salt homeostasis in renal collecting duct cells through
modulation by WNK4 and regulation of NaCl transport (75). These findings highlight the broad
involvement of CLDN7 in regulating various non-neoplastic
conditions.
Several agents modulate CLDN7 expression and
function. Compounds aiming to restore barrier integrity typically
upregulate CLDN7 expression. Metformin reverses TNF-α-induced
reductions in CLDN7 transcription and protein levels in intestinal
organoids by activating AMPK (122). Nobiletin restores CLDN7
expression and TJ integrity by recruiting HNF4α to directly bind
the CLDN7 promoter (109).
Polysaccharides from Atractylodes macrocephala Koidz
upregulates CLDN7 by simultaneously inhibiting MAPK-mediated
MMP-7/8 activity (reducing protein degradation) and activating the
JAK-STAT-IRF1 pathway (enhancing transcription), thus restoring TJ
function (123). Conversely,
prolonged treatment with the MAPK agonist anisomycin selectively
activates p38, causing removal of CLDN7 from TJs, reducing
transepithelial electrical resistance and increasing paracellular
permeability (124).
Furthermore, vitamin D upregulates CLDN7 expression in active
ulcerative colitis, enhancing barrier recovery and reducing
inflammatory infiltration (125). These agents remain in
preclinical stages, primarily tested in colitis or cancer models.
If clinical trials confirm their efficacy and safety, their
application may extend to other inflammatory or epithelial
barrier-related diseases beyond the gastrointestinal tract.
Conclusions and perspectives
CLDN7, a crucial member of the claudin family,
comprises four transmembrane domains, two extracellular loops and
N- and C-terminal regions. It shows high expression in various
epithelial tissues and is regulated by transcription factors,
post-translational modifications and epigenetic mechanisms. Beyond
its classical barrier function, CLDN7 also regulates stem cell fate
and ion transport, maintaining epithelial tissue homeostasis.
CLDN7 exhibits functional dysregulation in both
cancer and non-cancer diseases, displaying bidirectional roles and
distinct pathological outcomes. In tumors, CLDN7 expression loss
correlates with EMT and elevated metastatic risk. Conversely, its
overexpression promotes malignant progression through mechanisms
such as palmitoylation and MMP-CD147 complex formation, correlating
with poor prognosis in multiple malignancies. In non-cancer
diseases, CLDN7 expression is typically downregulated, leading to
increased epithelial/mucosal barrier permeability and contributing
to chronic inflammation. Mechanistically, both disease categories
involve CLDN7-mediated disruption of barrier integrity and related
signaling pathway alterations. Overall, the pathological roles of
CLDN7 depend on expression levels, post-translational modifications
and subcellular localization, reflecting context-dependent
functionality across diverse diseases. Although no CLDN7-selective
targeted drugs have entered clinical use, preclinical studies
demonstrate that agents, including epigenetic modulators,
glycolysis inhibitors, AMPK activators and natural products,
effectively influence epithelial barrier function, EMT and tumor
cell chemosensitivity by modulating CLDN7 expression.
However, clinical translation of CLDN7 still faces
several challenges: i) Current understanding of CLDN7's mechanisms
in tumors and non-neoplastic diseases remains insufficient,
necessitating further research to support drug development and
clinical application. ii) As CLDN7 is expressed in most normal
tissues, potential side effects require careful consideration in
targeted therapy design. iii) CLDN7 exhibits bidirectional
prognostic significance, high in some cancers and low in others,
even among different cancer subtypes, creating uncertainty in
indication selection and risk assessment. Additionally, several
unknowns and future research directions remain: Clarifying the
reasons underlying differential CLDN7 expression across tumors and
determining if CLDN7 regulatory signaling pathways are tissue- or
cell-type-specific in neoplastic and non-neoplastic diseases.
In summary, future studies should elucidate the
tissue-specific expression patterns of CLDN7 and regulatory
mechanisms underlying its bidirectional diseases roles, further
refining its pathological regulatory network. CLDN7 has the
potential to serve as a novel biomarker for tumor and non-tumor
inflammatory diseases, facilitating early diagnosis and prognosis.
Meanwhile, targeted drug development based on CLDN7's regulatory
mechanisms, combined with tissue-specific delivery technologies to
minimize off-target effects, may offer promising therapeutic
strategies, demonstrating considerable potential for clinical
translation.
Availability of data and materials
Not applicable.
Authors' contributions
XL and YY collected literatures, created the
figures and wrote the first draft of the manuscript. LS collected
literatures. ZL and YHY conceived the review, analyzed the relevant
literatures and critically revised the manuscript. Data
authentication is not applicable. All authors read and approved the
final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Acknowledgements
Not applicable.
Funding
The present study was supported by the National Natural Science
Foundation of China (grant no. 82171855) and the Key Field Special
Project for Colleges and Universities of Guangdong Province
(Biomedicine and Health; grant no. 2023ZDZX2030).
