Introduction
Gastric cancer (GC) is the fifth most frequently
diagnosed malignancy worldwide and represents the fourth leading
cause of cancer-related mortality (1,2).
Precancerous lesions of GC refer to pathological alterations of the
gastric mucosa that exhibit a high propensity for malignant
transformation, involving multiple risk factors and multistep
cellular changes (3). In-depth
investigation of these precancerous lesions is essential for
identifying key mechanisms underlying gastric carcinogenesis and
provides critical evidence for early prevention and effective
therapeutic intervention. Although advances in early screening and
diagnostic technologies have been achieved in recent years, most
patients with GC are still diagnosed at advanced stages of the
disease (4,5). Numerous studies have shown that
dietary factors play a significant role in the development of
gastrointestinal malignancies; however, reliable early diagnostic
biomarkers remain poorly defined (6,7).
N-methyl-N'-nitro-N-nitrosoguanidine (MNNG), composed primarily of
N-nitrosamines and N-nitrosamides, is a widely distributed
environmental chemical mutagen and carcinogen (8,9).
MNNG can form adducts with biological macromolecules, including DNA
and proteins (10). It has been
demonstrated that MNNG can induce precancerous lesions, tumor
formation and malignant cellular transformation in animal models.
For instance, MNNG exposure can lead to chronic atrophic gastritis,
esophageal cancer and malignant transformation of normal intestinal
epithelial cells in rats (11,12). Consequently, MNNG is commonly
employed as an experimental agent to model nitroso compound-induced
precancerous transformation of gastric mucosal cells (13,14). Previous studies have indicated
that MNNG exposure activates multiple oncogenic signaling pathways,
including NF-κB (15),
Wnt/β-catenin (16) and
PI3K/Akt/mTOR (14). However,
research focusing on the underlying molecular mechanisms remains
limited.
Circular RNAs (circRNAs) constitute a class of
non-coding RNA molecules characterized by covalently closed loop
structures, which confer greater stability compared with linear
RNAs. Initially, circRNAs were considered byproducts of aberrant
splicing events during transcription (17,18). However, accumulating evidence has
shown that circRNAs are predominantly generated through
back-splicing, a process in which exons from precursor mRNAs are
ligated to form circular transcripts (19). CircRNAs are widely expressed and
exhibit high abundance, remarkable stability and strong tissue- and
disease-specific expression patterns (20). An increasing number of studies
have revealed that circRNAs play diverse regulatory roles in both
physiological and pathological processes, including cancer
initiation and progression. For example, Lin et al (21) identified a novel circRNA,
circTFRC, which is markedly upregulated in GC tissues and cell
lines. Elevated plasma levels of circTFRC were closely associated
with tumor size and metastasis and mechanistically, circTFRC
promotes GC malignancy by impairing ferroptosis susceptibility. In
another study, circ_0005927 was shown to suppress proliferation,
colony formation, invasion and epithelial-mesenchymal transition
(EMT) in GC cells by sponging miR-570-3p and upregulating forkhead
box O3 (FOXO3), suggesting that the circ_0005927/miR-570-3p/FOXO3
axis represents a potential therapeutic target for GC (22). Collectively, these findings
highlight the critical roles of circRNAs in GC. Nevertheless, a
large number of circRNAs remain uncharacterized and their
biological functions and underlying mechanisms in GC require
further investigation.
The tumor microenvironment (TME) is a highly complex
and dynamic system formed through interactions among cancer cells,
stromal cells, immune cells and other surrounding components
(23). Exosomes are a subtype of
extracellular vesicles with diameters ranging from 30-200 nm. They
are released by nearly all cell types and have emerged as promising
biomarkers for the detection and diagnosis of various diseases
(24,25). Within the TME, exosomes function
as critical mediators of intercellular communication, acting as
'membrane messengers' that transmit biological signals between
cells (26). However, the roles
and regulatory mechanisms of exosome-derived circRNAs in malignant
cellular transformation remain largely unexplored.
The current study investigated the effects of
long-term MNNG exposure and MNNG-induced exosomes on the migratory,
proliferative and invasive capacities of normal GES-1 cells. By
analyzing data from the Gene Expression Omnibus (GEO) database
(https://www.ncbi.nlm.nih.gov/geo/;
accessed in May 2023), has_circ-0000549 (circ0000549, circSRSF5)
was identified as a candidate circRNA with markedly elevated
expression in GC tissues and plasma samples. Subsequent validation
confirmed that circ0000549 expression was markedly increased in
GES-1 cells following prolonged MNNG exposure. Moreover,
circ0000549 packaged within exosomes released from MNNG-induced
malignantly transformed cells could be transferred to neighboring
normal gastric epithelial cells. Functionally, exosomal circ0000549
enhanced invasion, migration and proliferation, while promoting
stemness and EMT in normal GES-1 cells through the
miR-15b-5p/KIF1B/PI3K/AKT signaling axis. These findings provide
novel insights into the mechanisms underlying MNNG-induced gastric
carcinogenesis.
Materials and methods
Cell culture
The human gastric epithelial cell line GES-1 was
obtained from Shanghai Enzyme Linked Biotechnology Co., Ltd. Cells
were cultured in Dulbecco's Modified Eagle's Medium (DMEM; Dalian
Meilun Biology Technology Co., Ltd.) enriched with 10% fetal bovine
serum (FBS; Gibco; Thermo Fisher Scientific, Inc.) and maintained
at 37°C in a humidified incubator containing 5% CO2. To
establish a malignant transformation model, GES-1 cells were
chronically exposed to MNNG. Based on preliminary cell survival
assays, MNNG concentrations of 0.5 and 1 μmol/l were
selected for subsequent experiments. When cells reached 80-90%
confluence, they were washed twice with phosphate-buffered saline
(PBS), digested with trypsin and centrifuged at 35 × g at room
temperature for 5 min. Cells were then passaged at a ratio of 1:3
and cultured in 8 ml DMEM containing 10% FBS under standard
conditions (37°C, 5% CO2), with the culture medium
replaced daily. On the day following cell passage, a 0.01 mol/l
MNNG stock solution was diluted with fresh DMEM to achieve final
MNNG concentrations of 0, 0.5 and 1 μmol/l. Cells were
continuously treated for ~30 passages to induce malignant
transformation and aliquots of cells were cryopreserved at passages
5, 10, 25 and 30 for subsequent analyses. The malignantly
transformed GES-1 cells were designated as TGES-1.
Bioinformatics analysis of differentially
expressed circRNAs
Bioinformatic screening was performed using two
publicly available GC-related circRNA datasets (GSE89143 and
GSE93541) from the GEO database. Differential expression analysis
was conducted using the GEO2R online tool (https://www.ncbi.nlm.nih.gov/geo/geo2r/), which is
based on the limma R package (Version 3.54.0; https://bioconductor.org/packages/limma/) for data
normalization and statistical evaluation. CircRNAs with an adjusted
P<0.05 and a log2 (fold change) >1 were defined as
markedly differentially expressed. From each dataset, the top 249
circRNAs meeting these criteria were selected for subsequent
intersection analysis. A Venn diagram was generated using the jvenn
online tool (https://jvenn.toulouse.inrae.fr/app/index.html) to
identify circRNAs shared between the two datasets. Expression data
corresponding to the overlapping circRNAs were then extracted from
both datasets and converted into circBase ID format for
standardization. Finally, a heatmap visualizing the expression
patterns of these shared circRNAs across different samples was
constructed using TBtools software (version 2.025, https://github.com/CJ-Chen/TBtools/releases). Based on
these analyses, the candidate differentially expressed circRNAs
were further validated and selected for downstream experimental
investigation.
CircInteractome-based prediction of
circ0000549-AGO2 binding interaction
The potential binding of circ0000549 to Argonaute 2
(AGO2) was predicted using the online bioinformatics tool
CircInteractome (https://circinteractome.nia.nih.gov/). The full
nucleotide sequence of circ0000549 was input into the analysis
module of the database, with the default algorithm parameters
retained for the AGO2 binding site prediction. The prediction was
performed based on data integrated in the database, and the results
were filtered to identify the reliable binding motifs of
circ0000549 to AGO2, so as to confirm the potential interaction
between circ0000549 and AGO2.
Target gene prediction for
miR-15b-5p
Potential target genes of miR-15b-5p were predicted
using five independent bioinformatics algorithms: miRDB (version
1.0; https://mirdb.org/), DIANA microT-CDS (version
5.0; https://www.microrna.gr/microt_webserver/), miRWalk
(version 2.0; http://mirwalk.umm.uni-heidelberg.de/), StarBase
(version 3.0, StarBase or ENCORI: Decoding the Encyclopedia of RNA
Interactomes, https://rnasysu.com/encori/) and TargetScan (version
8.0; https://www.targetscan.org/vert_80/). All predictions
were performed using the default parameters and score thresholds
provided by each database. All genes predicted by each tool were
collected without applying additional filters. The predicted gene
sets were then intersected and genes identified by at least three
of the five algorithms were defined as high-confidence target
candidates. Based on this intersection analysis, six candidate
target genes were screened and further subjected to multiple
validation assays to determine the bona fide target molecule.
Definition of high nitrite compound
exposure
Patients were categorized into the 'high nitrite
exposure' group based on a combined evaluation of dietary history
and relevant biomarkers.
Dietary Assessment: Patients were required to have a
confirmed history of long-term and regular consumption of foods
rich in N-nitroso compounds or their precursors. Dietary intake was
determined via a structured questionnaire, focusing on the
consumption of preserved foods, processed meats and high-salt
diets.
Biomarker Assessment: Serum levels of
nitrosation-related metabolites, including N-nitrosoproline, were
quantified using liquid chromatography-tandem mass spectrometry
(LC-MS/MS). Levels exceeding established reference ranges were
defined as indicative of high nitrite exposure.
Liquid chromatography-tandem mass
spectrometry (LC-MS/MS) quantification of serum
N-nitrosoproline
Serum levels of N-nitrosoproline were determined by
LC-MS/MS. Aliquots of serum underwent protein precipitation with
cold acetonitrile containing a stable isotope-labeled internal
standard. The processed samples were analyzed using an Acquity UPLC
system (Waters Corporation) equipped with an HSS T3 column. A
gradient elution was applied with mobile phases consisting of 0.1%
formic acid in water and acetonitrile. Detection was performed on a
triple quadrupole mass spectrometer operating in positive
electrospray ionization mode, with quantification achieved via
multiple reaction monitoring (MRM). The MRM transitions monitored
were m/z 157.1 to m/z 74.1 for N-nitrosoproline and m/z 158.1 to
m/z 75.1 for the internal standard. The mass spectrometer
parameters were set as follows: nitrogen gas temperature 500°C,
nebuliser pressure 7.0 psi, and desolvation gas flow rate 800 l/h.
A threshold corresponding to the 95th percentile of levels measured
in a reference cohort with low dietary nitrite intake defined the
high-exposure group.
Patients and clinical specimens
Patients with GC (n=40) with a documented history of
high nitrite exposure were randomly recruited from the Affiliated
Hospital of Jiangsu University. All patients were definitively
diagnosed by histopathological examination prior to surgical
intervention and had not received any anticancer treatment before
sample collection. Written informed consent was obtained from each
participant and the study protocol was approved by the Medical
Ethics Committee of Jiangsu University (ethics approval no.
2023182). Peripheral blood samples were collected from both
patients with GC and corresponding normal controls. Plasma was
separated from whole blood by centrifugation at 2,000 × g for 10
min at 4°C and stored for subsequent analyses. The inclusion
criteria for patients were: Being newly diagnosed and
histopathologically confirmed with primary gastric adenocarcinoma;
age between 25 and 65 years; having no history of other
malignancies; receiving no prior chemotherapy, radiotherapy, or
targeted therapy before sample collection; availability of complete
clinical and follow-up data; and for the 'high nitrite exposure'
subgroup, meeting the predefined criteria of combined dietary
history assessment and serum N-nitrosoproline level as described in
the 'Definition of high nitrite compound exposure' section.
Exclusion criteria comprised: Patients with recurrent or metastatic
gastric cancer at initial diagnosis; patients with severe
concurrent systemic infections, autoimmune diseases, or other major
organ dysfunction; pregnant or lactating women; and those with
incomplete dietary history or insufficient serum sample for
biomarker analysis. The demographic characteristics of the patients
were shown in Table SI. All
patients provided written informed consent for the use of their
samples in scientific research. Samples were collected between
March 2023 and March 2024 in accordance with the approved
protocol.
Exosome experiments
Cells were cultured in 10-cm culture dishes until
~50% confluence. The culture medium was then replaced with fresh
medium enriched with 10% exosome-depleted FBS following two washes
with PBS. After 48 h of incubation at 37°C, the conditioned medium
was collected and subjected to sequential centrifugation as
previously described (27,28) for exosome isolation. Briefly, the
conditioned medium was centrifuged at 2,000 × g at 4°C for 10 min
to remove intact cells, followed by centrifugation at 10,000 × g at
4°C for 30 min to clear cell debris and then ultracentrifuged at
100,000 × g at 4°C for 2 h (repeated once). Freshly isolated
exosomes were resuspended in PBS, filtered through a 0.2-μm
membrane and stored at −80°C until further use. Exosomes were
characterized and identified using a combination of western blot
analysis, nanoparticle tracking analysis (NTA) and transmission
electron microscopy (TEM). For TEM analysis (conducted by Zhenjiang
Danyang Zhuanbo Testing Technology Co., Ltd.), exosomes were fixed
with glutaraldehyde, dropped onto copper grids for 3 min adsorption
(residual liquid blotted with filter paper), negatively stained
with phosphotungstic acid (pH=6.8) at room temperature for 1 min,
dried under incandescent light, and then observed and imaged. For
NTA, the NanoSight instrument (Waters Corporation) was preheated
for ~30 min after power-on, cleaned with purified water, and
exosomes were diluted to an appropriate concentration with purified
water, mixed thoroughly, injected via a 1 ml syringe, and analyzed
with relevant parameters recorded. In addition, plasma-derived
exosomes from human samples were isolated using the ExoQuick Plasma
Prep and Exosome Precipitation Kit (System Biosciences, LLC).