References
|
1
|
Morita K, Furuse M, Fujimoto K and Tsukita
S: Claudin multigene family encoding four-transmembrane domain
protein components of tight junction strands. Proc Natl Acad Sci
USA. 96:511–516. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Kozieł MJ, Kowalska K and
Piastowska-Ciesielska AW: Claudins: New players in human fertility
and reproductive system cancers. Cancers (Basel). 12:7112020.
View Article : Google Scholar
|
|
3
|
Gonçalves A, Ambrósio AF and Fernandes R:
Regulation of claudins in blood-tissue barriers under physiological
and pathological states. Tissue Barriers. 1:e247822013. View Article : Google Scholar
|
|
4
|
Ouban A and Ahmed AA: Claudins in human
cancer: A review. Histol Histopathol. 25:83–90. 2010.
|
|
5
|
Akizuki R, Shimobaba S, Matsunaga T, Endo
S and Ikari A: Claudin-5, -7, and -18 suppress proliferation
mediated by inhibition of phosphorylation of Akt in human lung
squamous cell carcinoma. Biochim Biophys Acta Mol Cell Res.
1864:293–302. 2017. View Article : Google Scholar
|
|
6
|
Hou Y, Hou L, Liang Y, Zhang Q, Hong X,
Wang Y, Huang X, Zhong T, Pang W, Xu C, et al: The p53-inducible
CLDN7 regulates colorectal tumorigenesis and has prognostic
significance. Neoplasia. 22:590–603. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Xu C, Ding YH, Wang K, Hao M, Li H and
Ding L: Claudin-7 deficiency promotes stemness properties in
colorectal cancer through Sox9-mediated Wnt/β-catenin signalling. J
Transl Med. 19:3112021. View Article : Google Scholar
|
|
8
|
Wang K, Ding Y, Xu C, Hao M, Li H and Ding
L: Cldn-7 deficiency promotes experimental colitis and associated
carcinogenesis by regulating intestinal epithelial integrity.
Oncoimmunology. 10:19239102021. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Wang K, Liu Y, Li H, Liang X, Hao M, Yuan
D and Ding L: Claudin-7 is essential for the maintenance of colonic
stem cell homoeostasis via the modulation of Wnt/Notch signalling.
Cell Death Dis. 15:2842024. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Dong S, Li D and Shi D: Skin
barrier-inflammatory pathway is a driver of the psoriasis-atopic
dermatitis transition. Front Med (Lausanne). 11:13355512024.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Kirschner N, Poetzl C, von den Driesch P,
Wladykowski E, Moll I, Behne MJ and Brandner JM: Alteration of
tight junction proteins is an early event in psoriasis: Putative
involvement of proinflammatory cytokines. Am J Pathol.
175:1095–1106. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Aghapour M, Raee P, Moghaddam SJ, Hiemstra
PS and Heijink IH: Airway epithelial barrier dysfunction in chronic
obstructive pulmonary disease: Role of cigarette smoke exposure. Am
J Respir Cell Mol Biol. 58:157–169. 2018. View Article : Google Scholar
|
|
13
|
Lee YG, Lee SH, Hong J, Lee PH and Jang
AS: Titanium dioxide particles modulate epithelial barrier protein,
claudin 7 in asthma. Mol Immunol. 132:209–216. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Nur Husna SM, Siti Sarah CO, Tan HT, Md
Shukri N, Mohd Ashari NS and Wong KK: Reduced occludin and
claudin-7 expression is associated with urban locations and
exposure to second-hand smoke in allergic rhinitis patients. Sci
Rep. 11:12452021. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Angelow S, Ahlstrom R and Yu ASL: Biology
of claudins. Am J Physiol Renal Physiol. 295:F867–F876. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Alexandre MD, Jeansonne BG, Renegar RH,
Tatum R and Chen YH: The first extracellular domain of claudin-7
affects paracellular Cl− permeability. Biochem Biophys Res Commun.