DiL-labeled exosome uptake assay
Coverslips were attached to the bottom of 12-well
plates with medium. GES-1 cells in logarithmic phase were
trypsinized and seeded onto coverslips at 6×103
cells/well, then incubated at 37°C with 5% CO2. Exosomes
were labeled with DiL dye (MilliporeSigma; 1:1,000 dilution) for 30
min at 37°C in the dark, centrifuged at 3,000 × g for 15 min, and
filtered through a 0.22 μm filter (MilliporeSigma). Labeled
exosomes were added to adherent cells and cultured for 24 h at 37°C
with 5% CO2. Cells were rinsed twice with PBS, fixed
with 4% paraformaldehyde for 30 min at room temperature, and washed
three times with PBS at room temperature. Nuclei were stained with
DAPI (MilliporeSigma) for 20 min at room temperature, followed by
three PBS washes (5 min each). Coverslips were mounted with
glycerol and sealed, then imaged using a confocal laser scanning
microscope.
RNA extraction and reverse
transcription-quantitative (RT-q) PCR
Total RNA was extracted from 2×107 GES-1
and TGES-1 cells using RNA isolator Total RNA Extraction Reagent
(TRIzol® method; Vazyme Biotech Co., Ltd.) according to
the manufacturer's instructions. Complementary DNA (cDNA) was
synthesized using the HiScript QRT SuperMix for qPCR kit (Vazyme
Biotech Co., Ltd.) following the manufacturer's standard protocol.
RT-qPCR was subsequently performed on an Applied Biosystems
QuantStudio 3 Real-Time PCR System (Thermo Fisher Scientific, Inc.)
using SYBR Green PCR Master Mix (Vazyme Biotech Co., Ltd.)
according to the manufacturer's recommendations. The PCR cycling
conditions were as follows: pre-denaturation at 95°C for 5 min,
followed by 40 cycles of denaturation at 95°C for 20 sec, annealing
at 60°C for 30 sec and extension at 72°C for 20 sec. Relative gene
expression levels were calculated using the 2−ΔΔCq
method as previously described (29), with GAPDH used as the normalizer.
All experiments were independently replicated three times. The
primers used in RT-qPCR were shown in Table SII.
Cell counting kit-8 (CCK8) assay
Logarithmically growing cells were seeded into
96-well plates at a density of 1×103 cells per well.
Cell viability was assessed at 24, 48, 72 and 96 h after seeding.
At each time point, 10 μl of CCK-8 reagent was added to each
well, followed by incubation for an additional 3 h at 37°C.
Absorbance was measured at 450 nm using a microplate reader. All
experiments were performed in triplicate.
Colony formation assay
Logarithmically growing cells were seeded into
3.5-cm culture dishes at a density of 1×103 cells per
dish. The cells were cultured for 7-14 days until visible colonies
formed. The culture medium was then discarded and cells were gently
washed twice with PBS, fixed with 4% paraformaldehyde at room
temperature for 30 min and stained with crystal violet at room
temperature for 15 min. After washing, colonies were counted and
images captured.
Transwell assay
Logarithmically growing cells were harvested,
resuspended in serum-free DMEM and seeded into the upper chambers
of 24-well Transwell inserts at a density of 3×104 cells
per well. For invasion assays, the inserts were pre-coated with 50
μl of Matrigel diluted 1:4 in serum-free DMEM and incubated
at 37°C with 5% CO2 for 1 h to allow solidification,
whereas uncoated inserts were used for migration assays. Complete
culture medium containing 10% FBS was added to the lower chambers.
After 24 h of incubation at 37°C, non-migrated or non-invaded cells
on the upper surface of the membrane were removed. Cells on the
lower surface were fixed with paraformaldehyde at room temperature
for 15 min and stained with crystal violet at room temperature for
10 min. Migrated or invaded cells were counted under a light
microscope at 10× magnification.
Western blotting
Total protein was extracted from cells using RIPA
lysis buffer (Thermo Fisher Scientific, Inc.). Protein
concentration was determined using a BCA protein assay kit (Vazyme
Biotech Co., Ltd.). Equal amounts of protein (50-60 μg) were
loaded per lane separated by 10% sodium dodecyl
sulfate-polyacrylamide gel electrophoresis and transferred onto
polyvinylidene fluoride (PVDF) membranes. The membranes were
blocked with 5% skimmed milk for 1 h at room temperature and
incubated overnight at 4°C with primary antibodies against
N-cadherin (Cell Signaling Technology, Inc; cat. no. 13116S;
dilution: 1:1,000), E-cadherin (Cell Signaling Technology, Inc;
cat. no. 3195S; dilution: 1:1,000), PCNA (Cell Signaling
Technology, Inc.; cat. no. 13110S; dilution: 1:1,000), Nanog (Cell
Signaling Technology, Inc.; cat. no. 4903S; dilution: 1:1,000),
OCT4 (Cell Signaling Technology, Inc; cat. no. 2750; dilution:
1:1,000), Lin28 (Cell Signaling Technology, Inc.; cat. no. 8706S;
dilution: 1:1,000), SOX2 (Cell Signaling Technology, Inc; cat. no.
23064S; dilution: 1:1,000), Vimentin (Cell Signaling Technology,
Inc.; cat. no. 5741S; dilution: 1:1,000), HSP70 (Cell Signaling
Technology, Inc.; cat. no. 4872S; dilution: 1:1,000), Alix (Cell
Signaling Technology, Inc.; cat. no. 92880S; dilution: 1:1,000),
TSG101 (Proteintech, Inc; cat. no. 28283-1-AP; dilution: 1:1,000),
Calnexin (Cell Signaling Technology, Inc.; cat. no. 2679S;
dilution: 1:1,000), KIF1B (Abcam; cat. no. ab72108;
dilution:1:1,500) and GAPDH (Bioworld Technology, Inc.; cat. no.
BS65483M; dilution: 1:2,000). After washing with TBST containing
0.1% Tween-20, membranes were incubated with appropriate
horseradish peroxidase-conjugated secondary antibodies (Signalway
Antibody LLC; cat. no. L3012; dilution: 1:5,000). Protein bands
were visualized using an enhanced chemiluminescence (ECL) detection
system (Vazyme Biotech Co., Ltd.) and band intensities were
quantified using ImageJ software (version 1.53k, National
Institutes of Health).
In vivo xenograft experiments
Male nude mice (4-6 weeks old; weighing 18-22 g; SPF
grade; n=3 per group; total n=24) were purchased from Shanghai SLAC
Laboratory Animal Co., Ltd. All mice were housed under specific
pathogen-free conditions at 22-25°C and 50-60% humidity with a 12 h
light/dark cycle and randomly assigned to experimental groups.
TGES-1 cells, GES-1 cells, TGES-1 cells treated with
si-circ0000549-derived exosomes, or TGES-1 cells treated with
exosomes derived from circ0000549-overexpressing cells combined
with miR-15b-5p mimics were harvested by trypsinization and
resuspended in PBS at a concentration of 5×107 cells in
100 μl. The cell suspensions were then subcutaneously
inoculated into the right flank of each mouse. Mice were observed
daily for general health status and tumor growth was monitored
weekly by caliper measurements. The following humane endpoints were
predefined: Tumor volume >2,000 mm3, ulceration or
necrosis of the tumor, significant weight loss (>20% of body
weight), or signs of severe distress (such as lethargy, hunched
posture, impaired mobility). No animals reached these endpoints or
succumbed prior to the scheduled endpoint.
The experiment lasted for 4 weeks post-inoculation.
All animals were housed under standard conditions (12-h light/dark
cycle; controlled temperature and humidity) with ad libitum
access to food and water. To minimize distress, handling was
performed gently and cage enrichment was provided. No analgesics or
anesthetics were required during the tumor monitoring phase.
At the experimental endpoint, mice were sacrificed
by cervical dislocation under deep anesthesia induced by
intraperitoneal injection of sodium pentobarbital (80 mg/kg body
weight). Mortality was confirmed by the absence of a heartbeat,
cessation of breathing and loss of corneal reflex. Tumors were then
excised, images captured and stored at −80°C for subsequent
analysis.
Plasmid transfection
To inhibit the expression of circ0000549 and KIF1B,
specific small interfering RNAs (siRNAs) were synthesized and
transfected into cells, while a non-targeting siRNA served as the
negative control. miR-15b-5p expression was either enhanced or
suppressed using miR-15b-5p mimics or inhibitors, respectively. The
sequences of all siRNAs are in Table SIII. The siRNAs targeting
circ0000549 and KIF1B were designed and synthesized by Shanghai
GenePharma Co., Ltd. These siRNAs were designed based on the
sequences of the target transcripts retrieved from the circBase
(Version 0.1; https://www.circbase.org/) and NCBI GenBank database
(Version Release 263.0; https://www.ncbi.nlm.nih.gov/genbank/) (circ0000549,
accession number: NM_006925; KIF1B, accession number: accession
numbers: NM_001365952.1).
Cells in the logarithmic growth phase were digested
with trypsin, centrifuged at 35 × g for 5 min at room temperature,
resuspended in complete medium, and seeded into 6-well plates for
overnight incubation. After cell adhesion, the wells were washed
twice with PBS, and 1.5 ml of Opti-MEM serum-free medium (Gibco;
Thermo Fisher Scientific, Inc.) was added before the plates were
returned to a 37°C incubator. For transfection, 5 μl of
Lipofectamine 2000 (Invitrogen; Thermo Fisher Scientific, Inc.) was
mixed with 245 μl of Opti-MEM in the dark. In parallel, 5
μl of siRNA (final concentration: 50 nM) was diluted in 245
μl of Opti-MEM and incubated at room temperature for 5 min.
The two solutions were then combined to form a 500 μl
mixture, which was added to the cells and mixed gently. After
incubation at 37°C for 6 h, the medium was replaced with complete
medium and cells were cultured for another 24-36 h before further
experiments.
Generation of circ0000549
knockdown/overexpression stable cell lines
Cells in the logarithmic growth phase were
trypsinized and seeded into 6-well plates (3×104
cells/well). After overnight culture, the medium was replaced with
2 ml of fresh medium containing 2 μg/ml polybrene and the
appropriate volume of lentivirus, based on the predetermined
multiplicity of infection (MOI). Following 24-h incubation, the
lentivirus-containing medium was aspirated and replaced with
complete medium. The optimal MOI was determined by assessing
fluorescence intensity under a microscope and validating infection
efficiency via RT-qPCR. The lentivirus used was a third-generation
lentiviral expression system. For lentivirus production conducted
by HANBIO Biotechnology Co., Ltd., 293T cells (ATCC) were
transfected with lentiviral vector, packaging plasmid, and envelope
plasmid at a mass ratio of 5:3:2 (total plasmid amount per 10 cm
dish: 20 μg). Viral supernatants were harvested
post-transfection, filtered and concentrated before supply. The
optimal MOI for circ0000549 overexpression was 100. The
lentiviruses for overexpression were designated as OE-NC and
OE-circ549, while those for knockdown were designated as SH-NC and
SH-circ549.
Subsequently, cells were infected at the optimal MOI
and selected with 20 μg/ml puromycin. Stable cell pools were
obtained after selection, indicated by fluorescence in >95% of
adherent cells with minimal cell death. Successful circ0000549
knockdown or overexpression was confirmed by RT-qPCR.
Agarose gel electrophoresis
A 3% agarose gel was prepared by dissolving 1.8 g
agarose powder (Invitrogen; Thermo Fisher Scientific, Inc.) in 60
ml of 0.5XTAE buffer (prepared from 50XTAE buffer; MilliporeSigma),
followed by microwave heating. After solidification, the gel was
placed in an electrophoresis tank and submerged with 0.5XTAE
buffer. PCR products (1 μl) were mixed with loading buffer
at a ratio of 5:1 and loaded into wells. Electrophoresis was
carried out at 110 V until the marker migrated two-thirds of the
gel length. The gel was stained with SYBR Safe (Invitrogen; Thermo
Fisher Scientific, Inc.) and visualized using a gel imaging system
(Bio-Rad Laboratories, Inc.).
RNA fluorescence in situ hybridization
(FISH)
A Cy3-labeled probe specifically targeting
circ0000549 was synthesized by Shanghai GenePharma Co., Ltd. FISH
was performed using a commercially available FISH kit (Shanghai
GenePharma Co., Ltd.). Fluorescence signals were visualized and
captured using a fluorescence microscope.
Dual-luciferase reporter assay
Plasmids containing wild-type (WT) or mutant (MUT)
sequences of circ0000549, as well as miRNA mimics and corresponding
control constructs, were synthesized by Shanghai GenePharma Co.,
Ltd. Cells were co-transfected with the aforementioned plasmids
using Lipofectamine® 2000 Transfection Reagent
(Invitrogen; Thermo Fisher Scientific, Inc.) according to the
manufacturer's instructions. After 48 h of transfection, luciferase
activity was measured using the Dual-Luciferase Reporter Assay
System (Vazyme Biotech Co., Ltd.) strictly following the kit's
protocol.
GeneCards database analysis
To predict potential signaling pathways associated
with KIF1B, the GeneCards database (https://www.genecards.org/) was queried using 'KIF1B'
as the search term. All signaling pathways listed in the 'Pathways'
section of the KIF1B gene profile were retrieved. Pathways
previously reported to be associated with cancer-related biological
processes, including cell proliferation, apoptosis, migration and
differentiation, were prioritized for further analysis. Based on
their functional relevance to the cellular phenotypes examined in
the present study, as well as their well-established involvement in
tumor biology, the PI3K/AKT signaling pathway was selected for
subsequent investigation. This selection was further supported by
experimental validation performed in the present study.
In addition, other pathways predicted to be
associated with KIF1B included Golgi-to-ER retrograde transport,
vesicle-mediated transport, cytoskeletal signaling and response to
elevated platelet cytosolic Ca2+ levels.