357:87–91. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Bao L, Yang S, Zhao W and Zuo Y: Exploring
claudin proteins: From sequence motifs to their impact on tight
junction-mediated signaling pathways. Amino Acids. 57:482025.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Łukaszewicz-Zając M and Mroczko B:
Claudins-promising biomarkers for selected gastrointestinal (GI)
malignancies? Cancers (Basel). 16:1522023. View Article : Google Scholar
|
|
19
|
Hou J, Renigunta A, Konrad M, Gomes AS,
Schneeberger EE, Paul DL, Waldegger S and Goodenough DA: Claudin-16
and claudin-19 interact and form a cation-selective tight junction
complex. J Clin Invest. 118:619–628. 2008.PubMed/NCBI
|
|
20
|
Suzuki H, Nishizawa T, Tani K, Yamazaki Y,
Tamura A, Ishitani R, Dohmae N, Tsukita S, Nureki O and Fujiyoshi
Y: Crystal structure of a claudin provides insight into the
architecture of tight junctions. Science. 344:304–307. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Itoh M, Furuse M, Morita K, Kubota K,
Saitou M and Tsukita S: Direct binding of three tight
junction-associated MAGUKs, ZO-1, ZO-2, and ZO-3, with the COOH
termini of claudins. J Cell Biol. 147:1351–1363. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Hahn-Strömberg V, Askari S, Ahmad A,
Befekadu R and Nilsson TK: Expression of claudin 1, claudin 4, and
claudin 7 in colorectal cancer and its relation with CLDN DNA
methylation patterns. Tumour Biol. 39:10104283176975692017.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Süren D, Yıldırım M, Kaya V, Alikanoğlu
AS, Bülbüller N, Yıldız M and Sezer C: Loss of tight junction
proteins (claudin 1, 4, and 7) correlates with aggressive behavior
in colorectal carcinoma. Med Sci Monit. 20:1255–1262. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Bernardi MA, Logullo AF, Pasini FS,
Nonogaki S, Blumke C, Soares FA and Brentani MM: Prognostic
significance of CD24 and claudin-7 immunoexpression in ductal
invasive breast cancer. Oncol Rep. 27:28–38. 2012.
|
|
25
|
Dahiya N, Becker KG, Wood WH III, Zhang Y
and Morin PJ: Claudin-7 is frequently overexpressed in ovarian
cancer and promotes invasion. PLoS One. 6:e221192011. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Ding Y, Wang K, Xu C, Hao M, Li H and Ding
L: Intestinal claudin-7 deficiency impacts the intestinal
microbiota in mice with colitis. BMC Gastroenterol. 22:242022.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Pan Y, Tang S, Xu L, Zheng S, Qiao J and
Fang H: Expression and correlation of interleukin-36γ, claudin-1
and claudin-7 in psoriasis. Indian J Dermatol Venereol Leprol.
85:534–536. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Miao YS, Zhao YY, Zhao LN, Wang P, Liu YH,
Ma J and Xue Y: MiR-18a increased the permeability of BTB via RUNX1
mediated down-regulation of ZO-1, occludin and claudin-5. Cell
Signal. 27:156–167. 2015. View Article : Google Scholar
|
|
29
|
Markey GE, Ryan S, Furuta GT,
Menard-Katcher C, McNamee EN and Masterson JC: Hypoxia-inducible
microRNA-155 negatively regulates epithelial barrier in
eosinophilic esophagitis by suppressing tight junction claudin-7.
FASEB J. 38:e233582024. View Article : Google Scholar
|
|
30
|
Zhang B, Lin Y, Bao Q, Zheng Y and Lan L:
MiR-1193 inhibits the malignancy of cervical cancer cells by
targeting claudin 7 (CLDN7). Onco Targets Ther. 13:4349–4358. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Nakayama F, Semba S, Usami Y, Chiba H,
Sawada N and Yokozaki H: Hypermethylation-modulated downregulation
of claudin-7 expression promotes the progression of colorectal
carcinoma. Pathobiology. 75:177–185. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Li Y, Gong Y, Ning X, Peng D, Liu L, He S,
Gong K, Zhang C, Li X and Zhou L: Downregulation of CLDN7 due to
promoter hypermethylation is associated with human clear cell renal
cell carcinoma progression and poor prognosis. J Exp Clin Cancer
Res. 37:2762018. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Usami Y, Chiba H, Nakayama F, Ueda J,
Matsuda Y, Sawada N, Komori T, Ito A and Yokozaki H: Reduced
expression of claudin-7 correlates with invasion and metastasis in
squamous cell carcinoma of the esophagus. Hum Pathol. 37:569–577.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Wang K, Xu C, Li W and Ding L: Emerging
clinical significance of claudin-7 in colorectal cancer: A review.