Immunohistochemistry
Formalin-fixed, paraffin-embedded tumor tissues
obtained from nude mice were subjected to immunohistochemical
staining. Tissues were fixed in 4% neutral buffered formalin
(Sangon Biotech Co., Ltd.) for 24 h at room temperature, dehydrated
through a graded ethanol series (70, 80, 90, 95, and 100% ethanol,
15 min each), cleared in xylene (twice for 15 min each), and
embedded in paraffin wax. Sections (4 μm) were first
deparaffinized and rehydrated using standard procedures. Antigen
retrieval was performed by heating the sections in 0.01 M citrate
buffer (pH 6.0) for 2 min. Endogenous peroxidase activity was
blocked by incubation with 0.3% H2O2 for 10
min. Non-specific binding was then blocked with bovine serum
albumin (BSA; MedChemExpress) for 30 min. Subsequently, the
sections were incubated overnight at 4°C with primary antibodies
against Vimentin (Cell Signaling Technology, Inc; cat. no. 5741S;
dilution:1:500) and Nanog (Cell Signaling Technology, Inc; cat. no.
4903S; dilution:1:500). After incubation with a biotinylated
secondary antibody (Thermo Fisher Scientific, Inc.; cat. no. 31822;
dilution: 1:1,000; conjugate: biotin), immunoreactivity was
visualized using SABC detection system (Wuhan Boster Biological
Technology, Ltd.) for 30 min at room temperature, followed by
counterstaining with hematoxylin for 2 min at room temperature.
Images were captured with a light microscope (Olympus BX53; Olympus
Corporation) at 400× magnification.
Statistical analysis
All statistical analyses were performed with
GraphPad Prism version 8.0 (Dotmatics). Data were analyzed
utilizing either Student's t-test or one-way analysis of variance
(ANOVA) followed by Bonferroni's post hoc test, as appropriate.
P<0.05 was considered to indicate a statistically significant
difference.
Results
MNNG induces malignant transformation of
GES-1 cells
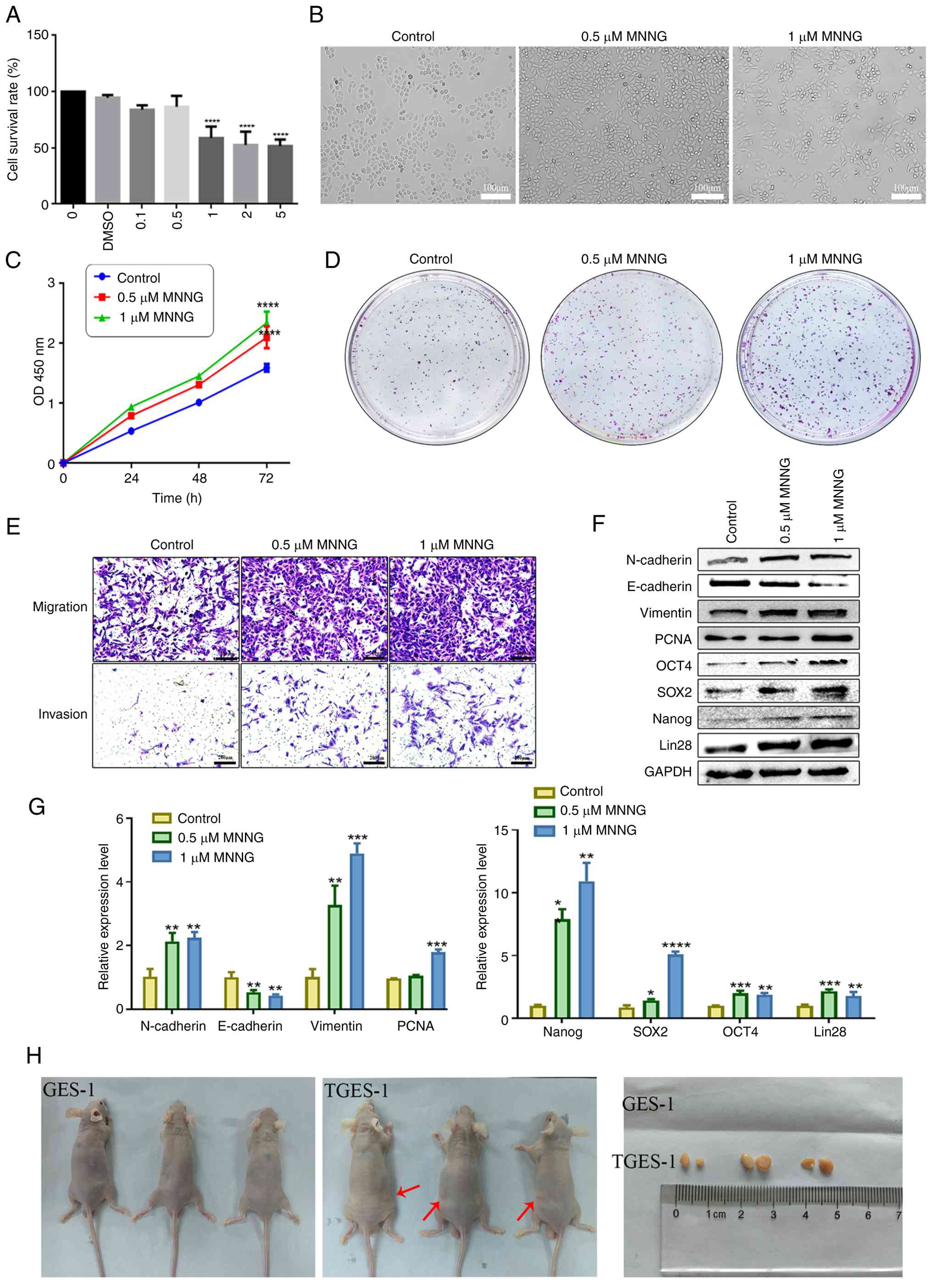
GES-1 cells were exposed to increasing
concentrations of MNNG (0, 0.1, 0.5, 1, 2 and 5 μM) for 72
h. Cell viability assessment using the CCK-8 assay revealed that a
statistically significant inhibitory effect was first observed at a
concentration of 1 μM MNNG (Fig. 1A). Based on these results, 0.5
μM and 1 μM MNNG were selected for long-term exposure
experiments to establish a malignant transformation model. GES-1
cells treated continuously with 1 μM MNNG for 30 passages
were defined as malignantly transformed GES-1 cells (TGES-1).
Morphological observation showed that prolonged MNNG exposure
markedly altered the appearance of GES-1 cells, which gradually
changed from a typical round or cobblestone-like morphology to a
spindle-shaped or cord-like phenotype (Fig. 1B). To determine whether long-term
MNNG exposure induced malignant transformation, a series of
functional assays, including migration and invasion assays, cell
proliferation assays and colony formation assays, were performed.
The results demonstrated that GES-1 cells treated with 1 μM
MNNG for 30 passages exhibited markedly enhanced proliferative
capacity, as well as increased migration and invasion abilities
(Fig. 1C-E). Furthermore,
RT-qPCR and western blot analyses showed that the expression levels
of mesenchymal markers and proliferation- and stemness-related
proteins, including N-cadherin, Vimentin, PCNA, OCT4, SOX2, Lin28
and NANOG, were markedly upregulated in MNNG-induced GES-1 cells,
whereas the epithelial marker E-cadherin was obviously
downregulated (Fig. 1F and
G).
In vivo tumorigenicity assays further
confirmed the malignant transformation of TGES-1 cells. The
incidence of xenograft tumors formed after subcutaneous injection
of TGES-1 cells, was 100% (6/6 per group), whereas no tumors were
observed in mice injected with control GES-1 cells (0/6; Fig. 1H). Collectively, these findings
demonstrated that both in vitro and in vivo exposure
to MNNG is sufficient to induce malignant transformation of normal
GES-1 cells.
TGES-1 cells affects the biological
characteristics of GES-1 cells via exosomes
Cancer cells are capable of inducing malignant
alterations in surrounding normal cells within the TME. To
investigate whether TGES-1 cells influenced the biological behavior
of normal gastric epithelial cells, GES-1 and TGES-1 cells were
co-cultured (Fig. 2A). The
proliferative, migratory and invasive capacities of GES-1 cells
before and after co-culture were evaluated using CCK-8 assays,
colony formation assays and Transwell assays. The results showed
that co-culture with TGES-1 cells clearly enhanced the
proliferative ability of GES-1 cells. In addition, the migration
and invasion capacities of GES-1 cells were markedly increased
following co-culture (Fig.
2B-D). Moreover, the mRNA expression levels of EMT markers and
stemness-associated genes were obviously upregulated (Fig. 2F and G).
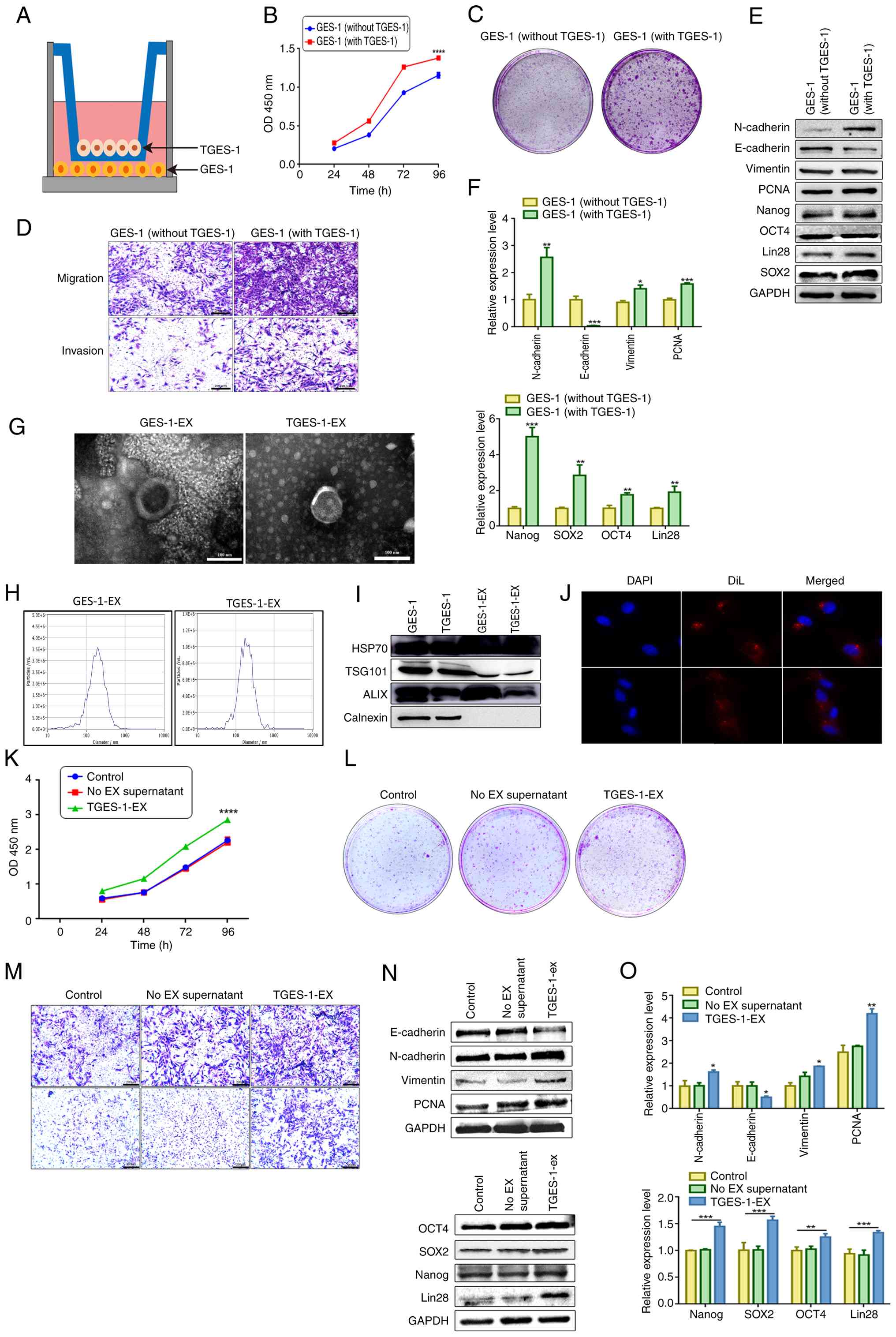 | Figure 2TGES-1 cells induce the biological
changes of GES-1 cells through exosomes. (A) Schematic
representation of GES-1 and TGES-1 cells in co-culture. (B) CCK8
assay was conducted to assess the proliferation of GES-1 cells
following co-culture with TGES-1 cells. (C) The cloning efficiency
of GES-1 cells was assessed following co-culture with TGES-1 cells.
(D) The capabilities for migration and invasion were analyzed after
co-culture. (E) Western blot analysis was performed to investigate
the protein expression of EMT and stemness markers in GES-1 cells
after co-culture with TGES-1 cells. (F) The mRNA expressions of
genes associated with EMT and stemness characteristics in GES-1
cells were analyzed after co-culture. (G) The morphological
characteristics of exosomes was characterized using TEM, scale bar:
100 nm. (H) The size of exosomes was detected by NTA. (I) The
surface marker proteins of exosomes were detected by western
blotting. (J) DAPI blue fluorescence labeled the nucleus and DiI
red fluorescence labeled the exosomes. The uptake of extracellular
vesicles by GES-1 cells was observed using confocal microscopy,
original magnification, ×60 (oil immersion). (K) The proliferation
of cells was detected after treatment with exosomes derived from
TGES-1 cells. (L) The cloning efficiency of cells was evaluated
following treatment with TGES-1-EX. (M) The effect of exosomes from
TGES-1 cells on the migration and invasion abilities of GES-1 cells
was investigated, scale bar: 200 μm. (N) Western blot
analysis was conducted to determine the protein levels of EMT and
stemness markers in GES-1 cells after treatment with TGES-1-EX. (O)
The mRNA levels of EMT and stemness markers were assessed after
treatment with TGES-1-EX using RT-qPCR. *P<0.05,
**P<0.01, ***P<0.001,
****P<0.0001. EMT, epithelial-mesenchymal transition;
TEM, transmission electron microscopy; NTA, nanoparticle tracking
analysis; RT-qPCR, reverse transcription-quantitative PCR; DiI,
1,1'-dioctadecyl-3,3,3',3'-tetramethylindocarbocyanine perchlorate;
EX, exosome; TGES-1-EX, exosome secreted by TGES-1 cells. |
Previous studies have demonstrated that cancer cells
can modulate the malignant transformation of adjacent or distant
normal epithelial cells through the secretion of exosomes (30,31). Based on this evidence, it was
hypothesized that TGES-1 cells exert their effects on GES-1 cells
via exosome-mediated communication. Exosomes were isolated from
TGES-1 cells that had been continuously exposed to 1 μM MNNG
for 30 passages (TGES-1-EX). The morphological features, particle
size distribution and surface marker expression of TGES-1-EX were
characterized using TEM, NTA and western blotting, respectively.
TEM imaging revealed typical cup-shaped, double-membrane vesicles
characteristic of exosomes (Fig.
2G). NTA analysis demonstrated that the mean particle diameters
were 197.6 and 188.7 nm (Fig.
2H). Western blot analysis confirmed the presence of exosomal
marker proteins, including HSP70, TSG101 and Alix, in both TGES-1
cells and their derived exosomes (Fig. 2I). To evaluate exosome uptake,
TGES-1-EX were labeled with the fluorescent dye DiI, while cell
nuclei were counterstained with DAPI. After incubation of labeled
exosomes with GES-1 cells for 24 h, fluorescence microscopy
revealed that DiI-labeled exosomes were efficiently internalized by
GES-1 cells (Fig. 2J).
GES-1 cells were subsequently treated with TGES-1-EX
for 72 h. CCK-8 assays showed that TGES-1-EX evidently promoted
GES-1 cell growth (Fig. 2K).
Colony formation and Transwell migration and invasion assays
further demonstrated that TGES-1-EX enhanced the proliferative,
migratory and invasive abilities of GES-1 cells (Fig. 2L and M). In addition, RT-qPCR and
western blot analyses revealed that TGES-1-EX induced EMT and
enhanced stemness-related properties in GES-1 cells (Fig. 2N and O). Collectively, these
results indicated that MNNG promoted gastric carcinogenesis and
that exosomes play a critical role in mediating intercellular
communication during this process.
CircRNA 0000549 is highly expressed in
TGES-1 cells, exosomes and serum exosomes
To further explore the molecular mechanisms
underlying MNNG-induced malignant transformation, circRNA
expression profiles were analyzed using two GEO datasets (GSE89143
and GSE93541) (Fig. 3A). Among
the differentially expressed circRNAs, hsa_circ_0000549
(circ0000549, circSRSF5) was identified as notably upregulated in
both GC tissues and plasma samples and has not been previously
characterized (Fig. 3B). A
heatmap showing the expression patterns of the shared circRNAs in
tissue samples is provided in Fig.
S1. According to circBase annotations (http://www.circbase.org/), circ0000549 is derived from
the SRSF5 gene and is located at chr14:70236227-70236759, generated
by circularization of exon 6 on human chromosome 14. Sanger
sequencing confirmed that the amplified product was fully
consistent with the predicted back-splice junction sequence
(Fig. 3C). Agarose gel
electrophoresis demonstrated that the RT-qPCR product of
circ0000549 appeared as a single band with the expected size and
showed no amplification from genomic DNA (gDNA), indicating that
the primers specifically amplified circ0000549 from cDNA (Fig. 3D). Furthermore, RNase R digestion
assays revealed that circ0000549 exhibited strong resistance to
RNase R, confirming its circular structure and enhanced stability
compared with its linear counterpart (Fig. 3E). Following treatment of TGES-1
cells with actinomycin D to inhibit transcription, the half-life of
linear SRSF5 mRNA was ~4 h, whereas circ0000549 remained stable
with a half-life of ~22 h (Fig.
3F). FISH analysis demonstrated that circ0000549 was
distributed in both the nucleus and cytoplasm of TGES-1 cells
(Fig. 3H). Collectively, these
results indicate that circ0000549 is a highly stable circular RNA
localized in both cellular compartments.
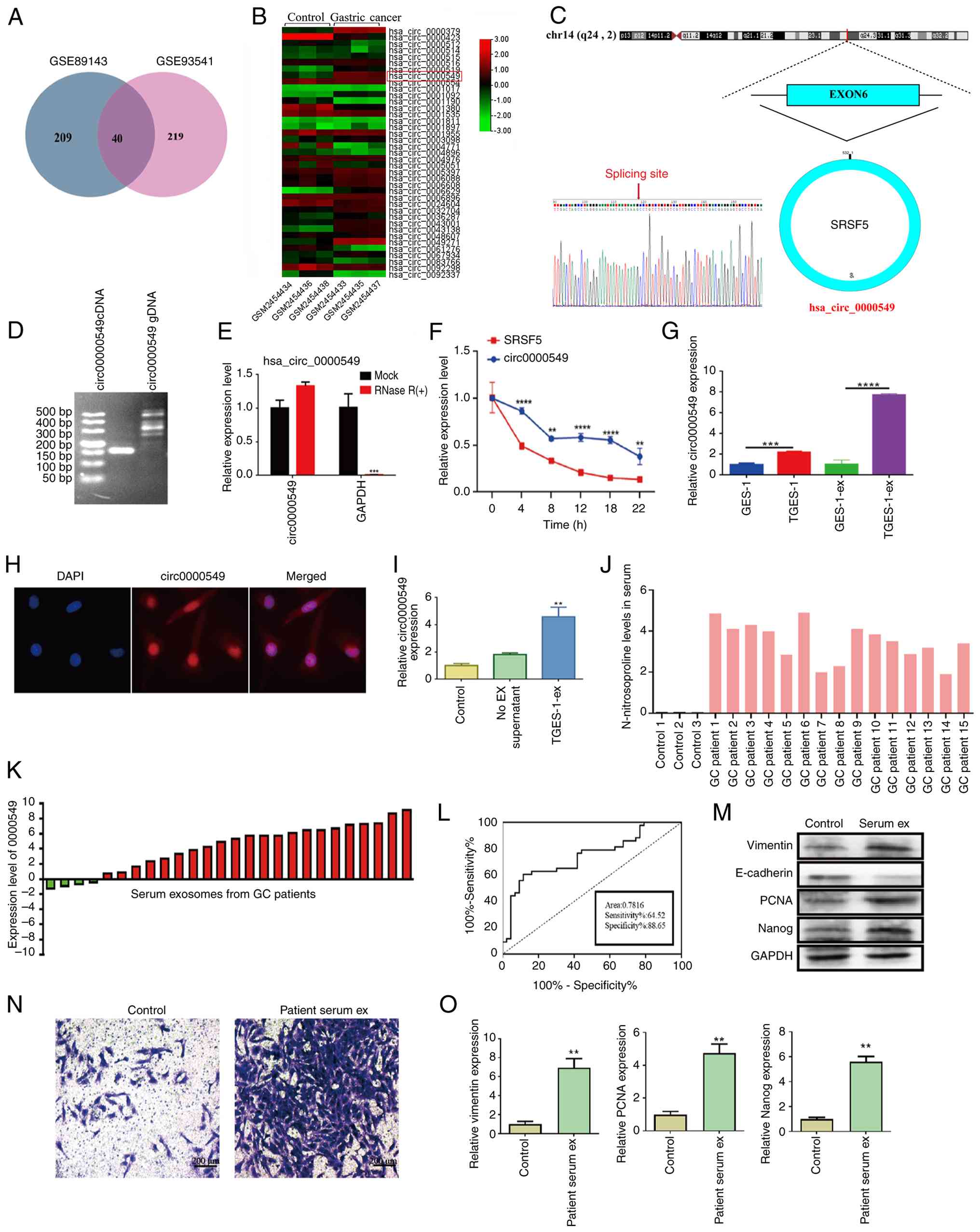 | Figure 3Characterization and expression of
circ0000549 in TGES-1 cells and exosomes. (A) Coincidence was
observed for the differentially expressed circular RNAs in the GEO
database. (B) Heat map illustrating the differential expression of
circRNAs, with circ0000549 standing out as highly expressed. (C)
Diagram of the cyclization mechanism of circ0000549. (D)
Verification through agarose gel electrophoresis showing the
specificity of circ0000549 amplification from cDNA. (E) RT-qPCR
results demonstrating the expression levels of circ0000549 before
and after RNase R treatment, confirming its resistance to RNase R
and stability compared to linear RNA. (F) The levels of circ0000549
and its parental gene SRSF5 were assessed after treatment with
actinomycin D. (G) The expression of circ0000549 in TGES-1 cells
and TGES-1-EX by RT-qPCR. (H) Localization of circ0000549
determined by FISH, original magnification, ×60 (oil immersion).
(I) The level of circ0000549 in GES-1 cells after TGES-1-EX
treatment by RT-qPCR. (J) Serum levels of N-nitrosoproline were
determined by LC-MS/MS. (K) The mRNA expression of circ0000549 in
serum exosomes, n=26. (L) Diagnostic efficacy of serum exosome
circ0000549, n=40. (M) Protein levels of EMT and stemness markers
in GES-1 cells after treatment with serum exosomes. (N) The
migration ability of GES-1 cells after treatment with serum
exosomes, scale bar, 200 μm. (O) The mRNA expression of EMT
and stemness markers in GES-1 cells after treatment with serum
exosomes. **P<0.01, ***P<0.001,
****P<0.0001. GEO, Gene Expression Omnibus; RT-qPCR,
reverse transcription-quantitative PCR; EMT, epithelial-mesenchymal
transition; GC, gastric cancer; FISH, fluorescence in situ
hybridization; Serum ex, serum derived exosomes from patients with
gastric cancer; DAPI, 4',6-diamidino-2-phenylindole. |
To confirm the expression of circ0000549 in TGES-1
cells, total RNA was extracted and analyzed by RT-qPCR, which
confirmed that circ0000549 expression was notably elevated in
TGES-1 cells. To determine whether MNNG-induced exosomes mediate
the transfer of circ0000549 during gastric carcinogenesis, total
RNA was isolated from exosomes derived from GES-1 and TGES-1 cells
and circ0000549 expression was assessed by RT-qPCR. Circ0000549 was
highly enriched in TGES-1 cells and their derived exosomes.
Moreover, treatment of GES-1 cells with TGES-1-derived exosomes for
72 h resulted in a marked increase in circ0000549 expression
(Fig. 3G and I).
To further evaluate the clinical relevance of
circ0000549 in nitrite-induced GC, patients with GC were stratified
based on dietary habits and exposure history to N-nitroso
compounds, including long-term consumption of pickled foods,
processed meat products and high-salt diets. Serum levels of
N-nitrosoproline were subsequently measured to identify patients
with high nitrate exposure (Fig.
3J). Notably, circ0000549 expression was elevated in serum
exosomes from patients with GC with a history of high nitrite
exposure (Fig. 3K). Receiver
operating characteristic (ROC) curve analysis was performed to
assess the diagnostic value of serum exosomal circ0000549 (n=40).
The area under the ROC curve (AUC) was 0.7816, with a sensitivity
of 64.52% and a specificity of 88.65% (Fig. 3L). In addition, serum-derived
exosomes from patients with GC facilitated EMT, proliferation and
stemness-related properties in GES-1 cells (Fig. 3M-O).
Circ0000549 promotes cell malignant
properties
To investigate the functional role of circ0000549 in
GC progression, two specific siRNAs targeting circ0000549 were
designed to reduce its expression. siRNA-1 and siRNA-2 both
efficiently suppressed circ0000549 expression in TGES-1 cells
(Fig. 4A). CCK-8 assays and
colony formation assays demonstrated that circ0000549 knockdown
notably inhibited cell proliferation (Fig. 4B and C). In addition, Transwell
assays revealed that silencing circ0000549 markedly reduced the
migratory and invasive abilities of TGES-1 cells (Fig. 4D). Consistent with these
functional changes, western blotting and RT-qPCR analyses showed
that circ0000549 downregulation resulted in decreased expression of
mesenchymal markers, proliferation-related proteins and
stemness-associated factors, including N-cadherin, Vimentin, PCNA,
OCT4, SOX2, Lin28 and Nanog, while the epithelial marker E-cadherin
was upregulated (Fig. 4E and
F).