Cancer Manag Res. 10:3741–3752. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Yuan M, Chen S, Lin Z, Yu R, Chao K, Ye S,
Li Q, Ke H, Zhang C, Huang J, et al: ACSS2-mediated histone H4
lysine 12 crotonylation (H4K12cr) alleviates colitis via enhancing
transcription of CLDN7. Adv Sci (Weinh). 12:e004612025. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Luk IY, Reehorst CM and Mariadason JM:
ELF3, ELF5, EHF and SPDEF transcription factors in tissue
homeostasis and cancer. Molecules. 23:21912018. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Kohno Y, Okamoto T, Ishibe T, Nagayama S,
Shima Y, Nishijo K, Shibata KR, Fukiage K, Otsuka S, Uejima D, et
al: Expression of claudin7 is tightly associated with epithelial
structures in synovial sarcomas and regulated by an Ets family
transcription factor, ELF3. J Biol Chem. 281:38941–38950. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Hirota C, Takashina Y, Ikumi N, Ishizuka
N, Hayashi H, Tabuchi Y, Yoshino Y, Matsunaga T and Ikari A:
Inverse regulation of claudin-2 and -7 expression by p53 and
hepatocyte nuclear factor 4α in colonic MCE301 cells. Tissue
Barriers. 9:18604092021. View Article : Google Scholar
|
|
39
|
Takashina Y, Ishizuka N, Ikumi N, Hayashi
H, Manabe A, Hirota C, Tabuchi Y, Matsunaga T and Ikari A:
Upregulation of claudin-7 expression by angiotensin II in colonic
epithelial cells of mice fed with NaCl-depleted diets. Int J Mol
Sci. 21:14422020. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Tamura T, Yanai H, Savitsky D and
Taniguchi T: The IRF family transcription factors in immunity and
oncogenesis. Annu Rev Immunol. 26:535–584. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Zhao GN, Jiang DS and Li H: Interferon
regulatory factors: At the crossroads of immunity, metabolism, and
disease. Biochim Biophys Acta. 1852:365–378. 2015. View Article : Google Scholar
|
|
42
|
Li X and Yang W: IRF2-induced claudin-7
suppresses cell proliferation, invasion and migration of oral
squamous cell carcinoma. Exp Ther Med. 23:72022. View Article : Google Scholar
|
|
43
|
Nieto MA: The snail superfamily of
zinc-finger transcription factors. Nat Rev Mol Cell Biol.
3:155–166. 2002. View
Article : Google Scholar : PubMed/NCBI
|
|
44
|
Usami Y, Satake S, Nakayama F, Matsumoto
M, Ohnuma K, Komori T, Semba S, Ito A and Yokozaki H:
Snail-associated epithelial-mesenchymal transition promotes
oesophageal squamous cell carcinoma motility and progression. J
Pathol. 215:330–339. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Mittal MK, Myers JN, Bailey CK, Misra S
and Chaudhuri G: Mode of action of the retrogene product SNAI1P, a
SNAIL homolog, in human breast cancer cells. Mol Biol Rep.
37:1221–1227. 2010. View Article : Google Scholar :
|
|
46
|
Li R, Zhang D, Cai C and Dong J: The
clinical significance of claudin-7 and slug expression in lung
squamous cell carcinoma and adenocarcinoma. Zhongguo Fei Ai Za Zhi.
14:492–496. 2011.In Chinese. PubMed/NCBI
|
|
47
|
Ling Y, Kang X, Yi Y, Feng S, Ma G and Qu
H: CLDN5: From structure and regulation to roles in tumors and
other diseases beyond CNS disorders. Pharmacol Res. 200:1070752024.
View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Heiler S, Mu W, Zöller M and Thuma F: The
importance of claudin-7 palmitoylation on membrane subdomain
localization and metastasis-promoting activities. Cell Commun
Signal. 13:292015. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Tatum R, Zhang Y, Lu Q, Kim K, Jeansonne
BG and Chen YH: WNK4 phosphorylates ser(206) of claudin-7 and
promotes paracellular Cl(−) permeability. FEBS Lett. 581:3887–3891.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Huang YY, Wang ZK, Li J, Bai SW, Shen B,
Du J, Xia XM and Wang FY: The effect of serine phosphorylated
claudin-7 on the epithelial barrier and the modulation by transient
receptor potential vanilloid 4 in human colonic cells. Biomed
Pharmacother. 108:540–546. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Huang YY, Li J, Zhang HR, Bai SW, Yang HY,
Shen B, Du J and Xia XM: The effect of transient receptor potential
vanilloid 4 on the intestinal epithelial barrier and human colonic
cells was affected by tyrosine-phosphorylated claudin-7. Biomed
Pharmacother. 122:1096972020. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Ren X, Jiang M, Ding P, Zhang X, Zhou X,
Shen J, Liu D, Yan X and Ma Z: Ubiquitin-specific protease 28: The
decipherment of its dual roles in cancer development. Exp Hematol
Oncol. 12:272023. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Kohn KW, Zeeberg BM, Reinhold WC and
Pommier Y: Gene expression correlations in human cancer cell lines
define molecular interaction networks for epithelial phenotype.
PLoS One. 9:e992692014. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Kido T and Lau YFC: Androgen receptor
variant 7 exacerbates hepatocarcinogenesis in a c-MYC-driven mouse
HCC model. Oncogenesis. 12:42023. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Nguyen N, Fernando SD, Biette KA, Hammer
JA, Capocelli KE, Kitzenberg DA, Glover LE, Colgan SP, Furuta GT
and Masterson JC: TGF-β1 alters esophageal epithelial barrier
function by attenuation of claudin-7 in eosinophilic esophagitis.