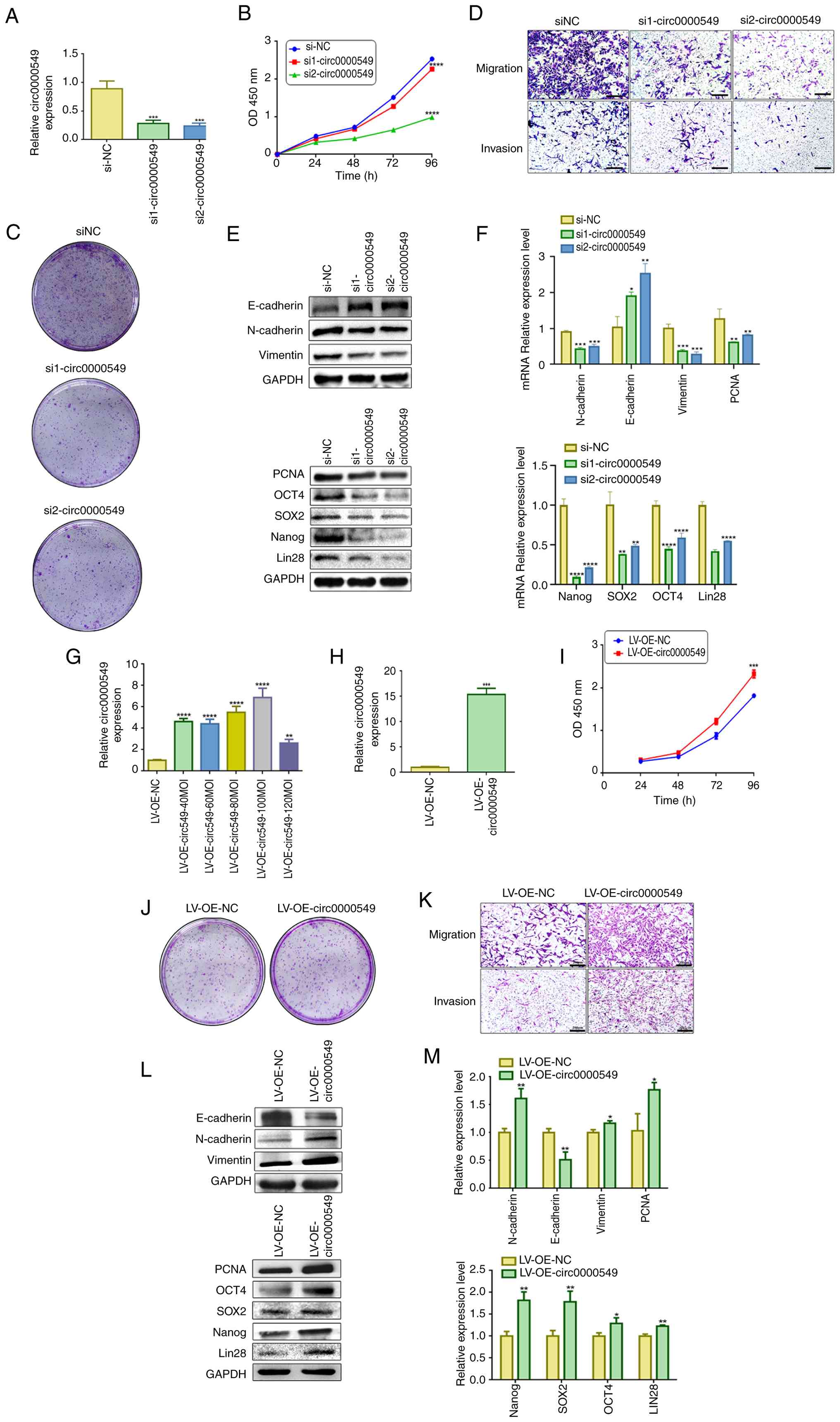 | Figure 4Circ0000549 promoted migration,
proliferation, invasion and EMT in GES-1 cells. (A) The
overexpression of circ0000549 in TGES-1 cells transfection with
si1-circ0000549 and si2-circ0000549 was detected by RT-qPCR. (B)
The effect of knocking down circ0000549 on the proliferation of
TGES-1 cells was determined through a CCK-8 assay. (C) The effect
of knocking down circ0000549 on the proliferative capacity of GES-1
cells was observed. (D) The abilities of TGES-1 cells to migrate
and invade were evaluated, scale bar, 200 μm. (E) The
protein levels of EMT-related and stemness were analyzed after
knocking down circ0000549. (F) The mRNAs expression of EMT-related
and stemness markers. (G) The level of circ0000549 in TGES-1 cells
was measured 48 h post-transfection with a lentivirus designed to
overexpress circ0000549. (H) The expression of circ0000549 after
puromycin screening. (I) Assessed the proliferative capacity of
TGES-1 cells following overexpression of circ0000549. (J) Changes
in the clonogenic ability of TGES-1 cells after overexpression of
circ0000549. (K) The effect of overexpression of circ0000549 on the
migration and invasion abilities in TGES-1 cells, scale bar, 200
μm. (L) Changes in the levels of EMT-related and stemness
protein after overexpression of circ0000549. (M) The expression of
EMT-related and stemness mRNA in TGES-1 cells after overexpression
of circ0000549. *P<0.05, **P<0.01,
***P<0.001, ****P<0.0001. EMT,
epithelial-mesenchymal transition; RT-qPCR, reverse
transcription-quantitative PCR; si1, siRNA1; si2, siRNA2; siNC,
siRNA negative control; LV-OE-NC, lentivirus overexpression
negative control; LV-OE, lentivirus overexpression; MOI,
multiplicity of infection. |
Next, TGES-1 cells were transfected with a
lentiviral vector overexpressing circ0000549 (Fig. 4G). Successful overexpression was
confirmed following puromycin selection (Fig. 4H). Functional assays demonstrated
that circ0000549 overexpression notably enhanced cell
proliferation, migration and invasion (Fig. 4I-K). Moreover, western blotting
and RT-qPCR analyses revealed that circ0000549 upregulation
increased the expression of N-cadherin, Vimentin, PCNA, OCT4, SOX2,
Lin28 and Nanog, while suppressing E-cadherin expression (Fig. 4L and M).
Exosomal circ0000549 facilitates
migration, invasion, proliferation, EMT and stemness of GES-1
cells
Previous studies have shown that exosomes serve as
important mediators of intercellular communication and can transfer
specific non-coding RNAs to recipient cells, thereby altering their
biological properties (32,33). To further explore whether
circ0000549 exerts its effects through exosome-mediated transfer,
TGES-1 cells were transfected with lentiviral vectors to knock down
circ0000549 expression (Fig.
5A). The efficiency of circ0000549 silencing was confirmed
after puromycin screening (Fig.
5B). Exosomes isolated from circ0000549 knockdown and
overexpression cells were subsequently characterized (Fig. 5C-E). Total RNA was extracted from
these exosomes and RT-qPCR analysis showed that circ0000549 levels
in exosomes derived from circ0000549-knockdown cells were reduced
by ~0.8-fold compared with control exosomes (sh-NC-EX). By
contrast, exosomes derived from circ0000549-overexpressing cells
exhibited an ~10-fold increase in circ0000549 expression relative
to control exosomes (OE-NC-EX; Fig.
5F). GES-1 cells were then treated with exosomes containing
either reduced or elevated circ0000549 levels. Functional assays
demonstrated that exosomes derived from circ0000549-knockdown cells
notably inhibited GES-1 cell migration, invasion, proliferation and
stemness (Fig. 5H-J).
Consistently, RT-qPCR and western blot analyses showed decreased
expression of N-cadherin, Vimentin, PCNA, OCT4, SOX2, Lin28 and
Nanog, accompanied by increased E-cadherin expression (Fig. 5J and K). By contrast, exosomes
enriched with circ0000549 exerted the opposite effects on GES-1
cells (Fig. 5K and L).
Collectively, these findings indicated that exosomal circ0000549
plays a critical role in promoting the malignant transformation of
normal gastric epithelial cells.
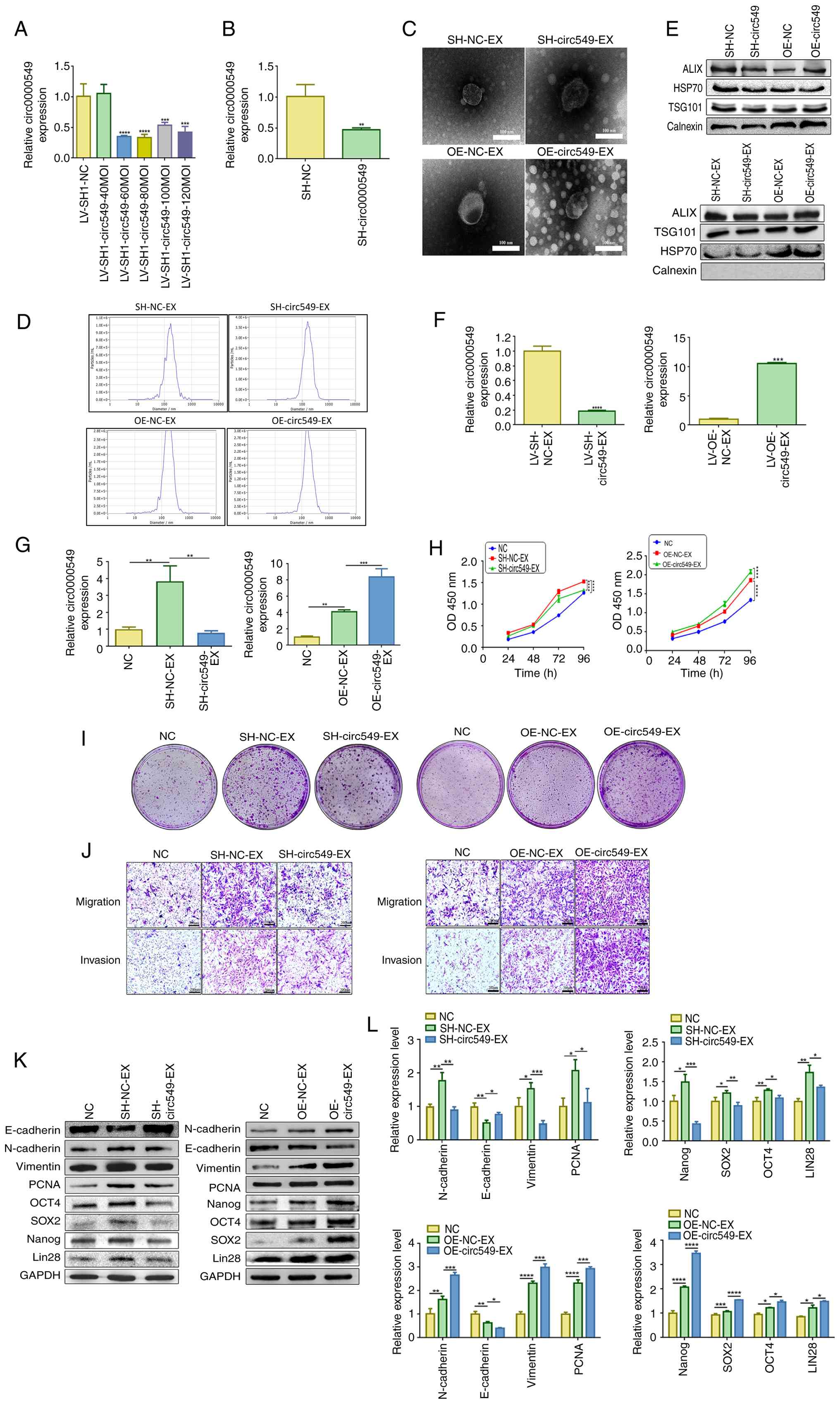 | Figure 5Exosomal circ0000549 promotes the
proliferation, migration, invasion, EMT and stemness in GES-1
cells. (A) Measure the level of circ0000549 in TGES-1 cells 48 h
post-transfection with a lentivirus designed for circ0000549
knockdown. (B) The level of circ0000549 after puromycin screening.
(C) The knockdown/overexpression of exosomes was detected by TEM,
scale bar, 100 nm. (D) The size of exosomes was determined using
NTA. (E) The protein levels of surface markers of exosomes were
detected. (F) The levels of circ0000549 in TGES-1-EX with
knockdown/overexpression of circ0000549. (G) The level of
circ0000549 in GES-1 cells after
SH-circ0000549-EX/OE-circ0000549-EX treatment. (H) Assessment of
GES-1 cell proliferation post-treatment with
SH-circ0000549-EX/OE-circ0000549-EX. (I) Evaluation of the
clonality of GES-1 cells after SH-circ0000549-EX/OE-circ0000549-EX
treatment. (J) Examination of the migratory and invasive
capabilities of GES-1 cells following
SH-circ0000549-EX/OE-circ0000549-EX treatment, scale bar, 200
μm. (K) Determination of EMT and stemness protein expression
levels in GES-1 cells treated SH-circ0000549-EX/OE-circ0000549-EX.
(L) Quantification of EMT and stemness mRNA levels in GES-1 cells
after SH-circ0000549-EX/OE-circ0000549-EX treatment using RT-qPCR.
*P<0.05, **P<0.01,
***P<0.001, ****P<0.0001. TEM,
transmission electron microscopy; NTA, nanoparticle tracking
analysis; RT-qPCR, reverse transcription-quantitative PCR; NC,
negative control; EX, exosome; OE-circ549-EX, exosomes from TGES-1
cells treated with circ0000549 lentivirus overexpression; OE-NC-EX,
exosomes from TGES-1 cells treated with circ0000549 lentivirus
overexpression negative control; SH-circ0000549-EX, exosomes from
TGES-1 cells treated with circ0000549 lentivirus knockdown;
SH-NC-EX; exosomes from TGES-1 cells treated with circ0000549
lentivirus knockdown negative control; MOI, multiplicity of
infection. |
miR-15b-5p serves as the target of
circ0000549 and attenuates its protumorigenic effects
To further elucidate the molecular mechanism
underlying circ0000549 function, the present study investigated its
potential role as a competing endogenous RNA. CircRNAs are known to
act as miRNA sponges in the cytoplasm and FISH analysis
demonstrated that circ0000549 was distributed in both the nucleus
and cytoplasm. In addition, CircInteractome analysis (https://circinteractome.nia.nih.gov/)
predicted that circ0000549 could bind to AGO2, suggesting its
involvement in miRNA-mediated regulation (Fig. 6A). It was therefore hypothesized
that circ0000549 may function as a ceRNA to modulate MNNG-induced
malignant phenotypes in GES-1 cells. Potential miRNAs interacting
with circ0000549 were predicted using the StarBase, circBank and
circAtlas databases. A total of six candidate miRNAs (miR-15b-5p,
miR-16-5p, miR-545-3p, miR-145-5p, miR-1307-5p and miR-330-5p) were
identified by overlapping predictions (Fig. 6B). Among these candidates, only
miR-15b-5p was notably upregulated following circ0000549 knockdown,
while circ0000549 overexpression resulted in a marked decrease in
miR-15b-5p expression (Fig. 6C and
D). In addition, miR-15b-5p was highly expressed in TGES-1
cells (Fig. 6E). To validate the
direct interaction between circ0000549 and miR-15b-5p,
circ0000549-WT and circ0000549-MUT binding sequences were cloned
into dual-luciferase reporter plasmids. Co-transfection of
circ0000549-WT with miR-15b-5p mimics notably reduced luciferase
activity, whereas no significant change was observed in cells
transfected with circ0000549-MUT or miR-NC (Fig. 6F). These results confirm that
miR-15b-5p directly binds to circ0000549.
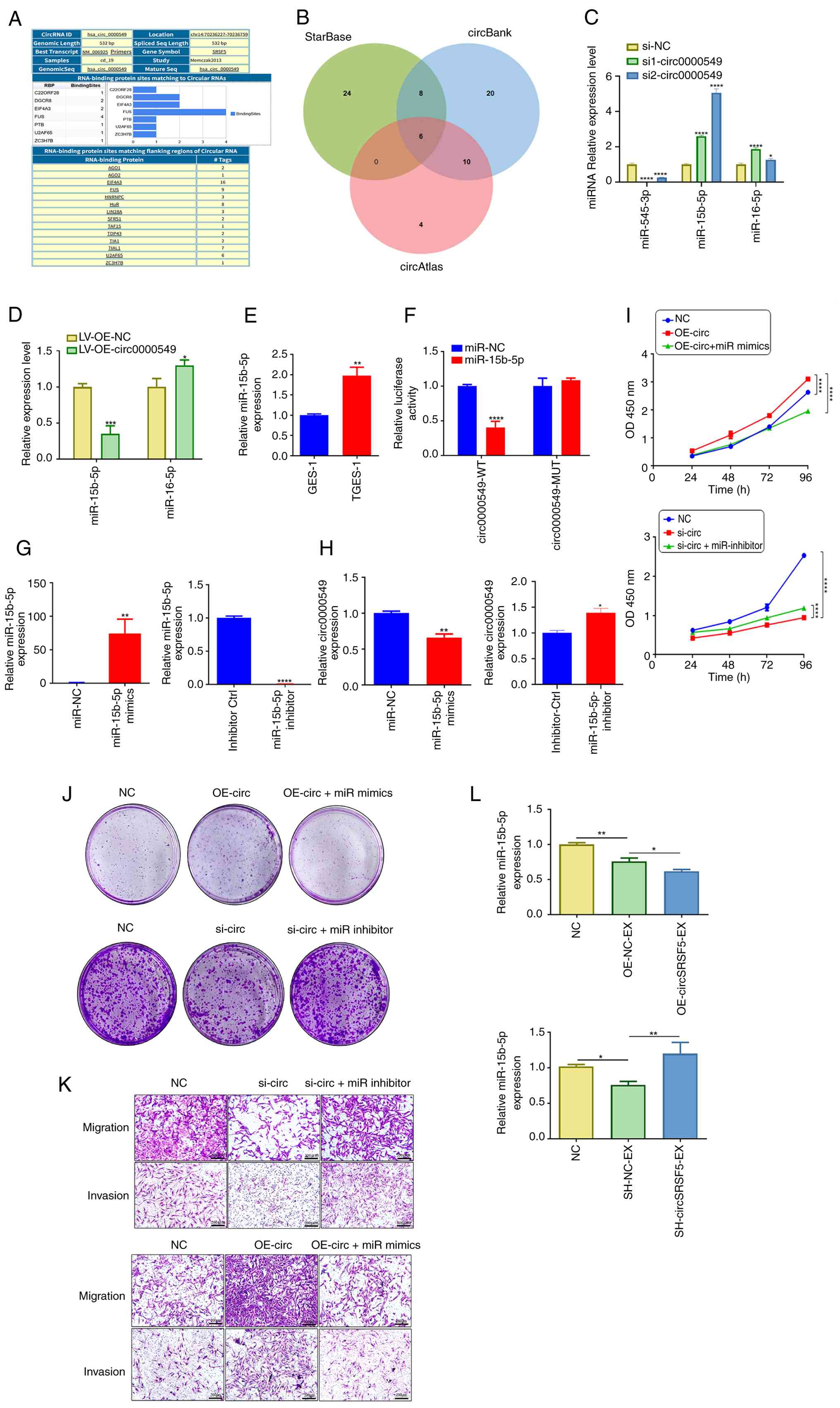 | Figure 6miR-15b-5p is the target of
circ0000549 and weakens the protumor effect of circ0000549. (A)
circInteractome database showed that AGO2 had binding ability with
the flanking region of circ0000549. (B) The online database
predicted the intersection of miRNA bound by circ0000549. (C) The
expression of miRNA after knocking down circ0000549 by RT-qPCR. (D)
The level of miRNA after overexpression of circ0000549 by RT-qPCR.
(E) The expression of miR-15b-5p was assessed. (F) A dual
luciferase reporter gene assay was conducted to investigate the
interaction between circ0000549 and miR-15b-5p. (G) The level of
miR-15b-5p expression in TGES-1 cells was assessed following
transfection with miR-15b-5p mimics or an inhibitor. (H) The level
of circ0000549 was assessed following transfection with miR-15b-5p
mimics or an inhibitor. (I) The proliferation of cells was
evaluated after co-transfecting them with OE-circ0000549 and
miR-15b-5p mimics, or with si-circ0000549 and miR-15b-5p inhibitor.
(J) Evaluation of the clonality of TGES-1 cells after
co-transfection of OE-circ0000549 and miR-15b-5p
mimics/si-circ0000549 and miR-15b-5p inhibitor. (K) The migration
and invasion capabilities of TGES-1 cells were assessed following
co-transfection with OE-circ0000549 and miR-15b-5p mimics, or with
si-circ0000549 and miR-15b-5p inhibitor, scale bar, 200 μm.
(L) Level of miR-15b-5p following
SH-circ0000549-EX/OE-circ0000549-EX treatment by RT-qPCR.
*P<0.05, **P<0.01,
***P<0.001, ****P<0.0001. RT-qPCR,
reverse transcription-quantitative PCR; si1, siRNA1; si2, siRNA2;
siNC, siRNA negative control; LV-OE-NC, lentivirus overexpression
negative control; LV-OE, lentivirus overexpression; WT, Wild Type;
MUT, Mutant; miR-NC, miR-15b-5p mimics negative control; inhibitor
Ctrl, miR-15b-5p inhibitor control; OE-circ, lentivirus
overexpression of circ0000549; miR mimics, miR-15b-5p mimics;
miR-inhibitor, miR-15b-5p inhibitor. |
RT-qPCR analysis verified the transfection
efficiency of miR-15b-5p mimics and inhibitors (Fig. 6G). Notably, circ0000549
expression was reduced following miR-15b-5p mimic transfection,
while inhibition of miR-15b-5p increased circ0000549 levels,
indicating a reciprocal regulatory relationship (Fig. 6H). Functionally, the promotive
effects of circ0000549 overexpression on cell proliferation,
migration and invasion were distinctly attenuated by miR-15b-5p
mimics (Fig. 6I and K).
Conversely, miR-15b-5p inhibition reversed the suppressive effects
induced by circ0000549 knockdown, restoring the malignant
phenotypes of GES-1 cells (Fig. 6I
and K). Consistently, miR-15b-5p expression was reduced in
GES-1 cells treated with circ0000549-overexpressing exosomes,
whereas circ0000549-depleted exosomes produced the opposite effect
(Fig. 6L). Collectively, these
results indicate that circ0000549 promotes malignant transformation
of GES-1 cells by sponging miR-15b-5p.
KIF1B is validated as a target gene of
miR-15b-5p
Candidate target genes of miR-15b-5p were identified
through intersection analysis of multiple online prediction
databases, yielding six potential targets (Fig. 7A). Among these candidates, KIF1B
was selected based on expression changes following circ0000549 and
miR-15b-5p overexpression or knockdown (Fig. 7B). RT-qPCR analysis confirmed
that KIF1B expression was distinctly elevated in TGES-1 cells
(Fig. 7C). A previous study has
reported that KIF1B functions as a downstream effector of oncogenic
circRNAs, such as CircPOSTN in glioma (34). Consistent with this,
co-transfection of circ0000549 overexpression plasmids and
miR-15b-5p mimics distinctly reduced KIF1B expression, whereas
co-transfection of circ0000549 siRNA and miR-15b-5p inhibitors
markedly increased KIF1B levels (Fig. 7D). Dual-luciferase reporter
assays further demonstrated that miR-15b-5p overexpression markedly
decreased luciferase activity in cells transfected with WT KIF1B
constructs, while no significant effect was observed in MUT
constructs (Fig. 7E).
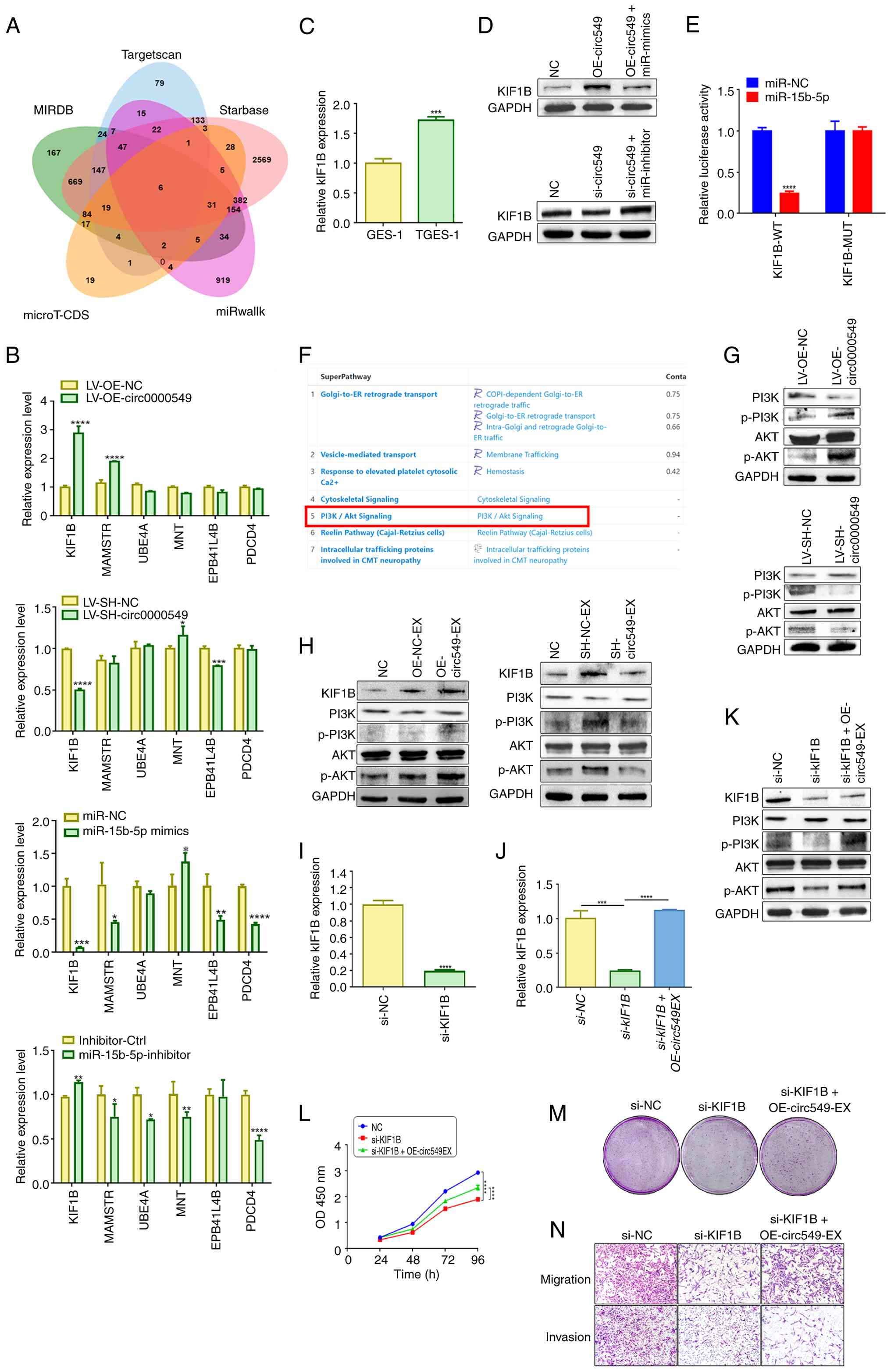 | Figure 7Circ0000549 in exosomes aids in
promoting the aggressive evolution of GES-1 cells, mainly through
miR-15b-5p/KIF1B/PI3K/AKT axis. (A) Internet databases were used to
predict which genes are regulated by miR-15b-5p. (B) The levels of
candidate target genes after knockdown/overexpression of
circ0000549 or transfection of miR-15b-5p mimics/inhibitor by
RT-qPCR. (C) mRNA level of KIF1B was detected. (D) The effect of co
transfection with OE circ0000549 and miR-15b-5p mimic/si
circ00000549 and miR-15b-5p inhibitor on the expression of KIF1B.
(E) A dual luciferase reporter gene assay was employed to elucidate
the interaction between KIF1B and miR-15b-5p. (F) GeneCards
database was used to predict KIF1B downstream pathways online. (G)
The pathway activity of the PI3K/AKT in TGES-1 cells after knocking
down/overexpressing circ0000549. (H) Changes in expression levels
of KIF1B and PI3K/AKT pathway proteins of GES-1 cells after
SH-circ0000549-EX/OE-circ0000549-EX treatment. (I) The effect on
KIF1B level in TGES-1 cells subsequent to KIF1B knockdown. (J) The
level of KIF1B in TGES-1 cells which was knocked down KIF1B after
treatment with OE-circ0000549 exosomes by RT-qPCR. (K) Detection of
changes in expression levels of KIF1B and PI3K/AKT pathway proteins
in TGES-1 cells which was knocked down KIF1B after treatment with
OE-circ0000549 exosomes. (L) Detection of changes in the
proliferation ability of TGES-1 cells. (M) Clonality of TGES-1
cells which was knocked down KIF1B after treatment with
OE-circ0000549 exosomes. (N) Alterations in the migratory and
invasive capabilities of TGES-1 cells treated with OE-circ0000549
exosomes and KIF1B knockdown in combination. *P<0.05,
**P<0.01, ***P<0.001,
****P<0.0001. miR, microRNA; RT-qPCR, reverse
transcription-quantitative PCR; NC, negative control; LV-OE-NC,
lentivirus overexpression negative control; LV-OE, lentivirus
overexpression; LV-SH-circ0000549, lentivirus knockdown of
circ0000549; LV-SH-NC; lentivirus knockdown of circ0000549 negative
control; miR-NC, miR-15b-5p mimics negative control; inhibitor
Ctrl, miR-15b-5p inhibitor control; OE-circ549, lentivirus
overexpression of circ0000549; OE-circ549-EX, exosomes from TGES-1
cells treated with circ0000549 lentivirus overexpression; siKIF1B,
siRNA of KIF1B; siNC, siRNA negative control; SH-circ549-EX,
exosomes from TGES-1 cells treated with circ0000549 lentivirus
knockdown; SH-NC-EX; exosomes from TGES-1 cells treated with
circ0000549 lentivirus knockdown negative control. |
Exosomal circ0000549 facilitates
malignant progression through the miR-15b-5p/KIF1B/PI3K/AKT
signaling pathway
GeneCards database analysis predicted that KIF1B is
functionally associated with the PI3K/AKT signaling pathway
(Fig. 7F). Western blot analysis
showed that circ0000549 overexpression activated the PI3K/AKT
pathway, whereas circ0000549 knockdown suppressed pathway
activation (Fig. 7G). Similarly,
exosomes enriched with circ0000549 increased the levels of
phosphorylated PI3K and AKT in GES-1 cells, while
circ0000549-depleted exosomes exerted the opposite effects
(Fig. 7H). To further verify the
involvement of KIF1B, siRNA targeting KIF1B was transfected into
TGES-1 cells (Fig. 7I).