Mucosal Immunol. 11:415–426. 2018. View Article : Google Scholar :
|
|
56
|
Ahlswede L, Siebenaller C, Junglas B,
Hellmann N and Schneider D: Human claudin-7 cis-interactions are
not crucial for membrane-membrane (trans-) interactions. Front Mol
Biosci. 9:9083832022. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Xiao Y, Lian H, Zhong XS, Krishnachaitanya
SS, Cong Y, Dashwood RH, Savidge TC, Powell DW, Liu X and Li Q:
Matrix metalloproteinase 7 contributes to intestinal barrier
dysfunction by degrading tight junction protein claudin-7. Front
Immunol. 13:10209022022. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Chen Y: The enzymes in ubiquitin-like
post-translational modifications. Biosci Trends. 1:16–25.
2007.PubMed/NCBI
|
|
59
|
Gehne N, Lamik A, Lehmann M, Haseloff RF,
Andjelkovic AV and Blasig IE: Cross-over endocytosis of claudins is
mediated by interactions via their extracellular loops. PLoS One.
12:e01821062017. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Kim J, Byun I, Kim DY, Joh H, Kim HJ and
Lee MJ: Targeted protein degradation directly engaging lysosomes or
proteasomes. Chem Soc Rev. 53:3253–3272. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Nübel T, Preobraschenski J, Tuncay H,
Weiss T, Kuhn S, Ladwein M, Langbein L and Zöller M: Claudin-7
regulates EpCAM-mediated functions in tumor progression. Mol Cancer
Res. 7:285–299. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Ladwein M, Pape UF, Schmidt DS, Schnölzer
M, Fiedler S, Langbein L, Franke WW, Moldenhauer G and Zöller M:
The cell-cell adhesion molecule EpCAM interacts directly with the
tight junction protein claudin-7. Exp Cell Res. 309:345–357. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Szabo R, Ward JM, Artunc F and Bugge TH:
EPCAM and TROP2 share a role in claudin stabilization and
development of intestinal and extraintestinal epithelia in mice.
Biol Open. 11:bio0594032022. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Lei Z, Maeda T, Tamura A, Nakamura T,
Yamazaki Y, Shiratori H, Yashiro K, Tsukita S and Hamada H: EpCAM
contributes to formation of functional tight junction in the
intestinal epithelium by recruiting claudin proteins. Dev Biol.
371:136–145. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Hagen SJ: Non-canonical functions of
claudin proteins: Beyond the regulation of cell-cell adhesions.
Tissue Barriers. 5:e13278392017. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Ding L, Lu Z, Foreman O, Tatum R, Lu Q,
Renegar R, Cao J and Chen YH: Inflammation and disruption of the
mucosal architecture in claudin-7-deficient mice. Gastroenterology.
142:305–315. 2012. View Article : Google Scholar
|
|
67
|
Xing T, Benderman LJ, Sabu S, Parker J,
Yang J, Lu Q, Ding L and Chen YH: Tight junction protein claudin-7
is essential for intestinal epithelial stem cell self-renewal and
differentiation. Cell Mol Gastroenterol Hepatol. 9:641–659. 2020.
View Article : Google Scholar :
|
|
68
|
Duckworth CA: Identifying key regulators
of the intestinal stem cell niche. Biochem Soc Trans. 49:2163–2176.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Hageman JH, Heinz MC, Kretzschmar K, van
der Vaart J, Clevers H and Snippert HJG: Intestinal regeneration:
Regulation by the microenvironment. Dev Cell. 54:435–446. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Naser AN, Xing T, Tatum R, Lu Q, Boyer PJ
and Chen YH: Colonic crypt stem cell functions are controlled by
tight junction protein claudin-7 through Notch/Hippo signaling. Ann
NY Acad Sci. 1535:92–108. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Li WY, Huey CL and Yu ASL: Expression of
claudin-7 and -8 along the mouse nephron. Am J Physiol Renal
Physiol. 286:F1063–F1071. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Alexandre MD, Lu Q and Chen YH:
Overexpression of claudin-7 decreases the paracellular
Cl-conductance and increases the paracellular Na+ conductance in
LLC-PK1 cells. J Cell Sci. 118:2683–2693. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Tatum R, Zhang Y, Salleng K, Lu Z, Lin JJ,
Lu Q, Jeansonne BG, Ding L and Chen YH: Renal salt wasting and
chronic dehydration in claudin-7-deficient mice. Am J Physiol Renal
Physiol. 298:F24–F34. 2010. View Article : Google Scholar
|
|
74
|
Kahle KT, Wilson FH, Leng Q, Lalioti MD,
O'Connell AD, Dong K, Rapson AK, MacGregor GG, Giebisch G, Hebert
SC and Lifton RP: WNK4 regulates the balance between renal NaCl
reabsorption and K+ secretion. Nat Genet. 35:372–376. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Fan J, Tatum R, Hoggard J and Chen YH:
Claudin-7 modulates Cl− and Na+ homeostasis
and WNK4 expression in renal collecting duct cells. Int J Mol Sci.
20:37982019. View Article : Google Scholar
|
|
76
|
Wang K, Li T, Xu C, Ding Y, Li W and Ding
L: Claudin-7 downregulation induces metastasis and invasion in
colorectal cancer via the promotion of epithelial-mesenchymal
transition. Biochem Biophys Res Commun. 508:797–804. 2019.
View Article : Google Scholar
|
|
77
|
Lu Z, Ding L, Hong H, Hoggard J, Lu Q and
Chen YH: Claudin-7 inhibits human lung cancer cell migration and
invasion through ERK/MAPK signaling pathway. Exp Cell Res.
317:1935–1946. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Singh AB and Dhawan P: Claudins and
cancer: Fall of the soldiers entrusted to protect the gate and keep
the barrier intact. Semin Cell Dev Biol. 42:58–65. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Ji W, Zhuang X, Jiang WG and Martin TA:
Tight junctional protein family, claudins in cancer and cancer
metastasis. Front Oncol. 15:15964602025. View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Tabariès S and Siegel PM: The role of
claudins in cancer metastasis. Oncogene. 36:1176–1190. 2017.
View Article : Google Scholar
|
|
81
|
Ji H, Ding X, Zhang W, Zheng Y, Du H,
Zheng Y, Song H, Li M, Jiang Y, Xie J, et al: Claudin-7 inhibits
proliferation and metastasis in salivary adenoid cystic carcinoma
through Wnt/β-catenin signaling. Cell Transplant.
29:9636897209435832020. View Article : Google Scholar
|
|
82
|
Liang X, Yuan D, Zhao S, Zhou J, Wang K,
Liu X, Liu Y, Li H, Hao M, Huang W, et al: Claudin-7 deficiency
induces metabolic reprogramming of neutrophils in the colorectal
cancer microenvironment. Cell Death Dis. 16:7282025. View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Lu Z, Kim DH, Fan J, Lu Q, Verbanac K,
Ding L, Renegar R and Chen YH: A non-tight junction function of
claudin-7-Interaction with integrin signaling in suppressing lung
cancer cell proliferation and detachment. Mol Cancer. 14:1202015.
View Article : Google Scholar : PubMed/NCBI
|
|
84
|
West JJ, Golloshi R, Cho CY, Wang Y,
Stevenson P, Stein-O'Brien G, Fertig EJ and Ewald AJ: Claudin 7
suppresses invasion and metastasis through repression of a smooth
muscle actin program. J Cell Biol. 223:e2023110022024. View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Zihni C, Mills C, Matter K and Balda MS:
Tight junctions: From simple barriers to multifunctional molecular
gates. Nat Rev Mol Cell Biol. 17:564–580. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Garcia-Hernandez V, Quiros M and Nusrat A:
Intestinal epithelial claudins: Expression and regulation in
homeostasis and inflammation. Ann N Y Acad Sci. 1397:66–79. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Li WJ, Xu C, Wang K, Li TY, Wang XN, Yang
H, Xing T, Li WX, Chen YH, Gao H and Ding L: Severe intestinal
inflammation in the small intestine of mice induced by controllable
deletion of claudin-7. Dig Dis Sci. 63:1200–1209. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Fan X, Qi A, Zhang M, Jia Y, Li S, Han D
and Liu Y: Expression and clinical significance of CLDN7 and its
immune-related cells in breast cancer. Diagn Pathol. 19:1132024.
View Article : Google Scholar :
|
|
89
|
Kim WK, Kwon Y, Jang M, Park M, Kim J, Cho
S, Jang DG, Lee WB, Jung SH, Choi HJ, et al: β-catenin activation
down-regulates cell-cell junction-related genes and induces
epithelial-to-mesenchymal transition in colorectal cancers. Sci
Rep. 9:184402019. View Article : Google Scholar
|
|
90
|
Wang W, Li X, Lee M, Jun S, Aziz KE, Feng
L, Tran MK, Li N, McCrea PD, Park JI and Chen J: FOXKs promote
Wnt/β-catenin signaling by translocating DVL into the nucleus. Dev
Cell. 32:707–718. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Alikanoglu AS, Gunduz S, Demirpence O,
Suren D, Gunduz UR, Sezer C, Yildiz M and Yildirim M: Expression
pattern and prognostic significance of claudin 1, 4 and 7 in
pancreatic cancer. Asian Pac J Cancer Prev. 16:4387–4392. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Jun KH, Kim JH, Jung JH, Choi HJ and Chin
HM: Expression of claudin-7 and loss of claudin-18 correlate with
poor prognosis in gastric cancer. Int J Surg. 12:156–162. 2014.
View Article : Google Scholar
|
|
93
|
Quan JC, Peng J, Guan X, Liu Z, Jiang Z,
Chen HP, Zhuang M, Wang S, Sun P, Wang HY, et al: Evaluation of
clinical significance of claudin 7 and construction of prognostic
grading system for stage II colorectal cancer. World J Clin Cases.