Overexpression of circ0000549-containing exosomes partially rescued
the reduction in KIF1B expression caused by KIF1B knockdown
(Fig. 7J). Western blot analysis
further revealed that KIF1B silencing inhibited PI3K/AKT signaling,
whereas circ0000549-overexpressing exosomes restored PI3K and p-AKT
expression (Fig. 7K).
Functionally, KIF1B knockdown remarkably suppressed cell migration,
invasion and proliferation, while circ0000549-enriched exosomes
alleviated these inhibitory effects (Fig. 7L and N). These findings suggested
that exosomal circ0000549 promotes malignant transformation of
GES-1 cells through the miR-15b-5p/KIF1B/PI3K/AKT signaling
axis.
Exosomal circ0000549 enhances
tumorigenicity, EMT, proliferation and stem-like properties in
vivo
As GES-1 cells lack intrinsic tumorigenicity, TGES-1
cells were used for in vivo validation. Exosomes with
circ0000549 knockdown evidently reduced tumor formation rate, tumor
size and tumor volume in nude mice (Fig. 8A-C). Analysis of tumor tissues
revealed that circ0000549 expression was markedly decreased in the
circ0000549 knockdown exosome group (Fig. 8D). Furthermore, western blotting
and RT-qPCR analyses showed reduced expression of Vimentin, PCNA
and Nanog, accompanied by increased E-cadherin expression (Fig. 8E and F). Immunohistochemical
staining yielded consistent results (Fig. 8G), indicating that circ0000549
regulates EMT, proliferation and stemness in vivo.
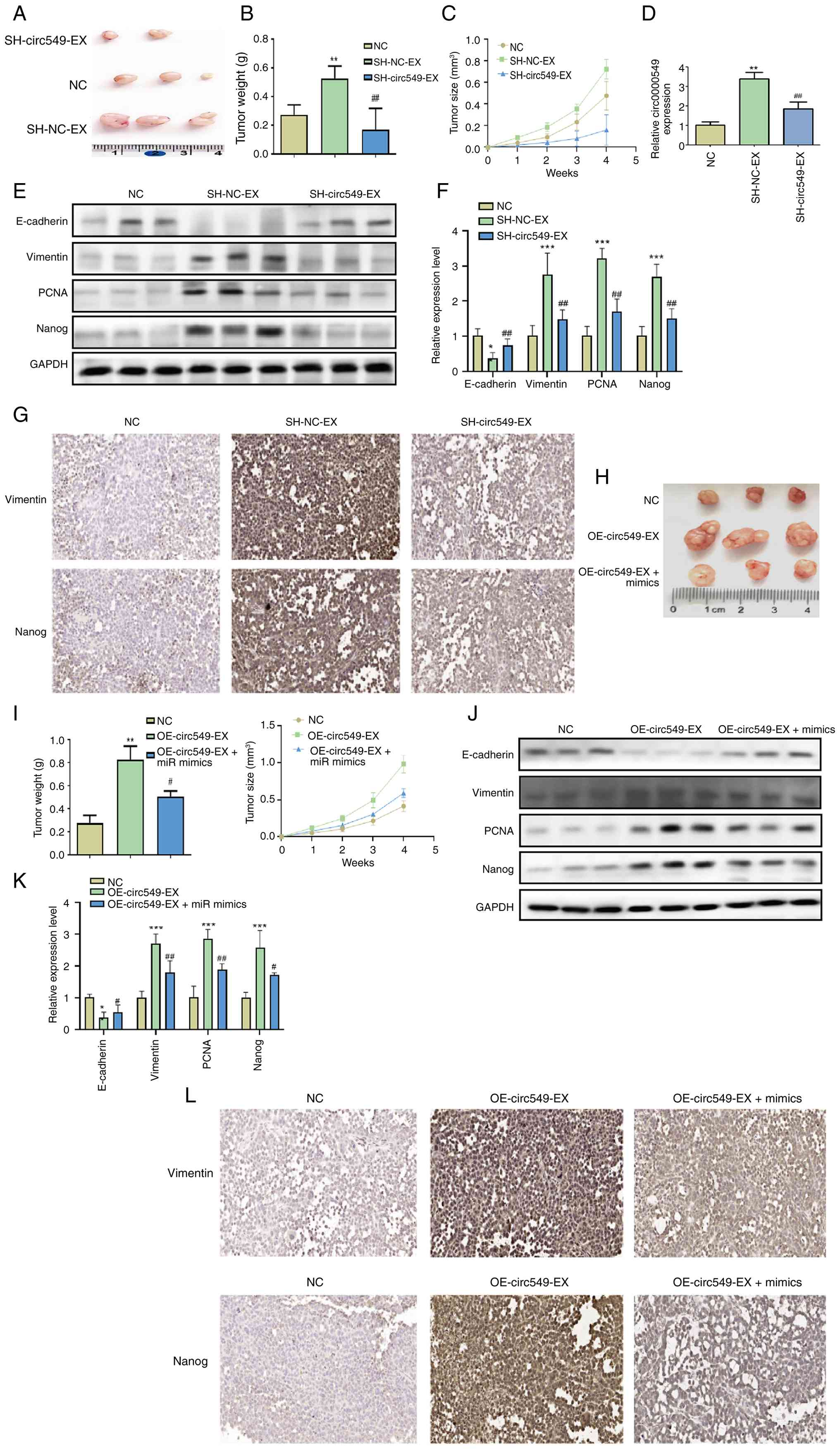 | Figure 8Exosomal circ0000549 tumorigenicity,
EMT, proliferation and stemness. (A) The downregulation of
circ0000549 impairs the tumorigenic capacity of TGES-1 cells. (B)
The effect of SH-circ0000549-EX treatment on tumor weight. (C) The
effect of SH-circ0000549-EX treatment on tumor volume. (D) The
level of circ0000549 in tumor tissues after SH-circ0000549-EX
treatment by RT-qPCR. (E) The levels of stemness, EMT and
proliferation proteins in tumor tissues after SH-circ0000549-EX
treatment. (F) The mRNA expression of stemness, EMT and
proliferation makers mRNA in tumor tissues after SH-circ0000549-EX
treatment. (G) Immunohistochemical analysis of tumor tissues
treated with SH-circ0000549-EX revealed alterations in the levels
of EMT and stemness markers, magnification, ×400. (H) The effect of
OE-circ0000549-EX alone or in combination with miR-15b-5p mimic on
the tumorigenicity of TGES-1 cells. (I) The effect of
OE-circ0000549-EX alone or in combination with miR-15b-5p mimic on
tumor weight and volume. (J) The levels of stemness, EMT and
proliferation-related markers in tumor tissues following treatment
with OE-circ0000549-EX alone or in combination with miR-15b-5p
mimic. (K) The expression of stemness, EMT and proliferation mRNA
in tumor tissues after treated with OE-circ0000549-EX alone or in
combination with miR-15b-5p mimic. (L) The levels of vimentin and
Nanog in tumor tissues after SH-circ0000549-EX treatment by
Immunohistochemistry, magnification, ×200. *P<0.05,
**P<0.01, ***P<0.001,
#P<0.05, ##P<0.01. EMT,
epithelial-mesenchymal transition; RT-qPCR, reverse
transcription-quantitative PCR; NC, negative control;
SH-circ549-EX, exosomes from TGES-1 cells treated with circ0000549
lentivirus knockdown; SH-NC-EX; exosomes from TGES-1 cells treated
with circ0000549 lentivirus knockdown negative control;
OE-circ549-EX, exosomes from TGES-1 cells treated with circ0000549
lentivirus overexpression. |
miR-15b-5p mediates circ0000549-driven
TGES-1 tumorigenesis in vivo
To further validate the circ0000549/miR-15b-5p axis
in vivo, nude mice were divided into three groups: TGES-1,
TGES-1 treated with circ0000549-overexpressing exosomes and TGES-1
treated with circ0000549-overexpressing exosomes combined with
miR-15b-5p mimics. Overexpression of circ0000549 evidently enhanced
the tumorigenicity of TGES-1 cells, whereas co-treatment with
miR-15b-5p mimics markedly attenuated this effect (Fig. 8H and I). Moreover, combined
treatment resulted in decreased expression of Vimentin, PCNA and
Nanog, along with increased E-cadherin expression, as confirmed by
western blotting, RT-qPCR and immunohistochemical analyses
(Fig. 8J and L). These results
demonstrate that circ0000549 promotes tumorigenicity, EMT,
proliferation and stemness of TGES-1 cells in a
miR-15b-5p-dependent manner.
Discussion
GC is one of the most prevalent malignant tumors
worldwide (35). Early GC is
often asymptomatic and is therefore frequently diagnosed at
advanced stages. The incidence of GC shows marked geographic
variation, which may be closely associated with genetic background,
lifestyle and environmental factors, particularly dietary habits.
Among these factors, dietary nitrite intake has been recognized as
one of the most important contributors to gastric carcinogenesis
(36). MNNG has been shown to
induce precancerous lesions, malignant transformation and tumor
formation in animal models (37). As a representative nitroso
compound, MNNG is widely used to simulate nitrosamine-induced
gastric mucosal carcinogenesis both in vivo and in
vitro (38). The present
study demonstrated that long-term exposure to MNNG is sufficient to
induce malignant transformation of human gastric epithelial cells.
However, despite extensive evidence supporting its carcinogenic
potential, the precise molecular mechanisms by which MNNG promotes
GC initiation and progression remain incompletely understood.
Within the TME, tumor cell-derived exosomes function
as important mediators of intercellular communication, facilitating
information exchange between malignant cells and neighboring normal
cells and thereby promoting tumor growth, invasion and metastasis
(39). Exosomes are nanoscale
extracellular vesicles with diameters ranging from 30-200 nm,
capable of selectively encapsulating a variety of bioactive
molecules from donor cells, including lipids, proteins, non-coding
RNAs, nucleic acids and metabolites (24,40). An increasing body of evidence has
demonstrated that circRNAs are involved in the initiation and
progression of numerous diseases and exert critical regulatory
roles in diverse physiological and pathological processes (41-44). In particular, exosome-mediated
transfer of circRNAs is critical in tumor development and
progression, including GC. For instance, Zheng et al
(45) reported that
exosome-transmitted circATG4B contributes to reduced
chemosensitivity in colorectal cancer cells, highlighting a
potential therapeutic strategy to overcome oxaliplatin resistance.
Similarly, Lin et al (46) demonstrated that hypoxia-induced
exosomes carrying circPDK1 enhance glycolysis in pancreatic cancer
cells by modulating the miR-628-3p/BPTF axis and promoting BIN1
degradation, thereby elevating c-Myc expression. Despite these
advances, the roles and molecular mechanisms of circRNAs in the
malignant transformation of normal cells induced by environmental
carcinogens remain poorly understood.
In the present study, normal GES-1 cells were
treated with MNNG-induced exosomes and it was observed that these
exosomes promoted malignant transformation of GES-1 cells in a
manner comparable to direct MNNG exposure. Analysis of GEO datasets
(GSE89143 and GSE93541) revealed that circ0000549 was highly
expressed in GC tissues, malignantly transformed GES-1 cells and
their derived exosomes, leading us to select circ0000549 as the
focus of this investigation. We further demonstrated that
circ0000549 carried by TGES-1-derived exosomes was capable of
inducing malignant transformation of GES-1 cells. To elucidate the
functional role and molecular mechanism of circ0000549 in GC
progression, the present study performed a series of experimental
analyses guided by bioinformatic predictions. The results showed
that circ0000549 promotes KIF1B expression through competitive
binding to miR-15b-5p. KIF1B has previously been reported to
regulate tumor progression in glioma and hepatocellular carcinoma
(34,47). Bioinformatic analysis further
suggested that KIF1B is functionally associated with the PI3K/AKT
signaling pathway. Consistent with these predictions, the findings
of the present study indicated that the MNNG-induced exosomal
circ0000549/miR-15b-5p/KIF1B axis regulates the proliferative,
migratory and invasive capacities of normal gastric epithelial
cells by activating the PI3K/AKT pathway and promoting the EMT
process, thereby facilitating malignant transformation and the
initiation of GC.
Taken together, the findings demonstrate that
exosomal circ0000549 promotes the malignant progression of
surrounding normal gastric epithelial cells through the
miR-15b-5p/KIF1B/PI3K/AKT signaling axis. These results suggested
that the exosomal circ0000549/miR-15b-5p/KIF1B axis may represent a
promising therapeutic target for the prevention and treatment of
nitrite-induced GC. Moreover, the present study provided important
mechanistic evidence supporting the clinical management of gastric
precancerous lesions and the potential delay of GC progression.
Although the present study focused on circ0000549, it does not
exclude the possibility that other exosomal components derived from
MNNG-transformed cells, such as oncogenic microRNAs or proteins,
may also contribute to the priming of recipient cells.
Nevertheless, the circ0000549 knockdown and rescue experiments
indicate that circ0000549 accounts for a substantial proportion of
the observed effects, underscoring its predominant role among the
oncogenic cargos carried by exosomes.
In conclusion, the present study demonstrated that
long-term exposure to MNNG induces malignant transformation of
GES-1 cells, with exosomes playing a pivotal role in mediating
intercellular communication during this process. Specifically,
exosomal circ0000549 was identified as a key driver of gastric
carcinogenesis by activating the PI3K/AKT signaling pathway and
promoting malignant transformation of normal gastric epithelial
cells through the miR-15b-5p/KIF1B axis. These findings revealed a
previously unrecognized regulatory mechanism involving exosomal
circRNAs in MNNG-induced GC development.
Supplementary Data
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author. The data of LC-MS/MS have
been submitted to the OMIX repository of the National Genomics Data
Center under accession number OMIX014373-02 (BioProject:
PRJCA055549) and are publicly accessible at: https://ngdc.cncb.ac.cn/omix/preview/tVgV3viP or
https://ngdc.cncb.ac.cn/gsub/submit/bioproject/PRJCA055549.