8:2190–2200. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Li XM, Wang H, Zhu LL, Zhao RZ and Ji HL:
Genes regulating epithelial polarity are critical suppressors of
esophageal oncogenesis. J Cancer. 6:694–700. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
95
|
Lioni M, Brafford P, Andl C, Rustgi A,
El-Deiry W, Herlyn M and Smalley KSM: Dysregulation of claudin-7
leads to loss of E-cadherin expression and the increased invasion
of esophageal squamous cell carcinoma cells. Am J Pathol.
170:709–721. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Flores AR, Rêma A, Carvalho F, Lopes G,
Faustino A and Dias Pereira P: Clinicopathological significance of
immunoexpression of claudin-1 and claudin-7 in feline mammary
carcinomas. J Comp Pathol. 151:339–346. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
97
|
Li X, Li Y, Qiu H and Wang Y:
Downregulation of claudin-7 potentiates cellular proliferation and
invasion in endometrial cancer. Oncol Lett. 6:101–105. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
98
|
Hoggard J, Fan J, Lu Z, Lu Q, Sutton L and
Chen YH: Claudin-7 increases chemosensitivity to cisplatin through
the upregulation of caspase pathway in human NCI-H522 lung cancer
cells. Cancer Sci. 104:611–618. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
99
|
Carino A, Graziosi L, Marchianò S,
Biagioli M, Marino E, Sepe V, Zampella A, Distrutti E, Donini A and
Fiorucci S: Analysis of gastric cancer transcriptome allows the
identification of histotype specific molecular signatures with
prognostic potential. Front Oncol. 11:6637712021. View Article : Google Scholar : PubMed/NCBI
|
|
100
|
Kwon MJ: Emerging roles of claudins in
human cancer. Int J Mol Sci. 14:18148–18180. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
101
|
Akimoto T, Takasawa A, Murata M, Kojima Y,
Takasawa K, Nojima M, Aoyama T, Hiratsuka Y, Ono Y, Tanaka S, et
al: Analysis of the expression and localization of tight junction
transmembrane proteins, claudin-1, -4, -7, occludin and JAM-A, in
human cervical adenocarcinoma. Histol Histopathol. 31:921–931.
2016.PubMed/NCBI
|
|
102
|
Thuma F, Heiler S, Schnölzer M and Zöller
M: Palmitoylated claudin7 captured in glycolipid-enriched membrane
microdomains promotes metastasis via associated transmembrane and
cytosolic molecules. Oncotarget. 7:30659–30677. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
103
|
Kim CJ, Lee JW, Choi JJ, Choi HY, Park YA,
Jeon HK, Sung CO, Song SY, Lee YY, Choi CH, et al: High claudin-7
expression is associated with a poor response to platinum-based
chemotherapy in epithelial ovarian carcinoma. Eur J Cancer.
47:918–925. 2011. View Article : Google Scholar
|
|
104
|
Bhat AA, Pope JL, Smith JJ, Ahmad R, Chen
X, Washington MK, Beauchamp RD, Singh AB and Dhawan P: Claudin-7
expression induces mesenchymal to epithelial transformation (MET)
to inhibit colon tumorigenesis. Oncogene. 34:4570–4580. 2015.
View Article : Google Scholar
|
|
105
|
Quansah E, Gardey E, Ramoji A,
Meyer-Zedler T, Goehrig B, Heutelbeck A, Hoeppener S, Schmitt M,
Waldner M, Stallmach A and Popp J: Intestinal epithelial barrier
integrity investigated by label-free techniques in ulcerative
colitis patients. Sci Rep. 13:26812023. View Article : Google Scholar : PubMed/NCBI
|
|
106
|
Büning C, Geissler N, Prager M, Sturm A,
Baumgart DC, Büttner J, Bühner S, Haas V and Lochs H: Increased
small intestinal permeability in ulcerative colitis: Rather genetic
than environmental and a risk factor for extensive disease? Inflamm
Bowel Dis. 18:1932–1939. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
107
|
Tanaka H, Takechi M, Kiyonari H, Shioi G,
Tamura A and Tsukita S: Intestinal deletion of claudin-7 enhances
paracellular organic solute flux and initiates colonic inflammation
in mice. Gut. 64:1529–1538. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
108
|
Li H, Wang K, Hao M, Liu Y, Liang X, Yuan
D and Ding L: Intestinal epithelial Cldn-7 regulates intestinal
inflammation by altering the gut microbiota. Pathol Res Pract.
260:1554482024. View Article : Google Scholar : PubMed/NCBI
|
|
109
|
Wen X, Zhao H, Wang L, Wang L, Du G, Guan
W, Liu J, Cao X, Jiang X, Tian J, et al: Nobiletin attenuates
DSS-induced intestinal barrier damage through the HNF4α-claudin-7
signaling pathway. J Agric Food Chem. 68:4641–4649. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
110
|
Goulet O, Pigneur B and Charbit-Henrion F:
Congenital enteropathies involving defects in enterocyte structure
or differentiation. Best Pract Res Clin Gastroenterol.
56-57:1017842022. View Article : Google Scholar : PubMed/NCBI
|
|
111
|
Wu CJ, Feng X, Lu M, Morimura S and Udey
MC: Matriptase-mediated cleavage of EpCAM destabilizes claudins and
dysregulates intestinal epithelial homeostasis. J Clin Invest.
127:623–634. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
112
|
Mueller JL, McGeough MD, Peña CA and
Sivagnanam M: Functional consequences of EpCam mutation in mice and
men. Am J Physiol Gastrointest Liver Physiol. 306:G278–G288. 2014.
View Article : Google Scholar
|
|
113
|
Zhou X, Chen Y, Cui L, Shi Y and Guo C:
Advances in the pathogenesis of psoriasis: From keratinocyte
perspective. Cell Death Dis. 13:812022. View Article : Google Scholar : PubMed/NCBI
|
|
114
|
Pawankar R: Allergic diseases and asthma:
A global public health concern and a call to action. World Allergy
Organ J. 7:122014. View Article : Google Scholar : PubMed/NCBI
|
|
115
|
Vareille M, Kieninger E, Edwards MR and
Regamey N: The airway epithelium: Soldier in the fight against
respiratory viruses. Clin Microbiol Rev. 24:210–229. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
116
|
Steelant B, Seys SF, Van Gerven L, Van
Woensel M, Farré R, Wawrzyniak P, Krohn IK, Bullens DM, Talavera K,
Raap U, et al: Histamine and T helper cytokine-driven epithelial
barrier dysfunction in allergic rhinitis. J Allergy Clin Immunol.
141:951–963.e8. 2018. View Article : Google Scholar
|
|
117
|
Shaikh SB, Gouda MM, Khandhal I, Rahman T,
Shetty A and Bhandary YP: Alterations in mRNA expression levels of
tight junction proteins in the blood cells of smokers with or
without COPD. Endocr Metab Immune Disord Drug Targets. 23:389–395.
2023. View Article : Google Scholar
|
|
118
|
Inoue H, Akimoto K, Homma T, Tanaka A and
Sagara H: Airway epithelial dysfunction in asthma: Relevant to
epidermal growth factor receptors and airway epithelial cells. J
Clin Med. 9:36982020. View Article : Google Scholar : PubMed/NCBI
|
|
119
|
Zhou J, Zhou XD, Xu R, Du XZ, Li Q, Li B,
Zhang GY, Chen LX, Perelman JM and Kolosov VP: The degradation of
airway epithelial tight junctions in asthma under high airway
pressure is probably mediated by Piezo-1. Front Physiol.
12:6377902021. View Article : Google Scholar : PubMed/NCBI
|
|
120
|
Lee PH, Park S, Lee YG, Choi SM, An MH and
Jang AS: The impact of environmental pollutants on barrier
dysfunction in respiratory disease. Allergy Asthma Immunol Res.
13:850–862. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
121
|
Lee PH, Kim BG, Park MK, Hong J, Lee YG
and Jang AS: The impact of diesel exhaust particles on tight
junctional proteins on nose and lung in a mouse model. Allergy
Asthma Immunol Res. 13:350–352. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
122
|
Hahn S, Kim G, Jin SM and Kim JH:
Protective effects of metformin in the pro-inflammatory cytokine
induced intestinal organoids injury model. Biochem Biophys Res
Commun. 690:1492912024. View Article : Google Scholar
|
|
123
|
Kong C, Zhou S, Xu C, Yang J, Chen J, Li
C, Yuan Y, Peng Z, Chang G, Yang Y and Lei Z: Polysaccharides of
Atractylodes macrocephala Koidz effectively improve the
EpCAM+/− genotype-related inflammatory bowel diseases
via modulating gut microbiota. Int J Biol Macromol. 333:1488552025.
View Article : Google Scholar
|
|
124
|
Nikaido M, Otani T, Kitagawa N, Ogata K,
Iida H, Anan H and Inai T: Anisomycin, a JNK and p38 activator,
suppresses cell-cell junction formation in 2D cultures of K38 mouse
keratinocyte cells and reduces claudin-7 expression, with an
increase of paracellular permeability in 3D cultures. Histochem
Cell Biol. 151:369–384. 2019. View Article : Google Scholar
|
|
125
|
Stio M, Retico L, Annese V and Bonanomi
AG: Vitamin D regulates the tight-junction protein expression in
active ulcerative colitis. Scand J Gastroenterol. 51:1193–1199.
2016. View Article : Google Scholar : PubMed/NCBI
|