Authors' contributions
ZL was responsible for conceptualization,
methodology, software, resources, data analysis, writing the
original draft and manuscript revision. ZG and YZ were responsible
for data curation, data analysis, methodology and writing the
original draft. JS was responsible for data analysis and
investigation. HQ was responsible for writing, reviewing and
editing and supervision. XX was responsible for writing, reviewing
and editing, conceptualization, methodology and resources. ZL and
YZ confirm the authenticity of all the raw data. All authors read
and approved the final manuscript.
Ethics approval and consent to
participate
All mouse experiments conducted in the present
study were approved by the Animal Care and Use Committee of Jiangsu
University (approval no. UJS-IACUC-AP-2023030801). Every effort was
made to minimize animal suffering and distress throughout the
experimental procedures. The collection of human samples in the
present study was approved by the Medical Ethics Committee of
Jiangsu University (ethics approval no. 2023182). Written informed
consent was obtained from each participant, who also agreed to the
use of the relevant experimental data for publication.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Acknowledgements
Not applicable.
Funding
The present study was generously supported by funding from the
National Natural Science Foundation of China (grant no. 81602883),
Technology Development Project of Jiangsu University (grant nos.
20220014 and 20220516) and Postgraduate Research and Practice
Innovation Program of Jiangsu Province (grant no. KYCX25_4287).
References
|
1
|
Siegel RL, Kratzer TB, Giaquinto AN, Sung
H and Jemal A: Cancer statistics, 2025. CA Cancer J Clin. 75:10–45.
2025.PubMed/NCBI
|
|
2
|
Diao X, Guo C, Jin Y, Li B, Gao X, Du X,
Chen Z, Jo M, Zeng Y, Ding C, et al: Cancer situation in China: An
analysis based on the global epidemiological data released in 2024.
Cancer Commun (Lond). 45:178–197. 2025. View Article : Google Scholar :
|
|
3
|
Malik TH, Sayahan MY, Al Ahmed HA and Hong
X: Gastric intestinal metaplasia: An intermediate precancerous
lesion in the cascade of gastric carcinogenesis. J Coll Physicians
Surg Pak. 27:166–172. 2017.PubMed/NCBI
|
|
4
|
Wang Q, Huang Y, Jiang M, Tang Y, Wang Q,
Bai L, Yu C, Yang X, Ding K, Wang W, et al: The demethylase ALKBH5
mediates ZKSCAN3 expression through the m6A modification
to activate VEGFA transcription and thus participates in
MNNG-induced gastric cancer progression. J Hazard Mater.
473:1346902024. View Article : Google Scholar
|
|
5
|
Xia C, Dong X, Li H, Cao M, Sun D, He S,
Yang F, Yan X, Zhang S, Li N and Chen W: Cancer statistics in China
and United States, 2022: Profiles, trends, and determinants. Chin
Med J (Engl). 135:584–590. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Chiavarini M, Bertarelli G, Minelli L and
Fabiani R: Dietary intake of meat cooking-related mutagens (HCAs)
and risk of colorectal adenoma and cancer: A systematic review and
meta-analysis. Nutrients. 9:5142017. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Wu X, Chen L, Cheng J, Qian J, Fang Z and
Wu J: Effect of dietary salt intake on risk of gastric cancer: A
systematic review and meta-analysis of case-control studies.
Nutrients. 14:42602022. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Schoental R: Carcinogenic activity of
N-methyl-N-nitroso-N'-nitroguanidine. Nature. 209:726–727. 1966.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Liu T, Feng YL, Wang RY, Yang S, Ge YL,
Zhang TY, Li J, Li CY, Ruan Y, Luo B and Liang GY: Long-term MNNG
exposure promotes gastric carcinogenesis by activating
METTL3/m6A/miR1184 axis-mediated epithelial-mesenchymal transition.
Sci Total Environ. 913:1697522024. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Wsierska-Gadek J, Gueorguieva M and
Wojciechowski J: MNNG induces dramatic DNA damage and non-apoptotic
changes in cervical carcinoma HeLa cells. Ann NY Acad Sci.
1010:278–282. 2003. View Article : Google Scholar
|
|
11
|
Wang Y, Chu F, Lin J, Li Y, Johnson N,
Zhang J, Gai C, Su Z, Cheng H, Wang L and Ding X: Erianin, the main
active ingredient of Dendrobium chrysotoxum Lindl, inhibits
precancerous lesions of gastric cancer (PLGC) through suppression
of the HRAS-PI3K-AKT signaling pathway as revealed by network
pharmacology and in vitro experimental verification. J
Ethnopharmacol. 279:1143992021. View Article : Google Scholar
|
|
12
|
Zhao Y, Sun Y, Wang G, Ge S and Liu H:
Dendrobium officinale polysaccharides protect against MNNG-induced
PLGC in rats via activating the NRF2 and antioxidant enzymes HO-1
and NQO-1. Oxid Med Cell Longev. 2019:93102452019. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Liang M, Liu Z, Lin H, Shi B, Li M, Chen
T, Qin L, Niu Q, Yu G, Jiang H and Zhou X: High-throughput
sequencing reveals circular RNA hsa_circ_0000592 as a novel player
in the carcinogenesis of gastric carcinoma. Biosci Rep.
39:BSR201819002019. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Zhu F, Xu Y, Pan J, Li M, Chen F and Xie
G: Epigallocatechin gallate protects against MNNG-induced
precancerous lesions of gastric carcinoma in rats via PI3K/Akt/mTOR
pathway. Evid Based Complement Alternat Med.
2021:88468132021.PubMed/NCBI
|
|
15
|
Xu J, Shen W, Pei B, Wang X, Sun D, Li Y,
Xiu L, Liu X, Lu Y, Zhang X and Yue X: Xiao Tan He Wei Decoction
reverses MNNG-induced precancerous lesions of gastric carcinoma in
vivo and vitro: Regulation of apoptosis through NF-κB pathway.
Biomed Pharmacother. 108:95–102. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Zhao Y, Li B, Wang G, Ge S, Lan X, Xu G
and Liu H: Dendrobium officinale polysaccharides inhibit
1-methyl-2-nitro-1-nitrosoguanidine induced precancerous lesions of
gastric cancer in rats through regulating Wnt/β-catenin pathway and
altering serum endogenous metabolites. Molecules. 24:26602019.
View Article : Google Scholar
|
|
17
|
Zhang Y, Chen Z, Song J, Qian H, Wang Y
and Liang Z: The role of m6A modified circ0049271 induced by MNNG
in precancerous lesions of gastric cancer. Heliyon. 10:e356542024.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Liu W, Niu J, Huo Y, Zhang L, Han L, Zhang
N and Yang M: Role of circular RNAs in cancer therapy resistance.
Mol Cancer. 24:552025. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Kristensen LS, Andersen MS, Stagsted LVW,
Ebbesen KK, Hansen TB and Kjems J: The biogenesis, biology and
characterization of circular RNAs. Nat Rev Genet. 20:675–691. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Conn VM, Chinnaiyan AM and Conn SJ:
Circular RNA in cancer. Nat Rev Cancer. 24:597–613. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Lin Z, Zhong C, Shi M, Long Q, Jing L, Yu
Y, Chou J, Chen M, Lan M and Long F: Circular RNA TFRC/SCD1 mRNA
interaction regulates ferroptosis and metastasis in gastric cancer.
Cell Death Dis. 16:4362025. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zhou G, Wen J, Lu F, Wang J and Tan S:
circRNA_0005927 inhibits gastric cancer metastasis by
downregulating the miR-570-3p/FOXO3 axis. Am J Transl Res.
17:1679–1693. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Glaviano A, Lau HS, Carter LM, Lee EHC,
Lam HY, Okina E, Tan DJJ, Tan W, Ang HL, Carbone D, et al:
Harnessing the tumor microenvironment: Targeted cancer therapies
through modulation of epithelial-mesenchymal transition. J Hematol
Oncol. 18:62025. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Li J, Wang A, Guo H, Zheng W, Chen R, Miao
C, Zheng D, Peng J, Wang J and Chen Z: Exosomes: Innovative
biomarkers leading the charge in non-invasive cancer diagnostics.
Theranostics. 15:5277–5311. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zheng Z, Zhai Y, Yan X, Wang Z, Zhang H,
Xu R, Liu X, Cai J, Zhang Z, Shang Y, et al: Functions and clinical
applications of exosomes in gastric cancer. Int J Biol Sci.
21:2330–2345. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Paskeh MDA, Entezari M, Mirzaei S,
Zabolian A, Saleki H, Naghdi MJ, Sabet S, Khoshbakht MA, Hashemi M,
Hushmandi K, et al: Emerging role of exosomes in cancer progression
and tumor microenvironment remodeling. J Hematol Oncol. 15:832022.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Yang R, Qu X, Zhi S, Wang J, Fu J, Tan C,
Chen H and Wang X: Exosomes derived from meningitic escherichia
coli-infected brain microvascular endothelial cells facilitate
astrocyte activation. Mol Neurobiol. 61:7195–7210. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Huang XJ, Wang Y, Wang HT, Liang ZF, Ji C,
Li XX, Zhang LL, Ji RB, Xu WR, Jin JH and Qian H: Exosomal
hsa_circ_000200 as a potential biomarker and metastasis enhancer of
gastric cancer via miR-4659a/b-3p/HBEGF axis. Cancer Cell Int.
23:1512023. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
30
|
Malyla V, Paudel KR, Rubis G, Hansbro NG,
Hansbro PM and Dua K: Extracellular vesicles released from cancer
cells promote tumorigenesis by inducing epithelial to mesenchymal
transition via β-catenin signaling. Int J Mol Sci. 24:35002023.
View Article : Google Scholar
|
|
31
|
Lin Y, Jiang H, Li J, Ren F, Wang Y, Qiu
Y, Li J, Li M, Wang Y, Yang L, et al: Microenvironment-induced
CREPT expression by cancer-derived small extracellular vesicles
primes field cancerization. Theranostics. 14:662–680. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Gao L, Tang X, He Q, Sun G, Wang C and Qu
H: Exosome-transmitted circCOG2 promotes colorectal cancer
progression via miR-1305/TGF-β2/SMAD3 pathway. Cell Death Discov.
7:2812021. View Article : Google Scholar
|
|
33
|
Zheng R, Zhang K, Tan S, Gao F, Zhang Y,
Xu W, Wang H, Gu D, Zhu L, Li S, et al: Exosomal circLPAR1
functions in colorectal cancer diagnosis and tumorigenesis through
suppressing BRD4 via METTL3-eIF3h interaction. Mol Cancer.
21:492022. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Guan Y, Yang W, Zhang F, Zhang L and Wang
L: CircPOSTN competes with KIF1B for miR-185-5p binding sites to
promote the tumorigenesis of glioma. Brain Res Bull. 180:86–96.
2022. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Sundar R, Nakayama I, Markar SR, Shitara
K, van Laarhoven HWM, Janjigian YY and Smyth EC: Gastric cancer.
Lancet. 405:2087–2102. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Wang XQ, Terry PD and Yan H: Review of
salt consumption and stomach cancer risk: Epidemiological and
biological evidence. World J Gastroenterol. 15:2204–2213. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Gunes-Bayir A, Guler EM, Bilgin MG, Ergun
IS, Kocyigit A and Dadak A: Anti-inflammatory and antioxidant
effects of carvacrol on N-methyl-N'-nitro-N-nitrosoguanidine (MNNG)
induced gastric carcinogenesis in Wistar rats. Nutrients.
14:28482022. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Li J, Chen X, Mao C, Xiong M, Ma Z, Zhu J,
Li Z, Chen W, Ma H and Ye X: Epiberberine ameliorates MNNG-induced
chronic atrophic gastritis by acting on the EGFR-IL33 axis. Int
Immunopharmacol. 145:1137182025. View Article : Google Scholar
|
|
39
|
Yang E, Wang X, Gong Z, Yu M, Wu H and
Zhang D: Exosome-mediated metabolic reprogramming: The emerging
role in tumor microenvironment remodeling and its influence on
cancer progression. Signal Transduct Target Ther. 5:2422020.
View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Huang S, Yan F, Qiu Y, Liu T, Zhang W,
Yang Y, Zhong R, Yang Y and Peng X: Exosomes in inflammation and
cancer: From bench to bedside applications. Mol Biomed. 6:412025.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Lv X, Liu W, Zhou X, Yang Y, Zhao W, Meng
L, Mu F, Zhang Z, Zhu S, Zhang S and Wang Y: Exosomes in systemic
autoimmune diseases: Recent advances in diagnostic biomarkers and
therapeutic applications. Int J Nanomedicine. 20:5137–5160. 2025.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Huang C, Li J, Xie Z, Hu X and Huang Y:
Relationship between exosomes and cancer: Formation, diagnosis, and
treatment. Int J Biol Sci. 21:40–62. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Wang J, Zhou L, Chen B, Yu Z, Zhang J,
Zhang Z, Hu C, Bai Y, Ruan X, Wang S, et al: Circular RNA circCSPP1
promotes the occurrence and development of colon cancer by sponging
miR-431 and regulating ROCK1 and ZEB1. J Transl Med. 20:582022.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
O'Leary E, Jiang Y, Kristensen LS, Hansen
TB and Kjems J: The therapeutic potential of circular RNAs. Nat Rev
Genet. 26:230–244. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Pan Z, Zheng J, Zhang J, Lin J, Lai J, Lyu
Z, Feng H, Wang J, Wu D and Li Y: A novel protein encoded by
exosomal CircATG4B induces oxaliplatin resistance in colorectal
cancer by promoting autophagy. Adv Sci (Weinh). 9:e22045132022.
View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Lin J, Wang X, Zhai S, Shi M, Peng C, Deng
X, Fu D, Wang J and Shen B: Hypoxia-induced exosomal circPDK1
promotes pancreatic cancer glycolysis via c-myc activation by
modulating miR-628-3p/BPTF axis and degrading BIN1. J Hematol
Oncol. 15:1282022. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Luo YY, Zhang HP, Huang AL and Hu JL:
Association between KIF1B rs17401966 genetic polymorphism and
hepatocellular carcinoma susceptibility: An updated meta-analysis.
BMC Med Genet. 20:592019. View Article : Google Scholar : PubMed/NCBI
|